Submitted:
09 March 2026
Posted:
09 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Direct Suppression of HIV-1 Transcriptional Activity
2.1. Tat-Mediated Control of HIV-1 Transcription as a Therapeutic Target for Block-and-Lock Strategies
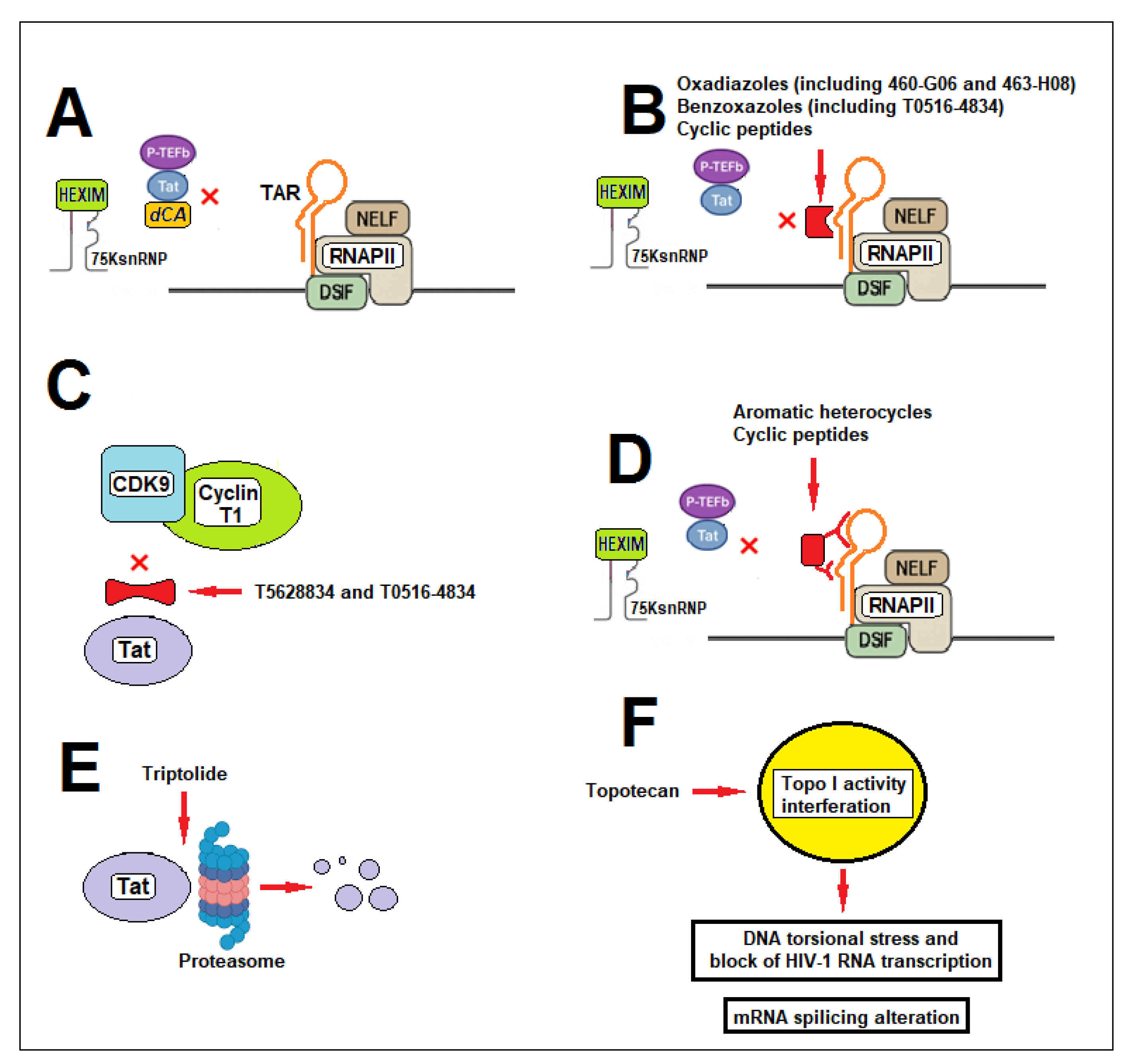
2.2. Didehydro-Cortistatin A: Mechanism and Preclinical Efficacy
2.3. dCA Pharmacokinetics, Bioavailability, and Safety
2.4. Novel Tat Inhibitors and Future Perspectives
2.5. Triptolide-Mediated Tat Degradation as a Block-and-Lock Strategy for HIV-1
2.6. Camptothecin Analogs
2.7. CRISPR–Cas Technologies as Block-and-Lock Strategies for HIV-1
3. Post-Transcriptional and Gene-Silencing Approaches
3.1. Splicing Inhibitors
3.2. RNA Interference Technologies as Block-and-Lock Strategies for HIV-1
4. Cellular Transcription Factor Modulators as Block-and-Lock Agents
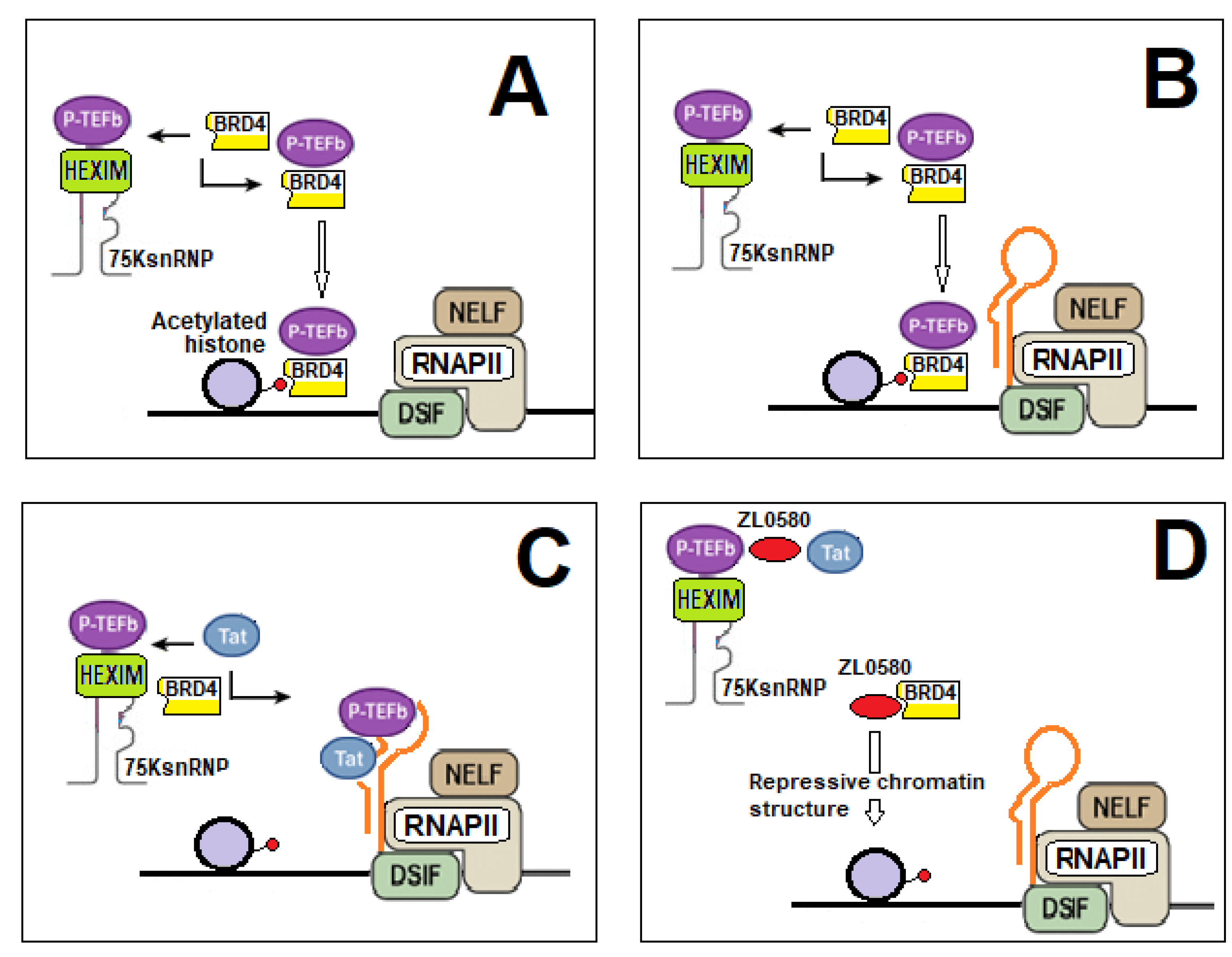
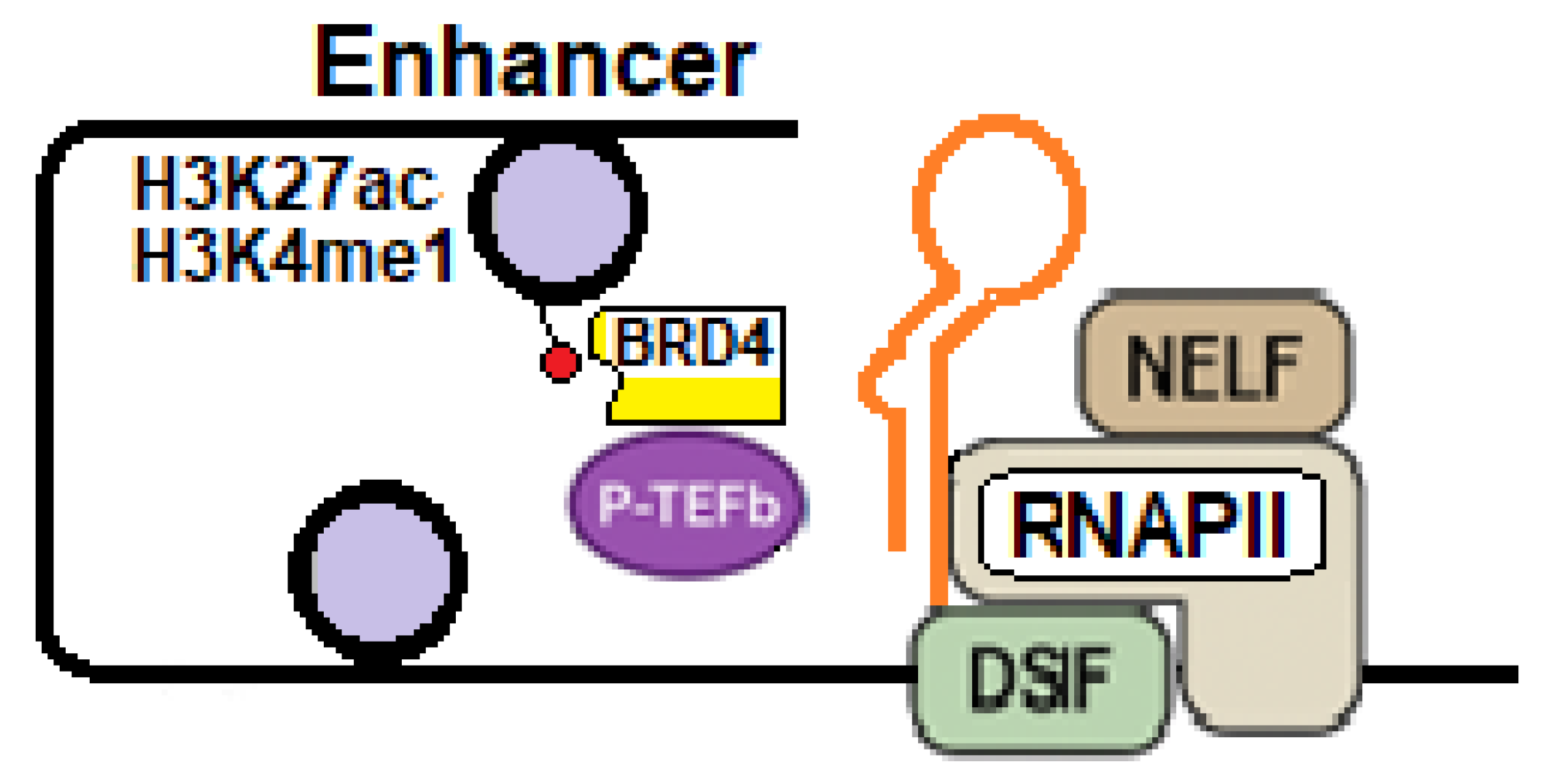
4.1. BDR4 Functions Under Physiologic Conditions and in HIV-1 Infection
4.2. Pharmacological Modulation of BRD4
4.3. NF-κB Inhibitors as Block-and-Lock Agents in HIV-1 Infection
5. Kinase Inhibitors in the Block-and-Lock Strategy
5.1. PI3K–AKT–mTOR Pathway Modulators as Latency-Promoting Agents in HIV-1 Block-and-Lock Strategies
5.2. Aurora Kinase and PAK1/2 Inhibitors
5.3. Protein Kinase C (PKC) Inhibitors
5.4. CDK9 Inhibitors
5.5. CDK8/19 Inhibitors
5.6. CDK7 Inhibitors
5.7. SR Kinase Inhibitors (CLK1/2, SRPK1)
6. Epigenetic and Chromatin-Based Silencing Approaches Within the Block-and-Lock Strategy
6.1. Direct Epigenetic Enzyme Inhibitors
6.2. Chromatin-Associated Transcriptional Modulators
6.3. Integration Site Modulators
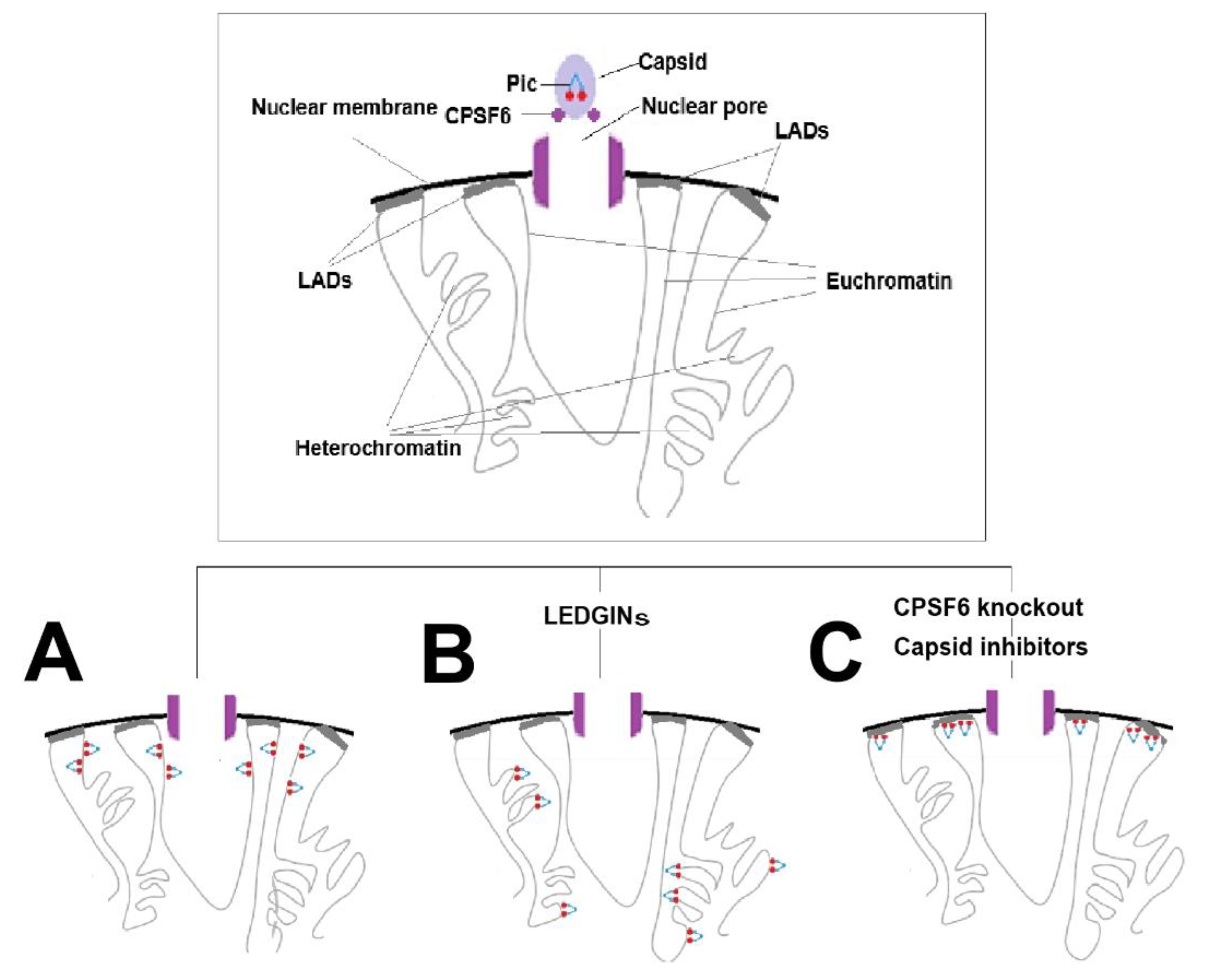
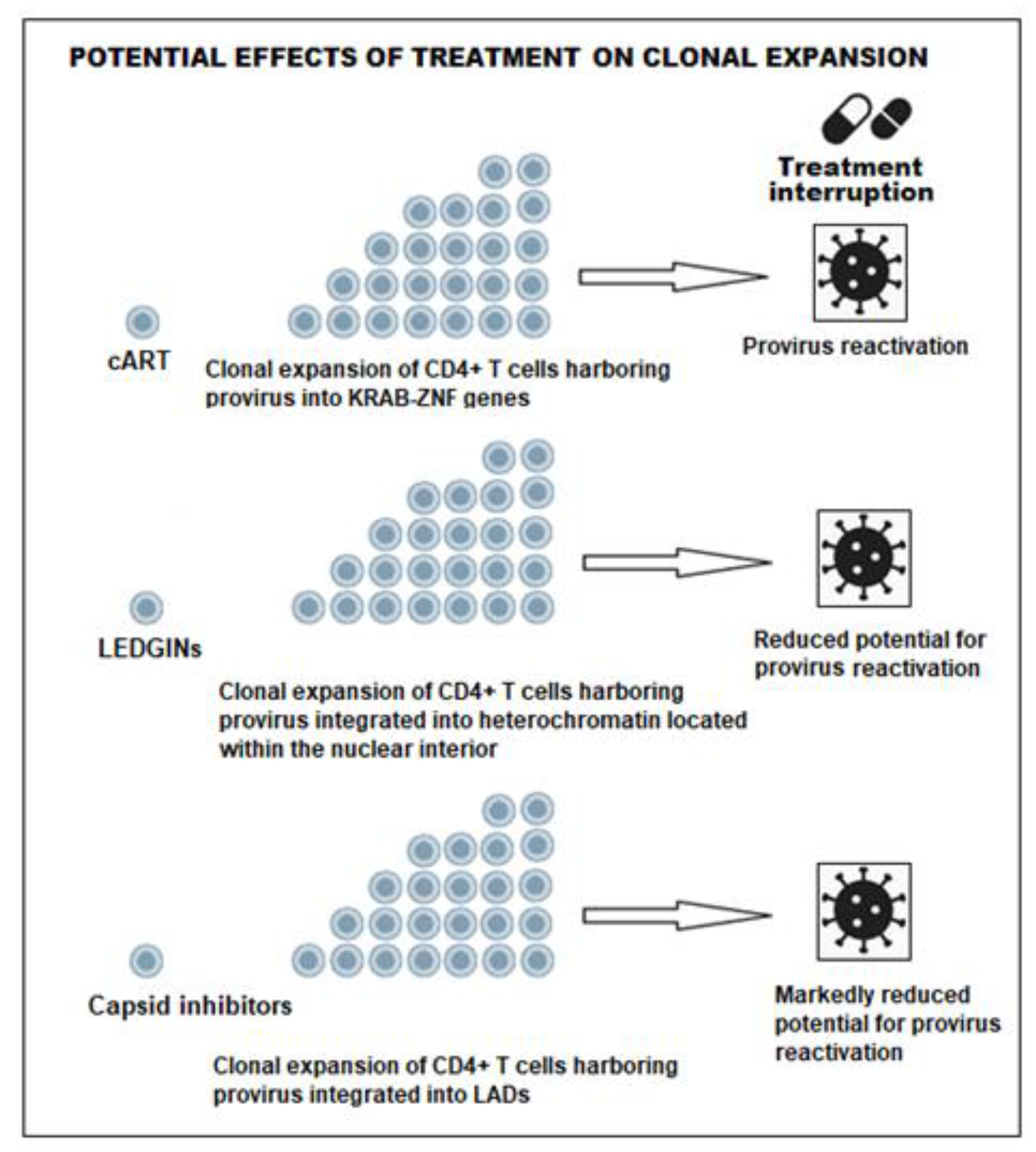
7. Redirecting HIV-1 Integration into LADs: CPSF6 Knockdown and Capsid Inhibitors as Block-and-Lock Strategies
7.1. Role of CPSF6 in Host Gene Regulation and HIV-1 Replication
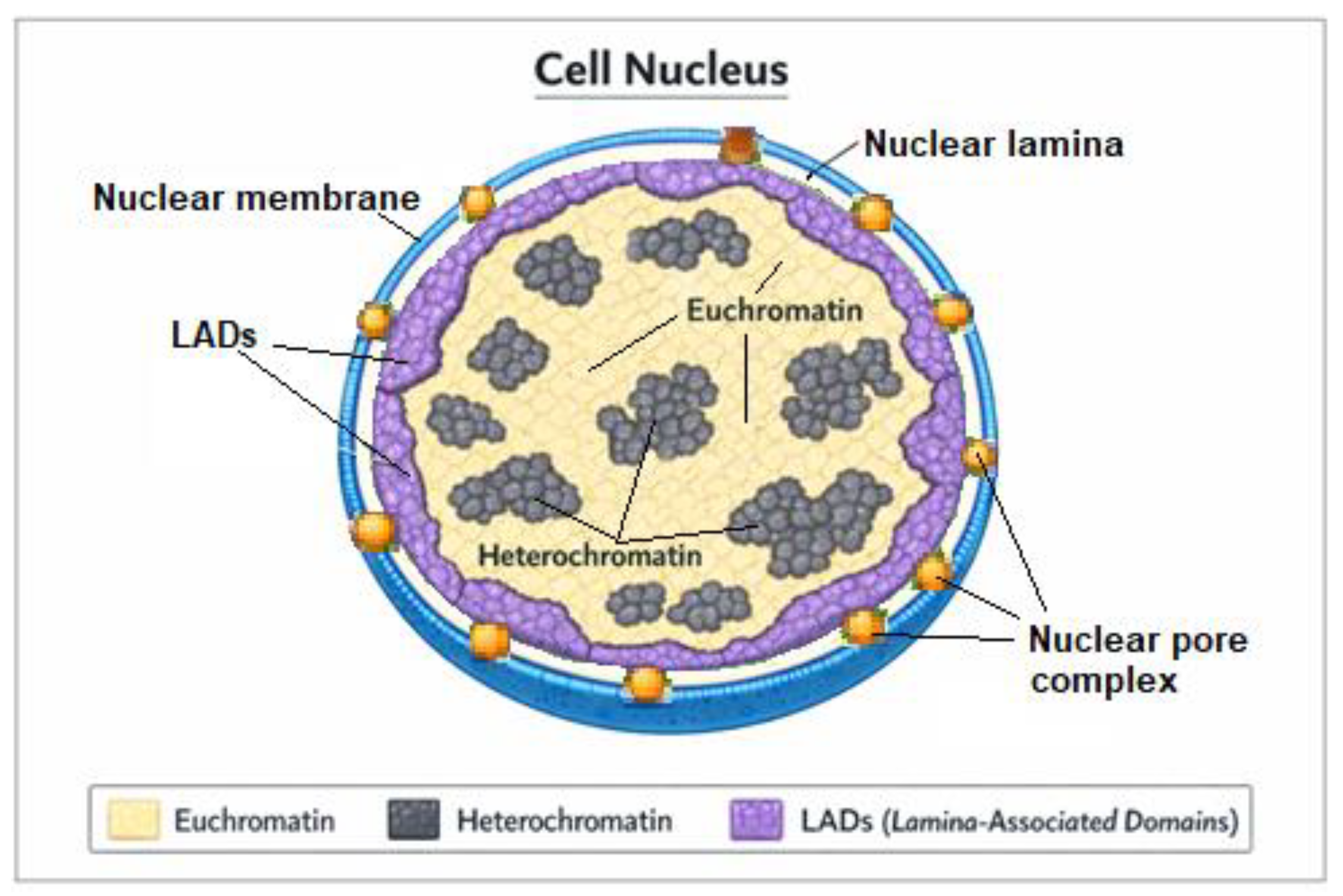
7.2. LADs Structure and Functions
7.3. LADs and HIV-1 Integration
7.4. Could Capsid–CPSF6 Interaction Inhibitors Enable a More Stable Block-and-Lock Strategy than LEDGINs?
7.5. CFIm, Alternative Polyadenylation Remodeling by HIV-1, and Implications for Capsid Inhibitors in Block-and-Lock Strategies
7.6. Capsid-Centered Modulation of HIV-1 Integration: Current Limits and Future Directions Toward LAD Targeting
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Tolomeo, M.; Cascio, A. The complex dysregulations of CD4 T cell subtypes in HIV infection. Int. J. Mol. Sci. 2024, 25, 7512. [Google Scholar] [CrossRef] [PubMed]
- Tolomeo, M.; Cascio, A. The STAT signaling pathway in HIV-1 infection: roles and dysregulation. Int J Mol Sci. 2025, 26, 9123. [Google Scholar] [CrossRef] [PubMed]
- Tolomeo, M.; Cavalli, A.; Cascio, A. Stat1 and its crucial role in the control of viral infections. Int. J. Mol. Sci. 2022, 23, 4095. [Google Scholar] [CrossRef] [PubMed]
- Tolomeo, M.; Tolomeo, F.; Cascio, A. The complex interactions between HIV-1 and human host cell genome: From molecular mechanisms to clinical practice. Int. J. Mol. Sci. 2025, 26, 3184. [Google Scholar] [CrossRef]
- Gunst, J.D.; Gohil, J.; Li, J.Z.; Bosch, R.J.; White, C.; Seamon, A.; Chun, T.W.; Mothe, B.; Gittens, K.; Praiss, L.; De Scheerder, M.A. Time to HIV viral rebound and frequency of post-treatment control after analytical interruption of antiretroviral therapy: an individual data-based meta-analysis of 24 prospective studies. Nat. Commun. 2025, 16, 906. [Google Scholar] [CrossRef]
- Kula-Pacurar, A.; Rodari, A.; Darcis, G.; Van Lint, C. Shocking HIV-1 with immunomodulatory latency reversing agents. Semin. Immunol. 2021, 51, 101478. [Google Scholar] [CrossRef]
- Sadowski, I.; Hashemi, F.B. Strategies to eradicate HIV from infected patients: elimination of latent provirus reservoirs. Cell Mol Life Sci. 2019, 76, 3583–3600. [Google Scholar] [CrossRef]
- Abner, E.; Jordan, A. HIV 'shock and kill' therapy: in need of revision. Antiviral Res. 2019, 166, 19–34. [Google Scholar] [CrossRef]
- Fidler, S.; Stöhr, W.; Pace, M.; Dorrell, L.; Lever, A.; Pett, S.; Kinloch-de Loes, S.; Fox, J.; Clarke, A.; Nelson, M.; et al. Antiretroviral therapy alone versus antiretroviral therapy with a kick and kill approach, on measures of the HIV reservoir in participants with recent HIV infection (The RIVER Trial): a phase 2, randomised trial. Lancet 2020, 395, 888–898. [Google Scholar] [CrossRef]
- Liu, G.; Liu, S.; Zhang, C.; Li, W.; Li, H. Modeling the effects of a shock-and-kill treatment for HIV: latency-reversing agents and natural killer cells. Bull. Math. Biol. 2025, 87, 116. [Google Scholar] [CrossRef]
- Ahlenstiel, C.L.; Symonds, G.; Kent, S.J.; Kelleher, A.D. Block and lock HIV cure strategies to control the latent reservoir. Front. Cell. Infect. Microbiol. 2020, 11 10, 424. [Google Scholar] [CrossRef] [PubMed]
- Vansant, G.; Bruggemans, A.; Janssens, J.; Debyser, Z. Block-and-lock strategies to cure HIV infection. Viruses 2020, 12, 84. [Google Scholar] [CrossRef] [PubMed]
- Moranguinho, I.; Valente, S.T. Block-and-lock: New horizons for a cure for HIV-1. Viruses 2020, 12, 1443. [Google Scholar] [CrossRef] [PubMed]
- Mediouni, S.; Lyu, S.; Schader, S.M.; Valente, S.T. Forging a functional cure for HIV: Transcription regulators and inhibitors. Viruses 2022, 14 14, 1980. [Google Scholar] [CrossRef]
- Li, C.; Mori, L.; Valente, S.T. The block-and-lock strategy for human immunodeficiency virus cure: Lessons learned from didehydro-cortistatin A. J. Infect. Dis. 2021, 223 (Suppl. 2), S46–S53. [Google Scholar] [CrossRef]
- Kessing, C.F.; Nixon, C.C.; Li, C.; Tsai, P.; Takata, H.; Mousseau, G.; Ho, P.T.; Honeycutt, J.B.; Fallahi, M.; Trautmann, L.; et al. In vivo suppression of HIV rebound by didehydro-cortistatin A, a "block-and-lock" strategy for HIV-1 treatment. Cell Rep. 2017, 21, 600–611. [Google Scholar] [CrossRef]
- Mediouni, S.; Kessing, C.F.; Jablonski, J.A.; Thenin-Houssier, S.; Clementz, M.; Kovach, M.D.; Mousseau, G.; de Vera, I.M.S.; Li, C.; Kojetin, D.J.; et al. The Tat inhibitor didehydro-cortistatin A suppresses SIV replication and reactivation. FASEB J. 2019, 33, 8280–8293. [Google Scholar] [CrossRef]
- Pellaers, E.; Janssens, J.; Wils, L.; Denis, A.; Bhat, A.; Van Belle, S.; Feng, D.; Christ, F.; Zhan, P.; Debyser, Z. BRD4 modulator ZL0580 and LEDGINs additively block and lock HIV-1 transcription. Nat. Commun. 2025, 16, 4226. [Google Scholar] [CrossRef]
- Niu, Q.; Liu, Z.; Alamer, E.; Fan, X; Chen, H.; Endsley, J.; Gelman, B.B.; Tian, B.; Kim, J.H.; Michael, N.L.; et al. Structure-guided drug design identifies a BRD4-selective small molecule that suppresses HIV. J. Clin. Investig. 2019, 129, 3361–3373. [Google Scholar] [CrossRef]
- Horvath, R.M.; Brumme, Z.L.; Sadowski, I. CDK8 inhibitors antagonize HIV-1 reactivation and promote provirus latency in T cells. J. Virol. 2023, 97, e0092323. [Google Scholar] [CrossRef]
