Submitted:
11 March 2026
Posted:
12 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Ethics Approval
2.2. Animals and Experimental Design
2.3. Sample Collection
2.4. Bacterial 16S rRNA Gene Sequencing and Analysis
2.5. LC-MS Non-Targeted Metabolomics Detection and Analysis
2.6. Statistical Analysis
3. Results
3.1. Analysis of 16S rRNA Gene Sequencing Results
3.2. Alpha Diversity Analysis
3.3. Beta Diversity Analysis
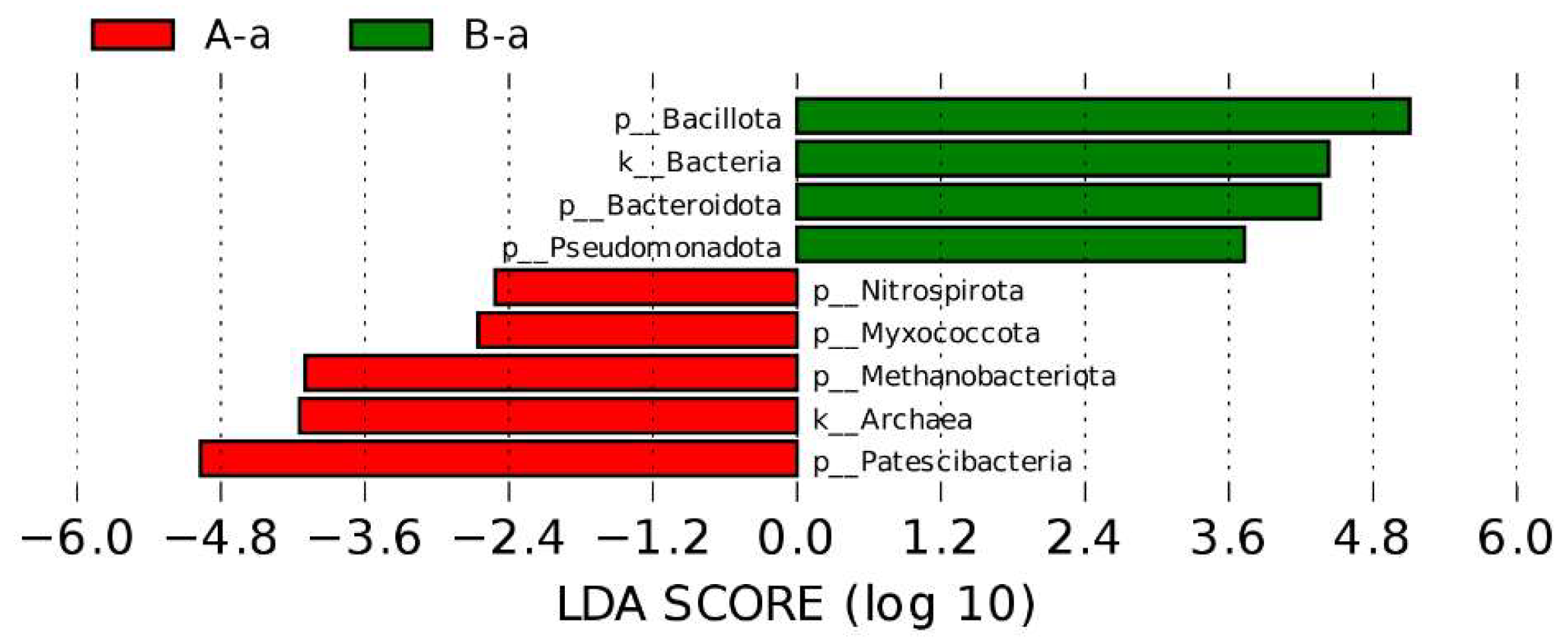
3.4. Differential Microbial Composition Analysis
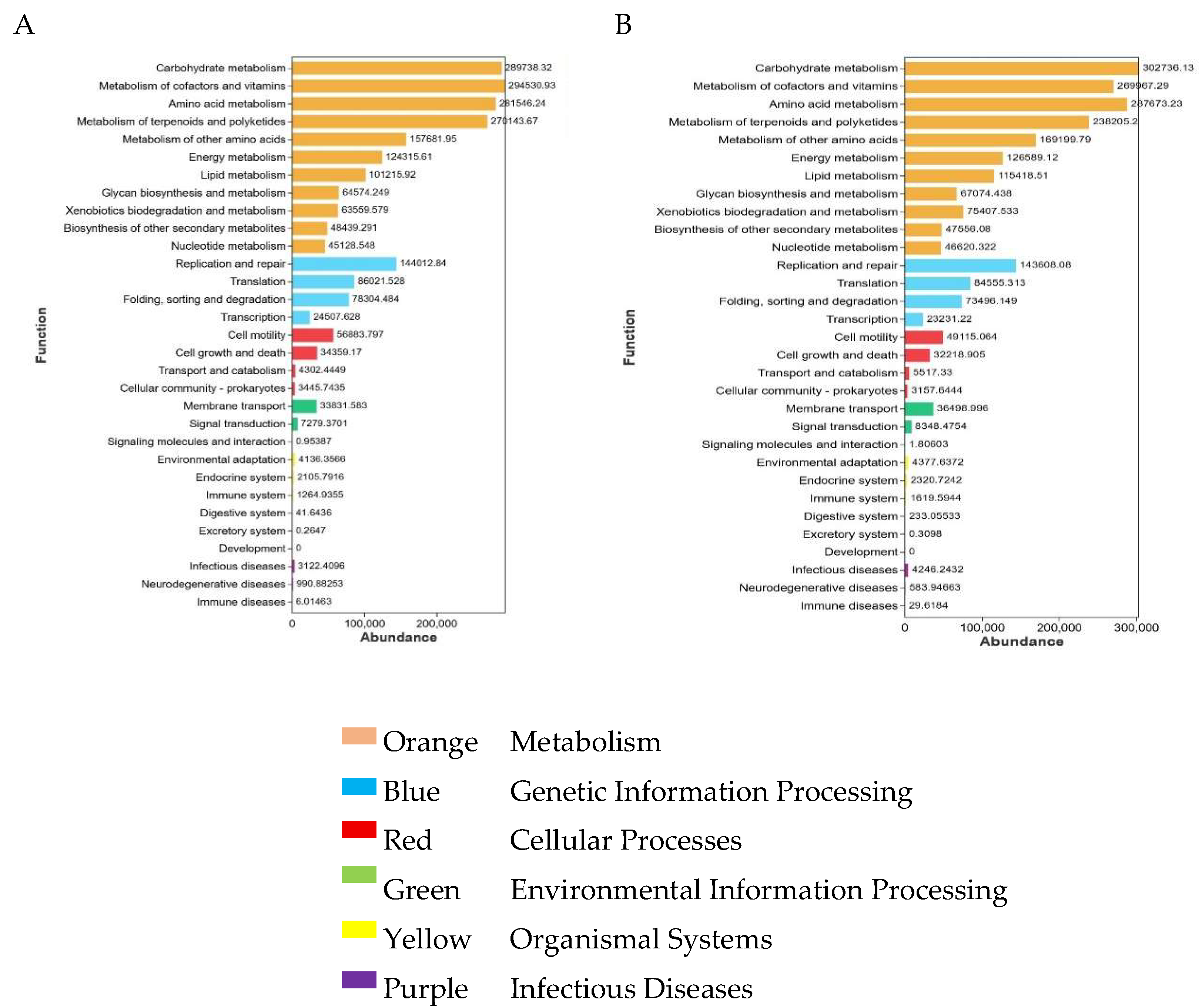
