Submitted:
20 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
Study Setting and Sample Collection
Generic E. coli by Membrane Filtration
3. GCR E. coli by Membrane Filtration
E. coli Isolate ESBL Phenotype Testing
E. coli Isolate Antibiotic Susceptibility Testing
Human Fecal Marker Testing by Digital PCR
Statistical Analysis
Results and Discussion
Method Comparisons for ESBL-Ec Screening from Water and Wastewater
3. GCR E. coli and ESBL-Ec in OWTS Effluent
3. GCR E. coli and ESBL-Ec in Surface Waters
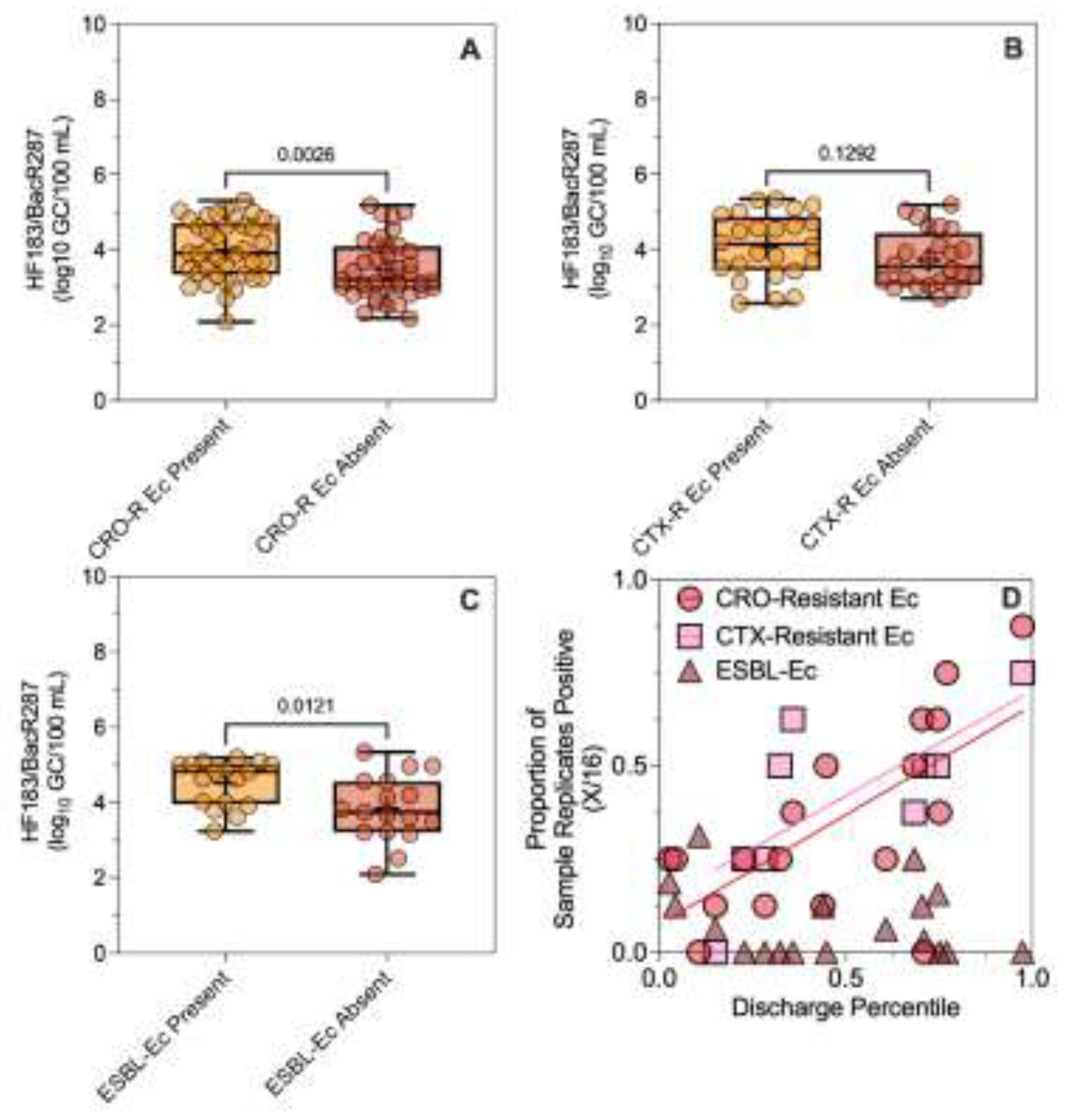
Associations between 3GCR E. coli, Human Sewage and, River Discharge
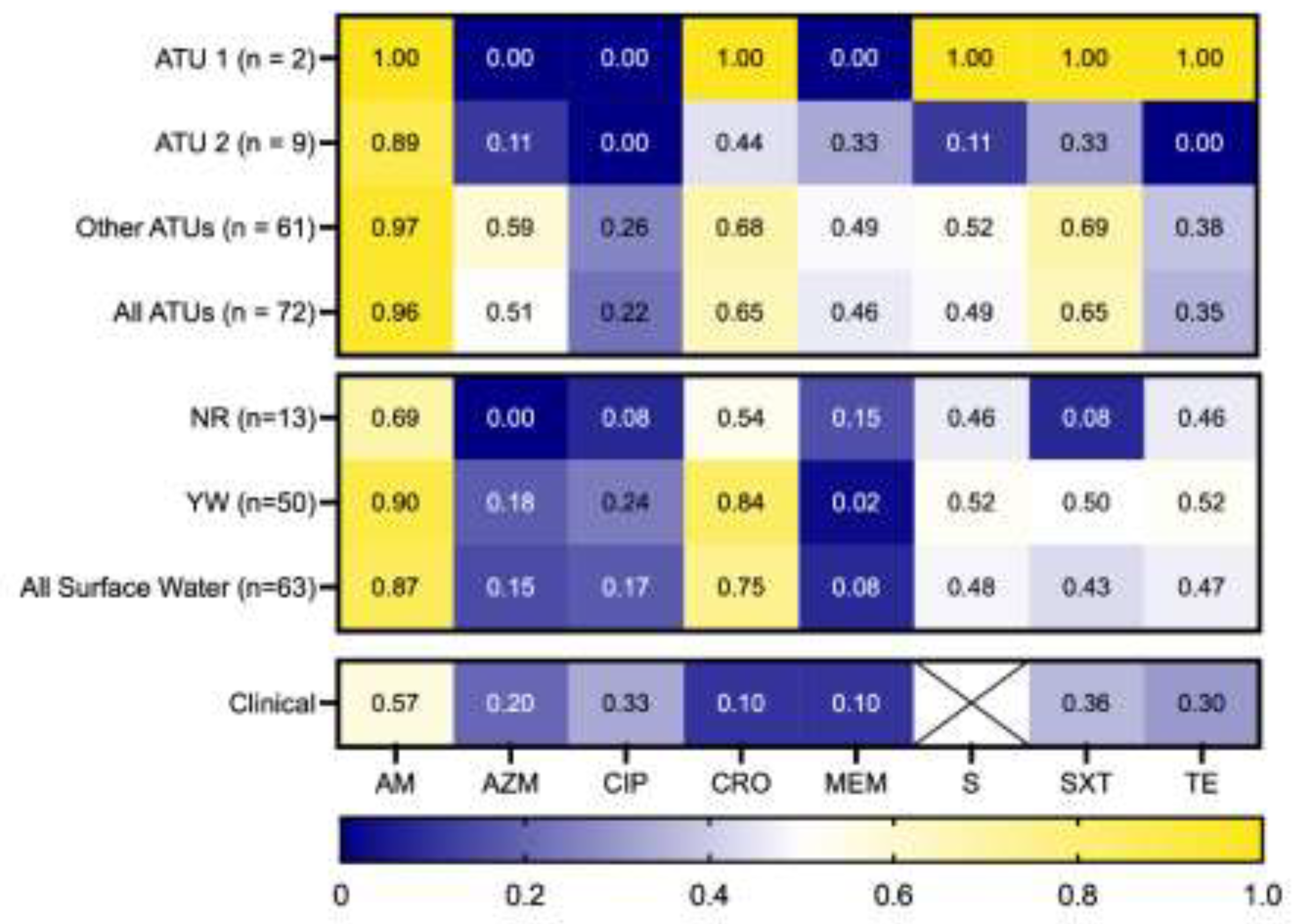
Antibiotic Susceptibility Among 3GCR and ESBL-Ec Isolates
Study Limitations
Conclusions
Supplementary Materials
Acknowledgments
References
- Keenum, I.; Berendonk, T. U.; Bonnedahl, J.; Cytryn, E.; Dagot, C.; Karkman, A.; Fatta-Kassinos, D.; Hayes, A.; Kirschner, A.; Kreft, J.-U.; Manaia, C. M.; Merlin, C.; Rahman, N.; Tipper, H.; Vass, M.; Zastepa, A. Towards One Health Action for Addressing Antimicrobial Resistance in the Age of Polycrisis. Nat Sustain 2026, 9(1), 24–34. [Google Scholar] [CrossRef]
- U.S. CDC. Antibiotic Resistance Threats in the United States 2019; U.S. Department of Health and Human Services: Atlanta, GA, 2019; p. 150. [Google Scholar] [CrossRef]
- Nelson, R. E.; Hatfield, K. M.; Wolford, H.; Samore, M. H.; Scott, R. D., II; Reddy, S. C.; Olubajo, B.; Paul, P.; Jernigan, J. A.; Baggs, J. National Estimates of Healthcare Costs Associated With Multidrug-Resistant Bacterial Infections Among Hospitalized Patients in the United States. Clin Infect Dis 2021, 72 (Supplement_1), S17–S26. [Google Scholar] [CrossRef]
- Murray, C. J. L.; Ikuta, K. S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; Johnson, S. C.; Browne, A. J.; Chipeta, M. G.; Fell, F.; Hackett, S.; Haines-Woodhouse, G.; Kashef Hamadani, B. H.; Kumaran, E. A. P.; McManigal, B.; Achalapong, S.; Agarwal, R.; Akech, S.; Albertson, S.; Amuasi, J.; Andrews, J.; Aravkin, A.; Ashley, E.; Babin, F.