- Janssens, J.; De Spiegelaere, W.; Trypsteen, W.; Van Nieuwerburgh, F.; De Paepe, A.; Vandekerckhove, L. Mechanisms and efficacy of small molecule latency-promoting agents to inhibit HIV reactivation ex vivo. JCI Insight 2024, 9, e183084. [Google Scholar] [CrossRef]
- Debyser, Z.; Vansant, G.; Bruggemans, A.; Janssens, J.; Christ, F. Insight in HIV integration site selection provides a block-and-lock strategy for a functional cure of HIV infection. Viruses 2018, 11, 12. [Google Scholar] [CrossRef] [PubMed]
- Bruggemans, A.; Vansant, G.; Balakrishnan, M.; Mitchell, M.L.; Cai, R.; Christ, F.; Debyser, Z. GS-9822, a preclinical LEDGIN candidate, displays a block-and-lock phenotype in cell culture. Antimicrob. Agents Chemother. 2023, 65, e02328-20. [Google Scholar] [CrossRef] [PubMed]
- Janssens, J.; De Wit, F.; Parveen, N.; Debyser, Z. Single-cell imaging shows that the transcriptional state of the HIV-1 provirus and its reactivation potential depend on the integration site. mBio 2022, 13, e0000722. [Google Scholar] [CrossRef] [PubMed]
- Vansant, G.; Chen, H.C.; Zorita, E.; Trejbalová, K.; Miklík, D.; Filion, G.; Debyser, Z. The chromatin landscape at the HIV-1 provirus integration site determines viral expression. Nucleic Acids Res. 2020, 48, 7801–7817. [Google Scholar] [CrossRef]
- Sowd, G.A.; Serrao, E.; Wang, H.; Wang, W.; Fadel, H.J.; Poeschla, E.M.; Engelman, A.N. A critical role for alternative polyadenylation factor CPSF6 in targeting HIV-1 integration to transcriptionally active chromatin. Proc. Natl. Acad. Sci. USA 2016, 26 113, E1054–E1063. [Google Scholar] [CrossRef]
- Achuthan, V.; Perreira, J.M.; Sowd, G.A.; Puray-Chavez, M.; McDougall, W.M.; Paulucci-Holthauzen, A.; Wu, X.; Fadel, H.J.; Poeschla, E.M.; Multani, A.S.; et al. Capsid-CPSF6 interaction licenses nuclear HIV-1 trafficking to sites of viral DNA integration. Cell Host Microbe 2018, 24, 392–404. [Google Scholar] [CrossRef]
- Bester, S.M.; Wei, G.; Zhao, H.; Adu-Ampratwum, D.; Iqbal, N.; Courouble, V.V.; Francis, A.C.; Annamalai, A.S.; Singh, P.K.; Shkriabai, N.; et al. Structural and mechanistic bases for a potent HIV-1 capsid inhibitor. Science 2020, 28 6514, 360–364. [Google Scholar] [CrossRef]
- Briand, N.; Collas, P. Lamina-associated domains: Peripheral matters and internal affairs. Genome Biol. 2020, 21, 85. [Google Scholar] [CrossRef]
- Hitchcock, A.M.; Kufel, W.D.; Dwyer, K.A.M.; Sidman, E.F. Lenacapavir: A novel injectable HIV-1 capsid inhibitor. Int. J. Antimicrob. Agents 2024, 63, 107009. [Google Scholar] [CrossRef]
- Engelman, A.N. HIV capsid and integration targeting. Viruses 2021, 13, 125. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Berhe, M.; Crofoot, G.; Benson, P.; Ramgopal, M.; Sims, J.; McDonald, C.; Ruane, P.; Sanchez, W.E.; Scribner, A.; et al. Lenacapavir administered every 26 weeks or daily in combination with oral ART for initial HIV treatment: A randomized, open-label, phase 2 trial. Lancet HIV 2023, 10, e15–e23. [Google Scholar] [CrossRef] [PubMed]
- Mounzer, K.; Slim, J.; Ramgopal, M.; Hedgcock, M.; Bloch, M.; Santana, J.; Mendes, I.; Guo, Y.; Arora, P.; Montezuma-Rusca, J.M.; et al. Phase 2 study of switch to daily BIC + LEN in individuals on a complex HIV treatment regimen. In Proceedings of the CROI Conference, Denver, CO, USA, 3–6 March 2024. [Google Scholar]
- Mounzer, K.; Slim, J.; Ramgopal, M.; Hedgcock, M.; Bloch, M.; Santana, J.; Mendes, I.; Guo, Y.; Arora, P.; Montezuma-Rusca, J.M.; et al. Efficacy and safety of switching to daily Bictegravir plus Lenacapavir from a complex HIV treatment regimen: A randomized, open-label, multicenter phase 2 Study (ARTISTRY-1). Clin. Infect. Dis. 2025, 80, 881–888. [Google Scholar] [CrossRef] [PubMed]
- Doan, J.; Brunzo-Hager, S.; Satterly, B.; Cory, T.J. Expanding therapeutic options: Lenacapavir + bictegravir as a potential treatment for HIV. Expert Opin. Pharmacother. 2023, 18, 1949–1956. [Google Scholar] [CrossRef]
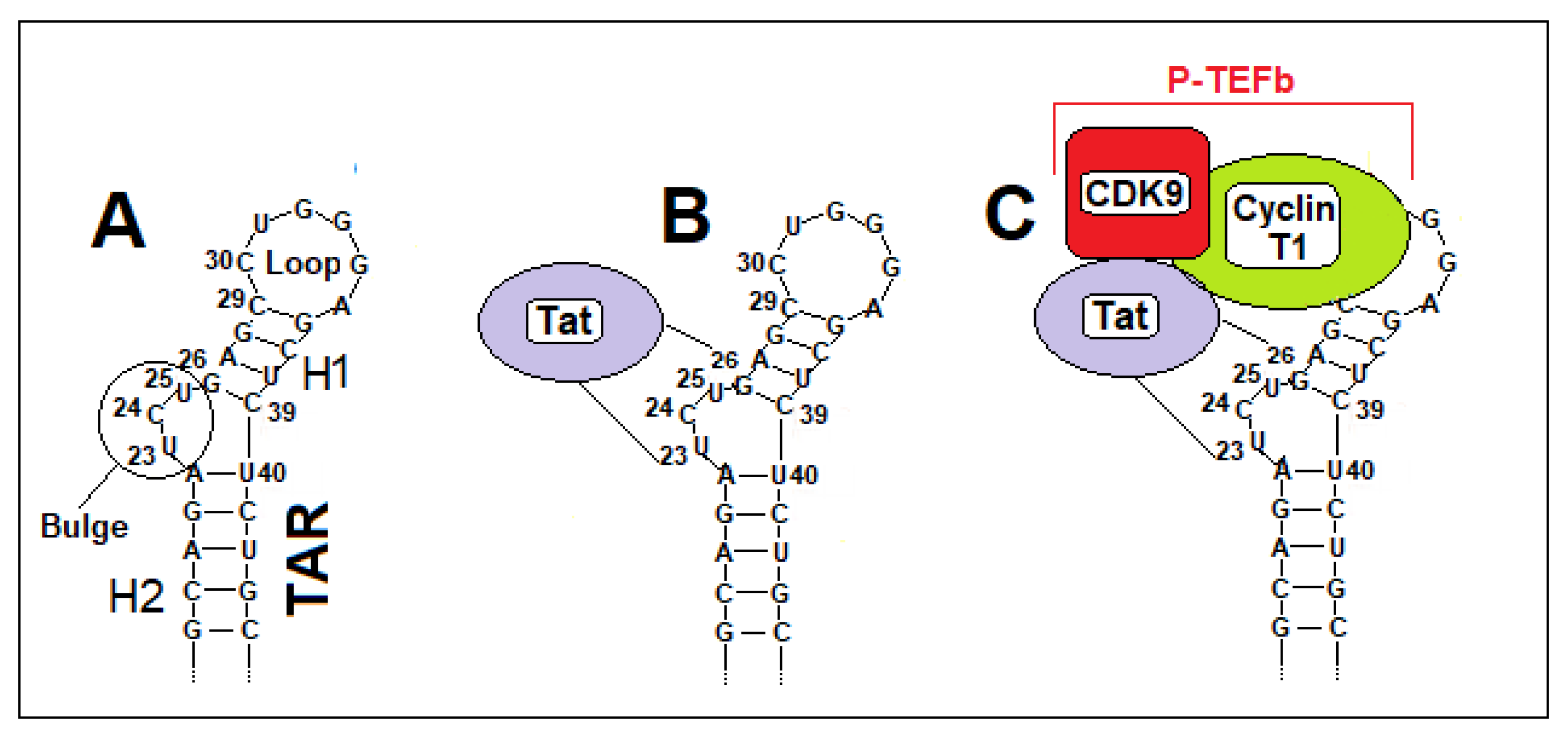
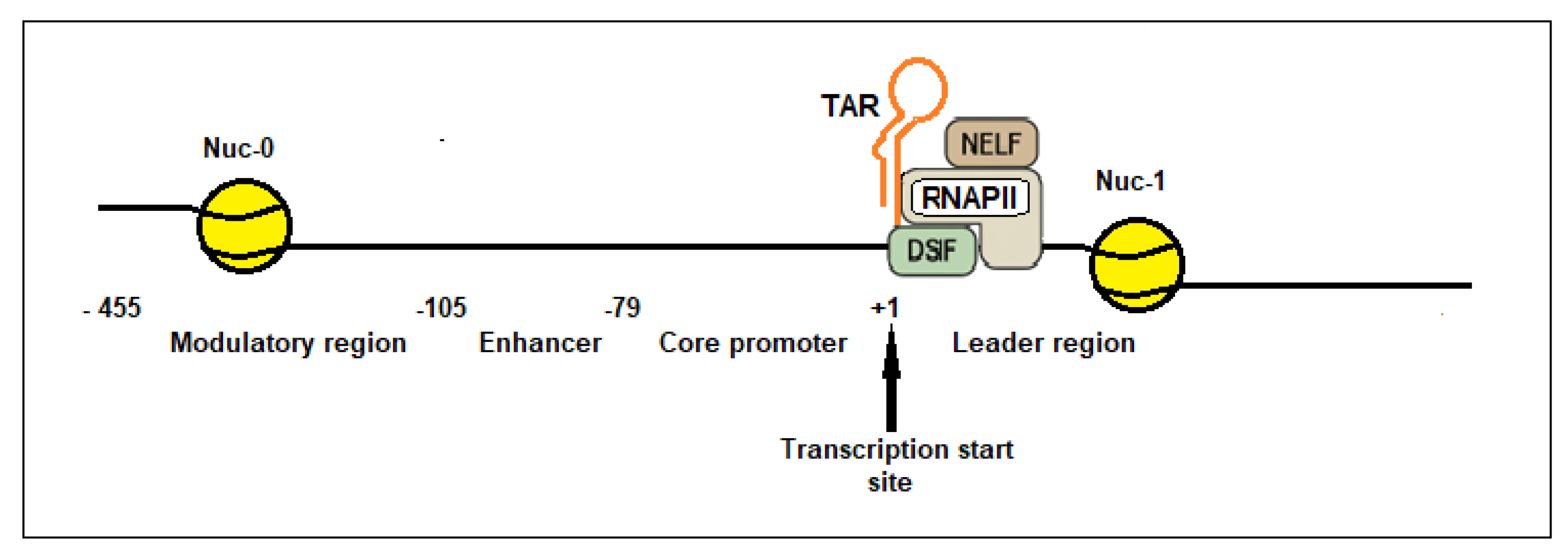
- Romani, B.; Engelbrecht, S.; Glashoff, R.H. Functions of Tat: the versatile protein of human immunodeficiency virus type 1. J. Gen. Virol. 2010, 91, 1–12. [Google Scholar] [CrossRef]
- Schulze-Gahmen, U.; Hurley, J.H. Structural mechanism for HIV-1 TAR loop recognition by Tat and the super elongation complex. Proc. Natl. Acad. Sci. USA 2018, 115, 12973–12978. [Google Scholar] [CrossRef]
- Chavali, S.; Bonn-Breach, R.; Wedekind, J. Face-time with TAR: portraits of an HIV-1 RNA with diverse modes of effector recognition relevant for drug discovery. J. Biol. Chem. 2019, 294, 9326–9341. [Google Scholar] [CrossRef]
- Cao, Y.; Qin, Y.; Zhang, W.; Tian, W.; Ren, Y.; Ren, J.; Wang, J.; Wang, M.; Jiang, J.; Wang, Z. Structural basis of the human negative elongation factor NELF-B/C/E ternary complex. Biochem. Biophys. Res. Commun. 2023, 677, 155–161. [Google Scholar] [CrossRef]
- Su, B.G.; Vos, S.M. Distinct negative elongation factor conformations regulate RNA polymerase II promoter-proximal pausing. Mol. Cell 2024, 84, 1243–1256.e5. [Google Scholar] [CrossRef]
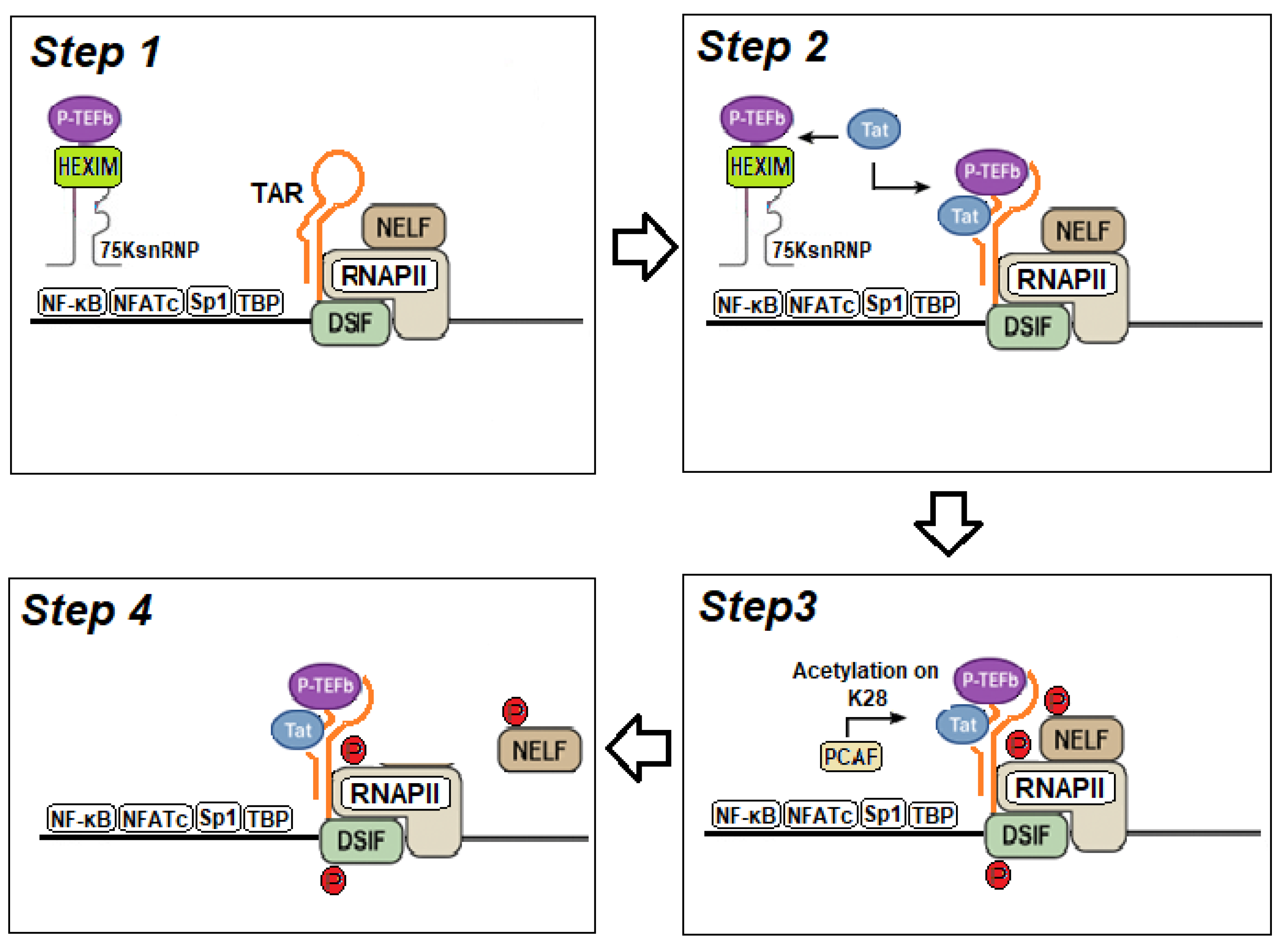
- Fujinaga, K.; Huang, F.; Peterlin, B.M. P-TEFb: the master regulator of transcription elongation. Mol. Cell 2023, 83, 393–403. [Google Scholar] [CrossRef]
- Pham, V.V.; Salguero, C.; Khan, S.N.; Meagher, J.L.; Brown, W.C.; Humbert, N.; de Rocquigny, H.; Smith, J.L.; D’Souza, V.M. HIV-1 Tat interactions with cellular 7SK and viral TAR RNAs identify dual structural mimicry. Nat. Commun. 2018, 9, 4266. [Google Scholar] [CrossRef]
- Barboric, M.; Yik, J.H.; Czudnochowski, N.; Yang, Z.; Chen, R.; Contreras, X.; Geyer, M.; Peterlin, B.M.; Zhou, Q. Tat competes with HEXIM1 to increase the active pool of P-TEFb for HIV-1 transcription. Nucleic Acids Res. 2007, 35, 2003–2012. [Google Scholar] [CrossRef] [PubMed]
- Schulte, A.; Czudnochowski, N.; Barboric, M.; Schönichen, A.; Blazek, D.; Peterlin, B.M.; Geyer, M. Identification of a cyclin T-binding domain in HEXIM1 and biochemical analysis of its binding competition with HIV-1 Tat. J. Biol. Chem. 2005, 280, 24968–24977. [Google Scholar] [CrossRef]
- Mousseau, G.; Valente, S. Strategies to block HIV transcription: focus on small molecule Tat inhibitors. Biology 2012, 1, 668–697. [Google Scholar] [CrossRef] [PubMed]
- Yu, K.; Liu, H.; Pan, T. HIV-1 Tat: molecular switch in viral persistence and emerging technologies for functional cure. Int. J. Mol. Sci. 2025, 26, 6311. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Li, D.; Lin, M.H.; Li, L.; Harrich, D. Tat-based therapies as an adjuvant for an HIV-1 functional cure. Viruses 2020, 12, 415. [Google Scholar] [CrossRef]
- Mediouni, S.; Chinthalapudi, K.; Ekka, M.K.; Usui, I.; Jablonski, J.A.; Clementz, M.A.; Mousseau, G.; Nowak, J.; Macherla, V.R.; Beverage, J.N. Didehydro-cortistatin A inhibits HIV-1 by specifically binding to the unstructured basic region of Tat. mBio 2019, 10, e02662-18. [Google Scholar] [CrossRef]
- Mousseau, G.; Kessing, C.F.; Fromentin, R.; Trautmann, L.; Chomont, N.; Valente, S.T. The Tat inhibitor didehydro-cortistatin A prevents HIV-1 reactivation from latency. mBio 2015, 6, e00465. [Google Scholar] [CrossRef]
