Submitted:
16 March 2026
Posted:
17 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
- Healthy individuals who understood the study purpose and procedures and voluntarily expressed willingness to participate;
- Individuals with a BMI exceeding the normal range according to institutional health screening standards;
- Individuals for whom both ultrasound imaging of the liver parenchyma and MRI DIXON examination were feasible.
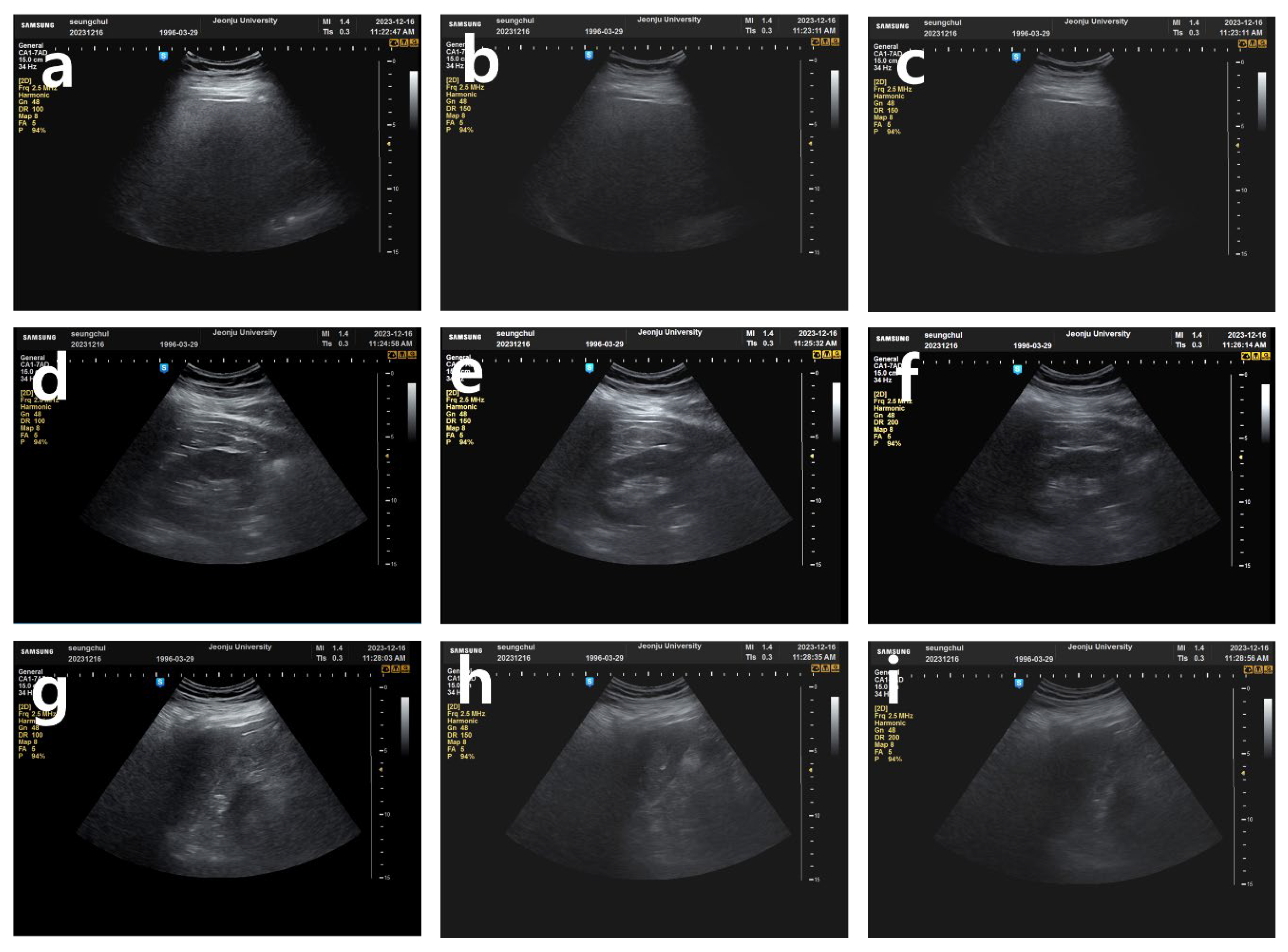
2.1. Ultrasound Imaging Acquisition Method
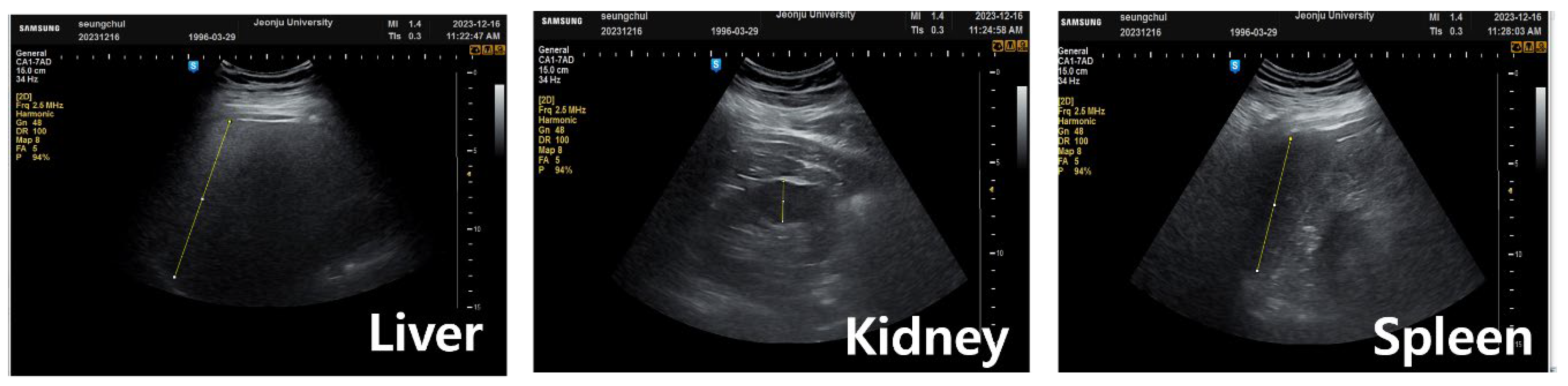
2.2. MRI Dixon-based FF Numerical Acquisition and Group Classification Criteria
2.3. Candidate Variable Design and Configuration
2.4. PCA, Statistical Analysis, and Key Variables
2.5. Machine Learning Prediction Model Development Procedure
3. Results
3.1. PCA and Derivation of Key Variables
3.2. Average Comparison of Ultrasound-Based Variables (Biomarkers)
3.2.1. Comparison of Mean Values for Key Variables Between MRI FF Groups
3.2.2. Summary of Correlation Analysis (Pearson's r) Results
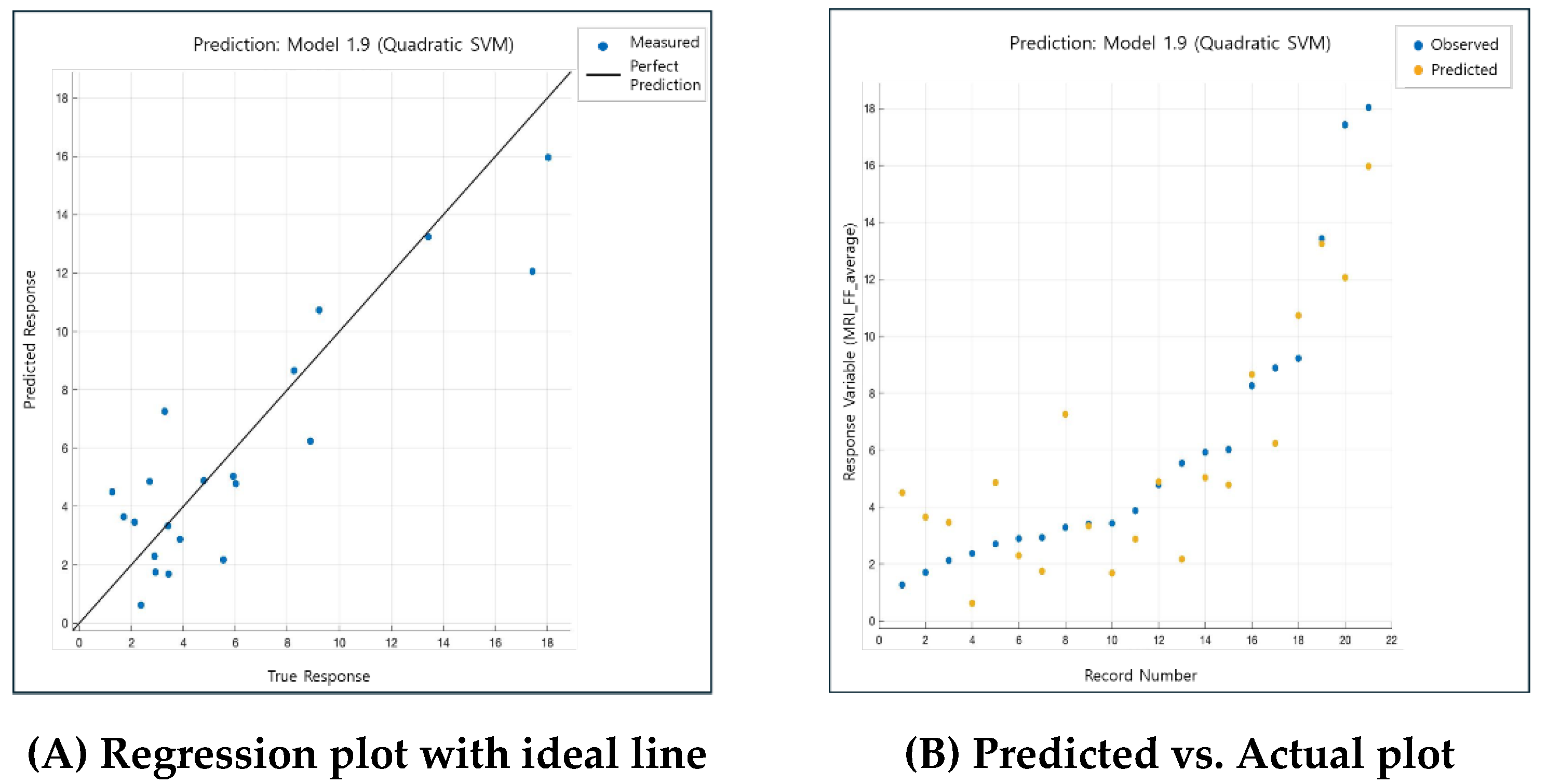
3.3. Predictive Model Performance Evaluation)
4. Discussion
4.1. Predictive Model Performance Evaluation)
4.2. Clinical Utility of Ultrasound-Based Variables
4.3. Limitations and Complementary Aspects of Machine Learning Predictive Models
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MRI | Magnetic Resonance Imaging |
| MRI-FF | MRI DIXON-based Fat Fraction |
| MRI-PDFF | MRI Proton Density Fat Fraction |
| FF | Fat Fraction |
| BMI | Body Mass Index |
| CT | Computed Tomography |
| DR | Dynamic Range |
| TGC | Time Gain Compensation |
| ROI | Region of Interest |
| PCA | Principal Component Analysis |
