Submitted:
09 March 2026
Posted:
11 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patient and Public Involvement
2.2. Sources of Data and the Search Strategy
2.3. Mapping Analysis
3. Results
3.1. General Data
3.2. Global Research Contributions and Collaborative Networks
3.3. Influential Journals and Citation Impact
3.4. Active Authors
3.5. Key Studies and Citation Dynamics
3.6. Thematic Evolution and Emerging Research Frontiers
3.7. Hotspot Analysis of microRNAs in Eye-Related Diseases
4. Discussion
4.1. Evolving Role of miRNAs in Eye-Related Diseases: Insights and Emerging Themes
4.2. MicroRNAs in Neurodegenerative Diseases (NDs)
4.3. MicroRNAs in Ophthalmic Diseases (ODs)
4.3.1. Corneal Disorders
Corneal squamous cell carcinoma
Corneal neovascularization
Endothelial dystrophy
Herpes Epithelial Keratitis (HEK)
Herpes Stromal Keratitis (HSK)
Fungal Keratitis (FK)
Bacterial Keratitis
4.3.2. Conjunctival Disorders
Pterygium
Allergic Conjunctivitis (AC)
Conjunctival Lymphoma
Conjunctival Melanoma
Vernal Keratoconjunctivitis
Trachoma
4.3.3. Lens Diseases and Refractive Error
Cataract
Myopia
4.3.4. Retinopathy
Diabetic retinopathy
Macular degeneration
Polypoidal Choroidal Vasculopathy
4.3.5. Optic Neuropathy
Glaucoma
Pseudoexfoliation Syndrome
Neuromyelitis Optica Spectrum Disorder (NMOSD)
4.3.6. Autoimmune Diseases
Graves’ Ophthalmopathy
Autoimmune Uveitis
Dry Eye Disease
Sjögren’s Syndrome
4.3.7. Hereditary Diseases
Keratoconus
EDICT Syndrome
Aniridia
Retinoblastoma
Retinitis pigmentosa
Leber Hereditary Optic Neuropathy
4.4. The Promise and Challenges of miRNA Biomarkers
3.7. Limitations and Future Directions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cai, Y.; Yu, X.; Hu, S.; Yu, J. A brief review on the mechanisms of miRNA regulation. Genomics Proteomics Bioinformatics 2009, 7, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Horvitz, H.R.; Sulston, J.E. Isolation and genetic characterization of cell-lineage mutants of the nematode Caenorhabditis elegans. Genetics 1980, 96, 435–454. [Google Scholar] [CrossRef]
- Ferguson, E.L.; Sternberg, P.W.; Horvitz, H.R. A genetic pathway for the specification of the vulval cell lineages of Caenorhabditis elegans. Nature 1987, 326, 259–267. [Google Scholar] [CrossRef]
- Backes, C.; Meese, E.; Keller, A. Specific miRNA Disease Biomarkers in Blood, Serum and Plasma: Challenges and Prospects. Mol Diagn Ther 2016, 20, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Vishnoi, A.; Rani, S. miRNA Biogenesis and Regulation of Diseases: An Updated Overview. Methods Mol Biol 2023, 2595, 1–12. [Google Scholar] [CrossRef]
- Wojciechowska, A.; Braniewska, A.; Kozar-Kamińska, K. MicroRNA in cardiovascular biology and disease. Adv Clin Exp Med 2017, 26, 865–874. [Google Scholar] [CrossRef]
- Wang, Y.; Niu, L.; Zhao, J.; Wang, M.; Li, K.; Zheng, Y. An update: mechanisms of microRNA in primary open-angle glaucoma. Brief Funct Genomics 2021, 20, 19–27. [Google Scholar] [CrossRef]
- Bagga, S.; Bracht, J.; Hunter, S.; Massirer, K.; Holtz, J.; Eachus, R.; Pasquinelli, A.E. Regulation by let-7 and lin-4 miRNAs results in target mRNA degradation. Cell 2005, 122, 553–563. [Google Scholar] [CrossRef]
- Meister, G. miRNAs get an early start on translational silencing. Cell 2007, 131, 25–28. [Google Scholar] [CrossRef]
- Fioravanti, A.; Pirtoli, L.; Giordano, A.; Dotta, F. Crosstalk between MicroRNA and Oxidative Stress in Physiology and Pathology. Int J Mol Sci 2020, 21. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.; Xu, S.Y.; Chen, L.S.; Liang, G.Y.; Pu, Y.P.; Yin, L.H. Trends of long noncoding RNA research from 2007 to 2016: a bibliometric analysis. Oncotarget 2017, 8, 83114–83127. [Google Scholar] [CrossRef]
- Shuaib, W.; Khan, M.S.; Shahid, H.; Valdes, E.A.; Alweis, R. Bibliometric analysis of the top 100 cited cardiovascular articles. Am J Cardiol 2015, 115, 972–981. [Google Scholar] [CrossRef] [PubMed]
- Narotsky, D.; Green, P.H.; Lebwohl, B. Temporal and geographic trends in celiac disease publications: a bibliometric analysis. Eur J Gastroenterol Hepatol 2012, 24, 1071–1077. [Google Scholar] [CrossRef]
- Geaney, F.; Scutaru, C.; Kelly, C.; Glynn, R.W.; Perry, I.J. Type 2 Diabetes Research Yield, 1951-2012: Bibliometrics Analysis and Density-Equalizing Mapping. PLoS One 2015, 10, e0133009. [Google Scholar] [CrossRef]
- van Eck, N.J.; Waltman, L. Software survey: VOSviewer, a computer program for bibliometric mapping. Scientometrics 2010, 84, 523–538. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Wang, R.; Jin, L.; Tian, Y.; Jin, H.; Han, Y.; Sun, C.; Ding, M.; Guo, H. MiR-223-3p Aggravates Ocular Inflammation in Sjögren's Syndrome. Endocr Metab Immune Disord Drug Targets 2023, 23, 1087–1095. [Google Scholar] [CrossRef]
- Plastino, F.; Pesce, N.A.; André, H. MicroRNAs and the HIF/VEGF axis in ocular neovascular diseases. Acta Ophthalmol 2021, 99, e1255–e1262. [Google Scholar] [CrossRef]
- Mead, B.; Cullather, E.; Nakaya, N.; Niu, Y.; Kole, C.; Ahmed, Z.; Tomarev, S. Viral delivery of multiple miRNAs promotes retinal ganglion cell survival and functional preservation after optic nerve crush injury. Exp Eye Res 2020, 197, 108071. [Google Scholar] [CrossRef]
- Piona, C.; Costantini, S.; Zusi, C.; Cozzini, T.; Pedrotti, E.; Marigliano, M.; Fornari, E.; Maguolo, A.; Morandi, A.; Maffeis, C. Early marker of ocular neurodegeneration in children and adolescents with type 1 diabetes: the contributing role of polymorphisms in mir146a and mir128a genes. Acta Diabetol 2022, 59, 1551–1561. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Wang, Y.; Chen, J.; Gao, X.; Pan, S.; Su, Y.; Zhou, X. Co-delivery of brinzolamide and miRNA-124 by biodegradable nanoparticles as a strategy for glaucoma therapy. Drug Deliv 2020, 27, 410–421. [Google Scholar] [CrossRef]
- Smit-McBride, Z.; Morse, L.S. MicroRNA and diabetic retinopathy-biomarkers and novel therapeutics. Ann Transl Med 2021, 9, 1280. [Google Scholar] [CrossRef]
- Kakan, S.S.; Edman, M.C.; Yao, A.; Okamoto, C.T.; Nguyen, A.; Hjelm, B.E.; Hamm-Alvarez, S.F. Tear miRNAs Identified in a Murine Model of Sjögren's Syndrome as Potential Diagnostic Biomarkers and Indicators of Disease Mechanism. Front Immunol 2022, 13, 833254. [Google Scholar] [CrossRef]
- Shalgi, R.; Lieber, D.; Oren, M.; Pilpel, Y. Global and local architecture of the mammalian microRNA-transcription factor regulatory network. PLoS Comput Biol 2007, 3, e131. [Google Scholar] [CrossRef]
- Nelson, P.T.; Wang, W.X.; Rajeev, B.W. MicroRNAs (miRNAs) in neurodegenerative diseases. Brain Pathol 2008, 18, 130–138. [Google Scholar] [CrossRef]
- Juźwik, C.A.; Drake, S.; Lécuyer, M.A.; Johnson, R.M.; Morquette, B.; Zhang, Y.; Charabati, M.; Sagan, S.M.; Bar-Or, A.; Prat, A.; et al. Neuronal microRNA regulation in Experimental Autoimmune Encephalomyelitis. Sci Rep 2018, 8, 13437. [Google Scholar] [CrossRef]
- Bredesen, D.E.; Rao, R.V.; Mehlen, P. Cell death in the nervous system. Nature 2006, 443, 796–802. [Google Scholar] [CrossRef]
- Glass, C.K.; Saijo, K.; Winner, B.; Marchetto, M.C.; Gage, F.H. Mechanisms underlying inflammation in neurodegeneration. Cell 2010, 140, 918–934. [Google Scholar] [CrossRef] [PubMed]
- Maciotta, S.; Meregalli, M.; Torrente, Y. The involvement of microRNAs in neurodegenerative diseases. Front Cell Neurosci 2013, 7, 265. [Google Scholar] [CrossRef] [PubMed]
- Swarbrick, S.; Wragg, N.; Ghosh, S.; Stolzing, A. Systematic Review of miRNA as Biomarkers in Alzheimer's Disease. Mol Neurobiol 2019, 56, 6156–6167. [Google Scholar] [CrossRef] [PubMed]
- Dolati, S.; Marofi, F.; Babaloo, Z.; Aghebati-Maleki, L.; Roshangar, L.; Ahmadi, M.; Rikhtegar, R.; Yousefi, M. Dysregulated Network of miRNAs Involved in the Pathogenesis of Multiple Sclerosis. Biomed Pharmacother 2018, 104, 280–290. [Google Scholar] [CrossRef]
- Juźwik, C.A.; S.D., S; Zhang, Y.; Paradis-Isler, N.; Sylvester, A.; Amar-Zifkin, A.; Douglas, C.; Morquette, B.; Moore, C.S.; Fournier, A.E. microRNA dysregulation in neurodegenerative diseases: A systematic review. Prog Neurobiol 2019, 182, 101664. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Ryan, D.G.; Getsios, S.; Oliveira-Fernandes, M.; Fatima, A.; Lavker, R.M. MicroRNA-184 antagonizes microRNA-205 to maintain SHIP2 levels in epithelia. Proc Natl Acad Sci U S A 2008, 105, 19300–19305. [Google Scholar] [CrossRef] [PubMed]
- Karali, M.; Peluso, I.; Gennarino, V.A.; Bilio, M.; Verde, R.; Lago, G.; Dollé, P.; Banfi, S. miRNeye: a microRNA expression atlas of the mouse eye. BMC Genomics 2010, 11, 715. [Google Scholar] [CrossRef]
- Zong, R.; Zhou, T.; Lin, Z.; Bao, X.; Xiu, Y.; Chen, Y.; Chen, L.; Ma, J.X.; Liu, Z.; Zhou, Y. Down-Regulation of MicroRNA-184 Is Associated With Corneal Neovascularization. Invest Ophthalmol Vis Sci 2016, 57, 1398–1407. [Google Scholar] [CrossRef]
- Zhang, X.; Di, G.; Dong, M.; Qu, M.; Zhao, X.; Duan, H.; Hu, X.; Liu, T.; Zhou, Q.; Shi, W. Epithelium-derived miR-204 inhibits corneal neovascularization. Exp Eye Res 2018, 167, 122–127. [Google Scholar] [CrossRef]
- Qian, J.; Yu, J.; Zhu, X.; Liang, S. MiR-335 promotes corneal neovascularization by Targeting EGFR. BMC Ophthalmol 2022, 22, 267. [Google Scholar] [CrossRef]
- Bai, Y.; Wang, W.; Sun, G.; Zhang, M.; Dong, J. Curcumin inhibits angiogenesis by up-regulation of microRNA-1275 and microRNA-1246: a promising therapy for treatment of corneal neovascularization. Cell Prolif 2016, 49, 751–762. [Google Scholar] [CrossRef]
- Adamis, A.P.; Filatov, V.; Tripathi, B.J.; Tripathi, R.C. Fuchs' endothelial dystrophy of the cornea. Surv Ophthalmol 1993, 38, 149–168. [Google Scholar] [CrossRef]
- Mimura, T.; Yamagami, S.; Amano, S. Corneal endothelial regeneration and tissue engineering. Prog Retin Eye Res 2013, 35, 1–17. [Google Scholar] [CrossRef]
- Pan, P.; Weisenberger, D.J.; Zheng, S.; Wolf, M.; Hwang, D.G.; Rose-Nussbaumer, J.R.; Jurkunas, U.V.; Chan, M.F. Aberrant DNA methylation of miRNAs in Fuchs endothelial corneal dystrophy. Sci Rep 2019, 9, 16385. [Google Scholar] [CrossRef] [PubMed]
- Matthaei, M.; Hu, J.; Kallay, L.; Eberhart, C.G.; Cursiefen, C.; Qian, J.; Lackner, E.M.; Jun, A.S. Endothelial cell microRNA expression in human late-onset Fuchs' dystrophy. Invest Ophthalmol Vis Sci 2014, 55, 216–225. [Google Scholar] [CrossRef]
- Toyono, T.; Usui, T.; Villarreal, G., Jr.; Kallay, L.; Matthaei, M.; Vianna, L.M.; Zhu, A.Y.; Kuroda, M.; Amano, S.; Jun, A.S. MicroRNA-29b Overexpression Decreases Extracellular Matrix mRNA and Protein Production in Human Corneal Endothelial Cells. Cornea 2016, 35, 1466–1470. [Google Scholar] [CrossRef] [PubMed]
- Kaye, S.; Choudhary, A. Herpes simplex keratitis. Prog Retin Eye Res 2006, 25, 355–380. [Google Scholar] [CrossRef]
- Azher, T.N.; Yin, X.T.; Tajfirouz, D.; Huang, A.J.; Stuart, P.M. Herpes simplex keratitis: challenges in diagnosis and clinical management. Clin Ophthalmol 2017, 11, 185–191. [Google Scholar] [CrossRef]
- Kim, Y.J.; Yeon, Y.; Lee, W.J.; Shin, Y.U.; Cho, H.; Lim, H.W.; Kang, M.H. Analysis of MicroRNA Expression in Tears of Patients with Herpes Epithelial Keratitis: A Preliminary Study. Invest Ophthalmol Vis Sci 2022, 63, 21. [Google Scholar] [CrossRef]
- Sharma, P.; Naqvi, R.A.; Borase, H.; Kapoor, D.; Valverde, A.; Capistrano, K.; Yadavalli, T.; Naqvi, A.R.; Shukla, D. Global MicroRNA Profiling of HSV-1 Infected Cornea Identifies miR-329 as a Novel Regulator of Virus Infection. Invest Ophthalmol Vis Sci 2025, 66, 61. [Google Scholar] [CrossRef]
- Cui, Y.H.; Liu, C.Q.; Song, X.L.; Yi, W.Z.; Liu, Q.; Liu, J.M.; Wu, Y.N.; Chen, J.Y.; Yang, L.J.; He, H.Y.; et al. Integrative Analysis of miRNA and circRNA Expression Profiles and Interaction Network in HSV-1-Infected Primary Corneal Epithelial Cells. Curr Eye Res 2024, 49, 368–379. [Google Scholar] [CrossRef] [PubMed]
- Bhela, S.; Mulik, S.; Gimenez, F.; Reddy, P.B.; Richardson, R.L.; Varanasi, S.K.; Jaggi, U.; Xu, J.; Lu, P.Y.; Rouse, B.T. Role of miR-155 in the pathogenesis of herpetic stromal keratitis. Am J Pathol 2015, 185, 1073–1084. [Google Scholar] [CrossRef] [PubMed]
