Submitted:
15 February 2026
Posted:
28 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Background
1.2. Related Works
1.3. Main Contributions
2. Mathematical Foundations
2.1. Variational Formulation of the Inverse Problem
2.2. Physical Isomorphism: Reaction–Diffusion Kinetics
3. The Screened Poisson Normalization (SPN) Model
4. Theoretical Analysis
5. Optimization and Implementation
5.1. Discrete and Non-Overlapping Domain Decomposition
5.2. Spectral Diagonalization and DCT Solver
Stability and Regularization.
5.3. Tiled SPN Implementation for Gigapixel WSIs
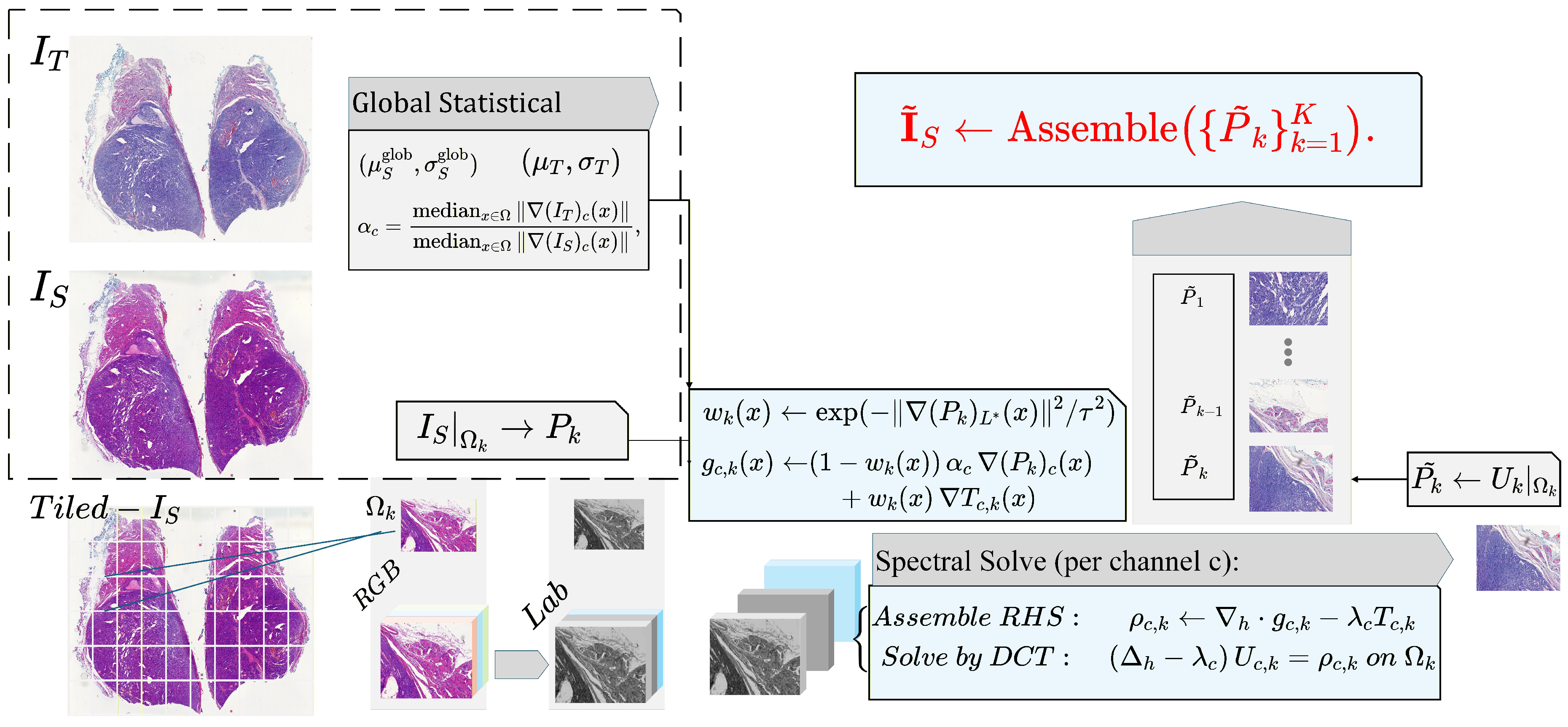
| Algorithm 1 Tiled Screened Poisson Normalization (SPN) |
|
5.4. Computational Complexity
- Assembly: Computing gradients, weights, and the right-hand side scales linearly, i.e., .
- Spectral Solve: The 2D DCT is implemented via separable 1D fast cosine transforms, with a cost of .