| PLS-DA | Partial Least Squares Discriminant Analysis |
| OPLS-DA | Orthogonal Partial Least Squares Discriminant Analysis |
| VIP | Variable Importance in Projection |
| SVM | Support Vector Machine |
| RMSE | Root Mean Square Error |
| R² | Coefficient of Determination |
| SD | Standard Deviation |
| TR | Repetition Time |
| TE | Echo Time |
| FOV | Field of View |
| IP | In-phase |
| OP | Out-of-phase |
| NASH | Non-Alcoholic Steatohepatitis |
| AI | Artificial Intelligence |
| IRB | Institutional Review Board |
| USA | United States of America |
References
- Chalasani, N.; Younossi, Z.; Lavine, J.E.; Diehl, A.M.; Brunt, E.M.; Cusi, K.; Charlton, M.; Sanyal, A.J. The diagnosis and management of non-alcoholic fatty liver disease: practice guideline by the American Association for the Study of Liver Diseases, American College of Gastroenterology, and the American Gastroenterological Association. Hepatology 2012, 55, 2005–2023. [Google Scholar] [CrossRef]
- Kitade, H.; Chen, G.; Ni, Y.; Ota, T. Nonalcoholic fatty liver disease and insulin resistance: new insights and potential new treatments. Nutrients 2017, 9, 387. [Google Scholar] [CrossRef]
- Fon Tacer, K.; Rozman, D. Nonalcoholic Fatty liver disease: focus on lipoprotein and lipid deregulation. Journal of lipids 2011, 2011, 783976. [Google Scholar] [CrossRef] [PubMed]
- Reeder, S.B.; Cruite, I.; Hamilton, G.; Sirlin, C.B. Quantitative assessment of liver fat with magnetic resonance imaging and spectroscopy. Journal of magnetic resonance imaging 2011, 34, 729–749. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.S.; Park, S.H. Radiologic evaluation of nonalcoholic fatty liver disease. World journal of gastroenterology: WJG 2014, 20, 7392. [Google Scholar] [CrossRef] [PubMed]
