Submitted:
08 March 2026
Posted:
10 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Filter for domain-specific relevance within medical IT systems.
- Precisely identify affected vendors and products to quantify vulnerability trends.
- Categorize vulnerabilities according to their position in the medical chain, from data generation to interoperability systems.
- Determine the specific component type and map the CVE to relevant MITRE ATT&CK categories.
2. Overview of Cybersecurity Concepts
2.1. CVE Catalog
- CVE identifier: A unique, standardized reference key (e.g., CVE-YYYY-NNNNN) assigned to a specific vulnerability for cross-database correlation.
- Description: A text summary explaining the vulnerability’s root cause, attack vector, and potential impact.
- Affected system: The specific vendor, product, and version(s) in which the vulnerability exists.
- Severity metrics (CVSS): A standardized framework that assigns a quantitative base score (0.0 to 10.0) reflecting the flaw’s intrinsic severity. This score is derived from exploitability metrics—such as the required attack vector, complexity, and privilege levels—as well as the potential impact.
- Administrative metadata: Details regarding the assigning CNA (CVE Numbering Authority) and timestamps for the record’s publication and most recent updates.
2.2. MITRE ATT&CK Framework
- Tactics: Represent the specific, short-term operational objectives an adversary seeks to achieve during the life-cycle of a cyberattack. The Enterprise matrix outlines 14 distinct tactics that encompass the entire attack continuum. These span from initial network compromise (Initial Access, Execution), to maintaining presence and stealth (Persistence, Privilege Escalation, Defense Evasion, Credential Access), navigating the environment (Discovery, Lateral Movement), and finally executing ultimate objectives against the target data or systems (Collection, Command and Control, Exfiltration, Impact).
- Techniques: Detail the precise technological or operational methods employed by an adversary to accomplish a given tactical objective. For example, to achieve Initial Access, an attacker might utilize the technique of Phishing or exploiting public-facing applications.
- Procedures: Document the exact operational steps, software tools, and malware signatures utilized by specific threat groups (e.g., Advanced Persistent Threats, or APTs) to execute a technique. For example, a given procedure might detail how a specific ransomware syndicate exploits a known vulnerability to encrypt file-systems.
3. Overview of Medical IT Systems
-
Data identity and management systems: These systems form the administrative backbone of healthcare organizations, managing patient identity and life-cycle. These systems include:
- HIS (Hospital Information System): Designed to manage medical, administrative, financial, and legal aspects of hospital operations, serving as the central integration point for departmental subsystems.
- EMR (Electronic Medical Record): Digital versions of traditional paper-based medical charts. EMRs are typically confined to a single healthcare provider and are not designed for extensive data sharing.
- EHR (Electronic Health Record): Electronic repositories of patient health information designed to support data sharing across multiple healthcare providers and organizations.
-
Departmental information systems: These systems support the operational workflows of specific clinical departments inside a medical organization, managing specialized ordering, tracking, and result-reporting processes, and acting as an intermediate operational layer between clinicians and the core medical records. These systems include:
- RIS (Radiology Information System): Systems for managing radiological workflows, often integrated with PACS and VNA solutions.
- LIS (Laboratory Information System): Managing laboratory operations such as order entry, specimen tracking, result reporting, and quality control.
- PIS (Pharmacy Information System): Supporting medication management, including drug dispensing, inventory control, and clinical checks.
- Data generation systems: Data generation systems comprise medical equipment and devices that produce clinical data, including diagnostic imaging modalities and bedside devices. These devices are typically integrated with medical IT systems via interoperability standards, such as DICOM or HL7. Examples of data generation systems include: CT (Computed Tomography), MRI (Magnetic Resonance Imaging), PET (Positron Emission Tomography), and X-Ray, among others.
-
Storage and archiving systems: These systems provide long-term storage and access to high-volume clinical data, particularly medical images, ensuring data integrity and availability. These systems include:
- PACS (Picture Archiving and Communication System): Providing efficient storage, retrieval, and distribution of medical images generated by multiple imaging modalities.
- VNA (Vendor Neutral Archive): Centralized archives that store medical images and documents in standardized formats.
