Submitted:
10 March 2026
Posted:
11 March 2026
You are already at the latest version
Abstract
Drug-induced toxicity remains a principal driver of attrition in pharmaceutical development, yet conventional screening paradigms typically address individual toxicity endpoints in isolation. Here, we introduce MultiEndpointTox, a chemoinformatics platform that simultaneously predicts seven critical drug toxicity endpoints—hERG cardiotoxicity, hepatotoxicity (DILI), nephrotoxicity (DIKI), Ames mutagenicity, skin sensitization, cytotoxicity, and reproductive toxicity (exploratory)—from molecular structures using curated datasets totaling over 18,000 compounds. The platform employs optimized classical machine learning models with systematic benchmarking of 2D topological descriptors (2240 features), enhanced multi-conformer 3D descriptors (1975 features from 5-conformer ensembles incorporating AUTOCORR3D, RDF, WHIM, and pharmacophore fingerprints), and hybrid representations. Under the tested conditions, 2D descriptors achieved the highest classification performance (AUC-ROC 0.859 ± 0.02), while enhanced 3D descriptors substantially narrowed the previously reported gap (AUC-ROC 0.833 ± 0.03 versus 0.69–0.73 for basic 14-feature 3D). Scaffold-based splitting provided rigorous generalization assessment, with an average performance reduction of approximately 8%. A multi-task learning framework via stacked generalization demonstrated cross-endpoint information sharing improves performance for 5 of 6 endpoints (average +2.1% AUC). The platform integrates leverage-based applicability domain assessment (31–100% coverage), SHAP-based feature importance