Submitted:
02 March 2026
Posted:
02 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Novel Architecture with PVT v2 Backbone: We propose a new deep learning framework for automatic skin lesion segmentation that incorporates the Pyramid Vision Transformer v2 (PVT v2) as the core encoder. This design effectively captures multi-scale contextual features from dermoscopic images, enhancing the model’s ability to recognize lesions of varying sizes and complexities [13].
- Dual-Module Cooperative Mechanism: The framework introduces a synergistic dual-module mechanism within the decoder pathway. The first module is dedicated to the sophisticated integration of low-level spatial details and high-level semantic features, ensuring a more discriminative and complete feature representation. The second module employs an enhanced skip-connection strategy for dynamic feature fusion between the encoder and decoder, significantly improving the precision of boundary delineation and the recovery of fine-grained structures [10,12].
- Comprehensive Evaluation on Benchmark Datasets: We conduct extensive experiments on publicly available datasets, such as ISIC 2016 and SISC 2018. The results demonstrate that our proposed method achieves state-of-the-art performance, particularly in handling boundary ambiguity and small lesions, outperforming existing leading approaches in terms of standard segmentation metrics like Dice Coefficient and IOU. This validates the practical effectiveness and robustness of our model in a clinical simulation setting.
2. Related Work
2.1. CNNs for Skin Lesion Segmentation
2.2. Transformers for Skin Lesion Segmentation
3. Materials and Methods
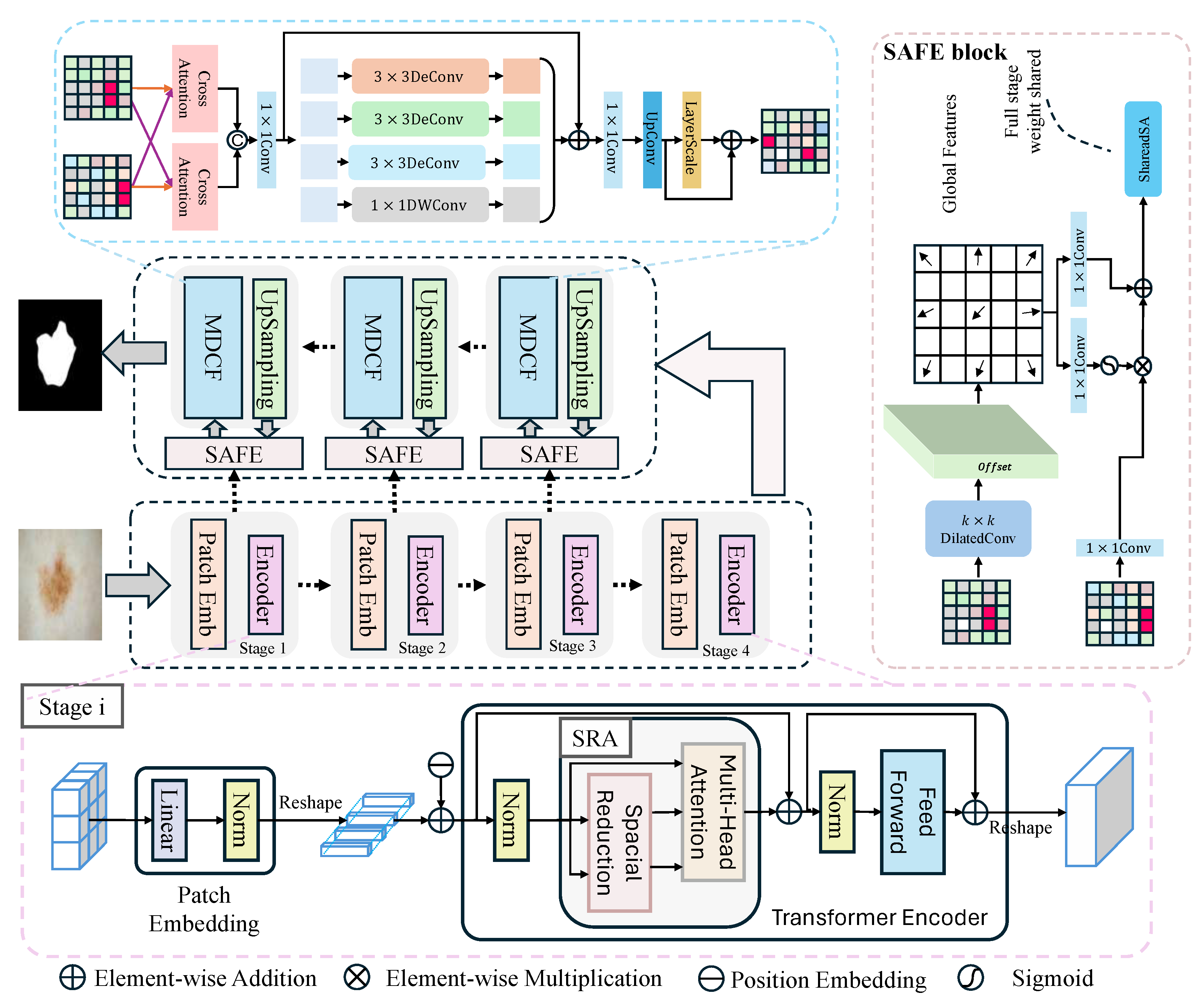
3.1. Architectural Overview
3.2. Encoder Model
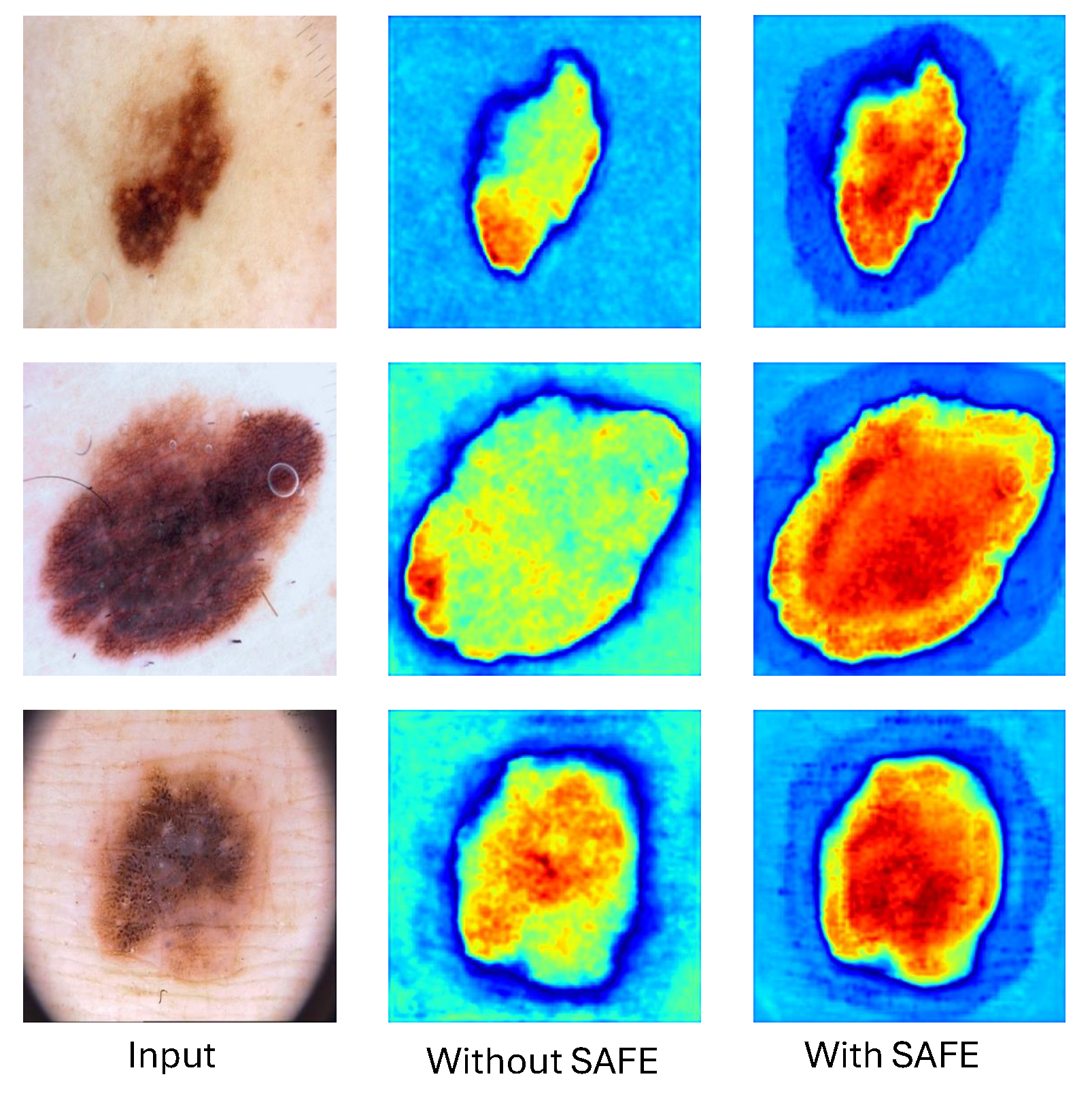
3.3. SAFE Model
3.4. Decoder Model
4. Results
4.1. Datasets
4.2. Evaluation Metrics
4.3. Implementation Details
4.4. Ablation Study
4.5. Comparison with the State-of-the-Art Methods
