Submitted:
04 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design and Population
- inclusion criteria: availability of baseline staging MRI, completion of neoadjuvant therapy, availability of restaging MRI, surgical resection.
- exclusion criteria: poor image quality (e.g. motion artifacts), missing clinical data.
2.2. MRI Acquisition Protocol
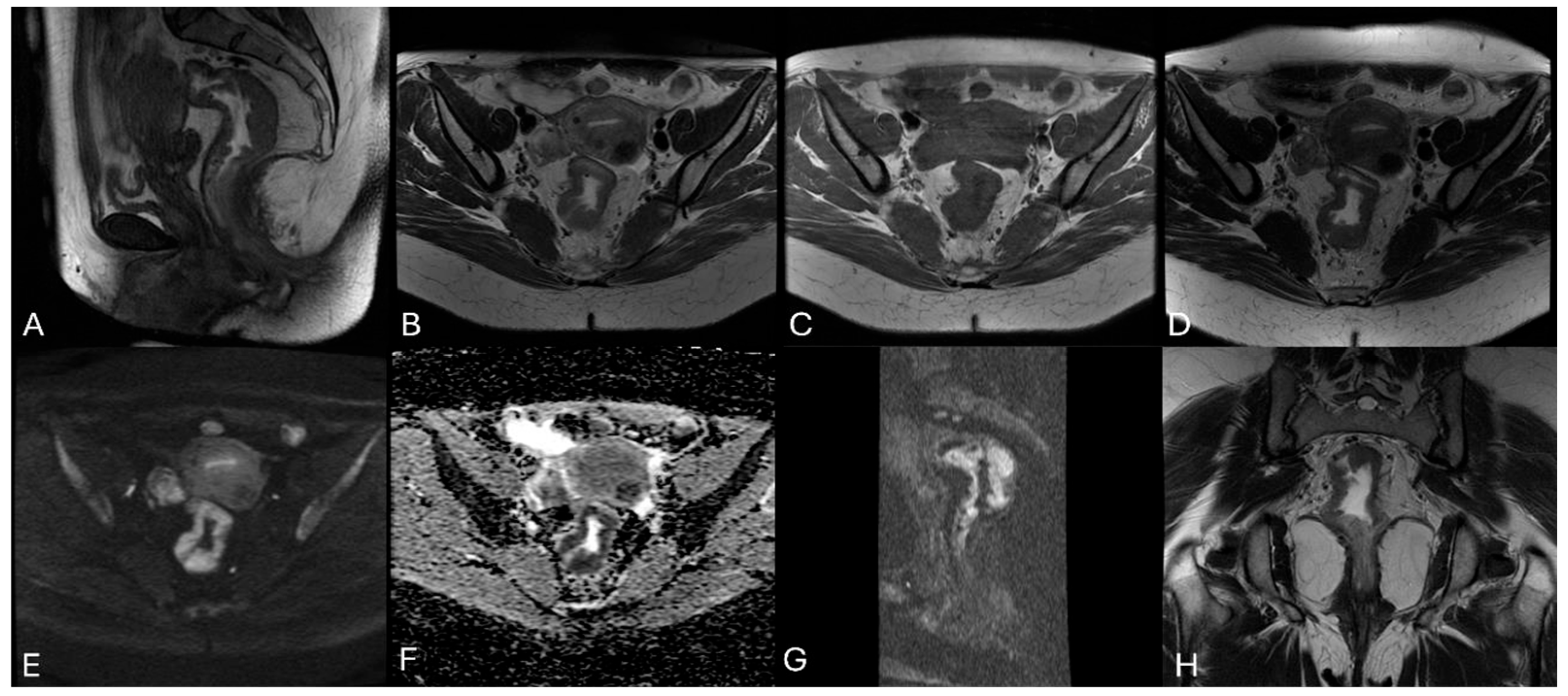
2.3. MRI-Based Tumor Regression Grading System (mrTRG)
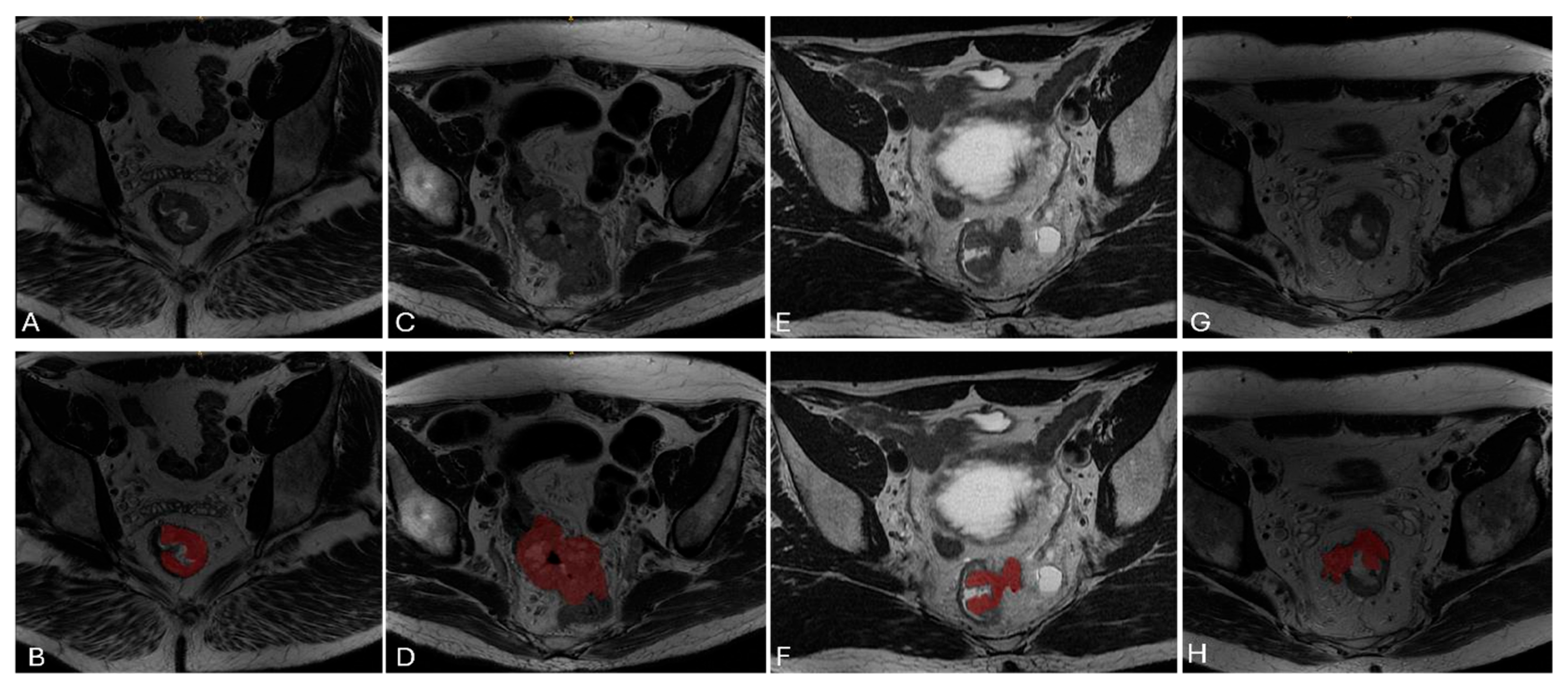
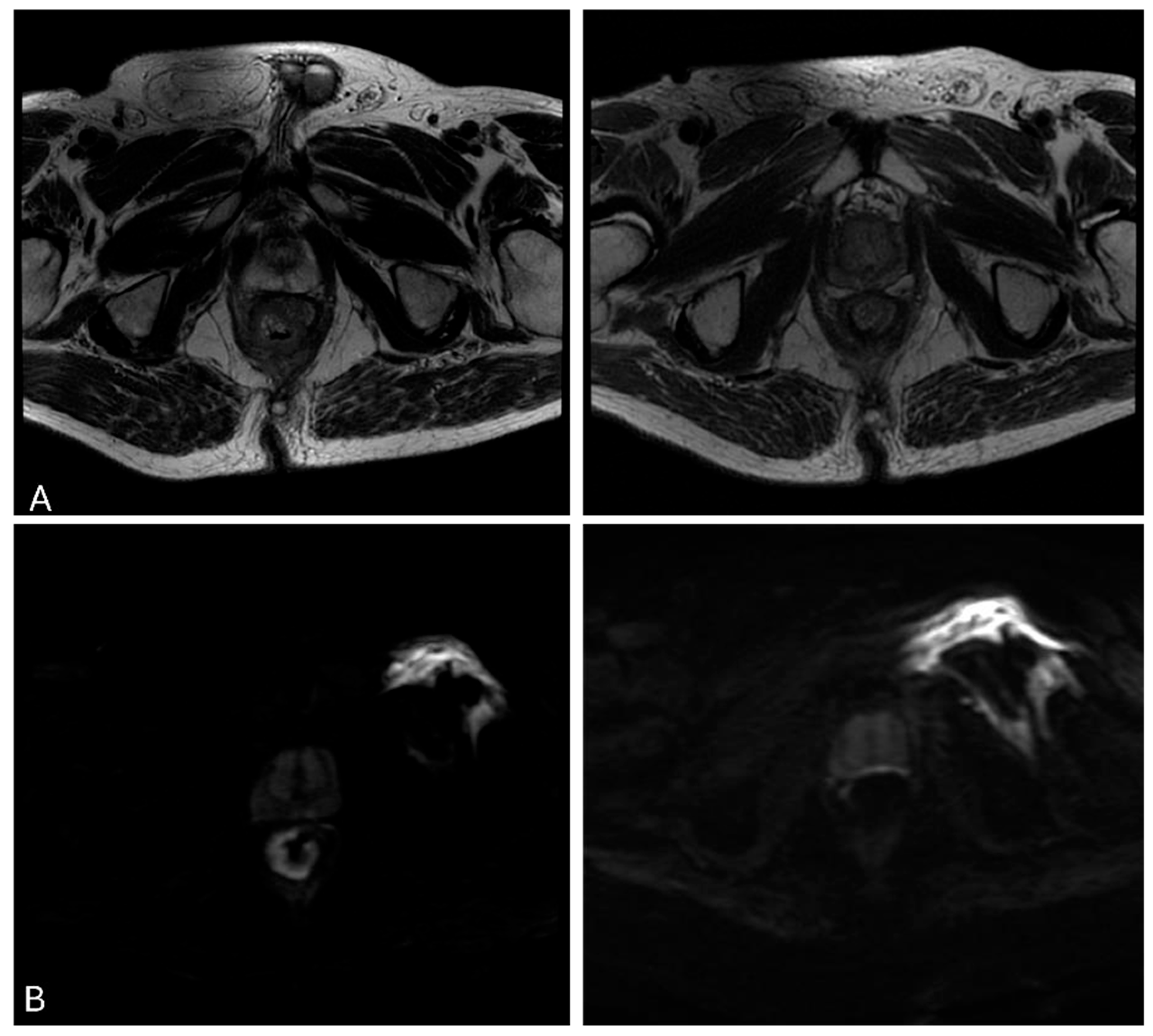
2.4. Tumor Segmentation and Pre-Processing
2.5. Radiomic Feature Extraction