-X.; Bailey, F.; Baker, S.; Basnyat, B.; Bekker, A.; Bender, R.; Berkley, J. A.; Bethou, A.; Bielicki, J.; Boonkasidecha, S.; Bukosia, J.; Carvalheiro, C.; Castañeda-Orjuela, C.; Chansamouth, V.; Chaurasia, S.; Chiurchiù, S.; Chowdhury, F.; Clotaire Donatien, R.; Cook, A. J.; Cooper, B.; Cressey, T. R.; Criollo-Mora, E.; Cunningham, M.; Darboe, S.; Day, N. P. J.; De Luca, M.; Dokova, K.; Dramowski, A.; Dunachie, S. J.; Duong Bich, T.; Eckmanns, T.; Eibach, D.; Emami, A.; Feasey, N.; Fisher-Pearson, N.; Forrest, K.; Garcia, C.; Garrett, D.; Gastmeier, P.; Giref, A. Z.; Greer, R. C.; Gupta, V.; Haller, S.; Haselbeck, A.; Hay, S. I.; Holm, M.; Hopkins, S.; Hsia, Y.; Iregbu, K. C.; Jacobs, J.; Jarovsky, D.; Javanmardi, F.; Jenney, A. W. J.; Khorana, M.; Khusuwan, S.; Kissoon, N.; Kobeissi, E.; Kostyanev, T.; Krapp, F.; Krumkamp, R.; Kumar, A.; Kyu, H. H.; Lim, C.; Lim, K.; Limmathurotsakul, D.; Loftus, M. J.; Lunn, M.; Ma, J.; Manoharan, A.; Marks, F.; May, J.; Mayxay, M.; Mturi, N.; Munera-Huertas, T.; Musicha, P.; Musila, L. A.; Mussi-Pinhata, M. M.; Naidu, R. N.; Nakamura, T.; Nanavati, R.; Nangia, S.; Newton, P.; Ngoun, C.; Novotney, A.; Nwakanma, D.; Obiero, C. W.; Ochoa, T. J.; Olivas-Martinez, A.; Olliaro, P.; Ooko, E.; Ortiz-Brizuela, E.; Ounchanum, P.; Pak, G. D.; Paredes, J. L.; Peleg, A. Y.; Perrone, C.; Phe, T.; Phommasone, K.; Plakkal, N.; Ponce-de-Leon, A.; Raad, M.; Ramdin, T.; Rattanavong, S.; Riddell, A.; Roberts, T.; Robotham, J. V.; Roca, A.; Rosenthal, V. D.; Rudd, K. E.; Russell, N.; Sader, H. S.; Saengchan, W.; Schnall, J.; Scott, J. A. G.; Seekaew, S.; Sharland, M.; Shivamallappa, M.; Sifuentes-Osornio, J.; Simpson, A. J.; Steenkeste, N.; Stewardson, A. J.; Stoeva, T.; Tasak, N.; Thaiprakong, A.; Thwaites, G.; Tigoi, C.; Turner, C.; Turner, P.; van Doorn, H. R.; Velaphi, S.; Vongpradith, A.; Vongsouvath, M.; Vu, H.; Walsh, T.; Walson, J. L.; Waner, S.; Wangrangsimakul, T.; Wannapinij, P.; Wozniak, T.; Young Sharma, T. E. M. W.; Yu, K. C.; Zheng, P.; Sartorius, B.; Lopez, A. D.; Stergachis, A.; Moore, C.; Dolecek, C.; Naghavi, M. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. The Lancet 2022, 399 (10325), 629–655.