- Hernaez, R.; Lazo, M.; Bonekamp, S.; Kamel, I.; Brancati, F.L.; Guallar, E.; Clark, J.M. Diagnostic accuracy and reliability of ultrasonography for the detection of fatty liver: a meta-analysis. Hepatology 2011, 54, 1082–1090. [Google Scholar] [CrossRef]
- Strauss, S.; Gavish, E.; Gottlieb, P.; Katsnelson, L. Interobserver and intraobserver variability in the sonographic assessment of fatty liver. Am J Roentgenol 2007, 189, W320–W323. [Google Scholar] [CrossRef]
- Strauss, S.; Gavish, E.; Gottlieb, P.; Katsnelson, L. Interobserver and intraobserver variability in the sonographic assessment of fatty liver. Am J Roentgenol 2007, 189, W320–W323. [Google Scholar] [CrossRef]
- Yajima, Y.; Ohta, K.; NARUI, T.; ABE, R.; SUZUKI, H.; OHTSUKI, M. Ultrasonographical diagnosis of fatty liver: significance of the liver-kidney contrast. Tohoku J Exp Med 1983, 139, 43–50. [Google Scholar] [CrossRef]
- Taylor, K.J.; Riely, C.A.; Hammers, L.; Flax, S.; Weltin, G.; Garcia-Tsao, G.; Conn, H.O.; Kuc, R.; Barwick, K.W. Quantitative US attenuation in normal liver and in patients with diffuse liver disease: importance of fat. Radiology 1986, 160, 65–71. [Google Scholar] [CrossRef]
- Zhang, Y.N.; Fowler, K.J.; Hamilton, G.; Cui, J.Y.; Sy, E.Z.; Balanay, M.; Hooker, J.C.; Szeverenyi, N.; Sirlin, C.B. Liver fat imaging—a clinical overview of ultrasound, CT, and MR imaging. Br J Radiol 2018, 91, 20170959. [Google Scholar] [CrossRef]
- Andre, M.P.; Han, A.; Heba, E.; Hooker, J.; Loomba, R.; Sirlin, C.B.; Erdman, J.W.; O'Brien, W.D. Accurate diagnosis of nonalcoholic fatty liver disease in human participants via quantitative ultrasound, 2014 IEEE International Ultrasonics Symposium, IEEE: 2014; , pp. 2375–2377.