analysis, and a confidence-weighted multi-endpoint risk scoring system validated on known drugs (AUC = 0.83, p = 4.06 × 10−14, Cliff’s δ = 0.66), with sensitivity analysis confirming robustness across five weight configurations (AUC range 0.72–0.98). External validation on independent benchmark datasets revealed the challenge of cross-dataset domain shift in computational toxicology. MultiEndpointTox is deployed as a production-ready REST API and publicly available at https://github.com/sharhabileltahir/MultiEndpointTox.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Collection and Curation
2.2. Molecular Descriptor Calculation
2.3. Machine Learning Models
2.4. Multi-Task Learning
2.5. Validation Strategy
2.6. Applicability Domain Assessment
2.7. Model Interpretability
2.8. Integrated Multi-Endpoint Risk Scoring
2.9. Software Implementation
3. Results
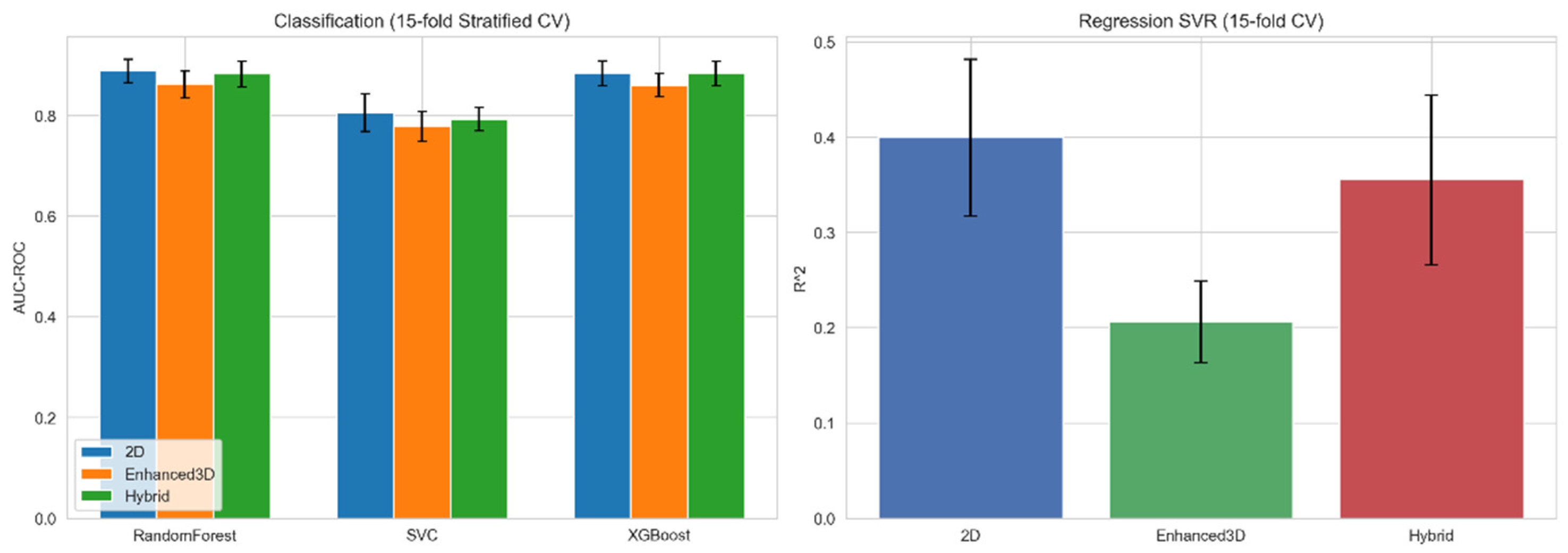
3.1. Enhanced 3D Descriptors Narrow the Representation Gap
3.2. Multi-Endpoint Model Performance
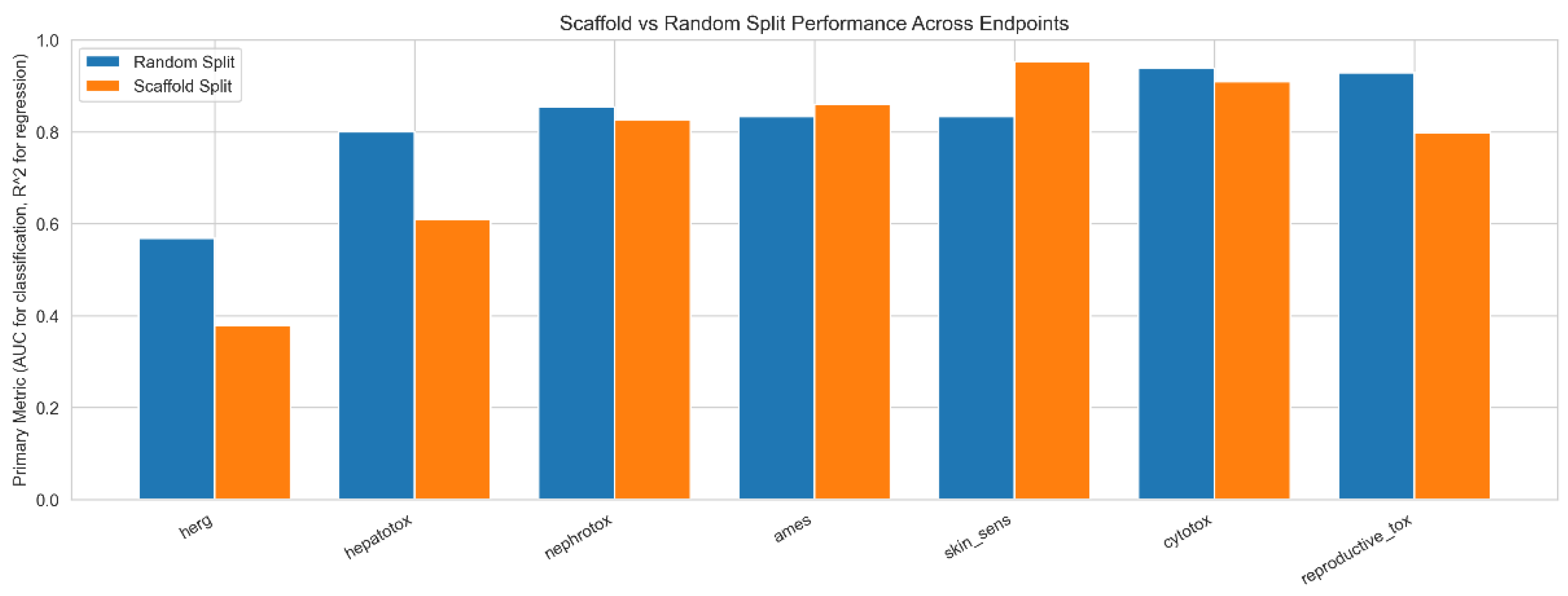
3.3. Scaffold-Based Validation Reveals Generalization Boundaries
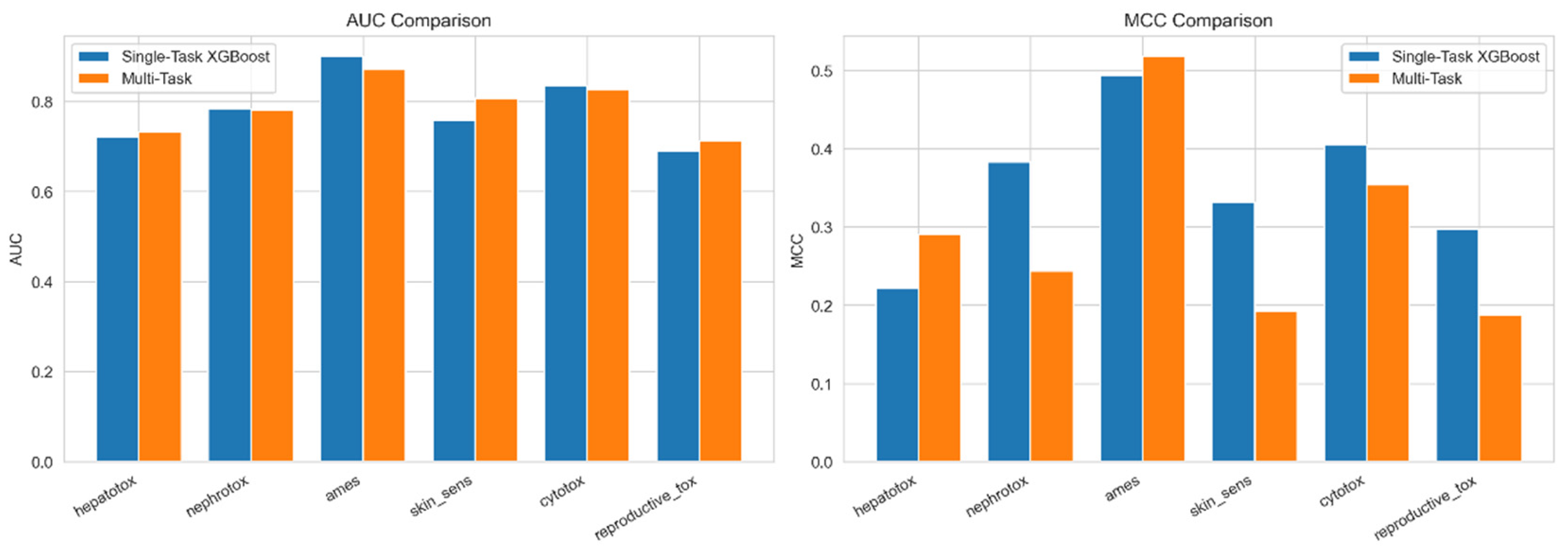
3.4. Multi-Task Learning Improves Data-Scarce Endpoints
| Endpoint | Single AUC | Multi AUC | Δ AUC |
| Hepatotox. | 0.691 | 0.713 | +0.022 |
| Nephrotox. | 0.694 | 0.717 | +0.023 |
| Ames | 0.772 | 0.779 | +0.007 |
| Skin Sens. | 0.791 | 0.820 | +0.029 |
| Cytotox. | 0.882 | 0.914 | +0.032 |
| Repro. Tox.† | 0.524 | 0.517 | −0.007 |
| Hepatotox. | 0.691 | 0.713 | +0.022 |
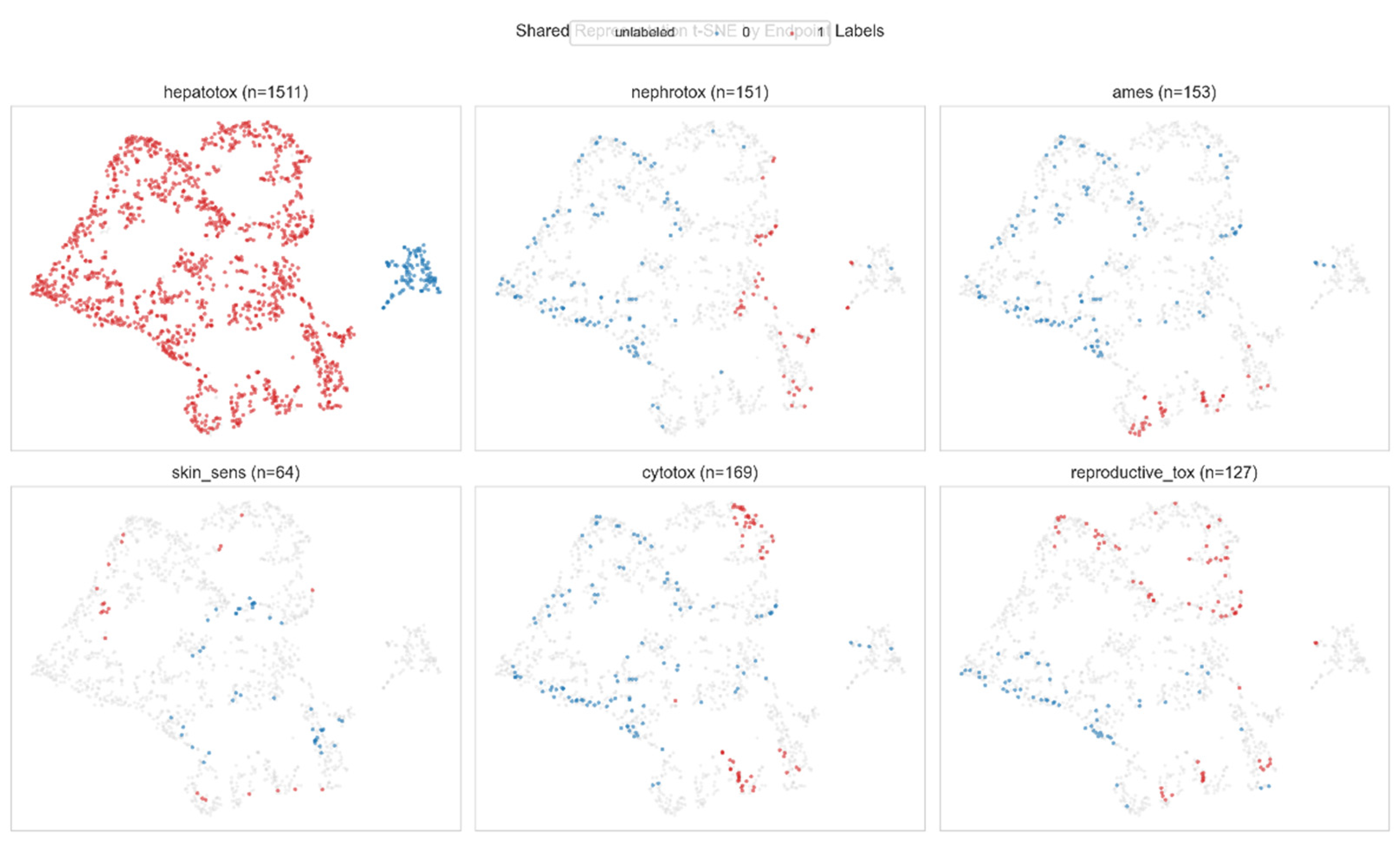
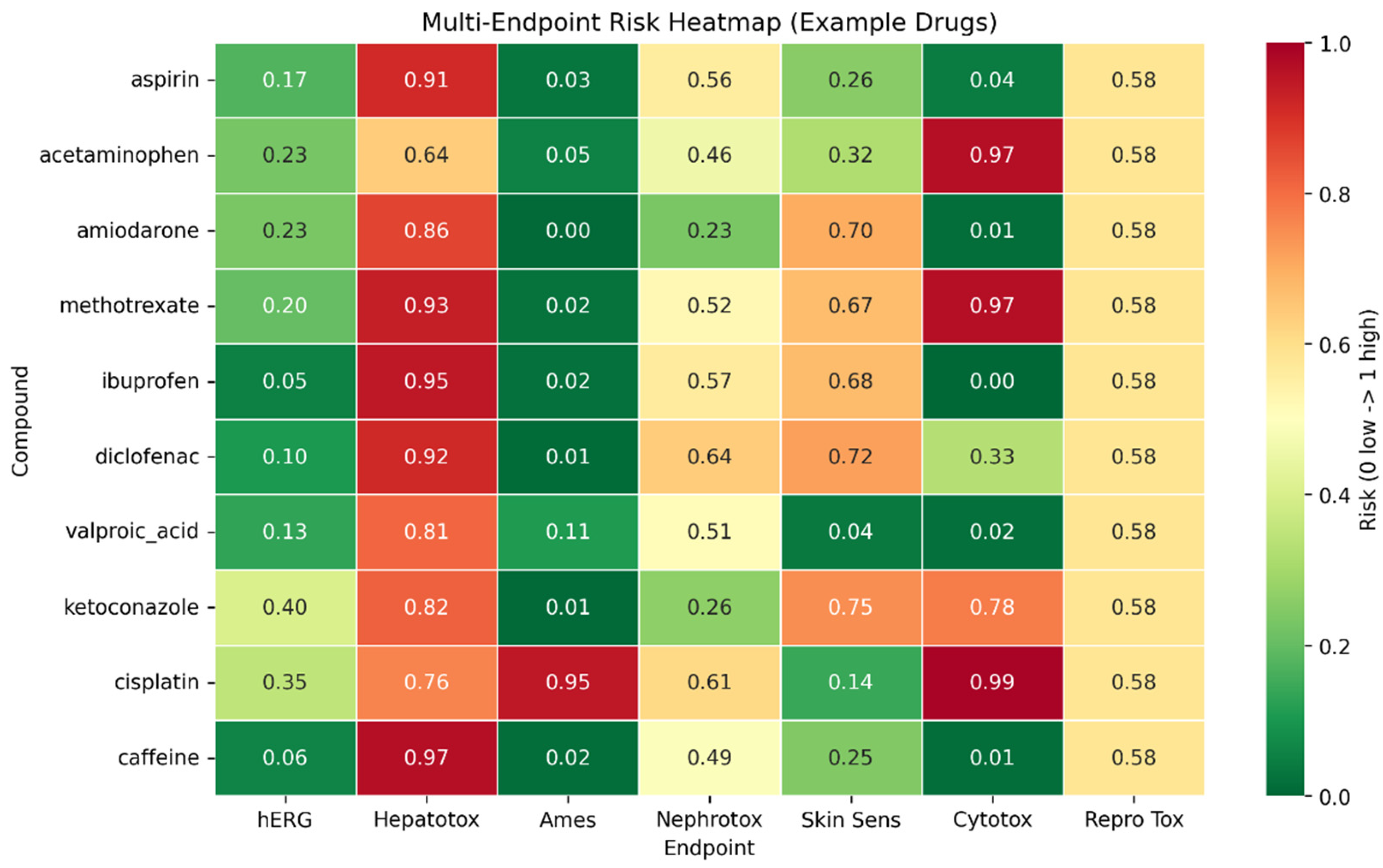
3.5. Integrated Risk Scoring with Sensitivity Analysis
