Submitted:
04 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
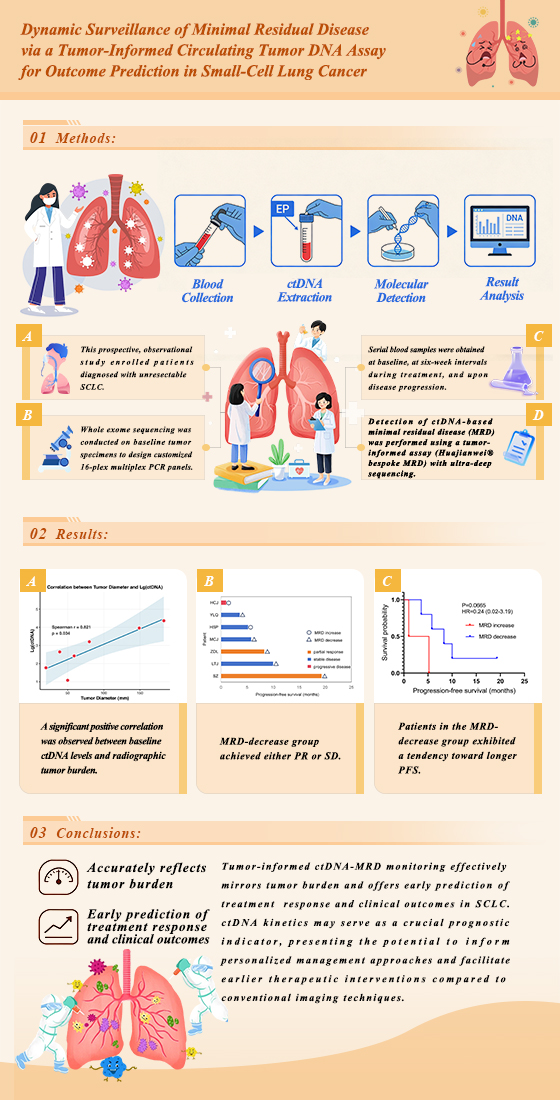
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design and Patients
2.2. Sample Collection
2.3. Efficacy Evaluation
2.4. Extraction of Cell-Free DNA (cfDNA)
2.5. WES at Baseline
2.6. Personalized Tumor-Informed ctDNA Detection
2.7. Statistic Analysis
3. Results
3.1. Patient Characteristics
3.2. Clinical Relevance of Baseline WES and ctDNA Analysis
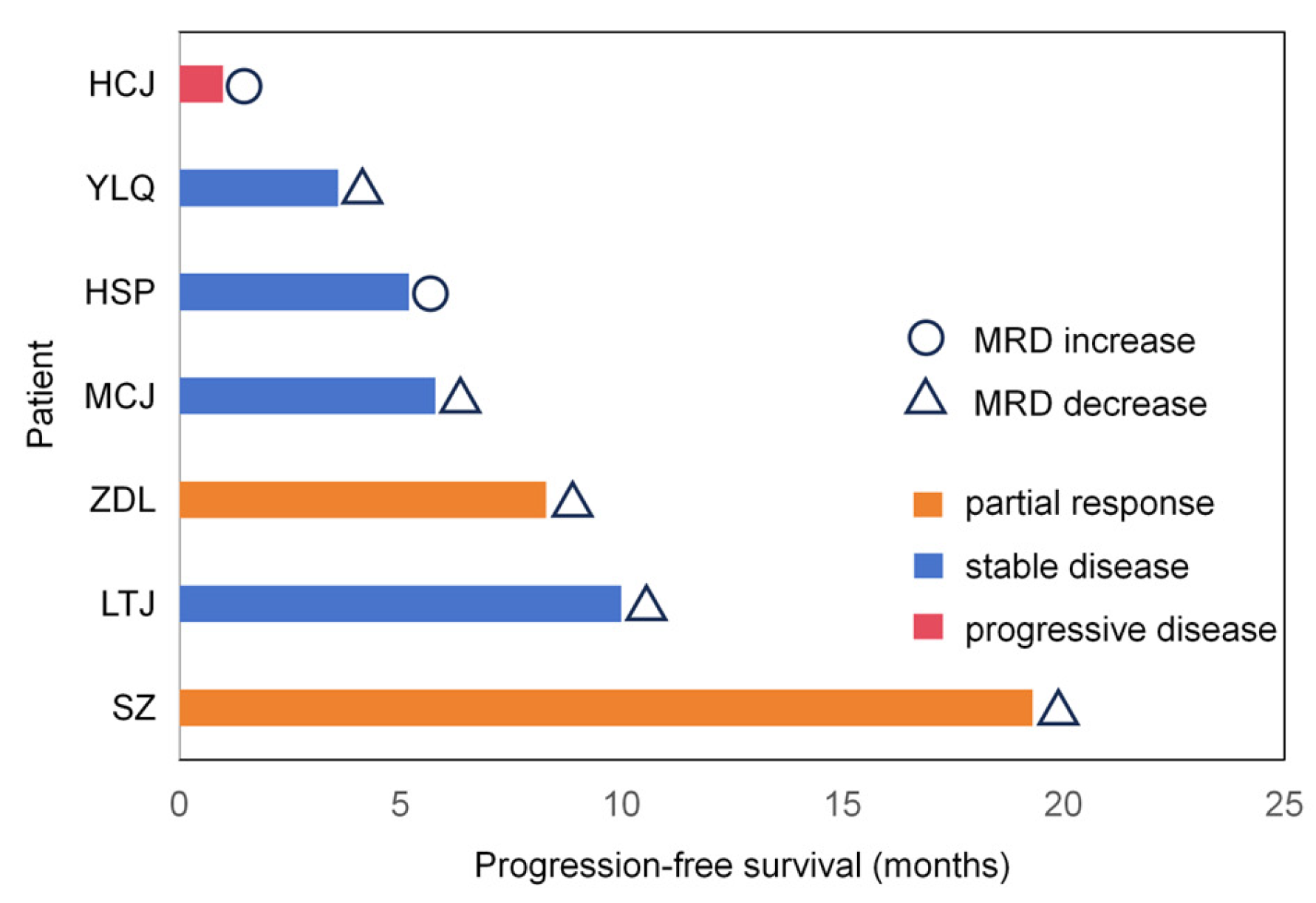
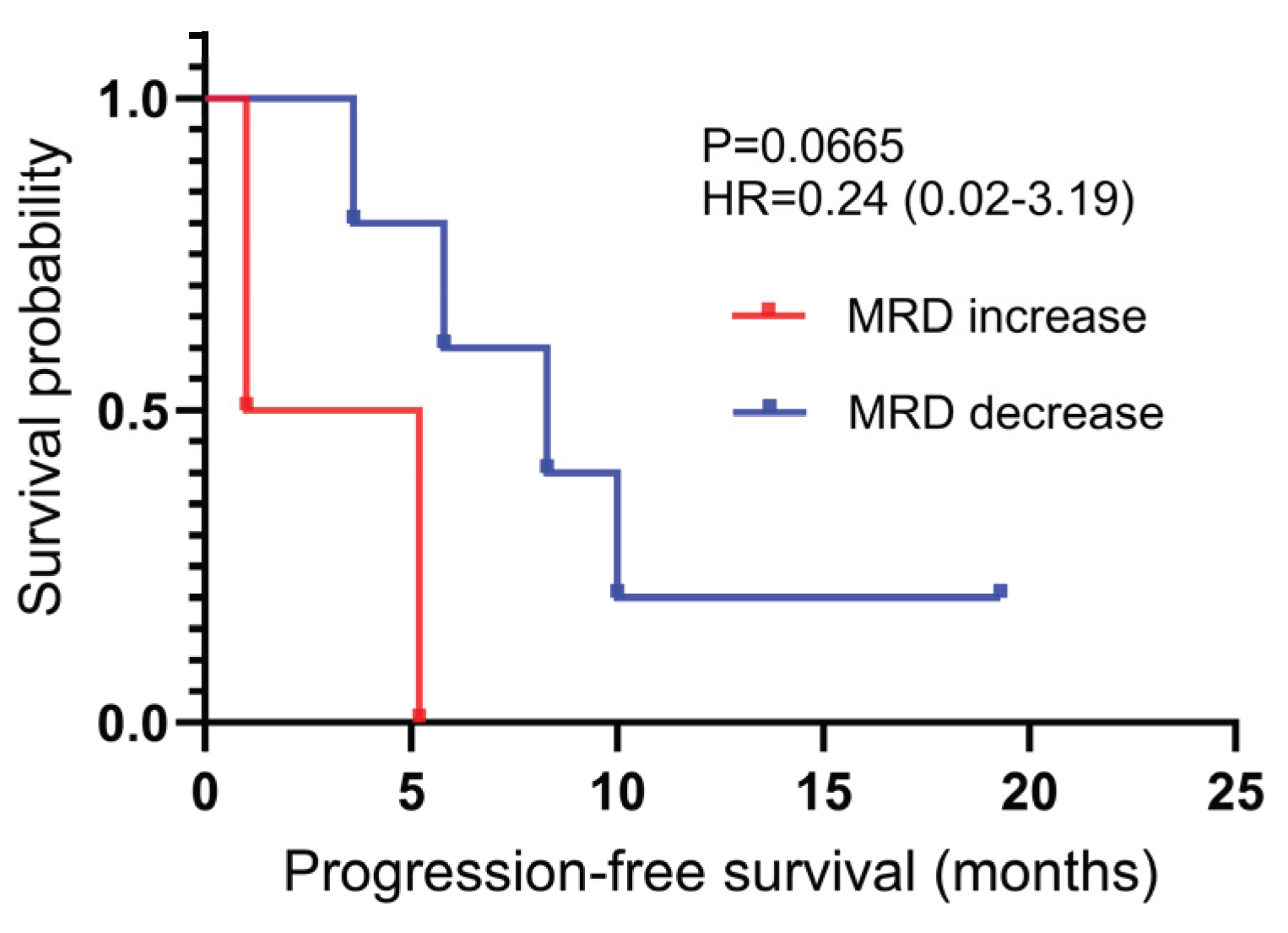
3.3. Longitudinal Assessment of ctDNA-MRD During Treatment
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| SCLC | Small-cell lung cancer |
| ctDNA | circulating tumor DNA |
| PFS | progression-free survival |
| OS | overall survival |
| mTBI | molecular tumor burden index |
| LS-SCLC | limited-stage small-cell lung cancer |
| MRD | minimal residual disease |
| NGS | next-generation sequencing |
| WES | whole exome sequencing |
| RECIST | Response Evaluation Criteria in Solid Tumors |
| FFPE | formalin-fixed paraffin-embedded |
| CT | computed tomography |
| MRI | magnetic resonance imaging |
| cfDNA | cell-free DNA |
| H&E | hematoxylin and eosin |
| gDNA | genomic DNA |
| SNVs | single nucleotide variants |
