Submitted:
14 March 2026
Posted:
23 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
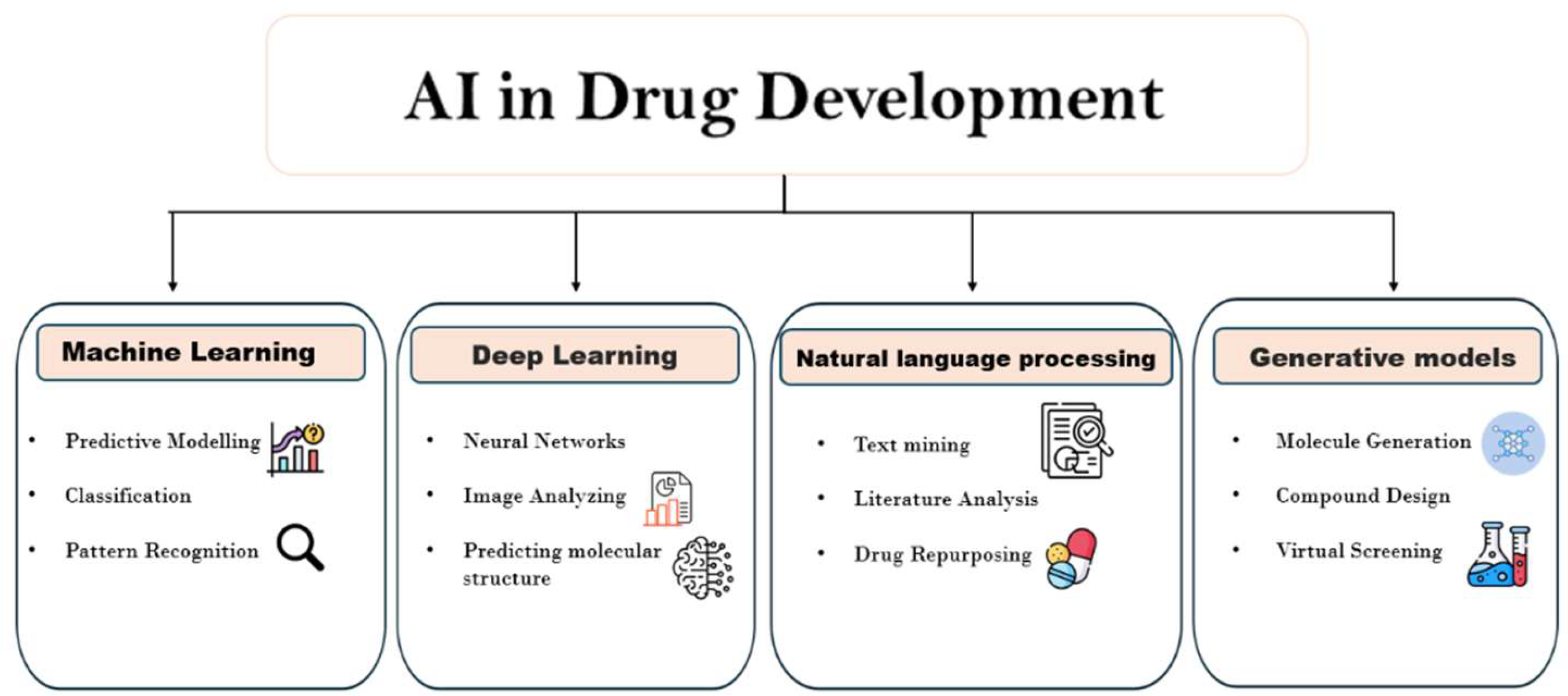
- This research provides an overview of the limitations of conventional drug discovery methods.
- In this research we discuss how AI-driven techniques, including machine learning, deep learning and natural language processing, allow researchers to rapidly analyze large biomedical datasets.
- We also examines how these technologies improve target identification, virtual screening and refine lead compounds.