| Metric | Value |
| Mann-Whitney U | 8,057 |
| p-value | 4.06 × 10−14 |
| Cliff’s δ | 0.66 (large effect) |
| Risk Score AUC | 0.83 |
| n (high-risk / safe) | 100 / 100 |
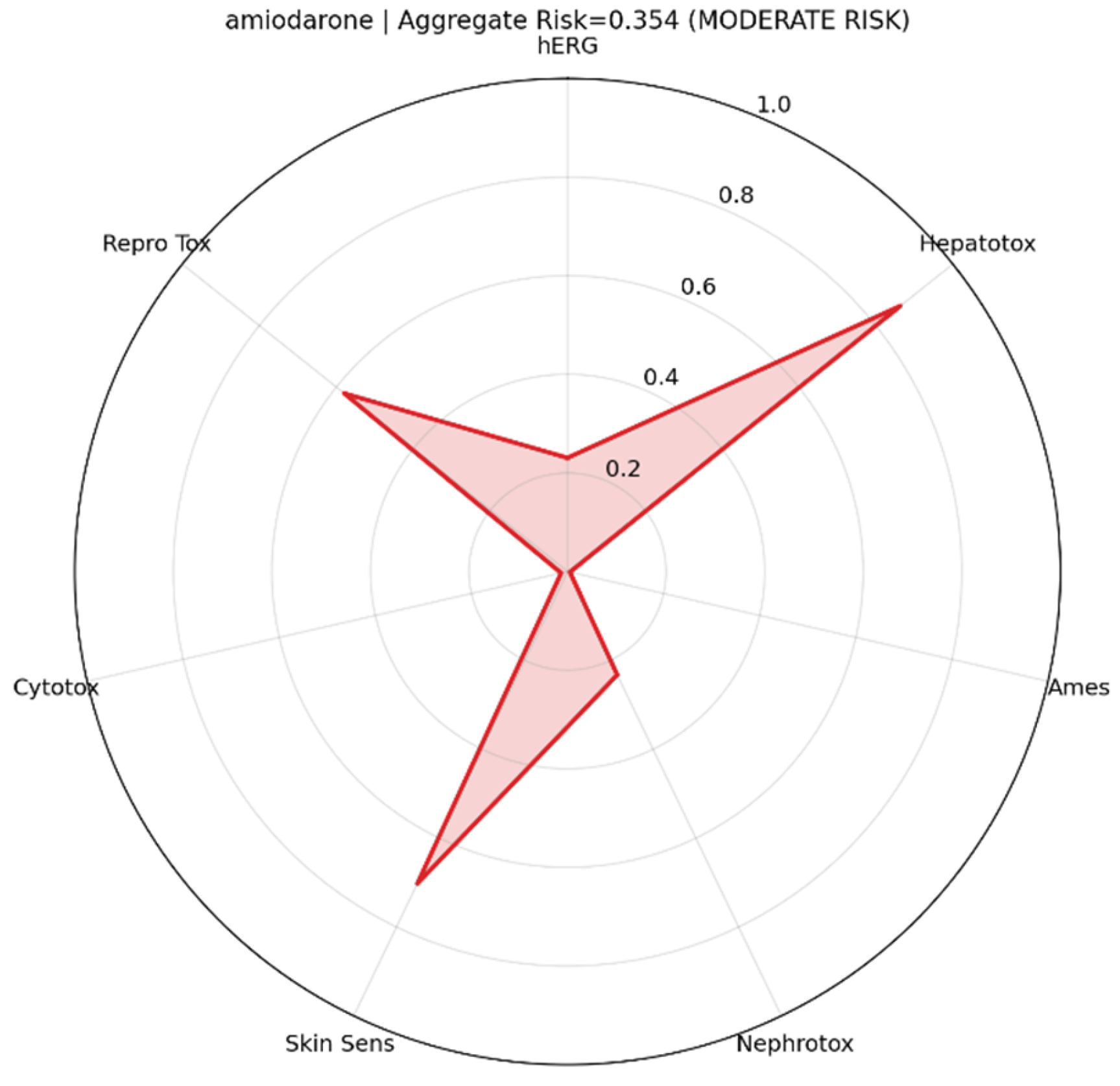
3.6. SHAP Feature Importance Analysis
3.7. External Validation on Independent Benchmark Datasets
4. Discussion
4.1. Multidimensional Toxicity Profiling in Context
4.2. Resolving the 2D versus 3D Descriptor Debate
4.3. Scaffold Validation and Generalization
4.4. Mechanistic Considerations for Hepatotoxicity Prediction
4.5. Cross-Endpoint Information Transfer
4.6. Risk Score Interpretation and Robustness
4.7. Applicability Domain Considerations
4.8. External Validation and Domain Shift
4.9. Limitations
4.10. Future Directions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Use of Artificial Intelligence
Conflicts of Interest
References
- Kola, I.; Landis, J. Can the pharmaceutical industry reduce attrition rates? Nat. Rev. Drug Discov. 2004, 3, 711–715. [Google Scholar] [CrossRef]
- Onakpoya, I.J.; Heneghan, C.J.; Aronson, J.K. Post-marketing withdrawal of 462 medicinal products. BMC Med. 2016, 14, 10. [Google Scholar] [CrossRef]
- Russell, W.M.S.; Burch, R.L. The Principles of Humane Experimental Technique; Methuen: London, UK, 1959. [Google Scholar]
- European Commission. Directive 2010/63/EU. Off. J. Eur. Union 2010, L276, 33–79. [Google Scholar]
- Raies, A.B.; Bajic, V.B. In silico toxicology: computational methods for chemical toxicity prediction. WIREs Comput. Mol. Sci. 2016, 6, 147–172. [Google Scholar] [CrossRef]
- Mayr, A.; Klambauer, G.; Unterthiner, T.; Hochreiter, S. DeepTox: Toxicity prediction using deep learning. Front. Environ. Sci. 2016, 3, 80. [Google Scholar] [CrossRef]