-
Interoperability standards: Define data communication and storage protocols that act as middleware to enable data exchange between heterogeneous healthcare systems. These standards include:
- DICOM (Digital Imaging and Communications in Medicine): International standard for the secure transmission, storage, and sharing of medical images (like MRIs or CT scans) and their associated data. It ensures interoperability between imaging equipment and healthcare IT systems across different manufacturers.
- HL7 (Health Level Seven): A framework of international standards governing the exchange, integration, and retrieval of electronic health information. It acts as a universal language that allows disparate medical software systems to share clinical and administrative data seamlessly.
- IT communication systems: The underlying network infrastructure that interconnects medical IT and clinical systems, enabling data exchange and service availability. They include wired and wireless networks, segmentation mechanisms such as VLANs, and security controls such as firewalls and access policies, among others.
4. Methodology
4.1. CVE Database Creation
4.2. LLM Setup and Configuration Parameters
4.3. LLM-Based Classification of CVEs
4.4. LLM Prompt Development
5. Results
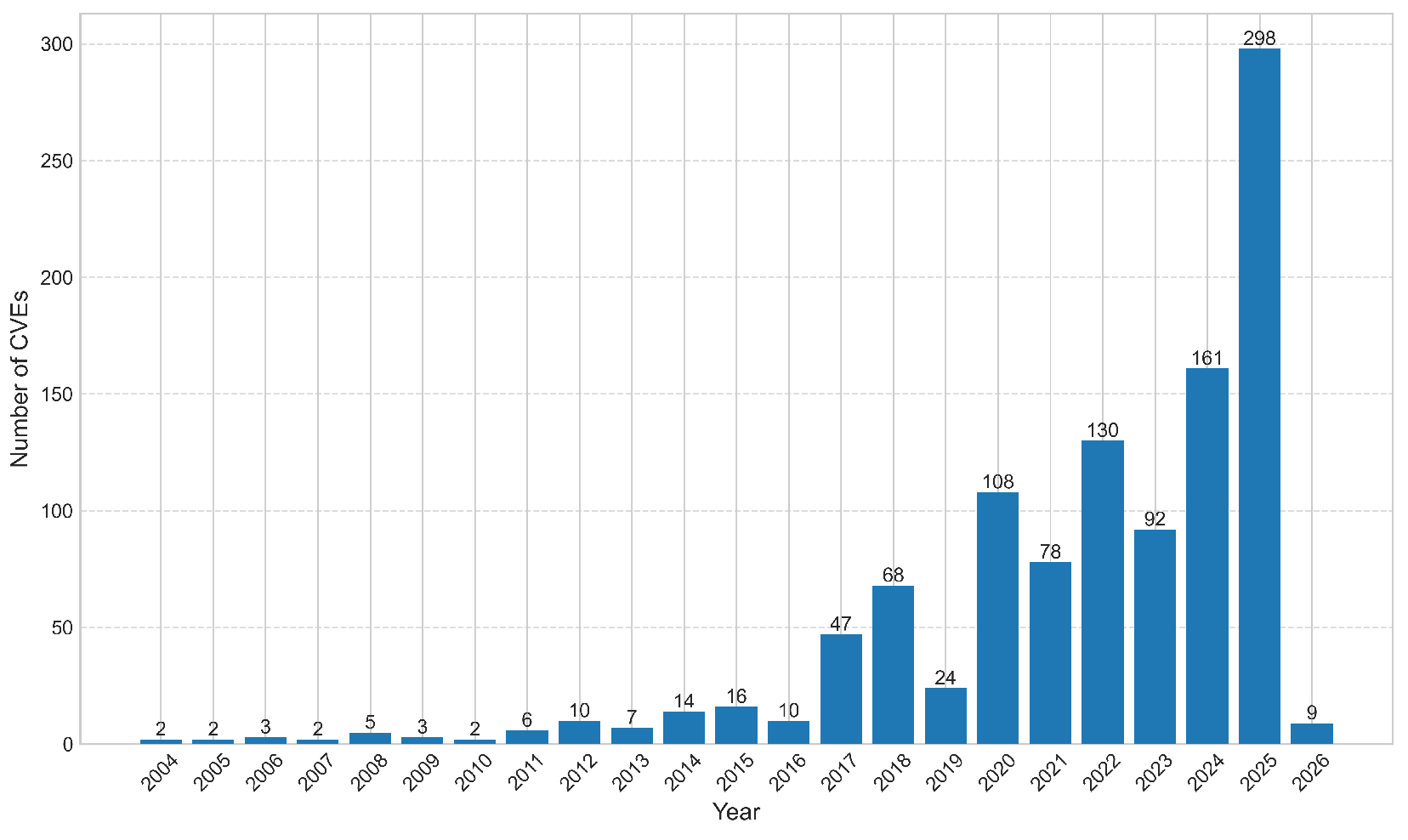
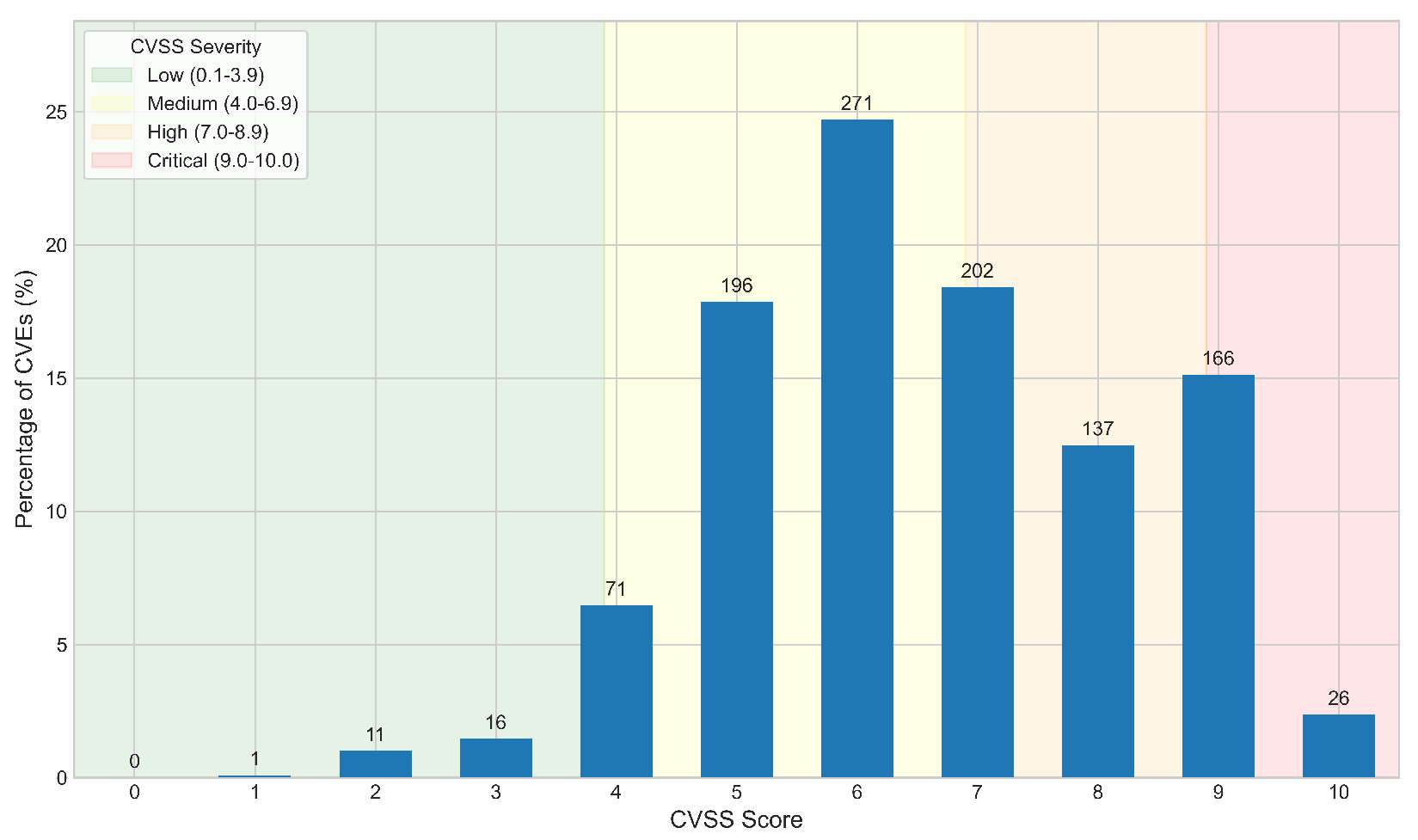
5.1. Preliminary Database Analysis
5.2. LLM-Based Database Analysis
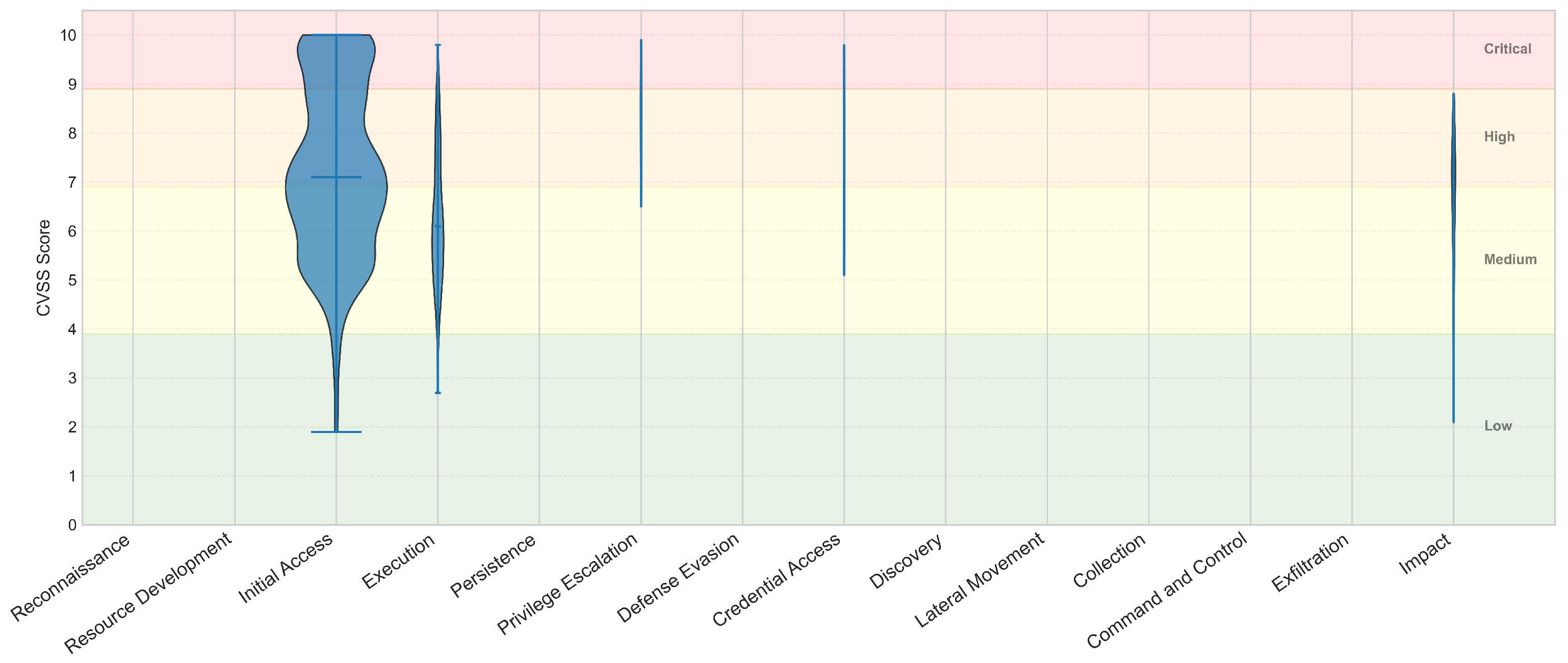
| ATT&CK Phase | Low (%) | Med (%) | High (%) | Crit (%) |
|---|---|---|---|---|
| Reconnaissance | - | - | - | - |
| Resource Development | - | - | - | - |
| Initial Access | 2.3% | 46.5% | 30.6% | 20.6% |
| Execution | 2.1% | 67.7% | 29.2% | 1.0% |
| Persistence | - | - | - | - |
| Privilege Escalation | 0.0% | 12.5% | 62.5% | 25.0% |
| Defense Evasion | - | - | - | - |
| Credential Access | 0.0% | 33.3% | 55.6% | 11.1% |
| Discovery | - | - | - | - |
| Lateral Movement | - | - | - | - |
| Collection | - | - | - | - |
| Command and Control | - | - | - | - |
| Exfiltration | - | - | - | - |
| Impact | 3.3% | 46.7% | 50.0% | 0.0% |
5.3. Discussion
6. Conclusions and Future Work
Acknowledgments
| 1 | Using such approach we retrieved 35 CVEs, which were also included to the CVE database. |
| 2 | The number in parenthesis represents the number of CVE entries that each keyword search returned. |
References
- Walkowski, Michał et al.. Vulnerability Management Models Using a Common Vulnerability Scoring System. 11, 8735. [CrossRef]
