Submitted:
17 March 2026
Posted:
19 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Literature Search Strategy
2.2. Inclusion and Exclusion Criteria
3. Results
3.1. Extracellular Vesicles (EVs)
3.1.1. Biology and Classification
3.1.2. EV Cargo as Rejection Biomarkers
3.1.3. Analytical Methodologies
3.2. Donor-Derived Cell-Free DNA (ddcfDNA)
3.2.1. Biological Basis and Analytical Platforms
3.2.2. Clinical Evidence
3.2.3. Limitations and Confounders
3.3. Donor-Specific Antibodies (DSA)
3.3.1. Immunobiology of DSA-Mediated Injury
3.3.2. DSA Characterization and Risk Stratification
3.3.3. Current Monitoring Guidelines
3.4. Combined Multi-Biomarker Approach
3.4.1. Rationale for Integration
3.4.2. Key Clinical Studies
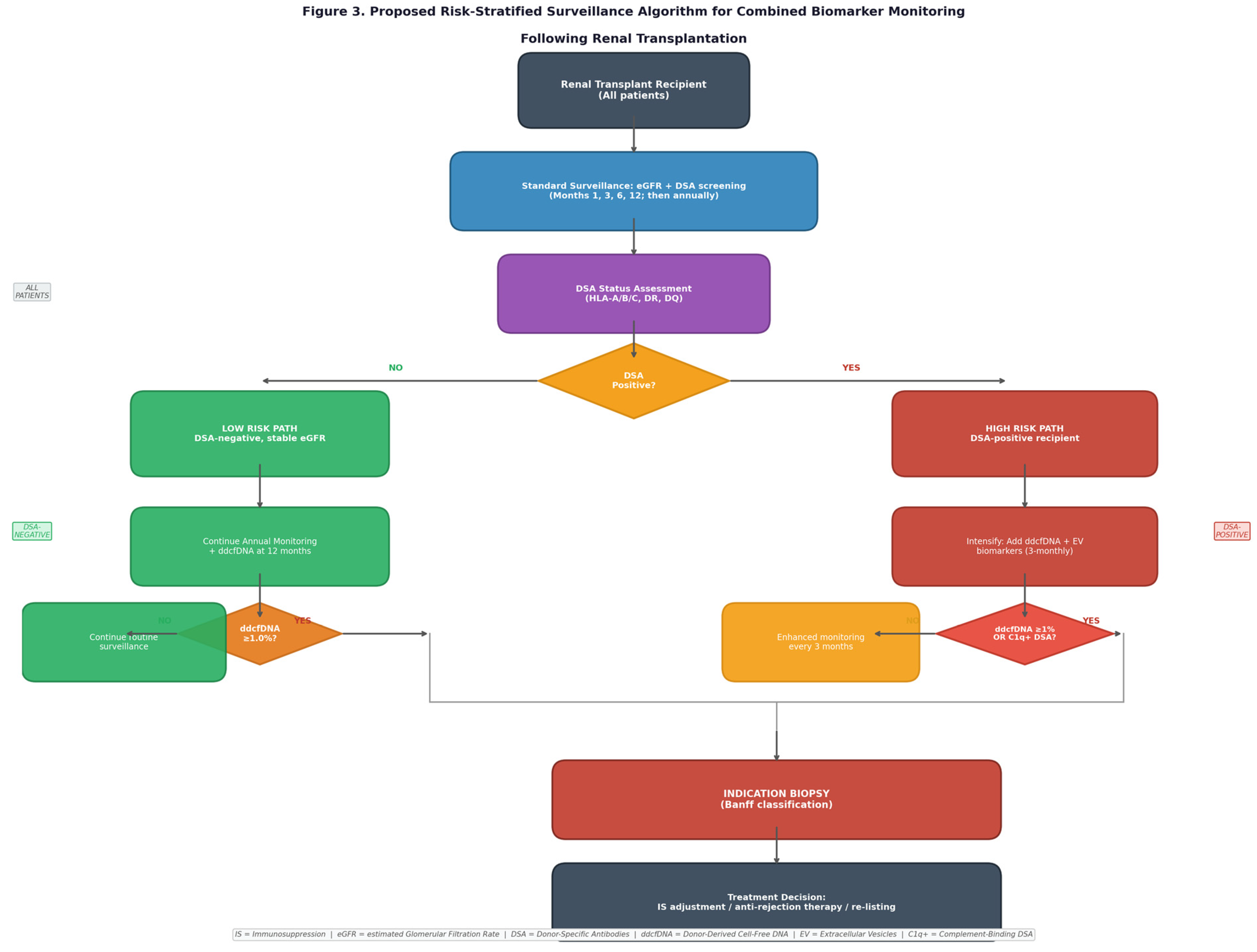
3.4.3. Proposed Surveillance Algorithm
- All three biomarkers negative: Continue routine surveillance at standard intervals.