| indels | insertions/deletions |
| CNVs | copy number variations |
| TMB | tumor mutational burden |
| MSI | microsatellite instability |
| MTM/mL | mean tumor molecules per milliliter |
| VAF | variant allele frequency |
| PD-(L)1 | programmed cell death (ligand) -1 |
| PR | partial response |
| SD | stable disease |
| PD | progressive disease |
| TPS | Tumor Proportion Scores |
| HR | hazard ratio |
| NSCLC | non-small-cell lung cancer |
References
- Sung, H.; Ferlay, J.; Siegel, R. L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA-Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Wood, D. E.; Kazerooni, E. A.; Aberle, D.; Berman, A.; Brown, L. M.; Eapen, G. A.; Ettinger, D. S.; Ferguson, J. S.; Hou, L.; Kadaria, D.; et al. NCCN Guidelines® Insights: Lung Cancer Screening, Version 1.2022. J. Natl. Compr. Cancer Netw. 2022, 20, 754–764. [Google Scholar] [CrossRef]
- Ganti, A. K. P.; Loo, B. W.; Bassetti, M.; Blakely, C.; Chiang, A.; D’Amico, T. A.; D’Avella, C.; Dowlati, A.; Downey, R. J.; Edelman, M. J.; et al. Small Cell Lung Cancer, Version 2.2022, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 2021, 19, 1441–1464. [Google Scholar] [CrossRef]