3.5. Functional Prediction of Microbial Communities
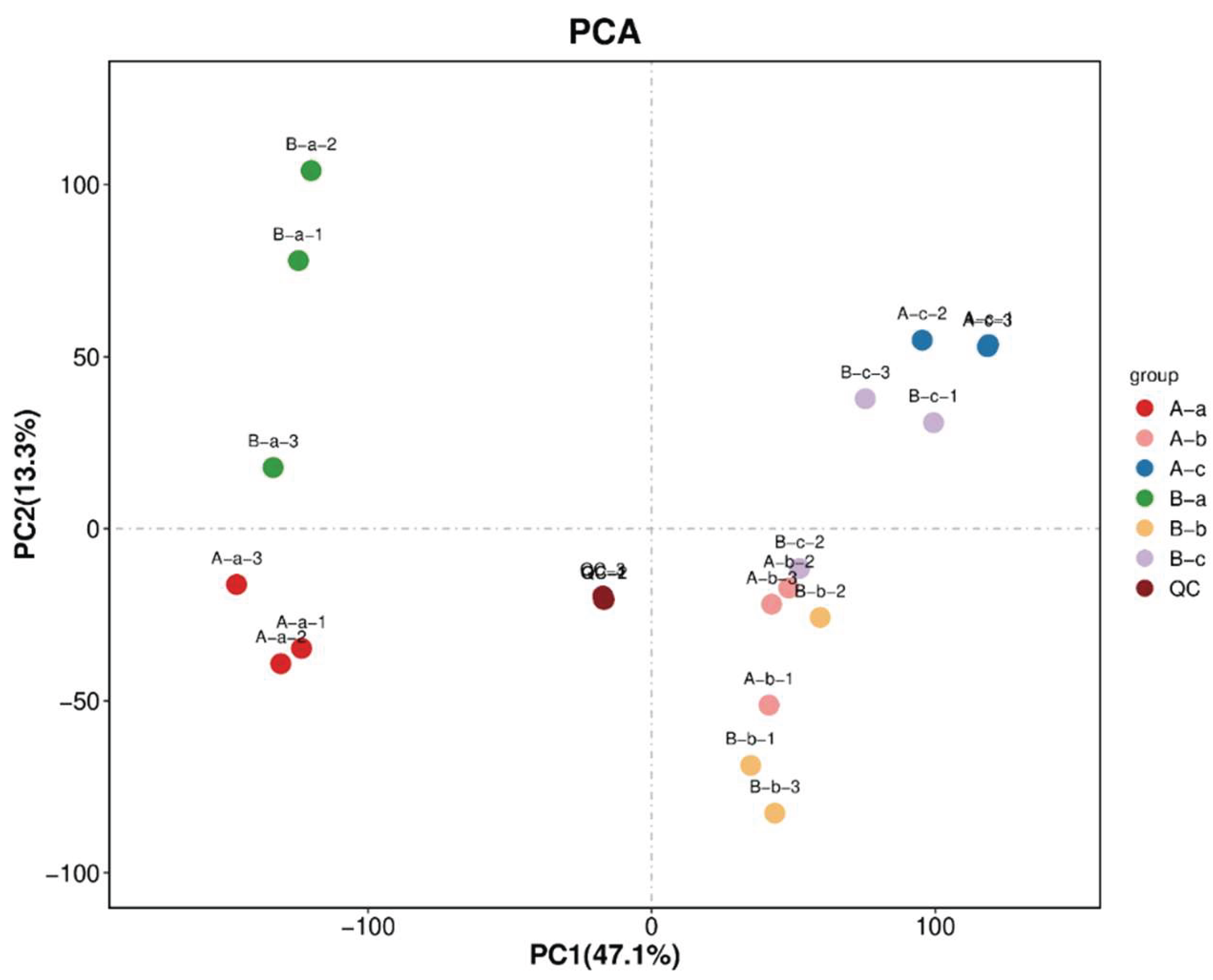
3.6. Analysis of Non-Targeted Metabolomics Results
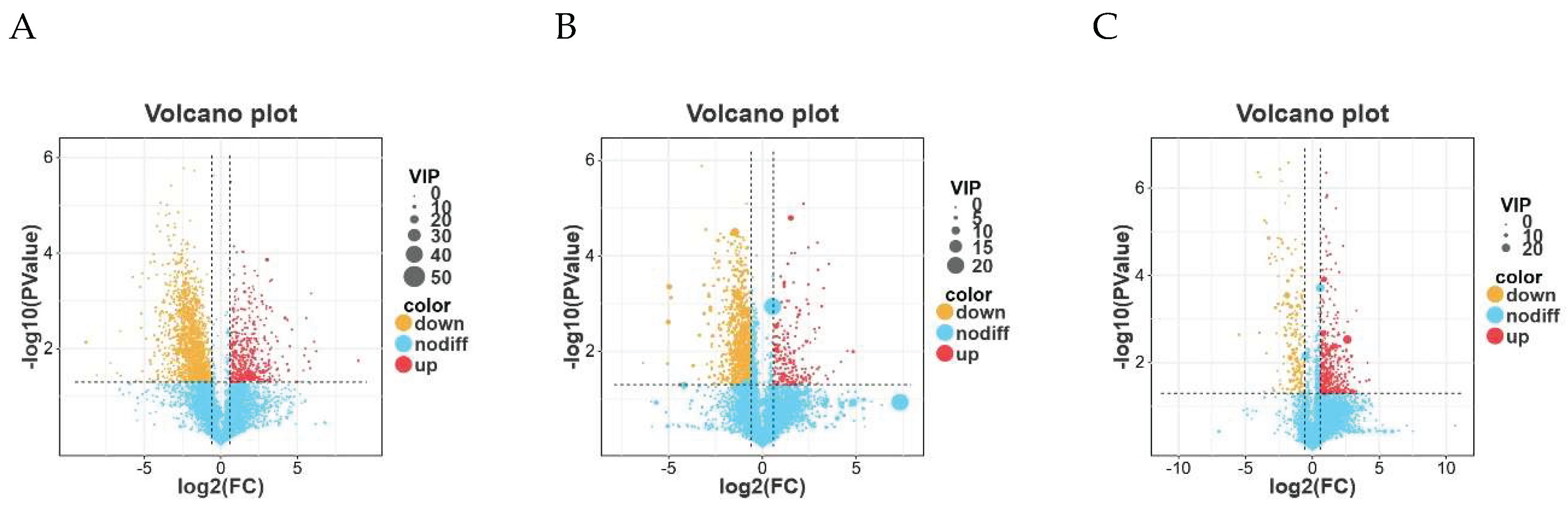
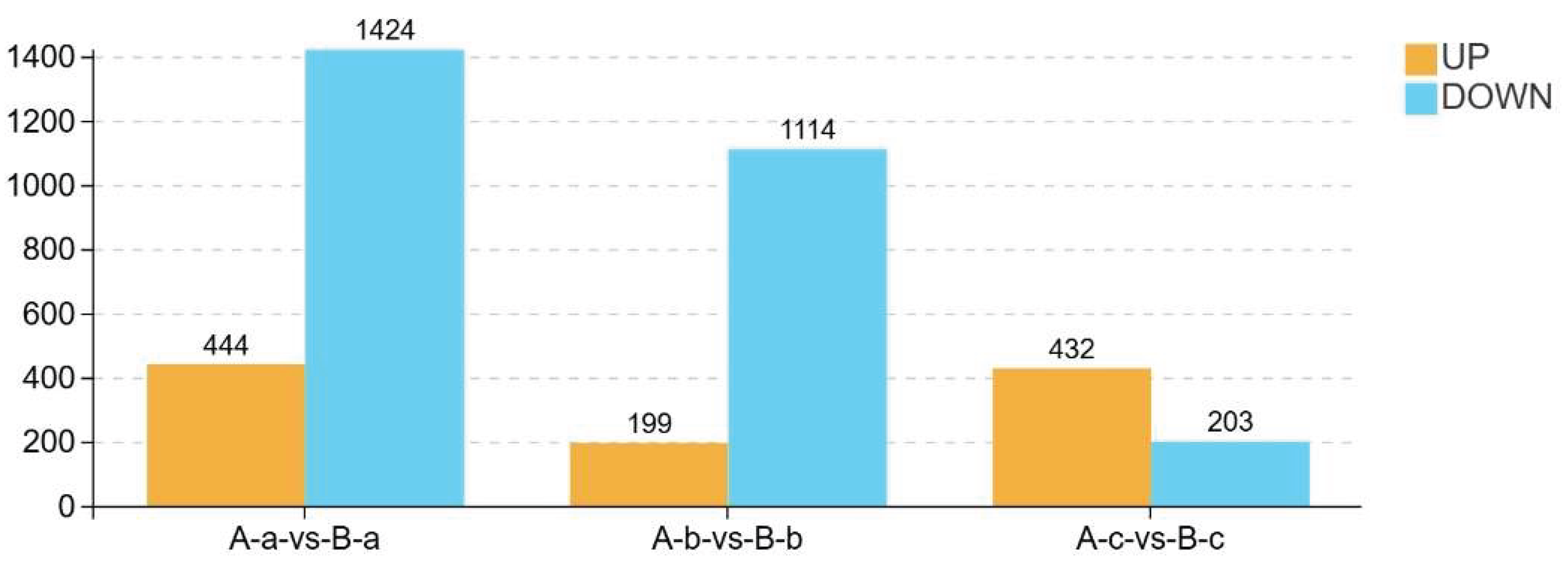
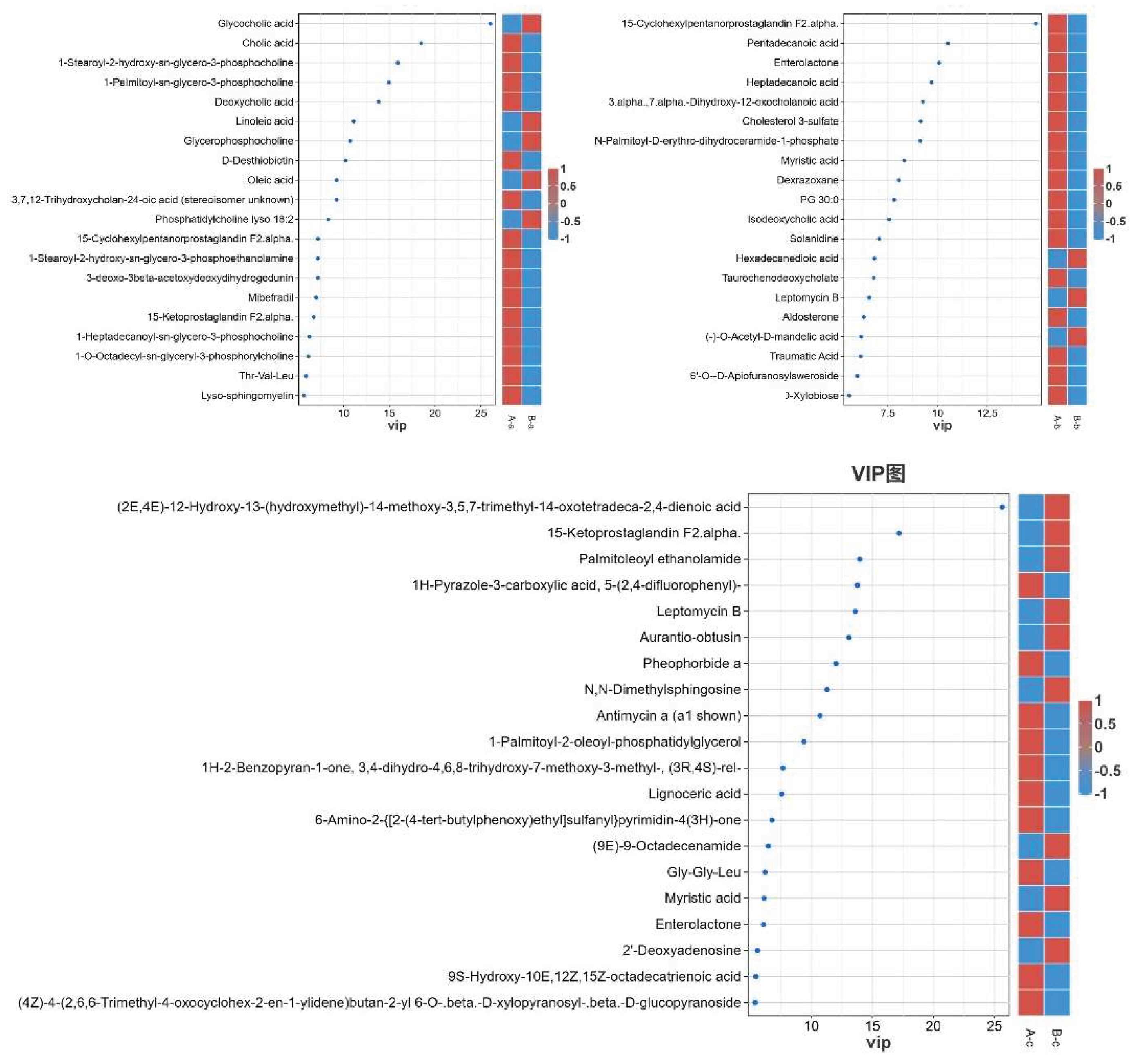
3.7. Differential Metabolite Profiling Between Groups
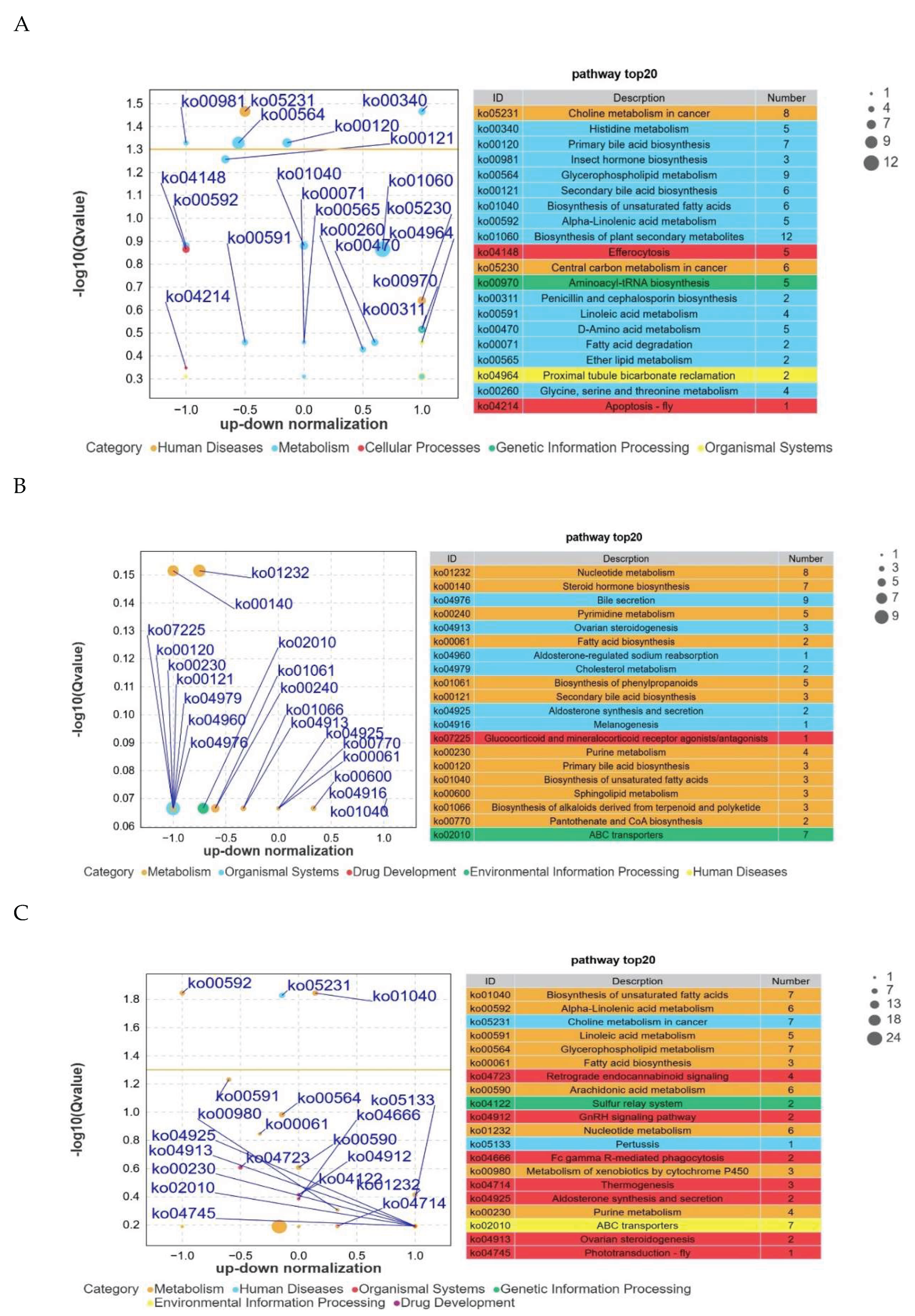
3.8. KEGG Pathway Enrichment Analysis of Differential Metabolites Between Groups
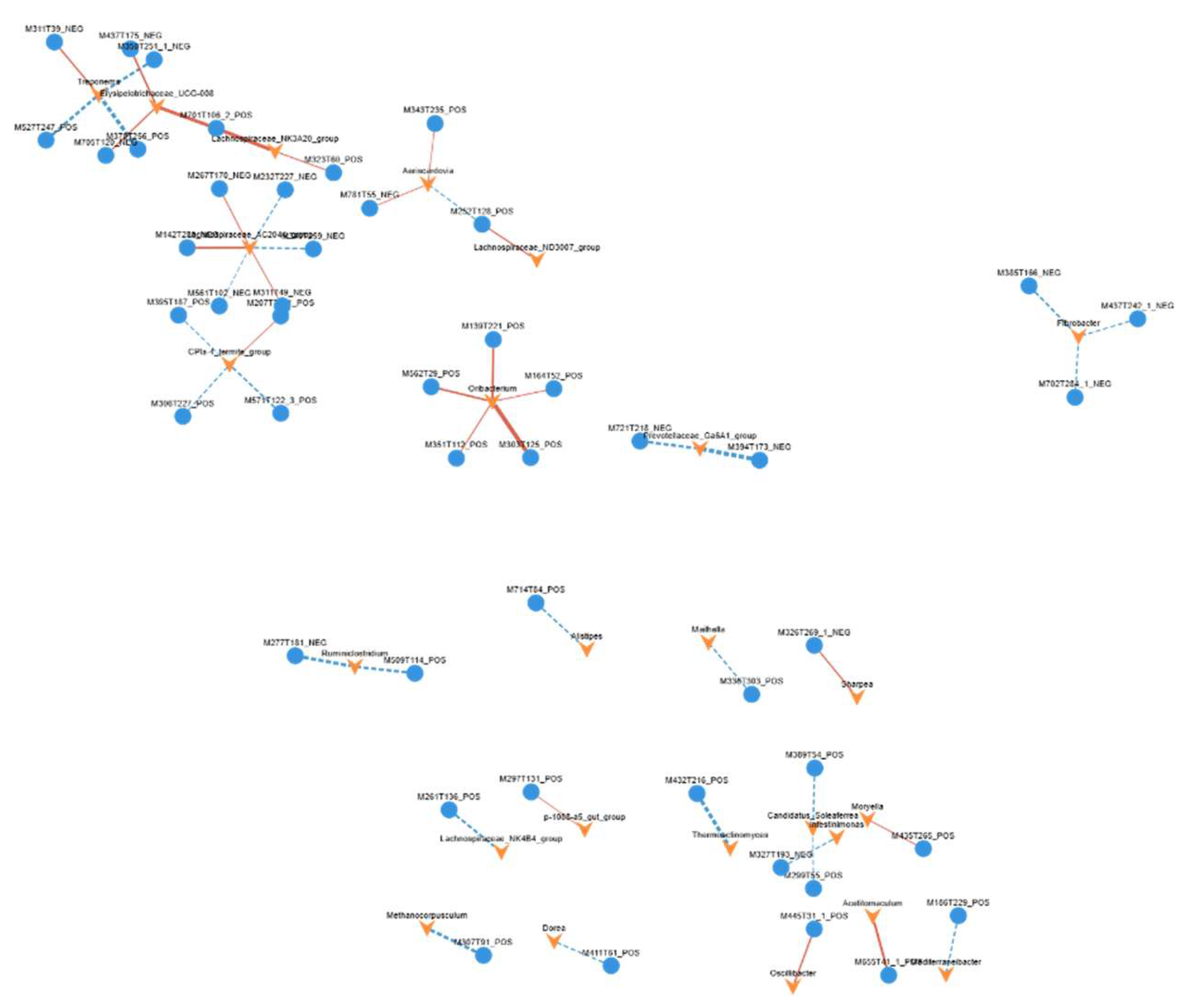
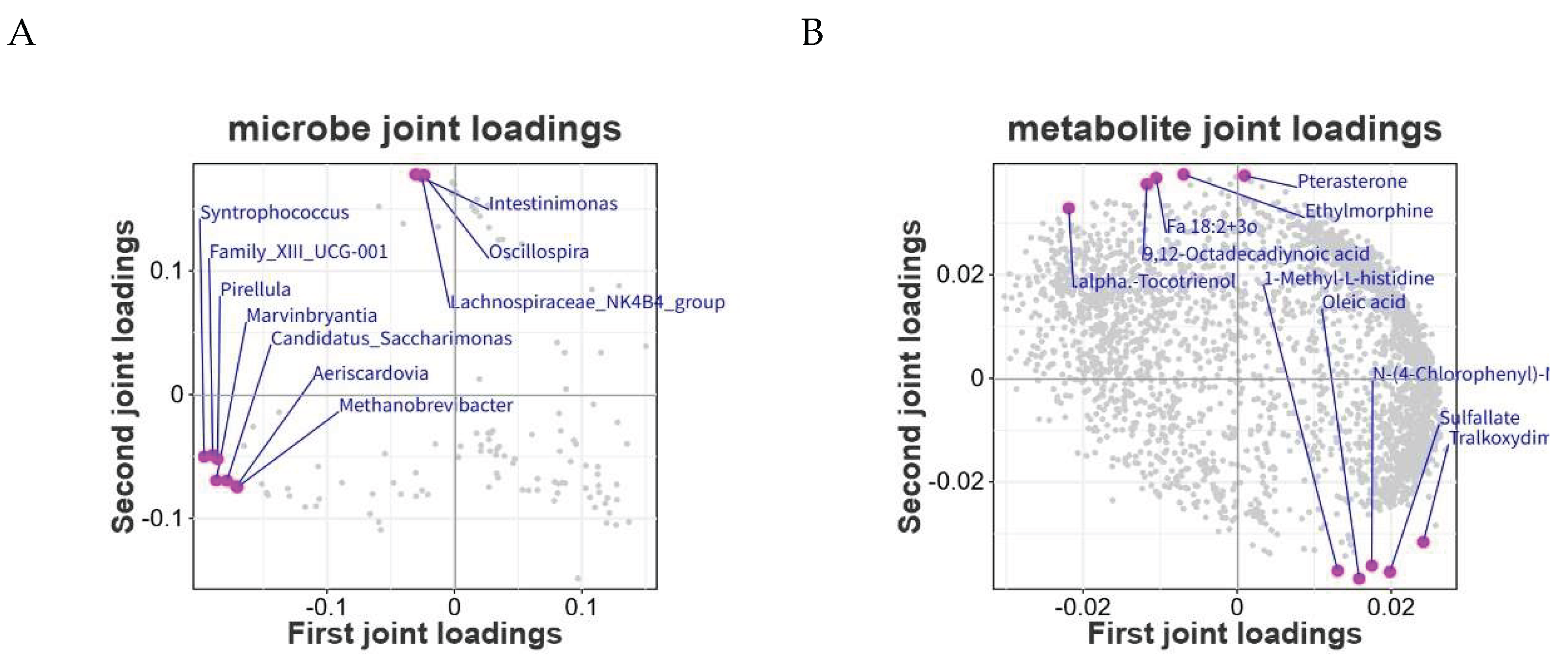
3.9. Integrated Analysis of 16S rRNA Gene Sequencing and Non-Targeted Metabolomics