- We conduct a computational case study in this research using machine learning to predict molecular solubility of selected drug molecules to show and analyze the practical application of AI techniques in evaluating physicochemical properties of potential drug candidates.
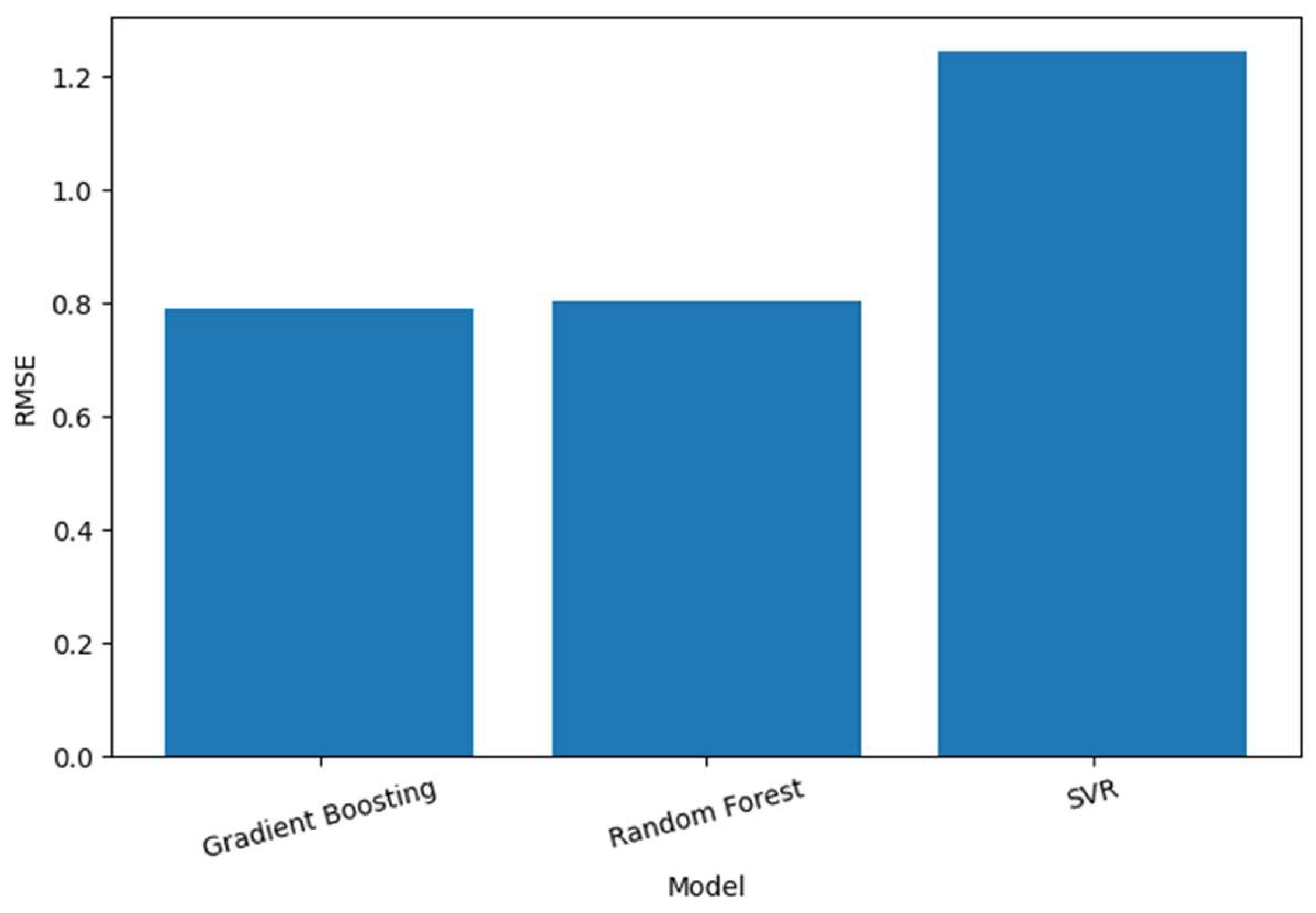
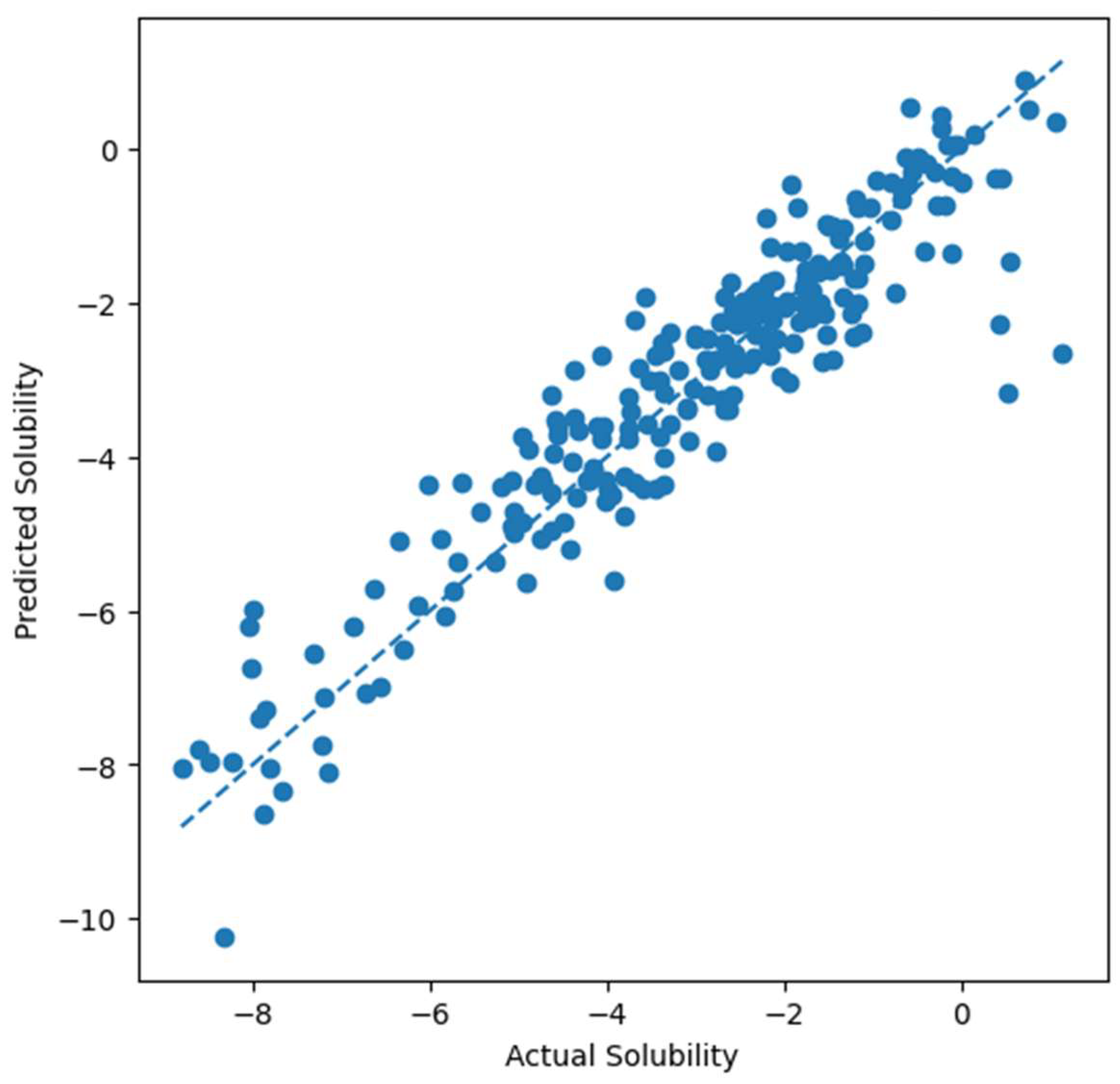
- Our obtained experimental results provide quantitative evidence of model performance, where the Gradient Boosting model achieved the best prediction accuracy (RMSE = 0.787 and R2 = 0.869).