2.6. Model Development and Validation
2.7. Statistical Analysis
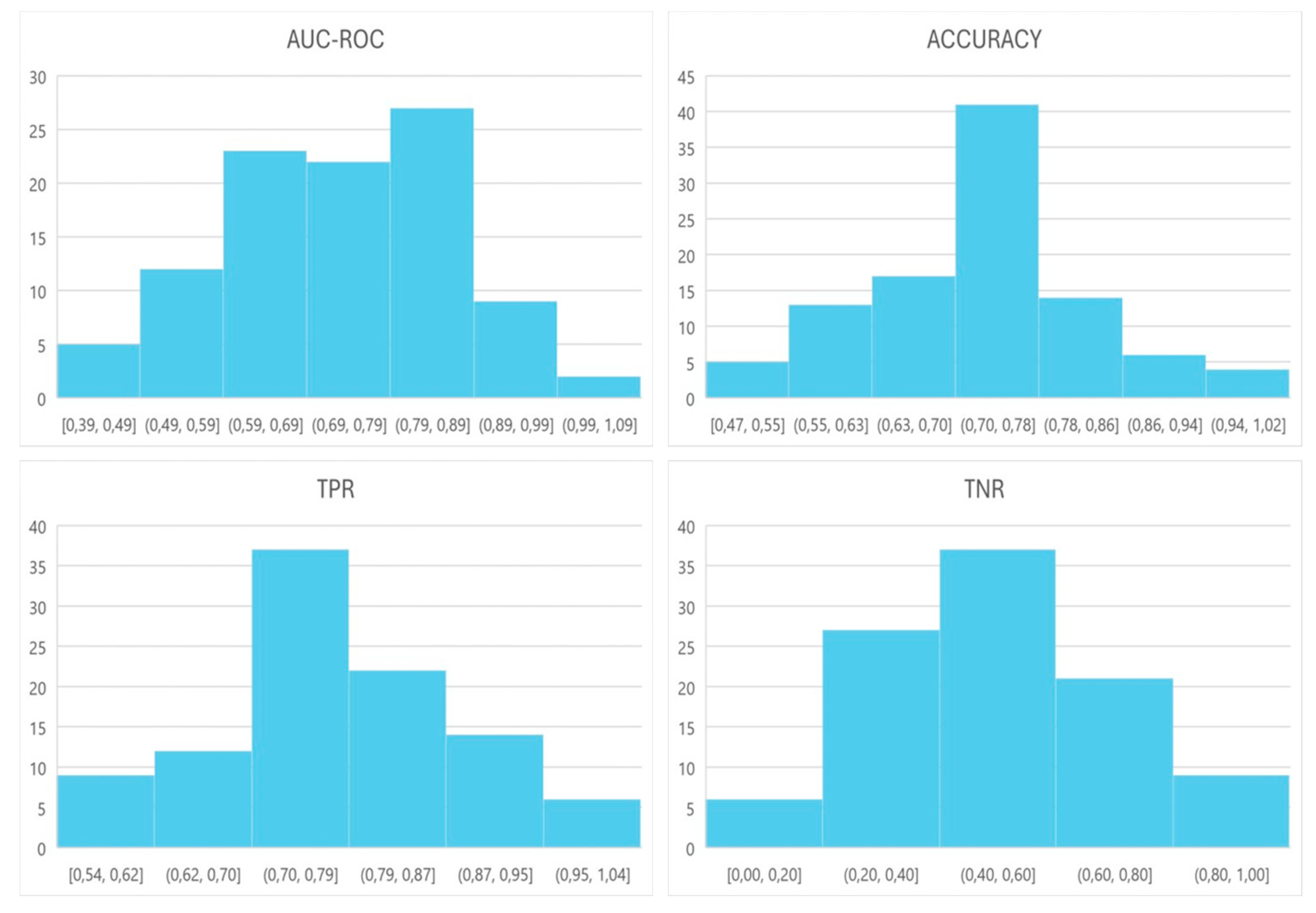
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ADC | Apparent Diffusion Coefficient |
| AUC-ROC | Area Under the Receiver Operating Characteristic Curve |
| CRC | Colorectal Cancer |
| CR | Complete Response |
| CT | Chemotherapy |
| CTRT | Chemoradiotherapy |
| CT IND>CTRT | Induction Chemotherapy followed by Chemoradiotherapy |
| DWI | Diffusion-Weighted Imaging |
| EMVI | Extramural Vascular Invasion |
| FOV | Field of View |
| FSE | Fast Spin Echo |
