Submitted:
26 February 2026
Posted:
02 March 2026
You are already at the latest version
Abstract
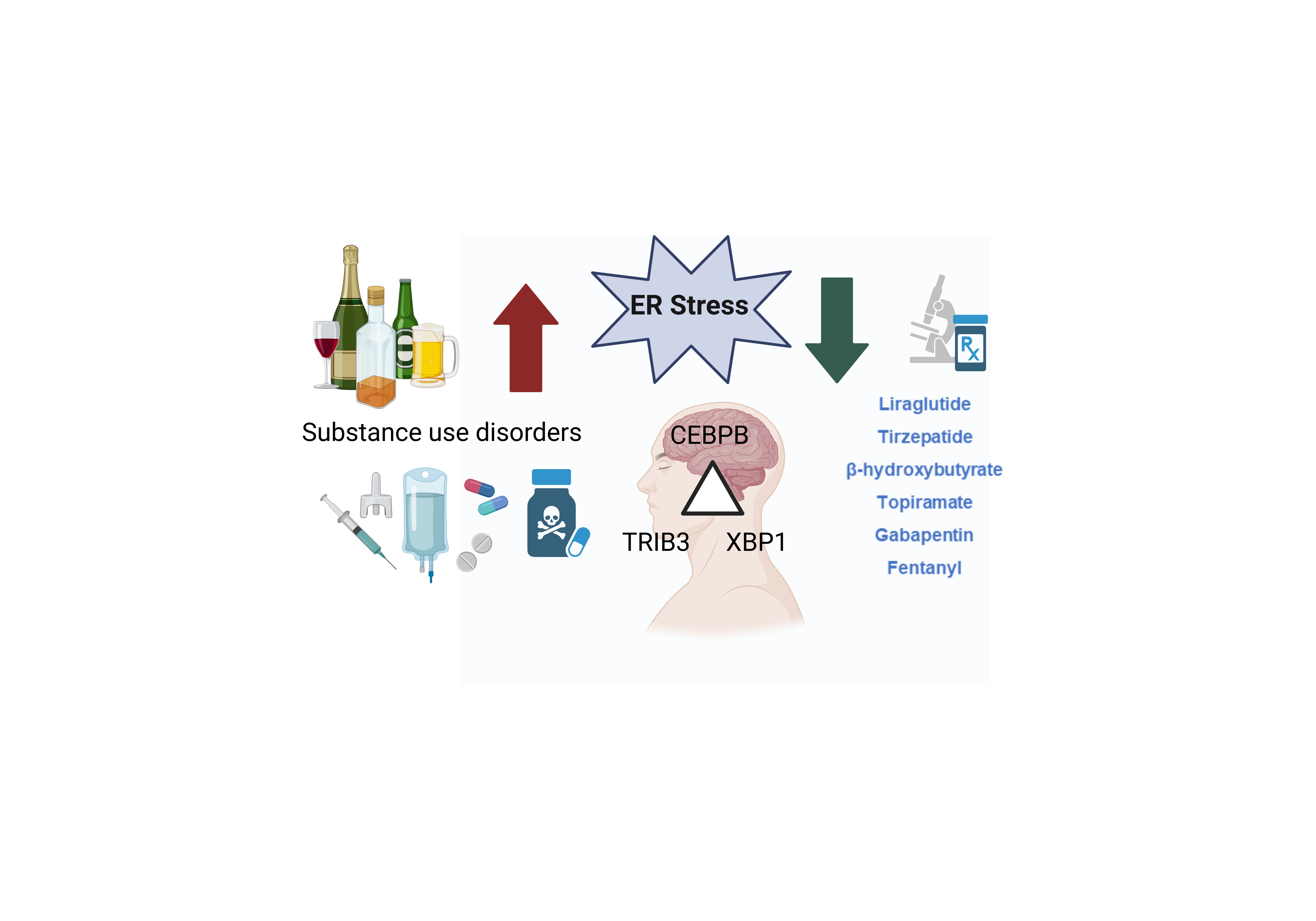
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Subjects and Ethics Statements
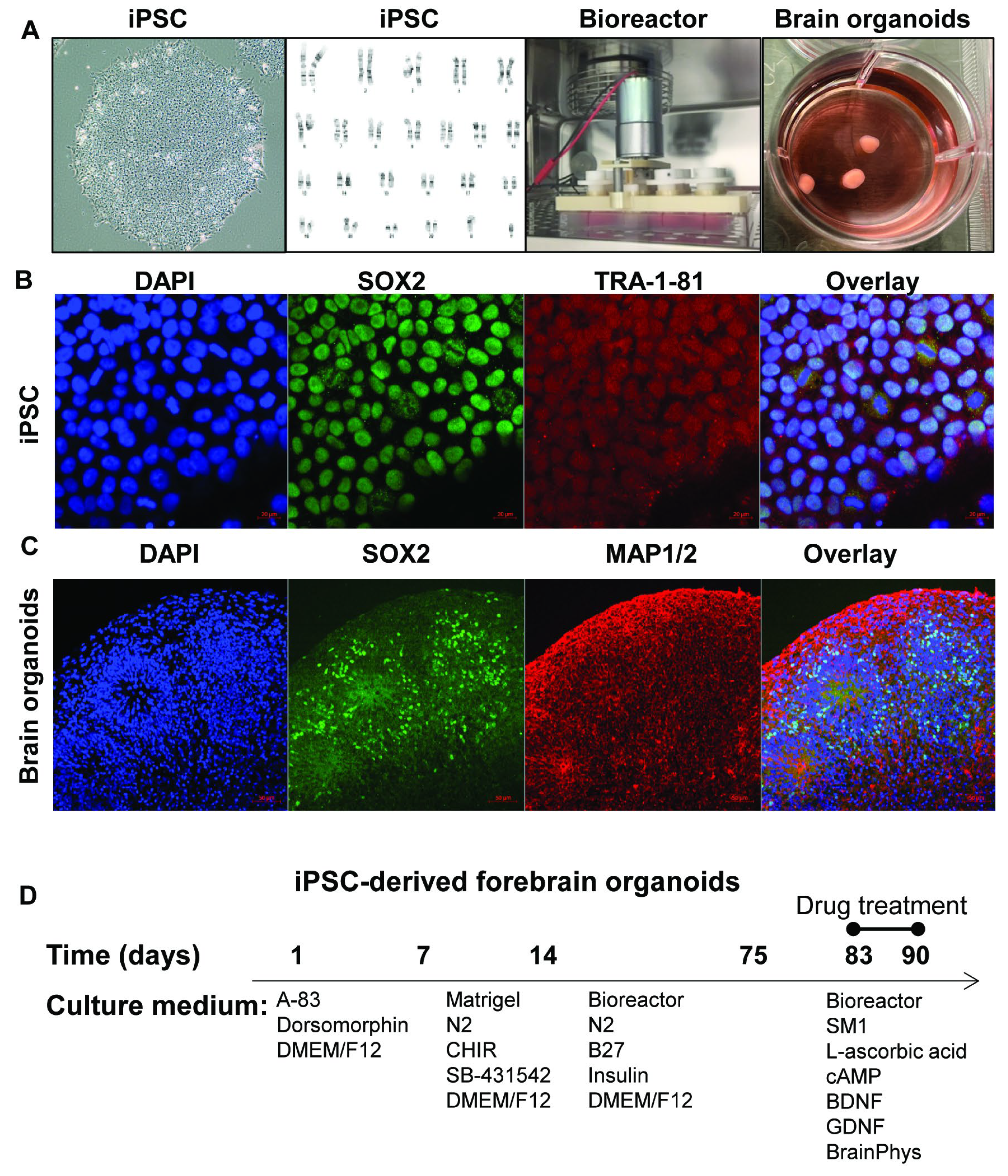
2.2. Generation of iPSCs, iPSC-Derived Forebrain Organoids and Neurons
2.3. Drug Treatment
2.4. Immunostaining and Confocal Microscopy
2.5. RNA Sequencing (RNA-seq) and Data Analysis
2.6. Co-Immunoprecipitation (co-IP) and Western Blot Analysis
2.7. CEBPB Chromatin Immunoprecipitation (ChIP) Assay
2.8. Statistical Analysis
2.9. Data Availability
3. Results
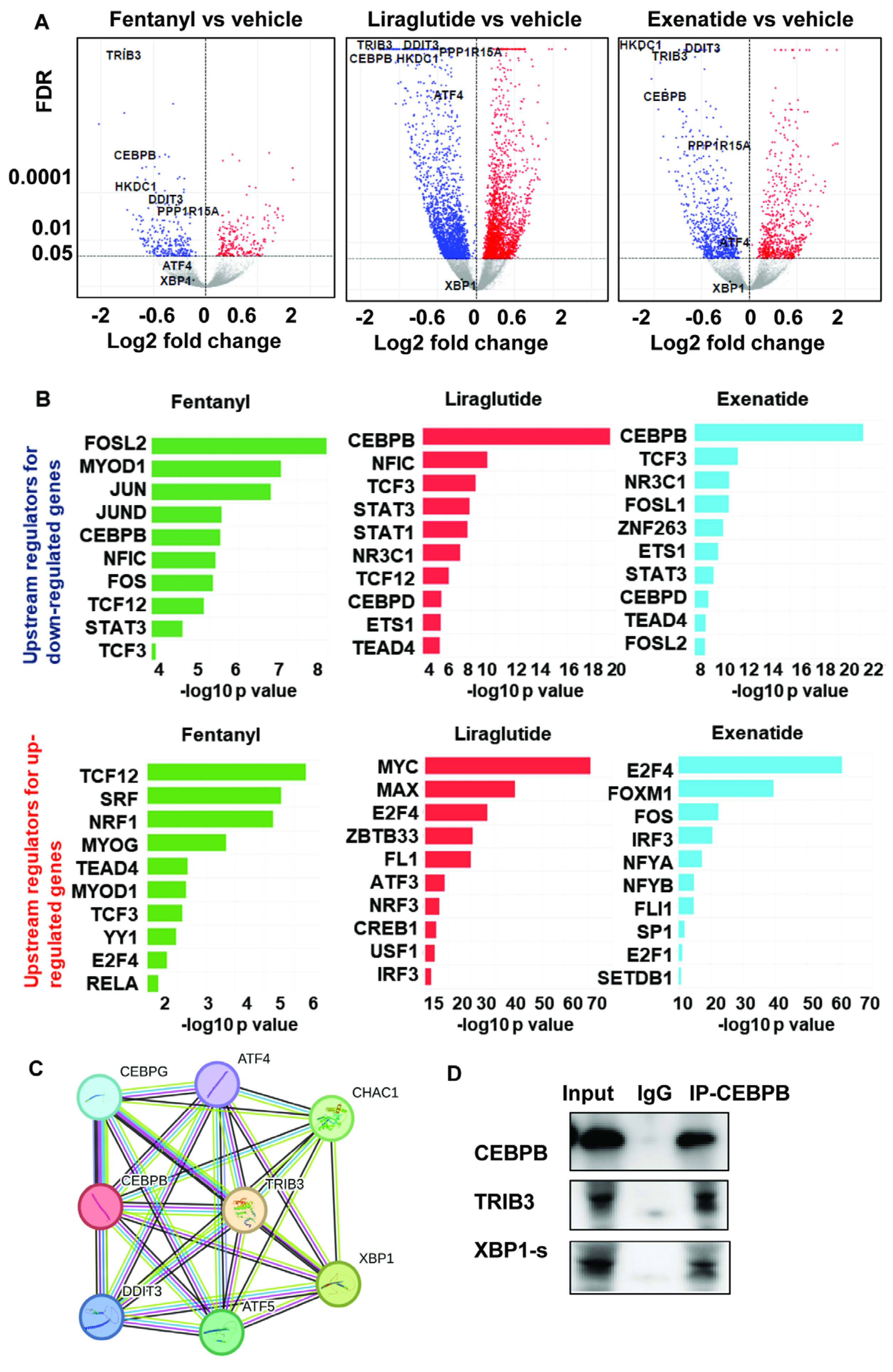
3.1. Transcriptomic Profiles in iPSC-Derived Forebrain Organoids
3.1.1. CEBPB as Upstream Regulator of ER Stress Genes
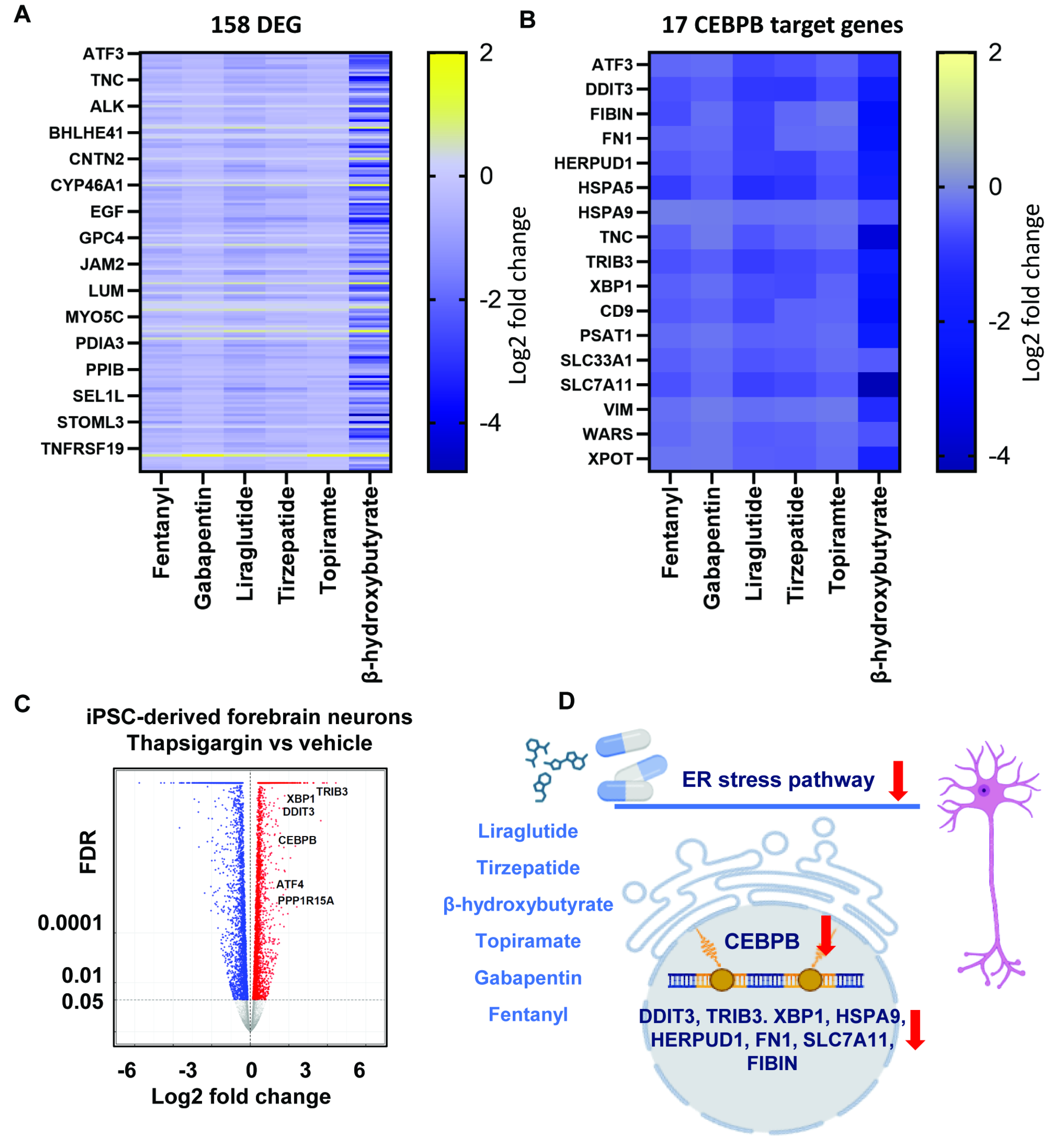