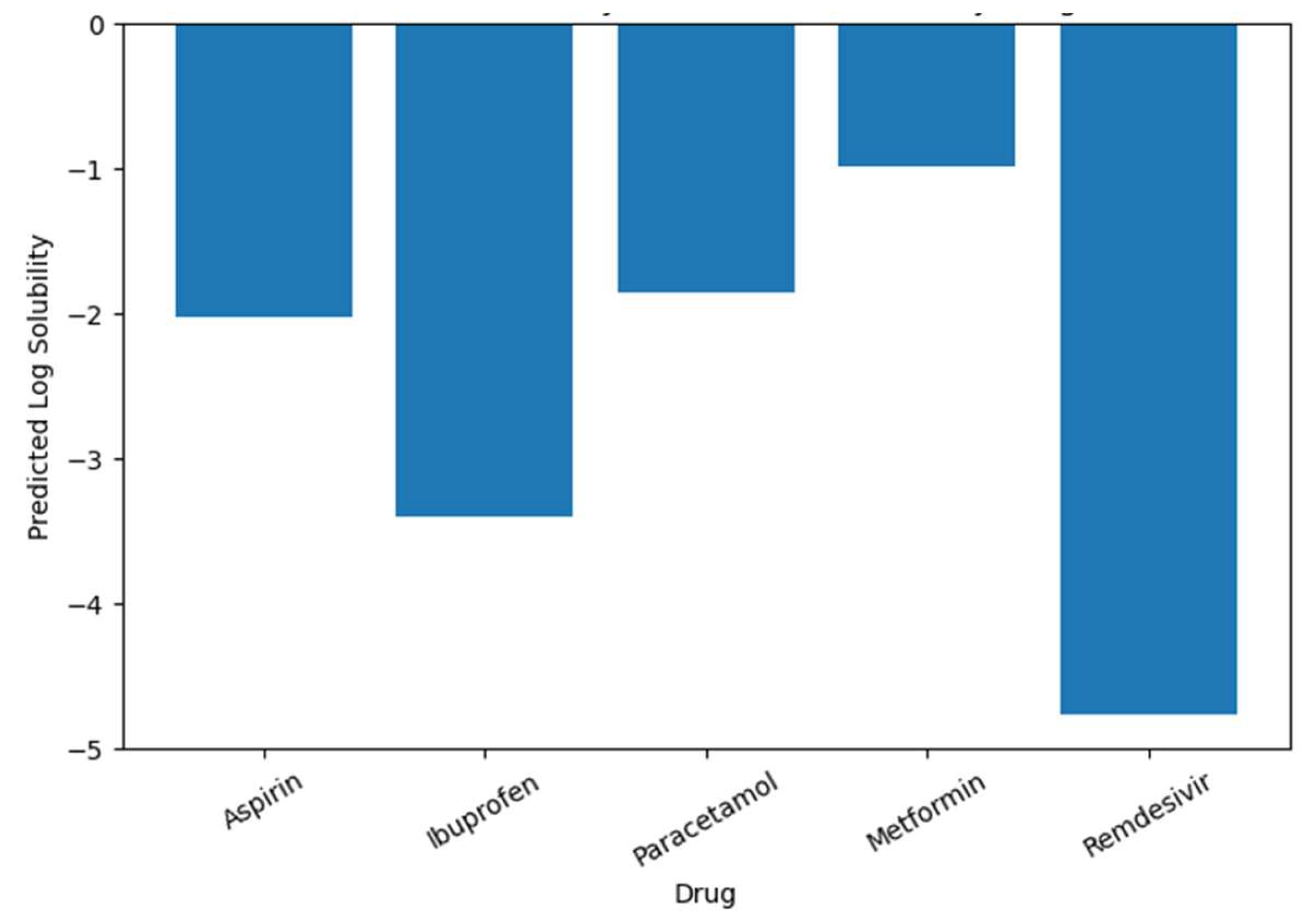
- We performed a case study analysis also where we show how AI-based prediction models can assist in screening and evaluating drug molecules such as Aspirin, Ibuprofen, Paracetamol, Metformin, and Remdesivir for supporting early-stage decision-making in drug discovery.
2. Literature Review
2.1. The Role of Proteomics in Disease Mechanisms and Drug Discovery.
2.2. Artificial Intelligence in Multi-Omics Data Analysis
2.3. De Novo Drug Design Using Artificial Intelligence
2.4. Databases in AI-Driven Drug Discovery
2.5. Challenges of AI-Driven Drug Discovery
3. Case Study
4. Methodology
4.1. Dataset Preparation
4.2. Feature Extraction
- Molecular Weight (MolWt)
- Octanol–Water Partition Coefficient (LogP)
- Topological Polar Surface Area (TPSA)
- Number of Hydrogen Bond Donors
- Number of Hydrogen Bond Acceptors
- Number of Rotatable Bonds
- Ring Count
4.3. Model Development and Evaluation
- Random Forest Regressor
- Gradient Boosting Regressor
- Support Vector Regressor (SVR)
- Root Mean Square Error (RMSE): This metric measures the average magnitude of prediction errors
- Coefficient of Determination (R2 score): This metric indicates the proportion of variance in the target variable explained by the model
4.4. Case Study Analysis
5. Results
| Model | RMSE | R2 Score |
| Gradient Boosting | 0.787 | 0.869 |
| Random Forest | 0.804 | 0.863 |
| SVR | 1.243 | 0.673 |
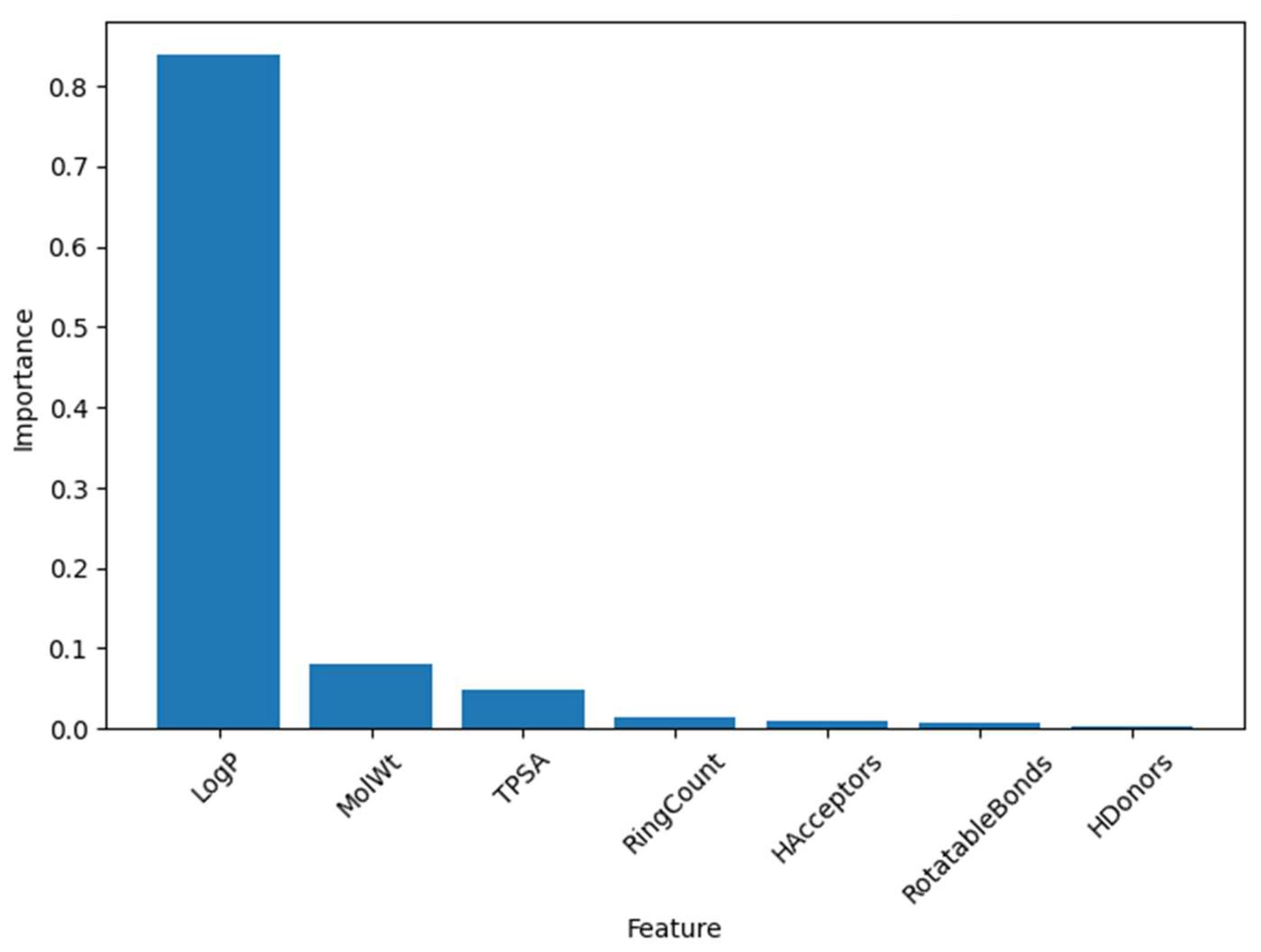
5.1. Feature Importance Analysis
5.2. Case Study Results
6. Discussion
7. Conclusion
References
