Submitted:
06 March 2026
Posted:
09 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Material and Methods
2.1. Instruments
2.2. Reagents and Chemicals
2.3. Procedures
2.3.1. Preparation of Solutions
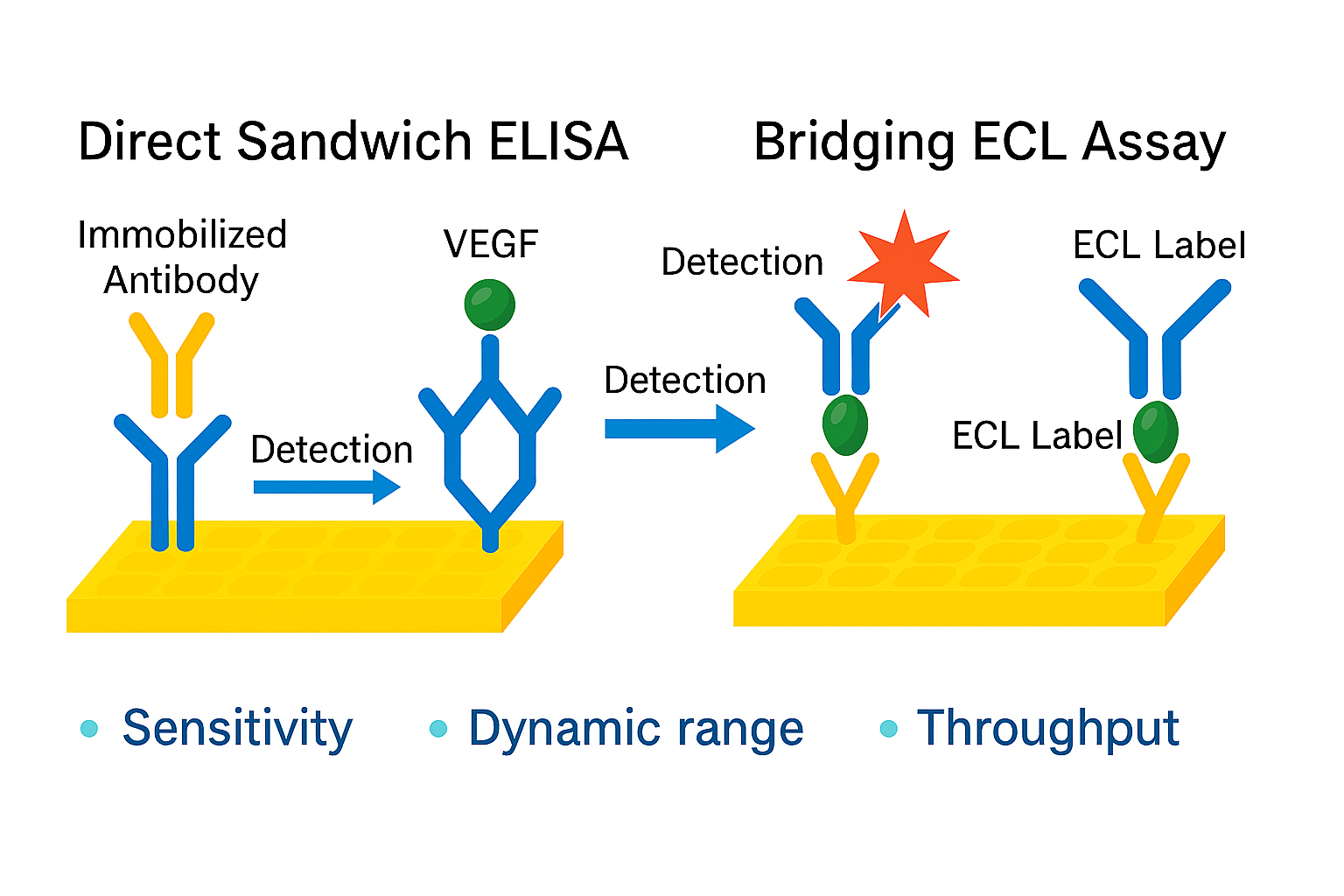
2.3.2. ELISA Methodology for VEGF Quantification [11,12]
2.3.3. ECL Methodology for VEGF Quantification [13,14]
2.3.4. Validation Parameters of Sandwich ELISA
2.3.4.1. Screening of Individual Plasma Samples
2.3.4.2. Data Analysis and Calibration Curve
2.3.4.3. Accuracy and Precision in Assay Buffer
2.3.4.4. Accuracy and Precision for Plasma Samples
2.3.4.5. Parallelism Evaluation
2.3.4.6. Minimal Required Dilution
2.3.4.7. Sensitivity
2.3.4.8. Stability Experiments
2.3.5. Study Sample Collection and Analysis
3. Results
3.1. ELISA Optimization for VEGF
3.1.1. Standard and Dilution Optimization
3.1.2. In-House Diluent Composition
3.1.3. Stop Solution and Wash Buffer Optimization
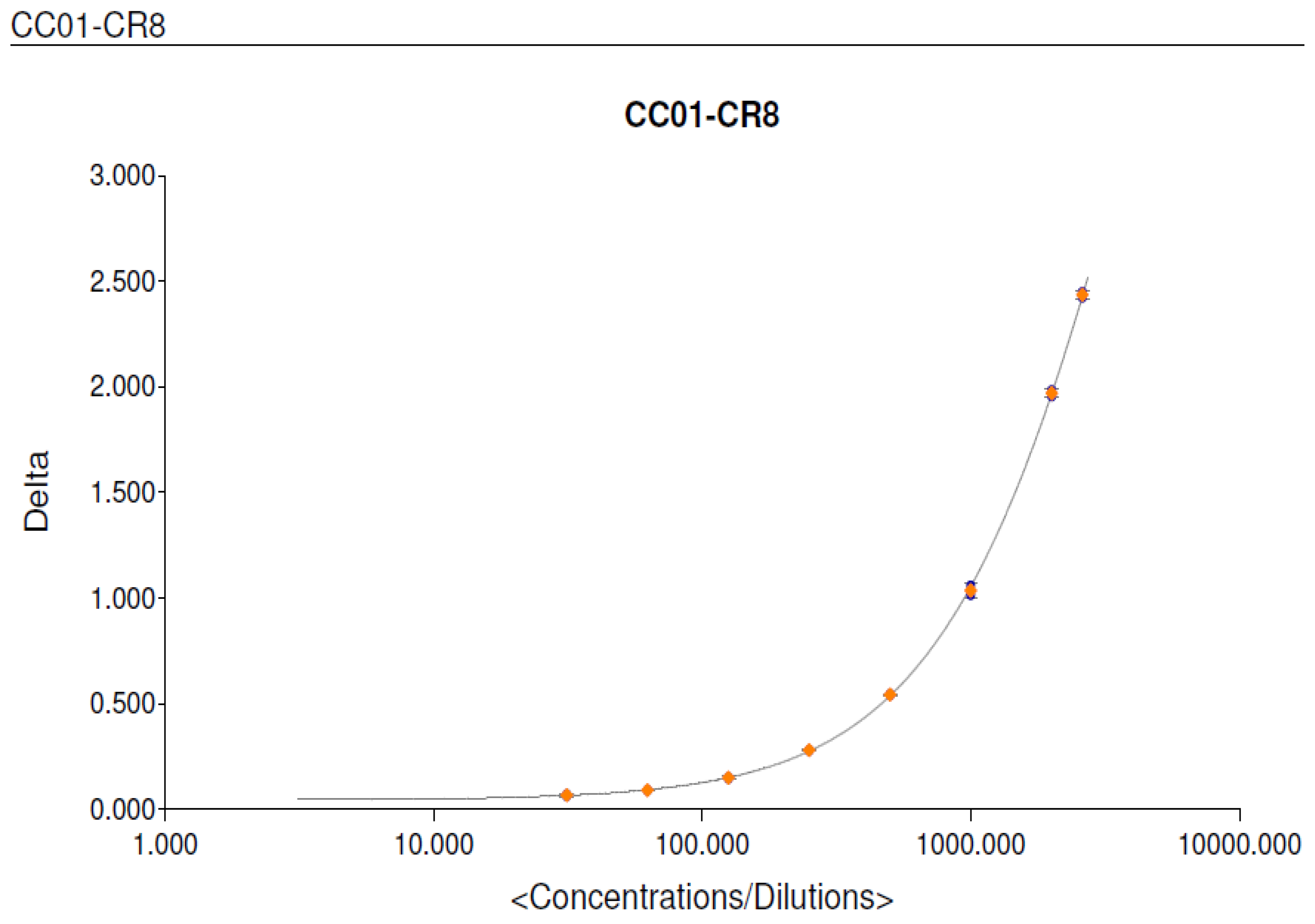
3.2. MSD ECL Assay Optimization for VEGF
3.3. Validation Results of the ELISA for VEGF
3.3.1. Screening of Individual Plasma Samples