3.1.2. Transcriptomic Profiles in iPSC-Derived Forebrain Neurons
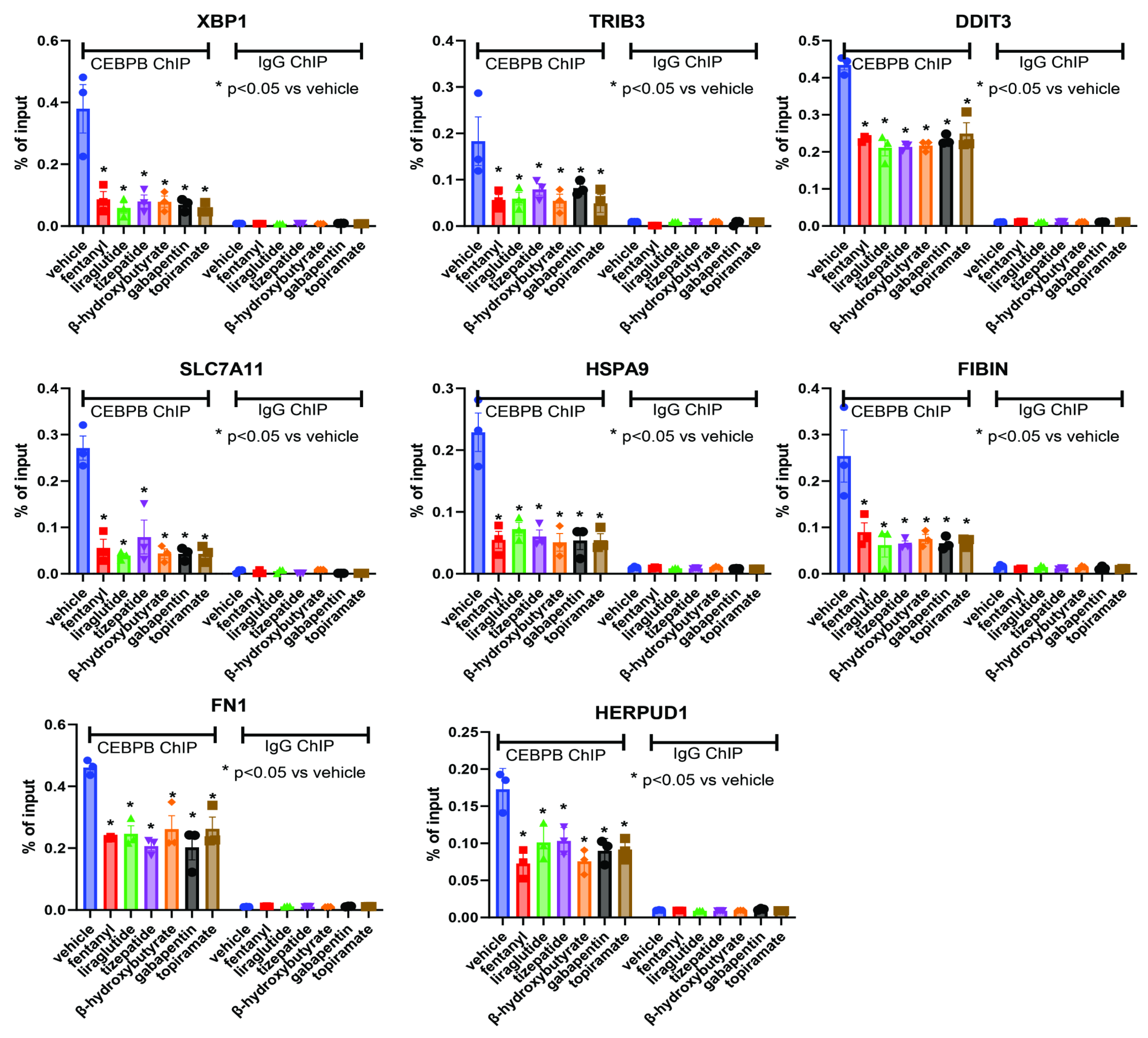
3.1.3. CEBPB Modulates ER Stress Related Genes by Decreasing Its DNA Binding
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AUD | alcohol use disorder (AUD) |
| CDC | Center for Disease Control (CDC) |
| ChIP | chromatin Immunoprecipitation (ChIP) |
| Co-IP | co-immunoprecipitation (co-IP) |
| DEGs | differentially expressed genes (DEGs) |
| EB | embryoid body (EB) |
| ER | endoplasmic reticulum (ER) |
| FDA | Food and Drug Administration (FDA) |
| GIP | glucose-dependent insulinotropic polypeptide (GIP) |
| GLP-1RA | GLP-1 receptor agonists |
| iPSC | Induced pluripotent stem cell |
| OUD | opioid use disorder (OUD) |
| SUD | substance use disorders (SUD) |
| TRIB3 | Tribbles Pseudokinase 3 |
| CEBPB | CCAAT/enhancer-binding protein beta |
