Submitted:
16 March 2026
Posted:
17 March 2026
You are already at the latest version
Abstract
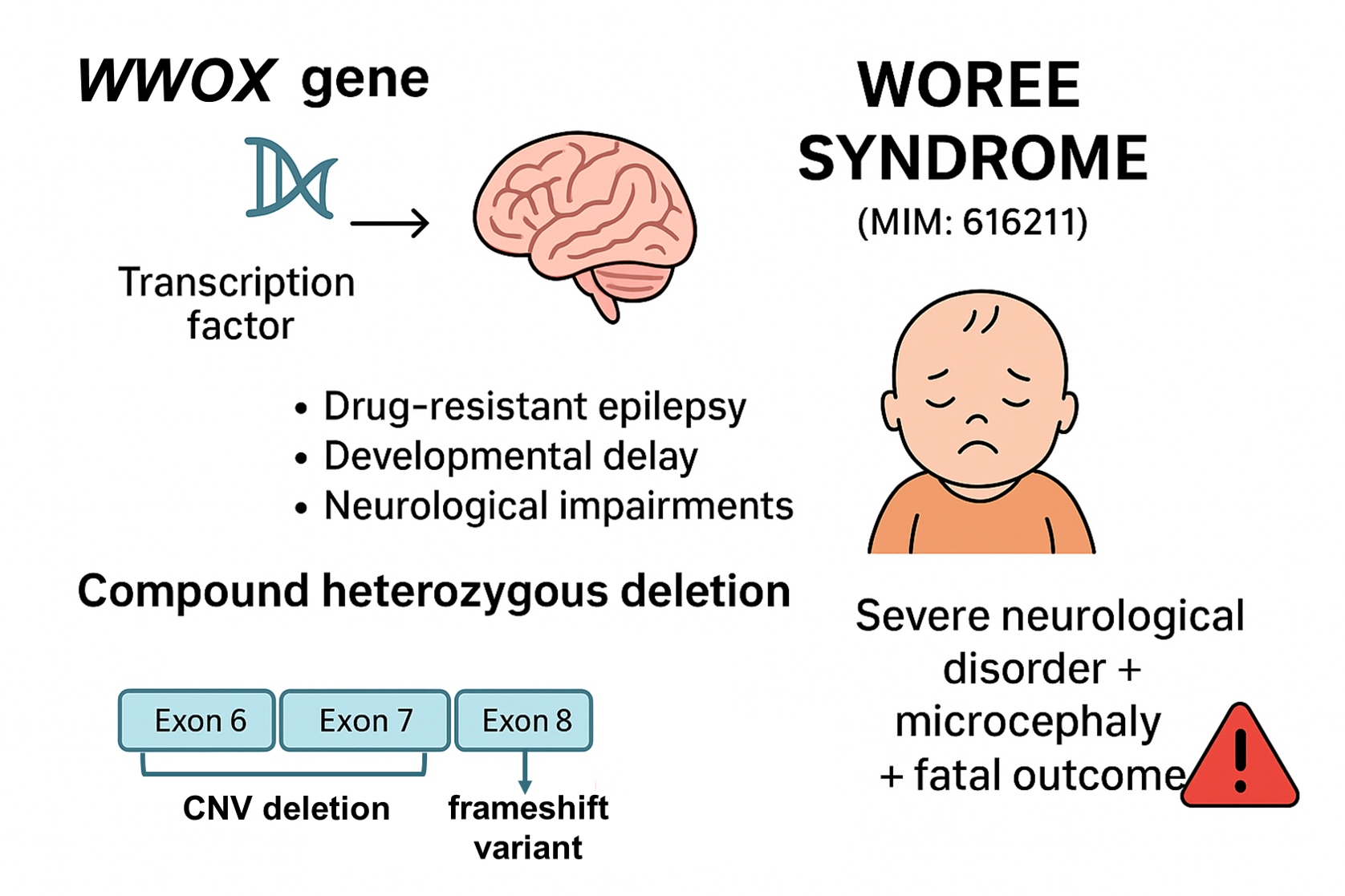
Keywords:
1. Introduction
2. Materials and Methods
2.1. Ethical Compliance
2.2. Genetic Testing and Genomic Data Analysis
3. Results
3.1. Case Presentation
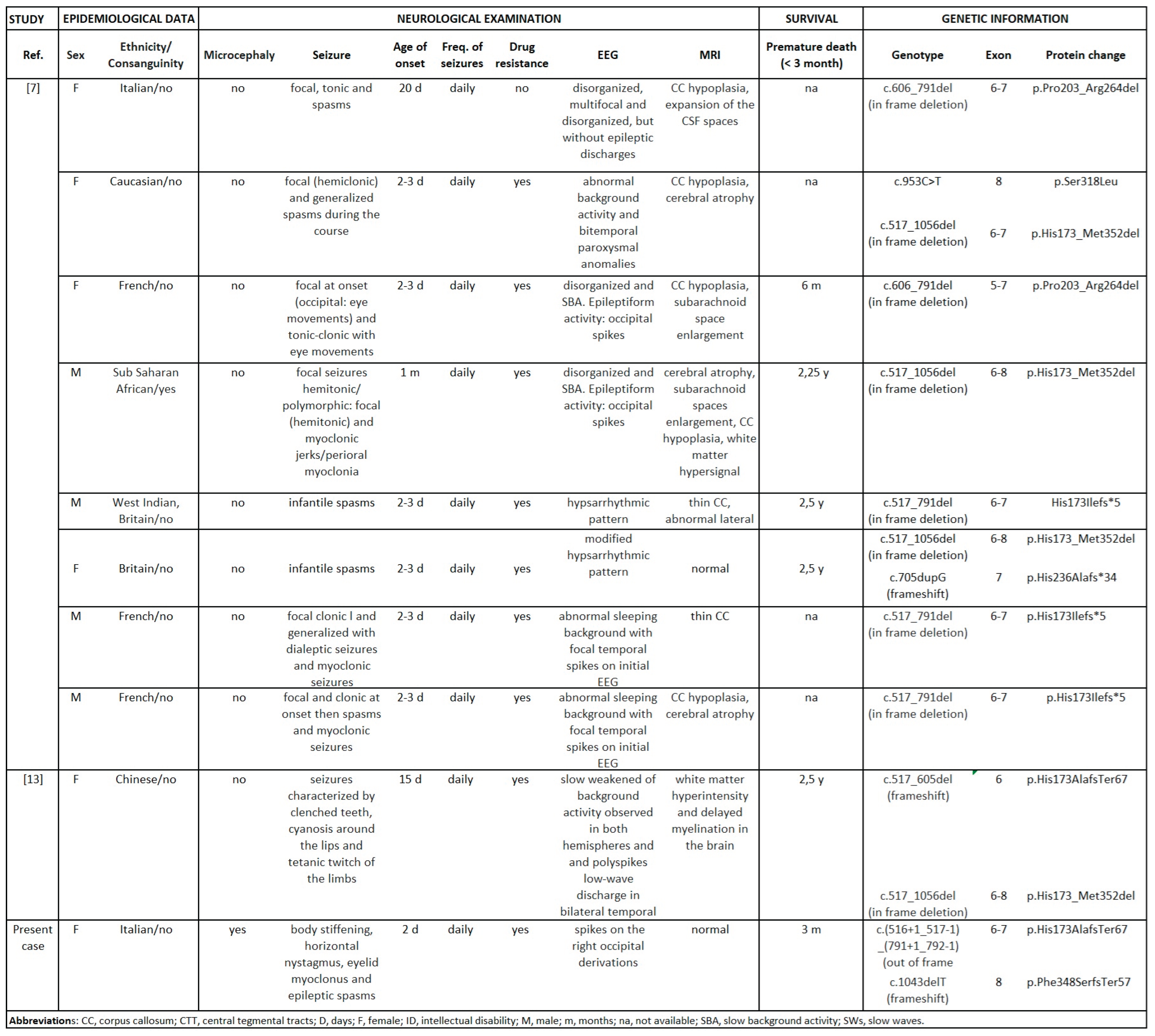
3.2. Mutational Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ADH/SDR | Short-chain Dehydrogenases/Reductases |
| ACMG | American College of Medical Genetics and Genomics |
| ASMs | Anti-Seizure Medications |
| bp | Base pairs |
| CNVs | Copy Number Variations |
| DD | Developmental Delay |
| DNA | Deoxyribonucleic Acid |
| EEG | Electroencephalogram |
| ExAC | Exome Aggregation Consortium |
| HGMD | Human Gene Mutation Database |
| IEE | Infantile Epileptic Encephalopathy |
| IEE28 | Infantile Epileptic Encephalopathy 28 |
| IED | Interictal Epileptiform Discharges |
| KO | Knockout |
| MIM | Mendelian Inheritance in Man |
| MRI | Magnetic Resonance Imaging |
| NICU | Neonatal Intensive Care Unit |
| PCR | Polymerase Chain Reaction |
| SCAR12 | Autosomal Recessive Spinocerebellar Ataxia |
| SDS | Standard Deviation Score |
| VEP | Variant Effect Predictor |
| WES | Whole Exome Sequencing |
| WOREE | WWOX-Related Epileptic Encephalopathy |
| WES | Whole Exome Sequencing |
References
- Hussain, T.; Liu, B.; Shrock, M.S.; Williams, T.; Aldaz, C.M. WWOX, the FRA16D gene: A target of and a contributor to genomic instability. Genes Chromosomes Cancer 2019, 58, 324–338. [Google Scholar] [CrossRef] [PubMed]
