Submitted:
17 March 2026
Posted:
18 March 2026
You are already at the latest version
Abstract
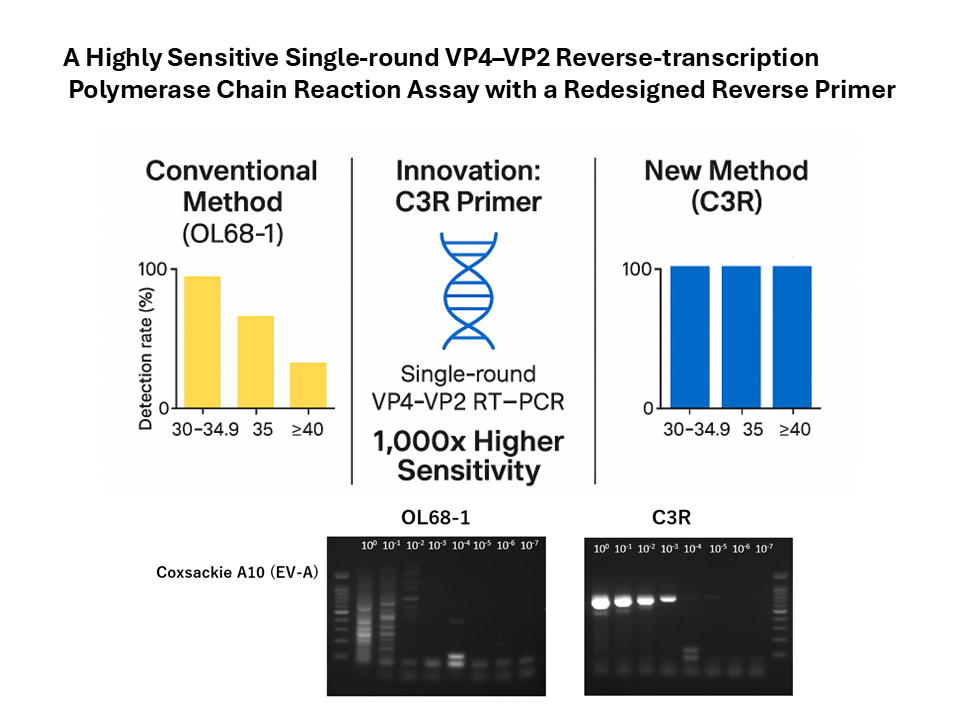
Keywords:
1. Introduction
2. Materials and Methods
2.1. Primers and Probes
2.2. Viral Isolates and Analytical Sensitivity
2.3. Reference Assay for Defining EV Positivity
2.4. Clinical Specimens
2.5. RNA Extraction
2.6. Pan-EV One-Step Real-Time RT-PCR
2.7. Complementary DNA Synthesis
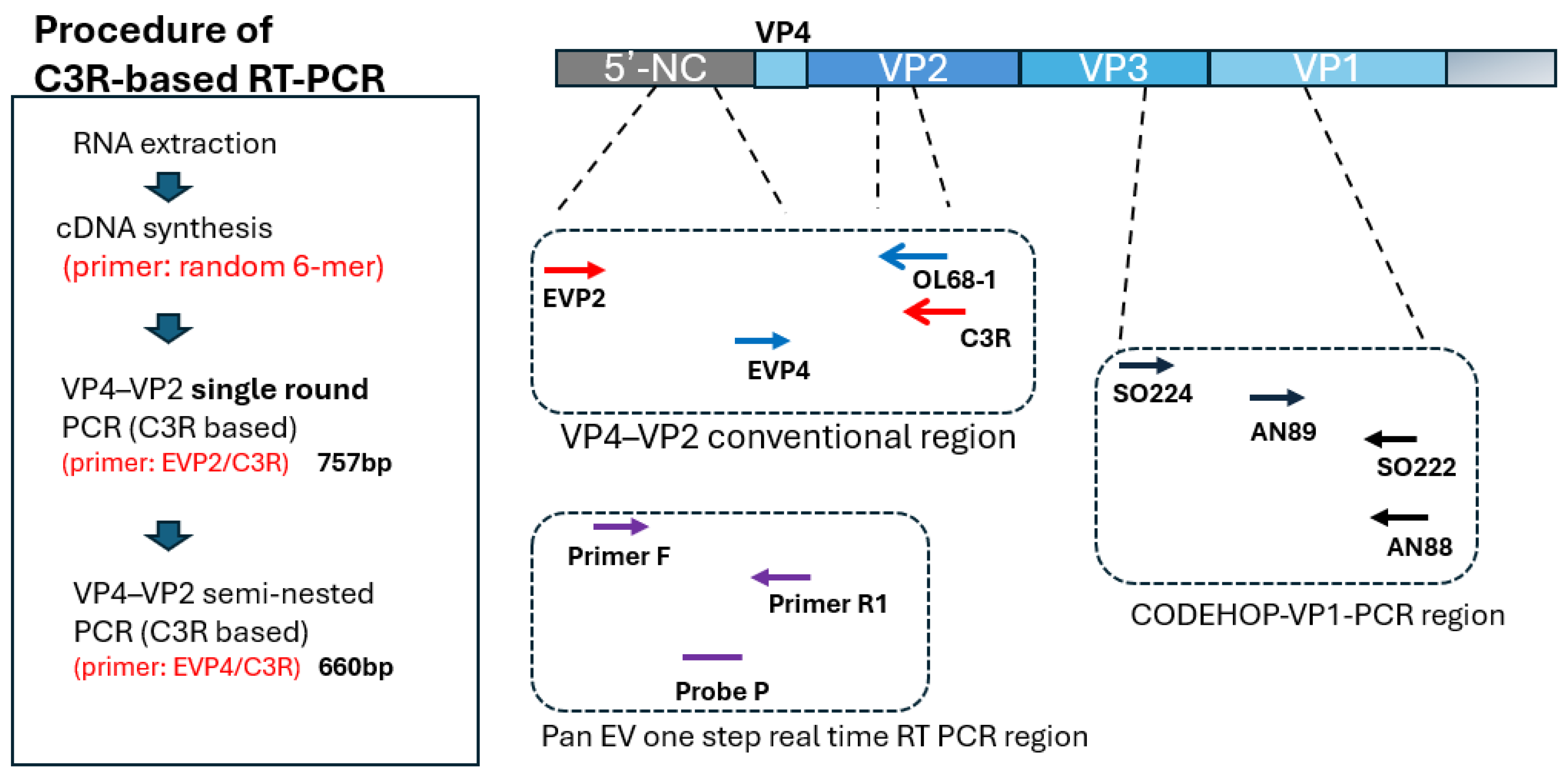
2.8. VP4–VP2 Single-Round PCR (C3R-Based and Conventional OL68-1-Based)
2.9. VP4–VP2 Semi-Nested PCR
2.10. VP1 CODEHOP PCR [17]
2.11. Gel Electrophoresis
2.12. Sanger Sequencing and EV Typing
2.13. In Silico Primer Coverage and 3′-End-Weighted Suitability Scoring (C3R vs OL68-1)
2.13.1. Panel and Alignment
2.13.2. International Union of Pure and Applied Chemistry (IUPAC) handling
2.13.3. Primer–Template Fit Score Calculation
2.13.4. Outputs and Interpretation
3. Results