- Tropsha, A.; Gramatica, P.; Gombar, V.K. The importance of being earnest: validation for QSPR models. QSAR Comb. Sci. 2003, 22, 69–77. [Google Scholar] [CrossRef]
- Netzeva, T.I.; Worth, A.; Aldenberg, T.; et al. Current status of methods for defining the applicability domain of (Q)SARs. ATLA 2005, 33, 155–173. [Google Scholar]
- Wu, Z.; Ramsundar, B.; Feinberg, E.N.; et al. MoleculeNet: a benchmark for molecular machine learning. Chem. Sci. 2018, 9, 513–530. [Google Scholar] [CrossRef]
- Axelrod, S.; Gomez-Bombarelli, R. GEOM, energy-annotated molecular conformations. Sci. Data 2022, 9, 185. [Google Scholar] [CrossRef] [PubMed]
- Chicco, D.; Jurman, G. The advantages of MCC over F1 score and accuracy. BMC Genomics 2020, 21, 6. [Google Scholar]
- OECD. Guidance Document on the Validation of (Q)SAR Models; No. 69; OECD Publishing: Paris, France, 2014; Volume No. 69. [Google Scholar]
- Yang, K.; Swanson, K.; Jin, W.; et al. Analyzing learned molecular representations. J. Chem. Inf. Model. 2019, 59, 3370–3388. [Google Scholar] [CrossRef] [PubMed]
- Withnall, M.; Lindelof, E.; Engkvist, O.; Chen, H. Building attention and edge MPNN. J. Cheminform. 2020, 12, 1. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Wu, Z.; Hsieh, C.Y.; et al. Could GNNs learn better molecular representation? J. Cheminform. 2021, 13, 12. [Google Scholar] [CrossRef]
- Walters, W.P.; Barzilay, R. Deep learning in molecule generation and property prediction. Acc. Chem. Res. 2021, 54, 263–270. [Google Scholar] [CrossRef]
- Huang, K.; Fu, T.; Gao, W.; et al. Therapeutics Data Commons. Proc. NeurIPS Track Datasets Benchmarks, 2021. [Google Scholar]