- Taylor, K.J.; Riely, C.A.; Hammers, L.; Flax, S.; Weltin, G.; Garcia-Tsao, G.; Conn, H.O.; Kuc, R.; Barwick, K.W. Quantitative US attenuation in normal liver and in patients with diffuse liver disease: importance of fat. Radiology 1986, 160, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Valckx, F.; Thijssen, J.M.; Van Geemen, A.J.; Rotteveel, J.J.; Mullaart, R. Calibrated parametric medical ultrasound imaging. Ultrason Imaging 2000, 22, 57–72. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.; Luo, F.; Liu, D. Automatic time gain compensation in ultrasound imaging system, 2009 3rd International Conference on Bioinformatics and Biomedical Engineering, IEEE: 2009; , pp. 1–4.
- Ivosev, G.; Burton, L.; Bonner, R. Dimensionality reduction and visualization in principal component analysis. Anal Chem 2008, 80, 4933–4944. [Google Scholar] [CrossRef] [PubMed]
- Abdi, H.; Williams, L.J. Principal component analysis. Wiley interdisciplinary reviews: computational statistics 2010, 2, 433–459. [Google Scholar] [CrossRef]
- Kurita, T. Principal component analysis (PCA). In Computer vision: a reference guide; Springer, 2021; pp. 1013–1016. [Google Scholar]
- Zhang, Z.; Castelló, A. Principal components analysis in clinical studies. Annals of translational medicine 2017, 5, 351. [Google Scholar] [CrossRef]
- Raptis, D.A.; Fischer, M.A.; Graf, R.; Nanz, D.; Weber, A.; Moritz, W.; Tian, Y.; Oberkofler, C.E.; Clavien, P. MRI: the new reference standard in quantifying hepatic steatosis? Gut 2012, 61, 117–127. [Google Scholar] [CrossRef]