- DSA positive only (ddcfDNA <1%, EVs normal): Increase monitoring frequency to monthly; assess adherence and immunosuppression trough levels; consider protocol biopsy within 4–6 weeks.
- ddcfDNA ≥1% only (DSA negative, EVs normal): Evaluate for non-immunological injury (BKPyVN screening, CNI trough levels, urinalysis); repeat ddcfDNA in 2–4 weeks.
- EV panel abnormal only: Enhanced surveillance at 4-week intervals; evaluate for subclinical tubular or glomerular injury.
- Any two biomarkers positive: Protocol biopsy within 2 weeks; nephrology review within 72 hours.
- All three biomarkers positive: Urgent biopsy within 48–72 hours; consider empirical immunosuppression adjustment while awaiting histology.
4. Discussion
5. Conclusions
| Biomarker | Current Limitations | Unresolved Challenges | Future Research Priorities |
|---|---|---|---|
| EVs | No standardized clinical-grade isolation protocol; high pre-analytical variability; no FDA clearance; complex and expensive workflow (proteomics/sequencing) | Identifying validated EV subpopulations with clinical-grade specificity; distinguishing donor-derived from recipient-derived EVs; reproducibility across platforms | Microfluidic point-of-care EV analysis; standardized urinary EV proteomics panels; MISEV2023 clinical translation; donor EV labeling strategies; health-economic evaluation |
| ddcfDNA | Elevated in non-immune injury (BKPyVN, ischemia, CNI toxicity, UTI); limited PPV (~40–50%); cost ($1,500–2,800/test); fraction confounded by total cfDNA background | Distinguishing rejection from non-immunological injury; optimal absolute vs. fractional thresholds; standardization of pre-analytical variables across platforms | cfDNA tissue-of-origin methylation mapping; combined cfRNA + cfDNA multi-analyte analysis; cost reduction via targeted sequencing panels; integration with molecular biopsy |
| DSA | Does not confirm active tissue injury; non-HLA antibodies (AT1R, ETAR, MICA) incompletely characterized; inter-laboratory MFI variability; no consensus on clinical intervention threshold | Defining MFI thresholds for clinical intervention; automated eplet analysis; integrating DSA trajectory data into clinical decisions; functional assays beyond C1q/C3d binding | Comprehensive non-HLA antibody panels; AI-based DSA trajectory modeling; NK cell activation assays; endothelial cell crossmatch assays; immunosuppression tailoring by DSA profile |
|
Combined Panel |
No prospective RCT demonstrating improved clinical outcomes with combined monitoring; complex and costly multi-assay workflow; health-economic evidence absent; decision algorithm not prospectively validated | Optimal biomarker weighting in composite risk score; defining intervention thresholds for each biomarker combination; regulatory pathway for multi-analyte panel claims | BEST and COSMOS-KT prospective randomized trials; AI/ML integration for panel interpretation and risk scoring; integration with digital pathology; clinician decision support tools; reimbursement pathway development |