- Banerjee, P.; Kemmler, E.; Dunkel, M.; Preissner, R. ProTox 3.0: prediction of toxicity of chemicals. Nucleic Acids Res. 2024, 52, W513–W520. [Google Scholar] [CrossRef] [PubMed]
- Xiong, G.; Wu, Z.; Yi, J.; et al. ADMETlab 3.0: an updated ADMET prediction platform. Nucleic Acids Res. 2024, 52, W422–W431. [Google Scholar]
- Pires, D.E.V.; Blundell, T.L.; Ascher, D.B. pkCSM: Predicting pharmacokinetic and toxicity properties using graph-based signatures. J. Med. Chem. 2015, 58, 4066–4072. [Google Scholar] [CrossRef] [PubMed]
- Sanguinetti, M.C.; Tristani-Firouzi, M. hERG potassium channels and cardiac arrhythmia. Nature 2006, 440, 463–469. [Google Scholar] [CrossRef]
- Chen, M.; Suzuki, A.; Thakkar, S.; et al. DILIrank. Drug Discov. Today 2016, 21, 648–653. [Google Scholar] [CrossRef]
- Hoste, E.A.J.; Kellum, J.A.; Selby, N.M.; et al. Global epidemiology of acute kidney injury. Nat. Rev. Nephrol. 2018, 14, 607–625. [Google Scholar] [CrossRef]
- Ames, B.N.; McCann, J.; Yamasaki, E. Methods for detecting carcinogens and mutagens. Mutat. Res. 1975, 31, 347–364. [Google Scholar] [CrossRef] [PubMed]
- Karlberg, A.T.; Bergstrom, M.A.; Borje, A.; et al. Allergic contact dermatitis. Chem. Res. Toxicol. 2008, 21, 53–69. [Google Scholar] [CrossRef] [PubMed]
- Riss, T.L.; Moravec, R.A.; Niles, A.L.; et al. Cell Viability Assays. In Assay Guidance Manual; Eli Lilly & NCATS: Bethesda, MD, USA, 2004. [Google Scholar]
- Daston, G.P. Laboratory models and teratogenesis. Am. J. Med. Genet. C 2011, 157C, 183–187. [Google Scholar] [CrossRef] [PubMed]