- Mulik, S.; Xu, J.; Reddy, P.B.; Rajasagi, N.K.; Gimenez, F.; Sharma, S.; Lu, P.Y.; Rouse, B.T. Role of miR-132 in angiogenesis after ocular infection with herpes simplex virus. Am J Pathol 2012, 181, 525–534. [Google Scholar] [CrossRef]
- Thomas, P.A. Fungal infections of the cornea. Eye (Lond) 2003, 17, 852–862. [Google Scholar] [CrossRef]
- Boomiraj, H.; Mohankumar, V.; Lalitha, P.; Devarajan, B. Human Corneal MicroRNA Expression Profile in Fungal Keratitis. Invest Ophthalmol Vis Sci 2015, 56, 7939–7946. [Google Scholar] [CrossRef]
- Vemuganti, G.K.; Garg, P.; Gopinathan, U.; Naduvilath, T.J.; John, R.K.; Buddi, R.; Rao, G.N. Evaluation of agent and host factors in progression of mycotic keratitis: A histologic and microbiologic study of 167 corneal buttons. Ophthalmology 2002, 109, 1538–1546. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Lin, Y.; Huang, L.; Hu, J. MiR-223-3p Regulates Autophagy and Inflammation by Targeting ATG16L1 in Fusarium solani-Induced Keratitis. Invest Ophthalmol Vis Sci 2022, 63, 41. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Lin, Y.; Hu, J. Inhibition of miR-665-3p Enhances Autophagy and Alleviates Inflammation in Fusarium solani-Induced Keratitis. Invest Ophthalmol Vis Sci 2021, 62, 24. [Google Scholar] [CrossRef] [PubMed]
- Hazlett, L.D. Corneal response to Pseudomonas aeruginosa infection. Prog Retin Eye Res 2004, 23, 1–30. [Google Scholar] [CrossRef]
- Wang, W.; Zhou, R.; Wu, Y.; Liu, Y.; Su, W.; Xiong, W.; Zeng, Z. PVT1 Promotes Cancer Progression via MicroRNAs. Front Oncol 2019, 9, 609. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhong, J.L.; Hou, N.; Sun, Y.; Ma, B.; Nisar, M.F.; Teng, Y.; Tan, Z.; Chen, K.; Wang, Y.; et al. MicroRNA Let-7b inhibits keratinocyte migration in cutaneous wound healing by targeting IGF2BP2. Exp Dermatol 2017, 26, 116–123. [Google Scholar] [CrossRef]
- Li, X.; Sun, M.; Long, Y. Cyanidin-3-O-Glucoside Attenuates Lipopolysaccharide-Induced Inflammation in Human Corneal Epithelial Cells by Inducing Let-7b-5p-Mediated HMGA2/PI3K/Akt Pathway. Inflammation 2020, 43, 1088–1096. [Google Scholar] [CrossRef]
- Mun, J.; Tam, C.; Chan, G.; Kim, J.H.; Evans, D.; Fleiszig, S. MicroRNA-762 is upregulated in human corneal epithelial cells in response to tear fluid and Pseudomonas aeruginosa antigens and negatively regulates the expression of host defense genes encoding RNase7 and ST2. PLoS One 2013, 8, e57850. [Google Scholar] [CrossRef]
- McClellan, S.; Pitchaikannu, A.; Wright, R.; Bessert, D.; Iulianelli, M.; Hazlett, L.D.; Xu, S. Prophylactic Knockdown of the miR-183/96/182 Cluster Ameliorates Pseudomonas aeruginosa-Induced Keratitis. Invest Ophthalmol Vis Sci 2021, 62, 14. [Google Scholar] [CrossRef]
- Shahraki, T.; Arabi, A.; Feizi, S. Pterygium: an update on pathophysiology, clinical features, and management. Ther Adv Ophthalmol 2021, 13, 25158414211020152. [Google Scholar] [CrossRef]
- Rezvan, F.; Khabazkhoob, M.; Hooshmand, E.; Yekta, A.; Saatchi, M.; Hashemi, H. Prevalence and risk factors of pterygium: a systematic review and meta-analysis. Surv Ophthalmol 2018, 63, 719–735. [Google Scholar] [CrossRef]
- Neale, R.E.; Lucas, R.M.; Byrne, S.N.; Hollestein, L.; Rhodes, L.E.; Yazar, S.; Young, A.R.; Berwick, M.; Ireland, R.A.; Olsen, C.M. The effects of exposure to solar radiation on human health. Photochem Photobiol Sci 2023, 22, 1011–1047. [Google Scholar] [CrossRef]
- Engelsvold, D.H.; Utheim, T.P.; Olstad, O.K.; Gonzalez, P.; Eidet, J.R.; Lyberg, T.; Trøseid, A.M.; Dartt, D.A.; Raeder, S. miRNA and mRNA expression profiling identifies members of the miR-200 family as potential regulators of epithelial-mesenchymal transition in pterygium. Exp Eye Res 2013, 115, 189–198. [Google Scholar] [CrossRef]
- Wu, C.W.; Peng, M.L.; Yeh, K.T.; Tsai, Y.Y.; Chiang, C.C.; Cheng, Y.W. Inactivation of p53 in pterygium influence miR-200a expression resulting in ZEB1/ZEB2 up-regulation and EMT processing. Exp Eye Res 2016, 146, 206–211. [Google Scholar] [CrossRef] [PubMed]
- Kato, N.; Shimmura, S.; Kawakita, T.; Miyashita, H.; Ogawa, Y.; Yoshida, S.; Higa, K.; Okano, H.; Tsubota, K. Beta-catenin activation and epithelial-mesenchymal transition in the pathogenesis of pterygium. Invest Ophthalmol Vis Sci 2007, 48, 1511–1517. [Google Scholar] [CrossRef]
- Wu, C.W.; Cheng, Y.W.; Hsu, N.Y.; Yeh, K.T.; Tsai, Y.Y.; Chiang, C.C.; Wang, W.R.; Tung, J.N. MiRNA-221 negatively regulated downstream p27Kip1 gene expression involvement in pterygium pathogenesis. Mol Vis 2014, 20, 1048–1056. [Google Scholar] [PubMed]
- İçme, G.; Yilmaz, A.; Dinç, E.; Görür, A.; Fidanci Ş, B.; Tamer, L. Assessment of miR-182, miR-183, miR-184, and miR-221 Expressions in Primary Pterygium and Comparison With the Normal Conjunctiva. Eye Contact Lens 2019, 45, 208–211. [Google Scholar] [CrossRef] [PubMed]
- Lan, W.; Chen, S.; Tong, L. MicroRNA-215 Regulates Fibroblast Function: Insights from a Human Fibrotic Disease. Cell Cycle 2015, 14, 1973–1984. [Google Scholar] [CrossRef]
- Han, S.; Chen, Y.; Gao, Y.; Sun, B.; Kong, Y. MicroRNA-218-5p inhibit the migration and proliferation of pterygium epithelial cells by targeting EGFR via PI3K/Akt/mTOR signaling pathway. Exp Eye Res 2019, 178, 37–45. [Google Scholar] [CrossRef]
- Cui, Y.H.; Li, H.Y.; Gao, Z.X.; Liang, N.; Ma, S.S.; Meng, F.J.; Li, Z.J.; Pan, H.W. Regulation of Apoptosis by miR-122 in Pterygium via Targeting Bcl-w. Invest Ophthalmol Vis Sci 2016, 57, 3723–3730. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Dai, Y.; Xu, J. MiR-21 promotes pterygium cell proliferation through the PTEN/AKT pathway. Mol Vis 2018, 24, 485–494. [Google Scholar] [PubMed]
- Zhong, X.; Tang, J.; Li, H.; Shi, X.; Wu, Y.; Xia, D.; Zhang, H.; Ye, J.; Wu, H. MiR-3175 promotes epithelial-mesenchymal transition by targeting Smad7 in human conjunctiva and pterygium. FEBS Lett 2020, 594, 1207–1217. [Google Scholar] [CrossRef]
- Chien, K.H.; Chen, S.J.; Liu, J.H.; Woung, L.C.; Chen, J.T.; Liang, C.M.; Chiou, S.H.; Tsai, C.Y.; Cheng, C.K.; Hu, C.C.; et al. Correlation of microRNA-145 levels and clinical severity of pterygia. Ocul Surf 2013, 11, 133–138. [Google Scholar] [CrossRef]
- Hwang, Y.H.; Jung, S.A.; Lyu, J.; Kim, Y.Y.; Lee, J.H. Transforming Growth Factor-β1-induced Human Subconjunctival Fibrosis is Mediated by MicroRNA 143/145 Expression. Invest Ophthalmol Vis Sci 2019, 60, 2064–2071. [Google Scholar] [CrossRef]
- Sun, W.; Sheng, Y.; Chen, J.; Xu, D.; Gu, Y. Down-Regulation of miR-146a Expression Induces Allergic Conjunctivitis in Mice by Increasing TSLP Level. Med Sci Monit 2015, 21, 2000–2007. [Google Scholar] [CrossRef]
- Yang, Y.; Yin, X.; Yi, J.; Peng, X. MiR-146a overexpression effectively improves experimental allergic conjunctivitis through regulating CD4(+)CD25(-)T cells. Biomed Pharmacother 2017, 94, 937–943. [Google Scholar] [CrossRef]
- Guo, C.; Liu, J.; Hao, P.; Wang, Y.; Sui, S.; Li, L.; Ying, M.; Han, R.; Wang, L.; Li, X. The Potential Inhibitory Effects of miR-19b on Ocular Inflammation are Mediated Upstream of the JAK/STAT Pathway in a Murine Model of Allergic Conjunctivitis. Invest Ophthalmol Vis Sci 2020, 61, 8. [Google Scholar] [CrossRef]
- Bai, M.; Li, Y.; Li, Y.; Hu, Z. Influence of miR155 on allergic conjunctivitis in mice via regulation of NF-κB signal pathway. Tropical Journal of Pharmaceutical Research 2019, 18. [Google Scholar]
- Zhang, S.; Sun, W. Clinical implications of miR-223 in allergic conjunctivitis and related factors affecting disease recurrence. Clin Immunol 2022, 237, 108966. [Google Scholar] [CrossRef] [PubMed]
- Dalvin, L.A.; Salomão, D.R.; Patel, S.V. Population-based incidence of conjunctival tumours in Olmsted County, Minnesota. Br J Ophthalmol 2018, 102, 1728–1734. [Google Scholar] [CrossRef]
- Cai, J.; Liu, X.; Cheng, J.; Li, Y.; Huang, X.; Li, Y.; Ma, X.; Yu, H.; Liu, H.; Wei, R. MicroRNA-200 is commonly repressed in conjunctival MALT lymphoma, and targets cyclin E2. Graefes Arch Clin Exp Ophthalmol 2012, 250, 523–531. [Google Scholar] [CrossRef]
- Hother, C.; Rasmussen, P.K.; Joshi, T.; Reker, D.; Ralfkiær, U.; Workman, C.T.; Heegaard, S.; Ralfkiær, E.; Grønbæk, K. MicroRNA profiling in ocular adnexal lymphoma: a role for MYC and NFKB1 mediated dysregulation of microRNA expression in aggressive disease. Invest Ophthalmol Vis Sci 2013, 54, 5169–5175. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.R.; Nanji, A.A.; Galor, A.; Karp, C.L. Management of conjunctival malignant melanoma: a review and update. Expert Rev Ophthalmol 2014, 9, 185–204. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, L.H.; Larsen, A.C.; von Buchwald, C.; Drzewiecki, K.T.; Prause, J.U.; Heegaard, S. Mucosal malignant melanoma - a clinical, oncological, pathological and genetic survey. Apmis 2016, 124, 475–486. [Google Scholar] [CrossRef] [PubMed]
- Larsen, A.C.; Dahl, C.; Dahmcke, C.M.; Lade-Keller, J.; Siersma, V.D.; Toft, P.B.; Coupland, S.E.; Prause, J.U.; Guldberg, P.; Heegaard, S. BRAF mutations in conjunctival melanoma: investigation of incidence, clinicopathological features, prognosis and paired premalignant lesions. Acta Ophthalmol 2016, 94, 463–470. [Google Scholar] [CrossRef]
- Larsen, A.C.; Mikkelsen, L.H.; Borup, R.; Kiss, K.; Toft, P.B.; von Buchwald, C.; Coupland, S.E.; Prause, J.U.; Heegaard, S. MicroRNA Expression Profile in Conjunctival Melanoma. Invest Ophthalmol Vis Sci 2016, 57, 4205–4212. [Google Scholar] [CrossRef]
- Mikkelsen, L.H.; Andersen, M.K.; Andreasen, S.; Larsen, A.C.; Tan, Q.; Toft, P.B.; Wadt, K.; Heegaard, S. Global microRNA profiling of metastatic conjunctival melanoma. Melanoma Res 2019, 29, 465–473. [Google Scholar] [CrossRef]
- Arif, A.S.; Aaqil, B.; Siddiqui, A.; Nazneen, Z.; Farooq, U. Corneal Complications And Visual Impairment In Vernal Keratoconjunctivitis Patients. J Ayub Med Coll Abbottabad 2017, 29, 58–60. [Google Scholar]
- Syed, N.H.; Shahidan, W.N.S.; Shatriah, I.; Zunaina, E. MicroRNA Profiling of the Tears of Children With Vernal Keratoconjunctivitis. Front Genet 2022, 13, 847168. [Google Scholar] [CrossRef]
- Jing, L.; Li, H.; Zhang, T.; Lu, J.; Zhong, L. MicroRNA-4530 suppresses cell proliferation and induces apoptosis by targeting RASA1 in human umbilical vein endothelial cells. Mol Med Rep 2019, 19, 3393–3402. [Google Scholar] [CrossRef]
- Pagliari, M.; Munari, F.; Toffoletto, M.; Lonardi, S.; Chemello, F.; Codolo, G.; Millino, C.; Della Bella, C.; Pacchioni, B.; Vermi, W.; et al. Helicobacter pylori Affects the Antigen Presentation Activity of Macrophages Modulating the Expression of the Immune Receptor CD300E through miR-4270. Front Immunol 2017, 8, 1288. [Google Scholar] [CrossRef]
- Ueta, M.; Nishigaki, H.; Komai, S.; Sotozono, C.; Kinoshita, S. Difference in the plasma level of miR-628-3p in atopic dermatitis patients with/without atopic keratoconjunctivitis. Immun Inflamm Dis 2021, 9, 1815–1819. [Google Scholar] [CrossRef]
- Mariotti, S.P.; Pascolini, D.; Rose-Nussbaumer, J. Trachoma: global magnitude of a preventable cause of blindness. Br J Ophthalmol 2009, 93, 563–568. [Google Scholar] [CrossRef]
- Burton, M.J.; Rajak, S.N.; Bauer, J.; Weiss, H.A.; Tolbert, S.B.; Shoo, A.; Habtamu, E.; Manjurano, A.; Emerson, P.M.; Mabey, D.C.; et al. Conjunctival transcriptome in scarring trachoma. Infect Immun 2011, 79, 499–511. [Google Scholar] [CrossRef] [PubMed]
- Derrick, T.; Last, A.R.; Burr, S.E.; Roberts, C.H.; Nabicassa, M.; Cassama, E.; Bailey, R.L.; Mabey, D.C.; Burton, M.J.; Holland, M.J. Inverse relationship between microRNA-155 and -184 expression with increasing conjunctival inflammation during ocular Chlamydia trachomatis infection. BMC Infect Dis 2016, 16, 60. [Google Scholar] [CrossRef] [PubMed]
- Derrick, T.; Roberts, C.; Rajasekhar, M.; Burr, S.E.; Joof, H.; Makalo, P.; Bailey, R.L.; Mabey, D.C.; Burton, M.J.; Holland, M.J. Conjunctival MicroRNA expression in inflammatory trachomatous scarring. PLoS Negl Trop Dis 2013, 7, e2117. [Google Scholar] [CrossRef] [PubMed]