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability
Competing Interests
References
- Liu X, Wei H, Shi H, Sha YZ, Wang JQ, Li SB, Lü WB, Guo XY, Pu XN, Luo YZ. Analysis of meat quality and fatty acid characteristics of grassland-grazing Tibetan sheep in different months [J]. Journal of Domestic Animal Ecology.
- Li Q. Screening, Analysis and Functional Research of Disease-Resistant Immune Genes in Tibetan Sheep [D]. Qinghai University, 2020.
- Johnson CL, Versalovic J. The human microbiome and its potential importance to pediatrics[J]. Pediatrics, 2012, 129(5): 950-960. [CrossRef]
- Mowat AM, Agace WW. Regional specialization within the intestinal immune system. Nat Rev Immunol. 2014 Oct;14(10):667-85. [CrossRef]
- ADOLPH TE, MAYR L, GRABHERR F, et al. Paneth cells and their antimicrobials in intestinal immunity[J]. CURR PHARM DESIGN, 2018, 24(10):1121-1129.
- Gong Z, Ye G, Xu S, He X. The characteristics of intestinal flora of Tibetan sheep in different regions at high altitude were revealed based on metagenomic technique[J]. Heliyon, 2024, 10(14):e34380.
- Zhao H, Mo Q, Kulyar M, Guan J, Zhang X, Luo X, Li J. Metagenomic analysis reveals a gut microbiota structure and function alteration between healthy and diarrheic juvenile yaks[J]. Animals, 2024, 14(8).
- Li H, Zhang XA. Research Progress on the Mechanism and Application of Pueraria lobata in the Treatment of Inflammatory Bowel Disease [J]. Chinese Herbal Medicines, 2025, 56(04): 1428-1439.
- Du Y, Gao Y, Hu M, Hou J, Yang L, Wang X, Du W, Liu J, Xu Q. Colonization and development of the gut microbiome in calves[J]. Journal of Animal Science and Biotechnology, 2023, 14: 46.