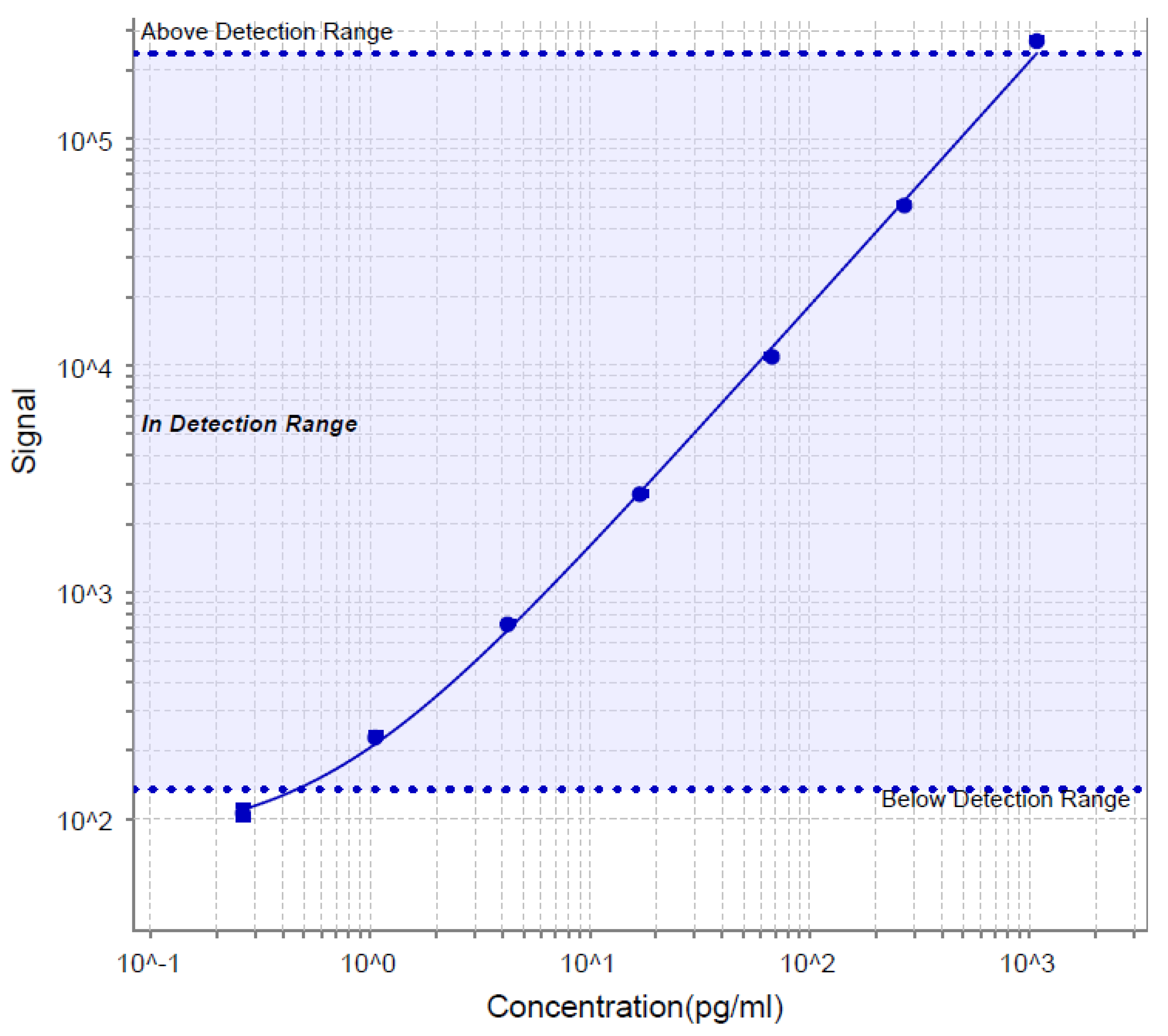
3.3.2. Calibration Curve (Linearity)
3.3.3. Accuracy and Precision in Assay Buffer
3.3.4. Accuracy and Precision for Plasma Samples
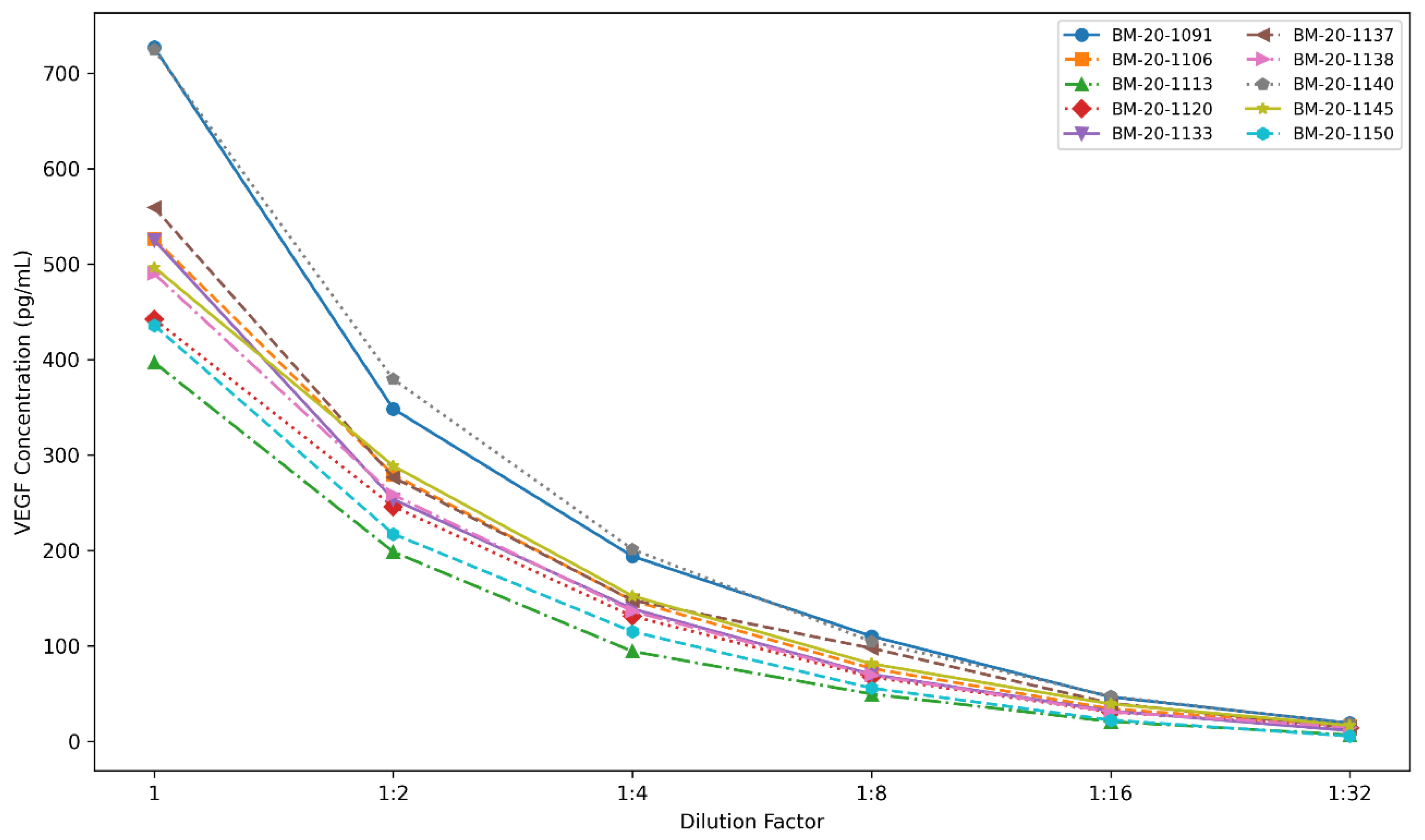
3.3.5. Parallelism Evaluation
3.3.6. Minimal Required Dilution
3.3.7. Sensitivity
3.3.8. Stability Experiments
3.4. Application of Validated Sandwich ELISA
4. Discussion
4.1. Strategy and Design for Assay Development
4.2. Comparative Assessment of PD Assays Developed for VEGF
5. Conclusions
Limitations and Future Directions
Author Contributions
Funding
Data Availability
Acknowledgments
Declaration of interests
Statement: on using Artificial Intelligence
References
- Zhao Y, Gu H, Postelnek J, et al (2020) Fit-for-purpose protein biomarker assay validation strategies using hybrid immunocapture-liquid chromatography-tandem-mass spectrometry platform: Quantitative analysis of total soluble cluster of differentiation 73. Anal Chim Acta 1126:144–153. [CrossRef]
