Submitted:
04 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Genetic Testing
2.2. Clinical Assessment
2.3. Literature Review
2.4. Ethical Considerations
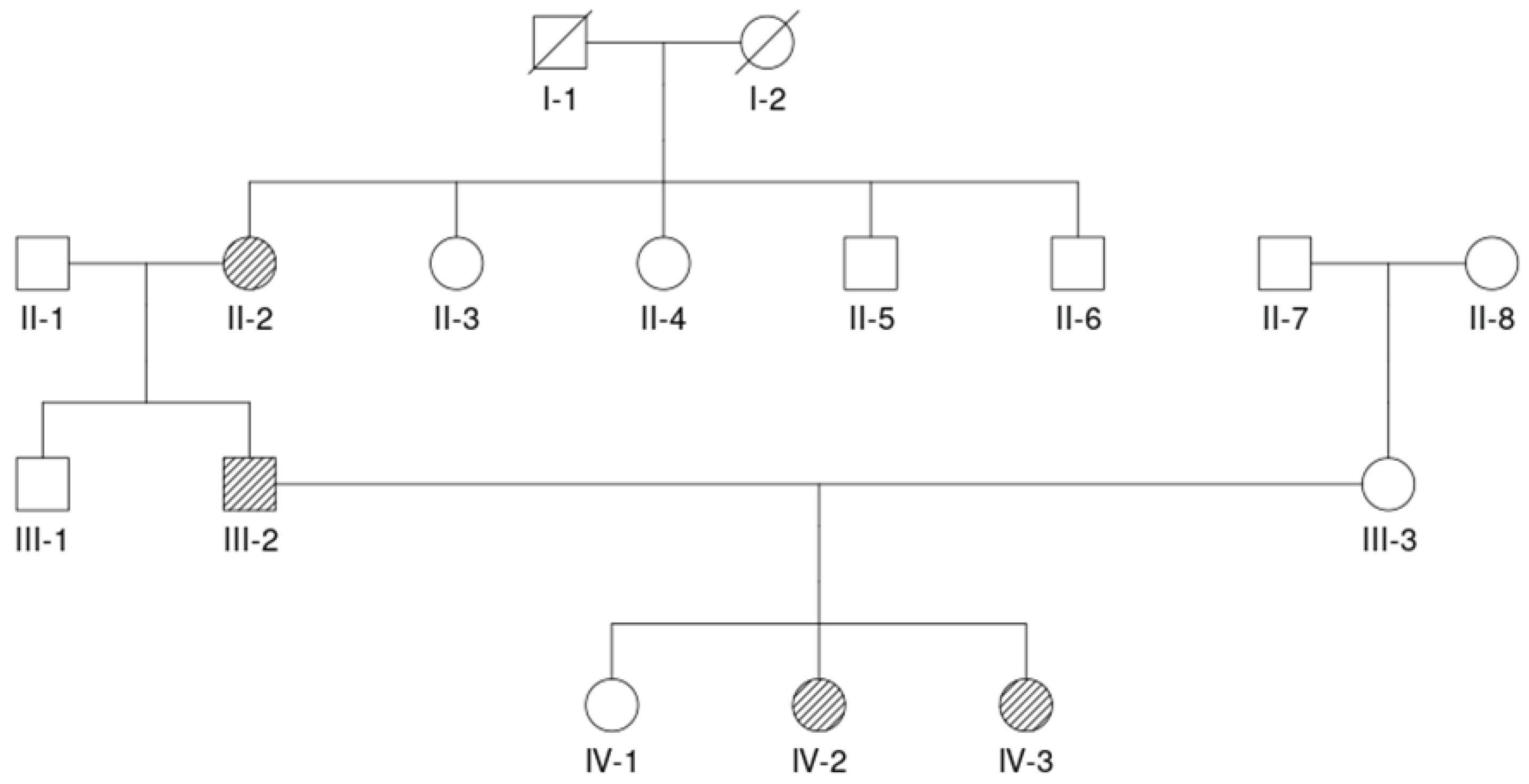
3. Case Report
3.1. Proband (IV-3)
3.2. Sister (IV-2)
3.3. Sister (IV-1)
3.4. Father (III-2)
3.5. Paternal Grandmother (II-2)
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| RNA | Ribonucleic acid |
| miRNA | Micro- Ribonucleic acid |
| ADHD | Attention Deficit Hyperactivity Disorder |
| CNV | Copy number variant |
| RISC | RNA-induced silencing complex |
| AGO | Argonaute |
| mRNA | Messenger Ribonucleic acid |
| GW | glycine–tryptophan |
| UBA | ubiquitin-associated |
| RRM | RNA recognition motif |
| DDD | Deciphering Developmental Disorders |
| CMA | Chromosomal microarray |
| MRI | Magnetic Resonance Imaging |
| SNP | Single Nucleotide Polymorphism |
| APGAR | Appearance, Pulse, Grimace, Activity, Respiration |
| IQ | Intelligence Quotient |
| WPSSI-R | Wechsler Preschool and Primary Scale of Intelligence—Revised |
| WISC-III | Wechsler Intelligence Scale for Children—Third Edition |
| ASD | autism spectrum disorder |
| ODD | Oppositional Defiant Disorder |
| ID | Intellectual disability |
| TOF | tetralogy of Fallot |
| PCGC | Paediatric Cardiac Genomics Consortium |
| PTCs | premature termination codons |
| NMD | nonsense-mediated decay |
References
- Chang, S.; Wen, S.; Chen, D.; Jin, P. Small regulatory RNAs in neurodevelopmental disorders. Hum Mol Genet 2009, 18, R18–26. [Google Scholar] [CrossRef] [PubMed]
- Goel, H.; Goel, A. MicroRNA and Rare Human Diseases. Genes (Basel) 2024, 15. [Google Scholar] [CrossRef]
- Johnson, S.T.; Chu, Y.; Liu, J.; Corey, D.R. Impact of scaffolding protein TNRC6 paralogs on gene expression and splicing. RNA 2021, 27, 1004–1016. [Google Scholar] [CrossRef]
- Yao, B.; La, L.B.; Chen, Y.C.; Chang, L.J.; Chan, E.K. Defining a new role of GW182 in maintaining miRNA stability. EMBO Rep 2012, 13, 1102–1108. [Google Scholar] [CrossRef]
- Yao, B.; Li, S.; Jung, H.M.; Lian, S.L.; Abadal, G.X.; Han, F.; Fritzler, M.J.; Chan, E.K. Divergent GW182 functional domains in the regulation of translational silencing. Nucleic Acids Res 2011, 39, 2534–2547. [Google Scholar] [CrossRef]