| FRFSE | Fast Recovery Fast Spin Echo |
| GLCM | Gray Level Co-occurrence Matrix |
| GLDM | Gray Level Dependence Matrix |
| GLRLM | Gray Level Run Length Matrix |
| GLSZM | Gray Level Size Zone Matrix |
| GRE | Gradient Echo |
| LARC | Locally Advanced Rectal Cancer |
| LASSO | Least Absolute Shrinkage and Selection Operator |
| MRI | Magnetic Resonance Imaging |
| mrTRG | Magnetic Resonance Imaging Tumor Regression Grade |
| NGTDM | Neighbouring Gray Tone Difference Matrix |
| NPV | Negative Predictive Value |
| pCR | Pathological Complete Response |
| PPV | Positive Predictive Value |
| ROI | Region of Interest |
| RT | Radiotherapy |
| RTSC | Short-Course Radiotherapy |
| TME | Total Mesorectal Excision |
| TNT | Total Neoadjuvant Therapy |
| TNM | Tumor–Node–Metastasis Staging System |
| T1w | T1-weighted |
| T2w | T2-weighted |
| TPR | True Positive Rate |
| TNR | True Negative Rate |
| TRG | Tumor Regression Grade |
References
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin 2018;68:7–30. [CrossRef]
- Fazeli MS, Keramati MR. Rectal cancer: a review. Med J Islam Repub Iran 2015;29:171.
- Calabria R. Linee guida Aiom NEOPLASIE DEL RETTO E ANO - Edizione 2017 n.d.