- Vamathevan, J.; Clark, D.; Czodrowski, P.; Dunham, I.; Ferran, E.; Lee, G.; Li, B.; Madabhushi, A.; Shah, P.; Spitzer, M.; Zhao, S. Applications of machine learning in drug discovery and development. Nature Reviews Drug Discovery 2019, 18, 463–477, https://www.nature.com/articles/s41573-019-0024-5. [Google Scholar] [CrossRef] [PubMed]
- Schneider, P.; Walters, W.P.; Plowright, A.T.; et al. Rethinking drug design in the artificial intelligence era. Nature Reviews Drug Discovery 2020, 19, 353–364, https://www.nature.com/articles/s41573-019-0050-3. [Google Scholar] [CrossRef] [PubMed]
- Pushpakom, S.; Iorio, F.; Eyers, P.A.; et al. Drug repurposing: progress, challenges, and recommendations. Nature Reviews Drug Discovery 2019, 18, 41–58, https://www.nature.com/articles/nrd.2018.168. [Google Scholar] [CrossRef]
- Ray, S.K.; Pawlikowski, K.; Sirisena, H. (2008, October). A fast MAC-layer handover for an IEEE 802.16 e-based WMAN. In International Conference on Access Networks (pp. 102–117). Berlin, Heidelberg: Springer Berlin Heidelberg.
- Mak, K.K.; Pichika, M.R. Artificial intelligence in drug development: present status and future prospects. Drug Discovery Today 2019, 24, 773–780. [Google Scholar] [CrossRef]
- Xu, Y.; Liu, X.; Cao, X.; Huang, C.; Liu, E.; Qian, S.; Liu, X.; Wu, Y.; Dong, F.; Qiu, C.; et al. Artificial intelligence: A powerful paradigm for scientific research. The innovation 2021, 2, 100179. [Google Scholar] [CrossRef]
- Samaras, V.; Daskapan, S.; Ahmad, R.; Ray, S.K. (2014, November). An enterprise security architecture for accessing SaaS cloud services with BYOD. In 2014 Australasian Telecommunication Networks and Applications Conference (ATNAC) (pp. 129–134). IEEE.