| ATF4 | activating transcription factor 4 |
| DDIT3 | DNA damage inducible transcript 3 |
| XBP1 | X-box binding protein 1 |
| SOX2 | SRY-box transcription factor 2 |
| PPP1R15A | Protein phosphatase 1 regulatory subunit 15A |
| GADD34 | growth arrest and DNA damage-inducible protein |
| HKDC1 | hexokinase domain containing 1 |
| TRA-1-81 | podocalyxin |
| SLC7A11 | solute carrier family 7 member 11 |
| FIBIN | fin bud initiation factor homolog |
| FN1 | fibronectin 1 |
| HERPUD1 | homocysteine inducible ER protein with ubiquitin like domain 1 |
| ENCODE | The Encyclopedia of DNA Elements |
| T2D | type 2 diabetes (T2D) |
References
- Mattson, C.L.; Tanz, L.J.; Quinn, K.; Kariisa, M.; Patel, P.; Davis, N.L. Trends and Geographic Patterns in Drug and Synthetic Opioid Overdose Deaths - United States, 2013-2019. MMWR. Morbidity and mortality weekly report 2021, 70, 202–207. [Google Scholar] [CrossRef]
- Blanco, C.; Volkow, N.D. Management of opioid use disorder in the USA: Present status and future directions. The Lancet 2019, 393, 1760–1772. [Google Scholar] [CrossRef]
- Scherrer, J.F.; Tucker, J.; Salas, J.; Zhang, Z.; Grucza, R. Comparison of Opioids Prescribed for Patients at Risk for Opioid Misuse Before and After Publication of the Centers for Disease Control and Prevention’s Opioid Prescribing Guidelines. JAMA Network Open 2020, 3, e2027481. [Google Scholar] [CrossRef]
- Ling, W.; Nadipelli, V.R.; Aldridge, A.P.; Ronquest, N.A.; Solem, C.T.; Chilcoat, H.; Albright, V.; Johnson, C.; Learned, S.M.; Mehra, V.; et al. Recovery From Opioid Use Disorder (OUD) After Monthly Long-acting Buprenorphine Treatment: 12-Month Longitudinal Outcomes From RECOVER, an Observational Study. Journal of Addiction Medicine 2020, 14, e233–e240. [Google Scholar] [CrossRef] [PubMed]
- Kosten, T.R.; Petrakis, I.L. The Hidden Epidemic of Opioid Overdoses During the Coronavirus Disease 2019 Pandemic. JAMA Psychiatry 2021, 78, 585–586. [Google Scholar] [CrossRef]
- Oesterle, T.S.; Thusius, N.J.; Rummans, T.A.; Gold, M.S. Medication-Assisted Treatment for Opioid-Use Disorder. Mayo Clinic Proceedings 2019, 94, 2072–2086. [Google Scholar] [CrossRef] [PubMed]
- NIDA. Only 1 in 5 U.S. adults with opioid use disorder received medications to treat it in 2021. National Institute on Drug Abuse website. Available online: https://nida.nih.gov/news-events/news-releases/2023/08/only-1-in-5-us-adults-with-opioid-use-disorder-received-medications-to-treat-it-in-2021 (accessed on 16 October 2025).