- Boateng, Gordon Owusu et al.. A Survey on Large Language Models for Communication, Network, and Service Management: Application Insights, Challenges, and Future Directions. IEEE Communications Surveys & Tutorials 2026, 28, 527–566. [CrossRef]
- Marchiori, Francesco et al.. Can LLMs Classify CVEs? Investigating LLMs Capabilities in Computing CVSS Vectors. In Proceedings of the 2025 IEEE Symposium on Computers and Communications (ISCC), 2025, pp. 1–6. [CrossRef]
- Miranda, Lucas et al.. Learning CNA-Oriented CVSS Scores. In Proceedings of the 2024 IEEE 13th International Conference on Cloud Networking (CloudNet), 2024, pp. 1–5. [CrossRef]
- Mirtaheri, Seyedeh Leili et al.. Knowledge-Driven Large Language Models for Automating CVSS Score Prediction. In Proceedings of the Proceedings of the 2025 Workshop on Research on Offensive and Defensive Techniques in the Context of Man At The End (MATE) Attacks, New York, NY, USA, 2025; CheckMATE ’25, p. 20–28. [CrossRef]
- Jafarikhah Sima, et al.. From Description to Score: Can LLMs Quantify Vulnerabilities?, 2026, [arXiv:cs.CR/2512.06781].