6. Experiments and Results
6.1. Experimental Setup
Datasets
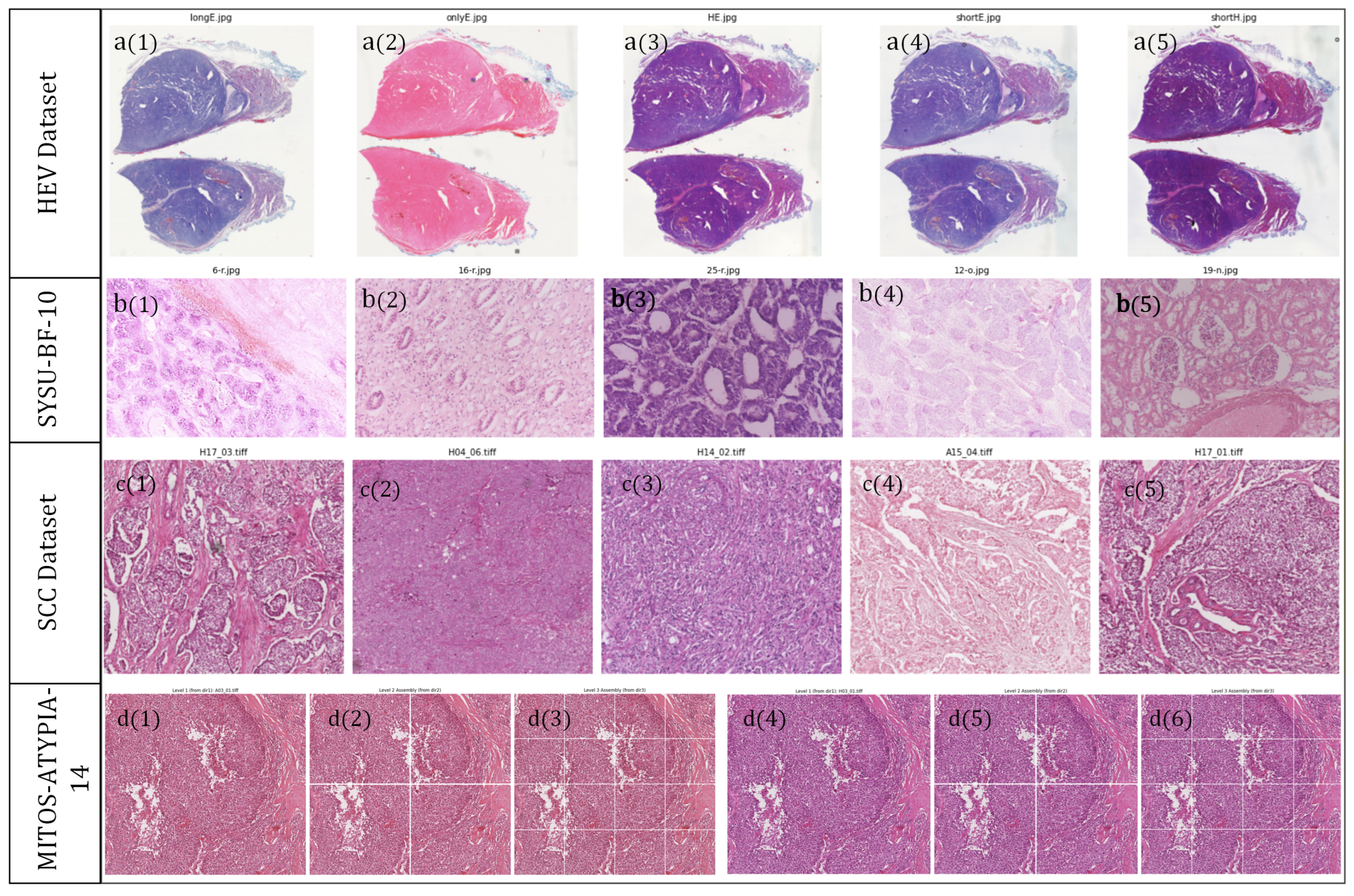
- Dataset 1: MITOS-ATYPIA-14 [35], released as part of the MITOS-ATYPIA Grand Challenge, was annotated by the Pathology Department of Pitié-Salpêtrière Hospital in Paris. It consists of 14 pairs of breast carcinoma slides, each scanned using two distinct digital pathology scanners—Aperio ScanScope XT and Hamamatsu NanoZoomer—allowing controlled analysis of inter-scanner color variation.
- Dataset 2: Multi-Scanner Squamous Cell Carcinoma (SCC) dataset [36] contains 44 samples of canine cutaneous squamous cell carcinoma digitized using 5 different scanning systems, producing a total of 220 WSIs.
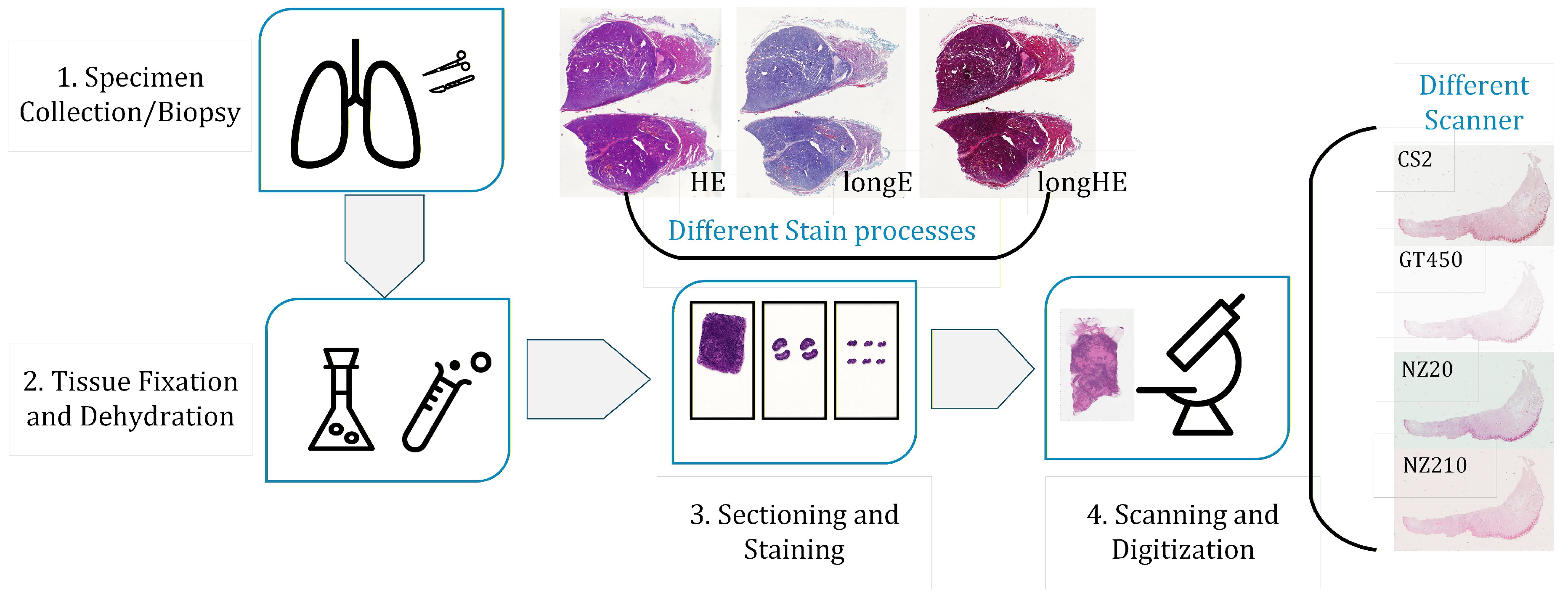
- Dataset 3: HE-Staining Variation (HEV) dataset [37] provided by Heidelberg University. The HEV dataset comprises follicular thyroid carcinoma slides prepared under nine distinct staining protocols: standard H&E, over-stained H&E (longHE), under-stained H&E (shortHE), hematoxylin-only (onlyH), eosin-only (onlyE), over-stained hematoxylin (longH), over-stained eosin (longE), under-stained hematoxylin (shortH), and under-stained eosin (shortE). These variations facilitate the evaluation of color normalization under chemically induced staining inconsistencies.
- Dataset 4: SUSY-BF-10 clinical dataset (local collection). To address the temporal color degradation that public datasets do not capture, we constructed a clinical dataset from Sun Yat-sen Memorial Hospital, Sun Yat-sen University. The SUSY-BF-10 dataset includes 25 long-term archived breast carcinoma slides, each preserved for over ten years and rescanned to analyze fading and chromatic shifts over time.
Implementation Details
- 1.
- Full-resolution WSI normalization. We evaluate the behavior of each method when applied directly to entire, full-size histopathology slides.
- 2.
- Tile-based reconstruction. We further consider the practically common pipeline in which a large WSI is first partitioned into tiles, normalized tile-wise, and then reassembled into a full-resolution slide. This setting is particularly relevant for memory-constrained or streaming-based deployments.
Baselines
- Reinhard. The classical global Reinhard color transfer algorithm [6] serves as our primary reference for global-statistics–based normalization. It matches low-order statistics between source and target images in a decorrelated color space and remains widely used in digital pathology.
- GCTI. We include a geometry-aware method with local alignment in the stain vector space (GCTI) [9], which explicitly exploits the underlying geometric structure of color distributions to compensate for scanner- and protocol-induced variations in microscopy images within a unified framework.
- CycleGAN. As a deep learning–based baseline, we employ a CycleGAN-style stain transfer model [10]. This method learns a non-linear mapping between source and target stain domains from unpaired data and represents a strong data-driven alternative to hand-crafted normalization schemes.
6.2. Evaluation Metrics
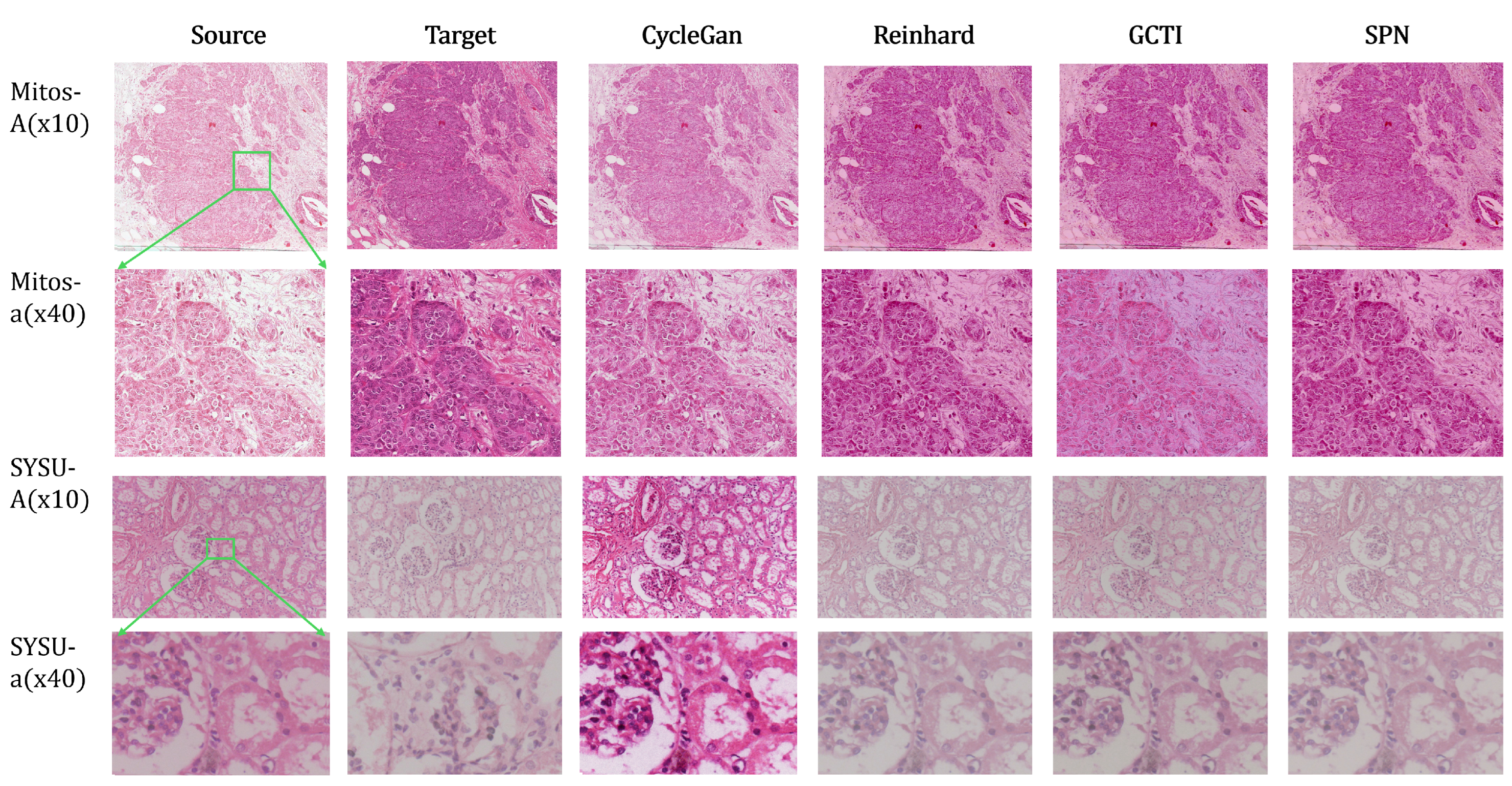
6.3. Results and Visual Analysis
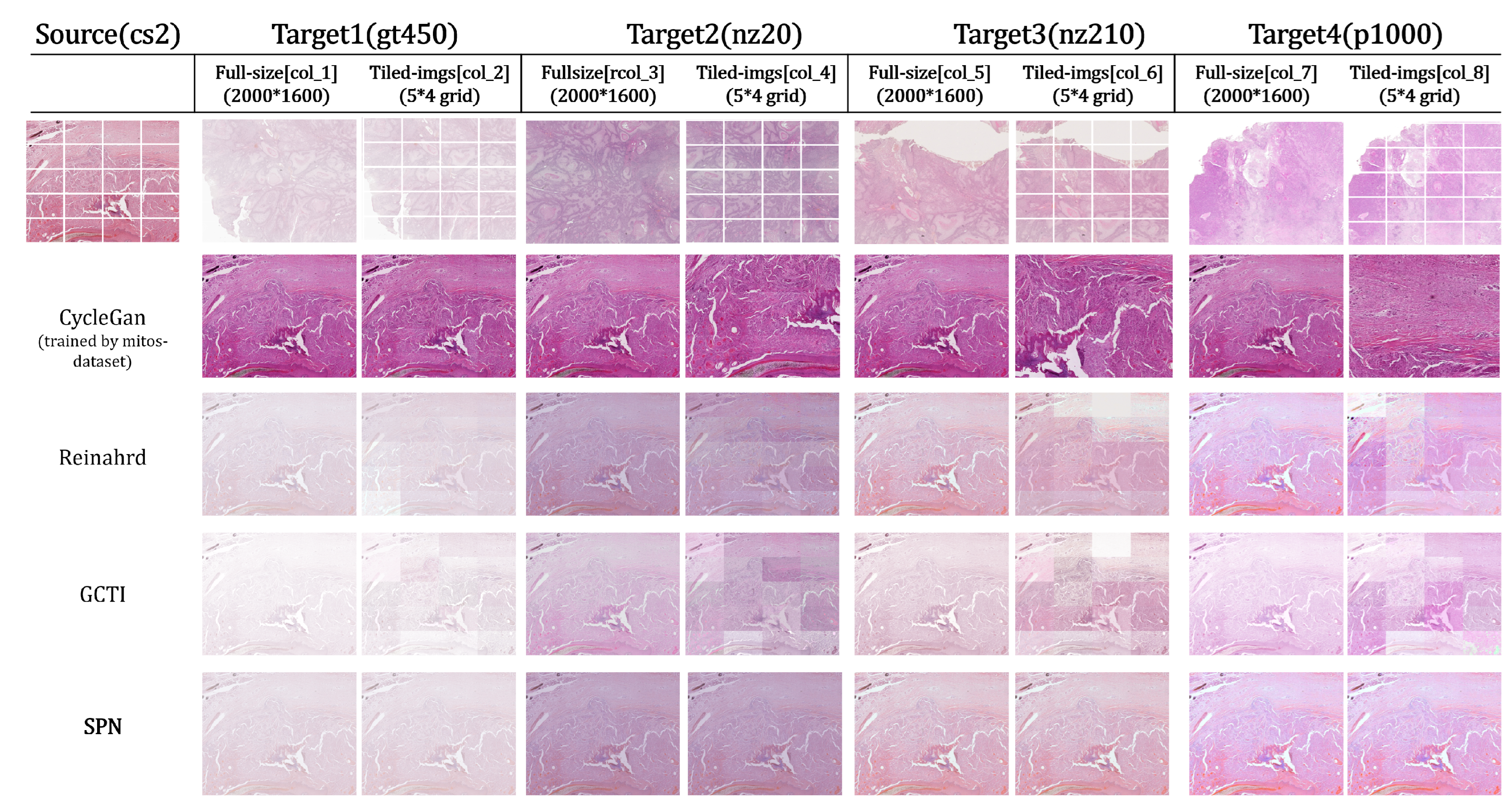
Full size baseline test
SCC dataset: full-size vs. tiled normalization.
