Submitted:
09 March 2026
Posted:
10 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Participants
2.2. Definitions of Metabolic Conditions
2.3. Pool Definitions for GWAS
2.4. Statistical Analyses
2.5. Functional Relevance of Strongly Associated Variants and Genes
3. Results
3.1. Study Participants and Pools
3.2. Heritability of Blood Pressure-Related Traits
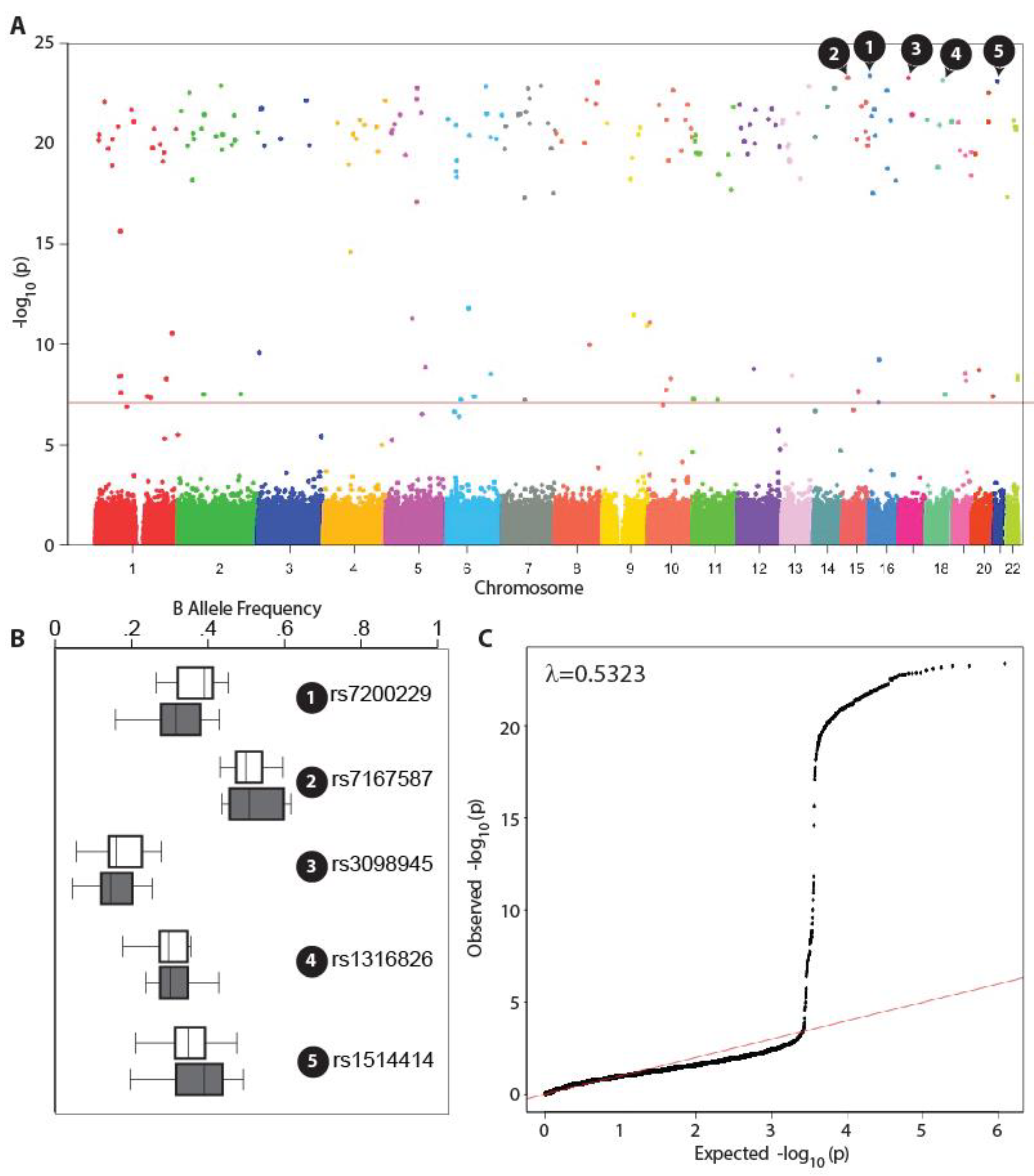
3.3. Pooled GWAS Results at the Level of Variant
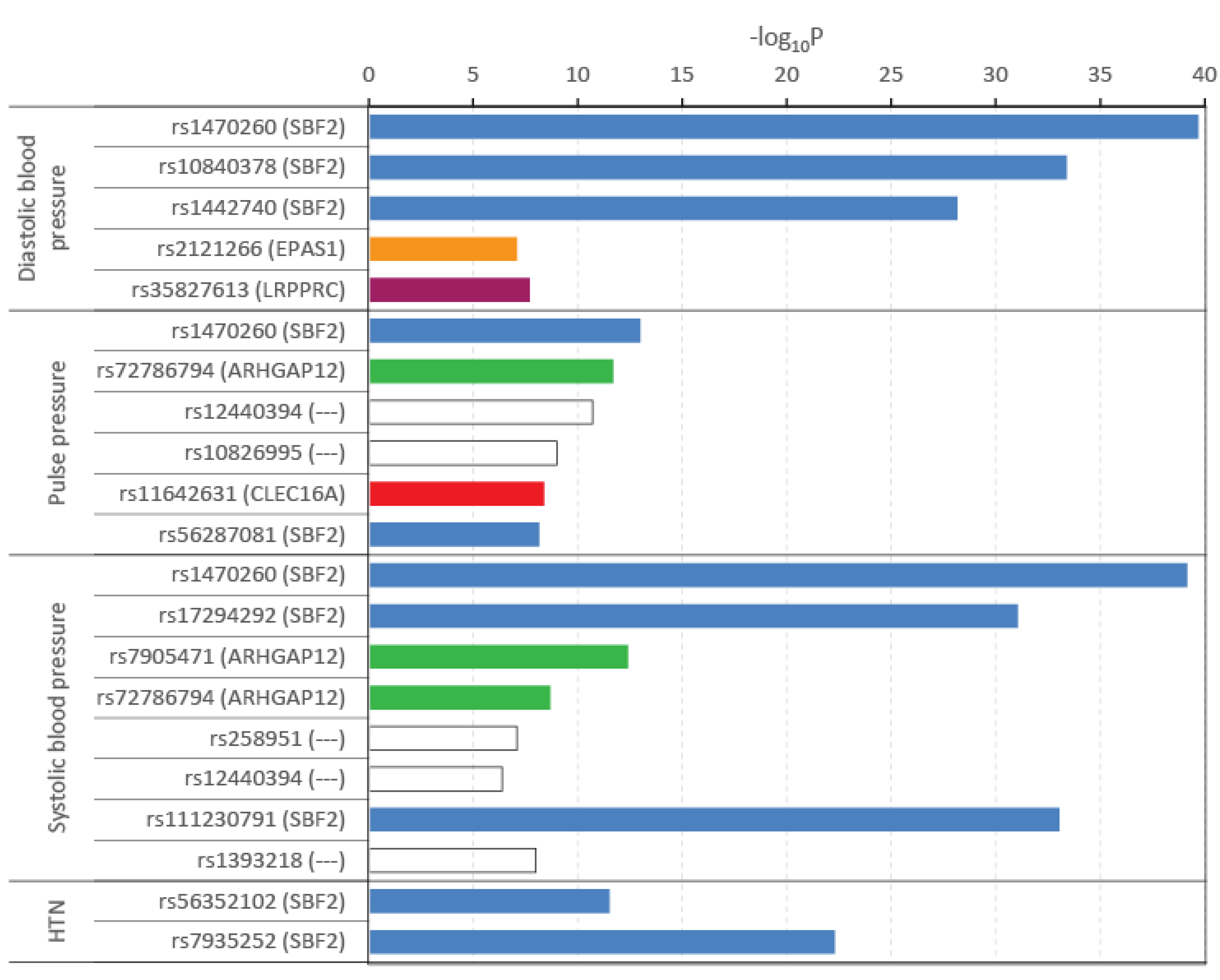
3.4. Pooled GWAS Results at the Level of Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World_Health_Organization. Global report on hypertension 2025: High stakes - turning evidence into action; World Health Organization: Geneva, Switzerland, 2025. [Google Scholar]
- Shih, P.A.; O'Connor, D.T. Hereditary determinants of human hypertension: strategies in the setting of genetic complexity. Hypertension 2008, 51, 1456–1464. [Google Scholar] [CrossRef]
- Kolifarhood, G.; Daneshpour, M.; Hadaegh, F.; Sabour, S.; Mozafar Saadati, H.; Akbar Haghdoust, A.; Akbarzadeh, M.; Sedaghati-Khayat, B.; Khosravi, N. Heritability of blood pressure traits in diverse populations: a systematic review and meta-analysis. J Hum Hypertens 2019, 33, 775–785. [Google Scholar] [CrossRef] [PubMed]
- Waken, R.J.; de las Fuentes, L.; Rao, D.C. A Review of the Genetics of Hypertension with a Focus on Gene-Environment Interactions. Curr Hypertens Rep 2017, 19, 23. [Google Scholar] [CrossRef] [PubMed]
- Keaton, J.M.; Kamali, Z.; Xie, T.; Vaez, A.; Williams, A.; Goleva, S.B.; Ani, A.; Evangelou, E.; Hellwege, J.N.; Yengo, L.; et al. Genome-wide analysis in over 1 million individuals of European ancestry yields improved polygenic risk scores for blood pressure traits. Nat Genet 2024, 56, 778–791. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Ram, C.V.S. Hypertension epidemiology in India: emerging aspects. Curr Opin Cardiol 2019, 34, 331–341. [Google Scholar] [CrossRef]