- WHO. Antimicrobial resistance. Available online: https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance (accessed on 2024-08-16).
- Vikesland, P. J.; Pruden, A.; Alvarez, P. J. J.; Aga, D.; Bürgmann, H.; Li, X.; Manaia, C. M.; Nambi, I.; Wigginton, K.; Zhang, T.; Zhu, Y.-G. Toward a Comprehensive Strategy to Mitigate Dissemination of Environmental Sources of Antibiotic Resistance. Environ. Sci. Technol. 2017, 51(22), 13061–13069. [Google Scholar] [CrossRef]
- Martinez, J. L. The Role of Natural Environments in the Evolution of Resistance Traits in Pathogenic Bacteria. Proc Biol Sci 2009, 276(1667), 2521–2530. [Google Scholar] [CrossRef]
- Huijbers, P. M. C.; Blaak, H.; de Jong, M. C. M.; Graat, E. A. M.; Vandenbroucke-Grauls, C. M. J. E.; de Roda Husman, A. M. Role of the Environment in the Transmission of Antimicrobial Resistance to Humans: A Review. Environ. Sci. Technol. 2015, 49(20), 11993–12004. [Google Scholar] [CrossRef] [PubMed]
- Graham, D. W.; Bergeron, G.; Bourassa, M. W.; Dickson, J.; Gomes, F.; Howe, A.; Kahn, L. H.; Morley, P. S.; Scott, H. M.; Simjee, S.; Singer, R. S.; Smith, T. C.; Storrs, C.; Wittum, T. E. Complexities in Understanding Antimicrobial Resistance across Domesticated Animal, Human, and Environmental Systems. Annals of the New York Academy of Sciences 2019, 1441(1), 17–30. [Google Scholar] [CrossRef]
- Wang, J.; Chu, L.; Wojnárovits, L.; Takács, E. Occurrence and Fate of Antibiotics, Antibiotic Resistant Genes (ARGs) and Antibiotic Resistant Bacteria (ARB) in Municipal Wastewater Treatment Plant: An Overview. Science of The Total Environment 2020, 744, 140997. [Google Scholar] [CrossRef]