- Tuset-Peiró, Pere et al.. Assessing cybersecurity of Internet-facing medical IT systems in Germany & Spain using OSINT tools. In Proceedings of the Actas de las X Jornadas Nacionales de Investigación en Ciberseguridad (JNIC), Zaragoza, Spain, 2025; pp. 190–197.
- HL7 International. HL7 Version 2 Product Suite. https://www.hl7.org/implement/standards/product_brief.cfm?product_id=185, 2023. Accessed: 2026-02.
- ISO/HL7 10781:2015; Health Informatics — Electronic Health Record System Functional Model (EHR-S FM). International Organization for Standardization: 2015.
- ISO 12967:2012; Health Informatics — Service Architecture (HISA). International Organization for Standardization: 2012.
- DICOM Standards Committee. Digital Imaging and Communications in Medicine (DICOM) Standard. https://www.dicomstandard.org, 2023. Accessed: 2026-02.
- Shortliffe, E.H.; Cimino, J.J. Biomedical Informatics: Computer Applications in Health Care and Biomedicine, 4th ed.; Springer, 2014.
- Huang, H.K. PACS and Imaging Informatics: Basic Principles and Applications, 2nd ed.; Wiley-Blackwell, 2010.
- Naveed, H.e.a. A Comprehensive Overview of Large Language Models. ACM Trans. Intell. Syst. Technol. 2025, 16. [CrossRef]
- Gemma Team et al.. Gemma 3 Technical Report, 2025, [arXiv:cs.CL/2503.19786].
