Submitted:
09 March 2026
Posted:
10 March 2026
You are already at the latest version
Abstract
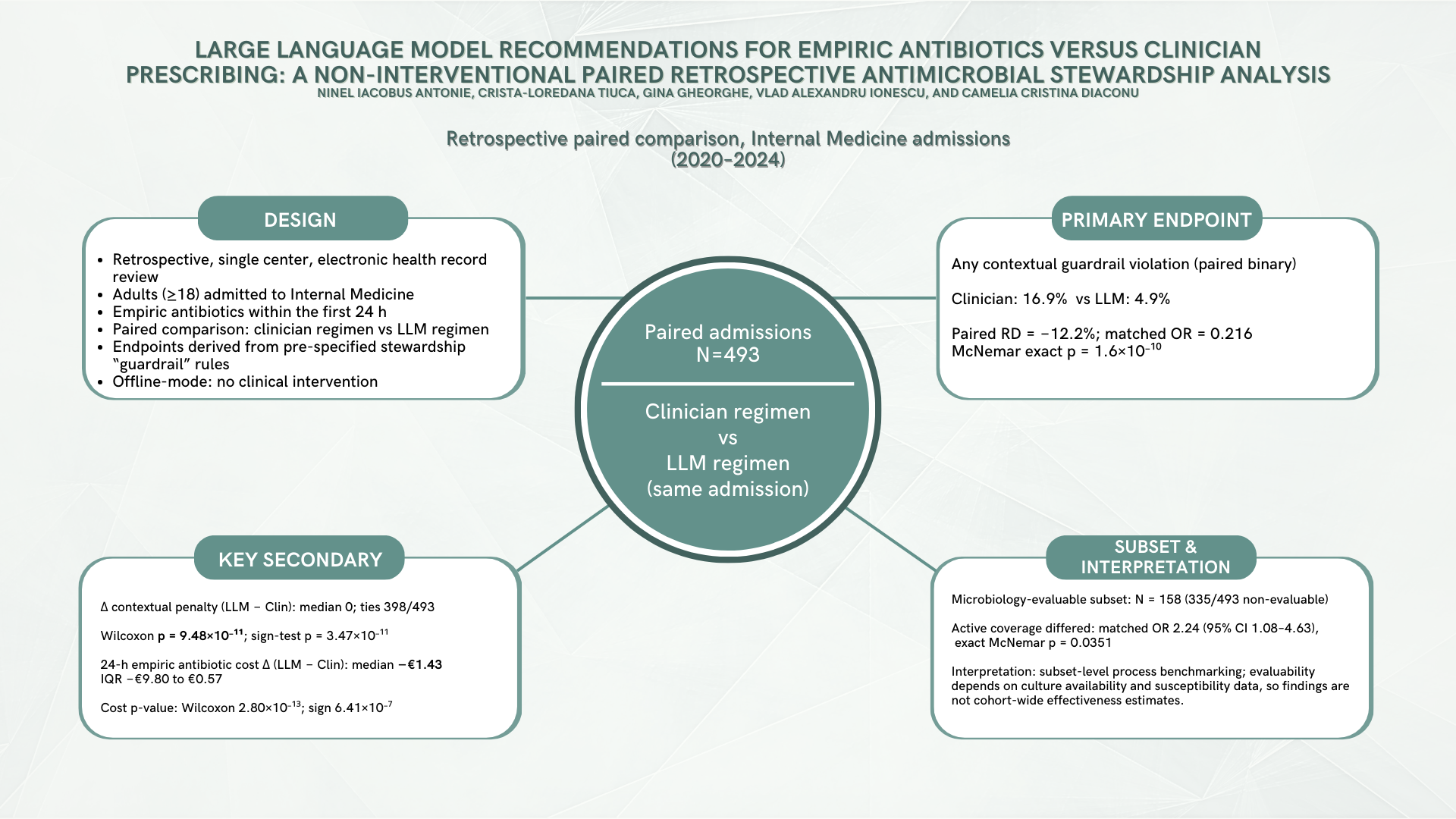
Background/Objectives: Antimicrobial resistance (AMR) remains a major global health threat, strengthening the case for antimicrobial stewardship that limits unnecessary broad-spectrum empiric therapy while preserving timely coverage in severe infection. Large language models (LLMs) are being explored for decision support, but require rigorous offline evaluation before any clinical implementation. Methods: Single-center retrospective paired evaluation at Clinical Emergency Hospital of Bucharest (Internal Medicine, 2020–2024). The unit of analysis was the admission (N = 493), with paired 24 h empiric regimens (clinician-prescribed vs post hoc LLM-recommended via OpenAI API; not visible to clinicians; no influence on care). Local laboratory-derived epidemiology was precomputed from microbiology exports and provided as structured prompt context to approximate information parity with clinicians’ implicit local ecology knowledge. Primary (prespecified) endpoint: any contextual guardrail violation (unjustified carbapenem/antipseudomonal/anti-MRSA under prespecified structured severity/MDR-risk rules), exact McNemar. Key secondary (prespecified): Δ contextual guardrail penalty (LLM − Clin), sign test and Wilcoxon signed-rank (ties reported). Ethics committee approval was obtained. Results: Guardrail violations occurred in 17.0% of clinician regimens vs 4.9% of LLM regimens (paired RD −12.2%; matched OR 0.216, 95% CI 0.127–0.367; McNemar exact p = 1.60 × 10⁻¹⁰). Δ penalty had median 0 with 398/493 ties; among non-ties, improvements (Δ < 0) exceeded adverse shifts (79 vs 16; sign-test p = 3.47 × 10⁻¹¹). Conclusions: In this offline, non-interventional paired evaluation, LLM regimens were associated with fewer prespecified contextual guardrail violations compared to clinician empiric regimens under a rule-based stewardship benchmarking framework. These endpoints strictly quantify concordance with stewardship constraints rather than patient outcomes, necessitating cautious interpretation of secondary and subset analyses. Ultimately, reproducible guardrail-based benchmarking may support subsequent prospective, safety-governed evaluations.