- Zugazagoitia, J.; Paz-Ares, L. Extensive-Stage Small-Cell Lung Cancer: First-Line and Second-Line Treatment Options. J. Clin. Oncol. 2022, 40, 671–680. [Google Scholar] [CrossRef]
- Abbosh, C.; Birkbak, N. J.; Wilson, G. A.; Jamal-Hanjani, M.; Constantin, T.; Salari, R.; Le Quesne, J.; Moore, D. A.; Veeriah, S.; Rosenthal, R.; et al. Phylogenetic ctDNA Analysis Depicts Early-Stage Lung Cancer Evolution. Nature 2017, 545, 446–451. [Google Scholar] [CrossRef]
- Chaudhuri, A. A.; Chabon, J. J.; Lovejoy, A. F.; Newman, A. M.; Stehr, H.; Azad, T. D.; Khodadoust, M. S.; Esfahani, M. S.; Liu, C. L.; Zhou, L.; et al. Early Detection of Molecular Residual Disease in Localized Lung Cancer by Circulating Tumor DNA Profiling. Cancer Discov. 2017, 7, 1394–1403. [Google Scholar] [CrossRef]
- Newman, A. M.; Bratman, S. V.; To, J.; Wynne, J. F.; Eclov, N. C. W.; Modlin, L. A.; Liu, C. L.; Neal, J. W.; Wakelee, H. A.; Merritt, R. E.; et al. An Ultrasensitive Method for Quantitating Circulating Tumor DNA with Broad Patient Coverage. Nat. Med. 2014, 20, 548–554. [Google Scholar] [CrossRef] [PubMed]
- Heeke, S.; Gay, C. M.; Estecio, M. R.; Tran, H.; Morris, B. B.; Zhang, B.; Tang, X.; Raso, M. G.; Rocha, P.; Lai, S.; et al. Tumor- and Circulating-Free DNA Methylation Identifies Clinically Relevant Small Cell Lung Cancer Subtypes. Cancer Cell 2024, 42, 225–237.e5. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Liu, Y.; Yuan, M.; Dong, G.; Zhang, H.; Zhang, T.; Chang, L.; Xia, X.; Li, L.; Zhu, H.; et al. The Feasibility of Using Biomarkers Derived from Circulating Tumor DNA Sequencing as Predictive Classifiers in Patients with Small-Cell Lung Cancer. Cancer Res. Treat. 2022, 54, 753–766. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Wu, Y.; Zhao, J.; Zhang, T.; Fei, K.; Zhao, X.; Deng, L.; Zhang, Z.; Jiang, Y.; Wang, J.; et al. Circulating Tumor DNA Refines Consolidation Immunotherapy for Limited-Stage Small Cell Lung Cancer Patients. Signal Transduct. Target. Ther. 2025, 10, 347. [Google Scholar] [CrossRef]
- Sivapalan, L.; Iams, W. T.; Belcaid, Z.; Scott, S. C.; Niknafs, N.; Balan, A.; White, J. R.; Kopparapu, P.; Cann, C.; Landon, B. V.; et al. Dynamics of Sequence and Structural Cell-Free DNA Landscapes in Small-Cell Lung Cancer. Clin. Cancer Res. 2023, 29, 2310–2323. [Google Scholar] [CrossRef]
- Herbreteau, G.; Langlais, A.; Greillier, L.; Audigier-Valette, C.; Uwer, L.; Hureaux, J.; Moro-Sibilot, D.; Guisier, F.; Carmier, D.; Madelaine, J.; et al. Circulating Tumor DNA as a Prognostic Determinant in Small Cell Lung Cancer Patients Receiving Atezolizumab. J. Clin. Med. 2020, 9, 3861. [Google Scholar] [CrossRef]