- Siegel R, DeSantis C, Jemal A. Colorectal cancer statistics, 2014. CA Cancer J Clin 2014;64:104–17. [CrossRef]
- Stefania G, Giuseppe A, Paolo A, Sergio B, Vanna Chiarion S, Alessandro C, et al. I numeri del cancro in Italia. 2017. http://www.registri-tumori.it/PDF/AIOM2014/I_numeri_del_cancro_2014.pdf.
- Garajová I, Di Girolamo S, De Rosa F, Corbelli J, Agostini V, Biasco G, et al. Neoadjuvant Treatment in Rectal Cancer: Actual Status. Chemother Res Pract 2011;2011:839742. [CrossRef]
- Jeong SY, Park JW, Nam BH, Kim S, Kang SB, Lim SB, et al. Open versus laparoscopic surgery for mid-rectal or low-rectal cancer after neoadjuvant chemoradiotherapy (COREAN trial): survival outcomes of an open-label, non-inferiority, randomised controlled trial. Lancet Oncol 2014;15:767–74. [CrossRef]
- Lindsetmo RO, Joh YG, Delaney CP. Surgical treatment for rectal cancer: an international perspective on what the medical gastroenterologist needs to know. World J Gastroenterol 2008;14:3281–9. [CrossRef]
- Benson AB, Venook AP, Al-Hawary MM, Azad N, Chen YJ, Ciombor KK, et al. Rectal Cancer, Version 2.2022, NCCN Clinical Practice Guidelines in Oncology. Journal of the National Comprehensive Cancer Network 2022;20:1139–67. [CrossRef]
- Kitz J, Fokas E, Beissbarth T, Ströbel P, Wittekind C, Hartmann A, et al. Association of Plane of Total Mesorectal Excision With Prognosis of Rectal Cancer: Secondary Analysis of the CAO/ARO/AIO-04 Phase 3 Randomized Clinical Trial. JAMA Surg 2018;153:e181607. [CrossRef]
- Peeters KCMJ, van de Velde CJH, Leer JWH, Martijn H, Junggeburt JMC, Kranenbarg EK, et al. Late Side Effects of Short-Course Preoperative Radiotherapy Combined With Total Mesorectal Excision for Rectal Cancer: Increased Bowel Dysfunction in Irradiated Patients—A Dutch Colorectal Cancer Group Study. Journal of Clinical Oncology 2005;23:6199–206. [CrossRef]
- Sauer R, Becker H, Hohenberger W, Rödel C, Wittekind C, Fietkau R, et al. Preoperative versus Postoperative Chemoradiotherapy for Rectal Cancer. New England Journal of Medicine 2004;351:1731–40. [CrossRef]
- Bujko K, Nowacki MP, Nasierowska-Guttmejer A, Michalski W, Bebenek M, Pudelko M, et al. Sphincter preservation following preoperative radiotherapy for rectal cancer: Report of a randomised trial comparing short-term radiotherapy vs. conventionally fractionated radiochemotherapy. Radiotherapy and Oncology 2004;72:15–24. [CrossRef]
- Glehen O, Chapet O, Adham M, Nemoz JC, Gerard JP. Long-term results of the Lyons R90-01 randomized trial of preoperative radiotherapy with delayed surgery and its effect on sphincter-saving surgery in rectal cancer. Br J Surg 2003;90:996–8. [CrossRef]
- Beets-Tan RGH, Lambregts DMJ, Maas M, Bipat S, Barbaro B, Curvo-Semedo L, et al. Magnetic resonance imaging for clinical management of rectal cancer: Updated recommendations from the 2016 European Society of Gastrointestinal and Abdominal Radiology (ESGAR) consensus meeting. Eur Radiol 2018;28:1465–75. [CrossRef]
- Beets-Tan RGH, Beets GL, Vliegen RFA, Kessels AGH, Van Boven H, De Bruine A, et al. Accuracy of magnetic resonance imaging in prediction of tumour-free resection margin in rectal cancer surgery. Lancet 2001;357:497–504. [CrossRef]
- Beets-Tan RGH, Lambregts DMJ, Maas M, Bipat S, Barbaro B, Caseiro-Alves F, et al. Magnetic resonance imaging for the clinical management of rectal cancer patients: recommendations from the 2012 European Society of Gastrointestinal and Abdominal Radiology (ESGAR) consensus meeting. Eur Radiol 2013;23:2522–31. [CrossRef]