- Feingold, K.R. Oral and Injectable (Non-Insulin) Pharmacological Agents for the Treatment of Type 2 Diabetes. In Endotext; Feingold, K.R., Ahmed, S.F., Anawalt, B., Blackman, M.R., Boyce, A., Chrousos, G., Corpas, E., de Herder, W.W., Dhatariya, K., Dungan, K., et al., Eds.; MDText.com, Inc. Copyright © 2000-2025, MDText.com, Inc.: South Dartmouth (MA), 2000. [Google Scholar]
- Wang, W.; Volkow, N.D.; Wang, Q.; Berger, N.A.; Davis, P.B.; Kaelber, D.C.; Xu, R. Semaglutide and Opioid Overdose Risk in Patients With Type 2 Diabetes and Opioid Use Disorder. JAMA Network Open 2024, 7, e2435247. [Google Scholar] [CrossRef] [PubMed]
- Qeadan, F.; McCunn, A.; Tingey, B. The association between glucose-dependent insulinotropic polypeptide and/or glucagon-like peptide-1 receptor agonist prescriptions and substance-related outcomes in patients with opioid and alcohol use disorders: A real-world data analysis. Addiction 2025, 120, 236–250. [Google Scholar] [CrossRef]
- Zhang, Y.; Kahng, M.W.; Elkind, J.A.; Weir, V.R.; Hernandez, N.S.; Stein, L.M.; Schmidt, H.D. Activation of GLP-1 receptors attenuates oxycodone taking and seeking without compromising the antinociceptive effects of oxycodone in rats. Neuropsychopharmacology 2020, 45, 451–461. [Google Scholar] [CrossRef]
- Ho, M.-F.; Zhang, C.; Zhang, L.; Wei, L.; Zhou, Y.; Moon, I.; Geske, J.R.; Choi, D.-S.; Biernacka, J.; Frye, M.; et al. TSPAN5 influences serotonin and kynurenine: Pharmacogenomic mechanisms related to alcohol use disorder and acamprosate treatment response. Molecular Psychiatry 2020, 26, 3122–3133. [Google Scholar] [CrossRef]
- Biernacka, J.M.; Coombes, B.J.; Batzler, A.; Ho, A.M.-C.; Geske, J.R.; Frank, J.; Hodgkinson, C.; Skime, M.; Colby, C.; Zillich, L.; et al. Genetic contributions to alcohol use disorder treatment outcomes: A genome-wide pharmacogenomics study. Neuropsychopharmacology 2021, 46, 2132–2139. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.-F.; Zhang, C.; Moon, I.; Zhu, X.; Coombes, B.J.; Biernacka, J.; Skime, M.; Oesterle, T.S.; Karpyak, V.M.; Schmidt, K.; et al. Single cell transcriptomics reveals distinct transcriptional responses to oxycodone and buprenorphine by iPSC-derived brain organoids from patients with opioid use disorder. Molecular Psychiatry 2022. [Google Scholar] [CrossRef]
- Qian, X.; Jacob, F.; Song, M.M.; Nguyen, H.N.; Song, H.; Ming, G.-l. Generation of human brain region–specific organoids using a miniaturized spinning bioreactor. Nature Protocols 2018, 13, 565. Available online: https://www.nature.com/articles/nprot.2017.152#supplementary-information. [CrossRef] [PubMed]
- Ho, M.-F.; Zhang, C.; Wei, L.; Zhang, L.; Moon, I.; Geske, J.R.; Skime, M.K.; Choi, D.-S.; Biernacka, J.M.; Oesterle, T.S.; et al. Genetic variants associated with acamprosate treatment response in alcohol use disorder patients: A multiple omics study. British journal of pharmacology 2022, 173, 16. [Google Scholar] [CrossRef]
- Ho, M.-F.; Zhang, L.; Moon, I.; Skime, M.; Ho, A.M.-C.; Choi, D.-S.; Biernacka, J.; Kaddurah-Daouk, R.; Wen, Z.; Frye, M.A.; et al. TSPAN5, an alcohol responsive gene that alters serotonin levels in human induced pluripotent stem cells: Novel molecular links to alcohol use disorder. Alcoholism: Clinical and Experimental Research 2019, 43, S786. [Google Scholar]
- Wen, Z.; Nguyen, H.N.; Guo, Z.; Lalli, M.A.; Wang, X.; Su, Y.; Kim, N.-S.; Yoon, K.-J.; Shin, J.; Zhang, C.; et al. Synaptic dysregulation in a human iPS cell model of mental disorders. Nature 2014, 515, 414–418. [Google Scholar] [CrossRef]
- Lötsch, J.; Walter, C.; Parnham, M.J.; Oertel, B.G.; Geisslinger, G. Pharmacokinetics of Non-Intravenous Formulations of Fentanyl. Clinical Pharmacokinetics 2013, 52, 23–36. [Google Scholar] [CrossRef]