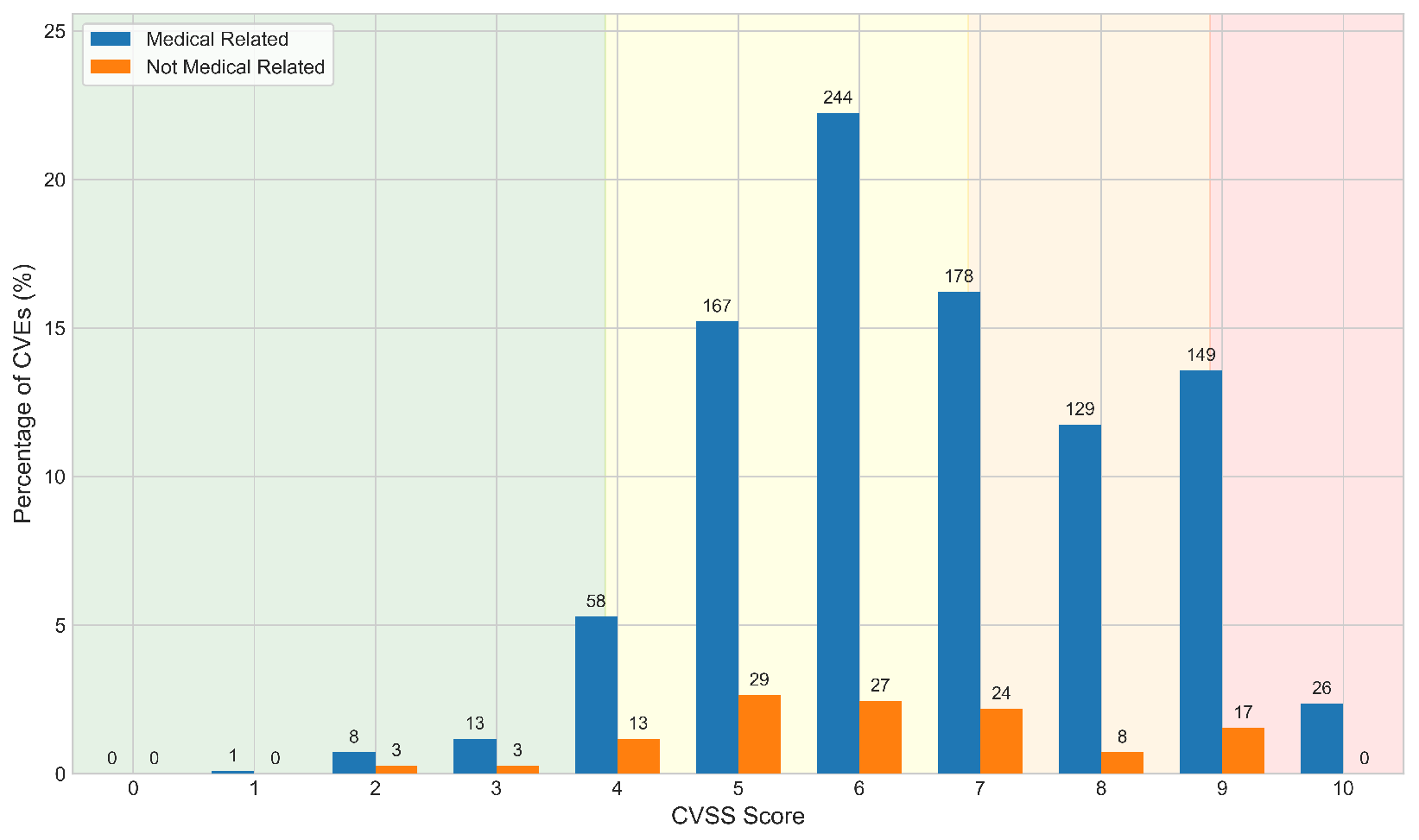
| Value | Med (N) | Med (%) | Non-Med (N) | Non-Med (%) |
|---|---|---|---|---|
| 0 | 0 | 0.0% | 0 | 0.0% |
| 1 | 1 | 0.1% | 0 | 0.0% |
| 2 | 8 | 0.7% | 3 | 0.3% |
| 3 | 13 | 1.2% | 3 | 0.3% |
| 4 | 58 | 5.3% | 13 | 1.2% |
| 5 | 167 | 15.2% | 29 | 2.6% |
| 6 | 244 | 22.2% | 27 | 2.5% |
| 7 | 178 | 16.2% | 24 | 2.2% |
| 8 | 129 | 11.8% | 8 | 0.7% |
| 9 | 149 | 13.6% | 17 | 1.5% |
| 10 | 26 | 2.4% | 0 | 0.0% |
| Total | 973 | 88.7% | 124 | 11.3% |
| Category | Count | Percentage |
|---|---|---|