- Gupta, A.; Dixit, R.; Prakash, A. Non-invasive hepatic fat quantification: Can multi-echo Dixon help? Radiologia Brasileira 2024, 57, e20230125. [Google Scholar] [CrossRef]
- Noh, H.; Song, X.; Heo, S.H.; Kim, J.W.; Shin, S.S.; Ahn, K.Y.; Jeong, Y.Y.; Kang, H. Comparative study of ultrasonography, computed tomography, magnetic resonance imaging, and magnetic resonance spectroscopy for the diagnosis of fatty liver in a rat model. Journal of the Korean Society of Radiology 2017, 76, 14–24. [Google Scholar] [CrossRef]
- Tang, A.N.; Tan, J.; Sun, M.; Hamilton, G.; Bydder, M.; Wolfson, T.; Gamst, A.C.; Middleton, M.; Brunt, E.M.; Loomba, R. Nonalcoholic fatty liver disease: MR imaging of liver proton density fat fraction to assess hepatic steatosis. Radiology 2013, 267, 422–431. [Google Scholar] [CrossRef]
- Sirlin, C.B. Noninvasive imaging biomarkers for steatosis assessment. Liver Transplantation 2009, 15, 1389–1391. [Google Scholar] [CrossRef]
- Lee, D.H.; Lee, J.Y.; Park, M.S.; Han, J.K. Non-invasive monitoring of hepatic steatosis via acoustic structure quantification of ultrasonography with MR spectroscopy as the reference standard. Ultrasonography 2020, 39, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Tang, A.N.; Tan, J.; Sun, M.; Hamilton, G.; Bydder, M.; Wolfson, T.; Gamst, A.C.; Middleton, M.; Brunt, E.M.; Loomba, R. Nonalcoholic fatty liver disease: MR imaging of liver proton density fat