- Solanki, P.; Baldaniya, D.; Jogani, D.; Chaudhary, B.; Shah, M.; Kshirsagar, A. Artificial intelligence: New age of transformation in petroleum upstream. Petroleum Research 2022, 7, 106–114. [Google Scholar] [CrossRef]
- Pandey, M.; Fernandez, M.; Gentile, F.; Isayev, O.; Tropsha, A.; Stern, A.C.; Cherkasov, A. The transformational role of GPU computing and deep learning in drug discovery. Nature Machine Intelligence 2022, 4, 211–221. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589, https://www.nature.com/articles/s41586-021-03819-2. [Google Scholar] [CrossRef]
- Esteva, A.; Robicquet, A.; Ramsundar, B.; et al. A guide to deep learning in healthcare. Nature Medicine 2019, 25, 24–29. [Google Scholar] [CrossRef]
- Dey, K.; Ray, S.; Bhattacharyya, P.K.; Gangopadhyay, A.; Bhasin, K.K.; Verma, R.D. Salicyladehyde 4-methoxybenzoylhydrazone and diacetylbis (4-methoxybenzoylhydrazone) as ligands for tin, lead and zirconium. J. Indian Chem. Soc. 1985, 62. [Google Scholar]
- Mohs, R.C.; Greig, N.H. Drug discovery and development: Role of basic biological research. Alzheimer's & Dementia: Translational Research & Clinical Interventions 2017, 3, 651–657. [Google Scholar] [CrossRef]
- De Moor, G.; Sundgren, M.; Kalra, D.; Schmidt, A.; Dugas, M.; Claerhout, B.; Karakoyun, T.; Ohmann, C.; Lastic, P.-Y.; Ammour, N.; et al. Using electronic health records for clinical research: the case of the EHR4CR project. Journal of biomedical informatics 2015, 53, 162–173. [Google Scholar] [CrossRef]
- Anwesa Chaudhuri, A.C.; Sanjib Ray, S.R. (2015). Antiproliferative activity of phytochemicals present in aerial parts aqueous extract of Ampelocissus latifolia (Roxb.) Planch. on apical meristem cells.
- Ray, S.K.; Sirisena, H.; Deka, D. (2013, October). LTE-Advanced handover: An orientation matching-based fast and reliable approach. In 38th annual IEEE conference on local computer networks (pp. 280–283). IEEE.
- Wilson, S.; Filipp, F.V. A network of epigenomic and transcriptional cooperation encompassing an epigenomic master regulator in cancer. Npj Systems Biology and Applications 2018, 4, 24. [Google Scholar] [CrossRef] [PubMed]
- Muzafar, S.; Jhanjhi, N.Z. (2020). Success stories of ICT implementation in Saudi Arabia. In Employing Recent Technologies for Improved Digital Governance (pp. 151–163). IGI Global Scientific Publishing.
- Hamet, P.; Tremblay, J. Artificial intelligence in medicine. Metabolism 2017, 69, S36–S40. [Google Scholar] [CrossRef] [PubMed]
- Curioni-Fontecedro, A. A new era of oncology through artificial intelligence. ESMO open 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Jabeen, T.; Jabeen, I.; Ashraf, H.; Ullah, A.; Jhanjhi, N.Z.; Ghoniem, R.M.; Ray, S.K. Smart wireless sensor technology for healthcare monitoring system using cognitive radio networks. Sensors 2023, 23, 6104. [Google Scholar] [CrossRef]
- Ivanisevic, T.; Sewduth, R.N. Multi-omics integration for the design of novel therapies and the identification of novel biomarkers. Proteomes 2023, 11, 3. [Google Scholar] [CrossRef]
- Pun, F.W.; Ozerov, I.V.; Zhavoronkov, A. AI-powered therapeutic target discovery. Trends in Pharmacological Sciences 2023, 44, 409–423. [Google Scholar] [CrossRef]
- Rasooly, D.; Peloso, G.M.; Pereira, A.C.; et al. Genome-wide association analysis and Mendelian randomization proteomics identify drug targets for heart failure. Nat. Commun. 2023, 14, 1–15. [Google Scholar] [CrossRef]
- Zhang, X.; Wu, F.; Yang, N.; Zhan, X.; Liao, J.; Mai, S. In silico methods for identification of potential therapeutic targets. Interdiscip. Sci. Comput. Life Sci. 2022, 14, 285–310. [Google Scholar] [CrossRef]
- Kaur, N.; Verma, S.; Jhanjhi, N.Z.; Singh, S.; Ghoniem, R.M.; Ray, S.K. Enhanced QoS-aware routing protocol for delay sensitive data in Wireless Body Area Networks. IEEE Access 2023, 11, 106000–106012. [Google Scholar] [CrossRef]
- Zhao, J.H.; Stacey, D.; Eriksson, N.; et al. Genetics of circulating inflammatory proteins identifies drivers of immune-mediated disease risk and therapeutic targets. Nat. Immunol. 2023, 24, 1540–1551. [Google Scholar] [CrossRef] [PubMed]
- Zou, M.; Zhou, H.; Gu, L.; Zhang, J.; Fang, L. Therapeutic target identification and drug discovery driven by chemical proteomics. Biology 2024, 13, 555. [Google Scholar] [CrossRef] [PubMed]
- Molajafari, F.; Li, T.; Abbasichaleshtori, M.; et al. Computational screening for prediction of co-crystals: Method comparison and experimental validation. CrystEngComm 2024, 26, 1620–1636. [Google Scholar] [CrossRef]
- Carvalho, V.; Gonçalves, I.M.; Rodrigues, N.; et al. Numerical evaluation and experimental validation of fluid flow behavior within an organ-on-a-chip model. Comput. Methods Programs Biomed. 2024, 243, 107883. [Google Scholar] [CrossRef]
- Shang, L.; Wang, Y.; Li, J.; et al. Mechanism of Sijunzi Decoction in the treatment of colorectal cancer based on network pharmacology and experimental validation. J. Ethnopharmacol. 2023, 302, 115876. [Google Scholar] [CrossRef]
- Vasconcelos, D.; Chaves, B.; Albuquerque, A.; Andrade, L. Development of new potential inhibitors of β1 integrins through in silico methods—screening and computational validation. Life 2022, 12, 932. [Google Scholar] [CrossRef]
- Muzammal, S.M.; Murugesan, R.K.; Jhanjhi, N.Z.; Jung, L.T. (2020, October). SMTrust: Proposing trust-based secure routing protocol for RPL attacks for IoT applications. In 2020 International Conference on Computational Intelligence (ICCI) (pp. 305–310). IEEE.