3.1. In Silico Evaluation of Reverse Primers (Summary)
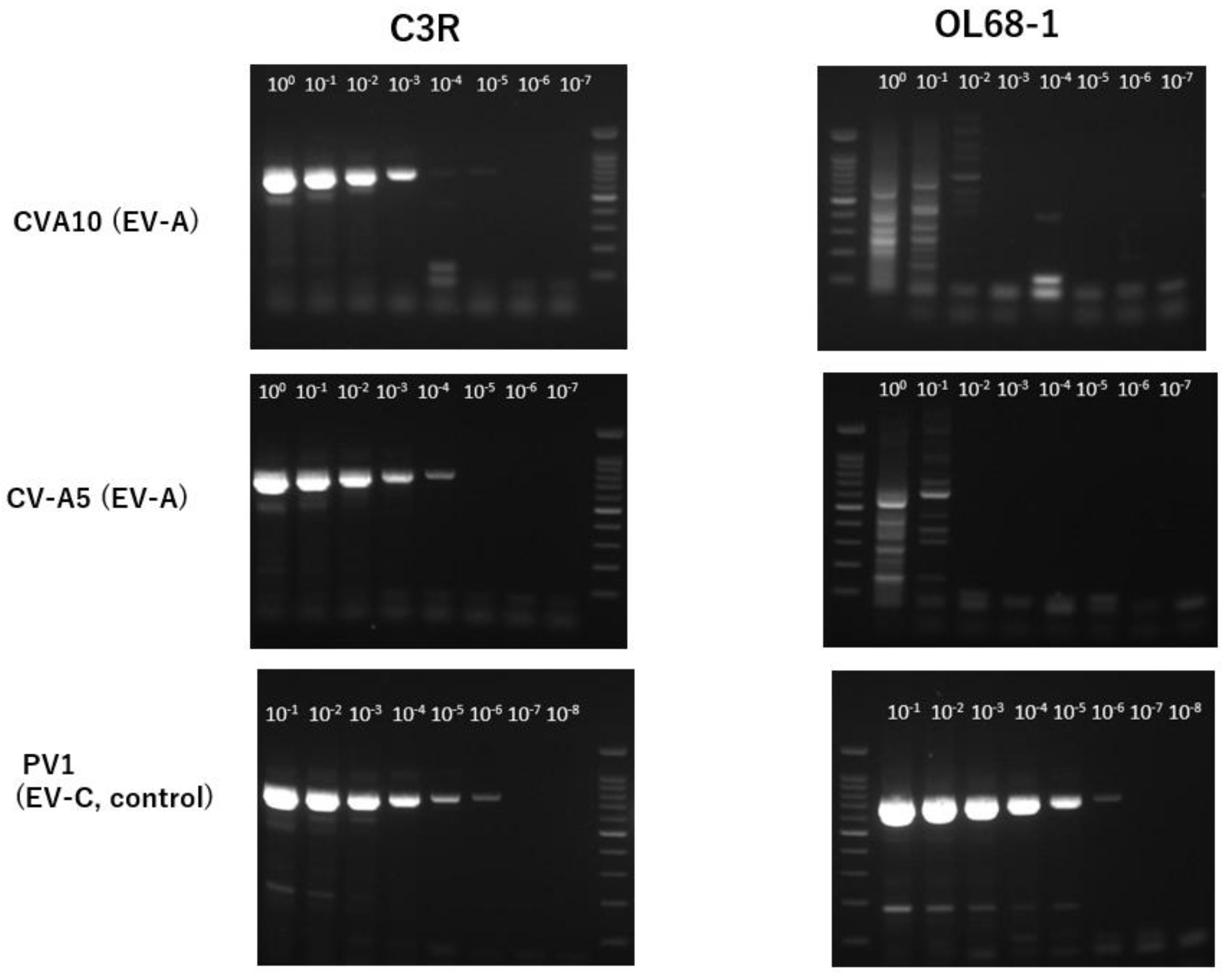
3.2. Analytical Sensitivity with Reference Isolates
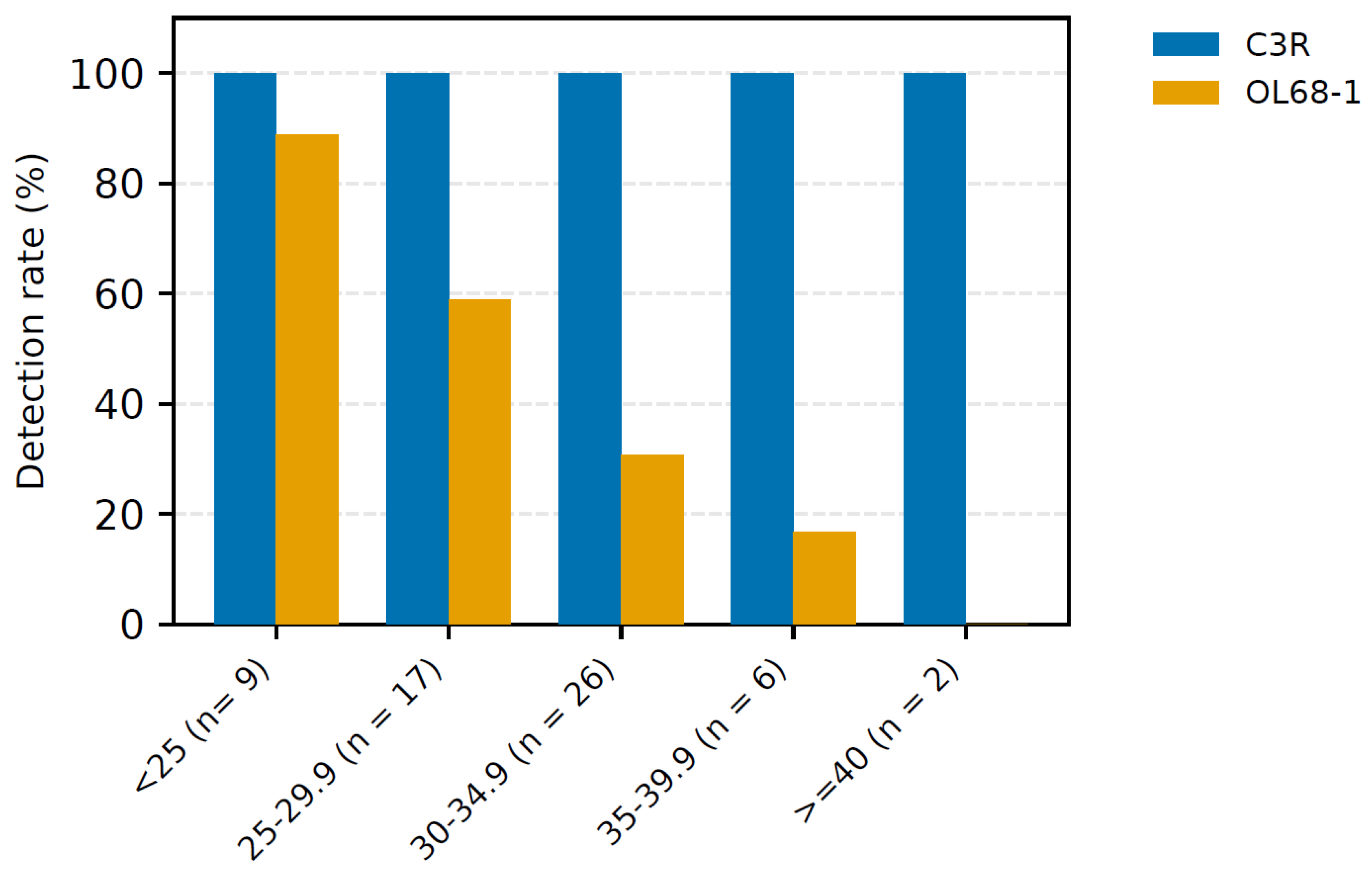
3.3. Clinical Performance in HFMD/Herpangina Cohort
3.4. In Silico Coverage and Fitness of C3R (vs. OL68-1)
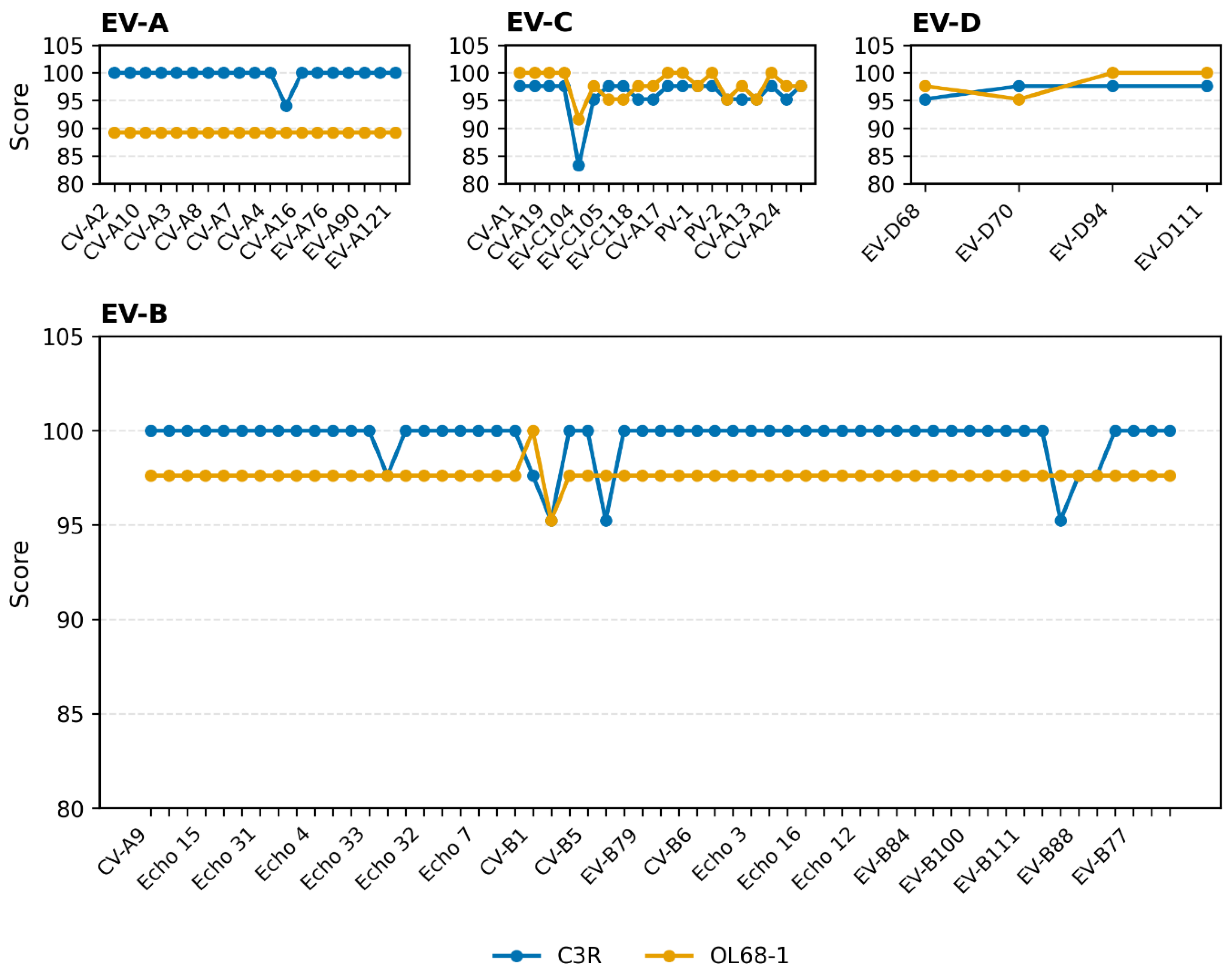
4. Discussion
4.1. Major Findings
4.2. Mechanistic Interpretation Anchored by Standard Strains
4.3. C3R-Based RT-PCR: Single-Round Alternative to Conventional VP4–VP2 RT-PCR
4.4. Future Application to Improve the Conventional VP4–VP2 Manual Protocol
4.5. Practical Implications for Typing Workflows.
4.6. Applicability to One-Step and Real-Time Formats.
4.7. Rationale for Selecting the VP4–VP2 Region and Positioning in Diagnostic Algorithms
4.8. Clinical and Public-Health Significance
4.9. Limitations and Future Directions
Abbreviations (Alphabetical Order)