- Strauss DG, Wang YM, Florian J, Zineh I (2023) Pharmacodynamic Biomarkers Evidentiary Considerations for Biosimilar Development and Approval. Clin Pharmacol Ther 113:55–61. [CrossRef]
- Ho QT, Kuo CJ (2007) Vascular endothelial growth factor: Biology and therapeutic applications. International Journal of Biochemistry and Cell Biology 39:1349–1357. [CrossRef]
- Zheng X, Gao S, Wu J, Hu X (2021) A Fluorescent Aptasensor Based on Assembled G-Quadruplex and Thioflavin T for the Detection of Biomarker VEGF165. Front Bioeng Biotechnol 9:764123. [CrossRef]
- Göpfert JC, Reiser A, Yañez VAC, et al (2019) Development and evaluation of an ultrasensitive free VEGF-A immunoassay for analysis of human aqueous humor. Bioanalysis 11:875–886. [CrossRef]
- Sumner G, Georgaros C, Rafique A, et al (2019) Anti-VEGF drug interference with VEGF quantitation in the R&D systems human quantikine VEGF ELISA kit. Bioanalysis 11:381–392. [CrossRef]
- Zhou Q, Yan H, Ran F, et al (2018) Ultrasensitive enzyme-free fluorescent detection of VEGF165 based on target-triggered hybridization chain reaction amplification. RSC Adv 8:25955–25960. [CrossRef]