- Brown G, Radcliffe AG, Newcombe RG, Dallimore NS, Bourne MW, Williams GT. Preoperative assessment of prognostic factors in rectal cancer using high-resolution magnetic resonance imaging. British Journal of Surgery 2003;90:355–64. [CrossRef]
- Fernandes MC, Gollub MJ, Brown G. The importance of MRI for rectal cancer evaluation. Surg Oncol 2022;43:101739. [CrossRef]
- Gollub MJ, Arya S, Beets-Tan RG, dePrisco G, Gonen M, Jhaveri K, et al. Use of magnetic resonance imaging in rectal cancer patients: Society of Abdominal Radiology (SAR) rectal cancer disease-focused panel (DFP) recommendations 2017. Abdominal Radiology 2018;43:2893–902. [CrossRef]
- Shihab OC, Moran BJ, Heald RJ, Quirke P, Brown G. MRI staging of low rectal cancer. Eur Radiol 2009;19:643–50. [CrossRef]
- Jhaveri KS, Hosseini-Nik H. MRI of rectal cancer: An overview and update on recent advances. American Journal of Roentgenology 2015;205:W42–55. [CrossRef]
- Taylor FGM, Swift RI, Blomqvist L, Brown G. A systematic approach to the interpretation of preoperative staging MRI for rectal cancer. American Journal of Roentgenology 2008;191:1827–35. [CrossRef]
- Horvat N, Rocha CCT, Oliveira BC, Petkovska I, Gollub MJ. MRI of rectal cancer: Tumor staging, imaging techniques, and management. Radiographics 2019;39:367–87. [CrossRef]
- Al-Sukhni E, Milot L, Fruitman M, Beyene J, Victor JC, Schmocker S, et al. Diagnostic accuracy of MRI for assessment of T category, lymph node metastases, and circumferential resection margin involvement in patients with rectal cancer: a systematic review and meta-analysis. Ann Surg Oncol 2012;19:2212–23. [CrossRef]
- Patel UB, Blomqvist LK, Taylor F, George C, Guthrie A, Bees N, et al. MRI after treatment of locally advanced rectal cancer: How to report tumor response - The MERCURY experience. American Journal of Roentgenology 2012;199. [CrossRef]
- Rullier A, Laurent C, Vendrely V, Le Bail B, Bioulac-Sage P, Rullier E. Impact of colloid response on survival after preoperative radiotherapy in locally advanced rectal carcinoma. Am J Surg Pathol 2005;29:602–6. [CrossRef]
- Compton CC. Key issues in reporting common cancer specimens: problems in pathologic staging of colon cancer. Arch Pathol Lab Med 2006;130:318–24. [CrossRef]
- Vecchio FM, Valentini V, Minsky BD, Padula GDA, Venkatraman ES, Balducci M, et al. The relationship of pathologic tumor regression grade (TRG) and outcomes after preoperative therapy in rectal cancer. Int J Radiat Oncol Biol Phys 2005;62:752–60. [CrossRef]
- Oberholzer K, Junginger T, Heintz A, Kreft A, Hansen T, Lollert A, et al. Rectal Cancer: MR imaging of the mesorectal fascia and effect of chemoradiation on assessment of tumor involvement. Journal of Magnetic Resonance Imaging 2012;36:658–63. [CrossRef]
- Thies S, Langer R. Tumor regression grading of gastrointestinal carcinomas after neoadjuvant treatment. Front Oncol 2013;3 OCT:64056. [CrossRef]
- Hou M, Sun JH. Emerging applications of radiomics in rectal cancer: State of the art and future perspectives. World J Gastroenterol 2021;27:3802–14. [CrossRef]
- Zhang S, Yu M, Chen D, Li P, Tang B, Li J. Role of MRI based radiomics in locally advanced rectal cancer (Review). Oncol Rep 2022;47. [CrossRef]
- Rizzo S, Botta F, Raimondi S, Origgi D, Fanciullo C, Morganti AG, et al. Radiomics: the facts and the challenges of image analysis. Eur Radiol Exp 2018;2:36. [CrossRef]
- Shur JD, Doran SJ, Kumar S, Ap Dafydd D, Downey K, O’connor JPB, et al. Radiomics in Oncology: A Practical Guide. Radiographics 2021;41:1717. [CrossRef]