- Pitman, M.C.; Lau, J.S.Y.; McMahon, J.H.; Lewin, S.R. Barriers and strategies to achieve a cure for HIV. Lancet HIV 2018, 5, e317–e328. [Google Scholar] [CrossRef]
- Ling, L.; Leda, A.R.; Begum, N.; Spagnuolo, R.A.; Wahl, A.; Garcia, J.V.; Valente, S.T. Loss of in vivo replication fitness of HIV-1 variants resistant to the Tat inhibitor dCA. Viruses 2023, 15, 950. [Google Scholar] [CrossRef]
- Mousseau, G.; Aneja, R.; Clementz, M.A.; Mediouni, S.; Lima, N.S.; Haregot, A.; Kessing, C.F.; Jablonski, J.A.; Thenin-Houssier, S.; Nagarsheth, N.; et al. Resistance to the Tat inhibitor didehydro-cortistatin A is mediated by heightened basal HIV-1 transcription. mBio 2019, 10, e01750-18. [Google Scholar] [CrossRef] [PubMed]
- Shin, Y.H.; Kim, D.E.; Yu, K.L.; Park, C.M.; Kim, H.G.; Kim, K.C.; Bae, S.; Yoon, C.H. Novel time-resolved fluorescence resonance energy transfer assay for the discovery of small-molecule inhibitors of HIV-1 Tat-regulated transcription. Int. J. Mol. Sci. 2023, 24, 9139. [Google Scholar] [CrossRef] [PubMed]
- Alanazi, A.; Ivanov, A.; Kumari, N.; Lin, X.; Wang, S.; Kovalskyy, D.; Nekhai, S. Targeting Tat–TAR RNA interaction for HIV-1 inhibition. Viruses 2021, 13, 2004. [Google Scholar] [CrossRef] [PubMed]
- Ronsard, L.; Rai, T.; Rai, D.; Ramachandran, V.G.; Banerjea, A.C. In silico analyses of subtype-specific HIV-1 Tat–TAR RNA interaction reveal the structural determinants for viral activity. Front. Microbiol. 2017, 8, 1467. [Google Scholar] [CrossRef]
- Aboul-Ela, F.; Karn, J.; Varani, G. Structure of HIV-1 TAR RNA in the absence of ligands reveals a novel conformation of the trinucleotide bulge. Nucleic Acids Res. 1996, 24, 3974–3981. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, Z.; Rana, T.M. Visualizing a specific contact in the HIV-1 Tat protein fragment and trans-activation responsive region RNA complex by photocross-linking. J. Biol. Chem. 1996, 271, 10391–10396. [Google Scholar] [CrossRef]
- Sheline, C.T.; Milocco, L.H.; Jones, K.A. Two distinct nuclear transcription factors recognize loop and bulge residues of the HIV-1 TAR RNA hairpin. Genes Dev. 1991, 5, 2508–2520. [Google Scholar] [CrossRef]
- Chavali, S.S.; Mali, S.M.; Bonn, R.; Saseendran Anitha, A.; Bennett, R.P.; Smith, H.C.; Fasan, R.; Wedekind, J.E. Cyclic peptides with a distinct arginine-fork motif recognize the HIV trans-activation response RNA in vitro and in cells. J. Biol. Chem. 2021, 297, 101390. [Google Scholar] [CrossRef]
- Maity, D.; Kumar, S.; Curreli, F.; Debnath, A.K.; Hamilton, A.D. α-Helix-mimetic foldamers for targeting HIV-1 TAR RNA. Chem. Eur. J. 2019, 25, 7265–7269. [Google Scholar] [CrossRef]
- Khatkar, P.; Mensah, G.; Ning, S.; Cowen, M.; Kim, Y.; Williams, A.; Abulwerdi, F.A.; Zhao, Y.; Zeng, C.; Le Grice; et al. HIV-1 transcription inhibition using small RNA-binding molecules. Pharmaceuticals 2023, 17, 33. [Google Scholar] [CrossRef]
- Davidson, A.; Leeper, T.C.; Athanassiou, Z.; Patora-Komisarska, K.; Karn, J.; Robinson, J.A.; Varani, G. Simultaneous recognition of HIV-1 TAR RNA bulge and loop sequences by cyclic peptide mimics of Tat protein. Proc. Natl. Acad. Sci. USA 2009, 106, 11931–11936. [Google Scholar] [CrossRef] [PubMed]
- Wan, Z.; Chen, X. Triptolide inhibits human immunodeficiency virus type 1 replication by promoting proteasomal degradation of Tat protein. Retrovirology 2014, 11, 88. [Google Scholar] [CrossRef] [PubMed]
- Gorrie, D.; Bravo, M.; Fan, L. The Yin and Yang of the natural product triptolide and its interactions with XPB, an essential protein for gene expression and DNA repair. Genes 2024, 15, 1287. [Google Scholar] [CrossRef] [PubMed]
- Lv, T.; Cao, W.; Xue, J.; Wei, Q.; Qiu, Z.; Han, Y.; Li, T. Therapeutic effect of (5R)-5-hydroxytriptolide (LLDT-8) in SIV infected rhesus monkeys. Int. Immunopharmacol. 2022, 110, 108932. [Google Scholar] [CrossRef]
- Liu, X.; Lv, T.; Li, X.; Xue, J.; Lin, L.; Lu, L.; Li, X.; Yang, Y.; Wu, Y.; Wei, Q.; et al. Comprehensive transcriptomic analyses identify the immunosuppressive effects of LLDT-8 in ART-treated SIV-infected rhesus macaques. Int. Immunopharmacol. 2024, 126, 111173. [Google Scholar] [CrossRef]
- Mukim, A.; Smith, D.M.; Deshmukh, S.; Qazi, A.A.; Beliakova-Bethell, N. A camptothecin analog, topotecan, promotes HIV latency via interference with HIV transcription and RNA splicing. J. Virol. 2023, 97, e01630-22. [Google Scholar] [CrossRef]
- Xu, Y.; Peng, X.; Zheng, Y.; Jin, C.; Lu, X.; Han, D.; Fu, H.; Chen, C.; Wu, N. Inactivation of latent HIV-1 proviral DNA using clustered regularly interspaced short palindromic repeats/Cas9 treatment and the assessment of off-target effects. Front. Microbiol. 2021, 12, 629153. [Google Scholar] [CrossRef]
- Yin, C.; Zhang, T.; Qu, X.; Zhang, Y.; Putatunda, R.; Xiao, X.; Li, F.; Xiao, W.; Zhao, H.; Dai, S.; et al. In vivo excision of HIV-1 provirus by saCas9 and multiplex single-guide RNAs in animal models. Mol. Ther. 2017, 25, 1168–1186. [Google Scholar] [CrossRef]
- Bhowmik, R.; Chaubey, B. CRISPR/Cas9: A tool to eradicate HIV-1. AIDS Res. Ther. 2022, 19, 58. [Google Scholar] [CrossRef]
- Burdo, T.H.; Chen, C.; Kaminski, R.; Sariyer, I.K.; Mancuso, P.; Donadoni, M.; Smith, M.D.; Sariyer, R.; Caocci, M.; Liao, S.; et al. Preclinical safety and biodistribution of CRISPR targeting SIV in non-human primates. Gene Ther. 2024, 31, 434–435. [Google Scholar] [CrossRef]
- Dash, P.K.; Chen, C.; Kaminski, R.; Su, H.; Mancuso, P.; Sillman, B.; Zhang, C.; Liao, S.; Sravanam, S.; Liu, H.; et al. CRISPR editing of CCR5 and HIV-1 facilitates viral elimination in antiretroviral drug-suppressed virus-infected humanized mice. Proc. Natl. Acad. Sci. U.S.A. 2023, 120, e2217887120. [Google Scholar] [CrossRef] [PubMed]
- Presti, R.; Kennedy, W.; Craddick, T.J.; Gordon, J. First-in-human study of EBT-101 in aviremic HIV-1 infected adults on stable ART. In Proceedings of the 27th Annual Meeting of the American Society of Gene & Cell Therapy, Baltimore, MD, USA, 7–11 May 2024. [Google Scholar]
- Kyei, G.B.; Meng, S.; Ramani, R.; Niu, A.; Lagisetti, C.; Webb, T.R.; Ratner, L. Splicing factor 3B subunit 1 interacts with HIV Tat and plays a role in viral transcription and reactivation from latency. mBio 2018, 9, e01423-18. [Google Scholar] [CrossRef] [PubMed]
- Swamy, M.N.; Wu, H.; Shankar, P. Recent advances in RNAi-based strategies for therapy and prevention of HIV-1/AIDS. Adv. Drug Deliv. Rev. 2016, 103, 174–186. [Google Scholar] [CrossRef] [PubMed]
- Herrera-Carrillo, E.; Berkhout, B. Attacking HIV-1 RNA versus DNA by sequence-specific approaches: RNAi versus CRISPR-Cas. Biochem. Soc. Trans. 2016, 44, 1355–1365. [Google Scholar] [CrossRef]
- Bobbin, M.L.; Burnett, J.C.; Rossi, J.J. RNA interference approaches for treatment of HIV-1 infection. Genome Med. 2015, 7, 50. [Google Scholar] [CrossRef]