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Declaration of Generative AI and AI-Assisted Technologies in the Writing Process
Abbreviations
References
- Lentine, KL; Smith, JM; Hart, A; et al. OPTN/SRTR 2020 Annual Data Report: Kidney. Am J Transplant. 2022, 22 (Suppl 2), 21–136. [Google Scholar] [CrossRef]
- Sellarés, J; de Freitas, DG; Mengel, M; et al. Understanding the causes of kidney transplant failure: the dominant role of antibody-mediated rejection and nonadherence. Am J Transplant. 2012, 12(2), 388–399. [Google Scholar] [CrossRef]
- Loupy, A; Haas, M; Roufosse, C; et al. The Banff 2019 Kidney Meeting Report (I): Updates on and clarification of criteria for T cell- and antibody-mediated rejection. Am J Transplant. 2020, 20(9), 2318–2331. [Google Scholar] [CrossRef]
- Gaston, RS; Fieberg, A; Helgeson, ES; et al. Late Graft Loss After Kidney Transplantation: Is “Death With Function” Really Death With a Functioning Allograft? Transplantation 2020, 104(7), 1483–1490. [Google Scholar] [CrossRef]
- Loga, L; Dican, L; Matei, HV; Mărunțelu, I; Constantinescu, I. Relevant biomarkers of kidney allograft rejection. J Med Life 2022, 15(11), 1330–1333. [Google Scholar] [CrossRef] [PubMed]
- Lazarou, C; Moysidou, E; Christodoulou, M; et al. Protocol Biopsies in Kidney Transplant Recipients: Current Practice After Much Discussion. Biomedicines 2025, 13(7), 1660. [Google Scholar] [CrossRef] [PubMed]
- Connal, S; Cameron, JM; Sala, A; et al. Liquid biopsies: the future of cancer early detection. J Transl Med. 2023, 21(1), 118. [Google Scholar] [CrossRef]
- Ciferri, MC; Quarto, R; Tasso, R. Extracellular Vesicles as Biomarkers and Therapeutic Tools: From Pre-Clinical to Clinical Applications. Biology (Basel) 2021, 10(5), 359. [Google Scholar] [CrossRef] [PubMed]
- Obrișcă, B; Butiu, M; Sibulesky, L; et al. Combining donor-derived cell-free DNA and donor specific antibody testing as non-invasive biomarkers for rejection in kidney transplantation. Sci Rep. 2022, 12(1), 15061. [Google Scholar] [CrossRef]
- Du, S; Guan, Y; Xie, A; et al. Extracellular vesicles: a rising star for therapeutics and drug delivery. J Nanobiotechnology 2023, 21(1), 231. [Google Scholar] [CrossRef]
- Venturella, M; Criscuoli, M; Carraro, F; Naldini, A; Zocco, D. Interplay between Hypoxia and Extracellular Vesicles in Cancer and Inflammation. Biology (Basel) 2021, 10(7), 606. [Google Scholar] [CrossRef]
- Karpman, D; Ståhl, AL; Arvidsson, I. Extracellular vesicles in renal disease. Nat Rev Nephrol. 2017, 13(9), 545–562. [Google Scholar] [CrossRef]