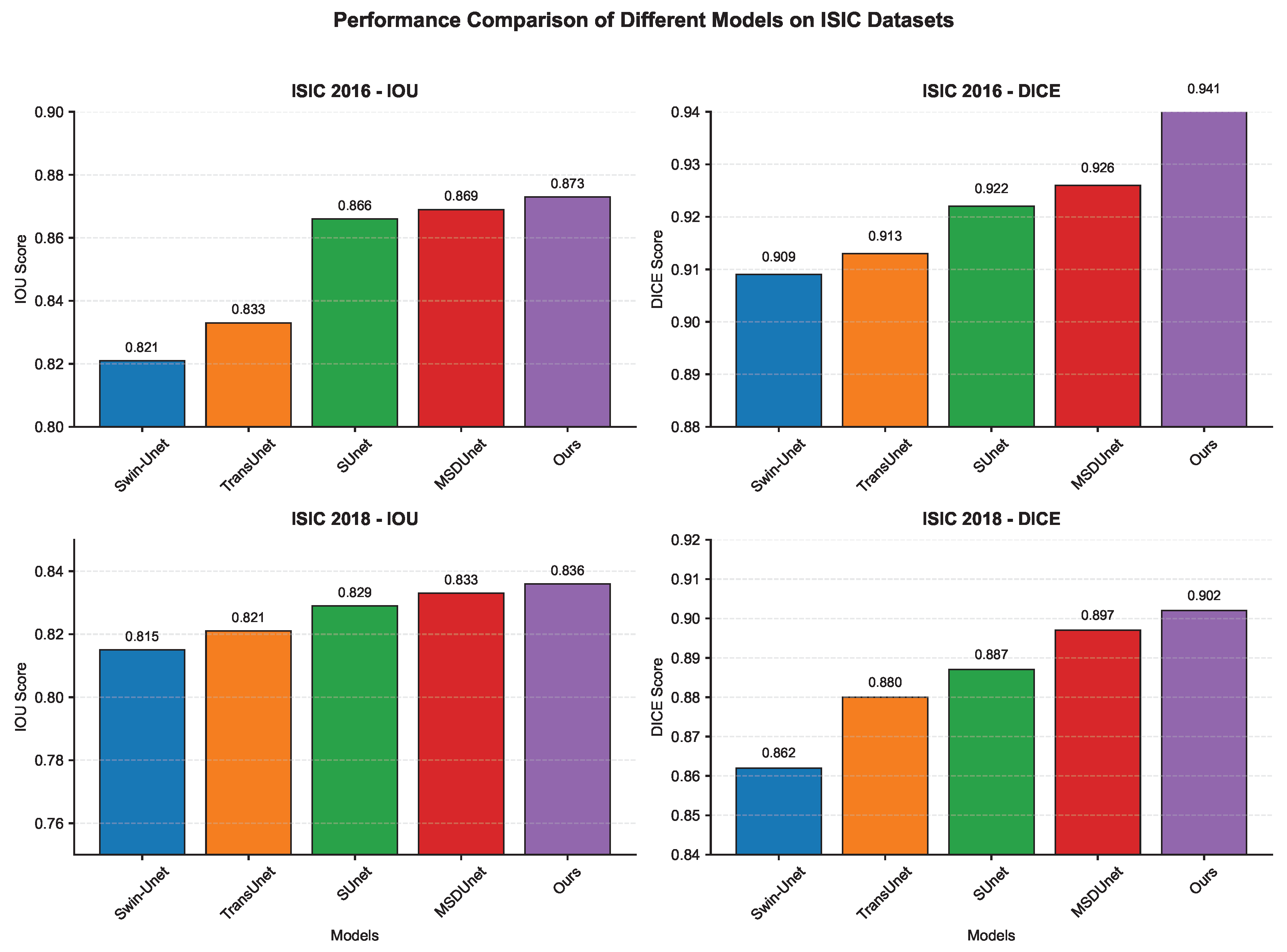
4.5.1. Quantitative Evaluation
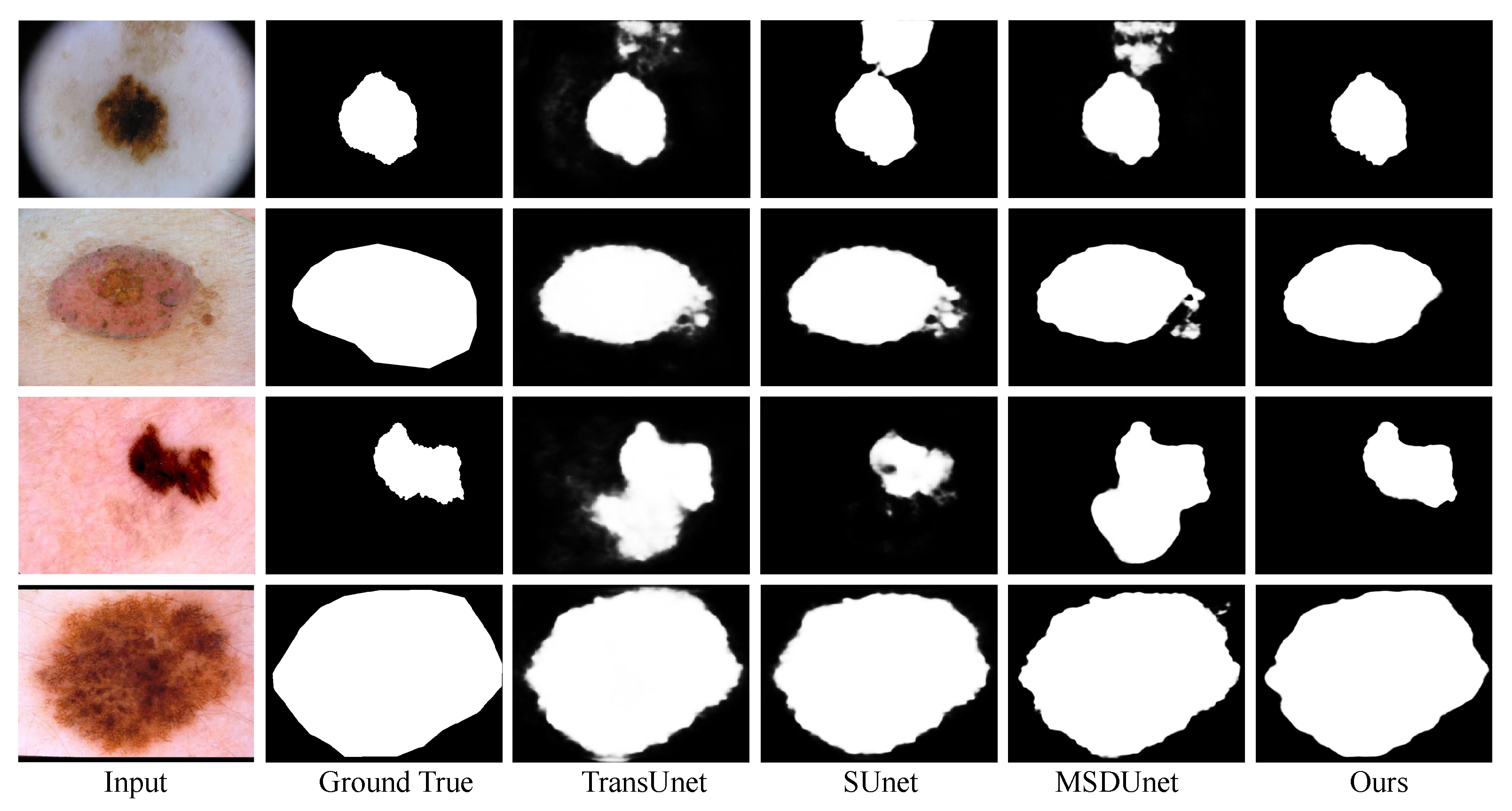
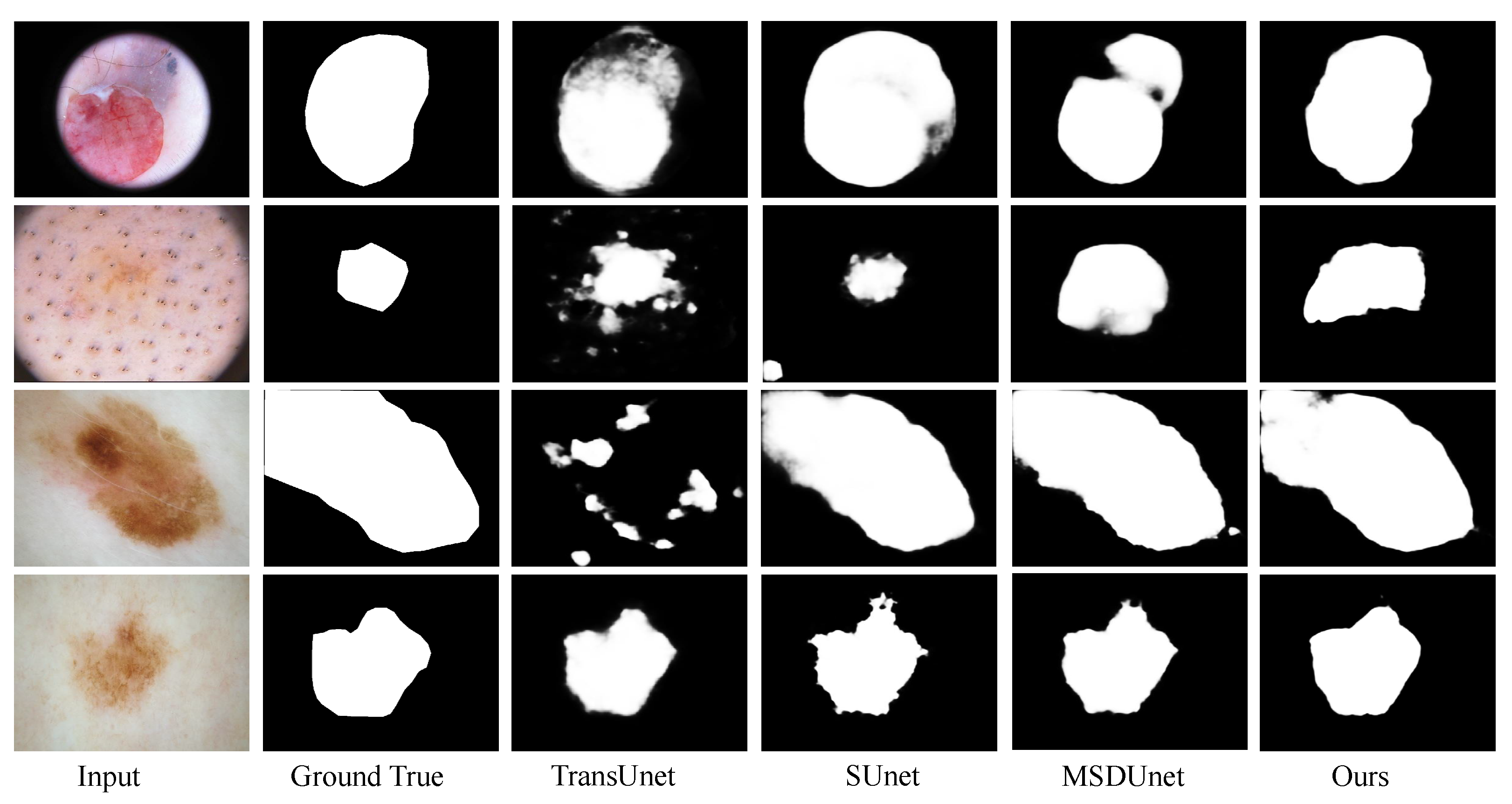
4.5.2. Qualitative Analysis
5. Discussion
6. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- R. L. Siegel et al., “Cancer statistics, 2022,” CA Cancer J. Clin., vol. 72, no. 1, pp. 7–33, 2022.
- H. Sung et al., “Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries,” CA Cancer J. Clin., vol. 71, no. 3, pp. 209–249, 2021.