- Derrick, T.; Ramadhani, A.M.; Mtengai, K.; Massae, P.; Burton, M.J.; Holland, M.J. miRNAs that associate with conjunctival inflammation and ocular Chlamydia trachomatis infection do not predict progressive disease. Pathog Dis 2017, 75. [Google Scholar] [CrossRef]
- Eldred, J.A.; Dawes, L.J.; Wormstone, I.M. The lens as a model for fibrotic disease. Philos Trans R Soc Lond B Biol Sci 2011, 366, 1301–1319. [Google Scholar] [CrossRef]
- Wei, C.; Kim, I.K.; Kumar, S.; Jayasinghe, S.; Hong, N.; Castoldi, G.; Catalucci, D.; Jones, W.K.; Gupta, S. NF-κB mediated miR-26a regulation in cardiac fibrosis. J Cell Physiol 2013, 228, 1433–1442. [Google Scholar] [CrossRef] [PubMed]
- Liang, H.; Gu, Y.; Li, T.; Zhang, Y.; Huangfu, L.; Hu, M.; Zhao, D.; Chen, Y.; Liu, S.; Dong, Y.; et al. Integrated analyses identify the involvement of microRNA-26a in epithelial-mesenchymal transition during idiopathic pulmonary fibrosis. Cell Death Dis 2014, 5, e1238. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, W.; Zang, X.; Chen, N.; Liu, T.; Tsonis, P.A.; Huang, Y. MicroRNA-204-5p regulates epithelial-to-mesenchymal transition during human posterior capsule opacification by targeting SMAD4. Invest Ophthalmol Vis Sci 2013, 54, 323–332. [Google Scholar] [CrossRef]
- Dong, N.; Xu, B.; Benya, S.R.; Tang, X. MiRNA-26b inhibits the proliferation, migration, and epithelial-mesenchymal transition of lens epithelial cells. Mol Cell Biochem 2014, 396, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Xiao, W.; Chen, W.; Liu, X.; Wu, M.; Bo, Q.; Luo, Y.; Ye, S.; Cao, Y.; Liu, Y. MicroRNA-26a and -26b inhibit lens fibrosis and cataract by negatively regulating Jagged-1/Notch signaling pathway. Cell Death Differ 2017, 24, 1431–1442. [Google Scholar] [CrossRef]
- Tian, L.; Huang, K.; DuHadaway, J.B.; Prendergast, G.C.; Stambolian, D. Genomic profiling of miRNAs in two human lens cell lines. Curr Eye Res 2010, 35, 812–818. [Google Scholar] [CrossRef]
- Wormstone, I.M. Posterior capsule opacification: a cell biological perspective. Exp Eye Res 2002, 74, 337–347. [Google Scholar] [CrossRef]
- Wormstone, I.M.; Wang, L.; Liu, C.S. Posterior capsule opacification. Exp Eye Res 2009, 88, 257–269. [Google Scholar] [CrossRef]
- Hoffmann, A.; Huang, Y.; Suetsugu-Maki, R.; Ringelberg, C.S.; Tomlinson, C.R.; Del Rio-Tsonis, K.; Tsonis, P.A. Implication of the miR-184 and miR-204 competitive RNA network in control of mouse secondary cataract. Mol Med 2012, 18, 528–538. [Google Scholar] [CrossRef]
- Hong, N.; Huang, B.S.; Tong, J.P. Primary silicone oil tamponade and internal limiting membrane peeling for retinal detachment due to macular hole in highly myopic eyes with chorioretinal atrophy. BMC Ophthalmol 2015, 15, 165. [Google Scholar] [CrossRef] [PubMed]
- Ohno-Matsui, K.; Lai, T.Y.; Lai, C.C.; Cheung, C.M. Updates of pathologic myopia. Prog Retin Eye Res 2016, 52, 156–187. [Google Scholar] [CrossRef]
- Xie, M.; Li, Y.; Wu, J.; Wu, J. Genetic variants in MiR-29a associated with high myopia. Ophthalmic Genet 2016, 37, 456–458. [Google Scholar] [CrossRef]
- Zhu, Y.; Zhang, Y.; Jiang, R.; Zhao, K.; Zhou, J. MicroRNA-29a May Influence Myopia Development by Regulating Collagen I. Curr Eye Res 2022, 47, 468–476. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Li, W.; Zhu, D.; Zhou, J. microRNA profiling in the aqueous humor of highly myopic eyes using next generation sequencing. Exp Eye Res 2020, 195, 108034. [Google Scholar] [CrossRef]
- Liu, S.; Chen, H.; Ma, W.; Zhong, Y.; Liang, Y.; Gu, L.; Lu, X.; Li, J. Non-coding RNAs and related molecules associated with form-deprivation myopia in mice. J Cell Mol Med 2022, 26, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Metlapally, R.; Park, H.N.; Chakraborty, R.; Wang, K.K.; Tan, C.C.; Light, J.G.; Pardue, M.T.; Wildsoet, C.F. Genome-Wide Scleral Micro- and Messenger-RNA Regulation During Myopia Development in the Mouse. Invest Ophthalmol Vis Sci 2016, 57, 6089–6097. [Google Scholar] [CrossRef] [PubMed]
- Tkatchenko, A.V.; Luo, X.; Tkatchenko, T.V.; Vaz, C.; Tanavde, V.M.; Maurer-Stroh, S.; Zauscher, S.; Gonzalez, P.; Young, T.L. Large-Scale microRNA Expression Profiling Identifies Putative Retinal miRNA-mRNA Signaling Pathways Underlying Form-Deprivation Myopia in Mice. PLoS One 2016, 11, e0162541. [Google Scholar] [CrossRef]
- Mei, F.; Wang, J.; Chen, Z.; Yuan, Z. Potentially Important MicroRNAs in Form-Deprivation Myopia Revealed by Bioinformatics Analysis of MicroRNA Profiling. Ophthalmic Res 2017, 57, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.Y.; Hsih, W.H.; Lin, Y.B.; Wen, C.Y.; Chang, T.J. Update in the epidemiology, risk factors, screening, and treatment of diabetic retinopathy. J Diabetes Investig 2021, 12, 1322–1325. [Google Scholar] [CrossRef] [PubMed]
- Ge, Y.; Zhang, R.; Feng, Y.; Lu, J.; Li, H. Mbd2 deficiency alleviates retinal cell apoptosisvia the miR-345-5p/Atf1 axis in high glucoseinjury and streptozotocin-induced diabetic mice. Mol Ther Nucleic Acids 2021, 26, 1201–1214. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Ding, L.; Liu, D.; Zhang, Q.; Zheng, G.; Xia, X.; Xiong, S. Molecular investigation of candidate genes for pyroptosis-induced inflammation in diabetic retinopathy. Front Endocrinol (Lausanne) 2022, 13, 918605. [Google Scholar] [CrossRef]
- Gu, C.; Draga, D.; Zhou, C.; Su, T.; Zou, C.; Gu, Q.; Lahm, T.; Zheng, Z.; Qiu, Q. miR-590-3p Inhibits Pyroptosis in Diabetic Retinopathy by Targeting NLRP1 and Inactivating the NOX4 Signaling Pathway. Invest Ophthalmol Vis Sci 2019, 60, 4215–4223. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Li, X.; Zhang, D.; Han, W. Astragaloside-IV alleviates high glucose-induced ferroptosis in retinal pigment epithelial cells by disrupting the expression of miR-138-5p/Sirt1/Nrf2. Bioengineered 2022, 13, 8240–8254. [Google Scholar] [CrossRef]
- Zeng, Q.; Luo, Y.; Fang, J.; Xu, S.; Hu, Y.H.; Yin, M. Circ_0000615 promotes high glucose-induced human retinal pigment epithelium cell apoptosis, inflammation and oxidative stress via miR-646/YAP1 axis in diabetic retinopathy. Eur J Ophthalmol 2022, 32, 1584–1595. [Google Scholar] [CrossRef]
- Jiang, Z.; Wu, J.; Ma, F.; Jiang, J.; Xu, L.; Du, L.; Huang, W.; Wang, Z.; Jia, Y.; Lu, L.; et al. MicroRNA-200a improves diabetic endothelial dysfunction by targeting KEAP1/NRF2. J Endocrinol 2020, 245, 129–140. [Google Scholar] [CrossRef]
- Hoorzad, P.; Mousavinasab, F.; Tofigh, P.; Kalahroud, E.M.; Aghaei-Zarch, S.M.; Salehi, A.; Fattahi, M.; Le, B.N. Understanding the lncRNA/miRNA-NFκB regulatory network in diabetes mellitus: From function to clinical translation. Diabetes Res Clin Pract 2023, 202, 110804. [Google Scholar] [CrossRef]
- Zhuang, P.; Muraleedharan, C.K.; Xu, S. Intraocular Delivery of miR-146 Inhibits Diabetes-Induced Retinal Functional Defects in Diabetic Rat Model. Invest Ophthalmol Vis Sci 2017, 58, 1646–1655. [Google Scholar] [CrossRef]
- Cui, C.; Li, Y.; Liu, Y. Down-regulation of miR-377 suppresses high glucose and hypoxia-induced angiogenesis and inflammation in human retinal endothelial cells by direct up-regulation of target gene SIRT1. Hum Cell 2019, 32, 260–274. [Google Scholar] [CrossRef] [PubMed]
- Qiu, H.; Liu, X.; Yao, S.; Zhou, J.; Zhang, X.; Du, J. Regulation and Mechanism of miR-518d through the PPARα-Mediated NF-κB Pathway in the Development of Gestational Diabetes Mellitus. J Diabetes Res 2020, 2020, 7019597. [Google Scholar] [CrossRef]
- Zhao, R.; Qian, L.; Jiang, L. miRNA-dependent cross-talk between VEGF and Ang-2 in hypoxia-induced microvascular dysfunction. Biochem Biophys Res Commun 2014, 452, 428–435. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Su, X.; Zhang, Q.Q.; Zhang, Y.Y.; Chu, Z.Y.; Zhang, J.L.; Ren, Q. MicroRNA-93-5p participates in type 2 diabetic retinopathy through targeting Sirt1. Int Ophthalmol 2021, 41, 3837–3848. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yao, Y.; Wang, K.; Li, J.; Chu, T.; Shen, H. MicroRNA-148a-3p alleviates high glucose-induced diabetic retinopathy by targeting TGFB2 and FGF2. Acta Diabetol 2020, 57, 1435–1443. [Google Scholar] [CrossRef]
- Lu, J.M.; Zhang, Z.Z.; Ma, X.; Fang, S.F.; Qin, X.H. Repression of microRNA-21 inhibits retinal vascular endothelial cell growth and angiogenesis via PTEN dependent-PI3K/Akt/VEGF signaling pathway in diabetic retinopathy. Exp Eye Res 2020, 190, 107886. [Google Scholar] [CrossRef]
- Zheng, Y.; Liu, Y.; Wang, L.; Xu, H.; Lu, Z.; Xuan, Y.; Meng, W.; Ye, L.; Fang, D.; Zhou, Y.; et al. MicroRNA-126 suppresses the proliferation and migration of endothelial cells in experimental diabetic retinopathy by targeting polo-like kinase 4. Int J Mol Med 2021, 47, 151–160. [Google Scholar] [CrossRef] [PubMed]
- Schober, A.; Nazari-Jahantigh, M.; Wei, Y.; Bidzhekov, K.; Gremse, F.; Grommes, J.; Megens, R.T.; Heyll, K.; Noels, H.; Hristov, M.; et al. MicroRNA-126-5p promotes endothelial proliferation and limits atherosclerosis by suppressing Dlk1. Nat Med 2014, 20, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Zampetaki, A.; Kiechl, S.; Drozdov, I.; Willeit, P.; Mayr, U.; Prokopi, M.; Mayr, A.; Weger, S.; Oberhollenzer, F.; Bonora, E.; et al. Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ Res 2010, 107, 810–817. [Google Scholar] [CrossRef]
- Jansen, F.; Yang, X.; Hoelscher, M.; Cattelan, A.; Schmitz, T.; Proebsting, S.; Wenzel, D.; Vosen, S.; Franklin, B.S.; Fleischmann, B.K.; et al. Endothelial microparticle-mediated transfer of MicroRNA-126 promotes vascular endothelial cell repair via SPRED1 and is abrogated in glucose-damaged endothelial microparticles. Circulation 2013, 128, 2026–2038. [Google Scholar] [CrossRef]
- Ye, E.A.; Liu, L.; Steinle, J.J. miR-15a/16 inhibits TGF-beta3/VEGF signaling and increases retinal endothelial cell barrier proteins. Vision Res 2017, 139, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Xie, S.C.; Ma, Y.C. Low expression of microRNA-15b promotes the proliferation of retinal capillary endothelial cells and pericytes by up-regulating VEGFA in diabetic rats. Eur Rev Med Pharmacol Sci 2019, 23, 6018–6025. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, Y.; Chen, Q.; Rajala, R.V.S.; Ma, J.X. MicroRNA-184 modulates canonical Wnt signaling through the regulation of frizzled-7 expression in the retina with ischemia-induced neovascularization. FEBS Lett 2015, 589, 1143–1149. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.B.; Moolani, H.V.; Sene, A.; Sidhu, R.; Kell, P.; Lin, J.B.; Dong, Z.; Ban, N.; Ory, D.S.; Apte, R.S. Macrophage microRNA-150 promotes pathological angiogenesis as seen in age-related macular degeneration. JCI Insight 2018, 3. [Google Scholar] [CrossRef] [PubMed]
- Dou, W.; Fu, C.; Jin, L.; Li, T.; Li, M. MiR-155-5p promotes endothelial-mesenchymal transition and oxidative stress in diabetic retinopathy. J Diabetes Investig 2026, 10.1111/jdi.70209. [Google Scholar] [CrossRef]
- Trotta, M.C.; Gesualdo, C.; Platania, C.B.M.; De Robertis, D.; Giordano, M.; Simonelli, F.; D'Amico, M.; Drago, F.; Bucolo, C.; Rossi, S. Circulating miRNAs in diabetic retinopathy patients: Prognostic markers or pharmacological targets? Biochem Pharmacol 2021, 186, 114473. [Google Scholar] [CrossRef] [PubMed]
- Prado, M.S.G.; de Jesus, M.L.; de Goes, T.C.; Mendonça, L.S.O.; Kaneto, C.M. Downregulation of circulating miR-320a and target gene prediction in patients with diabetic retinopathy. BMC Res Notes 2020, 13, 155. [Google Scholar] [CrossRef]
- Gemayel, M.C.; Bhatwadekar, A.D.; Ciulla, T. RNA therapeutics for retinal diseases. Expert Opin Biol Ther 2021, 21, 603–613. [Google Scholar] [CrossRef] [PubMed]
- Oltra, M.; Vidal-Gil, L.; Maisto, R.; Sancho-Pelluz, J.; Barcia, J.M. Oxidative stress-induced angiogenesis is mediated by miR-205-5p. J Cell Mol Med 2020, 24, 1428–1436. [Google Scholar] [CrossRef]