- Yang X, Fan X, Jiang H, Zhang Q, Dui B, Zhang Q, Dang S, Long R, Huang X. Simulated seasonal diets alter yak rumen microbiota structure and metabolic function[J]. Frontiers in Microbiology, 2022, 13: 1006285.
- Chen Y, Xiao L, Zhou M, Zhang H. The microbiota: a crucial mediator in gut homeostasis and colonization resistance[J]. Frontiers in Microbiology, 2024, 15: 1417864.
- Yang DX. Prevention and Control of Peste des Petits Ruminants in Tibetan Sheep [J]. Animal Husbandry and Veterinary Science, 2020, 79(19): 139-140.
- Xu BC, Li Z, Wang YZ, et al. Meta-Analysis of the Effects of Antimicrobial Peptides on Growth Performance, Diarrhea Rate and Immunoglobulin Levels in Piglets [J]. Chinese Journal of Animal Nutrition. 2020, 32(08):3584-3593.
- Wang ZX, Huang YQ, Liu YH, et al. Inhibitory Mechanism of Antimicrobial Peptides from Paenibacillus aiyuanensis on Candida albicans Biofilm [J]. Food Science, 2024, 45(21):176-184.
- Peng JJ, Chen YQ. Application of Insect-Derived Antimicrobial Peptides in Animal Husbandry Production [J]. Swine Industry Science, 2025, 42(02):87.
- Yu W, Guo X, Li X, Wei Y, Lyu Y, Zhang L, Wang J, Shan A. Novel multidomain peptide self-assembly biomaterials based on bola structure and terminal anchoring: Nanotechnology meets antimicrobial therapy. Mater Today Bio. 2024 Aug 6;28:101183.
- Ma X, Yang N, Mao R, Hao Y, Li Y, Guo Y, Teng D, Huang Y, Wang J. Self-assembly antimicrobial peptide for treatment of multidrug-resistant bacterial infection. J Nanobiotechnology. 2024 Oct 30;22(1):668.
- Zhang Z, Chen Y, Gao J, Yang M, Zhang D, Wang L, Zhang T, Cao Q, Mwangi J, He C, Li Y, Liu X, Jiang X, Kamau PM, Lai R. Orientational Nanoconjugation with Gold Endows Marked Antimicrobial Potential and Drugability of Ultrashort Dipeptides. Nano Lett. 2023 Dec 27;23(24):11874-11883.
- Aphale D, Kulkarni A. Modifications and optimization of manual methods for polymerase chain reaction and 16S rRNA gene sequencing quality community DNA extraction from goat rumen digesta. Vet World. 2018 Jul;11(7):990-1000.
- Mosley JD, Schock TB, Beecher CW, Dunn WB, Kuligowski J, Lewis MR, Theodoridis G, Ulmer Holland CZ, Vuckovic D, Wilson ID, Zanetti KA. Establishing a framework for best practices for quality assurance and quality control in untargeted metabolomics. Metabolomics. 2024 Feb 12;20(2):20.
- Chiang JY. Bile acid metabolism and signaling. Compr Physiol. 2013 Jul;3(3):1191-1212.
- Wang YN, Zhang ZH, Liu HJ, Guo ZY, Zou L, Zhang YM, Zhao YY. Integrative phosphatidylcholine metabolism through phospholipase A2 in rats with chronic kidney disease. Acta Pharmacol Sin. 2023 Feb;44(2):393-405.
- He M, Qin CX, Wang X, Ding NZ. Plant Unsaturated Fatty Acids: Biosynthesis and Regulation. Front Plant Sci. 2020 Apr 23;11:390.
- Law SH, Chan ML, Marathe GK, Parveen F, Chen CH, Ke LY. An Updated Review of Lysophosphatidylcholine Metabolism in Human Diseases. Int J Mol Sci. 2019 Mar 6;20(5):1149.
- Qin, J., Li, R., Raes, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
- Ren Q, Wang D, Han J, Liu Z, Wu Z. Lacrimispora sinapis sp. nov., isolated from pickled potherb mustard (Brassica juncea Coss.). Int J Syst Evol Microbiol. 2025 Feb;75(2):006675.
- Xie G, Jiang R, Wang X, Liu P, Zhao A, Wu Y, Huang F, Liu Z, Rajani C, Zheng X, Qiu J, Zhang X, Zhao S, Bian H, Gao X, Sun B, Jia W. Conjugated secondary 12α-hydroxylated bile acids promote liver fibrogenesis. EBioMedicine. 2021 Apr;66:103290.
- Jia W, Xie G, Jia W. Bile acid-microbiota crosstalk in gastrointestinal inflammation and carcinogenesis. Nat Rev Gastroenterol Hepatol. 2018 Feb;15(2):111-128.
- Chong J, Zhou Y, Li Z, Li X, Zhang J, Cao H, Ma J, Ge L, Zhong H, Sun J. Hyodeoxycholic acid modulates gut microbiota and bile acid metabolism to enhance intestinal barrier function in piglets. Front Vet Sci. 2025 Jun 20;12:1610956.
- Wang S, Zeng X, Yang Q, Qiao S. Antimicrobial Peptides as Potential Alternatives to Antibiotics in Food Animal Industry. Int J Mol Sci. 2016 May 3;17(5):603.
- Xu Z, Wang T, Wang Y, Li Y, Sun Y, Qiu HJ. Short-chain fatty acids: key antiviral mediators of gut microbiota. Front Immunol. 2025 Jul 25;16:1614879.
- Dias, B.d.C., Lamarca, A.P., Machado, D.T. et al. Metabolic pathways associated with Firmicutes prevalence in the gut of multiple livestock animals and humans. anim microbiome 7, 20 (2025).