- Li, J.; Wang, L.; Dong, Z.; Song, Q.; Wang, Z. A Meta-Analysis of Circulating Tumor DNA as a Survival Indicator in Small Cell Lung Cancer Patients. Clin. Exp. Med. 2023, 23, 3935–3945. [Google Scholar] [CrossRef]
- Pan, Y.; Zhang, J. T.; Gao, X.; Chen, Z. Y.; Yan, B.; Tan, P. X.; Yang, X. R.; Gao, W.; Gong, Y.; Tian, Z.; et al. Dynamic Circulating Tumor DNA during Chemoradiotherapy Predicts Clinical Outcomes for Locally Advanced Non-Small Cell Lung Cancer Patients. Cancer Cell 2023, 41, 1763–1773.e4. [Google Scholar] [CrossRef]
- Dong, S.; Wang, Z.; Zhang, J. T.; Yan, B.; Zhang, C.; Gao, X.; Sun, H.; Li, Y. S.; Yan, H. H.; Tu, H. Y.; et al. Circulating Tumor DNA-Guided De-Escalation Targeted Therapy for Advanced Non-Small Cell Lung Cancer: A Nonrandomized Controlled Trial. JAMA Oncol. 2024, 10, 932–940. [Google Scholar] [CrossRef] [PubMed]
- Dong, Q.; Chen, C.; Hu, Y.; Zhang, W.; Yang, X.; Qi, Y.; Zhu, C.; Chen, X.; Shen, X.; Ji, W.; et al. Clinical Application of Molecular Residual Disease Detection by Circulation Tumor DNA in Solid Cancers and a Comparison of Technologies: Review Article. Cancer Biol. Ther. 2023, 24, 2274123. [Google Scholar] [CrossRef]