- Takahashi H, Nomura Y, Nishida J, et al (2016) Vascular Endothelial Growth Factor (VEGF) Concentration Is Underestimated by Enzyme-Linked Immunosorbent Assay in the Presence of Anti-VEGF Drugs. Invest Ophthalmol Vis Sci 57:462–466. [CrossRef]
- Panoilia E, Schindler E, Samantas E, et al (2015) A pharmacokinetic binding model for bevacizumab and VEGF 165 in colorectal cancer patients. Cancer Chemother Pharmacol 3:791–803. [CrossRef]
- Avery RL, Castellarin AA, Steinle NC (2014) Systemic pharmacokinetics following intravitreal injections of ranibizumab, bevacizumab or aflibercept in patients with neovascular AMD. Br J Ophthalmol 98:1636–1641. [CrossRef]
- Usa, Canada, China Human VEGF Immunoassay Quantikine ® ELISA Catalog Number DVE00 Catalog Number SVE00 Catalog Number PDVE00 For the quantitative determination of human Vascular Endothelial Growth Factor (VEGF) concentrations in cell culture supernates, serum, and plasma. ….
- Quantikine® Colorimetric Sandwich ELISA Kit Assay Principle: R&D Systems. https://www.rndsystems.com/products/quantikine-colorimetric-sandwich-elisa-assay-principle. Accessed 2 Jan 2026.
