Submitted:
03 March 2026
Posted:
04 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Taiwan Pulmonary Specialist Writing Examination Questions
2.2. Question Categorization and Category Definition
2.3. Google Gemma3 family, OpenAI OSS-20B, and GPT-4Turbo
2.4. Prompt Input and Response Output
2.5. Performance Analysis and Statistics
2.5.1. Random Guessing
2.5.2. Intermodal Comparing
2.6. Software and Hardware
3. Results
3.1. Inference Latency
3.2. Instruction Adherence
3.3. Year-by-Year Performance by Model
| category | # | 1B | 4B | 12B | 27B | OSS-20B | GPT-4T |
|---|---|---|---|---|---|---|---|
| Lung cancer | 186 | 42 | 79 | 81 | 109 | 137 | 135 |
| Infection | 134 | 33 | 53 | 67 | 77 | 103 | 90 |
| Critical care | 103 | 25 | 34 | 41 | 58 | 66 | 70 |
| MV | 98 | 23 | 30 | 38 | 42 | 61 | 62 |
| Tuberculosis, | 81 | 18 | 30 | 39 | 48 | 49 | 47 |
| Asthma, | 69 | 11 | 24 | 32 | 38 | 43 | 43 |
| COPD | 67 | 16 | 25 | 30 | 34 | 45 | 36 |
| Esophageal disorders | 66 | 16 | 26 | 27 | 36 | 46 | 42 |
| PFT | 55 | 17 | 20 | 19 | 28 | 29 | 33 |
| Sleep medicine | 39 | 13 | 12 | 10 | 16 | 23 | 23 |
| Chest anatomy | 38 | 7 | 18 | 14 | 23 | 25 | 26 |
| Pharmacology | 37 | 8 | 16 | 17 | 23 | 28 | 25 |
| Interstitial lung disease | 35 | 7 | 13 | 11 | 17 | 30 | 24 |
| Pathophysiology | 30 | 8 | 11 | 18 | 19 | 22 | 19 |
| PE and DVT | 29 | 4 | 8 | 13 | 15 | 25 | 25 |
| Miscellaneous | 27 | 3 | 8 | 10 | 17 | 18 | 16 |
| Pneumothorax and others | 23 | 7 | 9 | 9 | 15 | 15 | 17 |
| Chest surgery | 22 | 4 | 9 | 10 | 15 | 15 | 15 |
| Autoimmune diseases | 14 | 5 | 3 | 8 | 7 | 10 | 10 |
| Bronchoscopy-related | 13 | 5 | 7 | 4 | 4 | 6 | 8 |
| Sarcoidosis and LAM | 11 | 2 | 2 | 3 | 6 | 7 | 10 |
| Pleural diseases | 9 | 2 | 4 | 2 | 4 | 3 | 5 |
| Vasculitis | 8 | 3 | 3 | 4 | 4 | 6 | 7 |
| Musculoskeletal issues | 3 | 0 | 1 | 2 | 2 | 3 | 2 |
| Tracheal disorders | 2 | 1 | 2 | 1 | 2 | 1 | 2 |
| Diaphragmatic problems | 1 | 0 | 0 | 1 | 1 | 1 | 1 |
4. Discussion
Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| LLM | Large language model |
| MCQ | Multiple-choice question |
| API | Application interface |
| AI | Artificial intelligence |
Appendix A. Definition of Clinical Categories
References