- Baillat, D.; Shiekhattar, R. Functional dissection of the human TNRC6 (GW182-related) family of proteins. Mol Cell Biol 2009, 29, 4144–4155. [Google Scholar] [CrossRef]
- Meister, G.; Landthaler, M.; Peters, L.; Chen, P.Y.; Urlaub, H.; Luhrmann, R.; Tuschl, T. Identification of novel argonaute-associated proteins. Curr Biol 2005, 15, 2149–2155. [Google Scholar] [CrossRef]
- Iossifov, I.; O'Roak, B.J.; Sanders, S.J.; Ronemus, M.; Krumm, N.; Levy, D.; Stessman, H.A.; Witherspoon, K.T.; Vives, L.; Patterson, K.E.; et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014, 515, 216–221. [Google Scholar] [CrossRef]
- Granadillo, J.L.; P.A.S., A; Guo, H.; Xia, K.; Angle, B.; Bontempo, K.; Ranells, J.D.; Newkirk, P.; Costin, C.; Viront, J.; et al. Pathogenic variants in TNRC6B cause a genetic disorder characterised by developmental delay/intellectual disability and a spectrum of neurobehavioural phenotypes including autism and ADHD. J Med Genet 2020, 57, 717–724. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.X.; Chen, Y.; Guo, H.R.; Chen, G.F. Systematic Review and Bioinformatic Analysis of microRNA Expression in Autism Spectrum Disorder Identifies Pathways Associated With Cancer, Metabolism, Cell Signaling, and Cell Adhesion. Front Psychiatry 2021, 12, 630876. [Google Scholar] [CrossRef] [PubMed]
- Harvey, D.C.; Verma, R.; Sedaghat, B.; Hjelm, B.E.; Morton, S.U.; Seidman, J.G.; Kumar, S.R. Mutations in genes related to myocyte contraction and ventricular septum development in non-syndromic tetralogy of Fallot. Front Cardiovasc Med 2023, 10, 1249605. [Google Scholar] [CrossRef]
- Takagi, M.; Ono, S.; Kumaki, T.; Nishimura, N.; Murakami, H.; Enomoto, Y.; Naruto, T.; Ueda, H.; Kurosawa, K. Complex congenital cardiovascular anomaly in a patient with AGO1-associated disorder. Am J Med Genet A 2023, 191, 882–892. [Google Scholar] [CrossRef]
- Niu, Y.; Qian, Q.; Li, J.; Gong, P.; Jiao, X.; Mao, X.; Xiao, B.; Long, L.; Yang, Z. De novo variants in AGO1 recapitulate a heterogeneous neurodevelopmental disorder phenotype. Clin Genet 2022, 101, 459–465. [Google Scholar] [CrossRef]