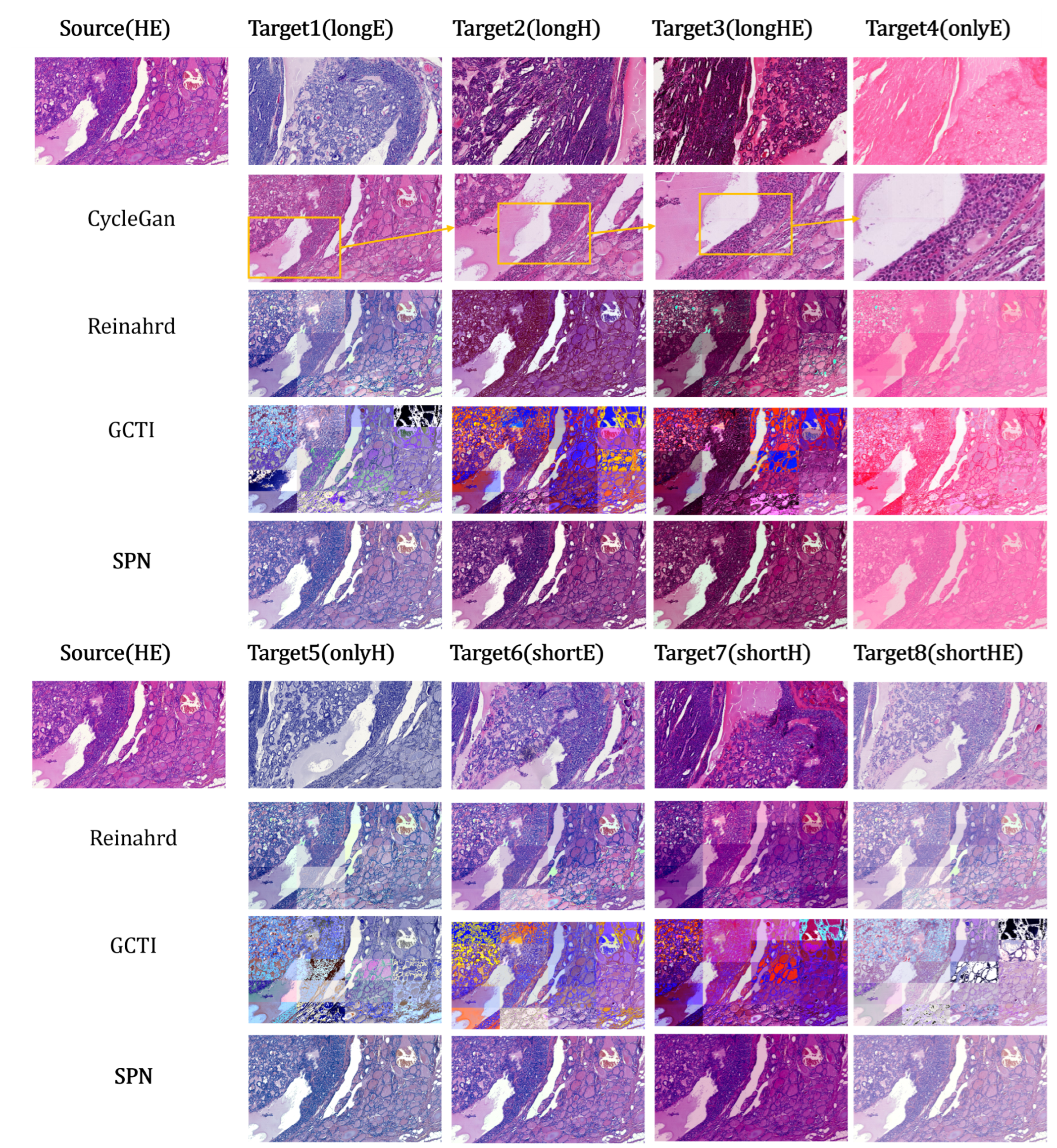
HEV dataset: Multi-stained pathology images.