- Chen, C.H.; Hsu, S.H.; Hsieh, K.Y.; Lai, H.Y. The two-stage detection-after-segmentation model improves the accuracy of identifying subdiaphragmatic lesions. Sci. Rep. 2024, 14, 25414. [CrossRef]
- Chung, Y.; Jin, J.; Jo H.I.; Lee, H.; Kim, S.H.; Chung, S.J.; Yoon, H.J.; Park, J.; Jeon, J.Y. Diagnosis of Pneumonia by Cough Sounds Analyzed with Statistical Features and AI. Sensors. 2021, 21(21), 7036. [CrossRef]
- Olszewski, R.; Brzeziński, J.; Watros, K.; Rysz, J. Quantifying Readability in Chatbot-Generated Medical Texts Using Classical Linguistic Indices: A Review. Appl. Sci. 2026, 16, 1423. [CrossRef]
- Chen, C.H.; Chen, C.W.; Hsieh, K.Y.; Huang, K.E.; Lai, H.-Y. Limitations in Chest X-Ray Interpretation by Vision-Capable Large Language Models, Gemini 1.0, Gemini 1.5 Pro, GPT-4 Turbo, and GPT-4o. Diagnostics 2026, 16, 376. [CrossRef]
- Khalid, N.; Qayyum, A.; Bilal, M.; Al-Fuqaha, A.; Qadir, J. Privacy-preserving artificial intelligence in healthcare: Techniques and applications. Computers in Biology and Medicine 2023, 158, 106848. [CrossRef]
- Dennstädt, F.; Hastings, J.; Putora, P.M.; Schmerder, M.; Cihoric, N. Implementing large language models in healthcare while balancing control, collaboration, costs and security. npj Digit. Med. 2025, 8, 143. [CrossRef]
- Chen, C.; Hsieh, K.; Huang, K.; Lai, H. Comparing Vision-Capable Models, GPT-4 and Gemini, With GPT-3.5 on Taiwan’s Pulmonologist Exam. Cureus. 2024, 16(8), e67641. [CrossRef]
- Tsai, C.Y.; Hsieh, S.J.; Huang, H.H.; Deng, J.H.; Huang, Y.Y.; Cheng, P.Y. Performance of ChatGPT on the Taiwan urology board examination: insights into current strengths and shortcomings. World J Urol. 2024, 42(1):250. [CrossRef]
- Kamath, G.T.; Ferret, J.; Pathak, S.; et al. Gemma 3 Technical Report. arXiv 2025. [CrossRef]
- OpenAI: Agarwal, S.; Ahmad, L.; Ai, J.; Altman, S.; Applebaum, A.; Arbus, E.; et al. gpt-oss-120b & gpt-oss-20b Model Card. arXiv 2025. [CrossRef]
- Ollama GitHub. https://github.com/ollama/ollama (accessed on 03 March 2026).
- Van Rossum, G.; Drake, F.L. Python 3 Reference Manual. Scotts Valley, CA: CreateSpace; 2009.
- Hunter, J.D. Matplotlib: A 2D graphics environment. Computing in Science & Engineering 2007, 9, 90-95. [CrossRef]