- J. E. Gershenwald et al., “Melanoma staging: Evidence-based changes in the American Joint Committee on Cancer eighth edition cancer staging manual,” CA Cancer J. Clin., vol. 67, no. 6, pp. 472–492, 2017.
- H. Kittler et al., “Diagnostic accuracy of dermoscopy,” Lancet Oncol., vol. 3, no. 3, pp. 159–165, 2002.
- G. Argenziano et al., “Dermoscopy of pigmented skin lesions: Results of a consensus net meeting conducted via the Internet,” J. Amer. Acad. Dermatol., vol. 48, no. 5, pp. 679–693, 2003.
- C. Barata, M. E. Celebi, and J. S. Marques, “A survey of feature extraction in dermoscopy image analysis of skin cancer,” IEEE J. Biomed. Health Inform., vol. 23, no. 3, pp. 1096–1109, May 2019.
- A. Lochan Sharma, K. Sharma, and P. Ghosal, “Skin lesion segmentation: A systematic review of computational techniques, tools, and future directions,” Comput. Biol. Med., vol. 196, 110842, 2025.
- M. A. M. Almeida and I. A. X. Santos, “Classification models for skin tumor detection using texture analysis in medical images,” J. Imaging, vol. 6, no. 6, p. 51, 2020.
- A. Karimi, K. Faez, and S. Nazari, ‘‘DEU-Net: Dual-encoder U-Net for automated skin lesion segmentation,’’ IEEE Access, vol. 11, pp. 134804–134821, 2023.
- O. Ronneberger, P. Fischer, and T. Brox, “U-Net: Convolutional Networks for Biomedical Image Segmentation,” in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015, N. Navab, J. Hornegger, W. Wells, and A. Frangi, Eds. Cham: Springer, 2015, pp. 234-241. [CrossRef]
- M. H. Jafari, N. Karimi, E. Nasr-Esfahani, S. M. R. Soroushmehr, S. Samavi, K. Ward, and K. Najarian, “Skin lesion segmentation in clinical images using deep learning,” in Proc. 23rd Int. Conf. Pattern Recognit. (ICPR), 2016, pp. 337–342. [CrossRef]
- J. Chen, Y. Lu, Q. Yu, et al., “TransUNet: Transformers Make Strong Encoders for Medical Image Segmentation,” arXiv preprint arXiv:2102.04306, 2021.
- W. Wang, E. Xie, X. Li, et al., “PVTv2: Improved Pyramid Vision Transformer for Dense Prediction,” in Proc. IEEE/CVF Conf. on Computer Vision and Pattern Recognition (CVPR), New Orleans, LA, USA, 2022, pp. 548–558. doi: 10.1109/CVPR52688.2022.00064.
- D. Gutman et al., “Skin lesion analysis toward melanoma detection: A challenge at the 2016 International Symposium on Biomedical Imaging (ISBI), hosted by the International Skin Imaging Collaboration (ISIC)” arXiv:1605.01397, 2016.
- N. C. Codella et al., “Skin lesion analysis toward melanoma detection 2018: A challenge hosted by the International Skin Imaging Collaboration (ISIC),” arXiv:1902.03368, 2019.
- V. Badrinarayanan, A. Kendall, and R. Cipolla, “SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation,” IEEE Trans. Pattern Anal. Mach. Intell., vol. 39, no. 12, pp. 2481–2495, Dec. 2017. [CrossRef]
- A. Azhari, N. Yudistira, A. Widodo, et al., “Two-stage CNN with weakly supervised segmentation for skin lesion classification,” Multimedia Tools and Applications, published online, 2025. doi: 10.1007/s11042-025-21091-8.
- F. Milletarì, N. Navab, and S.-A. Ahmadi, “V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation,” in Proc. Fourth International Conference on 3D Vision (3DV), Stanford, CA, USA, 2016, pp. 565-571. doi: 10.1109/3DV.2016.79.
- T.-Y. Lin, P. Goyal, R. Girshick, K. He, and P. Dollár, “Focal Loss for Dense Object Detection,” in Proc. IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 2017, pp. 2999–3007. doi: 10.1109/ICCV.2017.324.
- W. Wang, Y. Luo, and X. Wang, “BEFNet: A Hybrid CNN-Mamba Architecture for Accurate Skin Lesion Image Segmentation,” in Proc. IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Lisbon, Portugal, 2024, pp. 3795–3798. doi: 10.1109/BIBM62325.2024.10822106.