| True Positives (TP) | 954 | 86.96% |
| True Negatives (TN) | 55 | 5.01% |
| False Positives (FP) | 19 | 1.73% |
| False Negatives (FN) | 69 | 6.29% |
| Total | 1097 | 100% |
| Metric | Value |
|---|---|
| Precision | 98.05% |
| Recall (Sensitivity) | 93.25% |
| F1-score | 95.60% |
| Accuracy | 92.00% |
| Category | Low (%) | Med (%) | High (%) | Crit (%) |
|---|---|---|---|---|
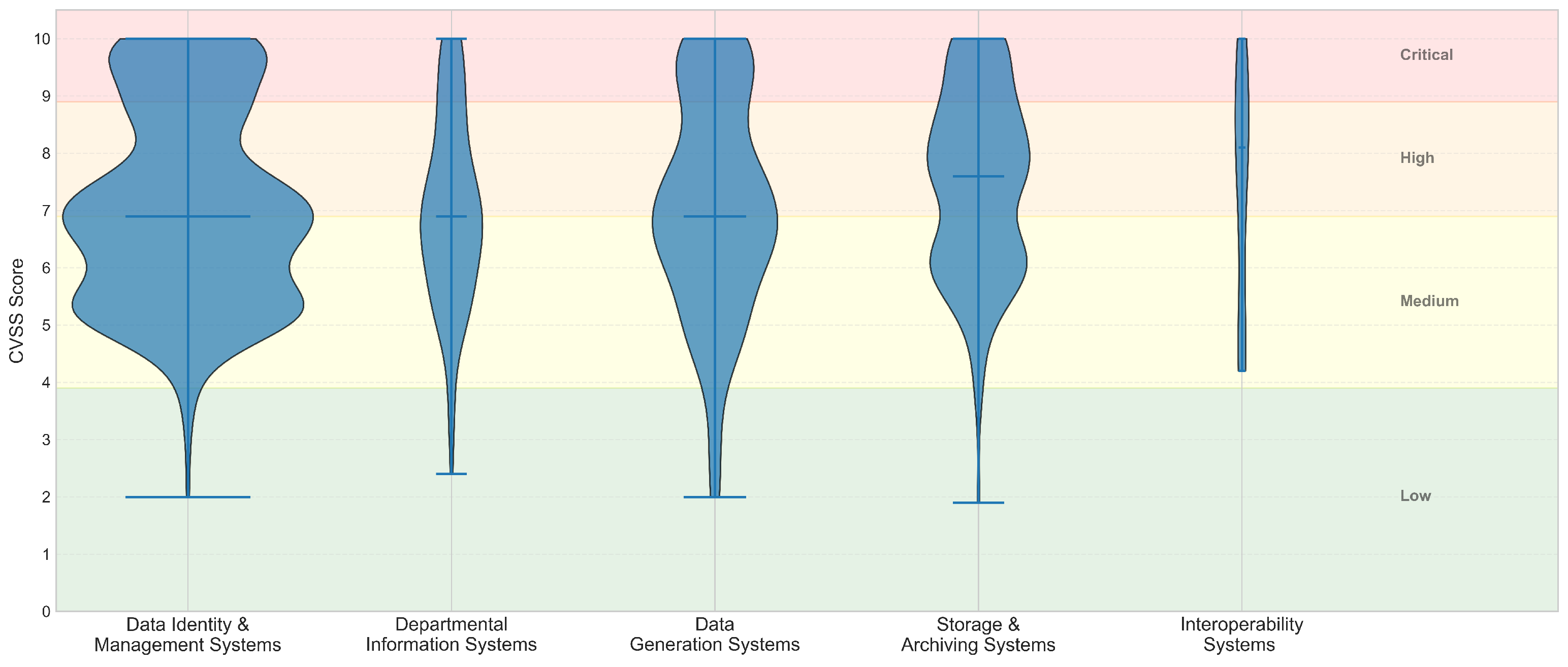
| Data Identity & Management Systems | 1.4% | 53.4% | 27.3% | 18.0% |
| Departmental Information Systems | 3.7% | 52.3% | 31.2% | 12.8% |