- Martínez-Castedo, B.; Camblor, D. G.; Martín-Arana, J.; Carbonell-Asins, J. A.; García-Micó, B.; Gambardella, V.; Huerta, M.; Roselló, S.; Roda, D.; Gimeno-Valiente, F.; et al. Minimal Residual Disease in Colorectal Cancer. Tumor-Informed versus Tumor-Agnostic Approaches: Unraveling the Optimal Strategy. Ann. Oncol. 2025, 36, 263–276. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Shields, M. D.; Chauhan, P. S.; Ramirez, R. J.; Harris, P. K.; Reimers, M. A.; Zevallos, J. P.; Davis, A. A.; Pellini, B.; Chaudhuri, A. A. Commercial ctDNA Assays for Minimal Residual Disease Detection of Solid Tumors. Mol. Diagn. Ther. 2021, 25, 757–774. [Google Scholar] [CrossRef]
- Jiang, S; Li, H; Zhang, L; Mu, W; Zhang, Y; Chen, T; Wu, J; Tang, H; Zheng, S; Liu, Y; Wu, Y; Luo, X; Xie, Y; Ren, J. Generic Diagramming Platform (GDP): a comprehensive database of high-quality biomedical graphics. Nucleic Acids Res. 2025, 53, D1670–D1676. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Abbosh, C.; Birkbak, N. J.; Wilson, G. A.; Jamal-Hanjani, M.; Constantin, T.; Salari, R.; Le Quesne, J.; Moore, D. A.; Veeriah, S.; Rosenthal, R.; et al. Phylogenetic ctDNA Analysis Depicts Early-Stage Lung Cancer Evolution. Nature 2017, 545, 446–451. [Google Scholar] [CrossRef] [PubMed]