- Jadeja, R.N.; Jones, M.A.; Abdelrahman, A.A.; Powell, F.L.; Thounaojam, M.C.; Gutsaeva, D.; Bartoli, M.; Martin, P.M. Inhibiting microRNA-144 potentiates Nrf2-dependent antioxidant signaling in RPE and protects against oxidative stress-induced outer retinal degeneration. Redox Biol 2020, 28, 101336. [Google Scholar] [CrossRef] [PubMed]
- Marneros, A.G. VEGF-A and the NLRP3 Inflammasome in Age-Related Macular Degeneration. Adv Exp Med Biol 2016, 854, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Penn, J.S.; Madan, A.; Caldwell, R.B.; Bartoli, M.; Caldwell, R.W.; Hartnett, M.E. Vascular endothelial growth factor in eye disease. Prog Retin Eye Res 2008, 27, 331–371. [Google Scholar] [CrossRef] [PubMed]
- Deissler, H.L.; Stutzer, J.N.; Lang, G.K.; Grisanti, S.; Lang, G.E.; Ranjbar, M. VEGF receptor 2 inhibitor nintedanib completely reverts VEGF-A(165)-induced disturbances of barriers formed by retinal endothelial cells or long-term cultivated ARPE-19 cells. Exp Eye Res 2020, 194, 108004. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.X.; Ma, D.Y.; Zhi, X.Y.; Wang, M.W.; Zhao, J.Y.; Qin, Y. MiR-125b attenuates retinal pigment epithelium oxidative damage via targeting Nrf2/HIF-1α signal pathway. Exp Cell Res 2022, 410, 112955. [Google Scholar] [CrossRef] [PubMed]
- Lukiw, W.J.; Surjyadipta, B.; Dua, P.; Alexandrov, P.N. Common micro RNAs (miRNAs) target complement factor H (CFH) regulation in Alzheimer's disease (AD) and in age-related macular degeneration (AMD). Int J Biochem Mol Biol 2012, 3, 105–116. [Google Scholar]
- Sticht, C.; De La Torre, C.; Parveen, A.; Gretz, N. miRWalk: An online resource for prediction of microRNA binding sites. PLoS One 2018, 13, e0206239. [Google Scholar] [CrossRef]
- Yang, L.; Liang, H.; Meng, X.; Shen, L.; Guan, Z.; Hei, B.; Yu, H.; Qi, S.; Wen, X. mmu_circ_0000790 Is Involved in Pulmonary Vascular Remodeling in Mice with HPH via MicroRNA-374c-Mediated FOXC1. Mol Ther Nucleic Acids 2020, 20, 292–307. [Google Scholar] [CrossRef]
- Wu, J.; Chen, J.; Hu, J.; Yao, M.; Zhang, M.; Wan, X.; Jia, H.; Wang, F.; Sun, X. CircRNA Uxs1/miR-335-5p/PGF axis regulates choroidal neovascularization via the mTOR/p70 S6k pathway. Transl Res 2023, 256, 41–55. [Google Scholar] [CrossRef]
- Choi, Y.A.; Jeong, A.; Woo, C.H.; Cha, S.C.; Park, D.Y.; Sagong, M. Aqueous microRNA profiling in age-related macular degeneration and polypoidal choroidal vasculopathy by next-generation sequencing. Sci Rep 2023, 13, 1274. [Google Scholar] [CrossRef]
- Behrman, S.; Acosta-Alvear, D.; Walter, P. A CHOP-regulated microRNA controls rhodopsin expression. J Cell Biol 2011, 192, 919–927. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Frost, R.J.A.; Anderson, C.; Zhao, F.; Ma, J.; Yu, B.; Wang, S. let-7 Contributes to Diabetic Retinopathy but Represses Pathological Ocular Angiogenesis. Mol Cell Biol 2017, 37. [Google Scholar] [CrossRef] [PubMed]
- Ménard, C.; Rezende, F.A.; Miloudi, K.; Wilson, A.; Tétreault, N.; Hardy, P.; SanGiovanni, J.P.; De Guire, V.; Sapieha, P. MicroRNA signatures in vitreous humour and plasma of patients with exudative AMD. Oncotarget 2016, 7, 19171–19184. [Google Scholar] [CrossRef] [PubMed]
- Ertekin, S.; Yıldırım, O.; Dinç, E.; Ayaz, L.; Fidancı, S.B.; Tamer, L. Evaluation of circulating miRNAs in wet age-related macular degeneration. Mol Vis 2014, 20, 1057–1066. [Google Scholar] [PubMed]
- Liu, J.; Ma, Z.; Ran, Z. MiR-21-3p modulates lipopolysaccharide-induced inflammation and apoptosis via targeting TGS4 in retinal pigment epithelial cells. Clin Exp Pharmacol Physiol 2019, 46, 883–889. [Google Scholar] [CrossRef]
- Wang, L.; Lee, A.Y.; Wigg, J.P.; Peshavariya, H.; Liu, P.; Zhang, H. miR-126 Regulation of Angiogenesis in Age-Related Macular Degeneration in CNV Mouse Model. Int J Mol Sci 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Chaikitmongkol, V.; Cheung, C.M.G.; Koizumi, H.; Govindahar, V.; Lai, T.Y.Y. Latest Developments in Polypoidal Choroidal Vasculopathy: Epidemiology, Etiology, Diagnosis, and Treatment. Asia-Pacific Journal of Ophthalmology 2020, 9. [Google Scholar] [CrossRef]
- Tasharrofi, N.; Kouhkan, F.; Soleimani, M.; Soheili, Z.S.; Kabiri, M.; Mahmoudi Saber, M.; Dorkoosh, F.A. Survival Improvement in Human Retinal Pigment Epithelial Cells via Fas Receptor Targeting by miR-374a. J Cell Biochem 2017, 118, 4854–4861. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.; Droppelmann, C.A.; Salsoso, R.; Westermeier, F.; Toledo, F.; Salomon, C.; Sanhueza, C.; Pardo, F.; Leiva, A.; Sobrevia, L. A Hypothesis for the Role of RECK in Angiogenesis. Curr Vasc Pharmacol 2016, 14, 106–115. [Google Scholar] [CrossRef]
- Weinreb, R.N.; Aung, T.; Medeiros, F.A. The pathophysiology and treatment of glaucoma: a review. Jama 2014, 311, 1901–1911. [Google Scholar] [CrossRef] [PubMed]
- Tham, Y.C.; Li, X.; Wong, T.Y.; Quigley, H.A.; Aung, T.; Cheng, C.Y. Global prevalence of glaucoma and projections of glaucoma burden through 2040: a systematic review and meta-analysis. Ophthalmology 2014, 121, 2081–2090. [Google Scholar] [CrossRef] [PubMed]
- Sharoukhov, D.; Bucinca-Cupallari, F.; Lim, H. Microtubule Imaging Reveals Cytoskeletal Deficit Predisposing the Retinal Ganglion Cell Axons to Atrophy in DBA/2J. Invest Ophthalmol Vis Sci 2018, 59, 5292–5300. [Google Scholar] [CrossRef] [PubMed]
- Martinez, B.; Peplow, P.V. MicroRNAs as biomarkers in glaucoma and potential therapeutic targets. Neural Regen Res 2022, 17, 2368–2375. [Google Scholar] [CrossRef] [PubMed]
- Weinreb, R.N.; Khaw, P.T. Primary open-angle glaucoma. Lancet 2004, 363, 1711–1720. [Google Scholar] [CrossRef]
- Xu, B.Y.; Liang, S.; Pardeshi, A.A.; Lifton, J.; Moghimi, S.; Lewinger, J.P.; Varma, R. Differences in Ocular Biometric Measurements among Subtypes of Primary Angle Closure Disease: The Chinese American Eye Study. Ophthalmol Glaucoma 2021, 4, 224–231. [Google Scholar] [CrossRef] [PubMed]
- Lobo, J.; Gillis, A.J.M.; van den Berg, A.; Dorssers, L.C.J.; Belge, G.; Dieckmann, K.P.; Roest, H.P.; van der Laan, L.J.W.; Gietema, J.; Hamilton, R.J.; et al. Identification and Validation Model for Informative Liquid Biopsy-Based microRNA Biomarkers: Insights from Germ Cell Tumor In Vitro, In Vivo and Patient-Derived Data. Cells 2019, 8. [Google Scholar] [CrossRef]
- Callaghan, B.; Lester, K.; Lane, B.; Fan, X.; Goljanek-Whysall, K.; Simpson, D.A.; Sheridan, C.; Willoughby, C.E. Genome-wide transcriptome profiling of human trabecular meshwork cells treated with TGF-β2. Sci Rep 2022, 12, 9564. [Google Scholar] [CrossRef] [PubMed]
- Seong, H.; Cho, H.K.; Kee, C.; Song, D.H.; Cho, M.C.; Kang, S.S. Profiles of microRNA in aqueous humor of normal tension glaucoma patients using RNA sequencing. Sci Rep 2021, 11, 19024. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Tsuda, S.; Kunikata, H.; Sato, J.; Kokubun, T.; Yasuda, M.; Nishiguchi, K.M.; Inada, T.; Nakazawa, T. Profiles of extracellular miRNAs in the aqueous humor of glaucoma patients assessed with a microarray system. Sci Rep 2014, 4, 5089. [Google Scholar] [CrossRef]
- Tabak, S.; Schreiber-Avissar, S.; Beit-Yannai, E. Crosstalk between MicroRNA and Oxidative Stress in Primary Open-Angle Glaucoma. Int J Mol Sci 2021, 22. [Google Scholar] [CrossRef]
- Drewry, M.; Helwa, I.; Allingham, R.R.; Hauser, M.A.; Liu, Y. miRNA Profile in Three Different Normal Human Ocular Tissues by miRNA-Seq. Invest Ophthalmol Vis Sci 2016, 57, 3731–3739. [Google Scholar] [CrossRef]
- Li, X.; Zhao, F.; Xin, M.; Li, G.; Luna, C.; Li, G.; Zhou, Q.; He, Y.; Yu, B.; Olson, E.; et al. Regulation of intraocular pressure by microRNA cluster miR-143/145. Sci Rep 2017, 7, 915. [Google Scholar] [CrossRef]
- Perkumas, K.M.; Hoffman, E.A.; McKay, B.S.; Allingham, R.R.; Stamer, W.D. Myocilin-associated exosomes in human ocular samples. Exp Eye Res 2007, 84, 209–212. [Google Scholar] [CrossRef]
- Cho, H.K.; Seong, H.; Kee, C.; Song, D.H.; Kim, S.J.; Seo, S.W.; Kang, S.S. MicroRNA profiles in aqueous humor between pseudoexfoliation glaucoma and normal tension glaucoma patients in a Korean population. Sci Rep 2022, 12, 6217. [Google Scholar] [CrossRef]
- Izzotti, A.; Ceccaroli, C.; Longobardi, M.G.; Micale, R.T.; Pulliero, A.; La Maestra, S.; Saccà, S.C. Molecular Damage in Glaucoma: from Anterior to Posterior Eye Segment. The MicroRNA Role. Microrna 2015, 4, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Yin, R.; Chen, X. Regulatory effect of miR-144-3p on the function of human trabecular meshwork cells and fibronectin-1. Exp Ther Med 2019, 18, 647–653. [Google Scholar] [CrossRef]
- Liu, H.; Xiu, Y.; Zhang, Q.; Xu, Y.; Wan, Q.; Tao, L. Silencing microRNA-29b-3p expression protects human trabecular meshwork cells against oxidative injury via upregulation of RNF138 to activate the ERK pathway. Int J Mol Med 2021, 47. [Google Scholar] [CrossRef]
- Tan, C.; Song, M.; Stamer, W.D.; Qiao, Y.; Chen, X.; Sun, X.; Lei, Y.; Chen, J. miR-21-5p: A viable therapeutic strategy for regulating intraocular pressure. Exp Eye Res 2020, 200, 108197. [Google Scholar] [CrossRef]
- Luna, C.; Li, G.; Qiu, J.; Epstein, D.L.; Gonzalez, P. MicroRNA-24 regulates the processing of latent TGFβ1 during cyclic mechanical stress in human trabecular meshwork cells through direct targeting of FURIN. J Cell Physiol 2011, 226, 1407–1414. [Google Scholar] [CrossRef]
- Kang, Y.; Jia, P.; Zhao, H.; Hu, C.; Yang, X. MicroRNA-26a overexpression protects RGC-5 cells against H2O2-induced apoptosis. Biochem Biophys Res Commun 2015, 460, 164–169. [Google Scholar] [CrossRef]
- Zhao, J.; Du, X.; Wang, M.; Yang, P.; Zhang, J. Salidroside mitigates hydrogen peroxide-induced injury by enhancement of microRNA-27a in human trabecular meshwork cells. Artif Cells Nanomed Biotechnol 2019, 47, 1758–1765. [Google Scholar] [CrossRef]
- Guo, J.; Liu, H.; Fu, L. MicroRNA-124 ameliorates autophagic dysregulation in glaucoma via regulation of P2X7-mediated Akt/mTOR signaling. Cutan Ocul Toxicol 2022, 41, 43–48. [Google Scholar] [CrossRef]
- Tomczyk-Socha, M.; Baczyńska, D.; Przeździecka-Dołyk, J.; Turno-Kręcicka, A. MicroRNA-125b overexpression in pseudoexfoliation syndrome. Adv Clin Exp Med 2020, 29, 1399–1405. [Google Scholar] [CrossRef]
- Wang, L.J.; Wang, X.Z.; Li, Z.M.; Kou, D.; Zhang, D.; Xu, Z.Z. MiR-126 facilitates apoptosis of retinal ganglion cells in glaucoma rats via VEGF-Notch signaling pathway. Eur Rev Med Pharmacol Sci 2020, 24, 8635–8641. [Google Scholar] [CrossRef]
- Zhang, L.Q.; Cui, H.; Yu, Y.B.; Shi, H.Q.; Zhou, Y.; Liu, M.J. MicroRNA-141-3p inhibits retinal neovascularization and retinal ganglion cell apoptosis in glaucoma mice through the inactivation of Docking protein 5-dependent mitogen-activated protein kinase signaling pathway. J Cell Physiol 2019, 234, 8873–8887. [Google Scholar] [CrossRef]
- Xi, X.; Ma, J.; Chen, Q.; Wang, X.; Xia, Y.; Wen, X.; Yuan, J.; Li, Y. Acteoside attenuates hydrogen peroxide-induced injury of retinal ganglion cells via the CASC2/miR-155/mTOR axis. Ann Transl Med 2022, 10, 5. [Google Scholar] [CrossRef]
- Liu, Y.; Bailey, J.C.; Helwa, I.; Dismuke, W.M.; Cai, J.; Drewry, M.; Brilliant, M.H.; Budenz, D.L.; Christen, W.G.; Chasman, D.I.; et al. A Common Variant in MIR182 Is Associated With Primary Open-Angle Glaucoma in the NEIGHBORHOOD Consortium. Invest Ophthalmol Vis Sci 2016, 57, 4528–4535. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, Y. RETRACTED: Lycium barbarum polysaccharides alleviate hydrogen peroxide-induced injury by up-regulation of miR-4295 in human trabecular meshwork cells. Exp Mol Pathol 2019, 106, 109–115. [Google Scholar] [CrossRef]
- Zou, Y.; Kim, J.; Kim, T.; Sung, S.E.; Han, J.C.; Cha, S.C.; Park, D.Y. A Pilot Study Profiling Aqueous Humor-Derived Exosomal MicroRNA in Primary Open-Angle and Exfoliation Glaucoma. Ophthalmol Sci 2026, 6, 100996. [Google Scholar] [CrossRef]
- Chatzikyriakidou, A.; Founti, P.; Melidou, A.; Minti, F.; Bouras, E.; Anastasopoulos, E.; Pappas, T.; Haidich, A.B.; Lambropoulos, A.; Topouzis, F. MicroRNA-related polymorphisms in pseudoexfoliation syndrome, pseudoexfoliative glaucoma, and primary open-angle glaucoma. Ophthalmic Genet 2018, 39, 603–609. [Google Scholar] [CrossRef]