- Mohammad, R.; Bansod, D.W. Hypertension in India: a gender-based study of prevalence and associated risk factors. BMC Public Health 2024, 24, 2681. [Google Scholar] [CrossRef]
- Venkatesan, V.; Lopez-Alvarenga, J.C.; Arya, R.; Ramu, D.; Koshy, T.; Ravichandran, U.; Ponnala, A.R.; Sharma, S.K.; Lodha, S.; Sharma, K.K.; et al. Burden of Type 2 Diabetes and Associated Cardiometabolic Traits and Their Heritability Estimates in Endogamous Ethnic Groups of India: Findings From the INDIGENIUS Consortium. Front Endocrinol (Lausanne) 2022, 13, 847692. [Google Scholar] [CrossRef]
- Newton-Cheh, C.; Johnson, T.; Gateva, V.; Tobin, M.D.; Bochud, M.; Coin, L.; Najjar, S.S.; Zhao, J.H.; Heath, S.C.; Eyheramendy, S.; et al. Genome-wide association study identifies eight loci associated with blood pressure. Nat Genet 2009, 41, 666–676. [Google Scholar] [CrossRef]
- Mamtani, M.; Jaisinghani, M.T.; Jaiswal, S.G.; Pipal, K.V.; Patel, A.A.; Kulkarni, H. Genetic association of anthropometric traits with type 2 diabetes in ethnically endogamous Sindhi families. PLoS One 2021, 16, e0257390. [Google Scholar] [CrossRef] [PubMed]
- Whelton, P.K.; Carey, R.M. The 2017 American College of Cardiology/American Heart Association Clinical Practice Guideline for High Blood Pressure in Adults. JAMA Cardiol 2018, 3, 352–353. [Google Scholar] [CrossRef]
- Kumar, R.; Nandhini, L.P.; Kamalanathan, S.; Sahoo, J.; Vivekanadan, M. Evidence for current diagnostic criteria of diabetes mellitus. World J Diabetes 2016, 7, 396–405. [Google Scholar] [CrossRef]
- Snehalatha, C.; Viswanathan, V.; Ramachandran, A. Cutoff values for normal anthropometric variables in asian Indian adults. Diabetes Care 2003, 26, 1380–1384. [Google Scholar] [CrossRef]
- Sharma, Y.K.; Prakash, N.; Gupta, A. Prevalence of metabolic syndrome as per the NCEP and IDF definitions vis-a-vis severity and duration of psoriasis in a semi-urban Maharashtrian population: A case control study. Diabetes Metab Syndr 2016, 10, S72–76. [Google Scholar] [CrossRef]