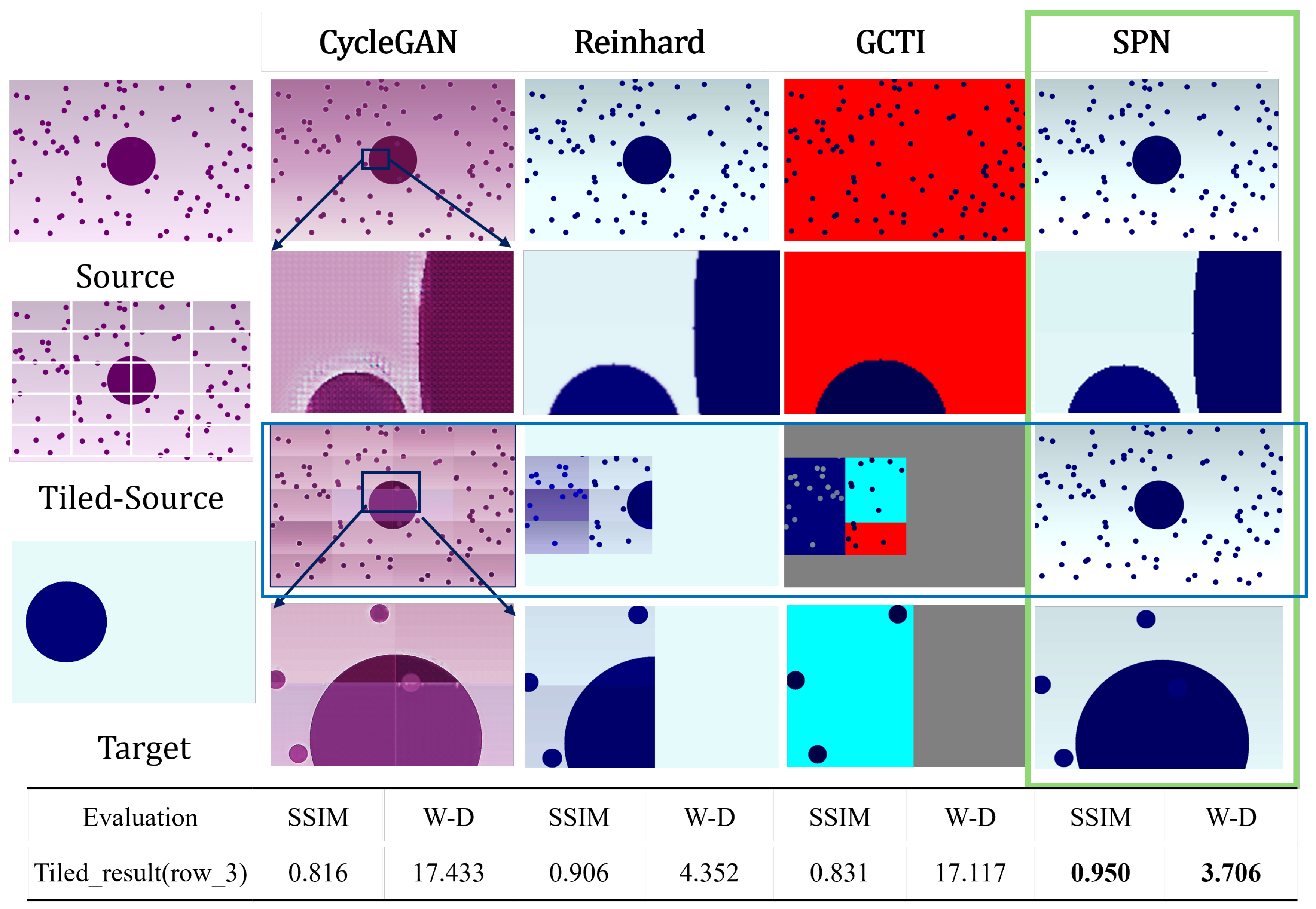
Evaluation on Synthetic Image
6.4. Quantitative Results and Discussion
7. Conclusions
- 1.
- Structural Fidelity and Scalability: Operating in the gradient domain allows SPN to preserve diagnostic morphological details (e.g., nuclear boundaries) often degraded by pure statistical alignment. Furthermore, the exponential decay of the screened Green’s function enables rigorous non-overlapping domain decomposition, permitting seamless gigapixel processing without tiling artifacts.
- 2.
- Determinism vs. Data-Dependence: In contrast to deep learning approaches (e.g., CycleGAN) which are prone to domain bias and generative hallucinations on out-of-distribution data, SPN is training-free and deterministic. This ensures that all output structures are causally linked to the input, providing the explainability and reproducibility essential for clinical diagnostics.
- 3.
- Robustness to Heterogeneity: SPN demonstrates consistent performance across diverse scanners, staining protocols, and archived slides. The global anchoring (screening) term effectively stabilizes the solution in tissue-sparse or background-dominated regions, overcoming the numerical brittleness observed in local geometry-based methods.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xu, C.; Sun, Y.; Zhang, Y.; Liu, T.; Wang, X.; Hu, D.; Huang, S.; Li, J.; Zhang, F.; Li, G. Stain Normalization of Histopathological Images Based on Deep Learning: A Review. Diagnostics 2025, 15(8), 1032. [Google Scholar] [CrossRef]
- Hetz, M.J.; Bucher, T.C.; Brinker, T.J. Multi-domain stain normalization for digital pathology: A cycle-consistent adversarial network for whole slide images. Medical Image Analysis 2024, 94, 103149. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, S.; Ke, J.; Zhang, C.; Shen, Y. RandStainNA++: Enhance random stain augmentation and normalization through foreground and background differentiation. IEEE Journal of Biomedical and Health Informatics 2024, 28, 3660–3671. [Google Scholar] [CrossRef] [PubMed]
- Baykal Kablan, E. Regional realness-aware generative adversarial networks for stain normalization. Neural Computation and Applications 2023, 35, 17915–17927. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, H.; Liu, N.; Shen, Y. (2023). Stain normalization and augmentation in frequency space for histology analysis. In 2023 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2031–2035.
- Reinhard, E.; Ashikhmin, M.; Gooch, B.; Shirley, P. Color transfer between images. IEEE Computer Graphics and Applications 2001, 21(5), 34–41. [Google Scholar] [CrossRef]
- Macenko, M.; Niethammer, M.; Marron, J.S.; Borland, D.; Woosley, J.T.; Guan, X.; Schmitt, C.; Thomas, N.E. (2009). A method for normalizing histology slides for quantitative analysis. In 2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro, 1107–1110.