fraction to assess hepatic steatosis. Radiology 2013, 267, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Boglárka, Z.; Zsombor, Z.; Rónaszéki, A.D.; Egresi, A.; Stollmayer, R.; Himsel, M.; Bérczi, V.; Kalina, I.; Werling, K.; Győri, G. Construction of a Compound Model to Enhance the Accuracy of Hepatic Fat Fraction Estimation with Quantitative Ultrasound. Diagnostics 2025, 15, 203. [Google Scholar] [CrossRef] [PubMed]
- Ferraioli, G.; Wong, V.W.; Castera, L.; Berzigotti, A.; Sporea, I.; Dietrich, C.F.; Choi, B.I.; Wilson, S.R.; Kudo, M.; Barr, R.G. Liver ultrasound elastography: an update to the world federation for ultrasound in medicine and biology guidelines and recommendations. Ultrasound Med Biol 2018, 44, 2419–2440. [Google Scholar] [CrossRef]
- Castera, L.; Friedrich-Rust, M.; Loomba, R. Noninvasive assessment of liver disease in patients with nonalcoholic fatty liver disease. Gastroenterology 2019, 156, 1264–1281. e4. [Google Scholar] [CrossRef]
- Ferraioli, G.; Monteiro, L.B.S. Ultrasound-based techniques for the diagnosis of liver steatosis. World journal of gastroenterology 2019, 25, 6053. [Google Scholar] [CrossRef]
- Dag, N.; Igci, G.; Yagin, F.H.; Hanci, M.S.; Kutlu, R. Interobserver Reproducibility of Ultrasound Attenuation Imaging Technology in Liver Fat Quantification. Journal of Clinical Ultrasound 2025, 53, 405–412. [Google Scholar] [CrossRef]