- Kim, S.; Yun, S.; Lee, H.; Gubri, M.; Yoon, S.; Oh, S.J. ProPILE: Probing Privacy Leakage in Large Language Models. arXiv 2023. [CrossRef]
- Yan, B.; Li, K.; Xu, M.; Dong, Y.; Zhang, Y.; Ren, Z.; Cheng, X. On Protecting the Data Privacy of Large Language Models (LLMs): A Survey. arXiv 2024. [CrossRef]
- Gulyamov, S.; Gulyamov, S.; Rodionov, A.; Khursanov, R.; Mekhmonov, K.; Babaev, D.; Rakhimjonov, A. Prompt Injection Attacks in Large Language Models and AI Agent Systems: A Comprehensive Review of Vulnerabilities, Attack Vectors, and Defense Mechanisms. Information 2026, 17, 54. [CrossRef]
- Thirunavukarasu, A.J.; Ting, D.S.J.; Elangovan, K.; Gutierrez, L.; Tan, T.F.; Ting, D.S.W. Large language models in medicine. Nat. Med. 2023, 29, 1930–1940. [CrossRef]
- Xu, H.; Zhang, Z.; Yu, X.; Wu, Y.; Zha, Z.; Xu, B.; Xu, W.; Hu, M.; Peng, K. Targeted Training Data Extraction—Neighborhood Comparison-Based Membership Inference Attacks in Large Language Models. Appl. Sci. 2024, 14, 7118. [CrossRef]
- Seh, A.H.; Zarour, M.; Alenezi, M.; Sarkar, A.K.; Agrawal, A.; Kumar, R.; Ahmad Khan, R. Healthcare Data Breaches: Insights and Implications. Healthcare 2020, 8, 133. [CrossRef]
- Jiang, J.X.; Ross, J.S.; Bai, G. Ransomware Attacks and Data Breaches in US Health Care Systems. JAMA Netw Open. 2025, 8, e2510180. [CrossRef]
- Feretzakis, G.; Papaspyridis, K.; Gkoulalas-Divanis, A.; Verykios, V.S. Privacy-Preserving Techniques in Generative AI and Large Language Models: A Narrative Review. Information 2024, 15, 697. [CrossRef]
- Huang, C.H.; Hsiao, H.J.; Yeh, P.C.; Wu, K.C.; Kao, C.H. Performance of ChatGPT on Stage 1 of the Taiwanese medical licensing exam. Digital Health 2024, 10. [CrossRef]
- Kao, Y.S.; Chuang, W.K.; Yang, J. Use of ChatGPT on Taiwan’s Examination for Medical Doctors. Ann Biomed Eng. 2024, 52, 455-457. [CrossRef]
- Liu, C.-L.; Ho, C.-T.; Wu, T.-C. Custom GPTs Enhancing Performance and Evidence Compared with GPT-3.5, GPT-4, and GPT-4o? A Study on the Emergency Medicine Specialist Examination. Healthcare 2024, 12, 1726. [CrossRef]
- Ting, Y.T.; Hsieh, T.C.; Wang, Y.F.; et al. Performance of ChatGPT incorporated chain-of-thought method in bilingual nuclear medicine physician board examinations. Digit Health 2024, 10. [CrossRef]