- Hindle, A.G.; Thoonen, R.; Jasien, J.V.; Grange, R.M.H.; Amin, K.; Wise, J.; Ozaki, M.; Ritch, R.; Malhotra, R.; Buys, E.S. Identification of Candidate miRNA Biomarkers for Glaucoma. Invest Ophthalmol Vis Sci 2019, 60, 134–146. [Google Scholar] [CrossRef]
- Zhao, J.; Sun, H.; Zhang, J.M.; Wang, M.; Du, X.J.; Zhang, J.L. Long non-coding RNA ANRIL down-regulates microRNA-7 to protect human trabecular meshwork cells in an experimental model for glaucoma. Eur Rev Med Pharmacol Sci 2019, 23, 3173–3182. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, S.; Wang, J.; Liu, Y.; Chen, Y.; Wen, T.; Fang, X.; Vidal-Sanz, M.; Jonas, J.B.; Zhang, X. MicroRNA-93/STAT3 signalling pathway mediates retinal microglial activation and protects retinal ganglion cells in an acute ocular hypertension model. Cell Death Dis 2021, 12, 41. [Google Scholar] [CrossRef]
- Kong, N.; Lu, X.; Li, B. Downregulation of microRNA-100 protects apoptosis and promotes neuronal growth in retinal ganglion cells. BMC Mol Biol 2014, 15, 25. [Google Scholar] [CrossRef]
- Zhang, Q.L.; Wang, W.; Li, J.; Tian, S.Y.; Zhang, T.Z. Decreased miR-187 induces retinal ganglion cell apoptosis through upregulating SMAD7 in glaucoma. Biomed Pharmacother 2015, 75, 19–25. [Google Scholar] [CrossRef]
- Zhang, Q.L.; Wang, W.; Alatantuya; Dongmei; Lu, Z.J.; Li, L.L.; Zhang, T.Z. Down-regulated miR-187 promotes oxidative stress-induced retinal cell apoptosis through P2X7 receptor. Int J Biol Macromol 2018, 120, 801–810. [Google Scholar] [CrossRef]
- Ghanbari, M.; Iglesias, A.I.; Springelkamp, H.; van Duijn, C.M.; Ikram, M.A.; Dehghan, A.; Erkeland, S.J.; Klaver, C.C.W.; Meester-Smoor, M.A. A Genome-Wide Scan for MicroRNA-Related Genetic Variants Associated With Primary Open-Angle Glaucoma. Invest Ophthalmol Vis Sci 2017, 58, 5368–5377. [Google Scholar] [CrossRef]
- Han, R.; Zhong, H.; Zhang, Y.; Yu, H.; Zhang, Y.; Huang, S.; Yang, Z.; Zhong, Y. MiR-146a reduces fibrosis after glaucoma filtration surgery in rats. J Transl Med 2024, 22, 440. [Google Scholar] [CrossRef]
- Sugali, C.K.; Gajula, N.N.; Chava, S.; Reddy, A.B.M. TGF-β2/OPTN/FOXC1/miR-200 axis regulates actin dynamics in human trabecular meshwork cells. BBA Adv 2026, 9, 100177. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, Y.; Chen, Y.; Fang, X.; Wen, T.; Xiao, M.; Chen, S.; Zhang, X. Discovery and Validation of Circulating Hsa-miR-210-3p as a Potential Biomarker for Primary Open-Angle Glaucoma. Invest Ophthalmol Vis Sci 2019, 60, 2925–2934. [Google Scholar] [CrossRef]
- Drewry, M.D.; Challa, P.; Kuchtey, J.G.; Navarro, I.; Helwa, I.; Hu, Y.; Mu, H.; Stamer, W.D.; Kuchtey, R.W.; Liu, Y. Differentially expressed microRNAs in the aqueous humor of patients with exfoliation glaucoma or primary open-angle glaucoma. Hum Mol Genet 2018, 27, 1263–1275. [Google Scholar] [CrossRef]
- Schlötzer-Schrehardt, U.; Naumann, G.O. Ocular and systemic pseudoexfoliation syndrome. Am J Ophthalmol 2006, 141, 921–937. [Google Scholar] [CrossRef]
- Rao, A.; Chakraborty, M.; Roy, A.; Sahay, P.; Pradhan, A.; Raj, N. Differential miRNA Expression: Signature for Glaucoma in Pseudoexfoliation. Clin Ophthalmol 2020, 14, 3025–3038. [Google Scholar] [CrossRef]
- Tomczyk-Socha, M.; Kręcicka, J.; Misiuk-Hojło, M.; Turno-Kręcicka, A. MicroRNA Expression in Pseudoexfoliation Syndrome with the Use of Next-Generation Sequencing. Genes (Basel) 2022, 13. [Google Scholar] [CrossRef]
- Kondkar, A.A.; Azad, T.A.; Sultan, T.; Radhakrishnan, R.; Osman, E.A.; Almobarak, F.A.; Lobo, G.P.; Al-Obeidan, S.A. Polymorphism rs3742330 in microRNA Biogenesis Gene DICER1 Is Associated with Pseudoexfoliation Glaucoma in Saudi Cohort. Genes (Basel) 2022, 13. [Google Scholar] [CrossRef]
- Banasaz, B.; Zamzam, R.; Aghadoost, D.; Golabchi, K.; Morshedi, M.; Bayat, M.; Sadri Nahand, J.; Sheida, A.; Eshraghi, R.; Rahimzadeh, Z.; et al. Evaluation of expression pattern of cellular miRNAs (let-7b, miR-29a, miR-126, miR-34a, miR-181a-5p) and IL-6, TNF-α, and TGF-β in patients with pseudoexfoliation syndrome. Pathol Res Pract 2023, 249, 154721. [Google Scholar] [CrossRef]
- Fujihara, K. Neuromyelitis optica and astrocytic damage in its pathogenesis. J Neurol Sci 2011, 306, 183–187. [Google Scholar] [CrossRef]
- Ratelade, J.; Asavapanumas, N.; Ritchie, A.M.; Wemlinger, S.; Bennett, J.L.; Verkman, A.S. Involvement of antibody-dependent cell-mediated cytotoxicity in inflammatory demyelination in a mouse model of neuromyelitis optica. Acta Neuropathol 2013, 126, 699–709. [Google Scholar] [CrossRef]
- Duan, T.; Smith, A.J.; Verkman, A.S. Complement-independent bystander injury in AQP4-IgG seropositive neuromyelitis optica produced by antibody-dependent cellular cytotoxicity. Acta Neuropathol Commun 2019, 7, 112. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, M.; Cui, C.; Zhao, Y.; Sun, X.; Wang, Y.; Liu, C.; Wu, H.; Zhong, X.; et al. Different Exosomal microRNA Profile in Aquaporin-4 Antibody Positive Neuromyelitis Optica Spectrum Disorders. Front Immunol 2020, 11, 1064. [Google Scholar] [CrossRef]
- Vaknin-Dembinsky, A.; Charbit, H.; Brill, L.; Abramsky, O.; Gur-Wahnon, D.; Ben-Dov, I.Z.; Lavon, I. Circulating microRNAs as biomarkers for rituximab therapy, in neuromyelitis optica (NMO). J Neuroinflammation 2016, 13, 179. [Google Scholar] [CrossRef]
- Keller, A.; Leidinger, P.; Meese, E.; Haas, J.; Backes, C.; Rasche, L.; Behrens, J.R.; Pfuhl, C.; Wakonig, K.; Gieß, R.M.; et al. Next-generation sequencing identifies altered whole blood microRNAs in neuromyelitis optica spectrum disorder which may permit discrimination from multiple sclerosis. J Neuroinflammation 2015, 12, 196. [Google Scholar] [CrossRef]
- Mangoud, N.; Hegazy, M.I.; Estfanous, S.; Ali, S.A. The lncRNA 74.1/miR-324-3p/NRG1 axis as a differential diagnostic and prognostic biomarker in neuromyelitis optica and multiple sclerosis. Biochim Biophys Acta Mol Basis Dis 2025, 1871, 167922. [Google Scholar] [CrossRef]
- Sharaf-Eldin, W.; Kishk, N.; Sakr, B.; El-Hariri, H.; Refeat, M.; ElBagoury, N.; Essawi, M. Potential Value of miR-23a for Discriminating Neuromyelitis Optica Spectrum Disorder from Multiple Sclerosis. Arch Iran Med 2020, 23, 678–687. [Google Scholar] [CrossRef]
- Chen, J.; Zhu, J.; Wang, Z.; Yao, X.; Wu, X.; Liu, F.; Zheng, W.; Li, Z.; Lin, A. MicroRNAs Correlate with Multiple Sclerosis and Neuromyelitis Optica Spectrum Disorder in a Chinese Population. Med Sci Monit 2017, 23, 2565–2583. [Google Scholar] [CrossRef]
- Teekaput, C.; Thiankhaw, K.; Tanprawate, S.; Apaijai, N.; Suparan, K.; Kumfu, S.; Chattipakorn, N.; Chattipakorn, S.C. Alterations in the Expression of Let-7i, miR-21-5p, and miR-30b-5p in Plasma-Derived Extracellular Vesicles as the Possible Prognostic Markers in Central Demyelinating Diseases. Mol Neurobiol 2025, 63, 267. [Google Scholar] [CrossRef]
- Vanderpump, M.P. The epidemiology of thyroid disease. Br Med Bull 2011, 99, 39–51. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Ueland, H.O.; Ueland, G.; Løvås, K.; Breivk, L.E.; Thrane, A.S.; Meling Stokland, A.E.; Rødahl, E.; Husebye, E.S. Novel inflammatory biomarkers in thyroid eye disease. Eur J Endocrinol 2022, 187, 293–300. [Google Scholar] [CrossRef]
- Woeller, C.F.; Roztocil, E.; Hammond, C.; Feldon, S.E. TSHR Signaling Stimulates Proliferation Through PI3K/Akt and Induction of miR-146a and miR-155 in Thyroid Eye Disease Orbital Fibroblasts. Invest Ophthalmol Vis Sci 2019, 60, 4336–4345. [Google Scholar] [CrossRef]
- Wei, H.; Guan, M.; Qin, Y.; Xie, C.; Fu, X.; Gao, F.; Xue, Y. Circulating levels of miR-146a and IL-17 are significantly correlated with the clinical activity of Graves' ophthalmopathy. Endocr J 2014, 61, 1087–1092. [Google Scholar] [CrossRef]
- Maitra, U.; Davis, S.; Reilly, C.M.; Li, L. Differential regulation of Foxp3 and IL-17 expression in CD4 T helper cells by IRAK-1. J Immunol 2009, 182, 5763–5769. [Google Scholar] [CrossRef]
- Martínez-Hernández, R.; Sampedro-Núñez, M.; Serrano-Somavilla, A.; Ramos-Leví, A.M.; de la Fuente, H.; Triviño, J.C.; Sanz-García, A.; Sánchez-Madrid, F.; Marazuela, M. A MicroRNA Signature for Evaluation of Risk and Severity of Autoimmune Thyroid Diseases. J Clin Endocrinol Metab 2018, 103, 1139–1150. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.S.; Lee, H.J.; Choi, S.H.; Chang, E.J.; Lee, S.Y.; Lee, E.J. Quercetin inhibits IL-1β-induced inflammation, hyaluronan production and adipogenesis in orbital fibroblasts from Graves' orbitopathy. PLoS One 2011, 6, e26261. [Google Scholar] [CrossRef]
- Zhang, L.; Masetti, G.; Colucci, G.; Salvi, M.; Covelli, D.; Eckstein, A.; Kaiser, U.; Draman, M.S.; Muller, I.; Ludgate, M.; et al. Combining micro-RNA and protein sequencing to detect robust biomarkers for Graves' disease and orbitopathy. Sci Rep 2018, 8, 8386. [Google Scholar] [CrossRef]
- Yang, W.J.; Ma, P.F.; Li, S.P.; Su, H.; Liu, Y.J. MicroRNA-146a contributes to CD4(+) T lymphocyte differentiation in patients with thyroid ophthalmopathy. Am J Transl Res 2017, 9, 1801–1809. [Google Scholar]
- Hu, Z.J.; He, J.F.; Li, K.J.; Chen, J.; Xie, X.R. Decreased microRNA-146a in CD4+T cells promote ocular inflammation in thyroid-associated ophthalmopathy by targeting NUMB. Eur Rev Med Pharmacol Sci 2017, 21, 1803–1809. [Google Scholar] [PubMed]
- Liberati, A.; Altman, D.G.; Tetzlaff, J.; Mulrow, C.; Gøtzsche, P.C.; Ioannidis, J.P.; Clarke, M.; Devereaux, P.J.; Kleijnen, J.; Moher, D. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate healthcare interventions: explanation and elaboration. Bmj 2009, 339, b2700. [Google Scholar] [CrossRef]
- Sun, J.; Wei, J.; Zhang, Y.; Li, J.; Li, J.; Yan, J.; Guo, M.; Han, J.; Qiao, H. Plasma Exosomes Transfer miR-885-3p Targeting the AKT/NFκB Signaling Pathway to Improve the Sensitivity of Intravenous Glucocorticoid Therapy Against Graves Ophthalmopathy. Front Immunol 2022, 13, 819680. [Google Scholar] [CrossRef]
- Craps, J.; Joris, V.; Baldeschi, L.; Daumerie, C.; Camboni, A.; Buemi, A.; Lengelé, B.; Behets, C.; Boschi, A.; Mourad, M.; et al. miR-199a Downregulation as a Driver of the NOX4/HIF-1α/VEGF-A Pathway in Thyroid and Orbital Adipose Tissues from Graves' Patients. Int J Mol Sci 2021, 23. [Google Scholar] [CrossRef]
- Ma, X.; Becker Buscaglia, L.E.; Barker, J.R.; Li, Y. MicroRNAs in NF-kappaB signaling. J Mol Cell Biol 2011, 3, 159–166. [Google Scholar] [CrossRef]
- Nahid, M.A.; Pauley, K.M.; Satoh, M.; Chan, E.K. miR-146a is critical for endotoxin-induced tolerance: IMPLICATION IN INNATE IMMUNITY. J Biol Chem 2009, 284, 34590–34599. [Google Scholar] [CrossRef]
- de Smet, M.D.; Taylor, S.R.; Bodaghi, B.; Miserocchi, E.; Murray, P.I.; Pleyer, U.; Zierhut, M.; Barisani-Asenbauer, T.; LeHoang, P.; Lightman, S. Understanding uveitis: the impact of research on visual outcomes. Prog Retin Eye Res 2011, 30, 452–470. [Google Scholar] [CrossRef] [PubMed]
- Rao, N.A.; Wu, G.S. Free radical mediated photoreceptor damage in uveitis. Prog Retin Eye Res 2000, 19, 41–68. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wu, L.Y.; Wu, G.S.; Rao, N.A. Differential expression of nitric oxide synthase in experimental uveoretinitis. Invest Ophthalmol Vis Sci 1999, 40, 1899–1905. [Google Scholar]
- Wu, G.S.; Lee, T.D.; Moore, R.E.; Rao, N.A. Photoreceptor mitochondrial tyrosine nitration in experimental uveitis. Invest Ophthalmol Vis Sci 2005, 46, 2271–2281. [Google Scholar] [CrossRef]
- Bowers, C.E.; Calder, V.L.; Greenwood, J.; Eskandarpour, M. Experimental Autoimmune Uveitis: An Intraocular Inflammatory Mouse Model. J Vis Exp 2022, 10.3791/61832. [Google Scholar] [CrossRef]
- Tsitsiou, E.; Lindsay, M.A. microRNAs and the immune response. Curr Opin Pharmacol 2009, 9, 514–520. [Google Scholar] [CrossRef]
- Seto, A.G. The road toward microRNA therapeutics. Int J Biochem Cell Biol 2010, 42, 1298–1305. [Google Scholar] [CrossRef]