- Read, D. S.; Gweon, H. S.; Bowes, M. J.; Anjum, M. F.; Crook, D. W.; Chau, K. K.; Shaw, L. P.; Hubbard, A.; AbuOun, M.; Tipper, H. J.; Hoosdally, S. J.; Bailey, M. J.; Walker, A. S.; Stoesser, N. Dissemination and Persistence of Antimicrobial Resistance (AMR) along the Wastewater-River Continuum. Water Research 2024, 264, 122204. [Google Scholar] [CrossRef] [PubMed]
- Sambaza, S. S.; Naicker, N. Contribution of Wastewater to Antimicrobial Resistance: A Review Article. Journal of Global Antimicrobial Resistance 2023, 34, 23–29. [Google Scholar] [CrossRef]
- LaPara, T. M.; Burch, T. R.; McNamara, P. J.; Tan, D. T.; Yan, M.; Eichmiller, J. J. Tertiary-Treated Municipal Wastewater Is a Significant Point Source of Antibiotic Resistance Genes into Duluth-Superior Harbor. Environ. Sci. Technol. 2011, 45(22), 9543–9549. [Google Scholar] [CrossRef]
- Mahaney, A. P.; Franklin, R. B. Persistence of Wastewater-Associated Antibiotic Resistant Bacteria in River Microcosms. Science of The Total Environment 2022, 819, 153099. [Google Scholar] [CrossRef] [PubMed]
- Maxcy-Brown, J.; Elliott, M. A.; Bearden, B. Household Level Wastewater Management and Disposal Data Collection in the U.S.: The History, Shortcomings, and Future Policy Implications. Water Policy 2023, 25(9), 927–947. [Google Scholar] [CrossRef]
- Brooks, B. W.; Callahan, T.; Stanley, J. K.; Holodak, J.; Stroski, K. M.; Cox, A. H.; Groves, T. W.; Jantrania, A.; Moeller, J. C.; Neset, K.; Walker, C.; Zhang, H.; Bakchan, A.; Alley, K. D.; Bell, J.; Blodig, A.; Casey, E.; Cosper, D.; D’Amato, V.; Elliott, M. A.; Graves, G.; Groover, R.; Himschoot, R.; Lusk, M. G.; Maxcy-Brown, J.; Meints, D.; Mejia, R.; Pace, M.; Ryan, B. J.; Scheffe, B.; Schimelfenig, T.; Wolfe, J. E.; Heger, S. F. Identifying Priority Research Questions for Decentralized Wastewater. Environ. Sci. Technol. 2026, 60(1), 49–63. [Google Scholar] [CrossRef] [PubMed]
- Connelly, K. N.; Wenger, S. J.; Gaur, N.; Bateman McDonald, J. M.; Occhipinti, M.; Capps, K. A. Assessing Relationships between Onsite Wastewater Treatment System Maintenance Patterns and System-Level Variables. Science of The Total Environment 2023, 870, 161851. [Google Scholar] [CrossRef] [PubMed]