- Yin, H.; Fan, Y.; Yu, J.; Xiong, B.; Zhou, B.; Sun, Y.; Wang, L.; Zhu, Y.; Xu, H. Quantitative US fat fraction for noninvasive assessment of hepatic steatosis in suspected metabolic-associated fatty liver disease. Insights Into Imaging 2024, 15, 159. [Google Scholar] [CrossRef]
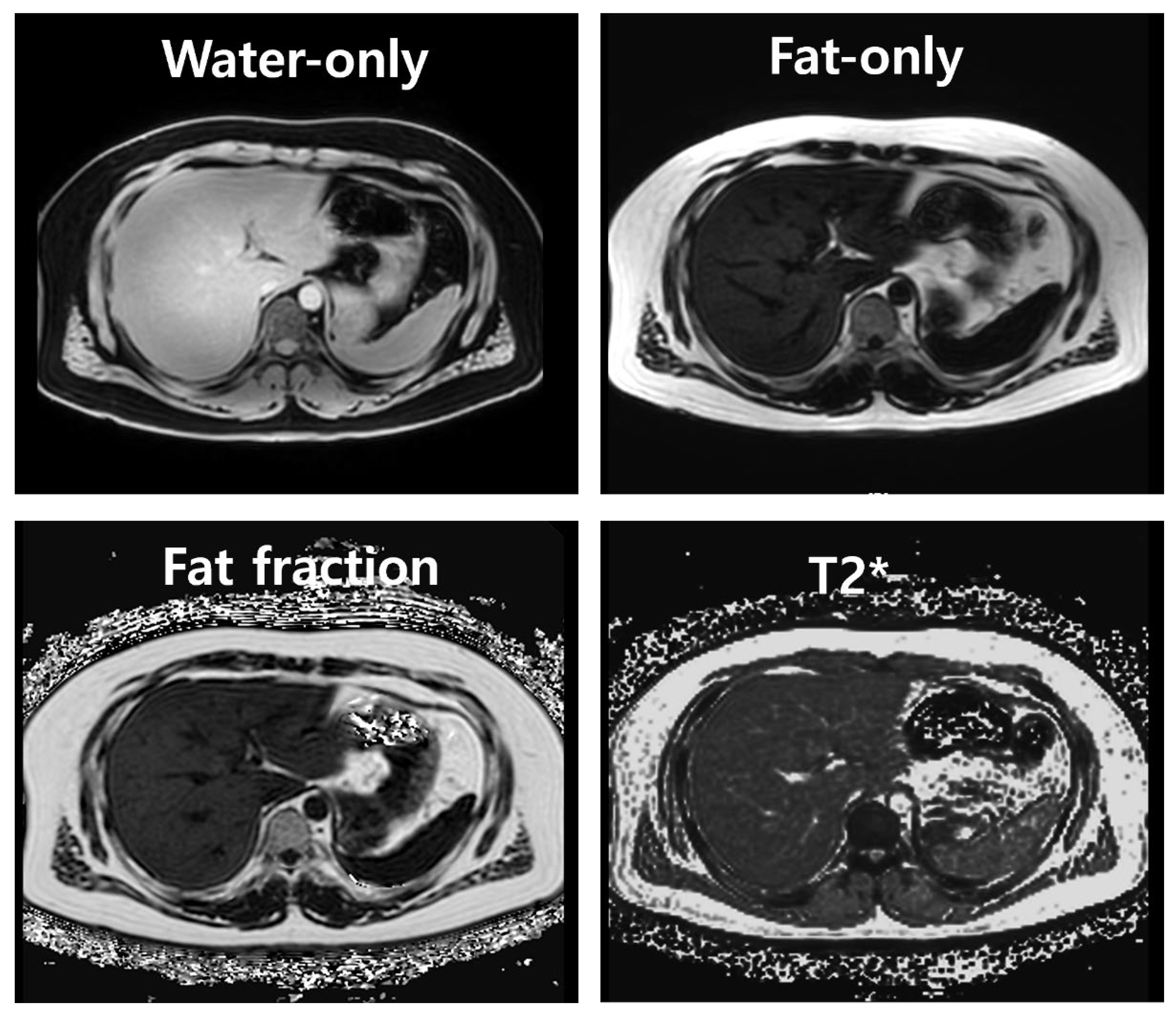
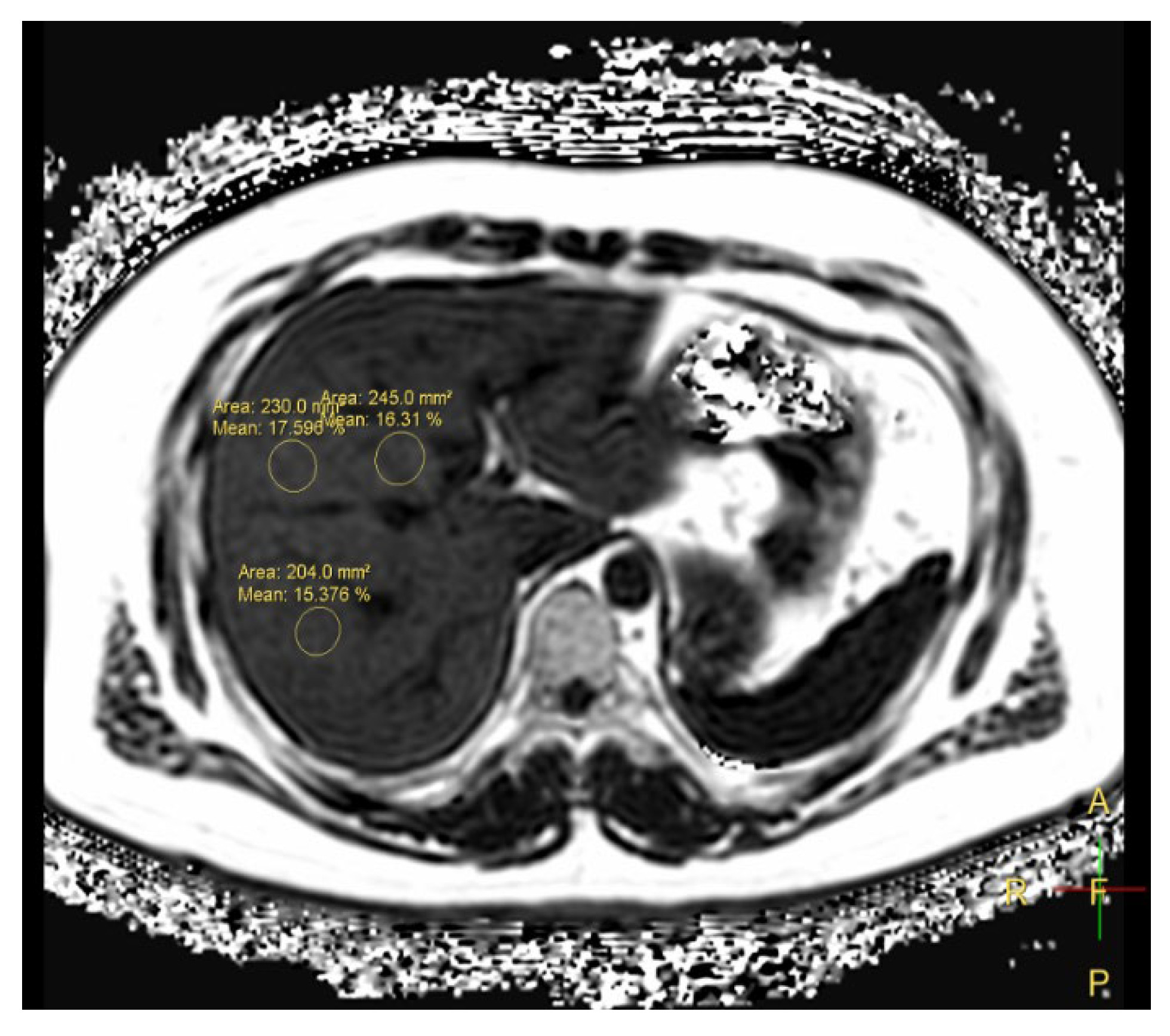
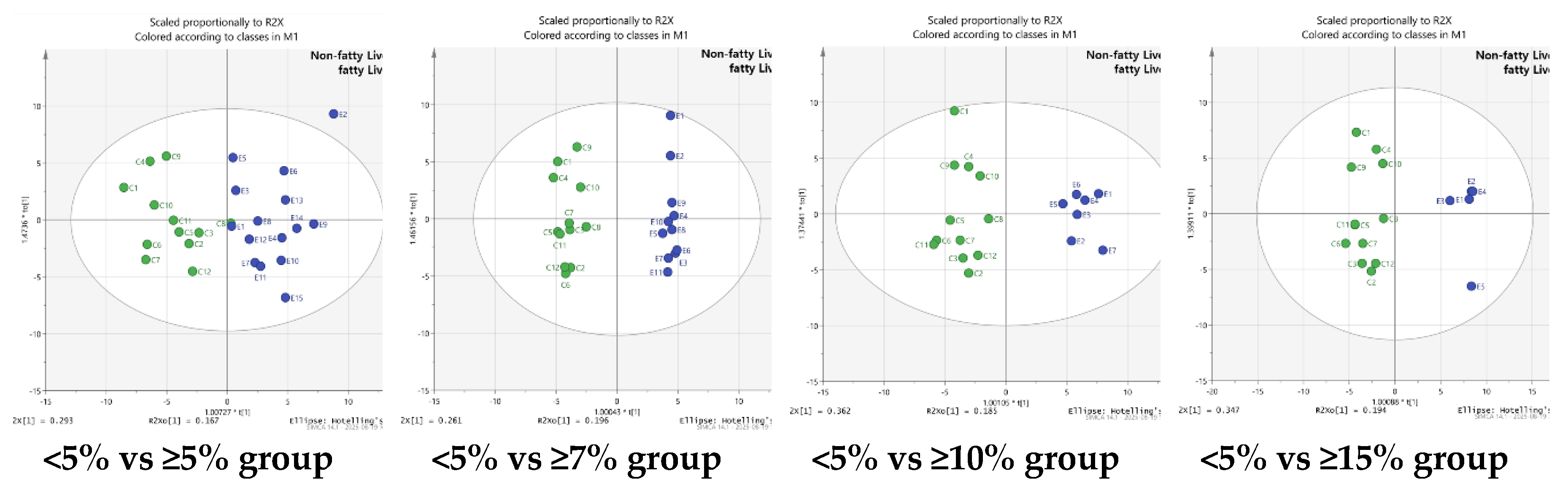
 : Non-fatty Liver,
: Non-fatty Liver,  : fatty Liver.
: fatty Liver.
 : Non-fatty Liver,
: Non-fatty Liver,  : fatty Liver.
: fatty Liver.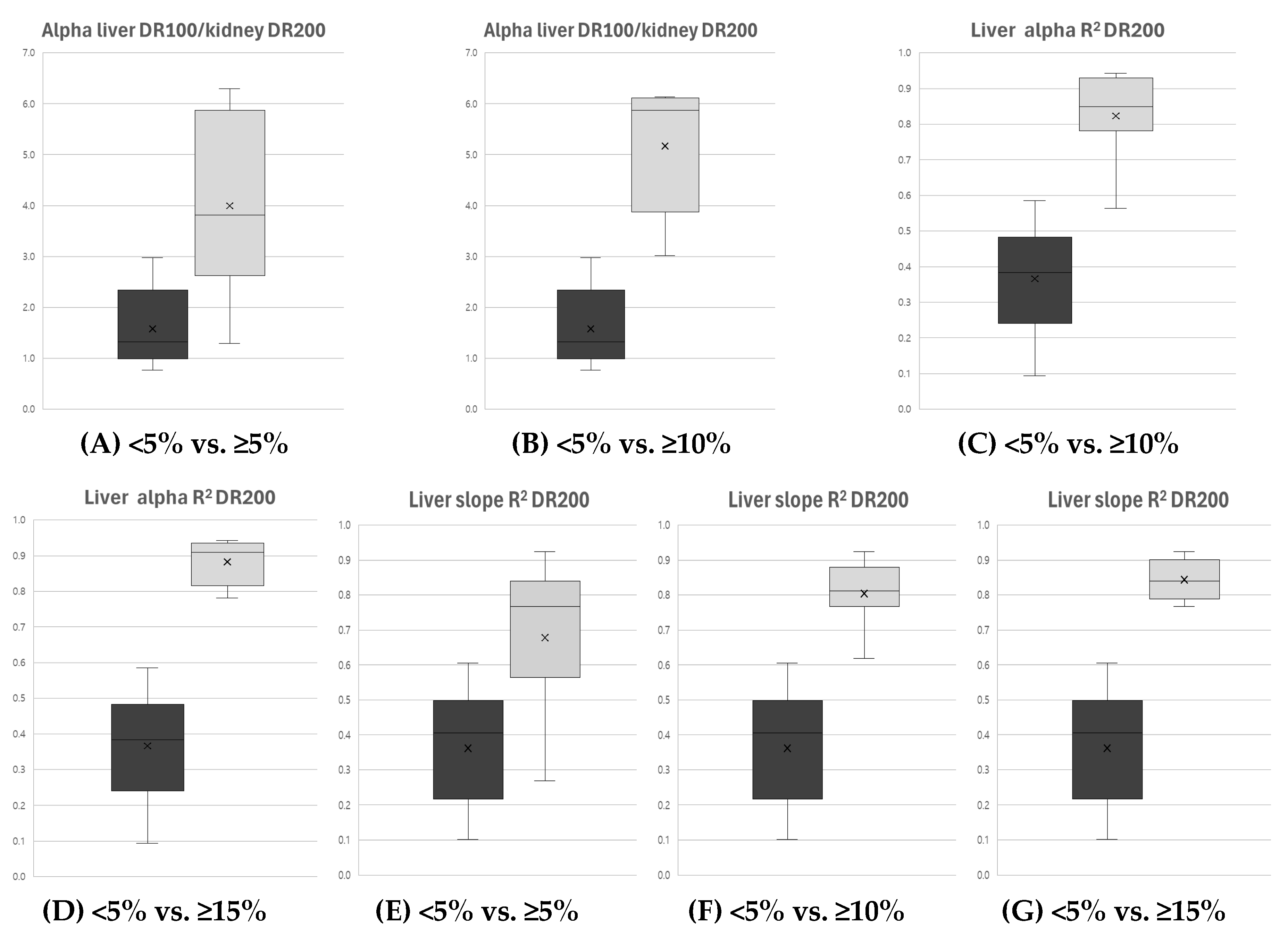
| MRI FF | VIP SCORE 1.0≥ | VIP SCORE 1.25≥ |
|---|---|---|
| <5% vs ≥5% | Alpha kidney200/liver100 Alpha kidney150/liver100 Alpha liver100/kidney150 Alpha liver100/kidney200 Liver alpha R2 DR100 Liver slope R2 DR200 Liver slope R2 DR100 Liver slope R2 DR150 Liver alpha R2 DR200 Alpha liver200/kidney150 Liver alpha DR100 Liver alpha R2 DR150 |
Alpha kidney200/liver100 Alpha kidney150/liver100 Alpha liver100/kidney150 Alpha liver100/kidney200 Liver alpha R2 DR100 Liver slope R2 DR200 Liver slope R2 DR100 Liver slope R2 DR150 |
| <5% vs ≥7% | Alpha kidney200/liver100 Alpha kidney150/liver100 Liver alpha R2 DR150 Liver slope R2 DR150 Alpha liver200/kidney150 Liver alpha R2 DR100 Alpha liver200/kidney100 Liver slope DR150 Liver slope R2 DR200 Alpha kidney200/liver150 Mean_ alpha _Liver Alpha liver150/kidney200 |
Alpha kidney200/liver100 Alpha kidney150/liver100 Liver alpha R2 DR150 Liver slope R2 DR150 |