- Saraswathy, S.; Rao, N.A. microRNA 146a ameliorates retinal damage in experimental autoimmune uveitis. Front Ophthalmol (Lausanne) 2023, 3, 1130202. [Google Scholar] [CrossRef]
- Carissimi, C.; Fulci, V.; Macino, G. MicroRNAs: novel regulators of immunity. Autoimmun Rev 2009, 8, 520–524. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.F.; Boldin, M.P.; Chaudhry, A.; Lin, L.L.; Taganov, K.D.; Hanada, T.; Yoshimura, A.; Baltimore, D.; Rudensky, A.Y. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 2010, 142, 914–929. [Google Scholar] [CrossRef] [PubMed]
- Louafi, F.; Martinez-Nunez, R.T.; Sanchez-Elsner, T. MicroRNA-155 targets SMAD2 and modulates the response of macrophages to transforming growth factor-{beta}. J Biol Chem 2010, 285, 41328–41336. [Google Scholar] [CrossRef]
- Hsu, Y.R.; Chang, S.W.; Lin, Y.C.; Yang, C.H. MicroRNA-146a Alleviates Experimental Autoimmune Anterior Uveitis in the Eyes of Lewis Rats. Mediators Inflamm 2017, 2017, 9601349. [Google Scholar] [CrossRef] [PubMed]
- Rovira, C.; Güida, M.C.; Cayota, A. MicroRNAs and other small silencing RNAs in cancer. IUBMB Life 2010, 62, 859–868. [Google Scholar] [CrossRef]
- Nicolas, F.E.; Pais, H.; Schwach, F.; Lindow, M.; Kauppinen, S.; Moulton, V.; Dalmay, T. Experimental identification of microRNA-140 targets by silencing and overexpressing miR-140. Rna 2008, 14, 2513–2520. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Gao, X.; Liu, K.; Bao, N.; Jiang, Z. MiR-379-5p aggravates experimental autoimmune uveitis in mice via the regulation of SEMA3A. Autoimmunity 2021, 54, 275–283. [Google Scholar] [CrossRef]
- Qu, R.; Peng, Y.; Zhou, M.; Xu, S.; Yin, X.; Qiu, Y.; Liu, B.; Gao, Y.; Bi, H.; Guo, D. MiR-223-3p attenuates M1 macrophage polarization via suppressing the Notch signaling pathway and NLRP3-mediated pyroptosis in experimental autoimmune uveitis. Eur J Pharmacol 2023, 960, 176139. [Google Scholar] [CrossRef]
- Liu, B.; Yin, X.; Wei, H.; Zhang, X.; Peng, Y.; Bi, H.; Guo, D. MiR-30b-5p ameliorates experimental autoimmune uveitis by inhibiting the Atg5/Atg12/Becn1 Axis. Int Immunopharmacol 2025, 151, 114370. [Google Scholar] [CrossRef]
- Yin, X.; Wei, H.; Guo, L.; Liu, B.; Peng, Y.; Zhou, M.; Qiu, Y.; Qu, R.; Gao, Y.; Wu, Q.; et al. Therapeutic effect of miR-30b-5p-loaded lentivirus on experimental autoimmune uveitis via inhibiting Notch signaling activation. J Transl Med 2025, 23, 426. [Google Scholar] [CrossRef]
- Zhang, L.; Sun, P.; Zhang, Y.; Xu, Y.; Sun, Y. miR-182-5p inhibits the pathogenic Th17 response in experimental autoimmune uveitis mice via suppressing TAF15. Biochem Biophys Res Commun 2020, 529, 784–792. [Google Scholar] [CrossRef] [PubMed]
- Craig, J.P.; Nichols, K.K.; Akpek, E.K.; Caffery, B.; Dua, H.S.; Joo, C.K.; Liu, Z.; Nelson, J.D.; Nichols, J.J.; Tsubota, K.; et al. TFOS DEWS II Definition and Classification Report. Ocul Surf 2017, 15, 276–283. [Google Scholar] [CrossRef]
- Wolffsohn, J.S.; Arita, R.; Chalmers, R.; Djalilian, A.; Dogru, M.; Dumbleton, K.; Gupta, P.K.; Karpecki, P.; Lazreg, S.; Pult, H.; et al. TFOS DEWS II Diagnostic Methodology report. Ocul Surf 2017, 15, 539–574. [Google Scholar] [CrossRef]
- Pucker, A.D.; Ngo, W.; Postnikoff, C.K.; Fortinberry, H.; Nichols, J.J. Tear Film miRNAs and Their Association With Human Dry Eye Disease. Curr Eye Res 2022, 47, 1479–1487. [Google Scholar] [CrossRef]
- Kim, Y.J.; Yeon, Y.; Lee, W.J.; Shin, Y.U.; Cho, H.; Sung, Y.K.; Kim, D.R.; Lim, H.W.; Kang, M.H. Comparison of MicroRNA Expression in Tears of Normal Subjects and Sjögren Syndrome Patients. Invest Ophthalmol Vis Sci 2019, 60, 4889–4895. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Fan, G.C. Extracellular/circulating microRNAs and their potential role in cardiovascular disease. Am J Cardiovasc Dis 2011, 1, 138–149. [Google Scholar] [PubMed]
- Jung, J.H.; Ji, Y.W.; Hwang, H.S.; Oh, J.W.; Kim, H.C.; Lee, H.K.; Kim, K.P. Proteomic analysis of human lacrimal and tear fluid in dry eye disease. Sci Rep 2017, 7, 13363. [Google Scholar] [CrossRef] [PubMed]
- Gopalan, V.; Pillai, S.; Ebrahimi, F.; Salajegheh, A.; Lam, T.C.; Le, T.K.; Langsford, N.; Ho, Y.H.; Smith, R.A.; Lam, A.K. Regulation of microRNA-1288 in colorectal cancer: altered expression and its clinicopathological significance. Mol Carcinog 2014, 53 Suppl 1, E36-44. [Google Scholar] [CrossRef]
- Zhao, G.; Zhang, J.G.; Shi, Y.; Qin, Q.; Liu, Y.; Wang, B.; Tian, K.; Deng, S.C.; Li, X.; Zhu, S.; et al. MiR-130b is a prognostic marker and inhibits cell proliferation and invasion in pancreatic cancer through targeting STAT3. PLoS One 2013, 8, e73803. [Google Scholar] [CrossRef]
- Krishnan, K.; Steptoe, A.L.; Martin, H.C.; Pattabiraman, D.R.; Nones, K.; Waddell, N.; Mariasegaram, M.; Simpson, P.T.; Lakhani, S.R.; Vlassov, A.; et al. miR-139-5p is a regulator of metastatic pathways in breast cancer. Rna 2013, 19, 1767–1780. [Google Scholar] [CrossRef]
- Xu, D.; Takeshita, F.; Hino, Y.; Fukunaga, S.; Kudo, Y.; Tamaki, A.; Matsunaga, J.; Takahashi, R.U.; Takata, T.; Shimamoto, A.; et al. miR-22 represses cancer progression by inducing cellular senescence. J Cell Biol 2011, 193, 409–424. [Google Scholar] [CrossRef]
- Wang, C.; Wang, X.; Liang, H.; Wang, T.; Yan, X.; Cao, M.; Wang, N.; Zhang, S.; Zen, K.; Zhang, C.; et al. miR-203 inhibits cell proliferation and migration of lung cancer cells by targeting PKCα. PLoS One 2013, 8, e73985. [Google Scholar] [CrossRef]
- Jin, Y.; Jiang, Z.; Guan, X.; Chen, Y.; Tang, Q.; Wang, G.; Wang, X. miR-450b-5p Suppresses Stemness and the Development of Chemoresistance by Targeting SOX2 in Colorectal Cancer. DNA Cell Biol 2016, 35, 249–256. [Google Scholar] [CrossRef]
- Ayaz, L.; Dinç, E. Evaluation of microRNA responses in ARPE-19 cells against the oxidative stress. Cutan Ocul Toxicol 2018, 37, 121–126. [Google Scholar] [CrossRef]
- Seen, S.; Tong, L. Dry eye disease and oxidative stress. Acta Ophthalmol 2018, 96, e412–e420. [Google Scholar] [CrossRef]
- Ying, H.; Kang, Y.; Zhang, H.; Zhao, D.; Xia, J.; Lu, Z.; Wang, H.; Xu, F.; Shi, L. MiR-127 modulates macrophage polarization and promotes lung inflammation and injury by activating the JNK pathway. J Immunol 2015, 194, 1239–1251. [Google Scholar] [CrossRef]
- Nakagawa, A.; Nakajima, T.; Azuma, M. Tear miRNA expression analysis reveals miR-203 as a potential regulator of corneal epithelial cells. BMC Ophthalmol 2021, 21, 377. [Google Scholar] [CrossRef]
- Liao, C.H.; Tseng, C.L.; Lin, S.L.; Liang, C.L.; Juo, S.H. MicroRNA Therapy for Dry Eye Disease. J Ocul Pharmacol Ther 2022, 38, 125–132. [Google Scholar] [CrossRef]
- Han, R.; Gao, J.; Wang, L.; Hao, P.; Chen, X.; Wang, Y.; Jiang, Z.; Jiang, L.; Wang, T.; Zhu, L.; et al. MicroRNA-146a negatively regulates inflammation via the IRAK1/TRAF6/NF-κB signaling pathway in dry eye. Sci Rep 2023, 13, 11192. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Zhu, Y.; Liu, Y.; Yang, M.; Zeng, L. Mesenchymal Stem Cells-Derived Exosomal miR-223-3p Alleviates Ocular Surface Damage and Inflammation by Downregulating Fbxw7 in Dry Eye Models. Invest Ophthalmol Vis Sci 2024, 65, 1. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, X.; Chen, Q.; Wei, Z.; Xu, X.; Han, D.; Zhang, Y.; Chen, Z.; Liang, Q. MicroRNAs of extracellular vesicles derived from mesenchymal stromal cells alleviate inflammation in dry eye disease by targeting the IRAK1/TAB2/NF-κB pathway. Ocul Surf 2023, 28, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Ji, H.; Zhang, W.; He, A.; Guo, C.; Ma, L.; Liu, Y. miR-214-3p inhibits LPS-induced macrophage inflammation and attenuates the progression of dry eye syndrome by regulating ferroptosis in cells. Genes Genomics 2025, 47, 183–195. [Google Scholar] [CrossRef]
- Patel, R.; Shahane, A. The epidemiology of Sjögren's syndrome. Clin Epidemiol 2014, 6, 247–255. [Google Scholar] [CrossRef]
- Cha, S.; Mona, M.; Lee, K.E.; Kim, D.H.; Han, K. MicroRNAs in Autoimmune Sjögren's Syndrome. Genomics Inform 2018, 16, e19. [Google Scholar] [CrossRef]
- Alevizos, I.; Alexander, S.; Turner, R.J.; Illei, G.G. MicroRNA expression profiles as biomarkers of minor salivary gland inflammation and dysfunction in Sjögren's syndrome. Arthritis Rheum 2011, 63, 535–544. [Google Scholar] [CrossRef]
- Chen, C.Z.; Li, L.; Lodish, H.F.; Bartel, D.P. MicroRNAs modulate hematopoietic lineage differentiation. Science 2004, 303, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Cortes-Troncoso, J.; Jang, S.I.; Perez, P.; Hidalgo, J.; Ikeuchi, T.; Greenwell-Wild, T.; Warner, B.M.; Moutsopoulos, N.M.; Alevizos, I. T cell exosome-derived miR-142-3p impairs glandular cell function in Sjögren's syndrome. JCI Insight 2020, 5. [Google Scholar] [CrossRef] [PubMed]
- Taheri, F.; Ebrahimi, S.O.; Shareef, S.; Reiisi, S. Regulatory and immunomodulatory role of miR-34a in T cell immunity. Life Sci 2020, 262, 118209. [Google Scholar] [CrossRef] [PubMed]
- Hart, M.; Walch-Rückheim, B.; Friedmann, K.S.; Rheinheimer, S.; Tänzer, T.; Glombitza, B.; Sester, M.; Lenhof, H.P.; Hoth, M.; Schwarz, E.C.; et al. miR-34a: a new player in the regulation of T cell function by modulation of NF-κB signaling. Cell Death Dis 2019, 10, 46. [Google Scholar] [CrossRef] [PubMed]
- Youinou, P.; Pennec, Y.L.; Blaschek, M.A.; Gentric, A.; Jouquan, J.; Lamour, A.; Angelidis, P. Activation of peripheral blood lymphocytes in patients with primary Sjögren's syndrome. Rheumatol Int 1988, 8, 125–130. [Google Scholar] [CrossRef]
- Zhou, H.; Yang, J.; Tian, J.; Wang, S. CD8(+) T Lymphocytes: Crucial Players in Sjögren's Syndrome. Front Immunol 2020, 11, 602823. [Google Scholar] [CrossRef]
- Yan, T.; Shen, J.; Chen, J.; Zhao, M.; Guo, H.; Wang, Y. Differential expression of miR-17-92 cluster among varying histological stages of minor salivary gland in patients with primary Sjögren's syndrome. Clin Exp Rheumatol 2019, 37 Suppl 118, 49–54. [Google Scholar]
- Xiao, C.; Srinivasan, L.; Calado, D.P.; Patterson, H.C.; Zhang, B.; Wang, J.; Henderson, J.M.; Kutok, J.L.; Rajewsky, K. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat Immunol 2008, 9, 405–414. [Google Scholar] [CrossRef]
- Wang-Renault, S.F.; Boudaoud, S.; Nocturne, G.; Roche, E.; Sigrist, N.; Daviaud, C.; Bugge Tinggaard, A.; Renault, V.; Deleuze, J.F.; Mariette, X.; et al. Deregulation of microRNA expression in purified T and B lymphocytes from patients with primary Sjögren's syndrome. Ann Rheum Dis 2018, 77, 133–140. [Google Scholar] [CrossRef]
- Mackay, F.; Woodcock, S.A.; Lawton, P.; Ambrose, C.; Baetscher, M.; Schneider, P.; Tschopp, J.; Browning, J.L. Mice transgenic for BAFF develop lymphocytic disorders along with autoimmune manifestations. J Exp Med 1999, 190, 1697–1710. [Google Scholar] [CrossRef]
- Zhang, Y.; Che, D.; Cao, Y.; Yue, Y.; He, T.; Zhu, Y.; Zhou, J. MicroRNA Profiling in the Aqueous Humor of Keratoconus Eyes. Transl Vis Sci Technol 2022, 11, 5. [Google Scholar] [CrossRef]
- Zhang, R.; Garrett, Q.; Zhou, H.; Wu, X.; Mao, Y.; Cui, X.; Xie, B.; Liu, Z.; Cui, D.; Jiang, L.; et al. Upregulation of miR-195 accelerates oxidative stress-induced retinal endothelial cell injury by targeting mitofusin 2 in diabetic rats. Mol Cell Endocrinol 2017, 452, 33–43. [Google Scholar] [CrossRef]
- Liu, X.; Gong, Q.; Yang, L.; Liu, M.; Niu, L.; Wang, L. microRNA-199a-5p regulates epithelial-to-mesenchymal transition in diabetic cataract by targeting SP1 gene. Mol Med 2020, 26, 122. [Google Scholar] [CrossRef] [PubMed]
- Ju, B.; Nie, Y.; Yang, X.; Wang, X.; Li, F.; Wang, M.; Wang, C.; Zhang, H. miR-193a/b-3p relieves hepatic fibrosis and restrains proliferation and activation of hepatic stellate cells. J Cell Mol Med 2019, 23, 3824–3832. [Google Scholar] [CrossRef] [PubMed]
- Tian, R.; Wang, L.; Zou, H.; Song, M.; Liu, L.; Zhang, H. Role of the XIST-miR-181a-COL4A1 axis in the development and progression of keratoconus. Mol Vis 2020, 26, 1–13. [Google Scholar]