- V-PLEX Plus Human VEGF Kit | Meso Scale Discovery. https://www.mesoscale.com/en/products/v-plex-plus-human-vegf-kit-k151rhg/. Accessed 2 Jan 2026.
- Why Electrochemiluminescence | Meso Scale Discovery. https://punchout.mesoscale.com/en/technical_resources/our_technology/ecl. Accessed 2 Jan 2026.
- Food and Drug Administration (FDA). Guidance for Industry, Bioanalytical Method Validation, Rockville, MD: US Department of Health and Human Services, FDA, Center for Drug Evaluation and Research: May 2018.
- (2022) ICH M10 Harmonised Guideline Bioanalytical Method Validation and Study sample analysis.
- Marini JC, Anderson M, Cai XY, et al (2014) Systematic Verification of Bioanalytical Similarity Between a Biosimilar and a Reference Biotherapeutic: Committee Recommendations for the Development and Validation of a Single Ligand-Binding Assay to Support Pharmacokinetic Assessments. Botanical Review 16:1149–1158. [CrossRef]
- Thway TM, Wang YM, Booth BP, et al (2020) Current Perspectives on Ligand-Binding Assay Practices in the Quantification of Circulating Therapeutic Proteins for Biosimilar Biological Product Development. AAPS J 22:. [CrossRef]
- Piccoli SP, Michael Sauer J, Ackermann B, et al (2019) Points to Consider Document: Scientific and Regulatory Considerations for the Analytical Validation of Assays Used in the Qualification of Biomarkers in Biological Matrices Biomarker Assay Collaborative Evidentiary Considerations Writing Group, Critical Path Institute (C-Path).
- Stevenson LF, Purushothama S (2014) Parallelism: Considerations for the development, validation and implementation of PK and biomarker ligand-binding assays. Bioanalysis 6:185–198. [CrossRef]
- Tu J, Bennett P (2017) Parallelism experiments to evaluate matrix effects, selectivity and sensitivity in ligand-binding assay method development: Pros and cons. Bioanalysis 9:1107–1122. [CrossRef]
- Islam R, Kar S, Ritzén H, et al (2019) Recommendations for classification of commercial LBA kits for biomarkers in drug development from the GCC for bioanalysis. Bioanalysis 11:645–653. [CrossRef]
- Rhyne PW, Wong OT, Zhang YJ, Weiner RS (2009) Electrochemiluminescence in bioanalysis. Bioanalysis 1:919–935. [CrossRef]
- Wyant T, Yang L, Rosario M (2021) Comparison of the ELISA and ECL Assay for Vedolizumab Anti-drug Antibodies: Assessing the Impact on Pharmacokinetics and Safety Outcomes of the Phase 3 GEMINI Trials. AAPS Journal 23:1–10. [CrossRef]
- Collet-Brose J, Couble P-J, Deehan MR, et al (2016) Evaluation of Multiple Immunoassay Technology Platforms to Select the Anti-Drug Antibody Assay Exhibiting the Most Appropriate Drug and Target Tolerance. [CrossRef]
| Calibration Standards | Spiked Concentration (pg/mL) | Measured Mean Concentration (pg/mL) | Inter-Assay Precision (% CV) | Inter-Assay Accuracy (% Bias) | Total Error (% TE) | |
|---|---|---|---|---|---|---|
| STD 1 | 2600 | Upper Anchor point | ||||
| STD 2 / ULOQ | 2000 | 1979.1 | 1.18 | -1.04 | 2.22 | |
| STD 3 | 1000 | 1002.0 | 1.94 | 0.21 | 2.15 | |
| STD 4 | 500 | 506.43 | 1.86 | 1.29 | 3.15 | |
| STD 5 | 250 | 248.8 | 1.61 | -0.46 | 2.07 | |
| STD 6 | 125 | 121.4 | 1.98 | -2.85 | 4.83 | |
| STD 7/ LLOQ | 62.5 | 65.8 | 5.01 | 5.39 | 10.40 | |
| STD 8 | 31.25 | Lower Anchor point | ||||
| Quality Control | Intra-run precision (% CV) | Intra-run accuracy | Intra-run % Bias |
|---|---|---|---|
| ULOQQC | 3.69% to 12.29% | 86.83% to 98.23% | 13.17% to -1.77% |