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xie, Z.; Khamrin, P.; Maneekarn, N.; Kumthip, K. Epidemiology of Enterovirus Genotypes in Association with Human Diseases. Viruses 2024, 16, 1165. [Google Scholar] [CrossRef] [PubMed]
- Jartti, M.; Flodström-Tullberg, M.; Hankaniemi, M.M. Enteroviruses: Epidemic Potential, Challenges and Opportunities with Vaccines. J. Biomed. Sci. 2024, 31, 73. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, G.; Carr, M.J.; Kobayashi, M.; Hanaoka, N.; Fujimoto, T. Enterovirus-Associated Hand-Foot-and-Mouth Disease and Neurological Complications in Japan and the Rest of the World. Int. J. Mol. Sci. 2019, 20, 5201. [Google Scholar] [CrossRef] [PubMed]
- Aswathyraj, S.; Arunkumar, G.; Alidjinou, E.K.; Hober, D. Hand, Foot and Mouth Disease (HFMD): Emerging Epidemiology and the Need for a Vaccine Strategy. Med. Microbiol. Immunol. 2016, 205, 397–407. [Google Scholar] [CrossRef]
- Yu, H.; Li, X.W.; Liu, Q.B.; Deng, H.L.; Liu, G.; Jiang, R.M.; et al. Diagnosis and Treatment of Herpangina: Chinese Expert Consensus. World J. Pediatr. 2020, 16, 129–134. [Google Scholar] [CrossRef]
- Takahashi, S.; Metcalf, C.J.E.; Arima, Y.; Fujimoto, T.; Shimizu, H.; Van Doorn, H.R.; et al. Epidemic Dynamics, Interactions and Predictability of Enteroviruses Associated with Hand, Foot and Mouth Disease in Japan. J. R. Soc. Interface 2018, 15, 20180507. [Google Scholar] [CrossRef]
- Ooi, M.H.; Wong, S.C.; Lewthwaite, P.; Cardosa, M.J.; Solomon, T. Clinical Features, Diagnosis, and Management of Enterovirus 71. Lancet Neurol. 2010, 9, 1097–1105. [Google Scholar] [CrossRef]
- Takechi, M.; Fukushima, W.; Nakano, T.; Inui, M.; Ohfuji, S.; Kase, T.; et al. Nationwide Survey of Pediatric Inpatients with Hand, Foot, and Mouth Disease, Herpangina, and Associated Complications during an Epidemic Period in Japan: Estimated Number of Hospitalized Patients and Factors Associated With Severe Cases. J. Epidemiol. 2019, 29, 354–362. [Google Scholar] [CrossRef]