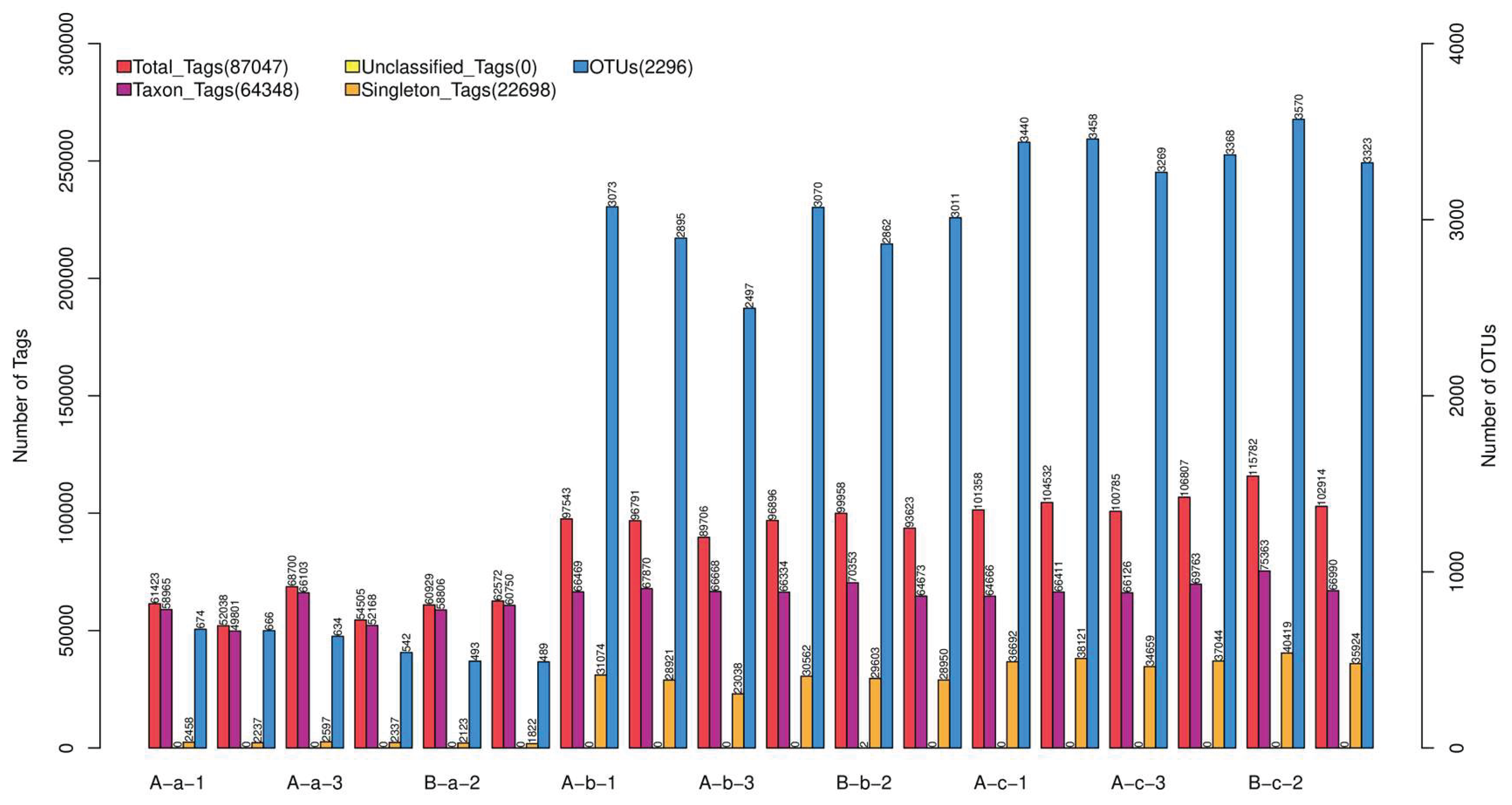
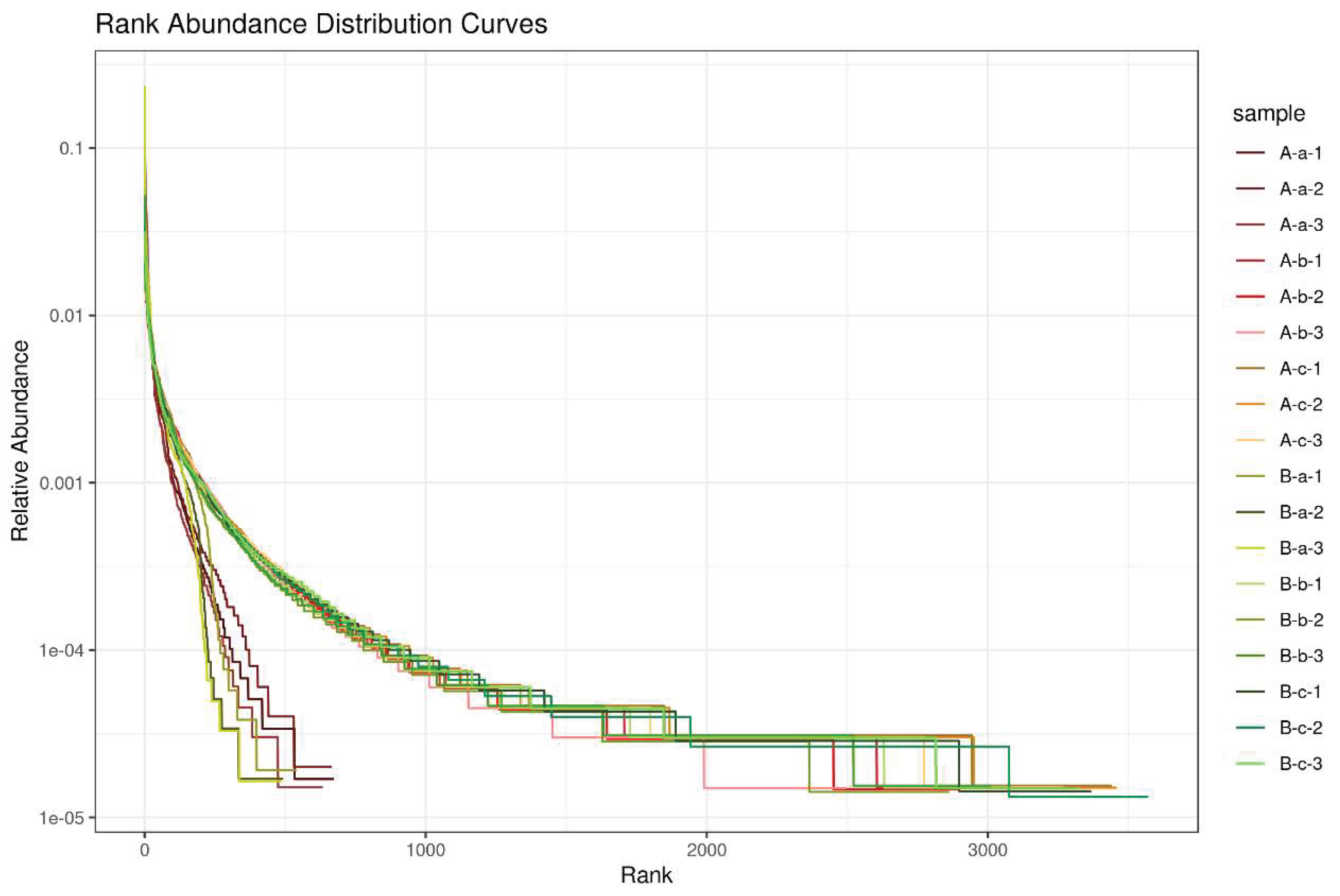
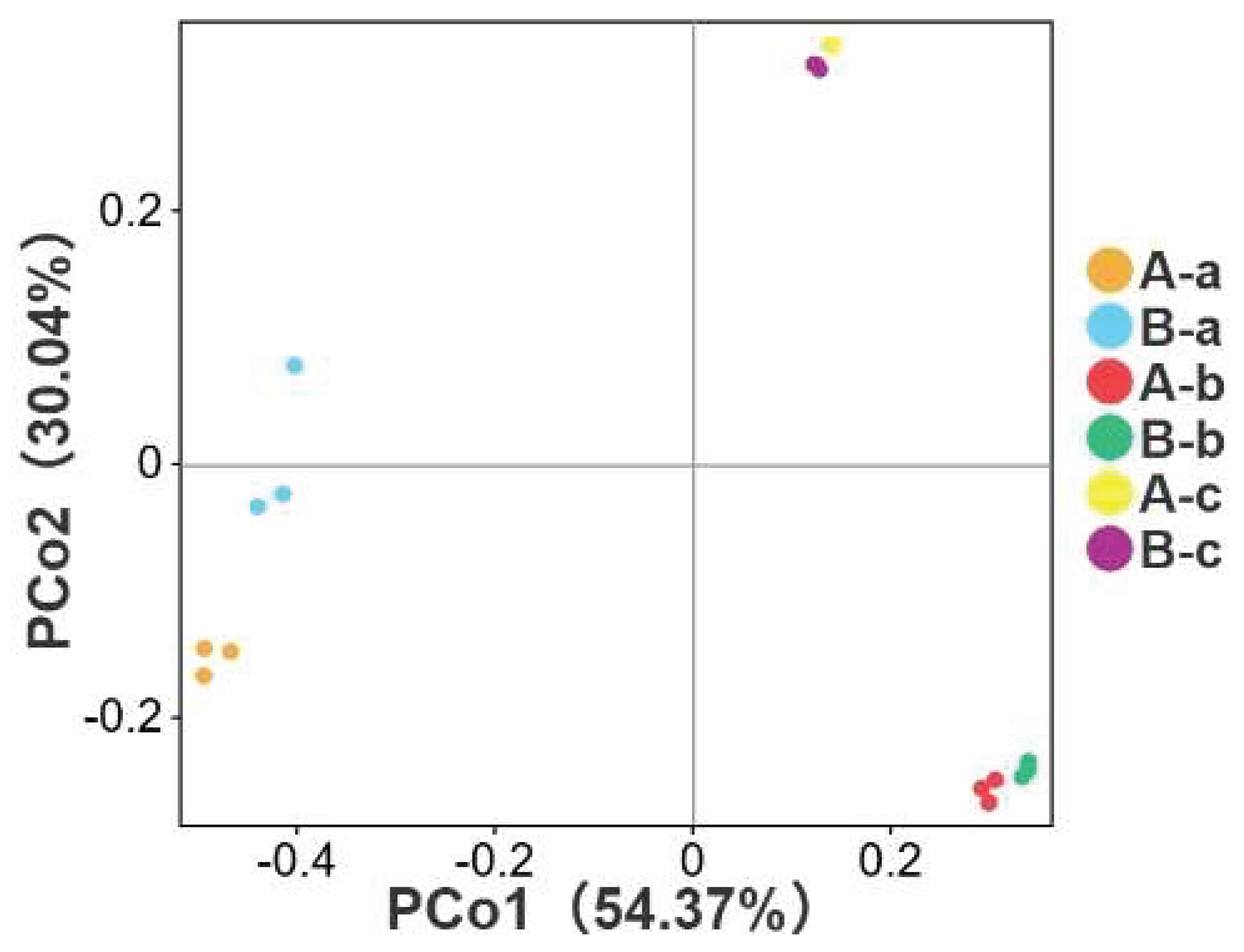

| Group | Sob (P value) | Chao1 (P value) | ACE (P value) |
| A-a vs B-a | 0.0029 | 0.0191 | 0.0062 |
| A-b vs B-b | 0.4549 | 0.4242 | 0.4097 |
| A-c- vs B-c | 0.7635 | 0.7670 | 0.8353 |
| Group | P value |
| A-a vs B-a | 0.03 |
| A-b vs B-b | 0.14 |
| A-c vs B-c | 0.07 |
| Genus | A-a-1/B-a-1 | A-a-2/B-a-2 | A-a-3/B-a-3 | Mean±SD |
| Group A | ||||
| Candidatus_Saccharimonas | 35.3091 | 28.1601 | 36.7487 | 33.4060 ± 4.5997 |
| Christensenellaceae_R-7_group | 3.0001 | 3.7911 | 3.0468 | 3.2793 ± 0.4438 |
| Ruminococcus | 2.7321 | 2.8092 | 1.8880 | 2.4764±0.5111 |
| NK4A214_group | 1.5246 | 1.5743 | 2.1102 | 1.7364±0.3247 |
| Saccharofermentans | 1.3109 | 1.0843 | 1.6868 | 1.3607 ± 0.3043 |
| Xylanibacter | 0.2273 | 0.9036 | 0.2723 | 0.4677 ± 0.3781 |
| UCG-005 | 0.0594 | 0.0161 | 0.0545 | 0.0433 ± 0.0237 |
| Rikenellaceae_RC9_gut_group | 0.0356 | 0.4377 | 0.0257 | 0.1663 ± 0.2351 |