- Mendez, D.; Gaulton, A.; Bento, A.P.; et al. ChEMBL: towards direct deposition of bioassay data. Nucleic Acids Res. 2019, 47, D930–D940. [Google Scholar] [CrossRef]
- Richard, A.M.; Judson, R.S.; Houck, K.A.; et al. ToxCast chemical landscape. Chem. Res. Toxicol. 2016, 29, 1225–1251. [Google Scholar] [CrossRef]
- Fourches, D.; Muratov, E.; Tropsha, A. Trust, but verify: chemical structure curation. J. Chem. Inf. Model. 2010, 50, 1189–1204. [Google Scholar] [CrossRef]
- Landrum, G. RDKit: Open-Source Cheminformatics. Available online: https://www.rdkit.org (accessed on 1 March 2026).
- Rogers, D.; Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 2010, 50, 742–754. [Google Scholar] [CrossRef]
- Durant, J.L.; Leland, B.A.; Henry, D.R.; Nourse, J.G. Reoptimization of MDL keys. J. Chem. Inf. Comput. Sci. 2002, 42, 1273–1280. [Google Scholar] [CrossRef]
- Riniker, S.; Landrum, G.A. Better informed distance geometry. J. Chem. Inf. Model. 2015, 55, 2562–2574. [Google Scholar] [CrossRef]
- Rappe, A.K.; Casewit, C.J.; Colwell, K.S.; et al. UFF force field. J. Am. Chem. Soc. 1992, 114, 10024–10035. [Google Scholar] [CrossRef]
- Stiefl, N.; Watson, I.A.; Baumann, K.; Bender, A. ErG: 2D pharmacophore descriptions for scaffold hopping. J. Chem. Inf. Model. 2006, 46, 208–220. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. XGBoost. Proc. 22nd ACM SIGKDD, 2016; ACM: New York, NY, USA; pp. 785–794. [Google Scholar]
- Ke, G.; Meng, Q.; Finley, T.; et al. LightGBM. Adv. Neural Inf. Process. Syst. 2017, 30, 3146–3154. [Google Scholar]
- Akiba, T.; Sano, S.; Yanase, T.; et al. Optuna. Proc. 25th ACM SIGKDD, 2019; ACM: New York, NY, USA; pp. 2623–2631. [Google Scholar]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Heid, E.; Greenman, K.P.; Chung, Y.; et al. Chemprop. J. Chem. Inf. Model. 2024, 64, 9–17. [CrossRef]