- Jacobsen, L.V.; Flint, A.; Olsen, A.K.; Ingwersen, S.H. Liraglutide in Type 2 Diabetes Mellitus: Clinical Pharmacokinetics and Pharmacodynamics. Clin Pharmacokinet 2016, 55, 657–672. [Google Scholar] [CrossRef]
- Cirincione, B.; Mager, D.E. Population pharmacokinetics of exenatide. Br J Clin Pharmacol 2017, 83, 517–526. [Google Scholar] [CrossRef]
- Schneck, K.; Urva, S. Population pharmacokinetics of the GIP/GLP receptor agonist tirzepatide. CPT Pharmacometrics Syst Pharmacol 2024, 13, 494–503. [Google Scholar] [CrossRef]
- Bae, E.-K.; Lee, J.; Shin, J.-W.; Moon, J.; Lee, K.-J.; Shin, Y.-W.; Kim, T.-J.; Shin, D.; Jang, I.-J.; Lee, S.K. Factors influencing topiramate clearance in adult patients with epilepsy: A population pharmacokinetic analysis. Seizure 2016, 37, 8–12. [Google Scholar] [CrossRef] [PubMed]
- Swearingen, D.; Aronoff, G.M.; Ciric, S.; Lal, R. Pharmacokinetics of immediate release, extended release, and gastric retentive gabapentin formulations in healthy adults. Int J Clin Pharmacol Ther 2018, 56, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Shivva, V.; Cox, P.J.; Clarke, K.; Veech, R.L.; Tucker, I.G.; Duffull, S.B. The Population Pharmacokinetics of D-β-hydroxybutyrate Following Administration of (R)-3-Hydroxybutyl (R)-3-Hydroxybutyrate. Aaps j 2016, 18, 678–688. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics (Oxford, England) 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology 2014, 15, 550. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Mootha, V.K.; Lindgren, C.M.; Eriksson, K.-F.; Subramanian, A.; Sihag, S.; Lehar, J.; Puigserver, P.; Carlsson, E.; Ridderstråle, M.; Laurila, E.; et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nature genetics 2003, 34, 267. Available online: https://www.nature.com/articles/ng1180#supplementary-information. [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 2022, 50, W216–W221. [Google Scholar] [CrossRef]
- Keenan, A.B.; Torre, D.; Lachmann, A.; Leong, A.K.; Wojciechowicz, M.L.; Utti, V.; Jagodnik, K.M.; Kropiwnicki, E.; Wang, Z.; Ma’ayan, A. ChEA3: Transcription factor enrichment analysis by orthogonal omics integration. Nucleic Acids Research 2019, 47, W212–W224. [Google Scholar] [CrossRef]
- Fang, N.; Zhang, W.; Xu, S.; Lin, H.; Wang, Z.; Liu, H.; Fang, Q.; Li, C.; Peng, L.; Lou, J. TRIB3 alters endoplasmic reticulum stress-induced β-cell apoptosis via the NF-κB pathway. Metabolism 2014, 63, 822–830. [Google Scholar] [CrossRef]
- Pitale, P.M.; Saltykova, I.V.; Adu-Agyeiwaah, Y.; Calzi, S.L.; Satoh, T.; Akira, S.; Gorbatyuk, O.; Boulton, M.E.; Pardue, M.T.; Garvey, W.T.; et al. Tribbles Homolog 3 Mediates the Development and Progression of Diabetic Retinopathy. Diabetes 2021, 70, 1738–1753. [Google Scholar] [CrossRef] [PubMed]
- Ohoka, N.; Yoshii, S.; Hattori, T.; Onozaki, K.; Hayashi, H. TRB3, a novel ER stress-inducible gene, is induced via ATF4-CHOP pathway and is involved in cell death. Embo j 2005, 24, 1243–1255. [Google Scholar] [CrossRef] [PubMed]
- Örd, D.; Örd, T. Characterization of human NIPK (TRB3, SKIP3) gene activation in stressful conditions. Biochemical and Biophysical Research Communications 2005, 330, 210–218. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Franceschini, A.; Wyder, S.; Forslund, K.; Heller, D.; Huerta-Cepas, J.; Simonovic, M.; Roth, A.; Santos, A.; Tsafou, K.P.; et al. STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 2015, 43, D447–D452. [Google Scholar] [CrossRef]
- Quddos, F.; Hubshman, Z.; Tegge, A.; Sane, D.; Marti, E.; Kablinger, A.S.; Gatchalian, K.M.; Kelly, A.L.; DiFeliceantonio, A.G.; Bickel, W.K. Semaglutide and Tirzepatide reduce alcohol consumption in individuals with obesity. Scientific Reports 2023, 13, 20998. [Google Scholar] [CrossRef]