| HQC | 1.70% to 3.99% | 89.76% to 94.24% | -10.24% to -5.76% |
| MQC | 2.39% to 4.45% | 90.91% to 105.85% | -9.09% to 5.85% |
| LQC | 3.77% to 7.14% | 91.62% to 107.41% | -8.38% to 7.41% |
| LLOQQC | 5.06% to14.39% | 91.32% to 106.21% | -8.68% to 6.21% |
| Quality Control | Spiked Concentration (pg/mL) | Measured Mean Concentration (pg/mL) | Inter-Assay Precision (% CV) |
Inter-Assay Accuracy (% Bias) | Total Error (% TE) |
|---|---|---|---|---|---|
| ULOQQC | 2000 | 1879.5 | 7.40 | -6.02 | 13.42 |
| HQC | 1500 | 1385.4 | 3.44 | -7.64 | 11.08 |
| MQC | 350 | 346.7 | 6.14 | -0.93 | 7.07 |
| LQC | 180 | 180.6 | 7.58 | 0.36 | 7.94 |
| LLOQQC | 62.5 | 61.9 | 10.01 | -0.91 | 10.92 |
| Level | PM-01 | PM-02 |
|---|---|---|
| Mean VEGF Conc. | 787.4 pg/mL | 468.5 pg/mL |
| SD | 44.9 | 31.6 |
| % CV | 5.70 | 6.74 |
| n | 36 | 36 |
| Mean CV | 6.22 | |
| Range (3* Mean CV) | 18.66 | |
| Upper Limit (100+Range) | 118.66% | |
| Lower Limit(100-Range) | 81.34% | |
| Analyte Name | VEGF | ||||||||||||||||||||
| Individual Plasma ID | BM-20-1091 | BM-20-1106 | BM-20-1113 | BM-20-1120 | BM-20-1133 | BM-20-1137 | BM-20-1138 | BM-20-1140 | BM-20-1145 | BM-20-1150 | |||||||||||
| Dilution Fold | Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold | %Accuracy | |
| % Accuracy using no Dilution | 1 | 727.0 | 100.00 | 526.6 | 100.00 | 397.0 | 100.00 | 442.4 | 100.00 | 525.0 | 100.00 | 559.3 | 100.00 | 489.8 | 100.00 | 724.5 | 100.00 | 496.7 | 100.00 | 435.9 | 100.00 |
| 2 | 696.7 | 95.82 | 558.9 | 106.14 | 397.3 | 100.09 | 492.1 | 111.25 | 507.3 | 96.63 | 553.3 | 98.94 | 516.8 | 105.50 | 758.8 | 104.74 | 577.2 | 116.22 | 434.8 | 99.76 | |
| 4 | 776.2 | 106.76 | 589.9 | 112.02 | 377.7 | 95.15 | 525.2 | 118.73 | 556.0 | 105.90 | 592.1 | 105.88 | 545.9 | 111.45 | 805.4 | 111.17 | 610.0 | 122.81 | 460.4 | 105.62 | |
| 8 | 882.5 | 121.38 | 612.5 | 116.31 | 397.6 | 100.15 | 540.2 | 122.13 | 563.3 | 107.29 | 781.7 | 139.78 | 555.5 | 113.41 | 837.4 | 115.59 | 651.7 | 131.21 | 447.6 | 102.69 | |
| 16 | 746.4 | 102.66 | 548.5 | 104.16 | 334.8 | 84.34 | 493.8 | 111.63 | 511.9 | 97.50 | 642.4 | 114.86 | 497.2 | 101.51 | 761.2 | 105.07 | 628.5 | 126.54 | 360.7 | 82.76 | |
| 32 | 618.5 | 85.07 | 518.7 | 98.51 | 221.9 | 55.90 | 457.1 | 103.33 | 363.2 | 69.19 | 460.3 | 82.31 | 479.8 | 97.96 | 600.6 | 82.90 | 539.8 | 108.68 | 179.2 | 41.11 | |
| % Accuracy using 1 in 2 Dilution | 1 | 104.36 | 94.22 | 99.91 | 89.89 | 103.49 | 101.07 | 94.79 | 95.47 | 86.04 | 100.24 | ||||||||||
| 2 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | |||||||||||
| 4 | 111.42 | 105.54 | 95.07 | 106.72 | 109.60 | 107.01 | 105.64 | 106.14 | 105.67 | 105.88 | |||||||||||
| 8 | 126.67 | 109.59 | 100.06 | 109.77 | 111.04 | 141.27 | 107.50 | 110.35 | 112.90 | 102.94 | |||||||||||
| 16 | 107.14 | 98.13 | 84.26 | 100.34 | 100.90 | 116.09 | 96.22 | 100.32 | 108.88 | 82.96 | |||||||||||
| 32 | 88.78 | 92.81 | 55.85 | 92.88 | 71.60 | 83.19 | 92.85 | 79.15 | 93.51 | 41.21 | |||||||||||
| % Accuracy using 1 in 4 Dilution |
1 | 93.67 | 89.27 | 105.09 | 84.23 | 94.43 | 94.45 | 89.73 | 89.95 | 81.42 | 94.68 | ||||||||||
| 2 | 89.75 | 94.75 | 105.19 | 93.71 | 91.24 | 93.45 | 94.66 | 94.22 | 94.63 | 94.44 | |||||||||||
| 4 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | |||||||||||
| 8 | 113.69 | 103.84 | 105.25 | 102.86 | 101.31 | 132.02 | 101.77 | 103.97 | 106.84 | 97.23 | |||||||||||
| 16 | 96.16 | 92.98 | 88.63 | 94.02 | 92.06 | 108.48 | 91.08 | 94.52 | 103.03 | 78.35 | |||||||||||