- Nagai, T.; Hanaoka, N.; Katano, H.; Konagaya, M.; Tanaka-Taya, K.; Shimizu, H.; et al. A Fatal Case of Acute Encephalopathy in a Child due to Coxsackievirus A2 Infection: A Case Report. BMC Infect. Dis. 2021, 21, 1167. [Google Scholar] [CrossRef]
- Ishii, M.; Hoshina, T.; Fujimoto, T.; Hanaoka, N.; Konagaya, M.; Shimbashi, R.; et al. A Pediatric Case of Encephalopathy with Hypoglycemia Induced by Coxsackievirus A4 Infection. Pediatr. Infect. Dis. J. 2024, 43, e124–e126. [Google Scholar] [CrossRef]
- Olive, D.M.; Al-Mufti, S.; Al-Mulla, W.; Khan, M.A.; Pasca, A.; Stanway, G.; Al-Nakib, W. Detection and Differentiation of Picornaviruses in Clinical Samples Following Genomic Amplification. J. Gen. Virol. 1990, 71, 2141–2147. [Google Scholar] [CrossRef]
- Ishiko, H.; Shimada, Y.; Yonaha, M.; Hashimoto, O.; Hayashi, A.; Sakae, K.; Takeda, N. Molecular Diagnosis of Human Enteroviruses by Phylogeny-Based Classification Using the VP4 Sequence. J. Infect. Dis. 2002, 185, 744–754. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, T.; Yoshida, S.; Munemura, T.; Taniguchi, K.; Shinohara, M.; Nishio, O.; et al. Detection and Quantification of Enterovirus 71 Genome from Cerebrospinal Fluid of an Encephalitis Patient by PCR. Jpn. J. Infect. Dis. 2008, 61, 497–499. Available online: https://www.jstage.jst.go.jp/article/yoken/61/6/61_JJID.2008.497/_article. [CrossRef] [PubMed]
- Kitamura, K.; Arita, M. Evaluation of VP4-VP2 Sequencing for Molecular Typing of Human Enteroviruses. PLoS ONE 2024, 19, e0311806. [Google Scholar] [CrossRef] [PubMed]
- National Institute of Infectious Diseases (NIID). Hand, Foot, and Mouth Disease Pathogen Detection Manual; NIID: Tokyo, Japan, 2023; Available online: https://id-info.jihs.go.jp/manuals/pathogen-detection/HFMdis20230704.pdf (accessed on 13 March 2026).