- Cisneros, E.; Sherwani, N.; Lanier, O.L.; Peppas, N.A. Targeted delivery methods for RNA interference are necessary to obtain a potential functional cure for HIV/AIDS. Adv. Drug Deliv. Rev. 2023, 199, 114970. [Google Scholar] [CrossRef]
- Suzuki, K.; Hattori, S.; Marks, K.; Ahlenstiel, C.; Maeda, Y.; Ishida, T.; Millington, M.; Boyd, M.; Symonds, G.; Cooper, D.A.; et al. Promoter targeting shRNA suppresses HIV-1 infection in vivo through transcriptional gene silencing. Mol. Ther. Nucleic Acids 2013, 2, e137. [Google Scholar] [CrossRef]
- Alpuche-Lazcano, S.P.; Scarborough, R.J.; Gatignol, A. MicroRNAs and long non-coding RNAs during transcriptional regulation and latency of HIV and HTLV. Retrovirology 2024, 21, 5. [Google Scholar] [CrossRef]
- Zhou, J.; Lazar, D.; Li, H.; Xia, X.; Satheesan, S.; Charlins, P.; O’Mealy, D.; Akkina, R.; Saayman, S.; Weinberg, M.S.; et al. Receptor-targeted aptamer-siRNA conjugate-directed transcriptional regulation of HIV-1. Theranostics 2018, 8, 1575–1590. [Google Scholar] [CrossRef]
- Zhao, N.; Wang, G.; Das, A.T.; Berkhout, B. Combinatorial CRISPR-Cas9 and RNA interference attack on HIV-1 DNA and RNA can lead to cross-resistance. Antimicrob. Agents Chemother. 2017, 61, e01486-17. [Google Scholar] [CrossRef]
- Herrera-Carrillo, E.; Gao, Z.; Berkhout, B. CRISPR therapy towards an HIV cure. Brief Funct. Genomics 2020, 19, 201–208. [Google Scholar]
- Hussein, M.; Molina, M.A.; Berkhout, B.; Herrera-Carrillo, E. A CRISPR-Cas cure for HIV/AIDS. Int. J. Mol. Sci. 2023, 24, 1563. [Google Scholar] [CrossRef] [PubMed]
- Maslennikova, A.; Mazurov, D. Application of CRISPR/Cas genomic editing tools for HIV therapy: Toward precise modifications and multilevel protection. Front. Cell Infect. Microbiol. 2022, 12, 880030. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Tian, J.; Wu, T. BRD4 in physiology and pathology: “BET” on its partners. BioEssays 2021, 43, e2100180. [Google Scholar] [CrossRef]
- Devaiah, B.N.; Case-Borden, C.; Gegonne, A.; Hsu, C.H.; Chen, Q.; Meerzaman, D.; Dey, A.; Ozato, K.; Singer, D.S. BRD4 is a histone acetyltransferase that evicts nucleosomes from chromatin. Nat. Struct. Mol. Biol. 2016, 23, 540–548. [Google Scholar] [CrossRef]
- Weissman, J.D.; Singh, A.K.; Devaiah, B.N.; Schuck, P.; LaRue, R.C.; Singer, D.S. The intrinsic kinase activity of BRD4 spans its BD2–B–BID domains. J. Biol. Chem. 2021, 297, 101326. [Google Scholar] [CrossRef]
- Devaiah, B.N.; Gegonne, A.; Singer, D.S. Bromodomain 4: a cellular Swiss Army knife. J. Leukoc. Biol. 2016, 100, 679–686. [Google Scholar] [CrossRef]
- Schröder, S.; Cho, S.; Zeng, L.; Zhang, Q.; Kaehlcke, K.; Mak, L.; Lau, J.; Bisgrove, D.; Schnölzer, M.; Verdin, E.; et al. Two-pronged binding with bromodomain-containing protein 4 liberates positive transcription elongation factor b from inactive ribonucleoprotein complexes. J. Biol. Chem. 2012, 287, 1090–1099. [Google Scholar] [CrossRef]
- Yang, Z.; Yik, J.H.N.; Chen, R.; He, N.; Jang, M.K.; Ozato, K.; Zhou, Q. Recruitment of P-TEFb for stimulation of transcriptional elongation by the bromodomain protein Brd4. Mol. Cell 2005, 19, 535–545. [Google Scholar] [CrossRef]
- Chen, R.; Yik, J.H.N.; Lew, Q.J.; Chao, S.H. Brd4 and HEXIM1: multiple roles in P-TEFb regulation and cancer. BioMed Res. Int. 2014, 2014, 232870. [Google Scholar] [CrossRef]
- Zhou, K.; Zhuang, S.; Liu, F.; Chen, Y.; Li, Y.; Wang, S.; Li, Y.; Wen, H.; Lin, X.; Wang, J.; et al. Disrupting the Cdk9/Cyclin T1 heterodimer of 7SK snRNP for the Brd4 and AFF1/4 guided reconstitution of active P-TEFb. Nucleic Acids Res. 2022, 50, 750–762. [Google Scholar] [CrossRef]
- Jang, M.K.; Mochizuki, K.; Zhou, M.; Jeong, H.S.; Brady, J.N.; Ozato, K. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol. Cell 2005, 19, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Bisgrove, D.A.; Mahmoudi, T.; Henklein, P.; Verdin, E. Conserved P-TEFb-interacting domain of BRD4 inhibits HIV transcription. Proc. Natl. Acad. Sci. USA 2007, 104, 13690–13695. [Google Scholar] [CrossRef] [PubMed]
- Muniz, L.; Egloff, S.; Ughy, B.; Jády, B.E.; Kiss, T. Controlling cellular P-TEFb activity by the HIV-1 transcriptional transactivator Tat. PLoS Pathog. 2010, 6, e1001152. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Li, Z.; Xue, Y.; Schulze-Gahmen, U.; Johnson, J.R.; Krogan, N.J.; Alber, T.; Zhou, Q. AFF1 is a ubiquitous P-TEFb partner to enable Tat extraction of P-TEFb from 7SK snRNP and formation of SECs for HIV transactivation. Proc. Natl. Acad. Sci. USA 2014, 111, E15–E24. [Google Scholar] [CrossRef]
- Itzen, F.; Greifenberg, A.K.; Bösken, C.A.; Geyer, M. Brd4 activates P-TEFb for RNA polymerase II CTD phosphorylation. Nucleic Acids Res. 2014, 42, 7577–7590. [Google Scholar] [CrossRef]
- Mbonye, U.; Wang, B.; Gokulrangan, G.; Shi, W.; Yang, S.; Karn, J. Cyclin-dependent kinase 7 (CDK7)-mediated phosphorylation of the CDK9 activation loop promotes P-TEFb assembly with Tat and proviral HIV reactivation. J. Biol. Chem. 2018, 293, 10009–10025. [Google Scholar] [CrossRef]
- Krueger, B.J.; Varzavand, K.; Cooper, J.J.; Price, D.H. The mechanism of release of P-TEFb and HEXIM1 from the 7SK snRNP by viral and cellular activators includes a conformational change in 7SK. PLoS ONE 2010, 5, e12335. [Google Scholar] [CrossRef]
- Conrad, R.J.; Fozouni, P.; Thomas, S.; Sy, H.; Zhang, Q.; Zhou, M.M.; Ott, M. The short isoform of BRD4 promotes HIV-1 latency by engaging repressive SWI/SNF chromatin-remodeling complexes. Mol. Cell 2017, 67, 1001–1012.e6. [Google Scholar] [CrossRef]
- Huang, W.; Zheng, X.; Yang, Y.; Wang, X.; Shen, Z. An overview on small molecule inhibitors of BRD4. Mini Rev. Med. Chem. 2016, 16, 1403–1414. [Google Scholar] [CrossRef]
- Jiang, G.; Deng, W.; Liu, Y.; Wang, C. General mechanism of JQ1 in inhibiting various types of cancer. Mol. Med. Rep. 2020, 21, 1021–1034. [Google Scholar] [CrossRef]
- Li, Z.; Guo, J.; Wu, Y.; Zhou, Q. The BET bromodomain inhibitor JQ1 activates HIV latency through antagonizing Brd4 inhibition of Tat transactivation. Nucleic Acids Res. 2013, 41, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Gaiha, G.D.; John, S.P.; Pertel, T.; Chin, C.R.; Gao, G.; Qu, H.; Walker, B.D.; Elledge, S.J.; Brass, A.L.; et al. Reactivation of latent HIV-1 by inhibition of BRD4. Cell Rep. 2012, 2, 807–816. [Google Scholar] [CrossRef] [PubMed]
- Bartholomeeusen, K.; Xiang, Y.; Fujinaga, K.; Peterlin, B.M. Bromodomain and extra-terminal (BET) bromodomain inhibition activates transcription via transient release of positive transcription elongation factor B (P-TEFb) from 7SK small nuclear ribonucleoprotein. J. Biol. Chem. 2012, 287, 36609–36616. [Google Scholar] [CrossRef]
- Kumar, N.; Ma, Z.; Long, F.; Bonam, S.R.; Lai, H.-T.; Wu, S.-Y.; Chen, H.; Hazell, N.C.; Bei, J.; Liu, X.; et al. Mechanistic insights and in vivo HIV suppression by the BRD4-targeting small molecule ZL0580. bioRxiv 2025, 2025.08.14.670267. [Google Scholar] [CrossRef] [PubMed]
- Alamer, E.; Zhong, C.; Liu, Z.; Niu, Q.; Long, F.; Guo, L.; Gelman, B.B.; Soong, L.; Zhou, J.; Hu, H. Epigenetic suppression of HIV in myeloid cells by the BRD4-selective small molecule modulator ZL0580. J. Virol. 2020, 94, e01880-19. [Google Scholar] [CrossRef]