- Vahadane, A.; et al. Structure-preserving color normalization and sparse stain separation. IEEE Transactions on Medical Imaging 2016, 35(8), 1962–1971. [Google Scholar] [CrossRef]
- Gupta, A.; Duggal, R.; Gehlot, S.; Gupta, R.; Mangal, A.; Kumar, L.; Thakkar, N.; Satpathy, D. GCTI-SN: Geometry-inspired chemical and tissue invariant stain normalization of microscopic medical images. Medical Image Analysis 2020, 65, 101788. [Google Scholar] [CrossRef]
- Runz, M.; Rusche, D.; Schmidt, S.; et al. Normalization of H&E-stained histological images using cycle consistent generative adversarial networks. Diagnostic Pathology 2021, 16, 71. [Google Scholar]
- Kang, H.; Luo, D.; Feng, W.; et al. StainNet: A fast and robust stain normalization network. Frontiers in Medicine 2021, 8, 746307. [Google Scholar] [CrossRef]
- Shaban, M.T.; Baur, C.; Navab, N.; Albarqouni, S. (2019). Staingan: Stain style transfer for digital histological images. In 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), 953–956.
- BenTaieb, A.; Hamarneh, G. Adversarial stain transfer for histopathology image analysis. IEEE Transactions on Medical Imaging 2017, 37(3), 792–802. [Google Scholar] [CrossRef]
- Zarella, M.D.; Yeoh, C.; Breen, D.E.; Garcia, F.U. An alternative reference space for H&E color normalization. PLoS One 2017, 12(3), e0174489. [Google Scholar]
- Magee, D.; Treanor, D.; Crellin, D.; Shires, M.; Smith, K.; Mohee, K.; Quirke, P. (2009). Colour normalisation in digital histopathology images. Proceedings of Optical Tissue Image Analysis in Microscopy, Histopathology and Endoscopy (MICCAI Workshop), 100–111.
- Alsubaie, N.; Trahearn, N.; Raza, S.E.A.; Snead, D.; Rajpoot, N.M. Stain deconvolution using statistical analysis of multi-resolution stain colour representation. PLoS One 2017, 12(1), e0169875. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Lal, S.; Kini, J.R. Novel color normalization method for Hematoxylin & Eosin stained histopathology images. IEEE Access 2019, 7, 28982–28998. [Google Scholar] [CrossRef]
- Vijh, S.; Saraswat, M.; Kumar, S. A new complete color normalization method for H&E stained histopathological images. Applied Intelligence 2021, 51, 1–14. [Google Scholar]
- Zheng, Y.; Jiang, Z.; Zhang, H.; Xie, F.; Shi, J.; Xue, C. Adaptive color deconvolution for histological WSI normalization. Computer Methods and Programs in Biomedicine 2019, 170, 107–120. [Google Scholar] [CrossRef]
- Nadeem, S.; Hollmann, T.; Tannenbaum, A. (2020). Multimarginal Wasserstein barycenter for stain normalization and augmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 362–371.
- Shrivastava, A.; Adorno, W.; Sharma, Y.; et al. (2021). Self-attentive adversarial stain normalization. In Pattern Recognition. ICPR Workshops and Challenges, 120–140. Springer.
- Nishar, H.; Chavanke, N.; Singhal, N. (2020). Histopathological stain transfer using style transfer network with adversarial loss. In Medical Image Computing and Computer Assisted Intervention (MICCAI), 330–340.
- Liang, H.; Plataniotis, K.N.; Li, X. (2020). Stain style transfer of histopathology images via structure-preserved generative learning. In Machine Learning for Medical Image Reconstruction, 153–162. Springer.
- Kausar, T.; Kausar, A.; Ashraf, M.A.; et al. SA-GAN: Stain acclimation generative adversarial network for histopathology image analysis. Applied Sciences 2021, 12, 288. [Google Scholar] [CrossRef]
- Cho, H.; Lim, S.; Choi, G.; Min, H. Neural stain-style transfer learning using GAN for histopathological images. arXiv 2017, arXiv:1710.08543. [Google Scholar] [CrossRef]
- Lee, C.C.; Kuo, P.T.P.; Peng, C.H. (2022). H&E stain normalization using U-net. In 2022 IEEE 22nd International Conference on Bioinformatics and Bioengineering (BIBE), 29–32.
- Menning, J.D.M.; Wallmersperger, T.; Meinhardt, M.; Ehrenhofer, A. Modeling and simulation of diffusion and reaction processes during the staining of tissue sections on slides. Histochemistry and Cell Biology 2022, 158, 137–148. [Google Scholar] [CrossRef]
- Gurina, T.S.; Simms, L. Histology, Staining. In StatPearls [Internet]; StatPearls Publishing, 2023. [Google Scholar]
- Suvarna, K.S.; Layton, C.; Bancroft, J.D. (Eds.) Bancroft’s Theory and Practice of Histological Techniques, 8th ed.; Elsevier, 2019. [Google Scholar]
- Mayerhöfer, T.G.; Pahlow, S.; Popp, J. The Bouguer-Beer-Lambert law: Shining light on the obscure. ChemPhysChem 2020, 21(18), 2029–2046. [Google Scholar] [CrossRef]
- Murray, J.D. Mathematical Biology I: An Introduction, 3rd ed.; Springer: New York, NY, USA, 2002. [Google Scholar]
- Sheehan, D.C.; Hrapchak, B.B. Theory and Practice of Histotechnology, 3rd ed.; Mosby, 2012. [Google Scholar]
- Hetz, M.J.; Bucher, T.C.; Brinker, T.J. Multi-domain stain normalization for digital pathology: A cycle-consistent adversarial network for whole slide images. Medical Image Analysis 2024, 94, 103149. [Google Scholar] [CrossRef]
- Landini, G. Colour deconvolution with ImageJ and Fiji: a bibliography. Bioinformatics 2021, 37, 4181–4182. [Google Scholar]
- Roux, L.; Racoceanu, D.; Loménie, N.; et al. Mitosis detection in breast cancer histological images: An ICPR 2012 contest. Journal of Pathology Informatics 2013, 4, 8. [Google Scholar] [CrossRef]
- Wilm, F.; Fragoso, M.; Bertram, C.A.; Stathonikos, N.; Öttl, M.; Qiu, J.; Klopfleisch, R.; Maier, A.; Breininger, K.; Aubreville, M. Multi-scanner canine cutaneous squamous cell carcinoma histopathology dataset. arXiv 2023, arXiv:2301.04423. [Google Scholar] [CrossRef]
- Runz, M.; Weis, C.-A. (2021). Normalization of HE-Stained Histological Images using Cycle Consistent Generative Adversarial Networks [Dataset]. heiDATA, V1. [CrossRef]
- Pérez, P.; Gangnet, M.; Blake, A. (2023). Poisson image editing. In Seminal Graphics Papers: Pushing the Boundaries, Volume 2 (pp. 577–582).