Keywords:
1. Introduction
2. Results
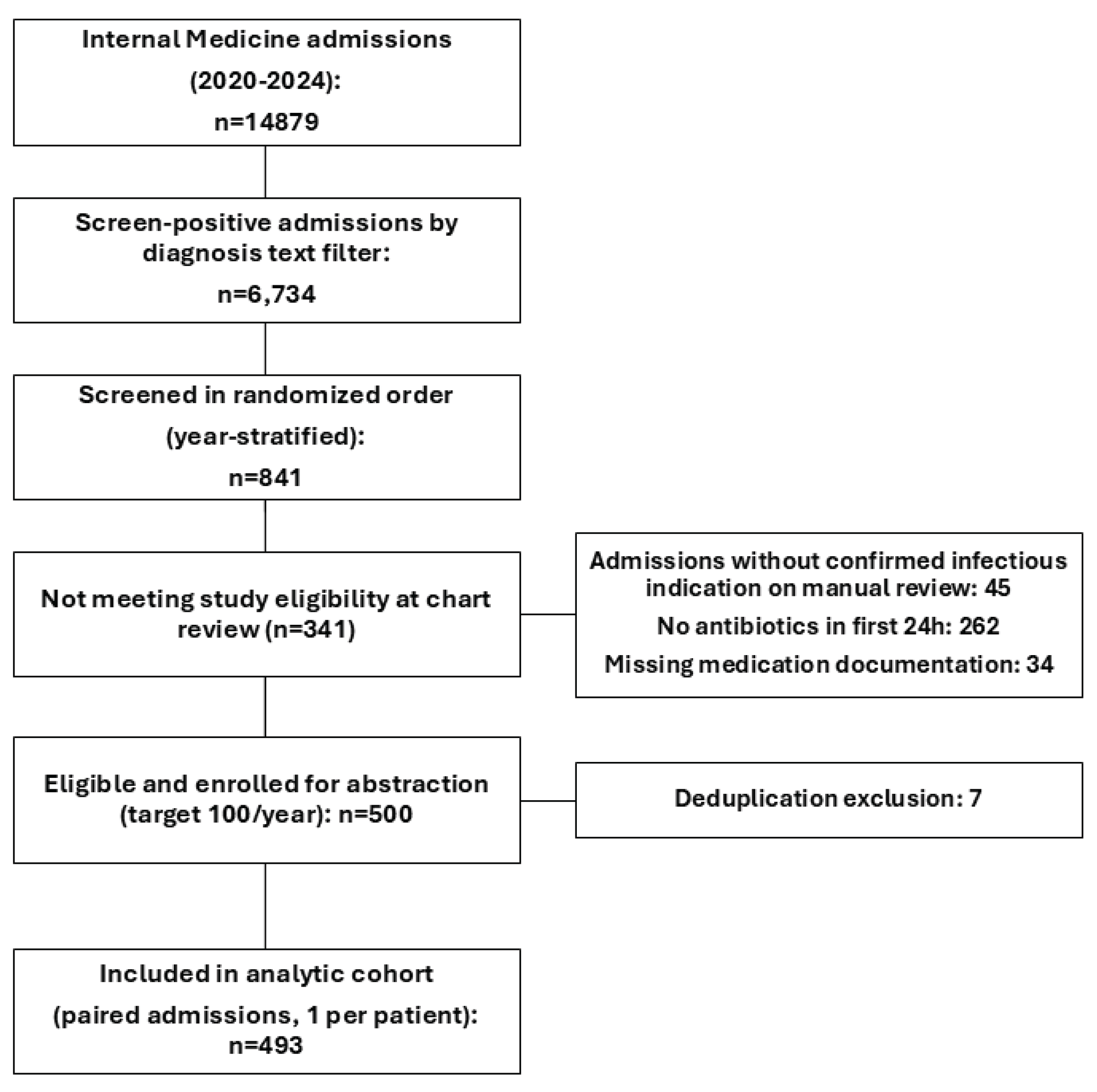
2.1. Cohort Flow
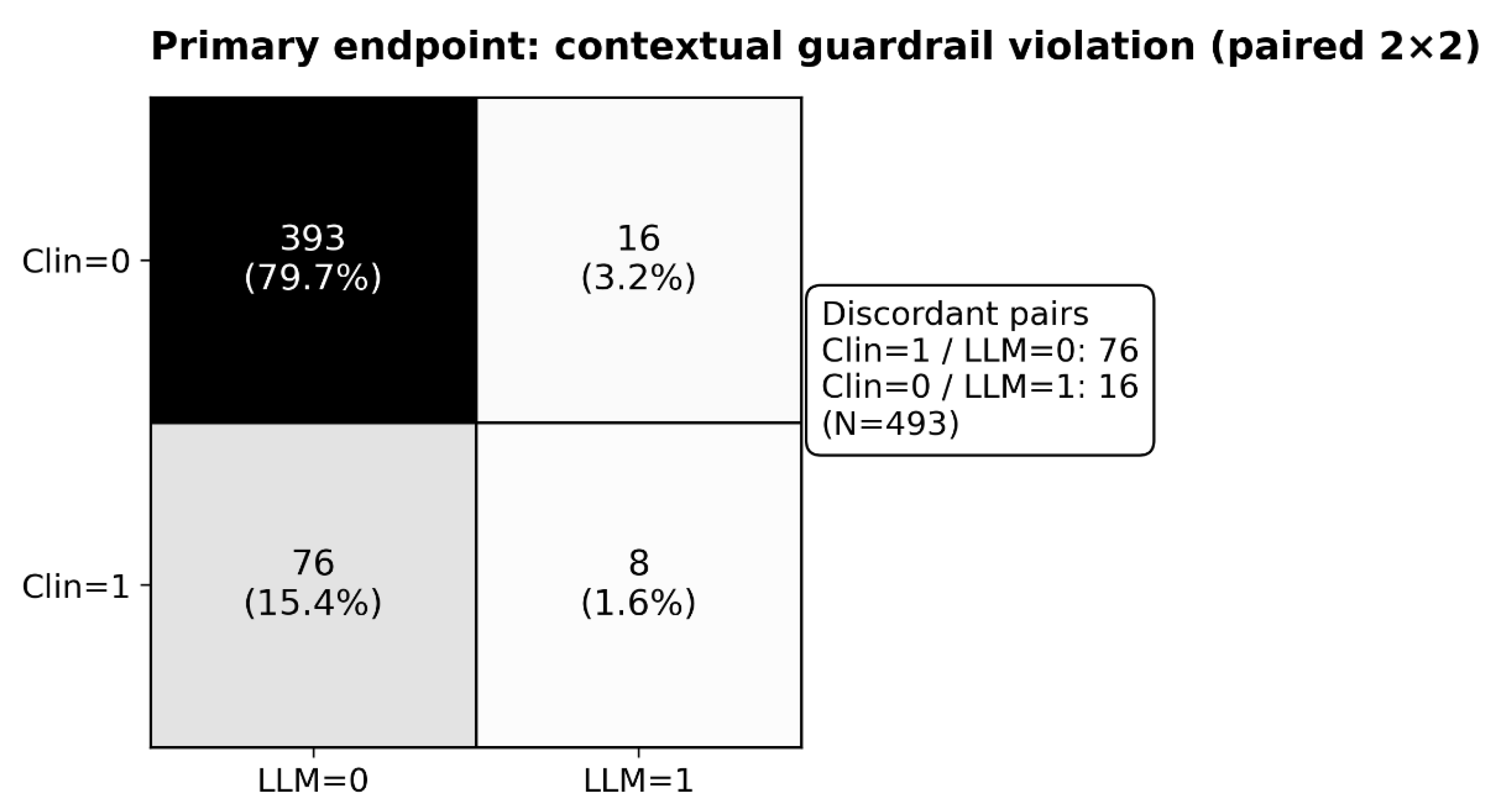
2.2. Primary Endpoint
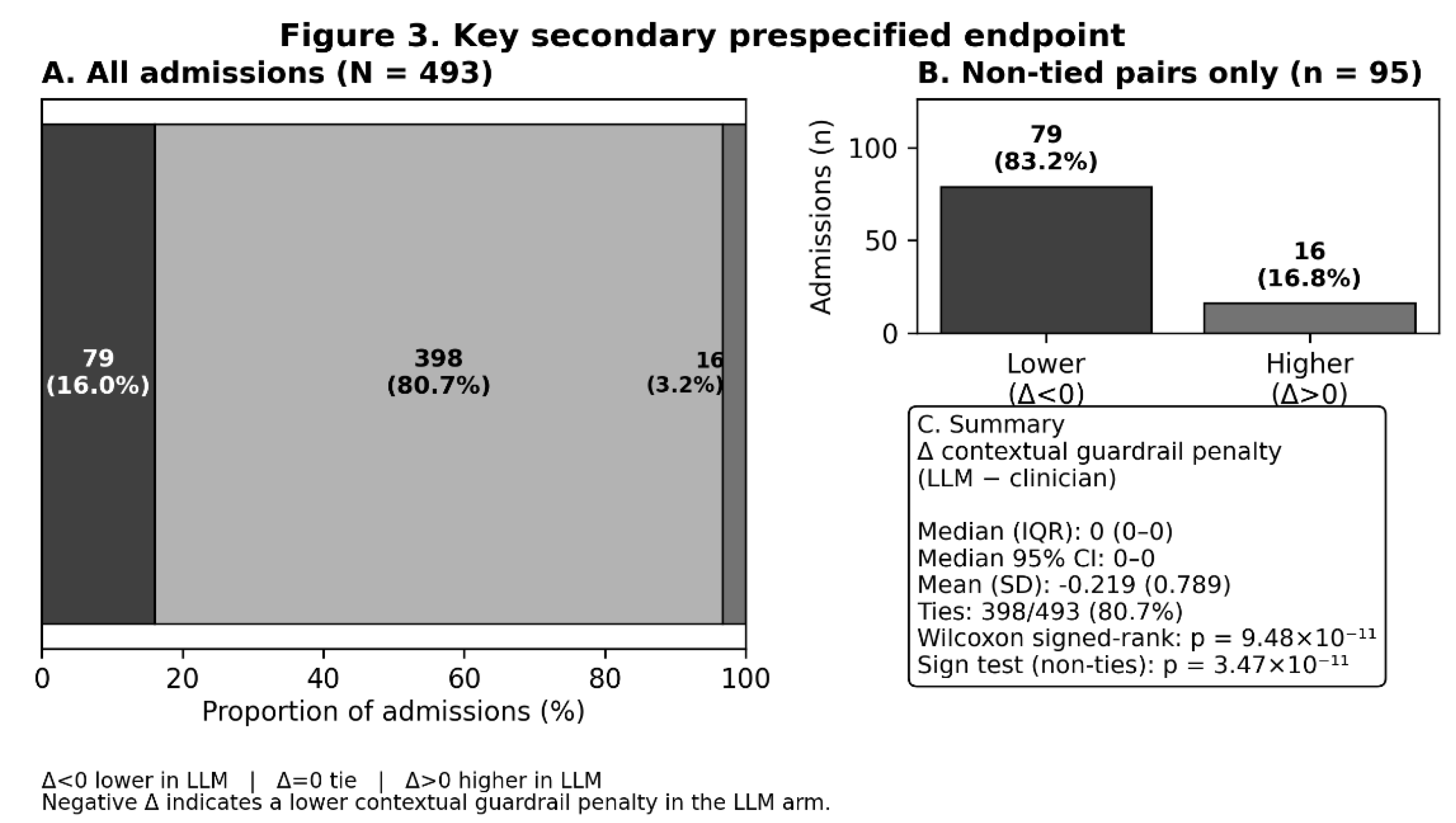
2.3. Key Secondary Endpoint
2.4. Secondary Endpoints (Multiplicity Caution)
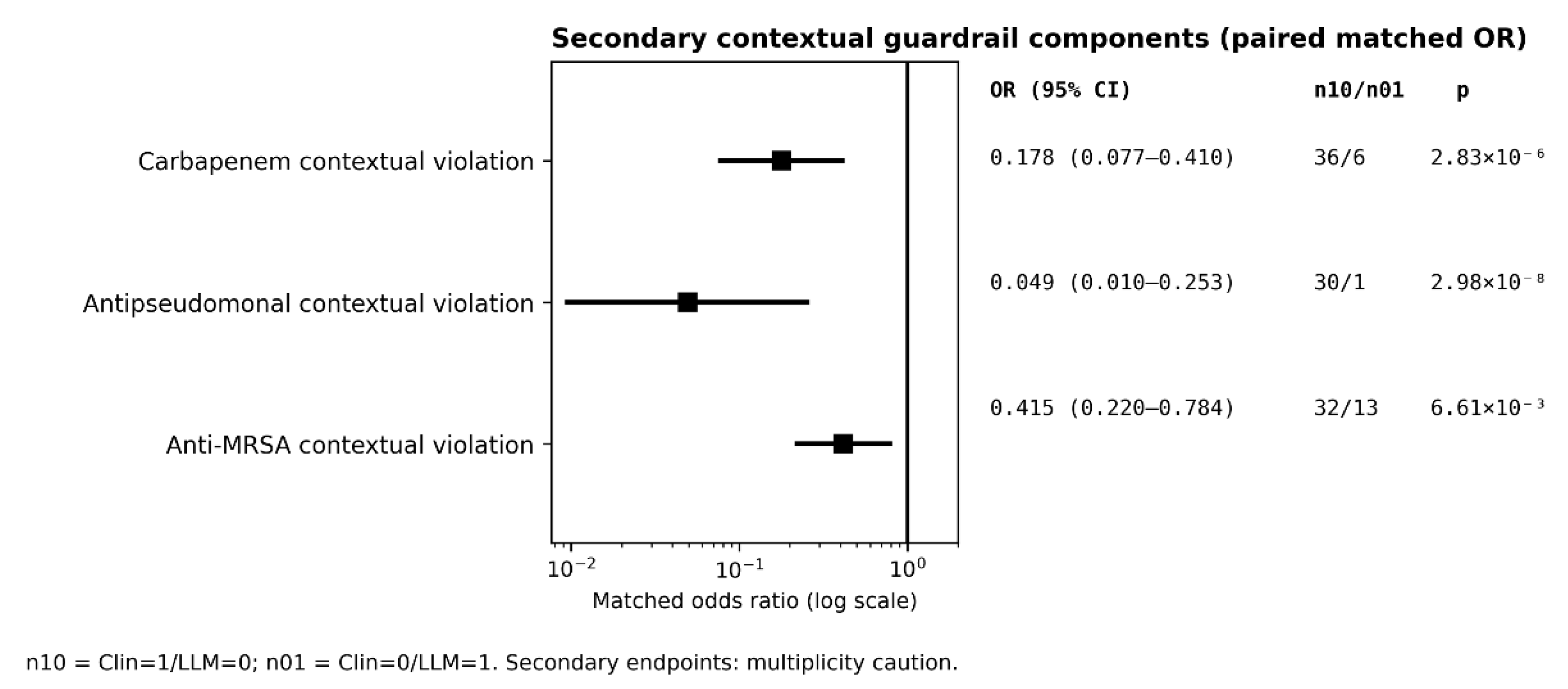
2.4.1. Contextual Guardrail Components
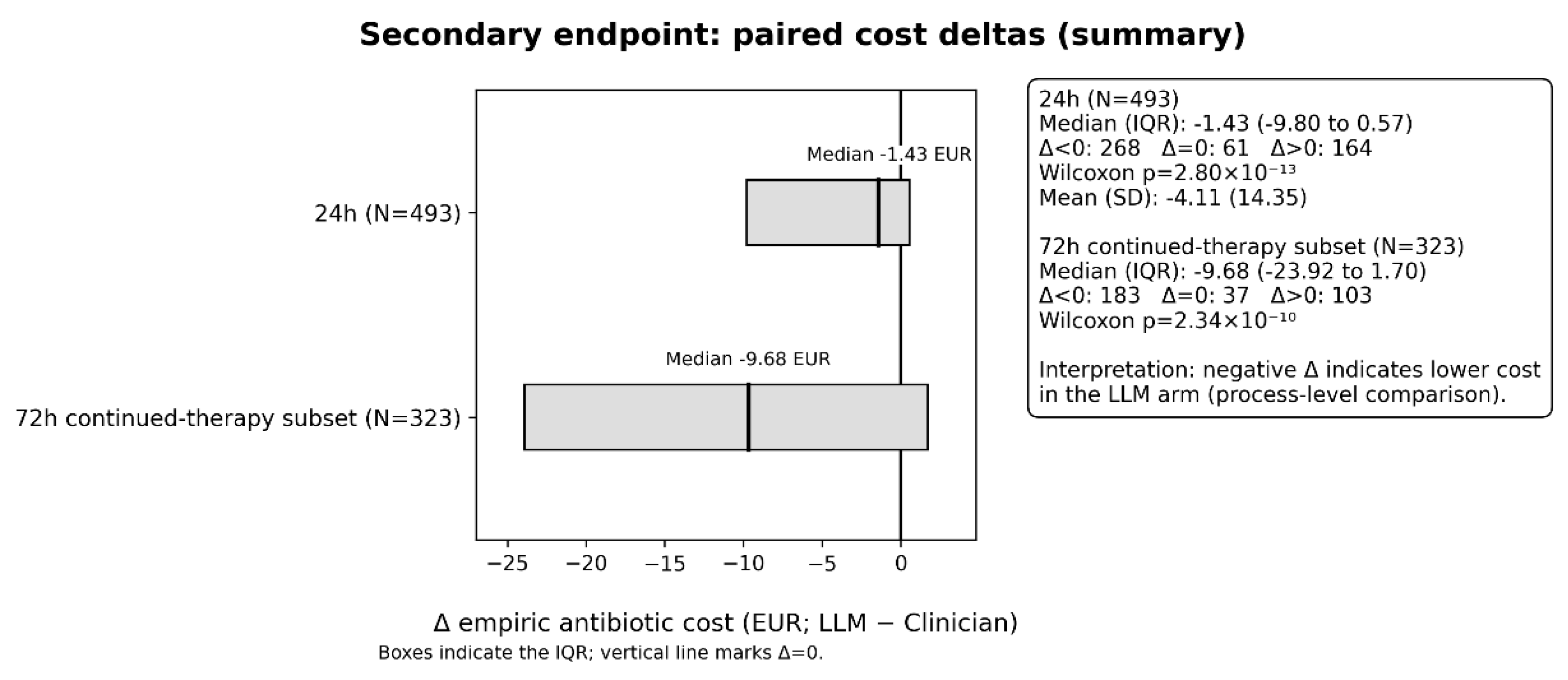
2.4.2. Costs (Secondary)
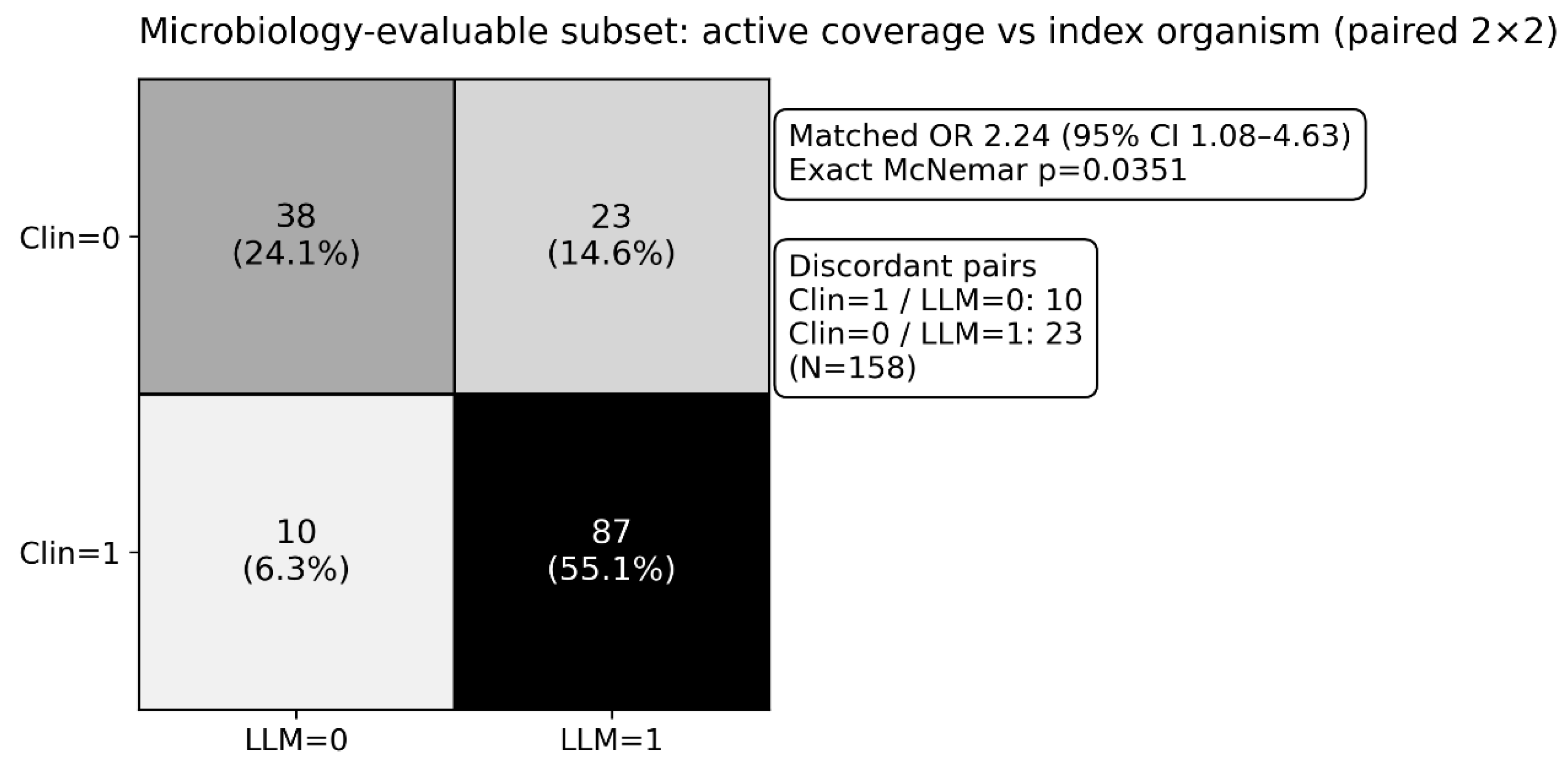
2.4.3. Microbiology-Evaluable Subset
2.4.4. Concordance (Supplementary Framing)
2.4.5. Supplementary QC Note (NO_ANTIBIOTIC)
2.4.6. Exploratory Modeling (Supplement Only)
3. Discussion
3.1. Principal Findings
3.2. Interpretation in Antimicrobial Stewardship Terms
3.3. Why Might the LLM Look Better on Guardrails?
3.4. Microbiology-Evaluable Subset: What It Means and What It Does Not Mean
3.5. Economic and Exposure Endpoints (Cost and DDD)
3.6. Comparison with Prior Work
3.7. Strengths
3.8. Limitations
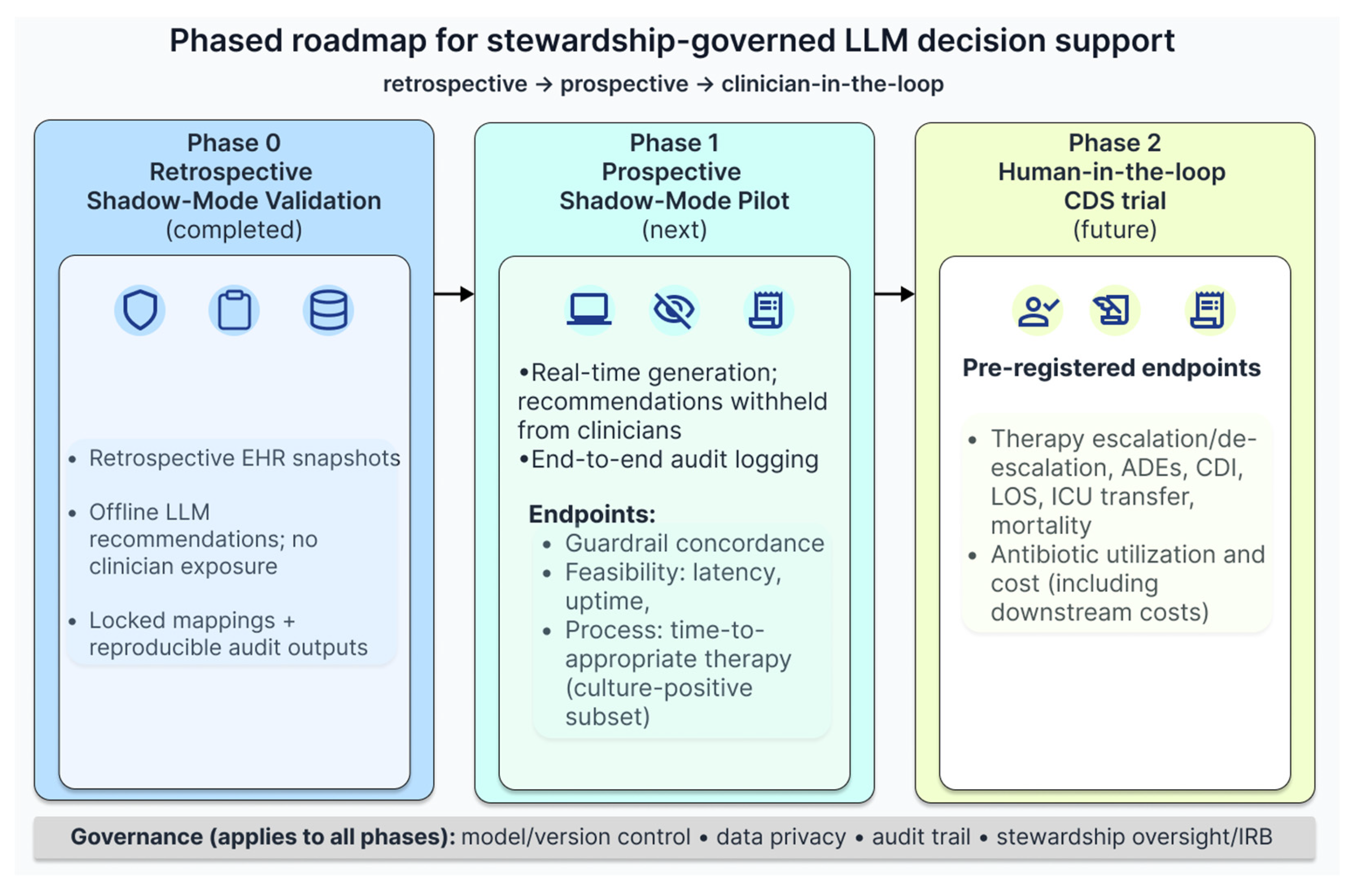