- Reinert, T.; Henriksen, T. V.; Christensen, E.; Sharma, S.; Salari, R.; Sethi, H.; Knudsen, M.; Nordentoft, I.; Wu, H. T.; Tin, A. S.; et al. Analysis of Plasma Cell-Free DNA by Ultradeep Sequencing in Patients with Stages I to III Colorectal Cancer. JAMA Oncol. 2019, 5, 1124–1131. [Google Scholar] [CrossRef]
- Moding, E. J.; Liu, Y.; Nabet, B. Y.; Chabon, J. J.; Chaudhuri, A. A.; Hui, A. B.; Bonilla, R. F.; Ko, R. B.; Yoo, C. H.; Gojenola, L.; et al. Circulating Tumor DNA Dynamics Predict Benefit from Consolidation Immunotherapy in Locally Advanced Non-Small Cell Lung Cancer. Nat. Cancer 2020, 1, 176–183. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Cuesta, L.; Perdomo, S.; Avogbe, P. H.; Leblay, N.; Delhomme, T. M.; Gaborieau, V.; Abedi-Ardekani, B.; Chanudet, E.; Olivier, M.; Zaridze, D.; et al. Identification of Circulating Tumor DNA for the Early Detection of Small-Cell Lung Cancer. EBioMedicine 2016, 10, 117–123. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Narayan, A.; Xu, Y.; Wolfe, J.; Vu, D.; Trinh, T.; Kantak, C.; Ivy, S. P.; Eder, J. P.; Deng, Y.; et al. Circulating Tumor DNA Dynamics Fail to Predict Efficacy of Poly(ADP-ribose) Polymerase/VEGFR Inhibition in Patients with Heavily Pretreated Advanced Solid Tumors. JCO Precis. Oncol. 2024, 8, e2300289. [Google Scholar] [CrossRef] [PubMed]