- Craig, J.E.; Hewitt, A.W.; McMellon, A.E.; Henders, A.K.; Ma, L.; Wallace, L.; Sharma, S.; Burdon, K.P.; Visscher, P.M.; Montgomery, G.W.; et al. Rapid inexpensive genome-wide association using pooled whole blood. Genome Res 2009, 19, 2075–2080. [Google Scholar] [CrossRef] [PubMed]
- Pipal, K.V.; Mamtani, M.; Patel, A.A.; Jaiswal, S.G.; Jaisinghani, M.T.; Kulkarni, H. Susceptibility Loci for Type 2 Diabetes in the Ethnically Endogamous Indian Sindhi Population: A Pooled Blood Genome-Wide Association Study. Genes (Basel) 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Turner, S. qqman: an R package for visualizing GWAS results using Q-Q and manhattan plots. The Journal of Open Source Software 2018. [Google Scholar] [CrossRef]
- Macgregor, S.; Zhao, Z.Z.; Henders, A.; Nicholas, M.G.; Montgomery, G.W.; Visscher, P.M. Highly cost-efficient genome-wide association studies using DNA pools and dense SNP arrays. Nucleic Acids Res 2008, 36, e35. [Google Scholar] [CrossRef]
- Ani, A.; Kamali, Z.; Vaez, A. SNPAnnotator: Automated Functionanl Annotation of Genetic Variants and Linked Proxies. 2026, 1.4.6. [Google Scholar]
- Santra, A.; Mallick, A. Prevalence of hypertension among individuals with diabetes and its determinants: evidences from the National Family Health Survey 2015-16, India. Ann Hum Biol 2022, 49, 133–144. [Google Scholar] [CrossRef]
- Anjana, R.M.; Unnikrishnan, R.; Deepa, M.; Pradeepa, R.; Tandon, N.; Das, A.K.; Joshi, S.; Bajaj, S.; Jabbar, P.K.; Das, H.K.; et al. Metabolic non-communicable disease health report of India: the ICMR-INDIAB national cross-sectional study (ICMR-INDIAB-17). Lancet Diabetes Endocrinol 2023, 11, 474–489. [Google Scholar] [CrossRef]
- Maheshwari, A.; Gupta, R.; Verma, N.; Narasingan, S.N.; Singh, R.B.; Saboo, B.; Kumar, C.H.V.; Gupta, A.; Srivastava, M.K.; Gupta, A.; et al. Position statement on hypertension by Indian Society of Hypertension, 2023. J Hum Hypertens 2024, 38, 736–744. [Google Scholar] [CrossRef]