- Victoriano, A.F.; Asamitsu, K.; Hibi, Y.; Imai, K.; Barzaga, N.G.; Okamoto, T. Inhibition of human immunodeficiency virus type 1 replication in latently infected cells by a novel IkappaB kinase inhibitor. Antimicrob. Agents Chemother. 2006, 50, 547–555. [Google Scholar] [CrossRef]
- de Jesus, M.S.M.; Macabeo, A.P.G.; Ramos, J.D.A.; de Leon, V.N.O.; Asamitsu, K.; Okamoto, T. Voacanga globosa spirobisindole alkaloids exert antiviral activity in HIV latently infected cell lines by targeting the NF-kB cascade: In vitro and in silico investigations. Molecules 2022, 27, 1078. [Google Scholar] [CrossRef]
- Pande, V.; Ramos, M.J. Nuclear factor kappa B: A potential target for anti-HIV chemotherapy. Curr. Med. Chem. 2003, 10, 1603–1615. [Google Scholar] [CrossRef]
- Huang, T.; Cai, J.; Wang, P.; Zhou, J.; Zhang, H.; Wu, Z.; Zhao, J.; Huang, Z.; Deng, K. Ponatinib represses latent HIV-1 by inhibiting AKT-mTOR. Antimicrob. Agents Chemother. 2023, 67, e0006723. [Google Scholar] [CrossRef]
- Cai, J.F.; Zhou, J.S.; Meng, Z.Y.; Wu, Z.Q.; Zhao, J.C.; Peng, H.X.; Liang, X.Y.; Chen, J.J.; Wang, P.P.; Deng, K. Ripretinib inhibits HIV-1 transcription through modulation of PI3K-AKT-mTOR. Acta Pharmacol. Sin. 2024, 45, 1632–1643. [Google Scholar] [CrossRef]
- Nunes, J.M.; Furtado, M.N.; de Morais Nunes, E.R.; Sucupira, M.C.A.; Diaz, R.S.; Janini, L.M.R. Modulation of epigenetic factors during the early stages of HIV-1 infection in CD4⁺ T cells in vitro. Virology 2018, 523, 41–51. [Google Scholar] [CrossRef] [PubMed]
- Vargas, B; Giacobbi, NS; Sanyal, A; et al. Inhibitors of Signaling Pathways That Block Reversal of HIV-1 Latency. Antimicrobial Agents and Chemotherapy 2019, 63(2), e01744-18. [Google Scholar] [CrossRef] [PubMed]
- Vargas, B.; Giacobbi, N.S.; Sanyal, A.; Venkatachari, N.J.; Han, F.; Gupta, P.; Sluis-Cremer, N. Inhibitors of signaling pathways that block reversal of HIV-1 latency. Antimicrob. Agents Chemother. 2019, 63, e01744-18. [Google Scholar] [CrossRef] [PubMed]
- Renkema, G.H.; Manninen, A.; Mann, D.A.; Harris, M.; Saksela, K. Identification of the Nef-associated kinase as p21-activated kinase 2. Curr. Biol. 1999, 9, 1407–1410. [Google Scholar] [CrossRef]
- Saksela, K. HIV-1 Nef and host cell protein kinases. Front. Biosci. 2011, 16, 176–191. [Google Scholar] [CrossRef]
- Vargas, B.; Boslett, J.; Yates, N.; Sluis-Cremer, N. Mechanism by which PF-3758309, a pan isoform inhibitor of p21-activated kinases, blocks reactivation of HIV-1 latency. Biomolecules 2023, 13, 100. [Google Scholar] [CrossRef]
- Pisell, T.L.; Ho, O.; Lee, G.; Butera, S.T. Spectrum of CDK9 inhibitor activity against HIV-1 replication among various models of chronic and latent infection. Antivir. Chem. Chemother. 2001, 12, 33–41. [Google Scholar]
- Horvath, R.M.; Brumme, Z.L.; Sadowski, I. Small molecule inhibitors of transcriptional cyclin-dependent kinases impose HIV-1 latency, presenting "block and lock" treatment strategies. Antimicrob. Agents Chemother. 2024, 68, e0107223. [Google Scholar] [CrossRef]
- Dahal, S.; Clayton, K.; Been, T.; Fernet-Brochu, R.; Ocando, A.V.; Balachandran, A.; Poirier, M.; Maldonado, R.K.; Shkreta, L.; Boligan, K.F.; et al. Opposing roles of CLK SR kinases in controlling HIV-1 gene expression and latency. Retrovirology 2022, 19, 18. [Google Scholar] [CrossRef]
- Quivy, V.; De Walque, S.; Van Lint, C. Chromatin-associated regulation of HIV-1 transcription: implications for the development of therapeutic strategies. Subcell. Biochem. 2007, 41, 371–396. [Google Scholar] [PubMed]
- Hokello, J.; Sharma, A.L.; Tyagi, M. Efficient non-epigenetic activation of HIV latency through the T-cell receptor signalosome. Viruses 2020, 12, 868. [Google Scholar] [CrossRef] [PubMed]
- Tripathy, M.K.; Abbas, W.; Herbein, G. Epigenetic regulation of HIV-1 transcription. Epigenomics 2011, 3, 487–502. [Google Scholar] [CrossRef] [PubMed]
- Imai, K.; Togami, H.; Okamoto, T. Involvement of histone H3 lysine 9 (H3K9) methyltransferase G9a in the maintenance of HIV-1 latency and its reactivation by BIX01294. J. Biol. Chem. 2010, 285, 16538–16545. [Google Scholar] [CrossRef]
- Matsuda, Y.; Kobayashi-Ishihara, M.; Fujikawa, D.; Ishida, T.; Watanabe, T.; Yamagishi, M. Epigenetic heterogeneity in HIV-1 latency establishment. Sci. Rep. 2015, 5, 7701. [Google Scholar] [CrossRef]
- Machida, S.; Depierre, D.; Chen, H.C.; Thenin-Houssier, S.; Petitjean, G.; Doyen, C.M.; Takaku, M.; Cuvier, O.; Benkirane, M. Exploring histone loading on HIV DNA reveals a dynamic nucleosome positioning between unintegrated and integrated viral genome. Proc. Natl. Acad. Sci. USA 2020, 117, 6822–6830. [Google Scholar] [CrossRef]
- Lusic, M.; Giacca, M. Regulation of HIV-1 latency by chromatin structure and nuclear architecture. J.Mol. Biol. 2015, 427, 688–694. [Google Scholar] [CrossRef]
- Widłak, P.; Garrard, W.T. Nucleosomes and regulation of gene expression: Structure of the HIV-1 5′ LTR. Acta Biochim. Pol. 1998, 45, 209–219. [Google Scholar] [CrossRef]
- Nguyen, K.; Dobrowolski, C.; Shukla, M.; Cho, W.-K.; Luttge, B.; Karn, J. Inhibition of the H3K27 demethylase UTX enhances the epigenetic silencing of HIV proviruses and induces HIV-1 DNA hypermethylation but fails to permanently block HIV reactivation. PLoS Pathog. 2021, 17, e1010014. [Google Scholar] [CrossRef]
- Balasubramanyam, K.; Altaf, M.; Varier, R.A.; Swaminathan, V.; Siddappa, N.B.; Ranga, U.; Kundu, T.K. Polyisoprenylated benzophenone, garcinol, a natural histone acetyltransferase inhibitor, represses chromatin transcription and alters global gene expression. J. Biol. Chem. 2004, 279, 33716–33726. [Google Scholar] [CrossRef]
- Mantelingu, K.; Reddy, B.A.A.; Swaminathan, V.; Meera, P.; Ramachandran, L.; Balasubramanyam, K.; Kundu, T.K. Specific inhibition of p300-HAT alters global gene expression and represses HIV replication. Chem. Biol. 2007, 14, 645–657. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.-H.; Ke, Y.-Y.; Su, C.-T.; Shiao, H.-Y.; Hsieh, H.-P.; Chao, Y.-K.; Lee, C.-N.; Kao, C.-L.; Chao, Y.-S.; Chang, S.-Y. Inhibition of HIV-1 Tat-mediated transcription by a coumarin derivative, BPRHIV001, through the Akt pathway. J. Virol. 2011, 85, 9114–9126. [Google Scholar] [CrossRef] [PubMed]
- Matysiak, J.; Lesbats, P.; Mauro, E.; Lapaillerie, D.; Dupuy, J.W.; Lopez, A.P.; Benleulmi, M.S.; Calmels, C.; Andreola, M.L.; Ruff, M.; et al. Modulation of chromatin structure by the FACT histone chaperone complex regulates HIV-1 integration. Retrovirology 2017, 14, 39. [Google Scholar] [CrossRef] [PubMed]
- Gallastegui, E.; Millán-Zambrano, G.; Terme, J.M.; Chávez, S.; Jordan, A. Chromatin reassembly factors are involved in transcriptional interference promoting HIV latency. J. Virol. 2011, 85, 3187–3202. [Google Scholar] [CrossRef]