- C. Yuan, D. Zhao, and S. S. Agaian, “UCM-Net: A lightweight and efficient solution for skin lesion segmentation using MLP and CNN,” Biomedical Signal Processing and Control, vol. 96, p. 106573, 2024. [CrossRef]
- H. Cao, Y. Wang, J. Chen, et al., “Swin-Unet: Unet-like Pure Transformer for Medical Image Segmentation,” arXiv preprint arXiv:2105.05537, 2021.
- E. Xie, W. Wang, Z. Yu, et al., “SegFormer: Simple and Efficient Design for Semantic Segmentation with Transformers,” in Advances in Neural Information Processing Systems (NeurIPS), 2021.
- T.-Y. Lin, P. Dollár, R. Girshick, K. He, and S. Belongie, “Feature Pyramid Networks for Object Detection,” in Proc. IEEE Conf. on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 2017, pp. 2117–2125. doi: 10.1109/CVPR.2017.106.
- L.-C. Chen, Y. Zhu, G. Papandreou, F. Schroff, and H. Adam, “Encoder-Decoder with Atrous Separable Convolution for Semantic Image Segmentation,” in Proc. European Conf. on Computer Vision (ECCV), Munich, Germany, 2018, pp. 801–818. doi: 10.1007/978-3-030-01234-2_49.
- A. He, K. Wang, T. Li, C. Du, S. Xia, and H. Fu, “H2Former: An efficient hierarchical hybrid Transformer for medical image segmentation,” IEEE Trans. Med. Imaging, vol. 42, no. 9, pp. 2763–2775, 2023. [CrossRef]
- C.-M. Fan, T.-J. Liu, and K.-H. Liu, “SUNet: Swin Transformer with UNet for image denoising,” in Proc. 2022 IEEE Int. Symp. Circuits Syst. (ISCAS), pp. 2333–2337, 2022. [CrossRef]
- X. Li, L. Linli, X. Xing, H. Liao, W. Wang, Q. Dong, and C. Yuan, “MSDUNet: A model based on feature multi-scale and dual-input dynamic enhancement for skin lesion segmentation,” IEEE Trans. Med. Imaging, vol. 44, no. 7, pp. 2819–2830, Jul. 2025. [CrossRef]
- A. Bilal, A. H. Khan, K. Almohammadi, S. A. A. Ghamdi, H. Long, and H. Malik, “PDCNET: Deep convolutional neural network for classification of periodontal disease using dental radiographs,” IEEE Access, vol. 12, pp. 150147–150168, 2024.
- L. Bi, J. Kim, E. Ahn, A. Kumar, M. Fulham, and D. Feng, “Dermoscopic image segmentation via multistage fully convolutional networks,” IEEE Trans. Biomed. Eng., vol. 64, no. 9, pp. 2065–2074, Sep. 2017.
- M. A. Khan, I. Haider, M. Nazir, A. Nabeeh, and E. S. M. El-Kenawy, “A multi-stage melanoma recognition framework with deep residual neural network and hyperparameter optimization-based decision support in dermoscopy images,” Expert Syst. Appl., vol. 215, Apr. 2023, Art. no. 119251.
- G. Zhang, X. Shen, S. Chen, L. Liang, Y. Luo, J. Yu, and J. Lu, “DSM: A deep supervised multi-scale network learning for skin cancer segmentation,” IEEE Access, vol. 7, pp. 140936–140945, 2019.
- A. Dosovitskiy et al., “An image is worth 16x16 words: Transformers for image recognition at scale,” in Proc. Int. Conf. Learn. Representations (ICLR), 2021. arXiv:2010.11929.
- J. Valanarasu, R. Sindagi, V. Hacihaliloglu, and V. Patel, “Medical Transformer: Gated Axial-Attention for Medical Image Segmentation,” in Proc. Int. Conf. on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2021, pp. 36–46.
- Z. Liu, Y. Lin, Y. Cao, H. Hu, Y. Wei, Z. Zhang, S. Lin, and B. Guo, “Swin Transformer: Hierarchical Vision Transformer using Shifted Windows,” in Proc. IEEE Int. Conf. on Computer Vision (ICCV), 2021, pp. 10012–10022.
- H. Zhao, J. Shi, X. Qi, X. Wang, and J. Jia, “Pyramid scene parsing network,” in Proc. IEEE Conf. Comput. Vis. Pattern Recognit. (CVPR), Jul. 2017, pp. 2881–2890.