- Welsh, JA; Goberdhan, DCI; O’Driscoll, L; et al. Minimal information for studies of extracellular vesicles (MISEV2023): From basic to advanced approaches. J Extracell Vesicles 2024, 13(2), e12404. [Google Scholar] [CrossRef]
- Suthanthiran, M; Schwartz, JE; Ding, R; et al. Urinary-cell mRNA profile and acute cellular rejection in kidney allografts. N Engl J Med. 2013, 369(1), 20–31. [Google Scholar] [CrossRef] [PubMed]
- Sigdel, TK; Ng, YW; Lee, S; et al. Perturbations in the urinary exosome in transplant rejection. Front Med (Lausanne) 2015, 1, 57. [Google Scholar] [CrossRef] [PubMed]
- Hricik, DE; Nickerson, P; Formica, RN; et al. Multicenter validation of urinary CXCL9 as a risk-stratifying biomarker for kidney transplant injury. Am J Transplant. 2013, 13(10), 2634–2644. [Google Scholar] [CrossRef]
- Stites, E; Renner, B; Laskowski, J; et al. Complement fragments are biomarkers of antibody-mediated endothelial injury. Mol Immunol. 2020, 118, 142–152. [Google Scholar] [CrossRef]
- Kuo, HH; Fan, R; Dvorina, N; Chiesa-Vottero, A; Baldwin, WM, 3rd. Platelets in early antibody-mediated rejection of renal transplants. J Am Soc Nephrol. 2015, 26(4), 855–863. [Google Scholar] [CrossRef] [PubMed]
- Boštjčančič, E; Večerič-Haler, Ž; Kojc, N. The Role of Immune-Related miRNAs in the Pathology of Kidney Transplantation. Biomolecules 2021, 11(8), 1198. [Google Scholar] [CrossRef]
- Oshikawa-Hori, S; Yokota-Ikeda, N; Sonoda, H; Ikeda, M. Urinary extracellular vesicular release of aquaporins in patients with renal transplantation. BMC Nephrol. 2019, 20(1), 216. [Google Scholar] [CrossRef]
- Witwer, KW; Buzás, EI; Bemis, LT; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J Extracell Vesicles 2013, 2, 20360. [Google Scholar] [CrossRef]
- Oellerich, M; Budde, K; Osmanodja, B; et al. Donor-derived cell-free DNA as a diagnostic tool in transplantation. Front Genet. 2022, 13, 1031894. [Google Scholar] [CrossRef]
- Snyder, TM; Khush, KK; Valantine, HA; Quake, SR. Universal noninvasive detection of solid organ transplant rejection. Proc Natl Acad Sci U S A 2011, 108(15), 6229–6234. [Google Scholar] [CrossRef]
- Grskovic, M; Hiller, DJ; Eubank, LA; et al. Validation of a Clinical-Grade Assay to Measure Donor-Derived Cell-Free DNA in Solid Organ Transplant Recipients. J Mol Diagn 2016, 18(6), 890–902. [Google Scholar] [CrossRef] [PubMed]
- Altuğ, Y; Liang, N; Ram, R; et al. Analytical Validation of a Single-nucleotide Polymorphism-based Donor-derived Cell-free DNA Assay for Detecting Rejection in Kidney Transplant Patients. Transplantation 2019, 103(12), 2657–2665. [Google Scholar] [CrossRef]
- Bloom, RD; Bromberg, JS; Poggio, ED; et al. Cell-Free DNA and Active Rejection in Kidney Allografts. J Am Soc Nephrol. 2017, 28(7), 2221–2232. [Google Scholar] [CrossRef] [PubMed]