- Vanti, M.; Gallastegui, E.; Respaldiza, I.; Rodríguez-Gil, A.; Gómez-Herreros, F.; Jimeno-González, S.; Jordan, A.; Chávez, S. Yeast genetic analysis reveals the involvement of chromatin reassembly factors in repressing HIV-1 basal transcription. PLoS Genet. 2009, 5, e1000339. [Google Scholar] [CrossRef]
- Jean, M.J.; Hayashi, T.; Huang, H.; Brennan, J.; Simpson, S.; Purmal, A.; Gurova, K.; Keefer, M.C.; Kobie, J.J.; Santoso, N.G.; et al. Curaxin CBL0100 blocks HIV-1 replication and reactivation through inhibition of viral transcriptional elongation. Front. Microbiol. 2017, 8, 2007. [Google Scholar] [CrossRef]
- Zhou, C.L.; Huang, Y.F.; Li, Y.B.; Liang, T.Z.; Zheng, T.Y.; Chen, P.; Wu, Z.Y.; Lai, F.Y.; Liu, S.W.; Xi, B.M.; et al. A new small molecule compound, Q308, silences latent HIV-1 provirus by suppressing Tat- and FACT-mediated transcription. Antimicrob. Agents Chemother. 2021, 65, e00470-21. [Google Scholar] [CrossRef]
- Chang, H.W.; Valieva, M.E.; Safina, A.; Chereji, R.V.; Wang, J.; Kulaeva, O.I.; Morozov, A.V.; Kirpichnikov, M.P.; Feofanov, A.V.; Gurova, K.V.; et al. Mechanism of FACT removal from transcribed genes by anticancer drugs curaxins. Sci. Adv. 2018, 4, eaav2131. [Google Scholar] [CrossRef]
- Huang, H.; Santoso, N.; Power, D.; Simpson, S.; Dieringer, M.; Miao, H.; Gurova, K.; Giam, C.Z.; Elledge, S.J.; Zhu, J. FACT proteins, SUPT16H and SSRP1, are transcriptional suppressors of HIV-1 and HTLV-1 that facilitate viral latency. J. Biol. Chem. 2015, 290, 27297–27310. [Google Scholar] [CrossRef]
- Debyser, Z.; Bruggemans, A.; Van Belle, S.; Janssens, J.; Christ, F. LEDGINs, inhibitors of the interaction between HIV-1 integrase and LEDGF/p75, are potent antivirals with a potential to cure HIV infection. Adv. Exp. Med. Biol. 2021, 1322, 97–114. [Google Scholar]
- Singh, P.K.; Li, W.; Bedwell, G.J.; Fadel, H.J.; Poeschla, E.M.; Engelman, A.N. Allosteric integrase inhibitor influences on HIV-1 integration and roles of LEDGF/p75 and HDGFL2 host factors. Viruses 2022, 14, 1883. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.P.; Ohguro, N.; Kikuchi, T.; Chylack, L.T., Jr.; Shinohara, T. Lens epithelium-derived growth factor: Effects on growth and survival of lens epithelial cells, keratinocytes, and fibroblasts. Biochem. Biophys. Res. Commun. 2000, 267, 373–381. [Google Scholar] [CrossRef] [PubMed]
- De Rijck, J.; Bartholomeeusen, K.; Ceulemans, H.; Debyser, Z.; Gijsbers, R. High-resolution profiling of the LEDGF/p75 chromatin interaction in the ENCODE region. Nucleic Acids Res. 2010, 38, 6135–6147. [Google Scholar] [CrossRef] [PubMed]
- Lapaillerie, D.; Lelandais, B.; Mauro, E.; Gozalo, A.; Gharbi, M.; Dufour, C.; Delelis, O.; Saïb, A.; Ruff, M.; Mély, Y.; et al. Modulation of the intrinsic chromatin binding property of HIV-1 integrase by LEDGF/p75. Nucleic Acids Res. 2021, 49, 11241–11256. [Google Scholar] [CrossRef]
- Eidahl, J.O.; Crowe, B.L.; North, J.A.; McKee, C.J.; Shkriabai, N.; Feng, L.; Plumb, M.; Graham, R.L.; Gorelick, R.J.; Hess, S.; et al. Structural basis for high-affinity binding of LEDGF PWWP to mononucleosomes. Nucleic Acids Res. 2013, 41, 3924–3936. [Google Scholar] [CrossRef]
- Maehigashi, T.; Ahn, S.; Kim, U.I.; Lindenberger, J.; Oo, A.; Koneru, P.C.; Mahboubi, B.; Engelman, A.N.; Kvaratskhelia, M.; Kim, K; et al. A highly potent and safe pyrrolopyridine-based allosteric HIV-1 integrase inhibitor targeting host LEDGF/p75 integrase interaction site. PLoS Pathog. 2021, 17, e1009671. [Google Scholar] [CrossRef]
- Christ, F.; Shaw, S.; Demeulemeester, J.; Desimmie, B.A.; Marchand, A.; Butler, S.; Smets, W.; Chaltin, P.; Westby, M.; Debyser, Z.; et al. Small-molecule inhibitors of the LEDGF/p75 binding site of integrase block HIV replication and modulate integrase multimerization. Antimicrob. Agents Chemother. 2012, 56, 4365–4374. [Google Scholar] [CrossRef]
- Xue, W.; Liu, H.; Yao, X. Molecular modeling study on the allosteric inhibition mechanism of HIV-1 integrase by LEDGF/p75 binding site inhibitors. PLoS ONE 2014, 9, e90799. [Google Scholar] [CrossRef]
- Bonnard, D.; Le Rouzic, E.; Singer, M.R.; Yu, Z.; Le Strat, F.; Batisse, C.; Batisse, J.; Amadori, C.; Chasset, S.; Pye, V.E.; et al. Biological and structural analyses of new potent allosteric inhibitors of HIV-1 integrase. Antimicrob. Agents Chemother. 2023, 67, e00462-23. [Google Scholar] [CrossRef]
- Vranckx, L.S.; Demeulemeester, J.; Saleh, S.; Boll, A.; Vansant, G.; Schrijvers, R.; Weydert, C.; Battivelli, E.; Verdin, E.; Cereseto, A.; Christ, F.; Gijsbers, R.; Debyser, Z. LEDGIN-mediated inhibition of integrase–LEDGF/p75 interaction reduces reactivation of residual latent HIV. EBioMedicine 2016, 8, 248–264. [Google Scholar] [CrossRef]
- Meng, X.; et al. Results from a proof-of-concept clinical trial of pirmitegravir, an investigational HIV-1 allosteric integrase inhibitor (ALLINI). Proceedings of IDWeek 2025, San Diego, CA, USA, 19–22 October 2025. [Google Scholar]
- HIVIH. A single ascending dose trial investigating the safety, tolerability and pharmacokinetics of orally administered BDM-2 in healthy male subjects. NCT03634085; ClinicalTrials.gov. 2020. [Google Scholar]
- Chhokar, N.; Kalra, S.; Chauhan, M.; Munshi, A.; Kumar, R. Quinoline-based protein-protein interaction inhibitors of LEDGF/p75 and HIV integrase: An in silico study. Curr. Top. Med. Chem. 2018, 18, 2800–2815. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Lin, H.Q.; Wang, P.; Hu, J.S.; Ip, T.M.; Yang, L.M.; Zheng, Y.T.; Wan, D.C.-C. Discovery of a novel HIV-1 integrase/p75 interacting inhibitor by docking screening, biochemical assay, and in vitro studies. J. Chem. Inf. Model. 2017, 57, 2336–2343. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.H.; Debnath, B.; Xu, Z.L.; Yang, L.M.; Song, L.R.; Zheng, Y.T.; Neamati, N.; Long, Y.Q. Discovery of novel 3-hydroxypicolinamides as selective inhibitors of HIV-1 integrase-LEDGF/p75 interaction. Eur. J. Med. Chem. 2017, 125, 1051–1063. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.W.; Yan, H.L.; Xu, X.S.; Xu, L.; Yin, Z.H.; Chang, S.; Luo, H. The selenium-containing drug ebselen potently disrupts LEDGF/p75 HIV-1 integrase interaction by targeting LEDGF/p75. J. Enzyme Inhib. Med. Chem. 2020, 35, 906–912. [Google Scholar] [CrossRef]
- Roberts, R.A.; Campbell, R.A.; Sikakana, P.; Sadler, C.; Osier, M.; Xu, Y.; Feng, J.Y.; Mitchell, M.; Sakowicz, R.; Chester, A.; et al. Species-specific urothelial toxicity with an anti-HIV noncatalytic site integrase inhibitor (NCINI) is related to unusual pH-dependent physicochemical changes. Toxicol. Sci. 2021, 183, 105–116. [Google Scholar] [CrossRef]
- Li, W.; Singh, P.K.; Sowd, G.A.; Fadel, H.J.; Cote, M.L.; Valente, S.T.; Engelman, A.N. CPSF6-dependent targeting of speckle-associated domains distinguishes primate from nonprimate lentiviral integration. mBio 2020, 11, e02254-20. [Google Scholar] [CrossRef]
- Chaudhuri, E.; Jang, S.; Chakraborty, R.; Li, W.; Singh, P.K.; Sowd, G.A.; Engelman, A.N. CPSF6 promotes HIV-1 preintegration complex function. J. Virol. 2025, 99, e00490-25. [Google Scholar] [CrossRef]