- Banne, E.; Abudiab, B.; Abu-Swai, S.; et al. Neurological Disorders Associated with WWOX Germline Mutations—A Comprehensive Overview. Cells 2021, 10, 824. [Google Scholar] [CrossRef]
- Aldaz, C.M.; Hussain, T. WWOX Loss of Function in Neurodevelopmental and Neurodegenerative Disorders. Int. J. Mol. Sci. 2020, 21, 8922. [Google Scholar] [CrossRef] [PubMed]
- Mallaret, M.; Synofzik, M.; Lee, J.; et al. The tumour suppressor gene WWOX is mutated in autosomal recessive cerebellar ataxia with epilepsy and mental retardation. Brain 2014, 137, 411–419. [Google Scholar] [CrossRef] [PubMed]
- Stenson, P.D.; Ball, E.V.; Mort, M.; et al. Human Gene Mutation Database (HGMD): 2003 update. Hum. Mutat. 2003, 21, 577–581. [Google Scholar] [CrossRef] [PubMed]
- Battaglia, L.; Scorrano, G.; Spiaggia, R.; et al. Neuroimaging features of WOREE syndrome: A mini-review of the literature. Front. Pediatr. 2023, 11, 1301166. [Google Scholar] [CrossRef] [PubMed]
- Piard, J.; Hawkes, L.; Milh, M.; et al. The phenotypic spectrum of WWOX-related disorders: 20 additional cases of WOREE syndrome and review of the literature. Genet. Med. 2019, 21, 1308–1318. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Salam, G.; Thoenes, M.; Afifi, H.H.; Körber, F.; Swan, D.; Bolz, H.J. The supposed tumor suppressor gene WWOX is mutated in an early lethal microcephaly syndrome with epilepsy, growth retardation and retinal degeneration. Orphanet J. Rare Dis. 2014, 9, 12. [Google Scholar] [CrossRef] [PubMed]