- Bemis, G.W.; Murcko, M.A. Properties of known drugs. 1. Molecular frameworks. J. Med. Chem. 1996, 39, 2887–2893. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.; Fu, T.; Gao, W.; et al. Therapeutics Data Commons. Proc. NeurIPS Track Datasets Benchmarks, 2021. [Google Scholar]
- Gramatica, P. Principles of QSAR models validation. QSAR Comb. Sci. 2007, 26, 694–701. [Google Scholar] [CrossRef]
- Lundberg, S.M.; Lee, S.I. A unified approach to interpreting model predictions. Adv. Neural Inf. Process. Syst. 2017, 30, 4765–4774. [Google Scholar]
- Aronov, A.M. Predictive in silico modeling for hERG blockers. Drug Discov. Today 2005, 10, 149–155. [Google Scholar] [CrossRef]
- Cramer, R.D.; Patterson, D.E.; Bunce, J.D. CoMFA. J. Am. Chem. Soc. 1988, 110, 5959–5967. [Google Scholar] [CrossRef] [PubMed]
- Thakkar, S.; Li, T.; Liu, Z.; et al. DILIst: binary classification of 1279 drugs. Drug Discov. Today 2020, 25, 201–208. [Google Scholar] [CrossRef] [PubMed]
| Endpoint | Task | n | Source | Class Distribution |
| hERG | Regression | 7889 | ChEMBL | pIC50: 3.0–10.0 |
| Hepatotoxicity | Cls. | 1597 | ChEMBL, FDA | 93% toxic / 7% non-toxic |
| Nephrotoxicity | Cls. | 565 | FDA DIRIL, Lit. | 58% toxic / 42% non-toxic |
| Ames Mutagenicity | Cls. | 6512 | ChEMBL, Lit. | 52% mut. / 48% non-mut. |
| Skin Sensitization | Cls. | 1100 | ChEMBL, LLNA | 55% sens. / 45% non-sens. |
| Cytotoxicity | Cls. | 8371 | ToxCast, ChEMBL | 62% toxic / 38% non-toxic |
| Repro. Tox.† | Cls. | 127 | Literature | 55% toxic / 45% non-toxic |