- Capps, K. A.; Bateman McDonald, J. M.; Gaur, N.; Parsons, R. Assessing the Socio-Environmental Risk of Onsite Wastewater Treatment Systems to Inform Management Decisions. Environ. Sci. Technol. 2020, 54(23), 14843–14853. [Google Scholar] [CrossRef]
- Iverson, G.; Humphrey, C. P.; O’Driscoll, M. A.; Sanderford, C.; Jernigan, J.; Serozi, B. Nutrient Exports from Watersheds with Varying Septic System Densities in the North Carolina Piedmont. Journal of Environmental Management 2018, 211, 206–217. [Google Scholar] [CrossRef]
- Teerlink, J.; Hering, A. S.; Higgins, C. P.; Drewes, J. E. Variability of Trace Organic Chemical Concentrations in Raw Wastewater at Three Distinct Sewershed Scales. Water Research 2012, 46(10), 3261–3271. [Google Scholar] [CrossRef]
- Conn, K. E.; Barber, L. B.; Brown, G. K.; Siegrist, R. L. Occurrence and Fate of Organic Contaminants during Onsite Wastewater Treatment. Environ. Sci. Technol. 2006, 40(23), 7358–7366. [Google Scholar] [CrossRef]
- Carrara, C.; Ptacek, C. J.; Robertson, W. D.; Blowes, D. W.; Moncur, M. C.; Sverko, E.; Backus, S. Fate of Pharmaceutical and Trace Organic Compounds in Three Septic System Plumes, Ontario, Canada. Environ. Sci. Technol. 2008, 42(8), 2805–2811. [Google Scholar] [CrossRef]
- Gao, Q.; Blum, K. M.; Gago-Ferrero, P.; Wiberg, K.; Ahrens, L.; Andersson, P. L. Impact of On-Site Wastewater Infiltration Systems on Organic Contaminants in Groundwater and Recipient Waters. Science of The Total Environment 2019, 651, 1670–1679. [Google Scholar] [CrossRef]
- Subedi, B.; Codru, N.; Dziewulski, D. M.; Wilson, L. R.; Xue, J.; Yun, S.; Braun-Howland, E.; Minihane, C.; Kannan, K. A Pilot Study on the Assessment of Trace Organic Contaminants Including Pharmaceuticals and Personal Care Products from On-Site Wastewater Treatment Systems along Skaneateles Lake in New York State, USA. Water Research 2015, 72, 28–39. [Google Scholar] [CrossRef]