- Oellerich, M; Shipkova, M; Asendorf, T; et al. Absolute quantification of donor-derived cell-free DNA as a marker of rejection and graft injury in kidney transplantation. Am J Transplant. 2019, 19(11), 3087–3099. [Google Scholar] [CrossRef] [PubMed]
- Bromberg, JS; Brennan, DC; Taber, DJ; et al. Donor-derived cell-free DNA significantly improves rejection yield in kidney transplant biopsies. Am J Transplant. 2025, 25(12), 2529–2542. [Google Scholar] [CrossRef]
- Bu, L; Gupta, G; Pai, A; et al. Clinical outcomes from the ADMIRAL study. Kidney Int. 2022, 101(4), 793–803. [Google Scholar] [CrossRef]
- Steggerda, JA; Pizzo, H; Garrison, J; et al. Use of a donor-derived cell-free DNA assay to monitor treatment response in pediatric renal transplant recipients with allograft rejection. Pediatr Transplant. 2022, 26(4), e14258. [Google Scholar] [CrossRef]
- Sun, K; Jiang, P; Chan, KC; et al. Plasma DNA tissue mapping by genome-wide methylation sequencing for noninvasive prenatal, cancer, and transplantation assessments. Proc Natl Acad Sci U S A 2015, 112(40), E5503–E5512. [Google Scholar] [CrossRef]
- Loupy, A; Lefaucheur, C. Antibody-Mediated Rejection of Solid-Organ Allografts. N Engl J Med. 2018, 379(12), 1150–1160. [Google Scholar] [CrossRef]
- Wiebe, C; Gibson, IW; Blydt-Hansen, TD; et al. Evolution and clinical pathologic correlations of de novo donor-specific HLA antibody post kidney transplant. Am J Transplant. 2012, 12(5), 1198–1211. [Google Scholar] [CrossRef]
- Abe, T; Murakami, M; Harada, H; et al. Molecular Mechanisms of Antibody-Mediated Rejection and Accommodation in Organ Transplantation. Nephron 2020, 144 (Suppl 1), 2–6. [Google Scholar] [CrossRef]
- Tait, BD; Süsal, C; Gebel, HM; et al. Consensus guidelines on the testing and clinical management issues associated with HLA and non-HLA antibodies in transplantation. Transplantation 2013, 95(1), 19–47. [Google Scholar] [CrossRef] [PubMed]
- Lefaucheur, C; Viglietti, D; Bentlejewski, C; et al. IgG Donor-Specific Anti-Human HLA Antibody Subclasses and Kidney Allograft Antibody-Mediated Injury. J Am Soc Nephrol. 2016, 27(1), 293–304. [Google Scholar] [CrossRef] [PubMed]
- Loupy, A; Lefaucheur, C; Vernerey, D; et al. Complement-binding anti-HLA antibodies and kidney-allograft survival. N Engl J Med. 2013, 369(13), 1215–1226. [Google Scholar] [CrossRef] [PubMed]
- Comoli, P; Cioni, M; Tagliamacco, A; et al. Acquisition of C3d-Binding Activity by De Novo Donor-Specific HLA Antibodies Correlates With Graft Loss in Nonsensitized Pediatric Kidney Recipients. Am J Transplant. 2016, 16(7), 2106–2116. [Google Scholar] [CrossRef]
- Crespo, M; Llinàs-Mallol, L; Redondo-Pachón, D; et al. Non-HLA Antibodies and Epitope Mismatches in Kidney Transplant Recipients With Histological Antibody-Mediated Rejection. Front Immunol. 2021, 12, 703457. [Google Scholar] [CrossRef]