- Miranda J, Horvat N, Araujo-Filho JAB, Albuquerque KS, Charbel C, Trindade BMC, et al. The Role of Radiomics in Rectal Cancer. J Gastrointest Cancer 2023;54:1158–80. [CrossRef]
- Ma X, Shen F, Jia Y, Xia Y, Li Q, Lu J. MRI-based radiomics of rectal cancer: preoperative assessment of the pathological features. BMC Med Imaging 2019;19. [CrossRef]
- Yu X, Song W, Guo D, Liu H, Zhang H, He X, et al. Preoperative Prediction of Extramural Venous Invasion in Rectal Cancer: Comparison of the Diagnostic Efficacy of Radiomics Models and Quantitative Dynamic Contrast-Enhanced Magnetic Resonance Imaging. Front Oncol 2020;10. [CrossRef]
- Zhou X, Yi Y, Liu Z, Cao W, Lai B, Sun K, et al. Radiomics-Based Pretherapeutic Prediction of Non-response to Neoadjuvant Therapy in Locally Advanced Rectal Cancer. Ann Surg Oncol 2019;26:1676–84. [CrossRef]
- Li ZY, Wang XD, Li M, Liu XJ, Ye Z, Song B, et al. Multi-modal radiomics model to predict treatment response to neoadjuvant chemotherapy for locally advanced rectal cancer. World J Gastroenterol 2020;26:2368–402. [CrossRef]
- Giannini V, Mazzetti S, Bertotto I, Chiarenza C, Cauda S, Delmastro E, et al. Predicting locally advanced rectal cancer response to neoadjuvant therapy with 18F-FDG PET and MRI radiomics features. Eur J Nucl Med Mol Imaging 2019;46:878–88. [CrossRef]
- Awiwi MO, Kaur H, Ernst R, Rauch GM, Morani AC, Stanietzky N, et al. Restaging MRI of Rectal Adenocarcinoma after Neoadjuvant Chemoradiotherapy: Imaging Findings and Potential Pitfalls. Radiographics 2023;43:e220135. [CrossRef]
- Sclafani F, Brown G, Cunningham D, Wotherspoon A, Mendes LST, Balyasnikova S, et al. Comparison between MRI and pathology in the assessment of tumour regression grade in rectal cancer. Br J Cancer 2017;117:1478–85. [CrossRef]
- Jankovic A, Kovac JD, Dakovic M, Mitrovic M, Saponjski D, Milicevic O, et al. MRI Tumor Regression Grade Combined with T2-Weighted Volumetry May Predict Histopathological Response in Locally Advanced Rectal Cancer following Neoadjuvant Chemoradiotherapy-A New Scoring System Proposal. Diagnostics (Basel) 2023;13. [CrossRef]
- Tustison NJ, Avants BB, Cook PA, Zheng Y, Egan A, Yushkevich PA, et al. N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 2010;29:1310–20. [CrossRef]
- Bonomo P, Socarras Fernandez J, Thorwarth D, Casati M, Livi L, Zips D, et al. Simulation CT-based radiomics for prediction of response after neoadjuvant chemo-radiotherapy in patients with locally advanced rectal cancer. Radiat Oncol 2022;17:84. [CrossRef]
- Petresc B, Lebovici A, Caraiani C, Feier DS, Graur F, Buruian MM. Pre-Treatment T2-WI Based Radiomics Features for Prediction of Locally Advanced Rectal Cancer Non-Response to Neoadjuvant Chemoradiotherapy: A Preliminary Study. Cancers (Basel) 2020;12:1–18. [CrossRef]
- Voogt ELK, Nordkamp S, Van Zoggel DMGI, Daniëls-Gooszen AW, Nieuwenhuijzen GAP, Bloemen JG, et al. MRI tumour regression grade in locally recurrent rectal cancer. BJS Open 2022;6. [CrossRef]
- Li Y, Liu W, Pei Q, Zhao L, Güngör C, Zhu H, et al. Predicting pathological complete response by comparing MRI-based radiomics pre- and postneoadjuvant radiotherapy for locally advanced rectal cancer. Cancer Med 2019;8:7244–52. [CrossRef]
- Chen H, Shi L, Nguyen KNB, Monjazeb AM, Matsukuma KE, Loehfelm TW, et al. MRI Radiomics for Prediction of Tumor Response and Downstaging in Rectal Cancer Patients after Preoperative Chemoradiation. Adv Radiat Oncol 2020;5:1286. [CrossRef]