3.9. Implications and Next Steps
4. Materials and Methods
4.1. Study Design, Setting, and Timeframe
4.2. Participants: Source Population, Sampling, and Flow
4.2.1. Source Population Sampling Frame
4.2.2. Random Sampling and Manual Screening (Year-Stratified)
4.2.3. Eligibility Assessment and Reasons for Non-Inclusion
4.2.4. Bias and Representativeness
4.3. Data Sources and Manual EHR Abstraction
4.3.1. Data Sources and Measurement
4.4. Local Epidemiology Module (Laboratory Exports → Structured Prompt Context)
4.5. LLM Recommendation Generation (OpenAI API) and Reproducibility/Auditability
4.6. Outcomes (Contextual Stewardship Guardrails)
4.6.1. Regimen Definition (24 H)
4.6.2. Primary (Prespecified) Endpoint
4.6.3. Secondary/Supplementary Endpoints (Multiplicity Caution)
- empiric antibiotic cost differences at 24 h (N = 493) and at 72 hours only in admissions where clinician empiric therapy was continued for ≥3 days (N = 323);
- ΔDDD/24 h (N = 493) calculated using the WHO ATC/DDD methodology [51];
- microbiology-evaluable coverage analyses (paired; N = 158; descriptive due to selection);
- regimen concordance measures;
- AWaRe class distributions (supplementary; interpretation-sensitive) based on the WHO AWaRe framework [15];
- exploratory multivariable models (complete-case; N = 493), reported as exploratory only
4.7. Statistical Analysis
4.7.1. Primary Endpoint (Paired Binary)
4.7.2. Key Secondary Endpoint (Paired Integer Deltas with Many Ties)
4.7.3. Secondary Analyses and Exploratory Modeling
4.7.4. Software and Reproducibility
4.8. Ethics, Data Governance, and Availability Statements
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Reproducibility/Code availability statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Abbreviation | Definition |
| AI | Artificial intelligence |
| AMR | Antimicrobial resistance |
| AMS | Antimicrobial stewardship |
| APS | Antipseudomonal broad-spectrum β-lactam |
| ASP | Antimicrobial stewardship program |
| AWaRe | Access, Watch, Reserve (WHO antibiotic classification) |
| CDI | Clostridioides difficile infection |
| CI | Confidence interval |
| DDD | Defined daily dose |
| EHR | Electronic health record |
| ICU | Intensive care unit |
| IQR | Interquartile range |
| LLM | Large language model |
| MDR | Multidrug-resistant |
| MRSA | Methicillin-resistant Staphylococcus aureus |
| ESBL | Extended-spectrum beta-lactamase |
| CRE | Carbapenem-resistant Enterobacterales |
| VRE | Vancomycin-resistant enterococci |
| RD | Risk difference |
References
- Murray, C.J.L.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. The Lancet 2022, 399, 629–655. [CrossRef]
- O’Neill, J. (2016) Tackling Drug-Resistant Infections Globally Final Report and Recommendations. Review on Antimicrobial Resistance. Wellcome Trust and HM Government. Available online: https://amr-review.org/sites/default/files/160525_Final%20paper_with%20cover.pdf (accessed on 18 February 2026).