- Rotbart, H.A. Enzymatic RNA Amplification of the Enteroviruses. J. Clin. Microbiol. 1990, 28, 438–442. [Google Scholar] [CrossRef]
- Nix, W.A.; Oberste, M.S.; Pallansch, M.A. Sensitive, Seminested PCR Amplification of VP1 Sequences for Direct Identification of All Enterovirus Serotypes from Original Clinical Specimens. J. Clin. Microbiol. 2006, 44, 2698–2704. [Google Scholar] [CrossRef]
- Wolffs, P.F.; Bruggeman, C.A.; van Well, G.T.; van Loo, I.H. Replacing Traditional Diagnostics of Fecal Viral Pathogens by a Comprehensive Panel of Real-Time PCRs. J. Clin. Microbiol. 2011, 49, 1926–1931. [Google Scholar] [CrossRef]
- Bustin, S.A.; Ruijter, J.M.; van den Hoff, M.J.B.; Kubista, M.; Pfaffl, M.W.; Shipley, G.L.; et al. MIQE 2.0: Revision of the Minimum Information for Publication of Quantitative Real-Time PCR Experiments Guidelines. Clin. Chem. 2025, 71, 634–651. [Google Scholar] [CrossRef]
- Johnson, A.D. An Extended IUPAC Nomenclature Code for Polymorphic Nucleic Acids. Bioinformatics 2010, 26, 1386–1389. [Google Scholar] [CrossRef]
- Stadhouders, R.; Pas, S.D.; Anber, J.; Voermans, J.; Mes, T.H.; Schutten, M. The Effect of Primer-Template Mismatches on Nucleic Acid Detection Using the 5′ Nuclease Assay. J. Mol. Diagn. 2010, 12, 109–117. [Google Scholar] [CrossRef]
- Rejali, N.A.; Moric, E.; Wittwer, C.T. The Effect of Single Mismatches on Primer Extension. Clin. Chem. 2018, 64, 801–809. [Google Scholar] [CrossRef]
- Thoelen, I.; Moes, E.; Lemey, P.; Mostmans, S.; Wollants, E.; Lindberg, A.M.; et al. Serotype and Genotype Correlation of VP1 and the 5′ NCR among Enterovirus B Species. J. Clin. Microbiol. 2004, 42, 963–971. [Google Scholar] [CrossRef]
- Fujimoto, T.; Chikahira, M.; Yoshida, S.; et al. Outbreak of Central Nervous System Disease Associated with Hand, Foot, and Mouth Disease in Japan during the Summer of 2000. Microbiol Immunol. 2002, 46, 621–627. [Google Scholar] [CrossRef]
- De Schrijver, S.; Vanhulle, E.; Ingenbleek, A.; Alexakis, L.; Johannesen, C.K.; Broberg, E.K.; et al. Epidemiological and Clinical Insights into Enterovirus Circulation in Europe, 2018–2023: A Multicenter Retrospective Surveillance Study. J. Infect. Dis. 2025, 232, e104–e115. [Google Scholar] [CrossRef]
- Whitehouse, E.R.; Lopez, A.; English, R.; Getachew, H.; Ng, T.F.F.; Emery, B.; et al. Surveillance for Acute Flaccid Myelitis, United States, 2018–2022. MMWR Morb. Mortal. Wkly. Rep. 2024, 73, 70–76. [Google Scholar] [CrossRef]
- Chong, P.F.; Kira, R.; Mori, H.; Okumura, A.; Torisu, H.; Yasumoto, S.; et al. Clinical Features of Acute Flaccid Myelitis Temporally Associated with an Enterovirus D68 Outbreak in Japan. Clin. Infect. Dis. 2018, 66, 653–664. [Google Scholar] [CrossRef]
| Primer/Probe name | Polarity | Sequence (5′→3′) | Position (V01149 PV1, Mahoney) | Primer length (bp) | Reference |
| EVP2 | + | CCTCCGGCCCCTGAATGCGGCTAAT | 444-468 | 25 | [16] |
| EVP4 | + | CTACTTTGGGTGTCCGTGTT | 541-560 | 20 | [12] |
| OL68-1 | − | GGTAAYTTCCACCACCANCC | 1178-1197 | 20 | [11] |
| C3R | - | TCNGGRAAYTTCCAVYACCA | 1181-1200 | 20 | This study |
| Primer F | + | CCCTGAATGCGGCTAATCC | 452-470 | 19 | [18] |
| Primer R1 | - | ATTGTCACCATAAGCAGCCA | 577-596 | 20 | [18] |
| Probe P(FAM) | + | AACCGACTACTTTGGGTGTCCGTGTTTC | 535-562 | 28 | [18] |
| SO224 | + | GCIATGYTIGGIACICAYRT | 1966-1977 | 20 | [17] |
| AN89 | + | CCAGCACTGACAGCAGYNGARAYNGG | 2603-2628 | 26 | [17] |
| SO222 | - | CICCIGGIGGIAYRWACAT | 2951-2969 | 19 | [17] |
| AN88 | - | TACTGGACCACCTGGNGGNAYRWACAT | 2951-2969 | 27 | [17] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