- Tan, L.; Zhang, C.; Liu, F.; Chen, P.; Wei, X.; Li, H.; Yi, G.; Xu, Y.; Zheng, X. Three-Compartment Septic Tanks as Sustainable on-Site Treatment Facilities? Watch out for the Potential Dissemination of Human-Associated Pathogens and Antibiotic Resistance. Journal of Environmental Management 2021, 300, 113709. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Chen, H.; Feng, Q.; Zhang, X.; Wu, D.; Feng, J.; Cheng, S.; Wang, D.; Liu, Z.; Zhong, Q.; Wei, J.; Liu, G. Dissemination of Antibiotic Resistance Genes through Fecal Sewage Treatment Facilities to the Ecosystem in Rural Area. Journal of Environmental Management 2023, 333, 117439. [Google Scholar] [CrossRef]
- Damashek, J.; Westrich, J. R.; McDonald, J. M. B.; Teachey, M. E.; Jackson, C. R.; Frye, J. G.; Lipp, E. K.; Capps, K. A.; Ottesen, E. A. Non-Point Source Fecal Contamination from Aging Wastewater Infrastructure Is a Primary Driver of Antibiotic Resistance in Surface Waters. Water Research 2022, 222, 118853. [Google Scholar] [CrossRef] [PubMed]
- Sidhu, A. S.; Mikolajczyk, F. N.; Fisher, J. C. Antimicrobial Resistance Linked to Septic System Contamination in the Indiana Lake Michigan Watershed. Antibiotics (Basel) 2023, 12(3), 569. [Google Scholar] [CrossRef]
- Carroll, S.; Hargreaves, M.; Goonetilleke, A. Sourcing Faecal Pollution from Onsite Wastewater Treatment Systems in Surface Waters Using Antibiotic Resistance Analysis*. J. Appl. Microbiol. 2005, 99(3), 471–482. [Google Scholar] [CrossRef]
- Heida, A.; Hamilton, M. T.; Gambino, J.; Sanderson, K.; Schoen, M. E.; Jahne, M. A.; Garland, J.; Ramirez, L.; Quon, H.; Lopatkin, A. J.; Hamilton, K. A. Population Ecology-Quantitative Microbial Risk Assessment (QMRA) Model for Antibiotic-Resistant and Susceptible E. Coli in Recreational Water. Environ. Sci. Technol. 2025, 59(9), 4266–4281. [Google Scholar] [CrossRef] [PubMed]
- WHO. WHO Integrated Global Surveillance on ESBL-Producing E. Coli Using a “One Health” Approach: Implementation and Opportunities; WHO, 2021; p. 76. Available online: https://apps.who.int/iris/rest/bitstreams/1335472/retrieve.