- Habring, A.; Holler, M. Neural-network-based regularization methods for inverse problems in imaging. GAMM-Mitteilungen 2024, 47(4), e202470004. [Google Scholar] [CrossRef]
- Hansen, P.C. (2010). Discrete Inverse Problems: Insight and Algorithms. SIAM.
- Benning, M.; Burger, M. Modern regularization methods for inverse problems. Acta Numerica 2018, 27, 1–111. [Google Scholar] [CrossRef]
- Fortunato, D.; Townsend, A.; Trefethen, L.N. A Spectral Method for Elliptic Problems in a Rectangle. IMA Journal of Numerical Analysis 2020, 40, 2250–2280. [Google Scholar]
- Klawonn, A.; Lanser, M.; Weber, J. Machine learning and domain decomposition methods - a survey. Computational Science and Engineering 2024, 1(1), 2. [Google Scholar] [CrossRef]
- Bakurov, I.; Buzzelli, M.; Schettini, R.; Bianco, S.; Raimondo, F. Structural similarity index (SSIM) revisited: A data-driven approach. Expert Systems with Applications 2022, 189, 116087. [Google Scholar] [CrossRef]
- Nguyen, K.; Ho, N. Revisiting sliced Wasserstein on images: From vectorization to convolution. Advances in Neural Information Processing Systems 2022, 35, 17788–17801. [Google Scholar]
| Dataset | #Samples | #WSIs | Magnification | Color variation source | Body part |
| MITOS-ATYPIA-14 | 14 | 28 | 40×/0.25m | Two imaging scanners | Breast |
| SCC dataset | 44 | 220 | 40×/0.25m | Five imaging scanners | Skin |
| HEV dataset | 1 | 9 | 40×/0.25m | Nine staining protocols | Thyroid |
| SUSY-BF-10 | 25 | 50 | 40×/0.25m | Color fading in 10 years | Breast |
| Dataset | Pair | CycleGAN† | Reinhard | GCTI | SPN (ours) | ||||
| SSIM ↑ | W-D ↓ | SSIM ↑ | W-D ↓ | SSIM ↑ | W-D ↓ | SSIM ↑ | W-D ↓ | ||
| SYSU-BF | 18-n → o | 0.742 | 18.572 | 0.981 | 0.751 | 0.970 | 1.104 | 0.875 | 0.689 |
| mitos-14 | A06_01 → H | 0.790 | 7.274 | 0.911 | 2.558 | 0.813 | 7.042 | 0.822 | 3.027 |
| HEV data | HE → longE | 0.957 | 15.392 | 0.920 | 4.280 | 0.736 | 6.338 | 0.983 | 3.524 |
| HE → onlyE | – | 12.134 | 0.729 | 2.464 | 0.832 | 6.485 | 0.718 | 1.940 | |
| HE → shortE | – | 11.902 | 0.932 | 5.577 | 0.832 | 13.999 | 0.992 | 3.705 | |
| HE → longHE | – | 5.892 | 0.765 | 3.370 | 0.656 | 10.915 | 0.814 | 4.255 | |
| HE → onlyH | – | 18.802 | 0.920 | 2.540 | 0.634 | 4.427 | 0.979 | 3.672 | |
| HE → shortHE | – | 15.113 | 0.873 | 3.488 | 0.667 | 4.096 | 0.896 | 3.118 | |
| HE → longH | – | 4.792 | 0.875 | 4.745 | 0.755 | 22.539 | 0.921 | 2.146 | |
| HE → shortH | – | 13.199 | 0.825 | 2.918 | 0.694 | 13.107 | 0.857 | 3.218 | |
| SCC data | cs2 → gt450 | 0.921 | 25.624 | 0.679 | 0.495 | 0.757 | 1.873 | 0.641 | 0.360 |
| cs2 → nz20 | – | 20.885 | 0.838 | 0.801 | 0.898 | 1.950 | 0.882 | 1.715 | |
| cs2 → nz210 | – | 16.042 | 0.880 | 0.524 | 0.917 | 2.074 | 0.827 | 0.581 | |
| cs2 → p1000 | – | 10.979 | 0.886 | 1.022 | 0.798 | 4.310 | 0.912 | 1.579 | |
| synthetic_image | 0.816 | 17.433 | 0.906 | 4.352 | 0.831 | 17.117 | 0.950 | 3.706 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