- Jang, S.; Cook, N.J.; Pye, V.E.; Regan, L.; Barford, D.; Yang, J.; Zhang, C.; Fischer, U.; Hennig, J. Differential role for phosphorylation in alternative polyadenylation function versus nuclear import of SR-like protein CPSF6. Nucleic Acids Res. 2019, 47, 4663–4683. [Google Scholar] [CrossRef]
- Liu, L.; Manley, J.L. Modulation of diverse biological processes by CPSF, the master regulator of mRNA 3' ends. RNA 2024, 30, 1122–1140. [Google Scholar] [CrossRef]
- Cornish, D.; Jackson-Jones, K.A.; Ling-Hu, T.; Simons, L.; Cisneros, W.; Kuffour, E.; Agnes, F.; Lee, Y.; Bieniasz, P.; Lorenzo-Redondo, R.; Hultquist, J. Alternative polyadenylation upon CPSF6 knock-out enhances HIV-1 infection in primary T cells. PLoS Pathog. 2025, 21, e1013745. [Google Scholar] [CrossRef]
- Bialas, K.; Diaz-Griffero, F. HIV-1-induced translocation of CPSF6 to biomolecular condensates. Trends Microbiol. 2024, 32, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Luchsinger, C.; Lee, K.; Mardones, G.A.; KewalRamani, V.N.; Diaz-Griffero, F. Formation of nuclear CPSF6/CPSF5 biomolecular condensates upon HIV-1 entry into the nucleus is important for productive infection. Sci. Rep. 2023, 13, 10974. [Google Scholar] [CrossRef] [PubMed]
- Ge, Y.; Huang, J.; Chen, R.; Fu, Y.; Ling, T.; Ou, X.; Rong, X.; Cheng, Y.; Lin, Y.; Zhou, F.; Lu, C.; Yuan, S.; Xu, A. Downregulation of CPSF6 leads to global mRNA 3' UTR shortening and enhanced antiviral immune responses. PLoS Pathog. 2024, 20, e1012061. [Google Scholar] [CrossRef] [PubMed]
- Rohlfes, N.; Radhakrishnan, R.; Singh, P.K.; Li, W.; Sowd, G.A.; Engelman, A.N. The nuclear localization signal of CPSF6 governs post-nuclear import steps of HIV-1 infection. PLoS Pathog. 2025, 21, e1012354. [Google Scholar] [CrossRef]
- Martin, C.J.; Oser, E.A.; Nagarajan, P.; Popova, L.V.; Sunkel, B.D.; Stanton, B.Z.; Parthun, M.R. Distinct classes of lamina-associated domains are defined by differential patterns of repressive histone methylation. Genome Res. 2025, 35, 1959–1974. [Google Scholar] [CrossRef]
- Lochs, S.J.A.; Kefalopoulou, S.; Kind, J. Lamina associated domains and gene regulation in development and cancer. Cells 2019, 8, 271. [Google Scholar] [CrossRef]
- van Steensel, B.; Belmont, A.S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 2017, 169, 780–791. [Google Scholar] [CrossRef]
- Yáñez-Cuna, J.O.; van Steensel, B. Genome-nuclear lamina interactions: from cell populations to single cells. Curr. Opin. Genet. Dev. 2017, 43, 67–72. [Google Scholar] [CrossRef]
- van Schaik, T.; Vos, M.; Peric-Hupkes, D.; Hn Celie, P.; van Steensel, B. Cell cycle dynamics of lamina-associated DNA. EMBO Rep. 2020, 21, e50193. [Google Scholar] [CrossRef]
- Kaczmarczyk, L.S.; Levi, N.; Segal, T.; Salmon-Divon, M.; Gerlitz, G. CTCF supports preferentially short lamina-associated domains. Chromosome Res. 2022, 30, 123–136. [Google Scholar] [CrossRef]
- Wilson, M.D.; Renault, L.; Maskell, D.P.; Vora, M.; Lehmann, L.C.; Halic, M.; Balasubramanian, S.; Cugliandolo, F.M.; Engelman, A.N.; Skalka, A.M.; et al. Retroviral integration into nucleosomes through DNA looping and sliding along the histone octamer. Nat. Commun. 2019, 10, 4189. [Google Scholar] [CrossRef] [PubMed]
- Emerson, R.O.; Thomas, J.H. Gypsy and the birth of the SCAN domain. J. Virol. 2011, 85, 12043–12052. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Schubert, H.L.; Singh, P.K.; He, S.; Kaur, S.; Chen, P.; Li, W.; Sowd, G.A.; Engelman, A.N. Cleavage and polyadenylation specificity factor 6 is required for efficient HIV-1 latency reversal. mBio 2021, 12, e01098-21. [Google Scholar] [CrossRef] [PubMed]
- Jang, S.; Engelman, A.N. Capsid-host interactions for HIV-1 ingress. Microbiol. Mol. Biol. Rev. 2023, 87, e00048-22. [Google Scholar] [CrossRef]
- Li, C.; Burdick, R.C.; Siddiqui, R.; Vora, M.; Renault, L.; Maskell, D.P.; Lehmann, L.C.; Halic, M.; Engelman, A.N.; Skalka, A.M.; et al. Lenacapavir disrupts HIV-1 core integrity while stabilizing the capsid lattice. Proc. Natl. Acad. Sci. USA 2025, 122, e2420497122. [Google Scholar] [CrossRef]
- Müller, T.G.; Klaus, S.; Zila, V.; Lucic, B.; Penzo, C.; Nopper, S.L.; Golani, G.; Anders-Össwein, M.; Sonntag-Buck, V.; Heuser, A.M.; et al. Lenacapavir-induced capsid damage uncovers HIV-1 genomes emanating from nuclear speckles. EMBO J. 2026, 45, 449–470. [Google Scholar] [CrossRef]
- Dragoni, F.; Kwaa, A.K.; Traut, C.C.; Veenhuis, R.T.; Woldemeskel, B.A.; Camilo-Contreras, A.; Raymond, H.E.; Dykema, A.G.; Scully, E.P.; Rosecrans, A.M.; et al. Proviral location affects cognate peptide-induced virus production and immune recognition of HIV-1-infected T cell clones. J. Clin. Invest. 2023, 133, e171097. [Google Scholar] [CrossRef]
- Halvas, E.K.; Joseph, K.W.; Brandt, L.D.; Guo, S.; Sobolewski, M.D.; Jacobs, J.L.; Tumiotto, C.; Bui, J.K.; Cyktor, J.C.; Keele, B.F.; et al. HIV-1 viremia not suppressible by antiretroviral therapy can originate from large T cell clones producing infectious virus. J. Clin. Invest. 2020, 130, 5847–5857. [Google Scholar] [CrossRef]
- Marini, B.; Kertesz-Farkas, A.; Ali, H.; Lucic, B.; Lisek, K.; Manganaro, L.; Pongor, S.; Luzzati, R.; Recchia, A.; Mavilio, F.; et al. Nuclear architecture dictates HIV-1 integration site selection. Nature 2015, 521, 227–231. [Google Scholar] [CrossRef]
- Battivelli, E.; Dahabieh, M.S.; Abdel-Mohsen, M.; Svensson, J.P.; Tojal Da Silva, I.; Cohn, L.B.; Gramatica, A.; Deeks, S.; Greene, W.C.; Pillai, S.K.; Verdin, E. Distinct chromatin functional states correlate with HIV latency reactivation in infected primary CD4+ T cells. eLife 2018, 7, e34655. [Google Scholar] [CrossRef]
- Jiang, C.; Lian, X.; Gao, C.; Sun, X.; Einkauf, K.B.; Chevalier, J.M.; Chen, S.M.Y.; Hua, S.; Rhee, B.; Chang, K.; et al. Distinct viral reservoirs in individuals with spontaneous control of HIV-1. Nature 2020, 585, 261–267. [Google Scholar] [CrossRef]
- Ecco, G.; Imbeault, M.; Trono, D. KRAB zinc finger proteins. Development 2017, 144, 2719–2729. [Google Scholar] [CrossRef] [PubMed]
- Manzo, S.G.; Mazouzi, A.; Leemans, C.; van Schaik, T.; Neyazi, N.; van Ruiten, M.S.; Rowland, B.D.; Brummelkamp, T.R.; van Steensel, B. Chromatin protein complexes involved in gene repression in lamina-associated domains. EMBO J. 2024, 43, 5260–5287. [Google Scholar] [CrossRef] [PubMed]
- Caruso, L.B.; Guo, R.; Keith, K.; Tempera, I.; Gewurz, B.E. The nuclear lamina binds the Epstein-Barr virus genome during latency and regulates viral gene expression. PLoS Pathog. 2022, 18, e1010400. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.D.; Tanizawa, H.; De Leo, A.; Luo, Y.; Huang, L.; Kossenkov, A.V.; Li, H.; Ishii, K.J.; Lieberman, P.M. Epigenetic specifications of host chromosome docking sites for latent Epstein-Barr virus. Nat. Commun. 2020, 11, 877. [Google Scholar] [CrossRef]
- Ramasubramanyan, S.; Osborn, K.; Flower, K.; Sinclair, A.J. Dynamic chromatin environment of key lytic cycle regulatory regions of the Epstein-Barr virus genome. J. Virol. 2012, 86, 1809–1819. [Google Scholar] [CrossRef]