- Naghavi, M.; Vollset, S.E.; Ikuta, K.S.; Swetschinski, L.R.; Gray, A.P.; Wool, E.E.; Aguilar, G.R.; Mestrovic, T.; Smith, G.; Han, C.; et al. Global Burden of Bacterial Antimicrobial Resistance 1990–2021: A Systematic Analysis with Forecasts to 2050. The Lancet 2024, 404, 1199–1226. [CrossRef]
- Uddin, T.M.; Chakraborty, A.J.; Khusro, A.; Zidan, B.R.M.; Mitra, S.; Emran, T.B.; Dhama, K.; Ripon, Md.K.H.; Gajdács, M.; Sahibzada, M.U.K.; et al. Antibiotic Resistance in Microbes: History, Mechanisms, Therapeutic Strategies and Future Prospects. J. Infect. Public Health 2021, 14, 1750–1766. [CrossRef]
- Oliveira, M.; Antunes, W.; Mota, S.; Madureira-Carvalho, Á.; Dinis-Oliveira, R.J.; Silva, D.D. da An Overview of the Recent Advances in Antimicrobial Resistance. Microorganisms 2024, 12. [CrossRef]
- Kanthali, M.; Bhagwat, G.; Pathak, A.; Purohit, M. Antibiotic Stewardship through Clinical Data Digitization: Perceived Opportunities and Obstructions by Medical Doctors from Semi-Urban Setting in Central India. Front. Digit. Health 2025, 7. [CrossRef]
- Yoon, Y.K.; Kwon, K.T.; Jeong, S.J.; Moon, C.; Kim, B.; Kiem, S.; Kim, H.; Heo, E.; Kim, S.-W. Guidelines on Implementing Antimicrobial Stewardship Programs in Korea. Infect. Chemother. 2021, 53, 617–659. [CrossRef]
- Amin, S.U.; Guizani, M.; Hossain, M.S. Advances, Evaluation, and Explainability of Large Language Models in Healthcare: A Systematic Review. ACM Trans Multimed. Comput Commun Appl 2026, 22, 60:1-60:32. [CrossRef]
- Artsi, Y.; Sorin, V.; Glicksberg, B.S.; Korfiatis, P.; Freeman, R.; Nadkarni, G.N.; Klang, E. Challenges of Implementing LLMs in Clinical Practice: Perspectives. J. Clin. Med. 2025, 14. [CrossRef]
- Pinto, A.; Pennisi, F.; Ricciardi, G.E.; Signorelli, C.; Gianfredi, V. Evaluating the Impact of Artificial Intelligence in Antimicrobial Stewardship: A Comparative Meta-Analysis with Traditional Risk Scoring Systems. Infect. Dis. Now 2025, 55, 105090. [CrossRef]
- Antonie, N.I.; Gheorghe, G.; Ionescu, V.A.; Tiucă, L.-C.; Diaconu, C.C. The Role of ChatGPT and AI Chatbots in Optimizing Antibiotic Therapy: A Comprehensive Narrative Review. Antibiotics 2025, 14. [CrossRef]
- Al Mazrouei, N.; Ahmed Elnour, A.; Badi, S.; Alsulami, F.T.; Awadallah Mohamed Saeed, A.; Awad Al-Kubaisi, K.; Menon, V.; Yousif Khidir, I.; Ismail, M.; Osman Mahagoub, M.M.; et al. The Impact of Artificial Intelligence on the Prescribing, Selection, Resistance, and Stewardship of Antimicrobials: A Scoping Review. BMC Infect. Dis. 2026, 26, 222. [CrossRef]
- Giamarellou, H.; Galani, L.; Karavasilis, T.; Ioannidis, K.; Karaiskos, I. Antimicrobial Stewardship in the Hospital Setting: A Narrative Review. Antibiotics 2023, 12. [CrossRef]
- Rapti, V.; Poulakou, G.; Mousouli, A.; Kakasis, A.; Pagoni, S.; Pechlivanidou, E.; Masgala, A.; Sympardi, S.; Apostolopoulos, V.; Giannopoulos, C.; et al. Assessment of De-Escalation of Empirical Antimicrobial Therapy in Medical Wards with Recognized Prevalence of Multi-Drug-Resistant Pathogens: A Multicenter Prospective Cohort Study in Non-ICU Patients with Microbiologically Documented Infection. Antibiotics 2024, 13. [CrossRef]
- WHO AWaRe System for Antimicrobial Stewardship Available online: https://www.who.int/teams/surveillance-prevention-control-AMR/control-and-response-strategies/AWaRe (accessed on 18 February 2026).
- Karaiskos, I.; Giamarellou, H. Carbapenem-Sparing Strategies for ESBL Producers: When and How. Antibiotics 2020, 9. [CrossRef]
- Parente, D.M.; Cunha, C.B.; Mylonakis, E.; Timbrook, T.T. The Clinical Utility of Methicillin-Resistant Staphylococcus Aureus (MRSA) Nasal Screening to Rule Out MRSA Pneumonia: A Diagnostic Meta-Analysis With Antimicrobial Stewardship Implications. Clin. Infect. Dis. 2018, 67, 1–7. [CrossRef]
- Metlay, J.P.; Waterer, G.W.; Long, A.C.; Anzueto, A.; Brozek, J.; Crothers, K.; Cooley, L.A.; Dean, N.C.; Fine, M.J.; Flanders, S.A.; et al. Diagnosis and Treatment of Adults with Community-Acquired Pneumonia. An Official Clinical Practice Guideline of the American Thoracic Society and Infectious Diseases Society of America. Am. J. Respir. Crit. Care Med. 2019, 200, e45–e67. [CrossRef]
- Trautner, B.W.; Cortés-Penfield, N.W.; Gupta, K.; Hirsch, E.B.; Horstman, M.; Moran, G.J.; Colgan, R.; O’Horo, J.C.; Ashraf, M.S.; Connolly, S.; et al. Clinical Practice Guideline by Infectious Diseases Society of America (IDSA): 2025 Guideline on Management and Treatment of Complicated Urinary Tract Infections: Selection of Antibiotic Therapy for Complicated UTI. Clin. Infect. Dis. 2025, ciaf460. [CrossRef]
- Rawson, T.M.; Moore, L.S.P.; Hernandez, B.; Charani, E.; Castro-Sanchez, E.; Herrero, P.; Hayhoe, B.; Hope, W.; Georgiou, P.; Holmes, A.H. A Systematic Review of Clinical Decision Support Systems for Antimicrobial Management: Are We Failing to Investigate These Interventions Appropriately? Clin. Microbiol. Infect. 2017, 23, 524–532. [CrossRef]
- Dhaliwal, M.; Elligsen, M.; Lam, P.W.; Daneman, N. Weighted-Incidence Syndromic Combination Antibiogram (WISCA) to Guide Antibiotic Regimens for Empiric Treatment of Prosthetic Joint Infections: A Retrospective Cohort Study. CMI Commun. 2026, 3, 105170. [CrossRef]
- Cook, A.; Sharland, M.; Yau, Y.; Bielicki, J. Improving Empiric Antibiotic Prescribing in Pediatric Bloodstream Infections: A Potential Application of Weighted-Incidence Syndromic Combination Antibiograms (WISCA). Expert Rev. Anti Infect. Ther. 2022, 20, 445–456. [CrossRef]
- Hebert, C.; Ridgway, J.; Vekhter, B.; Brown, E.C.; Weber, S.G.; Robicsek, A. Demonstration of the Weighted-Incidence Syndromic Combination Antibiogram: An Empiric Prescribing Decision Aid. Infect. Control Hosp. Epidemiol. 2012, 33, 381–388. [CrossRef]
- Vasey, B.; Nagendran, M.; Campbell, B.; Clifton, D.A.; Collins, G.S.; Denaxas, S.; Denniston, A.K.; Faes, L.; Geerts, B.; Ibrahim, M.; et al. Reporting Guideline for the Early-Stage Clinical Evaluation of Decision Support Systems Driven by Artificial Intelligence: DECIDE-AI. Nat. Med. 2022, 28, 924–933. [CrossRef]
- Gallifant, J.; Afshar, M.; Ameen, S.; Aphinyanaphongs, Y.; Chen, S.; Cacciamani, G.; Demner-Fushman, D.; Dligach, D.; Daneshjou, R.; Fernandes, C.; et al. The TRIPOD-LLM Reporting Guideline for Studies Using Large Language Models. Nat. Med. 2025, 31, 60–69. [CrossRef]