- Salehi, M.; Kheirabadi, G.R.; Maracy, M.R.; Ranjkesh, M. Importance of gabapentin dose in treatment of opioid withdrawal. J Clin Psychopharmacol 2011, 31, 593–596. [Google Scholar] [CrossRef]
- Mokhber, N.; Soltanifar, A.; Talebi, M. The Effect of Topiramate in the Treatment of Opioid (Heroin) Withdrawal. European Psychiatry 2009, 24, E424–E443. [Google Scholar] [CrossRef]
- Kong, D.; Sun, J.X.; Yang, J.Q.; Li, Y.S.; Bi, K.; Zhang, Z.Y.; Wang, K.H.; Luo, H.Y.; Zhu, M.; Xu, Y. Ketogenic diet: A potential adjunctive treatment for substance use disorders. Front Nutr 2023, 10, 1191903. [Google Scholar] [CrossRef]
- Trinko, R.; Diaz, D.M.; Foscue, E.; Thompson, S.L.; Taylor, J.R.; DiLeone, R.J. Ketogenic diet enhances the effects of oxycodone in mice. Scientific Reports 2023, 13, 7507. [Google Scholar] [CrossRef]
- Wiers, C.E.; Vendruscolo, L.F.; van der Veen, J.-W.; Manza, P.; Shokri-Kojori, E.; Kroll, D.S.; Feldman, D.E.; McPherson, K.L.; Biesecker, C.L.; Zhang, R.; et al. Ketogenic diet reduces alcohol withdrawal symptoms in humans and alcohol intake in rodents. Science Advances 2021, 7, eabf6780. [Google Scholar] [CrossRef] [PubMed]
- Williamson, J.; Kermanizadeh, A. A Review of Toxicological Profile of Fentanyl—A 2024 Update. Toxics 2024, 12, 690. [Google Scholar] [CrossRef] [PubMed]
- van der Krieken, S.E.; Popeijus, H.E.; Mensink, R.P.; Plat, J. CCAAT/Enhancer Binding Protein β in relation to ER Stress, Inflammation, and Metabolic Disturbances. BioMed Research International 2015, 2015, 324815. [Google Scholar] [CrossRef] [PubMed]
- Yao, Q.; Long, C.; Yi, P.; Zhang, G.; Wan, W.; Rao, X.; Ying, J.; Liang, W.; Hua, F. C/EBPβ: A transcription factor associated with the irreversible progression of Alzheimer's disease. CNS Neurosci Ther 2024, 30, e14721. [Google Scholar] [CrossRef]
- Griffin, H.; Sullivan, S.C.; Barger, S.W.; Phelan, K.D.; Baldini, G. Liraglutide Counteracts Endoplasmic Reticulum Stress in Palmitate-Treated Hypothalamic Neurons without Restoring Mitochondrial Homeostasis. International Journal of Molecular Sciences 2023, 24, 629. [Google Scholar] [CrossRef]
- Yang, B.; Zhang, R.; Leong Bin Abdullah, M.F.I. The association between neuropsychiatric effects of substance use and occurrence of endoplasmic reticulum and unfolded protein response: A systematic review. Toxicol Lett 2024, 391, 71–85. [Google Scholar] [CrossRef]
- Sprenkle, N.T.; Sims, S.G.; Sánchez, C.L.; Meares, G.P. Endoplasmic reticulum stress and inflammation in the central nervous system. Mol Neurodegener 2017, 12, 42. [Google Scholar] [CrossRef]
- Wang, W.; Volkow, N.D.; Berger, N.A.; Davis, P.B.; Kaelber, D.C.; Xu, R. Association of semaglutide with risk of suicidal ideation in a real-world cohort. Nat Med 2024, 30, 168–176. [Google Scholar] [CrossRef]
- Wium-Andersen, I.K.; Wium-Andersen, M.K.; Fink-Jensen, A.; Rungby, J.; Jørgensen, M.B.; Osler, M. Use of GLP-1 receptor agonists and subsequent risk of alcohol-related events. A nationwide register-based cohort and self-controlled case series study. Basic Clin Pharmacol Toxicol 2022, 131, 372–379. [Google Scholar] [CrossRef]
- Goldstein, R.Z.; Volkow, N.D. Drug Addiction and Its Underlying Neurobiological Basis: Neuroimaging Evidence for the Involvement of the Frontal Cortex. American Journal of Psychiatry 2002, 159, 1642–1652. [Google Scholar] [CrossRef]
- Ceceli, A.O.; Bradberry, C.W.; Goldstein, R.Z. The neurobiology of drug addiction: Cross-species insights into the dysfunction and recovery of the prefrontal cortex. Neuropsychopharmacology 2022, 47, 276–291. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