| Data Generation Systems | 4.5% | 45.9% | 29.1% | 20.5% |
| Storage & Archiving Systems | 1.1% | 37.8% | 43.3% | 17.8% |
| Interoperability Systems | 0.0% | 33.3% | 45.8% | 20.8% |
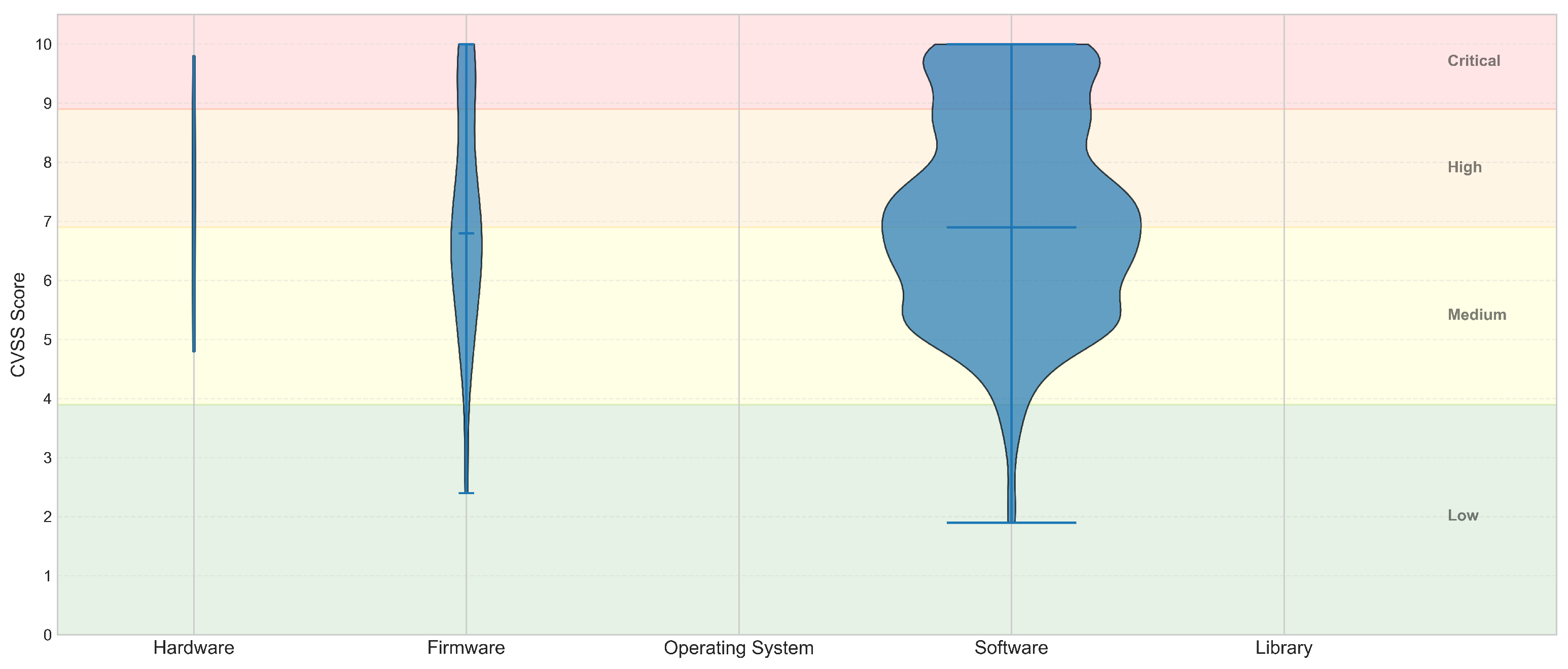
| Component | Low (%) | Med (%) | High (%) | Crit (%) |
|---|---|---|---|---|
| Hardware | 0.0% | 50.0% | 30.0% | 20.0% |
| Firmware | 3.9% | 48.5% | 27.2% | 20.4% |
| Operating System | - | - | - | - |
| Software | 2.1% | 48.1% | 32.1% | 17.7% |
| Library | - | - | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