- WHO. WHO Bacterial Priority Pathogens List, 2024: Bacterial Pathogens of Public Health Importance to Guide Research, Development and Strategies to Prevent and Control Antimicrobial Resistance; WHO: Geneva, 2024; Available online: https://www.who.int/publications/i/item/9789240093461 (accessed on 2024-08-16).
- CDC. Antimicrobial Resistance Threats in the United States. Available online: https://www.cdc.gov/antimicrobial-resistance/data-research/threats/update-2022.html (accessed on 2026-01-07).
- Belding, C.; Boopathy, R. Presence of Antibiotic-Resistant Bacteria and Antibiotic Resistance Genes in Coastal Recreational Waters of Southeast Louisiana, USA. Journal of Water Supply: Research and Technology-Aqua 2018, 67(8), 800–809. [Google Scholar] [CrossRef]
- Irion, K. A Deal with the Devil: How Politics, Expediency and Economics Resulted in the Proliferation of Onsite Wastewater Systems for Subdivisions in Louisiana and the Effect That the Adoption Is Having on Wastewater Treatment; New Orleans, LA, 2016; p 16.
- LDEQ. Bayou Lafourche Watershed Implementation Plan: Upper Bayou Lafourche Watershed Subsegment 020401. 2018. Available online: http://nonpoint.deq.louisiana.gov/wqa/links/watershedplan/Barataria/Bayou%20Lafourche_Final.pdf (accessed on 2026-01-07).
- EPA, U. How’s My Waterway? US EPA. Available online: https://mywaterway.epa.gov (accessed on 2026-01-12).
- Lay, J. O., Jr. MALDI-TOF Mass Spectrometry of Bacteria. Mass Spectrometry Reviews 2001, 20(4), 172–194. [Google Scholar] [CrossRef]
- Bivins, A.; Srivastava, A.; Katara, A.; Weise, K.; Kachhawaha, A. S. Extended-Spectrum Beta Lactamase-Producing E. Coli in Freshwater Lakes along an Anthropogenic Impact Gradient. ACS ES&T Water 2025, 5(3), 1321–1330. [Google Scholar] [CrossRef]
- CLSI. M100 Performance Standards for Antimicrobial Susceptibility Testing, 33rd Edition; CLSI, 2023; p 402. Available online: https://clsi.org/standards/products/microbiology/documents/m100/.
- Hudzicki, J. Kirby-Bauer Disk Diffusion Susceptibility Test Protocol; ASM. 2009. Available online: https://asm.org/getattachment/2594ce26-bd44-47f6-8287-0657aa9185ad/Kirby-Bauer-Disk-Diffusion-Susceptibility-Test-Protocol-pdf.pdf.
- US EPA. Method 1696.1: Characterization of Human Fecal Pollution in Water by HF183/BacR287 TaqMan® Quantitative Polymerase Chain Reaction (qPCR) Assay; EPA 821-R-22-001; Office of Water, 2022; p 56. Available online: https://www.epa.gov/system/files/documents/2022-01/draft_method-1696.1-hf183_01052022_508.pdf.