- Wang, Y.M.; Ng, T.K.; Choy, K.W.; Wong, H.K.; Chu, W.K.; Pang, C.P.; Jhanji, V. Histological and microRNA Signatures of Corneal Epithelium in Keratoconus. J Refract Surg 2018, 34, 201–211. [Google Scholar] [CrossRef]
- Stunf Pukl, S. Are miRNAs Dynamic Biomarkers in Keratoconus? A Review of the Literature. Genes (Basel) 2022, 13. [Google Scholar] [CrossRef]
- Abu-Amero, K.K.; Helwa, I.; Al-Muammar, A.; Strickland, S.; Hauser, M.A.; Allingham, R.R.; Liu, Y. Screening of the Seed Region of MIR184 in Keratoconus Patients from Saudi Arabia. Biomed Res Int 2015, 2015, 604508. [Google Scholar] [CrossRef]
- Hughes, A.E.; Bradley, D.T.; Campbell, M.; Lechner, J.; Dash, D.P.; Simpson, D.A.; Willoughby, C.E. Mutation altering the miR-184 seed region causes familial keratoconus with cataract. Am J Hum Genet 2011, 89, 628–633. [Google Scholar] [CrossRef]
- Dohlman, C. FAMILIAL CONGENITAL CORNEA GUTTATA IN ASSOCIATION WITH ANTERIOR POLAR CATARACT. Acta Ophthalmologica 1951, 29, 445–473. [Google Scholar] [CrossRef]
- Iliff, B.W.; Riazuddin, S.A.; Gottsch, J.D. A single-base substitution in the seed region of miR-184 causes EDICT syndrome. Invest Ophthalmol Vis Sci 2012, 53, 348–353. [Google Scholar] [CrossRef]
- Bykhovskaya, Y.; Caiado Canedo, A.L.; Wright, K.W.; Rabinowitz, Y.S. C. Ophthalmic Genet 2015, 36, 244–247. [CrossRef] [PubMed]
- Hughes, A.E.; Dash, D.P.; Jackson, A.J.; Frazer, D.G.; Silvestri, G. Familial keratoconus with cataract: linkage to the long arm of chromosome 15 and exclusion of candidate genes. Invest Ophthalmol Vis Sci 2003, 44, 5063–5066. [Google Scholar] [CrossRef]
- Akpek, E.K.; Jun, A.S.; Goodman, D.F.; Green, W.R.; Gottsch, J.D. Clinical and ultrastructural features of a novel hereditary anterior segment dysgenesis. Ophthalmology 2002, 109, 513–519. [Google Scholar] [CrossRef] [PubMed]
- Nelson, L.B.; Spaeth, G.L.; Nowinski, T.S.; Margo, C.E.; Jackson, L. Aniridia. A review. Surv Ophthalmol 1984, 28, 621–642. [Google Scholar] [CrossRef]
- Nishida, K.; Kinoshita, S.; Ohashi, Y.; Kuwayama, Y.; Yamamoto, S. Ocular surface abnormalities in aniridia. Am J Ophthalmol 1995, 120, 368–375. [Google Scholar] [CrossRef] [PubMed]
- Neuhaus, C.; Betz, C.; Bergmann, C.; Bolz, H.J. [Genetics of congenital aniridia]. Ophthalmologe 2014, 111, 1157–1163. [Google Scholar] [CrossRef]
- Ton, C.C.; Hirvonen, H.; Miwa, H.; Weil, M.M.; Monaghan, P.; Jordan, T.; van Heyningen, V.; Hastie, N.D.; Meijers-Heijboer, H.; Drechsler, M.; et al. Positional cloning and characterization of a paired box- and homeobox-containing gene from the aniridia region. Cell 1991, 67, 1059–1074. [Google Scholar] [CrossRef]
- Lee, H.; Khan, R.; O'Keefe, M. Aniridia: current pathology and management. Acta Ophthalmol 2008, 86, 708–715. [Google Scholar] [CrossRef]
- Landsend, E.C.S.; Lagali, N.; Utheim, T.P. Congenital aniridia - A comprehensive review of clinical features and therapeutic approaches. Surv Ophthalmol 2021, 66, 1031–1050. [Google Scholar] [CrossRef]
- Latta, L.; Ludwig, N.; Krammes, L.; Stachon, T.; Fries, F.N.; Mukwaya, A.; Szentmáry, N.; Seitz, B.; Wowra, B.; Kahraman, M.; et al. Abnormal neovascular and proliferative conjunctival phenotype in limbal stem cell deficiency is associated with altered microRNA and gene expression modulated by PAX6 mutational status in congenital aniridia. Ocul Surf 2021, 19, 115–127. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Yang, Y.; Wan, S.; Xu, Y.; Li, J.; Zhang, L.; Guo, W.; Zheng, Y.; Xiang, Y.; Xing, Y. Exploring the Mechanism of the miRNA-145/Paxillin Axis in Cell Metabolism During VEGF-A-Induced Corneal Angiogenesis. Invest Ophthalmol Vis Sci 2021, 62, 25. [Google Scholar] [CrossRef]
- Cao, Q.; Xu, W.; Chen, W.; Peng, D.; Liu, Q.; Dong, J.; Reinach, P.S.; Yan, D. MicroRNA-184 negatively regulates corneal epithelial wound healing via targeting CDC25A, CARM1, and LASP1. Eye Vis (Lond) 2020, 7, 35. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, G.; Wang, Q.; Jiang, R.B. Dexamethasone and MicroRNA-204 Inhibit Corneal Neovascularization. Mil Med 2024, 189, 374–378. [Google Scholar] [CrossRef]
- Lee, S.K.; Teng, Y.; Wong, H.K.; Ng, T.K.; Huang, L.; Lei, P.; Choy, K.W.; Liu, Y.; Zhang, M.; Lam, D.S.; et al. MicroRNA-145 regulates human corneal epithelial differentiation. PLoS One 2011, 6, e21249. [Google Scholar] [CrossRef]
- Nastaranpour, M.; Suiwal, S.; Stachon, T.; Fries, F.N.; Amini, M.; Seitz, B.; Meese, E.; Ludwig, N.; Szentmáry, N. miRNA Expression Profile in Primary Limbal Epithelial Cells of Aniridia Patients. Invest Ophthalmol Vis Sci 2025, 66, 20. [Google Scholar] [CrossRef] [PubMed]
- Yongblah, K.; Alford, S.C.; Ryan, B.C.; Chow, R.L.; Howard, P.L. Protecting Pax6 3' UTR from MicroRNA-7 Partially Restores PAX6 in Islets from an Aniridia Mouse Model. Mol Ther Nucleic Acids 2018, 13, 144–153. [Google Scholar] [CrossRef]
- Rojanaporn, D.; Boontawon, T.; Chareonsirisuthigul, T.; Thanapanpanich, O.; Attaseth, T.; Saengwimol, D.; Anurathapan, U.; Sujirakul, T.; Kaewkhaw, R.; Hongeng, S. Spectrum of germline RB1 mutations and clinical manifestations in retinoblastoma patients from Thailand. Mol Vis 2018, 24, 778–788. [Google Scholar]
- Liao, Y.; Yin, X.; Deng, Y.; Peng, X. MiR-140-5p suppresses retinoblastoma cell growth via inhibiting c-Met/AKT/mTOR pathway. Biosci Rep 2018, 38. [Google Scholar] [CrossRef]
- Liu, F.; Zhang, Q.; Liang, Y. MicroRNA-598 acts as an inhibitor in retinoblastoma through targeting E2F1 and regulating AKT pathway. J Cell Biochem 2020, 121, 2294–2302. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Sun, Y.; Chen, M.; Dai, Y.; Liu, Z. Downregulation of microRNA-320a inhibits proliferation and induces apoptosis of retinoblastoma cells via targeting TUSC3. Exp Ther Med 2020, 20, 9. [Google Scholar] [CrossRef] [PubMed]
- Ahangar Davoodi, N.; Najafi, S.; Naderi Ghale-Noie, Z.; Piranviseh, A.; Mollazadeh, S.; Ahmadi Asouri, S.; Asemi, Z.; Morshedi, M.; Tamehri Zadeh, S.S.; Hamblin, M.R.; et al. Role of non-coding RNAs and exosomal non-coding RNAs in retinoblastoma progression. Front Cell Dev Biol 2022, 10, 1065837. [Google Scholar] [CrossRef]
- Fernandez-Diaz, D.; Rodriguez-Vidal, C.; Silva-Rodríguez, P.; Paniagua, L.; Blanco-Teijeiro, M.J.; Pardo, M.; Piñeiro, A.; Bande, M. Applications of Non-Coding RNAs in Patients With Retinoblastoma. Front Genet 2022, 13, 842509. [Google Scholar] [CrossRef] [PubMed]
- Che, X.; Qian, Y.; Li, D. Suppression of Disheveled-Axin Domain Containing 1 (DIXDC1) by MicroRNA-186 Inhibits the Proliferation and Invasion of Retinoblastoma Cells. J Mol Neurosci 2018, 64, 252–261. [Google Scholar] [CrossRef]
- Khosravi, A.; Alizadeh, S.; Jalili, A.; Shirzad, R.; Saki, N. The impact of Mir-9 regulation in normal and malignant hematopoiesis. Oncol Rev 2018, 12, 348. [Google Scholar] [CrossRef]
- Gao, M.; Cui, Z.; Zhao, D.; Zhang, S.; Cai, Q. MicroRNA-9 inhibits proliferation and progression in retinoblastoma cells by targeting PTEN. Genes & Genomics 2021. [Google Scholar]
- Narayan, D.S.; Wood, J.P.; Chidlow, G.; Casson, R.J. A review of the mechanisms of cone degeneration in retinitis pigmentosa. Acta Ophthalmol 2016, 94, 748–754. [Google Scholar] [CrossRef]
- Napoli, D.; Biagioni, M.; Billeri, F.; Di Marco, B.; Orsini, N.; Novelli, E.; Strettoi, E. Retinal Pigment Epithelium Remodeling in Mouse Models of Retinitis Pigmentosa. Int J Mol Sci 2021, 22. [Google Scholar] [CrossRef]
- Kanow, M.A.; Giarmarco, M.M.; Jankowski, C.S.; Tsantilas, K.; Engel, A.L.; Du, J.; Linton, J.D.; Farnsworth, C.C.; Sloat, S.R.; Rountree, A.; et al. Biochemical adaptations of the retina and retinal pigment epithelium support a metabolic ecosystem in the vertebrate eye. Elife 2017, 6. [Google Scholar] [CrossRef]
- Boulton, M.; Dayhaw-Barker, P. The role of the retinal pigment epithelium: topographical variation and ageing changes. Eye (Lond) 2001, 15, 384–389. [Google Scholar] [CrossRef]
- Karali, M.; Persico, M.; Mutarelli, M.; Carissimo, A.; Pizzo, M.; Singh Marwah, V.; Ambrosio, C.; Pinelli, M.; Carrella, D.; Ferrari, S.; et al. High-resolution analysis of the human retina miRNome reveals isomiR variations and novel microRNAs. Nucleic Acids Res 2016, 44, 1525–1540. [Google Scholar] [CrossRef]
- Yin, J.; Shi, Z.; Wei, W.; Lu, C.; Wei, Y.; Yan, W.; Li, R.; Zhang, J.; You, Y.; Wang, X. MiR-181b suppress glioblastoma multiforme growth through inhibition of SP1-mediated glucose metabolism. Cancer Cell Int 2020, 20, 69. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Zheng, Y.; Huang, C.; Li, Y.; Gu, M.; Wu, Q. Downregulation of miR-181b-5p Inhibits the Viability, Migration, and Glycolysis of Gallbladder Cancer by Upregulating PDHX Under Hypoxia. Front Oncol 2021, 11, 683725. [Google Scholar] [CrossRef] [PubMed]
- Wei, Z.; Cui, L.; Mei, Z.; Liu, M.; Zhang, D. miR-181a mediates metabolic shift in colon cancer cells via the PTEN/AKT pathway. FEBS Lett 2014, 588, 1773–1779. [Google Scholar] [CrossRef]
- Carrella, S.; Di Guida, M.; Brillante, S.; Piccolo, D.; Ciampi, L.; Guadagnino, I.; Garcia Piqueras, J.; Pizzo, M.; Marrocco, E.; Molinari, M.; et al. miR-181a/b downregulation: a mutation-independent therapeutic approach for inherited retinal diseases. EMBO Mol Med 2022, 14, e15941. [Google Scholar] [CrossRef]
- Indrieri, A.; Carrella, S.; Romano, A.; Spaziano, A.; Marrocco, E.; Fernandez-Vizarra, E.; Barbato, S.; Pizzo, M.; Ezhova, Y.; Golia, F.M.; et al. miR-181a/b downregulation exerts a protective action on mitochondrial disease models. EMBO Mol Med 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Punzo, C.; Kornacker, K.; Cepko, C.L. Stimulation of the insulin/mTOR pathway delays cone death in a mouse model of retinitis pigmentosa. Nat Neurosci 2009, 12, 44–52. [Google Scholar] [CrossRef]
- Venkatesh, A.; Ma, S.; Le, Y.Z.; Hall, M.N.; Rüegg, M.A.; Punzo, C. Activated mTORC1 promotes long-term cone survival in retinitis pigmentosa mice. J Clin Invest 2015, 125, 1446–1458. [Google Scholar] [CrossRef]
- Donato, L.; Bramanti, P.; Scimone, C.; Rinaldi, C.; Giorgianni, F.; Beranova-Giorgianni, S.; Koirala, D.; D'Angelo, R.; Sidoti, A. miRNAexpression profile of retinal pigment epithelial cells under oxidative stress conditions. FEBS Open Bio 2018, 8, 219–233. [Google Scholar] [CrossRef]
- Shiels, A. TRPM3_miR-204: a complex locus for eye development and disease. Hum Genomics 2020, 14, 7. [Google Scholar] [CrossRef]
- Conte, I.; Hadfield, K.D.; Barbato, S.; Carrella, S.; Pizzo, M.; Bhat, R.S.; Carissimo, A.; Karali, M.; Porter, L.F.; Urquhart, J.; et al. MiR-204 is responsible for inherited retinal dystrophy associated with ocular coloboma. Proc Natl Acad Sci U S A 2015, 112, E3236-3245. [Google Scholar] [CrossRef]
- Zuzic, M.; Rojo Arias, J.E.; Wohl, S.G.; Busskamp, V. Retinal miRNA Functions in Health and Disease. Genes (Basel) 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Jedlickova, J.; Vajter, M.; Barta, T.; Black, G.C.M.; Perveen, R.; Mares, J.; Fichtl, M.; Kousal, B.; Dudakova, L.; Liskova, P. MIR204. Clin Genet 2023, 104, 418–426. [Google Scholar] [CrossRef] [PubMed]
- Lei, Z.; He-Lin, Z.; Hai-Yan, W.; Wei, J.; Ru, W.; Zhi-Li, C.; Qian-Feng, W. Retinitis pigmentosa with iris coloboma due to miR-204 gene variant in a Chinese family. Mol Genet Genomic Med 2024, 12, e2481. [Google Scholar] [CrossRef]
- Anasagasti, A.; Lara-López, A.; Milla-Navarro, S.; Escudero-Arrarás, L.; Rodríguez-Hidalgo, M.; Zabaleta, N.; González Aseguinolaza, G.; de la Villa, P.; Ruiz-Ederra, J. Inhibition of MicroRNA 6937 Delays Photoreceptor and Vision Loss in a Mouse Model of Retinitis Pigmentosa. Pharmaceutics 2020, 12. [Google Scholar] [CrossRef]
- Xu, X.; Lu, F.; Yu, D.; Wang, Y.; Chen, P.; Hu, W.; Wo, J.; Jia, S.; Liu, S. Cortex Dictamni induces retinitis pigmentosa in zebrafish by inhibiting pde6a post-transcriptional activity via mmu-mir-6240-p3_2. J Ethnopharmacol 2025, 340, 119282. [Google Scholar] [CrossRef]