- Senderek, J.; Bergmann, C.; Weber, S.; Ketelsen, U.P.; Schorle, H.; Rudnik-Schoneborn, S.; Buttner, R.; Buchheim, E.; Zerres, K. Mutation of the SBF2 gene, encoding a novel member of the myotubularin family, in Charcot-Marie-Tooth neuropathy type 4B2/11p15. Hum Mol Genet 2003, 12, 349–356. [Google Scholar] [CrossRef]
- Wang, J.; Guo, W.; Wang, Q.; Yang, Y.; Sun, X. Recent advances of myotubularin-related (MTMR) protein family in cardiovascular diseases. Front Cardiovasc Med 2024, 11, 1364604. [Google Scholar] [CrossRef]
- Dbouk, H.A.; Huang, C.L.; Cobb, M.H. Hypertension: the missing WNKs. Am J Physiol Renal Physiol 2016, 311, F16–27. [Google Scholar] [CrossRef]
- Luque, Y.; Lenoir, O.; Bonnin, P.; Hardy, L.; Chipont, A.; Placier, S.; Vandermeersch, S.; Xu-Dubois, Y.C.; Robin, B.; Lazareth, H.; et al. Endothelial Epas1 Deficiency Is Sufficient To Promote Parietal Epithelial Cell Activation and FSGS in Experimental Hypertension. J Am Soc Nephrol 2017, 28, 3563–3578. [Google Scholar] [CrossRef] [PubMed]
- Barry, A.; McNulty, M.T.; Jia, X.; Gupta, Y.; Debiec, H.; Luo, Y.; Nagano, C.; Horinouchi, T.; Jung, S.; Colucci, M.; et al. Multi-population genome-wide association study implicates both immune and non-immune factors in the etiology of pediatric steroid sensitive nephrotic syndrome. medRxiv 2022, 2022.2009.2013.22279644. [CrossRef]
- Proust, C.; Empana, J.P.; Boutouyrie, P.; Alivon, M.; Challande, P.; Danchin, N.; Escriou, G.; Esslinger, U.; Laurent, S.; Li, Z.; et al. Contribution of Rare and Common Genetic Variants to Plasma Lipid Levels and Carotid Stiffness and Geometry: A Substudy of the Paris Prospective Study 3. Circ Cardiovasc Genet 2015, 8, 628–636. [Google Scholar] [CrossRef]
- Gao, G.; Chen, A.; Gong, J.; Lin, W.; Wu, W.; Mohammad Ismail Hajary, S.; Lian, G.; Luo, L.; Xie, L. Comprehensive analyses of m6A RNA methylation patterns and related immune microenvironment in idiopathic pulmonary arterial hypertension. Front Genet 2023, 14, 1222368. [Google Scholar] [CrossRef] [PubMed]