| Repr. | n Features | Cls. AUC-ROC | Reg. R2 (SVR) | Repr. |
| 2D | 2240 | 0.859 ± 0.02 | 0.399 ± 0.04 | 2D |
| Enhanced 3D | 1975 | 0.833 ± 0.03 | 0.206 ± 0.05 | Enhanced 3D |
| Hybrid (2D+3D) | 4215 | 0.853 ± 0.02 | 0.355 ± 0.05 | Hybrid (2D+3D) |
| Endpoint | Best Model | Features | CV AUC/R2 | AD (%) |
| hERG | SVR | 500 | R2 = 0.57 ± 0.03 | 98.9 |
| Hepatotox. | XGBoost | 500 | AUC = 0.80 ± 0.04 | 78.9 |
| Nephrotox. | RF | 500 | AUC = 0.91 ± 0.05 | 93.5 |
| Ames | XGBoost | 500 | AUC = 0.92 ± 0.02 | 100 |
| Skin Sens. | RF | 318 | AUC = 0.87 ± 0.06 | 31.0 |
| Cytotox. | SVC | 500 | AUC = 0.89 ± 0.03 | 100 |
| Repro. Tox.† | SVC | 500 | AUC = 0.77 ± 0.10 | 100 |
| Endpoint | Random | Scaffold | Δ |
| hERG (R2) | 0.568 | 0.378 | −0.190 |
| Hepatotox. (AUC) | 0.801 | 0.764 | −0.037 |
| Nephrotox. (AUC) | 0.909 | 0.827 | −0.082 |
| Ames (AUC) | 0.921 | 0.839 | −0.082 |
| Skin Sens. (AUC) | 0.867 | 0.821 | −0.046 |
| Cytotox. (AUC) | 0.894 | 0.901 | +0.007 |
| Repro. Tox.† (AUC) | 0.772 | 0.588 | −0.184 |
| Scenario | AUC | p-value | Cliff’s δ |
| Baseline | 0.83 | 4.4 × 10−16 | 0.66 |
| Equal weights | 0.72 | 5.4 × 10−8 | 0.44 |
| Cardiac-focused | 0.98 | 8.5 × 10−32 | 0.96 |
| Hepatic-focused | 0.78 | 4.2 × 10−12 | 0.56 |
| Genotox-focused | 0.79 | 5.1 × 10−13 | 0.58 |
| Endpoint | External Dataset | Type | n | AUC/R2 | MCC |
| Ames | TDC Ames benchmark | External | 7183 | 0.603 | 0.131 |
| Hepatotox. | TDC DILI benchmark | External | 3* | n/a | n/a |
| Cytotox. | Tox21 SR-MMP | External | 5646 | 0.473 | −0.019 |
| hERG | ChEMBL post-2020 | Pseudo-ext. | 2419 | 0.038 | n/a |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