- Lefaucheur, C; Louis, K; Morris, AB; et al. Clinical recommendations for posttransplant assessment of anti-HLA donor-specific antibodies: A Sensitization in Transplantation: Assessment of Risk consensus document. Am J Transplant. 2023, 23(1), 115–132. [Google Scholar] [CrossRef]
- Viglietti, D; Loupy, A; Aubert, O; et al. Dynamic Prognostic Score to Predict Kidney Allograft Survival in Patients with Antibody-Mediated Rejection. J Am Soc Nephrol. 2018, 29(2), 606–619. [Google Scholar] [CrossRef]
- Oellerich, M; Sherwood, K; Keown, P; et al. Liquid biopsies: donor-derived cell-free DNA for the detection of kidney allograft injury. Nat Rev Nephrol. 2021, 17(9), 591–603. [Google Scholar] [CrossRef]
- Halloran, PF; Reeve, J; Madill-Thomsen, KS; et al. Combining Donor-derived Cell-free DNA Fraction and Quantity to Detect Kidney Transplant Rejection. Transplantation 2022, 106(12), 2435–2442. [Google Scholar] [CrossRef]
- Mayer, KA; Doberer, K; Tillgren, A; et al. Diagnostic value of donor-derived cell-free DNA to predict antibody-mediated rejection in donor-specific antibody-positive renal allograft recipients. Transpl Int. 2021, 34(9), 1689–1702. [Google Scholar] [CrossRef]
- Mertens, I; Willems, H; Van Loon, E; et al. Urinary Protein Biomarker Panel for the Diagnosis of Antibody-Mediated Rejection in Kidney Transplant Recipients. Kidney Int Rep. 2020, 5(9), 1448–1458. [Google Scholar] [CrossRef] [PubMed]
- Puliyanda, DP; Swinford, R; Pizzo, H; Garrison, J; De Golovine, AM; Jordan, SC. Donor-derived cell-free DNA (dd-cfDNA) for detection of allograft rejection in pediatric kidney transplants. Pediatr Transplant. 2021, 25(2), e13850. [Google Scholar] [CrossRef] [PubMed]
- Halloran, PF; Madill-Thomsen, KS; Reeve, J. The Molecular Phenotype of Kidney Transplants: Insights From the MMDx Project. Transplantation 2024, 108(1), 45–71. [Google Scholar] [CrossRef]
- Loupy, A; Vernerey, D; Tinel, C; et al. Subclinical Rejection Phenotypes at 1 Year Post-Transplant and Outcome of Kidney Allografts. J Am Soc Nephrol. 2015, 26(7), 1721–1731. [Google Scholar] [CrossRef]
- Mayer, KA; Schrezenmeier, E; Diebold, M; et al. A Randomized Phase 2 Trial of Felzartamab in Antibody-Mediated Rejection. N Engl J Med. 2024, 391(2), 122–132. [Google Scholar] [CrossRef] [PubMed]
- Huang, E; Haas, M; Gillespie, M; et al. An Assessment of the Value of Donor-derived Cell-free DNA Surveillance in Patients With Preserved Kidney Allograft Function. Transplantation 2023, 107(1), 274–282. [Google Scholar] [CrossRef]