- Hsieh, C.H.; Hsieh, H.Y.; Lin, H.P. Evaluating the performance of ChatGPT-3.5 and ChatGPT-4 on the Taiwan plastic surgery board examination. Heliyon 2024, 10, e34851. [CrossRef]
- Tkachenko, Alexei V. Thermodynamic Bound on Energy and Negentropy Costs of Inference in Deep Neural Networks. arXiv 2025. [CrossRef]
- Fernandez, J.; Na, C.; Tiwari, V.; Bisk, Y.; Luccioni, S.; Strubell, E. Energy considerations of large language model inference and efficiency optimizations. arXiv 2025. [CrossRef]
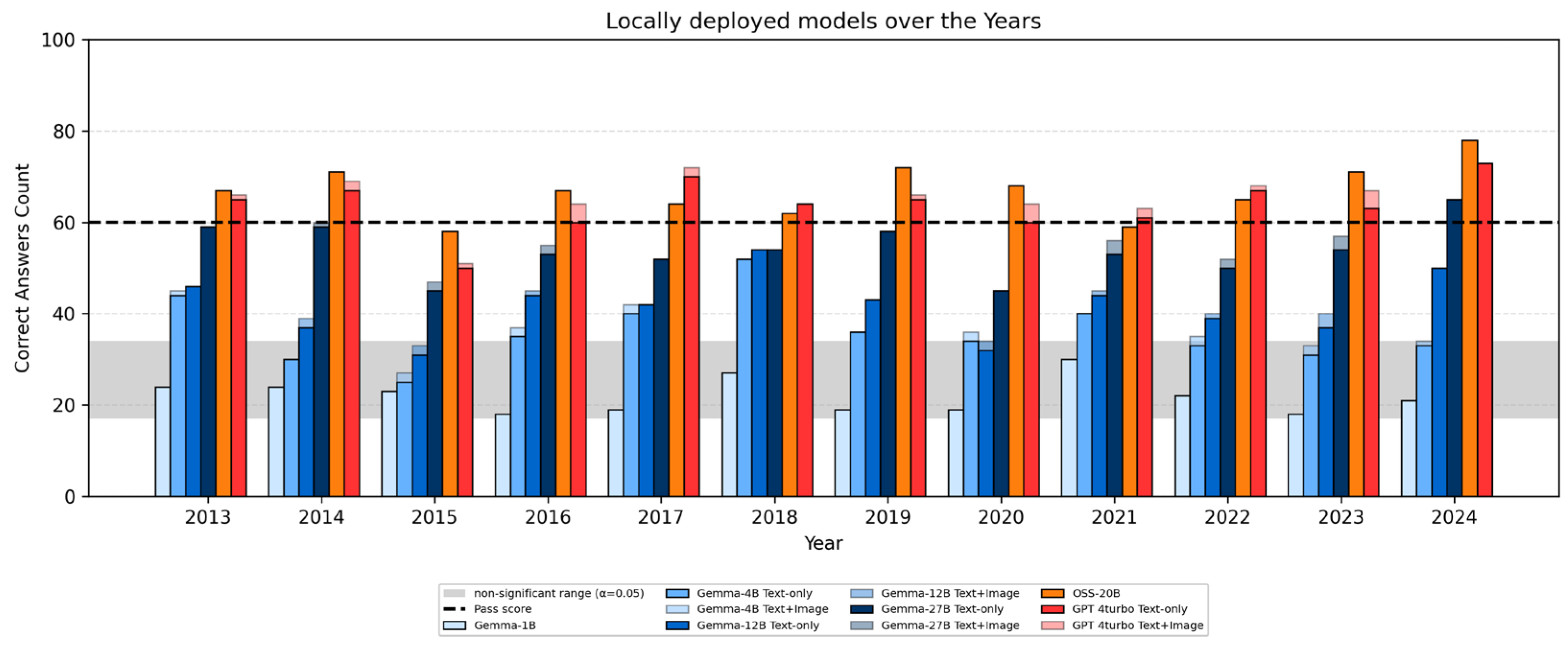
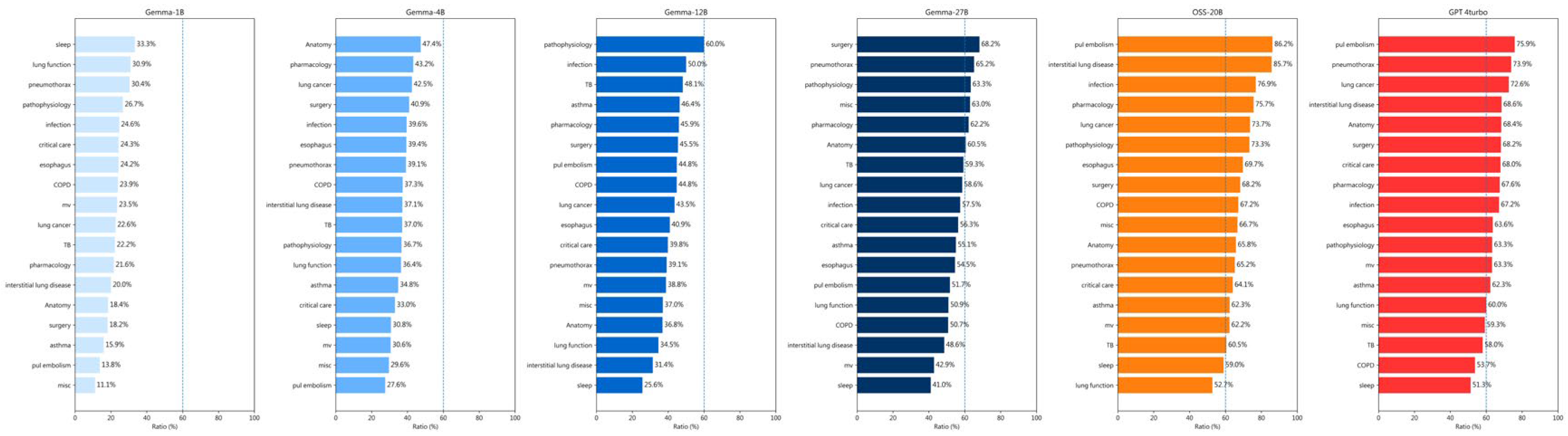
| 2013 | 2014 | 2015 | 2016 | 2017 | 2018 | 2019 | 2020 | 2021 | 2022 | 2023 | 2024 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T(V) | 99(1) | 95(5) | 96(4) | 95(5) | 97(3) | 99(1) | 99(1) | 94(6) | 96(4) | 96(4) | 93(7) | 97(3) |
| 27B | 59(0) | 59(1) | 45(2) | 53(2) | 52(0) | 54(0) | 58(0) | 45(0) | 53(3) | 50(2) | 54(3) | 65(0) |
| 12B | 46(0) | 37(2) | 31(2) | 44(1) | 42(0) | 54(0) | 43(0) | 32(2) | 44(1) | 39(1) | 37(3) | 50(0) |
| 4B | 44(1) | 30(0) | 25(2) | 35(2) | 40(2) | 52(0) | 36(0) | 34(2) | 40(0) | 33(2) | 31(2) | 33(1) |
| 1B | 24(0) | 24(2) | 23(2) | 18(2) | 19(1) | 27(1) | 19(0) | 19(0) | 30(1) | 22(2) | 18(3) | 21(2) |
| OSS | 67(1) | 71(3) | 58(2) | 67(1) | 64(1) | 62(0) | 72(1) | 68(2) | 59(1) | 65(0) | 71(3) | 78(0) |
| GPT | 65(1) | 67(2) | 50(1) | 60(4) | 70(2) | 64(0) | 65(1) | 60(4) | 61(2) | 67(1) | 63(4) | 73(0) |
| A | B | Mean Difference (T) | p-corr | Hedges’ g |
|---|---|---|---|---|
| OSS-20B | Gemma3-27B | -8.674031 | 1.501442e-05 | -2.025969 |
| OSS-20B | Gemma3-12B | -11.115588 | 1.781392e-06 | -3.976852 |
| OSS-20B | Gemma3-4B | -10.395901 | 3.005323e-06 | -4.558086 |
| OSS-20B | Gemma3-1B | -18.542707 | 1.201425e-08 | -8.906559 |
| Gemma3-27B | Gemma3-12B | -8.511444 | 1.501442e-05 | -2.068610 |
| Gemma3-27B | Gemma3-4B | 7.254426 | 3.269766e-05 | 2.773100 |
| Gemma3-27B | Gemma3-1B | -17.952853 | 1.526909e-08 | -6.710339 |
| Gemma3-12B | Gemma3-4B | 3.854888 | 2.676820e-03 | 0.803686 |
| Gemma3-12B | Gemma3-1B | 12.022477 | 9.128954e-07 | 3.942880 |
| Gemma3-4B | Gemma3-1B | -8.184753 | 1.576218e-05 | -2.687739 |
| GPT-4T | OSS-20B | -0.946449 | 3.642470e-01 | -0.213263 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