- Ayon, S.I.; Islam, M.M.; Hossain, M.R. Coronary artery heart disease prediction: A comparative study of computational intelligence techniques. IETE J. Res. 2022, 68, 2488–2507. [Google Scholar] [CrossRef]
- Crucitti, D.; Pérez Míguez, C.; Díaz Arias, J.Á.; et al. De novo drug design through artificial intelligence: An introduction. Front. Hematol. 2024, 3, 1305741. [Google Scholar] [CrossRef]
- Azeem, M.; Ullah, A.; Ashraf, H.; Jhanjhi, N.Z.; Humayun, M.; Aljahdali, S.; Tabbakh, T.A. Fog-oriented secure and lightweight data aggregation in iomt. IEEE Access 2021, 9, 111072–111082. [Google Scholar] [CrossRef]
- Khater, T.; Alkhatib, S.A.; AlShehhi, A.; Pitsalidis, C. Generative artificial intelligence based models optimization towards molecule design enhancement. J. Chemin 2025, 17, 116. [Google Scholar] [CrossRef] [PubMed]
- Xie, S.; Zhu, H.; Huang, N. AI-Designed molecules in drug discovery: Structural novelty evaluation and implications. J. Chem. Inf. Model. 2025, 65, 8924–8933. [Google Scholar] [CrossRef] [PubMed]
- Hunter, F.M.I.; Ioannidis, H.; Bento, A.P.; et al. Drug and clinical candidate drug data in ChEMBL. J. Med. Chem. 2025, 68, 19800–19827. [Google Scholar] [CrossRef] [PubMed]
- Shah, I.A.; Jhanjhi, N.Z.; Amsaad, F.; Razaque, A. (2022). The role of cutting-edge technologies in Industry 4.0. In Cyber Security Applications for Industry 4.0 (pp. 97–109). Chapman and Hall/CRC.
- Kim, S.; Bolton, E.E. (2024). PubChem: A large-scale public chemical database for drug discovery. Wiley.
- Lee, S.; Abdullah, A.; Jhanjhi, N.Z. A review on honeypot-based botnet detection models for smart factory. International Journal of Advanced Computer Science and Applications 2020, 11. [Google Scholar] [CrossRef]
- Lyubishkin, N.R.; Kardash, O.V.; Klenina, O.V. (2022). Virtual databases for drug discovery.
- de Azevedo, D.Q.; Castilho, R.O.; Gómez-García, A. (2025). Molecular databases in computer-aided drug design. Springer.