- Akifova, A; Budde, K; Amann, K; et al. Donor-derived cell-free DNA monitoring for early diagnosis of antibody-mediated rejection after kidney transplantation: a randomized trial. Nephrol Dial Transplant. 2025, 40(7), 1384–1395. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | Extracellular Vesicles (EVs) | Donor-Derived cfDNA (ddcfDNA) | Donor-Specific Antibodies (DSA) |
|---|---|---|---|
| Biological source | All cell types; tubular, glomerular, endothelial, immune cells | Apoptotic/necrotic graft parenchymal cells | Recipient B cells / plasma cells sensitized to donor HLA |
| Biofluid(s) | Plasma, urine (urine preferred for renal monitoring) | EDTA plasma (cfDNA-stabilizing tubes required) | Serum or EDTA plasma |
| Analyte | Membrane vesicles 40–1000 nm; protein, RNA, lipid cargo | Short dsDNA fragments ~165 bp of donor genomic origin | IgG (subclasses 1–4) directed against HLA class I/II |
| Pathophysiological signal | Cell activation, stress, tubular/endothelial injury, immune effector activity | Active cell death (apoptosis/necrosis) within graft parenchyma | Upstream humoral alloimmune sensitization; risk of ABMR |
| Detection method | NTA, TEM, Western blot (CD9/63/81); proteomics, miRNA-seq, flow cytometry | SNP-based NGS (AlloSure); genotype-informed NGS (Prospera); ddPCR | Single antigen bead Luminex (MFI); C1q/C3d complement-binding assays |
| Diagnostic threshold | Variable, assay- and cargo-dependent | ≥1.0% (DART trial); absolute >10 cp/mL (emerging) | >500 MFI (detection); >5000–10,000 MFI (high-risk); C1q+ threshold lab-dependent |
| Sensitivity (rejection) | 70–85% (EV CXCL9, CD3ε mRNA) | 59–89% (DART, meta-analyses) | 55–80% (ABMR-specific; lower for TCMR) |
| Specificity (rejection) | 72–88% | 73–92% | 75–92% |
| AUC (best reported) | 0.85–0.88 (EV cargo panels) | 0.74–0.82 | 0.79–0.87 (C1q+ DSA for graft loss) |
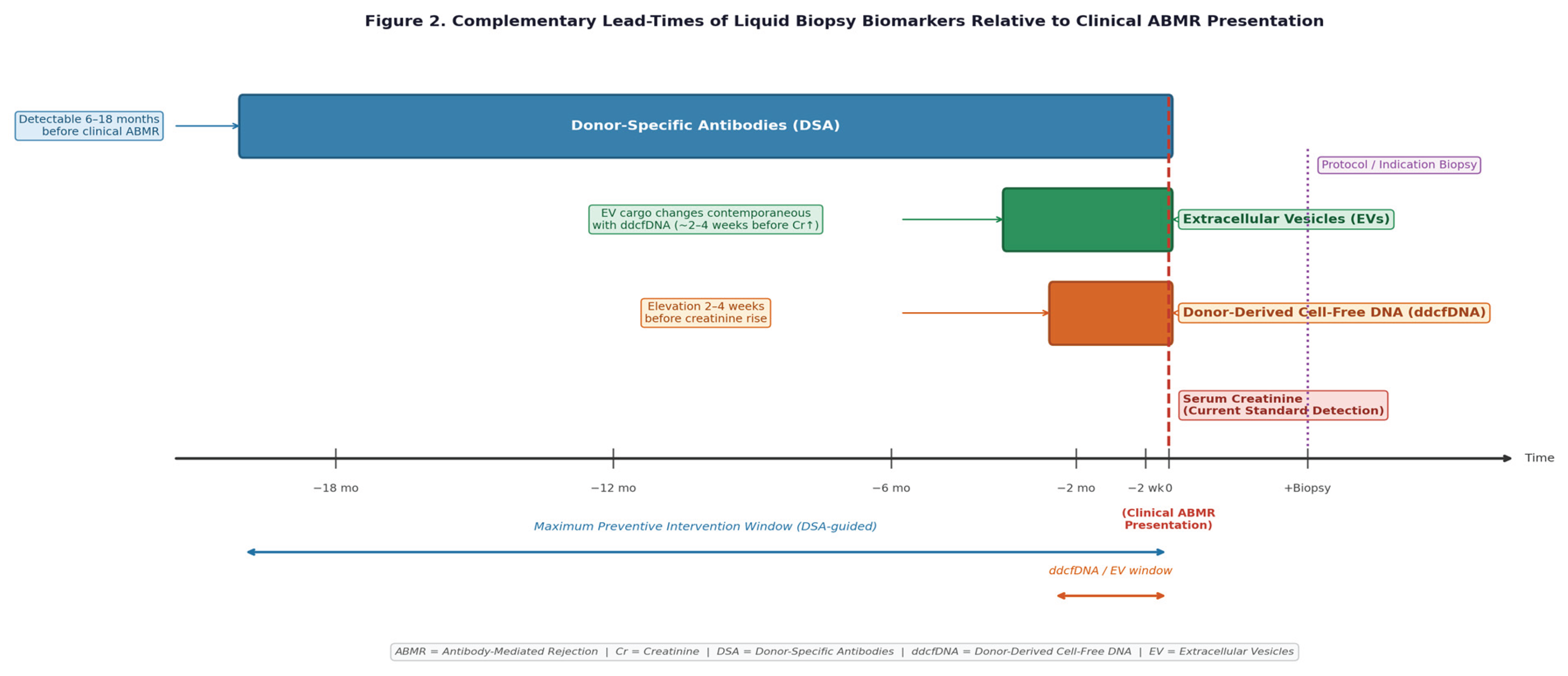
| Lead time vs. creatinine | Days to 1–2 weeks before creatinine rise | 1–4 weeks before clinical presentation | 6–18 months before clinical ABMR (dnDSA) |
| FDA clearance | No (research use only) | Yes (AlloSure 2020, Prospera 2021) | No (lab-developed tests; CAP-accredited labs) |
| Standardization status | Low (MISEV2023 guidelines; no clinical-grade assay) | Moderate (cleared assays; pre-analytical standards required) | Moderate–High (SAB-Luminex standardized; inter-lab MFI variability remains) |
| Approximate cost (USD) | $300–800 (research proteomics/sequencing) | $1,500–2,800 per test (commercial) | $200–600 per panel |
| Key limitation | No standardized clinical assay; complex workflow; pre-analytical variability | Elevated in non-rejection injury (BKPyVN, CNI toxicity, UTI); low PPV (~40–50%) | Does not confirm active injury; non-HLA antibodies not covered; inter-lab MFI variability |
| Best clinical utility | Subclinical tubular/endothelial injury; rejection subtyping; pediatric monitoring | Active rejection detection; ABMR/mixed rejection; serial surveillance | Pre-transplant risk stratification; de novo sensitization monitoring; ABMR diagnosis |