| <5% vs ≥10% | Liver slope R2 DR200 Liver alpha R2 DR200 Alpha liver200/kidney100 Alpha liver200/kidney150 Alpha liver150/kidney100 Liver slope R2 DR150 Mean_ alpha _Liver Alpha kidney150/liver100 Liver alpha R2 DR150 Liver alpha DR100 Alpha kidney200/liver100 Alpha liver150/kidney200 |
Liver slope R2 DR200 Liver alpha R2 DR200 Alpha liver200/kidney100 Alpha liver200/kidney150 Alpha liver150/kidney100 Liver slope R2 DR150 |
| ⫶ | ||
| <5% vs ≥15% | Liver alpha R2 DR200 Alpha liver100/kidney200 Alpha liver100/kidney150 Liver slope R2 DR200 Liver alpha R2 DR150 Liver slope R2 DR150 Alpha liver200/kidney100 Alpha liver200/kidney150 Liver alpha R2 DR100 Alpha liver150/kidney200 Mean_ alpha _Liver Alpha kidney200/liver100 |
Liver alpha R2 DR200 Alpha liver100/kidney200 Alpha liver100/kidney150 Liver slope R2 DR200 Liver alpha R2 DR150 Liver slope R2 DR150 |
| Biomarkers | Correlation coefficient value | p-value (two-tailed) |
|---|---|---|
| Alpha liver 100/kidney 200 Liver alpha R2 DR200 Liver slope R2 DR200 Alpha liver 100/kidney 150 Liver slope R2 DR150 Liver alpha R2 DR150 Liver alpha DR100 Alpha kidney200/liver100 Alpha kidney150/liver100 Liver alpha R2 DR100 Liver alpha DR150 |
.814** .753** .724** .724** .687** .681** .680** -.651** -.644** .637** .633** |
<.001 <.001 <.001 <.001 <.001 <.001 <.001 <.001 <.001 <.001 <.001 |
| Observed | Predicted | |
|---|---|---|
| T1 T2 T3 T4 T5 T6 |
16.42666667 12.155 6.996666667 7.781333333 26.39166667 27.37166667 |
18.7186 16.5497 11.1050 9.3975 24.5986 34.1854 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