- Andreassen, O.A.; McEvoy, L.K.; Thompson, W.K.; Wang, Y.; Reppe, S.; Schork, A.J.; Zuber, V.; Barrett-Connor, E.; Gautvik, K.; Aukrust, P.; et al. Identifying common genetic variants in blood pressure due to polygenic pleiotropy with associated phenotypes. Hypertension 2014, 63, 819–826. [Google Scholar] [CrossRef]
- Mani, N.; Nygaard, R.; Mancia, F. Gone with the Wnt(less): a mechanistic perspective on the journey of Wnt. Biochem Soc Trans 2022, 50, 1763–1772. [Google Scholar] [CrossRef] [PubMed]
- Williams, H.; Mill, C.A.; Monk, B.A.; Hulin-Curtis, S.; Johnson, J.L.; George, S.J. Wnt2 and WISP-1/CCN4 Induce Intimal Thickening via Promotion of Smooth Muscle Cell Migration. Arterioscler Thromb Vasc Biol 2016, 36, 1417–1424. [Google Scholar] [CrossRef] [PubMed]
- van der Harst, P.; Verweij, N. Identification of 64 Novel Genetic Loci Provides an Expanded View on the Genetic Architecture of Coronary Artery Disease. Circ Res 2018, 122, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Miller, C.L.; Bentley, A.M.; Brown, M.R.; Nagarajan, P.; Noordam, R.; Morrison, J.; Schwander, K.; Westerman, K.; Kho, M.; et al. A Large-Scale Genome-wide Association Study of Blood Pressure Accounting for Gene-Depressive Symptomatology Interactions in 564,680 Individuals from Diverse Populations. Research Square (Preprint) 2025. [Google Scholar] [CrossRef]
- Kunnas, T.; Nikkari, S.T. Contribution of syndecan-4 genetic variants to hypertension, the TAMRISK study. BMC Res Notes 2014, 7, 815. [Google Scholar] [CrossRef]
- Vargas, J.D.; Abbas, M.; Goodney, G.; Le, H.; Hinton, A.O.; Gaye, A. Regulatory Roles of Long Noncoding RNAs in Arterial Stiffness and Hypertension. Hypertension 2025, 82, 1195–1207. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Ning, Q. Long noncoding RNAs as novel players in the pathogenesis of hypertension. Hypertens Res 2020, 43, 597–608. [Google Scholar] [CrossRef] [PubMed]
- Gluba-Sagr, A.; Franczyk, B.; Rysz-Gorzynska, A.; Olszewski, R.; Rysz, J. The Role of Selected lncRNAs in Lipid Metabolism and Cardiovascular Disease Risk. Int J Mol Sci 2024, 25. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