| 32 | 79.68 | 87.94 | 58.74 | 87.03 | 65.33 | 77.74 | 87.90 | 74.57 | 88.49 | 38.92 | |||||||||||
| Individual ID | BM-20-1091 | BM-20-1106 | BM-20-1113 | BM-20-1120 | BM-20-1133 | BM-20-1137 | BM-20-1138 | BM-20-1140 | BM-20-1145 | BM-20-1150 | |||||||||||
| Dilution Fold | Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold | %Accuracy | Conc. (pg/mL)* Dilution Fold |
% Accuracy |
Conc. (pg/mL)* Dilution Fold | %Accuracy | |
| % Accuracy using 1 in 8 Dilution | 1 | 82.38 | 85.98 | 99.85 | 81.88 | 93.20 | 71.54 | 88.17 | 86.52 | 76.21 | 97.38 | ||||||||||
| 2 | 78.94 | 91.25 | 99.94 | 91.10 | 90.06 | 70.79 | 93.02 | 90.62 | 88.58 | 97.14 | |||||||||||
| 4 | 87.95 | 96.31 | 95.01 | 97.22 | 98.70 | 75.75 | 98.27 | 96.18 | 93.60 | 102.85 | |||||||||||
| 8 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | |||||||||||
| 16 | 84.58 | 89.55 | 84.21 | 91.40 | 90.87 | 82.17 | 89.50 | 90.90 | 96.44 | 80.59 | |||||||||||
| 32 | 70.09 | 84.69 | 55.81 | 84.61 | 64.48 | 58.88 | 86.37 | 71.72 | 82.83 | 40.03 | |||||||||||
| % Accuracy using 1 in 16 Dilution | 1 | 97.41 | 96.01 | 118.57 | 89.58 | 102.57 | 87.06 | 98.51 | 95.17 | 79.03 | 120.83 | ||||||||||
| 2 | 93.34 | 101.90 | 118.68 | 99.66 | 99.11 | 86.14 | 103.93 | 99.69 | 91.85 | 120.54 | |||||||||||
| 4 | 103.99 | 107.55 | 112.83 | 106.36 | 108.62 | 92.18 | 109.79 | 105.80 | 97.06 | 127.63 | |||||||||||
| 8 | 118.24 | 111.67 | 118.76 | 109.40 | 110.05 | 121.69 | 111.73 | 110.01 | 103.69 | 124.09 | |||||||||||
| 16 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | |||||||||||
| 32 | 82.87 | 94.58 | 66.28 | 92.57 | 70.96 | 71.66 | 96.50 | 78.90 | 85.89 | 49.67 | |||||||||||
| % Accuracy using 1 in 32 Dilution | 1 | 117.55 | 101.51 | 178.90 | 96.78 | 144.54 | 116.55 | 102.08 | 120.62 | 92.01 | 243.25 | ||||||||||
| 2 | 112.64 | 107.75 | 179.06 | 107.66 | 139.66 | 115.32 | 107.70 | 126.34 | 106.94 | 242.66 | |||||||||||
| 4 | 125.50 | 113.71 | 170.23 | 114.90 | 153.07 | 123.41 | 113.77 | 134.09 | 113.00 | 256.93 | |||||||||||
| 8 | 142.68 | 118.07 | 179.18 | 118.19 | 155.08 | 162.92 | 115.78 | 139.42 | 120.73 | 249.80 | |||||||||||
| 16 | 120.68 | 105.73 | 150.88 | 108.03 | 140.92 | 133.88 | 103.62 | 126.74 | 116.43 | 201.31 | |||||||||||
| 32 | 100.00 | 100.00 | 100.00 | 100.00 | 100.00 | 95.93 | 100.00 | 100.00 | 100.00 | 100.00 | |||||||||||
| Analyte Name: | VEGF | ||||||||||
| Dilution Fold | BM-20-1091 | BM-20-1106 | BM-20-1113 | BM-20-1120 | BM-20-1133 | BM-20-1137 | BM-20-1138 | BM-20-1140 | BM-20-1145 | BM-20-1150 | |
| Obtained concentration in pg/mL | |||||||||||
| Neat | 727.0 | 526.6 | 397.0 | 442.4 | 525.0 | 559.3 | 489.8 | 724.5 | 496.7 | 435.9 | |
| 1:2 | 348.3 | 279.4 | 198.6 | 246.0 | 253.6 | 276.6 | 258.4 | 379.4 | 288.6 | 217.4 | |
| 1:4 | 194.0 | 147.4 | 94.4 | 131.3 | 139.0 | 148.0 | 136.4 | 201.3 | 152.5 | 115.1 | |
| 1:8 | 110.3 | 76.5 | 49.7 | 67.5 | 70.4 | 97.7 | 69.4 | 104.6 | 81.4 | 55.9 | |
| 1:16 | 46.6 | 34.2 | 20.9 | 30.8 | 31.9 | 40.1 | 31.0 | 47.5 | 39.2 | 22.5 | |
| 1:32 | 19.3 | 16.2 | 6.9 | 14.2 | 11.3 | 14.3 | 14.9 | 18.7 | 16.8 | 5.6 | |
| Stability Condition & Temperature | QC Level | % Difference (n=3) | Duration | ||
|---|---|---|---|---|---|
| FT Stability II cycle & -75 °C | PM-01 | -0.19 | 7.78 | 12.72 | Up to six FT cycles |
| PM-02 | -4.70 | -9.54 | -4.05 | ||
| FT Stability IV cycle & -75 °C | PM-01 | 11.50 | 9.07 | 9.36 | |