- O. Oktay, J. Schlemper, L. L. Folgoc, M. Lee, M. Heinrich, K. Misawa, et al., “Attention U-Net: Learning where to look for the pancreas,” 2018, arXiv preprint arXiv:1804.03999. [Online]. Available: https://arxiv.org/abs/1804.03999.
- S. Qamar, S. F. Qadri, R. Alroobaea, G. M. Alshmrani, and R. Jiang, “ScaleFusionNet: Transformer-guided multi-scale feature fusion for skin lesion segmentation,” Sci. Rep., vol. 15, Art. 34393, 2025. [CrossRef]
- W. Huang, X. Cai, Y. Yan, and Y. Kang, “MA-DenseUNet: A skin lesion segmentation method based on multi-scale attention and bidirectional LSTM,” Appl. Sci., vol. 15, no. 12, Art. 6538, Jun. 2025. [CrossRef]
- Y. Yin, J. Li, F. Zhang, et al., “Skin lesion segmentation with a multiscale input fusion U-Net incorporating Res2-SE and pyramid dilated convolution,” Sci. Rep., vol. 15, Art. 92447, 2025. [CrossRef]
- D. Li, X. Wu, and Q. Wei, “MSDTCN-Net: A multi-scale dual-encoder network for skin lesion segmentation,” Diagnostics, vol. 15, no. 22, Art. 2924, Nov. 2025. [CrossRef]
- S. Shahin, X. Zhao, Y. … et al., “CTH-Net: A CNN and Transformer hybrid network for skin lesion segmentation,” iScience, vol. 27, Art. 11042008, 2024. [CrossRef]
- Y. Li, T. Tian, J. Hu, and C. Yuan, “SUTrans-NET: A hybrid transformer approach to skin lesion segmentation,” PeerJ Comput. Sci., vol. 10, p. e1935, 2024. [CrossRef]
- M. Li, Y. Jiang, G. Cao, T. Xu, and R. Guo, “HyperFusionNet combines vision transformer for early melanoma detection and precise lesion segmentation,” Sci. Rep., vol. 15, Art. 30184, Nov. 2025. [CrossRef]
- S. Perera, Y. Erzurumlu, D. Gulati, and A. Yilmaz, “MobileUNETR: A lightweight end-to-end hybrid vision transformer for efficient medical image segmentation,” in Proc. ECCV Workshops, 2024.
| Methods | ACC | IoU | DC | SEN |
|---|---|---|---|---|
| Baseline | 0.959 | 0.869 | 0.926 | 0.912 |
| PVT v2 backbone | 0.963 | 0.870 | 0.931 | 0.915 |
| PVT v2 backbone+SAFE | 0.966 | 0.872 | 0.936 | 0.918 |
| PVT v2 backbone+MDCF | 0.968 | 0.869 | 0.938 | 0.920 |
| PVT v2 backbone+SAFE+MDCF | 0.970 | 0.873 | 0.941 | 0.921 |
| Methods | ACC | IoU | DC | SEN |
|---|---|---|---|---|
| U-Net [10] | 0.902 | 0.803 | 0.882 | 0.886 |
| Swin-Unet [22] | 0.923 | 0.821 | 0.909 | 0.899 |
| TransUNet [12] | 0.932 | 0.833 | 0.913 | 0.901 |
| H2Former [26] | 0.956 | 0.852 | 0.918 | 0.906 |
| SUnet [27] | 0.954 | 0.866 | 0.922 | 0.909 |
| MSDUnet [28] | 0.959 | 0.869 | 0.926 | 0.912 |
| Ours | 0.970 | 0.873 | 0.941 | 0.921 |
| Methods | ACC | IOU | DC | SEN |
|---|---|---|---|---|
| U-Net [10] | 0.921 | 0.806 | 0.858 | 0.899 |
| Swin-Unet [22] | 0.933 | 0.815 | 0.862 | 0.906 |
| TransUNet [12] | 0.946 | 0.821 | 0.880 | 0.912 |
| H2Former [26] | 0.959 | 0.825 | 0.877 | 0.910 |
| SUnet [27] | 0.958 | 0.829 | 0.887 | 0.916 |
| MSDUnet [28] | 0.962 | 0.833 | 0.897 | 0.921 |
| Ours | 0.970 | 0.836 | 0.902 | 0.928 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