- Lessel, D.; Zeitler, D.M.; Reijnders, M.R.F.; Kazantsev, A.; Hassani Nia, F.; Bartholomaus, A.; Martens, V.; Bruckmann, A.; Graus, V.; McConkie-Rosell, A.; et al. Germline AGO2 mutations impair RNA interference and human neurological development. Nat Commun 2020, 11, 5797. [Google Scholar] [CrossRef]
- Ishiura, H.; Doi, K.; Mitsui, J.; Yoshimura, J.; Matsukawa, M.K.; Fujiyama, A.; Toyoshima, Y.; Kakita, A.; Takahashi, H.; Suzuki, Y.; et al. Expansions of intronic TTTCA and TTTTA repeats in benign adult familial myoclonic epilepsy. Nat Genet 2018, 50, 581–590. [Google Scholar] [CrossRef] [PubMed]
- Eising, E.; Carrion-Castillo, A.; Vino, A.; Strand, E.A.; Jakielski, K.J.; Scerri, T.S.; Hildebrand, M.S.; Webster, R.; Ma, A.; Mazoyer, B.; et al. A set of regulatory genes co-expressed in embryonic human brain is implicated in disrupted speech development. Mol Psychiatry 2019, 24, 1065–1078. [Google Scholar] [CrossRef] [PubMed]
- Yahia, A.; Li, D.; Lejerkrans, S.; Rajagopalan, S.; Kalnak, N.; Tammimies, K. Whole exome sequencing and polygenic assessment of a Swedish cohort with severe developmental language disorder. Hum Genet 2024, 143, 169–183. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Zhong, H.; Wu, B.; Cui, Y.; Li, J.; Jia, X.; Yu, C.; Li, D.; Shu, J.; Cai, C. Identification of the synonymous variant c.3141G > A in TNRC6B gene that altered RNA splicing by minigene assay. Mol Biol Rep 2024, 51, 899. [Google Scholar] [CrossRef]
- Bellido-Cuellar, S.; Perez de la Fuente, R.; Lezana-Rosales, J.M.; Sanchez-Calvin, M.T.; Saiz-Diaz, R.A.; Gonzalez de la Aleja, J. Epilepsy and Autism spectrum disorder caused by a pathogenic variant in TNRC6B. Seizure 2023, 110, 117–118. [Google Scholar] [CrossRef]
- Yang, Q.; Ou, S.; Zhou, X.; Yi, S.; Lin, L.; Yi, S.; Zhang, S.; Qin, Z.; Luo, J. Novel variants in TNRC6B cause global developmental delay with speech and behavioral abnormalities, short stature, low body weight, cafe-au-lait spots, and metabolic abnormality. Mol Genet Genomic Med 2024, 12, e2408. [Google Scholar] [CrossRef]
- Mangkalaphiban, K.; Fu, L.; Du, M.; Thrasher, K.; Keeling, K.M.; Bedwell, D.M.; Jacobson, A. Extended stop codon context predicts nonsense codon readthrough efficiency in human cells. Nat Commun 2024, 15, 2486. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