- Brohi, S.N.; Jhanjhi, N.Z.; Brohi, N.N.; Brohi, M.N. (2023). Key applications of state-of-the-art technologies to mitigate and eliminate COVID-19. Authorea Preprints.
- Blanco-Gonzalez, A.; Cabezon, A.; Seco-Gonzalez, A.; et al. The role of AI in drug discovery: Challenges, opportunities, and strategies. Pharmaceuticals 2023, 16, 891. [Google Scholar] [CrossRef]
- Pun, F.W.; Ozerov, I.V.; Zhavoronkov, A. AI-powered therapeutic target discovery. Trends in Pharmacological Sciences 2023, 44, 561–572. [Google Scholar] [CrossRef]
- You, Y.; Lai, X.; Pan, Y.; Zheng, H.; Vera, J.; Liu, S. Artificial intelligence in cancer target identification and drug discovery. Signal Transduct. Target. Ther. 2022, 7, 1–24. [Google Scholar] [CrossRef]
- Zhang, X.; et al. In silico methods for identification of potential therapeutic targets. Interdiscip. Sci. Comput. Life Sci. 2022, 14, 285–310. [Google Scholar] [CrossRef]
- Schneider, P.; Walters, W.P.; Plowright, A.T.; et al. Rethinking drug design in the artificial intelligence era. Nat. Rev. Drug Discov. 2020, 19, 353–364. [Google Scholar] [CrossRef]
- Vamathevan, J.; et al. Applications of machine learning in drug discovery and development. Nat. Rev. Drug Discov. 2019, 18, 463–477. [Google Scholar] [CrossRef]
- Zhavoronkov, A.; et al. Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nat. Biotechnol. 2019, 37, 1038–1040. [Google Scholar] [CrossRef]
- Mak, K.K.; Pichika, M.R. Artificial intelligence in drug development: present status and future prospects. Drug Discovery Today. Drug Discov. Today 2019, 24, 773–780. [Google Scholar] [CrossRef]
- Topol, E. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 2019, 25, 44–56. [Google Scholar] [CrossRef]
- Pushpakom, S.; et al. Drug repurposing: progress, challenges and recommendations. Nature Reviews Drug Discovery 2019. [CrossRef]
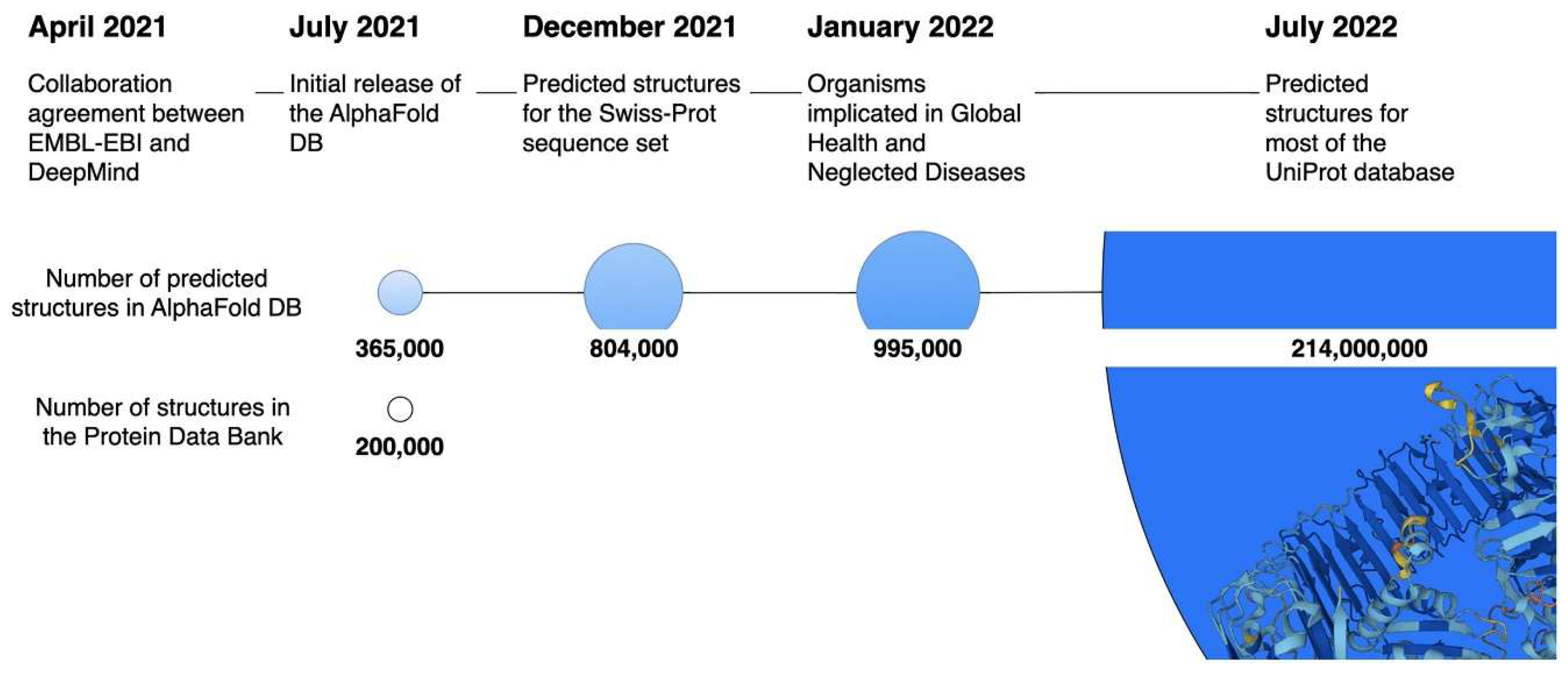