| Treponema | 0.0153 | 0.0281 | 0.0076 | 0.0170 ± 0.0104 |
| Bacteroides | 0.0034 | 0.0080 | 0.0045 | 0.0053 ± 0.0024 |
| Group B | ||||
| Saccharofermentans | 12.7933 | 20.9417 | 23.6148 | 19.1166 ± 5.6369 |
| Candidatus_Saccharimonas | 16.1843 | 11.4155 | 9.0831 | 12.2276 ± 3.6196 |
| Christensenellaceae_R-7_group | 3.1475 | 4.6747 | 5.0815 | 4.3012 ± 1.0197 |
| NK4A214_group | 3.0114 | 2.2583 | 2.5926 | 2.6208 ± 0.3774 |
| Xylanibacter | 3.0133 | 0.5459 | 0.2699 | 1.2764 ± 1.5105 |
| Ruminococcus | 1.6600 | 0.94038 | 1.0140 | 1.2048 ± 0.3959 |
| Treponema | 0.9929 | 0.0002 | 0.0082 | 0.3338 ± 0.5708 |
| Rikenellaceae_RC9_gut_group | 0.7150 | 0.0238 | 0.0280 | 0.2556 ± 0.3979 |
| UCG-005 | 0.1591 | 0.0357 | 0.0807 | 0.0918 ± 0.0624 |
| Bacteroides | 0.0077 | 0.0017 | 0.0066 | 0.0053 ± 0.003 |
| Type | All | known | Unknown |
| POS | 9793 | 3916 | 5877 |
| NEG | 8309 | 3521 | 4788 |
| Model | R2X | R2Y | R2Xcorr | R2Ycorr |
| Genus | 0.838 | 0.873 | 0.664 | 0.838 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