| PM-02 | 0.02 | -2.24 | -5.00 | ||
| FT Stability VI cycle & -75 °C | PM-01 | 7.22 | -0.09 | 6.55 | |
| PM-02 | -3.90 | -0.69 | -3.60 | ||
| FT Stability II cycle & -20 °C | PM-01 | -0.39 | 6.97 | 13.10 | Up to six FT cycles |
| PM-02 | 8.38 | 6.25 | 2.00 | ||
| FT Stability IV cycle & -20 °C | PM-01 | 9.34 | 6.02 | 0.78 | |
| PM-02 | -1.09 | 1.45 | -0.96 | ||
| FT Stability VI cycle & -20 °C | PM-01 | 4.80 | 1.97 | 1.60 | |
| PM-02 | -4.33 | 2.23 | 5.75 | ||
| Bench Top Stability at Room Temperature |
PM-01 | 6.57 | 9.98 | 6.65 | 27 Hours |
| PM-02 | 8.93 | -3.17 | -4.89 | ||
| Refrigerator Stability at 2-8 °C |
PM-01 | 9.00 | 12.33 | 9.59 | 74 hours |
| PM-02 | -9.98 | -11.57 | -10.53 | ||
| Subject ID | Day 1- Pre Dose | Day 1- Post Dose | Day 3 | Day 7 | Day 15 | Day 30 | Day 60 | Day 60- Post Dose | Day 67 | Day 90 |
|---|---|---|---|---|---|---|---|---|---|---|
| 11-202 | Missing | Missing | 71.5 | BLQ | BLQ | 70.6 | BLQ | 72.5 | BLQ | 59.5 |
| 11-203 | BLQ | Missing | BLQ | BLQ | BLQ | BLQ | 70.2 | BLQ | BLQ | 84.4 |
| 11-204 | 127.3 | 235.2 | 67.1 | 174.3 | 158.1 | 137.1 | 103.3 | 156.7 | Missing | 123.8 |
| 11-205 | 195.5 | 286.8 | 89.5 | BLQ | BLQ | BLQ | 90.2 | Missing | Missing | Missing |
| 11-206 | 62.7 | 73.6 | BLQ | BLQ | BLQ | BLQ | 69.9 | BLQ | BLQ | BLQ |
| 11-210 | 74.8 | 50.4 | 50.4 | BLQ | BLQ | 68.4 | BLQ | BLQ | BLQ | Missing |
| 21-001 | BLQ | BLQ | BLQ | BLQ | 54.3 | BLQ | Missing | |||
| 21-002 | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | |||
| 21-003 | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | |||
| 21-004 | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | |||
| 21-005 | BLQ | 54.8 | BLQ | BLQ | 83.0 | BLQ | BLQ | |||
| 21-006 | 65.9 | BLQ | BLQ | BLQ | BLQ | Missing | BLQ | |||
| 21-007 | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | |||
| 21-008 | BLQ | BLQ | BLQ | BLQ | BLQ | 230.6 | BLQ | |||
| 21-009 | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | |||
| 21-010 | 121.7 | 59.6 | 126.0 | BLQ | BLQ | BLQ | BLQ | |||
| 21-011 | 80.3 | BLQ | BLQ | BLQ | BLQ | 75.6 | BLQ | |||
| 21-012 | 133.2 | 101.5 | 82.9 | 138.3 | 90.2 | BLQ | 116.9 | |||
| 21-013 | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | |||
| 21-014 | BLQ | BLQ | BLQ | BLQ | BLQ | BLQ | Missing |
| PD assays for VEGF in plasma | |||
|---|---|---|---|
| Parameter | ELISA | MSD | |
| Biological matrix | : | Normal Human plasma | Normal Human plasma |
| Plasma Anticoagulant | : | Sodium Citrate | Sodium Citrate |
| Standard | : | Recombinant Human VEGF (CR08) | Recombinant Human VEGF (CR08) |
| Calibration curve range | : | 62.5 pg/mL –2000.0 pg/mL | 7.7 - 562 pg/mL |
| Anchor points | : | Upper – 2600.0 pg/mL, Lower-31.25 pg/mL, MRD: 4 | 1.12- 1080 pg/mL, MRD: 2 |
| Analytical technique | : | Direct Sandwich ELISA- Dual Colorimetric Detection | Sandwich ECL Detection |
| Regression | : | Non-Linear (4PL) | Non-Linear (4PL) |
| Weighing factor | : | 1/Y² | 1/Y² |
| Sample Volume | : | 100 µL | 50 µL |
| Time required | : | 05 Hours | 05 Hours |
| Wash steps | : | 2 | 3 |
| Kit / Vendor | : | Human VEGF Quantikine ELISA kit, R&D systems |
V-PLEX Plus Human VEGF Kit, MSD |
| Pre-coated plate | : | mAb specific for Sf21-expressed recombinant human VEGF-A165 | Anti-hu VEGF Antibody |
| Detection Antibody | : | peroxidase-conjugated Anti-human VEGF Conjugate polyclonal Ab | SULFO-TAG™ Anti-hVEGF Antibody |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