| Study (Year) | Biomarker(s) Evaluated | Study Design / Sample Size | Primary Outcome / Key Finding | AUC (95% CI) / NPV |
|---|---|---|---|---|
| Suthanthiran et al. (CTOT-04), 2013 [14] | Urinary EV CD3ε mRNA | Prospective; n = 85 recipients | Urinary exosomal CD3ε mRNA elevated in TCMR, preceding creatinine by 5–7 days | AUC 0.81 NPV 89% |
| Sigdel et al., 2015 [15] | Urinary EV proteome (CXCL9, CXCL10, GzmB) | Cross-sectional; n = 120 | CXCL9, CXCL10, granzyme B most discriminatory for acute rejection vs. stable function | AUC 0.85 — |
| Loupy et al., 2013 [37] | DSA (C1q-binding) | Prospective multicenter; n = 1016 | C1q+ DSA: 5-year graft survival 54% vs. 93% in C1q-neg; HR 4.78 for graft loss | — HR 4.7 |
| Bloom et al. (DART), 2017 [26] | ddcfDNA (AlloSure) | Prospective multicenter; n = 102 (107 biopsies) | ddcfDNA ≥1.0% identified active rejection; superior to creatinine/eGFR; NPV 84% | AUC 0.74 NPV 84% |
| Oellerich et al., 2019 [27] | ddcfDNA (absolute, copies/mL) | Prospective; n = 217 | Absolute ddcfDNA superior to fractional measurement for ABMR vs. non-rejection injury | AUC 0.82 — |
| Bromberg et al. (KOAR), 2025 [28] | ddcfDNA (AlloSure serial) | Registry; n = 1092 (14 centers) | Serial monitoring changed management in 30%; rejection detected ~3 weeks early | — PPV 45% |
| Mertens et al. (BIOMARGIN), 2020 [45] | ddcfDNA + Urinary EV CXCL9 | Multicenter; n = 388 | ddcfDNA + DSA-MFI combined panel; AUC 0.97 for graft injury in DSA-positive recipients; confirms complementarity of both markers | AUC 0.97 (combined panel) |
| Mayer et al., 2021 [44] | DSA + ddcfDNA + EV proteome | Prospective; n = 210 | Combined panel: sensitivity 89%, specificity 91% for ABMR; all three markers independently predictive | AUC 0.94 NPV 93% |
| Puliyanda et al., 2021 [46] | ddcfDNA (AlloSure) alone | Pediatric cohort; n = 78 | ddcfDNA associated with biopsy-confirmed rejection in pediatric cohort; performance consistent with adult data | Performance consistent with adult data |
| Bu et al. (ADMIRAL), 2022 [29] | ddcfDNA (AlloSure) | Multicenter; n = 315 | ddcfDNA correlated with Banff injury score (r = 0.71) and predicted eGFR recovery post-treatment | AUC 0.79 NPV 87% |
| Halloran et al., 2024 [47] | Blood EV miRNA + molecular biopsy | Prospective; n = 195 | EV miR-142-3p + miR-223-3p enhanced ABMR vs. TCMR discrimination in histologically ambiguous cases | AUC 0.87 — |
| Viglietti et al., 2018 [41] | DSA (serial trajectory, MFI + IgG subclass) | Prospective; n = 412 | Rising DSA MFI trajectory independently predicted ABMR progression regardless of absolute MFI level | — HR 3.2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