- Buschle, A.; Hammerschmidt, W. Epigenetic lifestyle of Epstein-Barr virus. Semin. Immunopathol. 2020, 42, 131–142. [Google Scholar] [CrossRef]
- Murata, T.; Sugimoto, A.; Inagaki, T.; Yanagi, Y.; Watanabe, T.; Sato, Y.; Kimura, H. Molecular basis of Epstein-Barr virus latency establishment and lytic reactivation. Viruses 2021, 13, 2344. [Google Scholar] [CrossRef]
- De Leo, A.; Calderon, A.; Lieberman, P.M. Control of viral latency by episome maintenance proteins. Trends Microbiol. 2020, 28, 150–162. [Google Scholar] [CrossRef]
- Cohen, J.I. Herpesvirus Latency. J. Clin. Investig. 2020, 130, 3361–3369. [Google Scholar] [CrossRef]
- Lieberman, P.M. Epigenetics and genetics of viral latency. Cell Host Microbe 2016, 19, 619–628. [Google Scholar] [CrossRef]
- Luchsinger, C.; Dai, A.Z.; Yalamanchili, H.; Lee, K.; Mardones, G.A.; KewalRamani, V.N.; Diaz-Griffero, F. HIV-1 infection regulates gene expression by altering alternative polyadenylation correlated with CPSF6 and CPSF5 redistribution. mBio 2025, 16, e02865-25. [Google Scholar] [CrossRef]
- Tomasini, C.; Cuche, C.; Ay, S.; Collard, M.; Cui, B.; Rashid, M.; Bhattacharjee, S.; Tello-Rubio, B.; Buchrieser, J.; Luchsinger, C.; et al. Decoding the biogenesis of HIV-induced CPSF6 puncta and their fusion with nuclear speckles. Elife 2026, 13, RP103725. [Google Scholar] [CrossRef]
- Wei, G.; Iqbal, N.; Courouble, V.V.; Francis, A.C.; Singh, P.K.; Hudait, A.; Annamalai, A.S.; Bester, S.; Huang, S.W.; Shkriabai, N.; et al. Prion-like low complexity regions enable avid virus-host interactions during HIV-1 infection. Nat. Commun. 2022, 13, 5879. [Google Scholar] [CrossRef]
- Ding, D.; Xu, S.; Zhang, X.; Jiang, X.; Cocklin, S.; Dick, A.; Zhan, P.; Liu, X. The discovery and design of novel HIV-1 capsid modulators and future perspectives. Expert Opin. Drug Discov. 2023, 18, 5–12. [Google Scholar] [CrossRef]
- McFadden, W.M.; Faerch, M.; Kirby, K.A.; Dick, R.A.; Torbett, B.E.; Sarafianos, S.G. Considerations for capsid-targeting antiretrovirals in pre-exposure prophylaxis. Trends Mol. Med. 2025, 31, 801–813. [Google Scholar] [CrossRef]
| Group | Block-and-lock agent's name or strategy name | Mechanism/Target | References |
|---|---|---|---|
| Direct suppressors of HIV-1 transcriptional activity | Tat inhibitors, Triptolide, Camptothecin analogs, CRISPR–Cas systems. | Direct inhibition of viral transcription, splicing, or RNA stability. | [16,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73] |
| Post-transcriptional agents | Splicing inhibitors, RNA interference technologies. |
Sequence-specific silencing of HIV-1 gene expression through RNA degradation or targeted transcriptional repression, promoting durable viral latency. | [74,75,76,77,78,79,80,81,82,83,84,85] |
| Cellular transcription factor modulators. | BRD4 modulators, NF-κB inhibitors. | Inhibition of host transcription factors that activate HIV-1 LTR. | [86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111] |
| Cellular Kinase inhibitors. | PI3K–AKT–mTOR pathway modulators, Aurora kinase and PAK1/2 inhibitors, PKC inhibitors, CDK inhibitors, SR kinase inhibitors. | Block of cellular kinases/signaling required for HIV-1 transcription. | [112,113,114,115,116,117,118,119,120,121,122] |
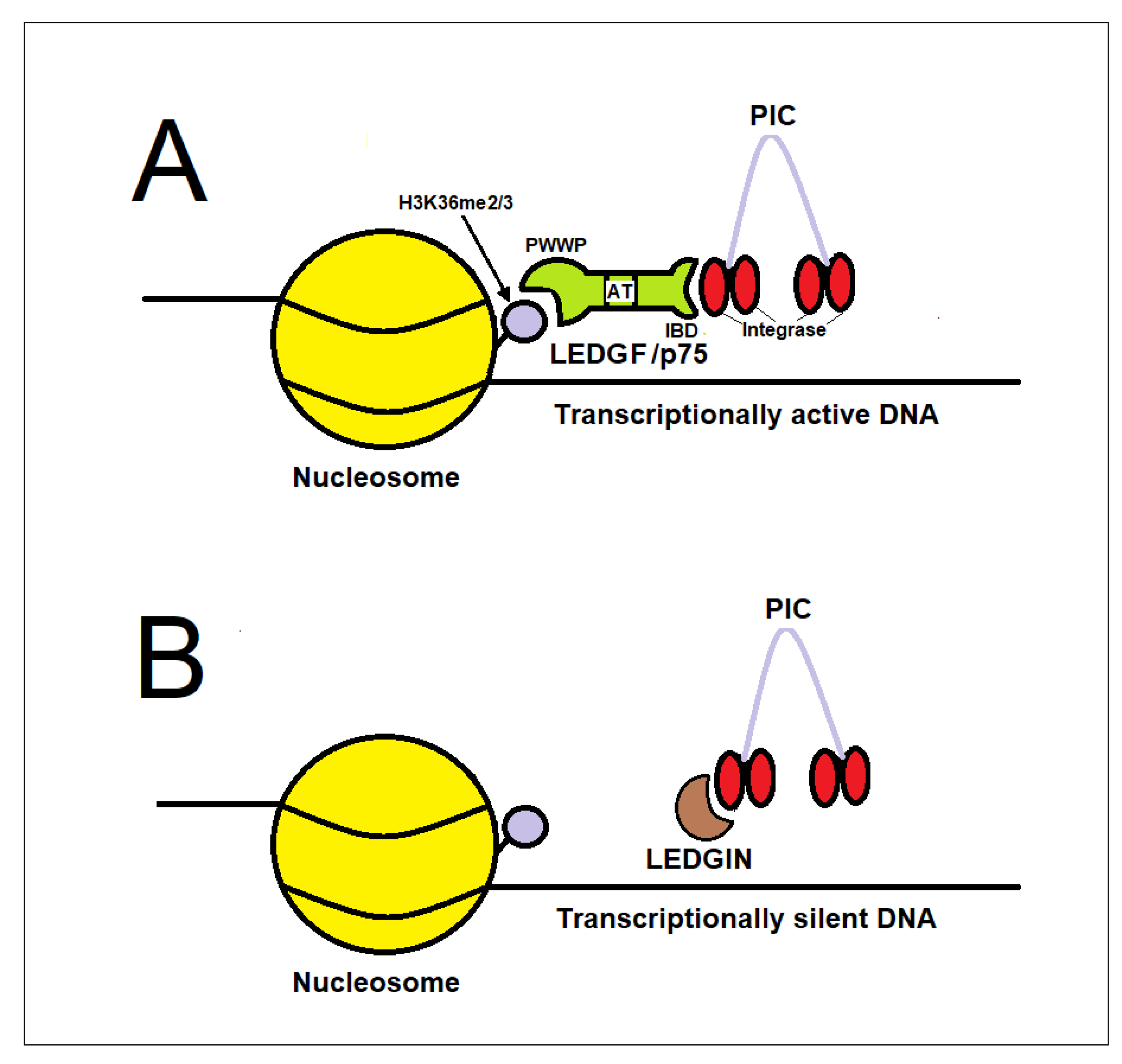
| Epigenetic, chromatin, and integration site modulators. | H3K27 demethylase inhibitors, Histone acetyltransferase (HAT) inhibitors, FACT targeting compounds, LEDGF/p75–integrase inhibitors (LEDGINs) | Induction of repressive chromatin and epigenetic silencing of provirus. Integration in transcriptionally silent chromatin | [123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159] |
| Intervention | Time to Viral Rebound | References |
|---|---|---|
| cART alone | 2–4 weeks (median time 16 days) in humans. 3–10 days in humanized mice of HIV-1 infection. |
[5,49] |
| dCA + cART | 10–19 days in humanized mice of HIV-1 infection. | [16,49] |
| Novel Tat inhibitors (1,3,4-oxadiazole derivatives, benzoxazole compounds, cyclic peptides, aromatic heterocycles) | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [53,54,55,56,57,58,59,60,61,62] |
| LLDT + cART | LLDT-8 + cART does not delay viral rebound in SIV-infected macaques after cART discontinuation. | [65,66] |
| Topotecan | No in vivo studies. In in vitro studies, suppression of HIV-1 reactivation persisted for ~72 hours after the drug was removed from the culture medium. | [67] |
| CRISPR–Cas (Targeted provirus editing) | Quantitative rebound kinetics are not yet well defined. | [68,69,70,71,72] |
| EBT-101 infusion (CRISPR excising HIV DNA) + cART in humans | In humans, ~2–4 weeks; one individual, ~16 weeks. | [73] |
| SF3B1 inhibitors | No in vivo studies. In in vitro studies, suppression of HIV-1 reactivation by sudemycin D6 persisted for ~72 hours after the drug was removed from the culture medium. | [74] |
| RNAi technologies | Quantitative rebound kinetics are not yet well defined. | [75,76,77,78,79,80,81,82,83,84,85] |
| ZL0580 | ~4 weeks in a humanized mouse model of HIV-1 infection. | [107] |
| NF-κB Inhibitors | No in vivo studies. Precise kinetics have not been systematically reported in standard in vitro latency models. | [109] |
| PI3K–AKT–mTOR Pathway Modulators | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [112,113] |
| Danusertib | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [116] |
| PF-3758309 | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [119] |
| Protein kinase C inhibitors | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [120] |
| CDK9 Inhibitors | No in vivo studies. In in vitro studies, the transcriptional suppression persists for at least 24 h after drug removal. | [121] |
| CDK8/19 Inhibitors | No in vivo studies. In in vitro studies, the transcriptional suppression persists for at least 7 days after drug removal. | [121] |
| CDK7 Inhibitors | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [121] |
| SR Kinase Inhibitors | No in vivo studies. No in vitro studies calculating the time of transcriptional suppression after drug removal. | [122] |
| H3K27 demethylase inhibitors, histone acetyltransferase (HAT) inhibitors | No in vivo studies. In in vitro studies, the transcriptional suppression lasts less than 72 hours after GSK-J4 removal. | [131,132,133,134] |
| CBL0100 | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [138] |
| Q308 | No in vivo studies. No in vitro studies have calculated the time of transcriptional suppression after drug removal. | [139] |
| LEDGINs | No in vivo or in vitro studies calculating the time of transcriptional suppression after drug removal. | [148,149,150,151,152,153,154,155,156,157,158,159] |
| Chromatin type | H3K9me2/3 | Gene density | Enrichment in nuclear B compartment* |
Compact/ structurally constrained environment |
|---|---|---|---|---|
| Euchromatin | ↓ | ↑↑↑ | ↓ | ↓ |
| Intranuclear heterochromatin | ↑↑ | ↓↓ | ↑↑ | ↑↑ |
| LADs | ↑↑↑ | ↓↓↓ | ↑↑↑ | ↑↑↑ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