- Su, H.; Liang, Z.; Weng, S.; Sun, C.; Huang, J.; Zhang, T.; Wang, X.; Wu, S.; Zhang, Z.; Zhang, Y.; et al. miR-9-5p regulates immunometabolic and epigenetic pathways in β-glucan-trained immunity via IDH3α. JCI Insight 2021, 6. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Smith, S.B. A Novel Mechanism of Sigma 1 Receptor Neuroprotection: Modulation of miR-214-3p. Adv Exp Med Biol 2019, 1185, 463–467. [Google Scholar] [CrossRef]
- Wallace, D.C. A new manifestation of Leber's disease and a new explanation for the agency responsible for its unusual pattern of inheritance. Brain 1970, 93, 121–132. [Google Scholar] [CrossRef]
- Bi, R.; Logan, I.; Yao, Y.G. Leber Hereditary Optic Neuropathy: A Mitochondrial Disease Unique in Many Ways. Handb Exp Pharmacol 2017, 240, 309–336. [Google Scholar] [CrossRef]
- Majander, A.; Robson, A.G.; João, C.; Holder, G.E.; Chinnery, P.F.; Moore, A.T.; Votruba, M.; Stockman, A.; Yu-Wai-Man, P. The pattern of retinal ganglion cell dysfunction in Leber hereditary optic neuropathy. Mitochondrion 2017, 36, 138–149. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.P.; Chang, Y.L.; Chiou, G.Y.; Lee, M.S.; Wu, Y.R.; Chen, P.W.; Lin, Y.Y.; Lai, W.Y.; Liu, Y.H.; Hwang, D.K.; et al. Dysregulation of the circRNA_0087207/miR-548c-3p/PLSR1-TGFB2 axis in Leber hereditary optic neuropathy in vitro. J Chin Med Assoc 2024, 87, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lim, D.; Yao, Y.; Dong, C.; Feng, Z. Global Research Trends in Radiotherapy for Gliomas: A Systematic Bibliometric Analysis. World Neurosurg 2022, 161, e355–e362. [Google Scholar] [CrossRef]
- Fu, H.Z.; Wang, M.H.; Ho, Y.S. The most frequently cited adsorption research articles in the Science Citation Index (Expanded). J Colloid Interface Sci 2012, 379, 148–156. [Google Scholar] [CrossRef] [PubMed]
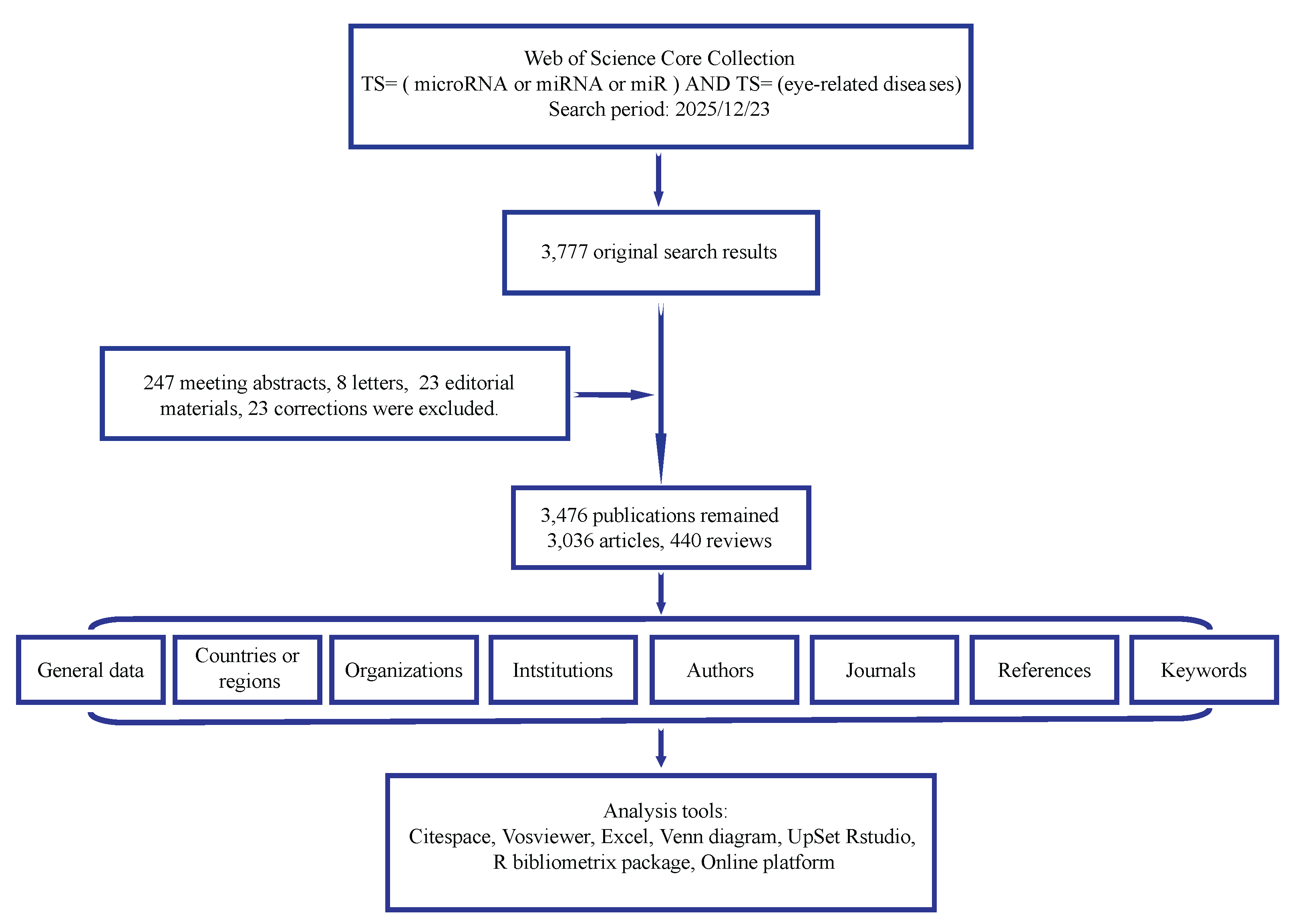
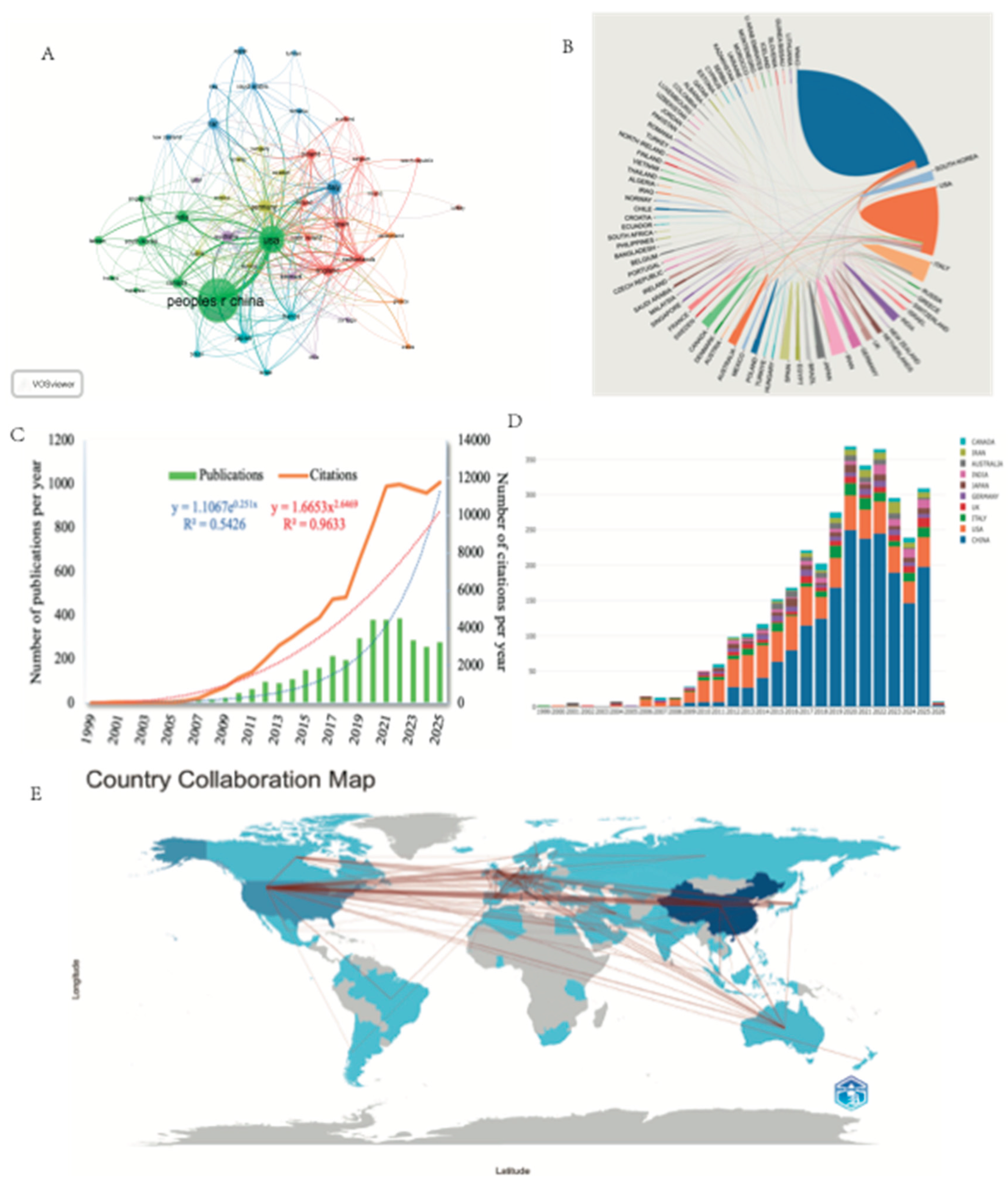
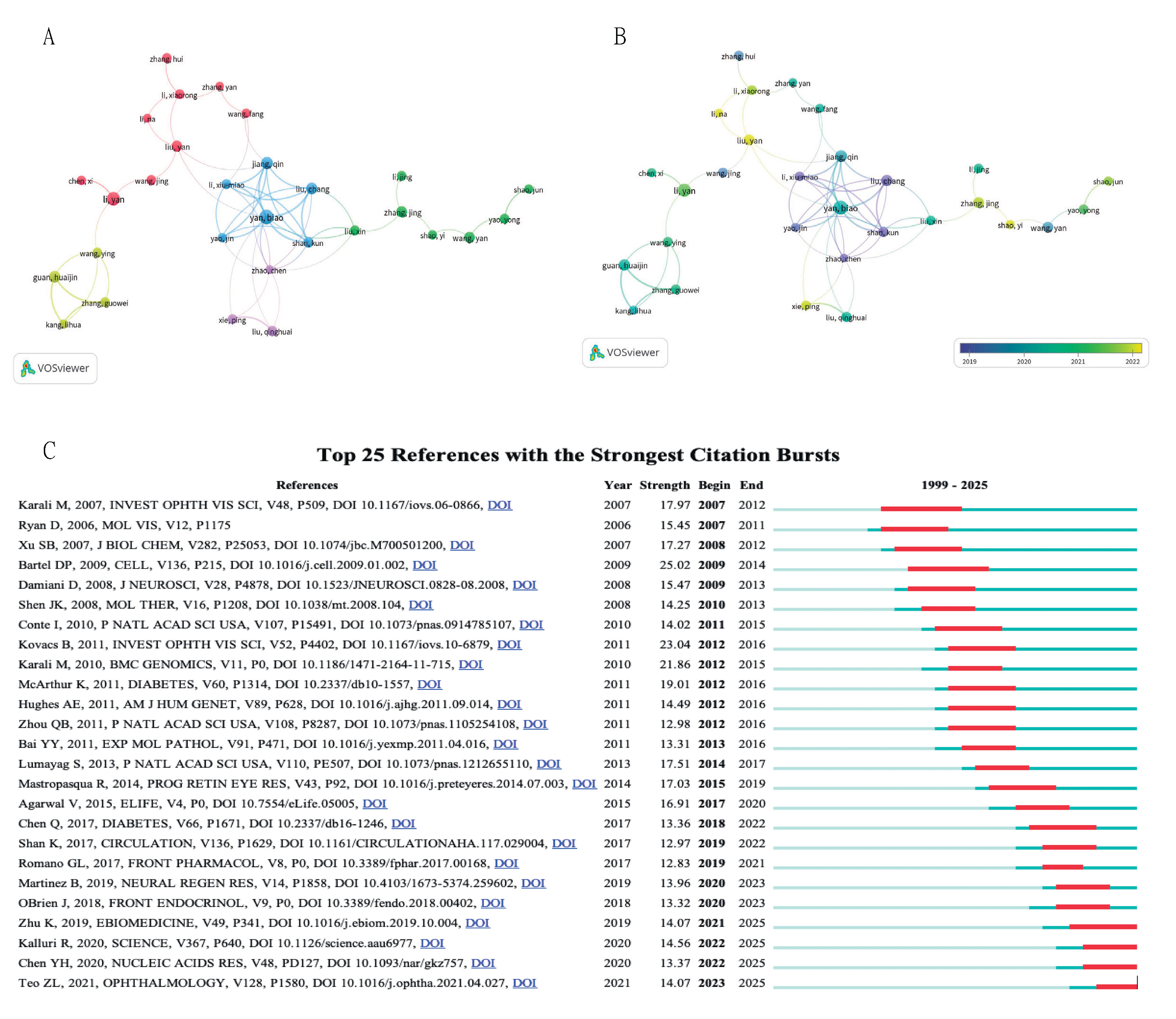
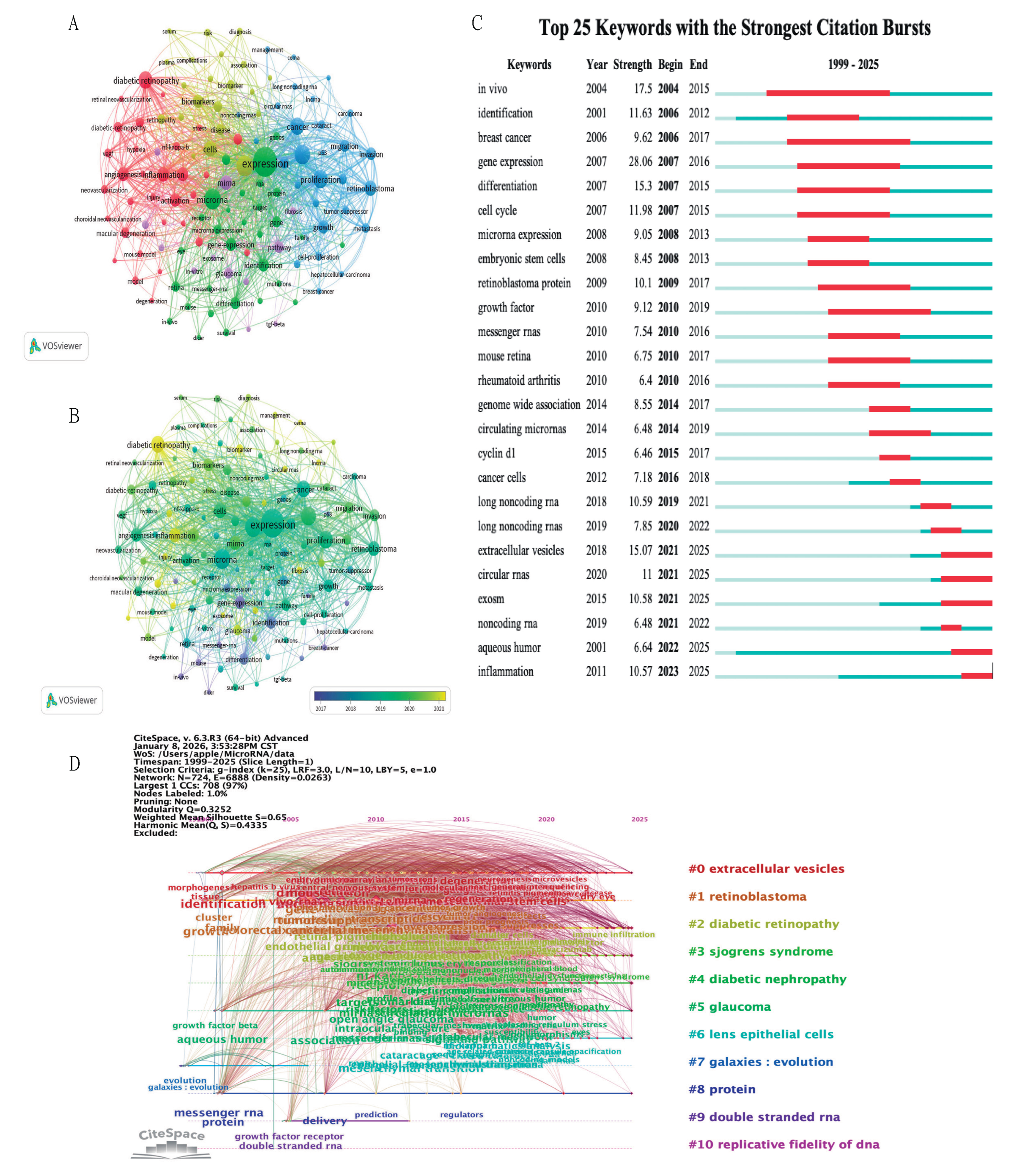
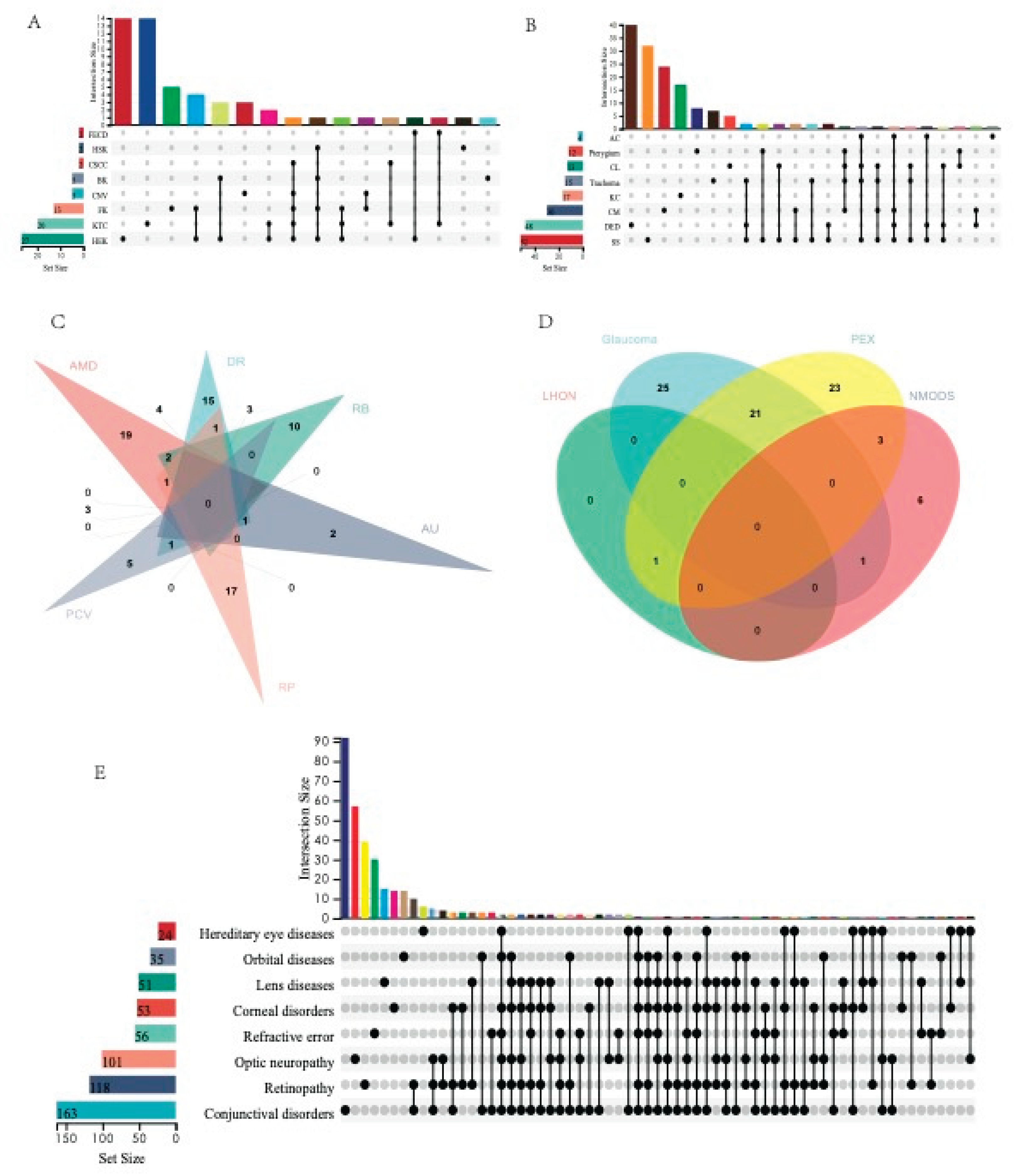
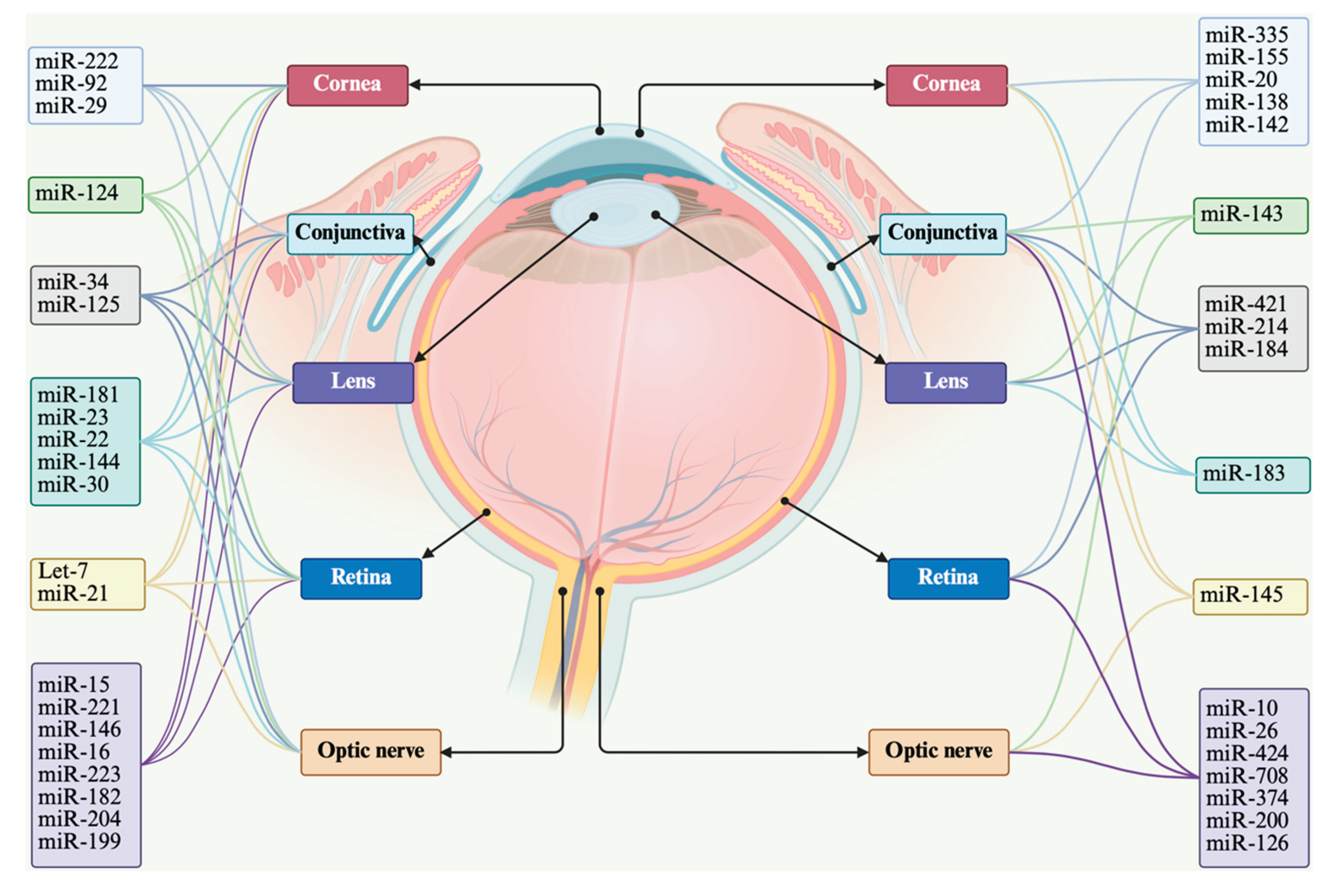
| Year | Publications | Citations |
| 1999 | 2 | 16 |
| 2000 | 1 | 23 |
| 2001 | 5 | 28 |
| 2002 | 1 | 35 |
| 2003 | 1 | 34 |
| 2004 | 5 | 43 |
| 2005 | 1 | 49 |
| 2006 | 12 | 99 |
| 2007 | 12 | 269 |
| 2008 | 15 | 555 |
| 2009 | 24 | 867 |
| 2010 | 46 | 1,334 |
| 2011 | 63 | 1,662 |
| 2012 | 96 | 2,327 |
| 2013 | 91 | 3,055 |
| 2014 | 108 | 3,532 |
| 2015 | 150 | 4,068 |
| 2016 | 159 | 4,533 |
| 2017 | 215 | 5,532 |
| 2018 | 196 | 5,651 |
| 2019 | 297 | 7,610 |
| 2020 | 382 | 9,559 |
| 2021 | 382 | 11,538 |
| 2022 | 386 | 11,640 |
| 2023 | 286 | 11,401 |
| 2024 | 258 | 11,181 |
| 2025 | 277 | 11,745 |
| Total | 3,476 | 108,708 |
| A | ||||||||||
| Rank | Country | Documents | Citations | Citation/ Document (%) | Centrality | Total link strength | ||||
| 1 | China | 1,896 | 40,916 | 21.58 | 0.18 | 252 | ||||
| 2 | USA | 696 | 41,142 | 59.11 | 0.37 | 433 | ||||
| 3 | Italy | 170 | 11,111 | 65.35 | 0.21 | 135 | ||||
| 4 | Germany | 102 | 4,882 | 47.86 | 0.09 | 127 | ||||
| 5 | Japan | 99 | 3,701 | 37.38 | 0.02 | 68 | ||||
| 6 | England | 94 | 4,058 | 43.17 | 0.27 | 143 | ||||
| 7 | India | 94 | 2,422 | 25.76 | 0.10 | 50 | ||||
| 8 | Australia | 85 | 3,035 | 35.70 | 0.06 | 90 | ||||
| 9 | Iran | 82 | 1,599 | 19.50 | 0.10 | 46 | ||||
| 10 | Canada | 77 | 4,873 | 63.28 | 0.04 | 74 | ||||
| B | ||||||||||
| Rank | Organization | Documents | Citations | Citation/ Document (%) (%) | Centrality | |||||
| 1 | Nanjing Medical University | 107 | 3,797 | 35.48 | 0.07 | |||||
| 2 | Shanghai Jiao Tong University | 106 | 2,627 | 24.78 | 0.20 | |||||
| 3 | Fudan University | 96 | 3,089 | 32.17 | 0.07 | |||||
| 4 | Sun Yat Sen University | 87 | 2,297 | 26.40 | 0.12 | |||||
| 5 | Central South University | 60 | 932 | 15.53 | 0.02 | |||||
| 6 | Wen Zhou Medical University | 60 | 1,192 | 19.86 | 0.03 | |||||
| 7 | Zheng Zhou University | 60 | 961 | 16.01 | 0.01 | |||||
| 8 | Tian Jin Medical University | 55 | 1,296 | 23.56 | 0.05 | |||||
| 9 | Ji Lin University | 51 | 1,334 | 26.15 | 0.02 | |||||
| 10 | Capital Medical University | 50 | 766 | 15.32 | 0.06 | |||||
| A | |||||||||||
| Rank | Author | Documents | Citations | Total link strength | Co-cited author | Citations | Total link strength | ||||
| 1 | Yan Biao | 27 | 2,084 | 138 | Bartel DP | 672 | 13,333 | ||||
| 2 | Li Yan | 22 | 346 | 80 | Wang Y | 378 | 7,474 | ||||
| 3 | Jiang Qin | 19 | 1,508 | 86 | Zhang Y | 369 | 7,537 | ||||
| 4 | Zhang Hong | 19 | 350 | 23 | Karali M | 301 | 9,305 | ||||
| 5 | Guan Huai Jin | 18 | 251 | 112 | Livak KJ | 285 | 3,568 | ||||
| 6 | Banfi Sandro | 18 | 978 | 82 | Li Y | 248 | 5,428 | ||||
| 7 | Liu Chang | 17 | 2,003 | 96 | Liu Y | 237 | 5,057 | ||||
| 8 | Zhang Jing | 16 | 207 | 35 | Xu SB | 236 | 6,464 | ||||
| 9 | Gonzalez Pedro | 15 | 1,073 | 45 | Zhang L | 224 | 4,430 | ||||
| 10 | Wang Yan | 15 | 417 | 29 | Ambros V | 224 | 4,339 | ||||
| B | |||||||||||
| Rank | Journal | Country | Documents (%)2024. | Citations | Total link strength | IF (2024) | Co-cited journal | Citations | Total link strength | IF (2024) | |
| 1 | IOVS | USA | 150 (4.31) | 5,432 | 1976 | 4.7 | IOVS | 7,324 | 541,486 | 4.7 | |
| 2 | Experimental Eye Research | USA | 119 (3.42) | 2,308 | 1115 | 2.7 | Plos One | 4,301 | 333,398 | 2.6 | |
| 3 | International Journal of Molecular Sciences | USA | 86 (2.47) | 1,459 | 798 | 4.9 | PNAS | 4,043 | 322,014 | 9.1 | |
| 4 | Plos One | USA | 79 (2.27) | 3,036 | 670 | 2.6 | CELL | 3,610 | 274,554 | 42.5 | |
| 5 | Scientific Reports | England | 69 (1.98) | 1,342 | 623 | 3.9 | NATURE | 3,397 | 276,337 | 48.5 | |
| 6 | Current Eye Research | Netherlands | 61 (1.75) | 595 | 539 | 2.0 | Experimental Eye Research | 2,852 | 220,606 | 2.7 | |
| 7 | Molecular Medicine Reports | Greece | 52 (1.49) | 750 | 332 | 3.5 | Journal of Biological Chemistry | 2,447 | 209,467 | 3.9 | |
| 8 | Molecular Vision | USA | 43 (1.23) | 1,757 | 604 | 1.4 | Nucleic Acids Research | 2,365 | 158,511 | 13.1 | |
| 9 | International Journal of Ophthalmology | China | 33 (0.94) | 380 | 253 | 1.8 | Scientific Reports | 2,234 | 171,979 | 3.9 | |
| 10 | Frontiers in Immunology | Switzerland | 31 (0.89) | 1,120 | 259 | 5.9 | International Journal of Molecular Sciences | 2,122 | 165,181 | 4.9 | |
| Rank | Cited reference | Title | Journal | Citations | Total link strength |
|---|---|---|---|---|---|
| 1 | bartel dp, 2004, cell, v116, p281, doi 10.1016/s0092-8674(04)00045-5 | MicroRNAs: genomics, biogenesis, mechanism, and function | CELL | 399 | 4,628 |
| 2 | livak kj, 2001, methods, v25, p402, doi 10.1006/meth.2001.1262 | Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method | Methods | 284 | 2,001 |
| 3 | bartel dp, 2009, cell, v136, p215, doi 10.1016/j.cell.2009.01.002 | MicroRNAs: target recognition and regulatory functions | CELL | 209 | 2,669 |
| 4 | ambros v, 2004, nature, v431, p350, doi 10.1038/nature02871 | The functions of animal microRNAs | NATURE | 155 | 1,754 |
| 5 | lewis bp, 2005, cell, v120, p15, doi 10.1016/j.cell.2004.12.035 | Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets | CELL | 149 | 2,111 |
| 6 | kovacs b, 2011, invest ophth vis sci, v52, p4402, doi 10.1167/iovs.10-6879 | MicroRNAs in early diabetic retinopathy in streptozotocin-induced diabetic rats | IOVS | 141 | 2,817 |
| 7 | xu sb, 2007, j biol chem, v282, p25053, doi 10.1074/jbc.m700501200 | MicroRNA (miRNA) transcriptome of mouse retina and identification of a sensory organ-specific miRNA cluster | Journal of Biological Chemistry | 136 | 3,068 |
| 8 | shen jk, 2008, mol ther, v16, p1208, doi 10.1038/mt.2008.104 | MicroRNAs regulate ocular neovascularization | Molecular Therapy | 126 | 2,606 |
| 9 | lee rc, 1993, cell, v75, p843, doi 10.1016/0092-8674(93)90529-y | The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14 | CELL | 122 | 2,819 |
| 10 | mcarthur k, 2011, diabetes, v60, p1314, doi 10.2337/db10-1557 | MicroRNA-200b regulates vascular endothelial growth factor-mediated alterations in diabetic retinopathy |
Diabetes | 113 | 2,437 |
| A | |||||||||||||
| Disease | microRNA | Disease | microRNA | Disease | microRNA | Disease | microRNA | ||||||
| AMD|DR | let-7, miR-21, miR-200, miR-138 | AMD|PCV | miR-374, miR-22, miR-502 | TRC|DED|SS | miR-142, miR-203 | HEK|BK | let-7, miR-182, miR-183 | ||||||
| AMD|RB | miR-708, miR-330 | AMD|RP|PCV | miR-335 | CL|TRC|SS | miR-150 | HEK|KTC | miR-181, miR-222 | ||||||
| AMD|RP | miR-25 | AMD|RB|PCV | miR-140 | AC|DED|SS | miR-223 | CNV|FK | miR-204 | ||||||
| DR|RB | miR-363, miR-320, miR-98 | AMD|RB|RP|PCV | miR-9, miR-181 | CL|DED|SS | let-7 | CSCC|KTC | miR-205 | ||||||
| DR|AU | miR-30 | AMD|DR|PCV | miR-20 | PTG|CL|CM|TRC | miR-184 | KTC|FECD | miR-199 | ||||||
| DR|RP | miR-126 | PTG|SS | miR-21, miR-145 | AC|CL|TRC|SS | miR-155 | LHON|PEX | miR-548 | ||||||
| AMD|DR|AU | miR-155 | CL|SS | miR-16, miR-222 | CL|CM|TRC|SS | miR-30 | GLC|NMODS | miR-200 | ||||||
| AMD|DR|RP | miR-27 | CM|SS | miR-34, miR-302 | AC|CM|DED|SS | miR-146 | PEX|NMODS | miR-126, miR-122, miR-30 | ||||||
| AMD|RB|RP | miR-34 | TRC|SS | miR-181, miR-147 | CSCC|CNV|HEK|FK|KTC | miR-184 | GLC|PEX | let-7, miR-4667, miR-637, miR-4490, miR-1253, miR-3190, miR-3173, miR-608, miR-4725, miR-4448, miR-323, miR-4538, miR-3913, miR-3159, miR-4663, miR-4767, miR-4724, miR-1306, miR-181, miR-433, miR-6777 | ||||||
| AMD|AU|RP | miR-223 | DED|SS | miR-130, miR-20 | HEK|FK|KTC | miR-146 | ||||||||
| AMD|DR|AU|RP | miR-146 | PTG|CL | miR-200 | HEK|FECD | miR-29 | ||||||||
| DR|PCV | miR-195 | CM|DED | miR-4427 | HEK|FK | miR-21, miR-124, miR-142, miR-223 | ||||||||
| B | |||||||||||||
| Disease | MicroRNA | Disease | MicroRNA | Disease | MicroRNA | ||||||||
| CJD|RD | miR-218, miR-150, miR-342, miR-485, miR-153, miR-365, miR-503, miR-106, miR-186, miR-191 | ON|HED | miR-375 | CND|CJD|LD|OD | miR-183 | ||||||||
| RD|ON | miR-1307, miR-637, miR-382, miR-1253 | CJD|RD|ON | miR-10, miR-26, miR-424, miR-708, miR-374 | CND|RD|ON|OD | miR-320 | ||||||||
| CND|RD | miR-195, miR-363, miR-25 | CND|CJD|RD | miR-335, miR-155, miR-20 | CJD|LD|RD|RE | miR-214 | ||||||||
| LD|RD | miR-377, miR-148, miR-24 | CJD|RD|OD | miR-27, miR-502 | CND|CJD|ON|RE | miR-145 | ||||||||
| LD|ON | miR-210, miR-630 | CJD|ON|RE | miR-122, miR-302 | CJD|LD|ON|RE | miR-143 | ||||||||
| CJD|OD | miR-130, miR-484 | CJD|LD|RD | miR-421 | CND|CJD|RD|HED | miR-138 | ||||||||
| CJD|RE | miR-500, miR-539, miR-18 | CND|RD|ON | miR-451 | CJD|LD|RD|HED | miR-184 | ||||||||
| CND|CJD | miR-1246, miR-132 | RD|ON|OD | miR-192 | CND|CJD|LD|RD|ON | miR-181, miR-23 | ||||||||
| CJD|LD | miR-19, miR-4728 | CND|CJD|RE | miR-127 | CND|CJD|LD|RD|RE | miR-16, miR-223 | ||||||||
| ON|RE | miR-1306, miR-671 | CND|LD|RE | miR-338 | CND|CJD|LD|RD|OD | miR-182 | ||||||||
| CJD|HED | miR-7977, miR-31 | CND|CJD|HED | miR-205 | CJD|LD|RD|ON|RE | miR-125 | ||||||||
| CJD|ON | miR-548 | CND|LD|HED | miR-193 | CND|CJD|RD|OD|RE | miR-142 | ||||||||
| CND|OD | miR-96 | LD|RD|HED | miR-224 | CND|CJD|LD|RD|HED | miR-204 | ||||||||
| RD|OD | miR-129 | CJD|ON|HED | miR-1260 | CND|CJD|LD|RD|ON|OD | miR-22, miR-144 | ||||||||
| LD|RE | miR-216 | CND|CJD|LD|RD | miR-15, miR-221 | CND|CJD|LD|RD|OD|RE | miR-199 | ||||||||
| RD|RE | miR-98 | CND|CJD|LD|ON | miR-222, miR-92 | CND|CJD|LD|ON|OD|RE | miR-29 | ||||||||
| OD|RE | miR-101 | CJD|RD|ON|RE | miR-200, miR-126 | CND|CJD|LD|RD|ON|HED | miR-30 | ||||||||
| CND|HED | miR-7 | CND|LD|RD|ON | miR-124 | CND|CJD|RD|ON|OD|RE|HED | let-7, miR-21 | ||||||||
| LD|HED | miR-211 | CJD|LD|RD|ON | miR-34 | CND|CJD|LD|RD|OD|RE|HED | miR-146 | ||||||||
| MircoRNA | MircoRNA | MircoRNA |
|---|---|---|
| miR-146 (76) | let-7 (28) | miR-150 (7) |
| miR-126 (53) | miR-223 (15) | miR-192 (7) |
| miR-155 (52) | miR-182 (13) | miR-30 (7) |
| miR-34 (45) | miR-214 (12) | miR-132 (6) |
| miR-21 (43) | miR-9 (12) | miR-23 (5) |
| miR-204 (41) | miR-181 (11) | |
| miR-29 (39) | miR-96 (9) | |
| miR-184 (24) | miR-183 (8) | |
| miR-200 (23) | miR-142 (7) | |
| miR-124 (31) | miR-145 (7) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