- Asgari, E.; Montaña-Brown, N.; Dubois, M.; Khalil, S.; Balloch, J.; Yeung, J.A.; Pimenta, D. A Framework to Assess Clinical Safety and Hallucination Rates of LLMs for Medical Text Summarisation. Npj Digit. Med. 2025, 8, 274. [CrossRef]
- Barlam, T.F.; Cosgrove, S.E.; Abbo, L.M.; MacDougall, C.; Schuetz, A.N.; Septimus, E.J.; Srinivasan, A.; Dellit, T.H.; Falck-Ytter, Y.T.; Fishman, N.O.; et al. Implementing an Antibiotic Stewardship Program: Guidelines by the Infectious Diseases Society of America and the Society for Healthcare Epidemiology of America. Clin. Infect. Dis. 2016, 62, e51–e77. [CrossRef]
- Schoffelen, T.; Papan, C.; Carrara, E.; Eljaaly, K.; Paul, M.; Keuleyan, E.; Martin Quirós, A.; Peiffer-Smadja, N.; Palos, C.; May, L.; et al. European Society of Clinical Microbiology and Infectious Diseases Guidelines for Antimicrobial Stewardship in Emergency Departments (Endorsed by European Association of Hospital Pharmacists). Clin. Microbiol. Infect. 2024, 30, 1384–1407. [CrossRef]
- Baur, D.; Gladstone, B.P.; Burkert, F.; Carrara, E.; Foschi, F.; Döbele, S.; Tacconelli, E. Effect of Antibiotic Stewardship on the Incidence of Infection and Colonisation with Antibiotic-Resistant Bacteria and Clostridium Difficile Infection: A Systematic Review and Meta-Analysis. Lancet Infect. Dis. 2017, 17, 990–1001. [CrossRef]
- Ray, M.J.; Strnad, L.C.; Tucker, K.J.; Furuno, J.P.; Lofgren, E.T.; McCracken, C.M.; Park, H.; Gerber, J.S.; McGregor, J.C. Influence of Antibiotic Exposure Intensity on the Risk of Clostridioides Difficile Infection. Clin. Infect. Dis. 2024, 79, 1129–1135. [CrossRef]
- Bassetti, M.; Rello, J.; Blasi, F.; Goossens, H.; Sotgiu, G.; Tavoschi, L.; Zasowski, E.J.; Arber, M.R.; McCool, R.; Patterson, J.V.; et al. Systematic Review of the Impact of Appropriate versus Inappropriate Initial Antibiotic Therapy on Outcomes of Patients with Severe Bacterial Infections. Int. J. Antimicrob. Agents 2020, 56, 106184. [CrossRef]
- Paul, M.; Shani, V.; Muchtar, E.; Kariv, G.; Robenshtok, E.; Leibovici, L. Systematic Review and Meta-Analysis of the Efficacy of Appropriate Empiric Antibiotic Therapy for Sepsis. Antimicrob. Agents Chemother. 2010, 54, 4851–4863. [CrossRef]
- Bosetti, D.; Grant, R.; Catho, G. Computerized Decision Support for Antimicrobial Prescribing: What Every Antibiotic Steward Should Know. Antimicrob. Steward. Healthc. Epidemiol. 2025, 5, e210. [CrossRef]
- Hatton, C.; Quarton, S.; Livesey, A.; Alenazi, B.A.; Jeff, C.; Sapey, E. Impact of Clinical Decision Support Software on Empirical Antibiotic Prescribing and Patient Outcomes: A Systematic Review and Meta-Analysis. BMJ Open 2025, 15, e099100. [CrossRef]
- Fitzpatrick, F.; Tarrant, C.; Hamilton, V.; Kiernan, F.M.; Jenkins, D.; Krockow, E.M. Sepsis and Antimicrobial Stewardship: Two Sides of the Same Coin. BMJ Qual. Saf. 2019, 28, 758–761. [CrossRef]
- Mcleod, M.; Campbell, A.; Hayhoe, B.; Borek, A.J.; Tonkin-Crine, S.; Moore, M.V.; Butler, C.C.; Walker, A.S.; Holmes, A.; Wong, G. How, Why and When Are Delayed (Back-up) Antibiotic Prescriptions Used in Primary Care? A Realist Review Integrating Concepts of Uncertainty in Healthcare. BMC Public Health 2024, 24, 2820. [CrossRef]
- Richards, A.R.; Linder, J.A. Behavioral Economics and Ambulatory Antibiotic Stewardship: A Narrative Review. Clin. Ther. 2021, 43, 1654–1667. [CrossRef]
- Schisterman, E.F.; Cole, S.R.; Platt, R.W. Overadjustment Bias and Unnecessary Adjustment in Epidemiologic Studies. Epidemiol. Camb. Mass 2009, 20, 488–495. [CrossRef]
- Hernán, M.A.; Hernández-Díaz, S.; Robins, J.M. A Structural Approach to Selection Bias. Epidemiology 2004, 15, 615. [CrossRef]
- Nathwani, D.; Varghese, D.; Stephens, J.; Ansari, W.; Martin, S.; Charbonneau, C. Value of Hospital Antimicrobial Stewardship Programs [ASPs]: A Systematic Review. Antimicrob. Resist. Infect. Control 2019, 8, 35. [CrossRef]
- Malone, D.C.; Armstrong, E.P.; Gratie, D.; Pham, S.V.; Amin, A. A Systematic Review of Real-World Healthcare Resource Use and Costs of Clostridioides Difficile Infections. Antimicrob. Steward. Healthc. Epidemiol. 2023, 3, e17. [CrossRef]
- Giacobbe, D.R.; Marelli, C.; La Manna, B.; Padua, D.; Malva, A.; Guastavino, S.; Signori, A.; Mora, S.; Rosso, N.; Campi, C.; et al. Advantages and Limitations of Large Language Models for Antibiotic Prescribing and Antimicrobial Stewardship. Npj Antimicrob. Resist. 2025, 3, 14. [CrossRef]
- Ngoc Nguyen, O.; Amin, D.; Bennett, J.; Hetlevik, Ø.; Malik, S.; Tout, A.; Vornhagen, H.; Vellinga, A. GP or ChatGPT? Ability of Large Language Models (LLMs) to Support General Practitioners When Prescribing Antibiotics. J. Antimicrob. Chemother. 2025, 80, 1324–1330. [CrossRef]
- Williams, C.Y.K.; Miao, B.Y.; Kornblith, A.E.; Butte, A.J. Evaluating the Use of Large Language Models to Provide Clinical Recommendations in the Emergency Department. Nat. Commun. 2024, 15, 8236. [CrossRef]
- Vo, T.; Dahal, K.; Klepser, M.; Pontefract, B.; Caniff, K.E.; Sohn, M. Evaluation of Large Language Models for Antimicrobial Classification: Implications for Antimicrobial Stewardship Programs. Antimicrob. Steward. Healthc. Epidemiol. 2025, 5, e324. [CrossRef]
- von Elm, E.; Altman, D.G.; Egger, M.; Pocock, S.J.; Gøtzsche, P.C.; Vandenbroucke, J.P. The Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) Statement: Guidelines for Reporting Observational Studies. The Lancet 2007, 370, 1453–1457. [CrossRef]
- Holm, S. A Simple Sequentially Rejective Multiple Test Procedure. Scand. J. Stat. 1979, 6, 65–70.
- Cole, S.R.; Platt, R.W.; Schisterman, E.F.; Chu, H.; Westreich, D.; Richardson, D.; Poole, C. Illustrating Bias Due to Conditioning on a Collider. Int. J. Epidemiol. 2010, 39, 417–420. [CrossRef]
- Randhawa, V.; Sarwar, S.; Walker, S.; Elligsen, M.; Palmay, L.; Daneman, N. Weighted-Incidence Syndromic Combination Antibiograms to Guide Empiric Treatment of Critical Care Infections: A Retrospective Cohort Study. Crit. Care 2014, 18, R112. [CrossRef]
- API Platform Available online: https://openai.com/api/ (accessed on 18 February 2026).