- Oliver, K.L.; Trivisano, M.; Mandelstam, S.A.; et al. WWOX developmental and epileptic encephalopathy: Understanding the epileptology and the mortality risk. Epilepsia 2023, 64, 1351–1367. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Aziz, N.; Bale, S.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.Y.; Chou, Y.T.; Lai, F.J.; et al. WWOX deficiency leads to neurodevelopmental and degenerative neuropathies and glycogen synthase kinase 3β-mediated epileptic seizure activity in mice. Acta Neuropathol. Commun. 2020, 8, 6. [Google Scholar] [CrossRef] [PubMed]
- Chong, S.C.; Cao, Y.; Fung, E.L.W.; et al. Expansion of the clinical and molecular spectrum of WWOX-related epileptic encephalopathy. Am. J. Med. Genet. A 2023, 191, 776–785. [Google Scholar] [CrossRef] [PubMed]
- Mignot, C.; Lambert, L.; Pasquier, L.; et al. WWOX-related encephalopathies: Delineation of the phenotypical spectrum and emerging genotype-phenotype correlation. J. Med. Genet. 2015, 52, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.S.; Wen, X.J.; Zhang, S.; Wang, D.G.; Xiong, Y.; Li, Z.M. Identification of compound heterozygous deletion of the WWOX gene in WOREE syndrome. BMC Med. Genom. 2023, 16, 291. [Google Scholar] [CrossRef] [PubMed]
- Vetri, L.; Calì, F.; Saccone, S.; et al. Whole Exome Sequencing as a First-Line Molecular Genetic Test in Developmental and Epileptic Encephalopathies. Int. J. Mol. Sci. 2024, 25, 1146. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