- Varadi, M.; Bertoni, D.; Magana, P.; Paramval, U.; Pidruchna, I.; Radhakrishnan, M.; Tsenkov, M.; Nair, S.; Mirdita, M.; Yeo, J.; et al. AlphaFold Protein Structure Database in 2024: Providing structure coverage for over 214 million protein sequences. Nucleic Acids Research 2024, 52, D368–D375. [Google Scholar] [CrossRef]
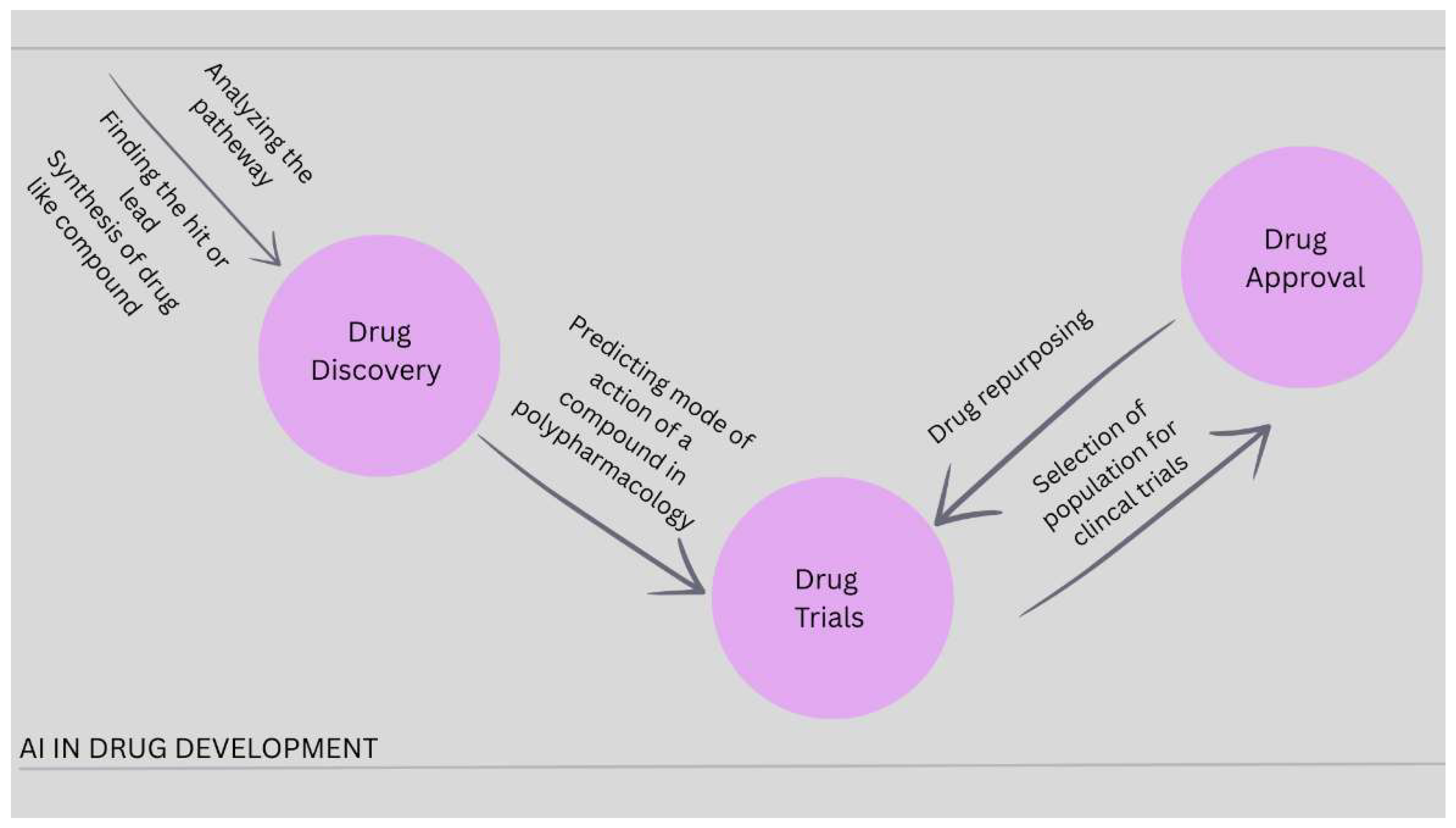
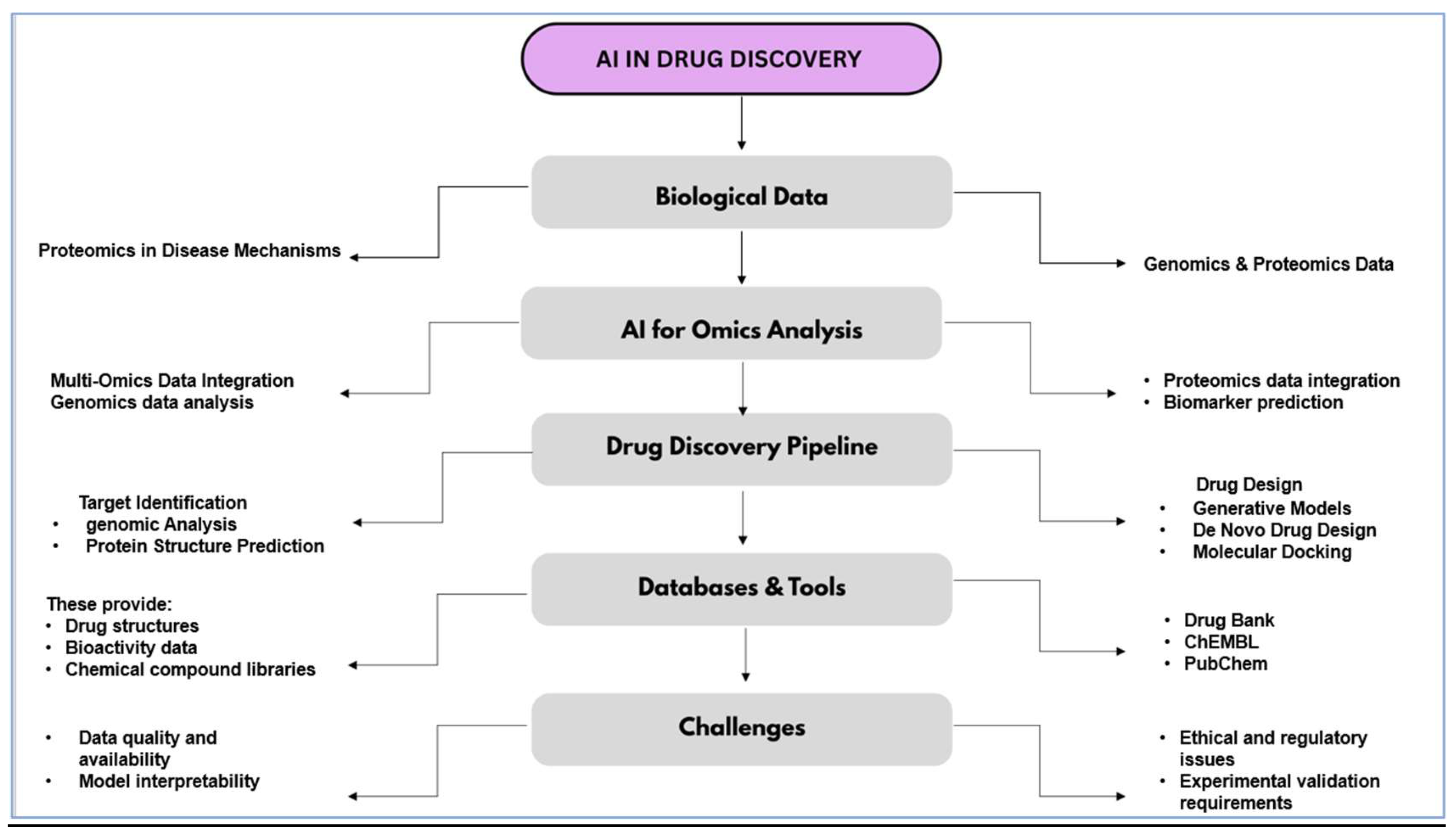
| Drug Discovery Stage | AI Application | Description | References |
| Target Identification | Genomic and proteomic data analysis | AI models analyze large-scale genomic and proteomic datasets to identify disease-associated genes, proteins, and potential therapeutic targets. | [34,35] |
| Target Validation | Biological pathway analysis | Machine learning algorithms validate targets by predicting interactions taking place between genes, proteins, and diseases. | [36] |
| Hit Identification | Virtual screening | AI screens large chemical libraries this saves time and cost. | [37] |
| Lead Optimization | Molecular property prediction | Deep learning models predict molecular properties that enhance drug efficacy and selectivity. | [38] |
| Drug Design | De novo molecule generation | Generative AI designs novel drug molecules with desired pharmacological properties. | [39] |
| Toxicity Prediction | ADMET prediction | AI evaluates absorption, distribution, metabolism, excretion, and toxicity profiles to improve safety. | [40] |
| Clinical Trials | Patient recruitment and trial optimization | AI analyzes patient data and electronic health records to recruit suitable participants. | [41] |
| Drug Repurposing | Identification of new therapeutic uses | AI analyzes biomedical literature and drug databases. | [42] |
| Feature | Importance Score |
| LogP | 0.837791 |
| MolWt | 0.080135 |
| TPSA | 0.049242 |
| RingCount | 0.014979 |
| HAcceptors | 0.009607 |
| RotatableBonds | 0.006056 |
| HDonors | 0.002190 |
| Drug | Predicted Log Solubility |
| Aspirin | -2.02 |
| Ibuprofen | -3.40 |
| Paracetamol | -1.86 |
| Metformin | -0.99 |
| Remdesivir | -4.76 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