- Cui Y, Yang X, Shi Z, Yang Z, Du X, Zhao Z, et al. Radiomics analysis of multiparametric MRI for prediction of pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Eur Radiol 2019;29:1211–20. [CrossRef]
- Kocak B, Akinci D’Antonoli T, Mercaldo N, Alberich-Bayarri A, Baessler B, Ambrosini I, et al. METhodological RadiomICs Score (METRICS): a quality scoring tool for radiomics research endorsed by EuSoMII. Insights Imaging 2024;15:8. [CrossRef]
- Chiloiro G, Cusumano D, de Franco P, Lenkowicz J, Boldrini L, Carano D, et al. Does restaging MRI radiomics analysis improve pathological complete response prediction in rectal cancer patients? A prognostic model development. Radiol Med 2022;127:11–20. [CrossRef]
- Petkovska I, Tixier F, Ortiz EJ, Golia Pernicka JS, Paroder V, Bates DD, et al. Clinical utility of radiomics at baseline rectal MRI to predict complete response of rectal cancer after chemoradiation therapy. Abdom Radiol (NY) 2020;45:3608–17. [CrossRef]
- Li Y, Liu W, Pei Q, Zhao L, Güngör C, Zhu H, et al. Predicting pathological complete response by comparing MRI-based radiomics pre- and postneoadjuvant radiotherapy for locally advanced rectal cancer. Cancer Med 2019;8:7244–52. [CrossRef]
- Boldrini L, Cusumano D, Chiloiro G, Casà C, Masciocchi C, Lenkowicz J, et al. Delta radiomics for rectal cancer response prediction with hybrid 0.35 T magnetic resonance-guided radiotherapy (MRgRT): a hypothesis-generating study for an innovative personalized medicine approach. Radiol Med 2019;124:145–53. [CrossRef]
- Song M, Li S, Wang H, Hu K, Wang F, Teng H, et al. MRI radiomics independent of clinical baseline characteristics and neoadjuvant treatment modalities predicts response to neoadjuvant therapy in rectal cancer. Br J Cancer 2022;127:249. [CrossRef]
| MRI Scanner |
Echo time (ms) |
Repetition Time (ms) |
Slice Thickness (mm) |
FOV (mm) |
Acquisition Matrix |
Flip Angle |
|
DISCOVERY MR750 |
61,307 | 1400 | 2 | 100 | 0/224/224/0 | 90 |
| Signa HDxt | 65 | 1200 | 2 | 100 | 0/288/256 | 90 |
| Symphony Tim | 84 | 4200 | 4 | 100 | 320/0/0/256 | 150 |
| Grade | Radiologic Response | Description |
| mrTRG 1 | Complete Response (CR) | Complete regression: no evidence of tumor signal or barely visible linear fibrotic scar (low signal intensity) in the mucosa/submucosal layer of previous tumor site |
| mrTRG 2 | Near complete Response (n-CR) | Good regression: predominant low signal intensity fibrotic scar with no obvious residual tumor signal |
| mrTRG 3 | Moderate Response | Moderate regression: >50% low signal intensity fibrosis/mucin areas, but there are obvious areas of intermediate signal intensity |
| mrTRG 4 | Mild Response | Slight regression: few areas of low signal intensity fibrosis or mucin but mostly tumor signal intensity on T2w images |
| mrTRG 5 | No Response | Intermediate signal intensity, same appearances as the original tumor on T2w images |
| Population Characteristics | N (%) |
| Age (mean ± standard deviation) | 63 ± 12 years |
| Male | 65 |
| Female | 35 |
| T | T2 8,14% |
| T3 9,3% | |
| T3a 10,47% | |
| T3b 37,21% | |
| T3c 15,12% | |
| T3d 3,49% | |
| T4 10,47% | |
| T4a 3,49% | |
| T4b 2,33% | |
| N | N0 3,49% |
| N1 31,4% | |
| N1b 2,33% | |
| N2 62,8% | |
| M | M0 86,05% |
| M1 13,95% | |
| EMVI | absent 63,95% |
| present 36,05% | |
| CT | 1,16% |
| CTRT | 61,63% |
| RTSC | 19,77% |
| CT IND>CTRT | 16,28% |
| RT | 1,16% |
| mrTRG | mrTRG1 1,15% |
| mrTRG2 19,54% | |
| mrTRG3 67,82% | |
| mrTRG4 4,60% | |
| mrTRG5 2,30% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