- ATCDDD - Guidelines Available online: https://atcddd.fhi.no/atc_ddd_index_and_guidelines/guidelines/ (accessed on 18 February 2026).
- Dixon, W.J.; Mood, A.M. The Statistical Sign Test. J. Am. Stat. Assoc. 1946, 41, 557–566. [CrossRef]
- Efron, B.; Tibshirani, R. An Introduction to the Bootstrap; Monographs on statistics and applied probability; Nachdr.; Chapman & Hall: Boca Raton, Fla., 1998; ISBN 978-0-412-04231-7.
- Regulation (EU) 2016/679 of the European Parliament and of the Council of 27 April 2016 on the protection of natural persons with regard to the processing of personal data and on the free movement of such data, and repealing Directive 95/46/EC (General Data Protection Regulation). Off. J. Eur. Union 2016, L 119, 1–88.
- Data Controls in the OpenAI Platform Available online: https://developers.openai.com/api/docs/guides/your-data/ (accessed on 20 February 2026).
| Characteristic | Total (N = 493) |
|---|---|
| Demographics | |
| Age, years, median (IQR) | 72 (64–82) |
| Female sex, n (%) | 250 (50.7%) |
| Length of stay, days, median (IQR) | 9 (5–15) |
| Study period | |
| Cohort year 2020, n (%) | 98 (19.9%) |
| Cohort year 2021, n (%) | 100 (20.3%) |
| Cohort year 2022, n (%) | 100 (20.3%) |
| Cohort year 2023, n (%) | 98 (19.9%) |
| Cohort year 2024, n (%) | 97 (19.7%) |
| Acquisition setting | |
| Community-onset, n (%) | 459 (93.1%) |
| Healthcare-associated, n (%) | 34 (6.9%) |
| Index syndrome (top categories) | |
| Community-acquired pneumonia, n (%) | 311 (63.1%) |
| Urinary tract infection - unspecified site, n (%) | 56 (11.4%) |
| Bloodstream infection / sepsis, n (%) | 50 (10.1%) |
| COPD infectious exacerbation, n (%) | 32 (6.5%) |
| Urinary tract infection – pyelonephritis, n (%) | 29 (5.9%) |
| Skin and soft-tissue infection, n (%) | 7 (1.4%) |
| Other syndromes, n (%) | 8 (1.6%) |
| Severity / support within 24 h | |
| Sepsis documented, n (%) | 183 (37.1%) |
| Septic shock documented, n (%) | 129 (26.2%) |
| Respiratory failure documented, n (%) | 374 (75.9%) |
| Mechanical ventilation, n (%) | 147 (29.8%) |
| Vasopressors, n (%) | 127 (25.8%) |
| ICU transfer, n (%) | 141 (28.6%) |
| Prior exposure / MDR-risk proxies | |
| Antibiotics in prior 90 days, n (%) | 75 (15.2%) |
| Hospitalization in prior 90 days, n (%) | 70 (14.2%) |
| Long-term care facility resident, n (%) | 32 (6.5%) |
| Prior MRSA colonization/infection, n (%) | 6 (1.2%) |
| Prior ESBL/CRE/VRE history, n (%) | 22 (4.5%) |
| Home antibiotics before admission: Yes, n (%) | 65 (13.2%) |
| Home antibiotics before admission: No, n (%) | 428 (86.8%) |
| Comorbidities | |
| Hypertension, n (%) | 374 (75.9%) |
| Diabetes mellitus, n (%) | 160 (32.5%) |
| COPD, n (%) | 94 (19.1%) |
| Chronic kidney disease, n (%) | 137 (27.8%) |
| Congestive heart failure, n (%) | 241 (48.9%) |
| Atrial fibrillation, n (%) | 180 (36.5%) |
| Prior stroke, n (%) | 98 (19.9%) |
| Cirrhosis, n (%) | 27 (5.5%) |
| Malignancy, n (%) | 90 (18.3%) |
| Immunosuppression, n (%) | 29 (5.9%) |
| Panel | Endpoint | N | Clinician | LLM | Discordant pairs | Effect | P value | Median 95% CI (bootstrap) |
|---|---|---|---|---|---|---|---|---|
| A (prespecified) | Primary: Any contextual guardrail violation (composite) | 493 | 84/493 (17.0%) | 24/493 (4.9%) | Clin = 1/LLM = 0: 76; Clin = 0/LLM = 1: 16 | Matched OR 0.216 (95% CI 0.127–0.367) RD (LLM−Clin) −0.122 |
p = 1.60 × 10⁻¹⁰ | |
| B (prespecified) | Key secondary: Δ contextual guardrail penalty (LLM−Clin) | 493 | — | — | Δ < 0 (lower): 79; Δ > 0 (higher): 16; Δ = 0: 398 | Median 0 (IQR 0–0) Mean −0.219 (SD 0.789) |
Wilcoxon p = 9.48 × 10⁻¹¹; Sign p = 3.47 × 10⁻¹¹ |
Median 0 (IQR 0–0); 95% CI 0–0 |
| C (Secondary; multiplicity caution) | Any broad-spectrum class used (carb OR APS OR anti-MRSA) | 493 | 289/493 (58.6%) | 195/493 (39.6%) | Clin = 1/LLM = 0: 143; Clin = 0/LLM = 1: 49 | Matched OR 0.345 (95% CI 0.250–0.477) RD (LLM−Clin) −0.191 |
p = 7.26 × 10⁻¹² | |
| C (Secondary; multiplicity caution) | Carbapenem contextual violation | 493 | 39/493 (7.9%) | 9/493 (1.8%) | Clin = 1/LLM = 0: 36; Clin = 0/LLM = 1: 6 | Matched OR 0.178 (95% CI 0.077–0.410) RD (LLM−Clin) −0.061 |
p = 2.83 × 10⁻⁶ | |
| C (Secondary; multiplicity caution) | Antipseudomonal contextual violation | 493 | 31/493 (6.3%) | 2/493 (0.4%) | Clin = 1/LLM = 0: 30; Clin = 0/LLM = 1: 1 | Matched OR 0.049 (95% CI 0.010–0.253) RD (LLM−Clin) −0.059 |
p = 2.98 × 10⁻⁸ | |
| C (Secondary; multiplicity caution) | Anti-MRSA contextual violation | 493 | 36/493 (7.3%) | 17/493 (3.4%) | Clin = 1/LLM = 0: 32; Clin = 0/LLM = 1: 13 | Matched OR 0.415 (95% CI 0.220–0.784) RD (LLM−Clin) −0.039 |
p = 6.61 × 10⁻³ | |
| C (Secondary; multiplicity caution) | Δ empiric antibiotic cost, 24 h (EUR; LLM−Clin) | 493 | — | — | Δ < 0 (lower): 268; Δ > 0 (higher): 164; Δ = 0: 61 | Median -1.43 (EUR) (IQR -9.80–0.57) Mean −4.11 (EUR) (SD 14.35) |
Wilcoxon p = 2.80 × 10⁻¹³; Sign p = 6.41 × 10⁻⁷ |
Median 95% CI (bootstrap): −4.09 to -0.07 |
| Metric | n | % |
|---|---|---|
| Exact identical regimen set | 57 | 11.6% |
| Same primary agent (first-listed antibiotic) | 137 | 27.8% |
| Any overlap (≥1 shared agent) | 148 | 30.0% |
| No overlap (0 shared agents) | 345 | 70.0% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