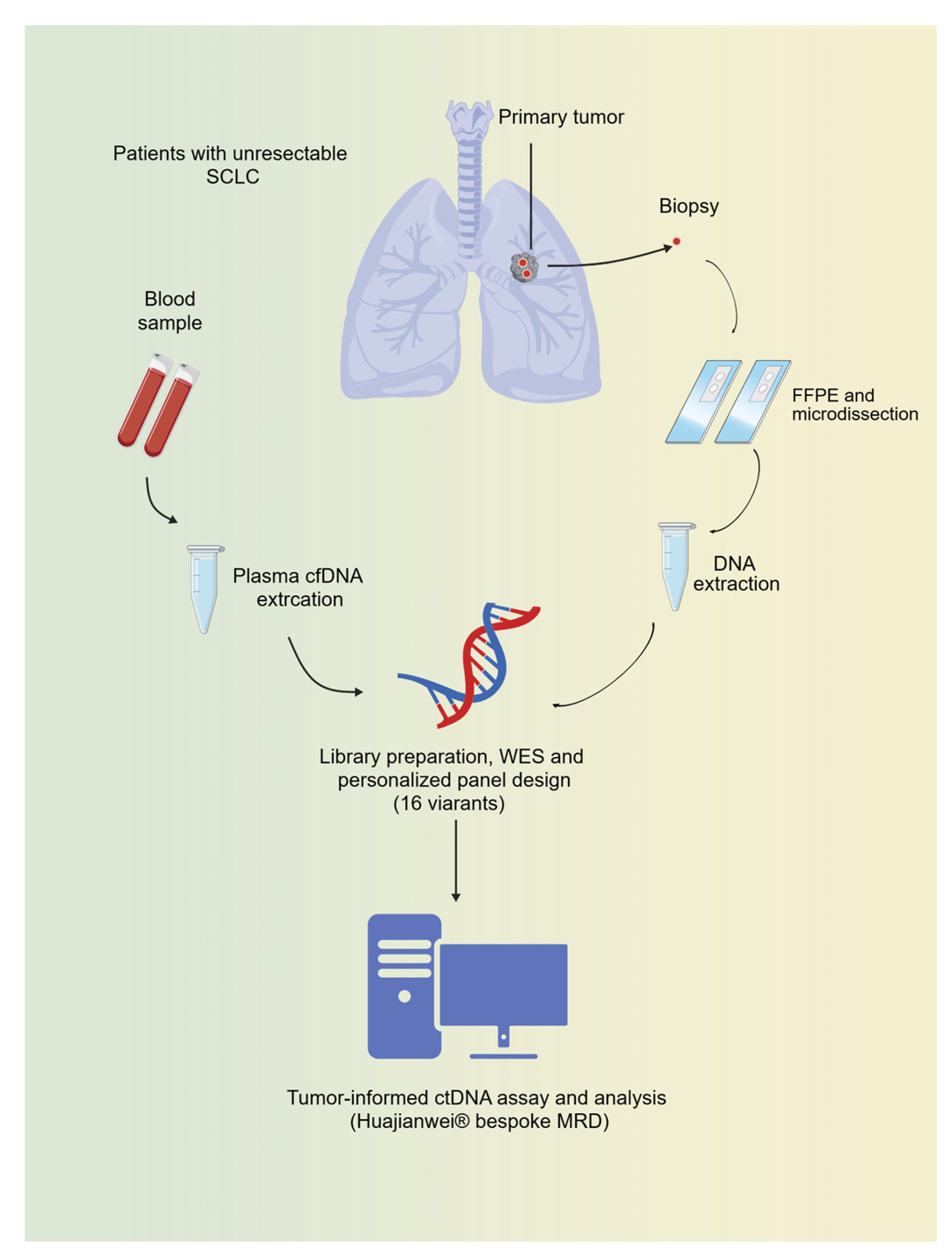
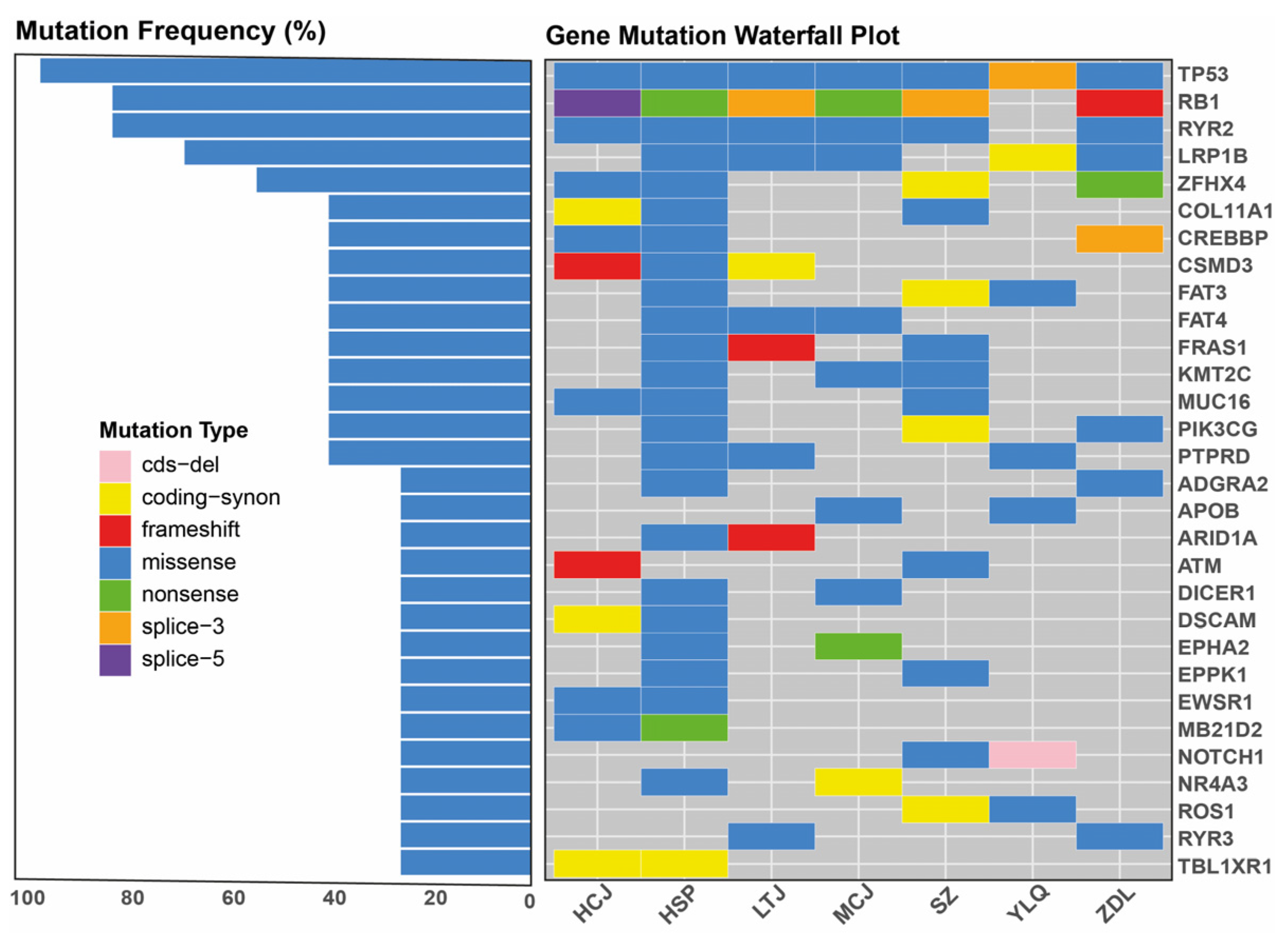
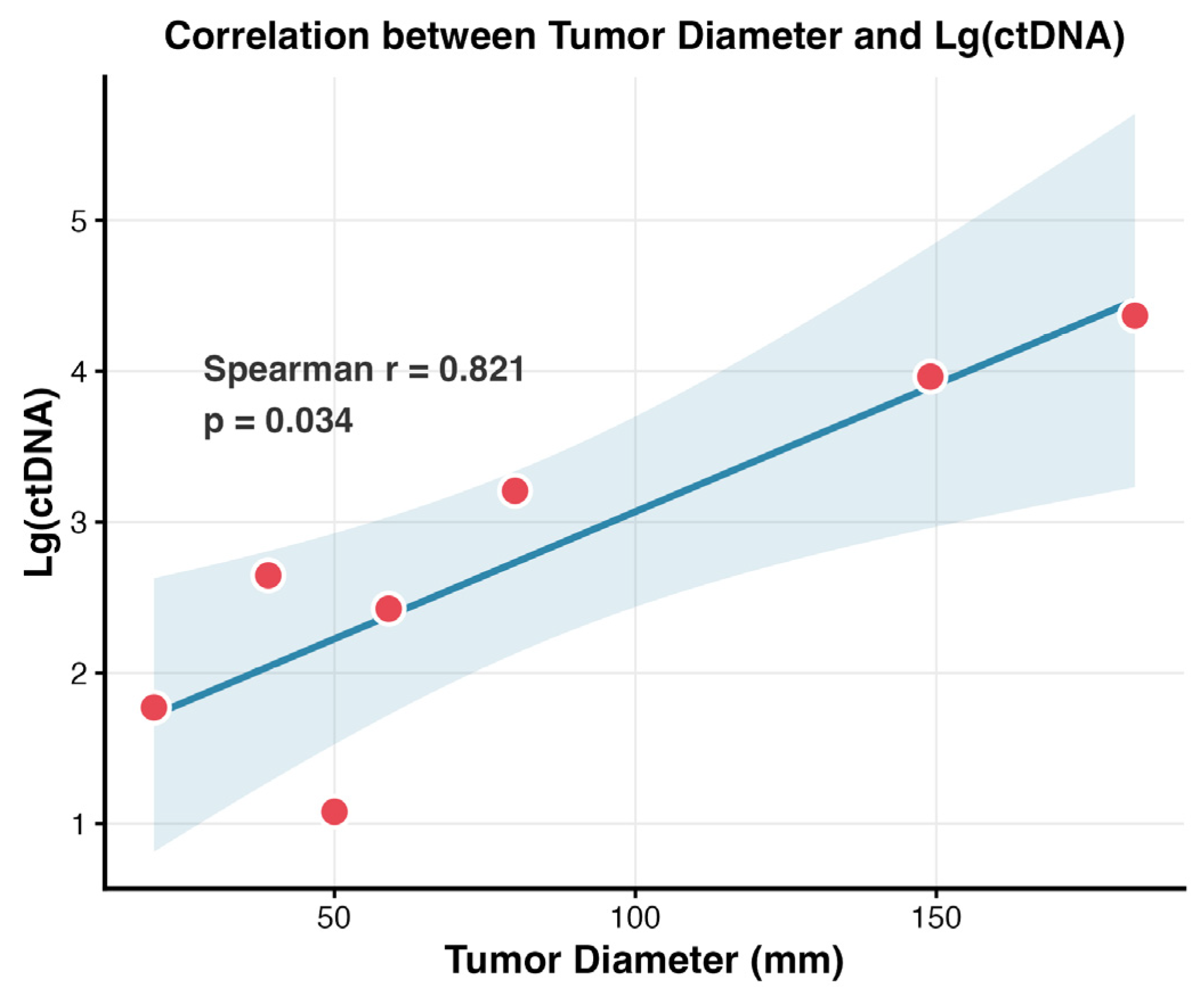
| Patient | Age | Sex | Smoking, pack-years | Pathological diagnosis | Tumor stage | Treatment | Lines | PFS (months) | Tumor response | TMB (Muts/Mb) |
PD-L1 (TPS) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| LTJ | 58 | Male | 80 | SCLC | LS | platinum-etoposide with thoracic radiotherapy | 1st | 10.0 | SD | 7.97 | NA |
| MCJ | 60 | Male | 40 | SCLC | ES | platinum-etoposide with PD-L1 inhibitor | 1st | 5.8 | SD | 5.19 | 0% |
| ZDL | 59 | Male | 40 | SCLC | ES | platinum-etoposide with thoracic radiotherapy | 2nd | 8.3 | PR | 5.01 | NA |
| HSP | 60 | Male | 122 | SCLC | ES | platinum-etoposide with PD-1 inhibitor | 3rd | 5.2 | SD | 61.8 | NA |
| YLQ | 61 | Male | 60 | SCLC | ES | anlotinib | 5th | 3.6 | PD | 4.81 | <1% |
| HCJ | 60 | Male | 15 | C-SCLC | ES | platinum-etoposide | 3rd | 1.0 | PD | 8.25 | <1% |
| SZ | 74 | Male | 30 | SCLC | LS | platinum-etoposide with thoracic radiotherapy | 1st | 19.3 | PR | 11.17 | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