- Ghosh, S.; Bivins, A. Optimizing Digital PCR Assays for Multiplexed Microbial Source Tracking in Surface Waters. In Digital PCR: Methods and Protocols; Pavlovic, G., Ed.; Springer US: New York, NY, 2026; pp. 61–85. [Google Scholar] [CrossRef]
- Cohen, J. A Coefficient of Agreement for Nominal Scales. Educational and Psychological Measurement 1960, 20(1), 37–46. [Google Scholar] [CrossRef]
- Upton, G. J. G. Fisher’s Exact Test. J R Stat Soc Ser A Stat Soc 1992, 155(3), 395–402. [Google Scholar] [CrossRef]
- Agresti, A.; Coull, B. A. Approximate Is Better than “Exact” for Interval Estimation of Binomial Proportions. The American Statistician 1998, 52(2), 119–126. [Google Scholar] [CrossRef] [PubMed]
- Spearman, C. The Proof and Measurement of Association between Two Things. The American Journal of Psychology 1904, 15(1), 72–101. [Google Scholar] [CrossRef]
- Kruskal, W. H.; Wallis, W. A. Use of Ranks in One-Criterion Variance Analysis. Journal of the American Statistical Association 1952, 47(260), 583–621. [Google Scholar] [CrossRef]
- Banu, R. A.; Alvarez, J. M.; Reid, A. J.; Enbiale, W.; Labi, A.-K.; Ansa, E. D. O.; Annan, E. A.; Akrong, M. O.; Borbor, S.; Adomako, L. A. B.; Ahmed, H.; Mustapha, M. B.; Davtyan, H.; Owiti, P.; Hedidor, G. K.; Quarcoo, G.; Opare, D.; Kikimoto, B.; Osei-Atweneboana, M. Y.; Schmitt, H. Extended Spectrum Beta-Lactamase Escherichia Coli in River Waters Collected from Two Cities in Ghana, 2018–2020. Tropical Medicine and Infectious Disease 2021, 6(2), 105. [Google Scholar] [CrossRef]
- Pek, H. B.; Kadir, S. A.; Arivalan, S.; Osman, S.; Mohamed, R.; Ng, L. C.; Wong, J. C.; Octavia, S. Screening for Extended-Spectrum Beta-Lactamase Escherichia Coli in Recreational Beach Waters in Singapore. Future Microbiology 2023, 18(13), 867–873. [Google Scholar] [CrossRef]
- Water Quality. Available online: https://mics.unicef.org/methodological-work/water-quality (accessed on 2026-02-11).
- Hornsby, G.; Ibitoye, T. D.; Keelara, S.; Harris, A. Validation of a Modified IDEXX Defined-Substrate Assay for Detection of Antimicrobial Resistant E. Coli in Environmental Reservoirs. Environ. Sci.: Processes Impacts 2023, 25(1), 37–43. [Google Scholar] [CrossRef] [PubMed]
- Mizuochi, S.; Nelson, M.; Baylis, C.; Green, B.; Jewell, K.; Monadjemi, F.; Chen, Y.; Salfinger, Y.; Fernandez, M. C. Matrix Extension Study: Validation of the Compact Dry EC Method for Enumeration of Escherichia Coli and Non-E. Coli Coliform Bacteria in Selected Foods. J AOAC Int 2016, 99(2), 451–460. [Google Scholar] [CrossRef]
- Galvin, S.; Boyle, F.; Hickey, P.; Vellinga, A.; Morris, D.; Cormican, M. Enumeration and Characterization of Antimicrobial-Resistant Escherichia Coli Bacteria in Effluent from Municipal, Hospital, and Secondary Treatment Facility Sources. Applied and Environmental Microbiology 2010, 76(14), 4772–4779. [Google Scholar] [CrossRef] [PubMed]
- Korzeniewska, E.; Harnisz, M. Extended-Spectrum Beta-Lactamase (ESBL)-Positive Enterobacteriaceae in Municipal Sewage and Their Emission to the Environment. Journal of Environmental Management 2013, 128, 904–911. [Google Scholar] [CrossRef]
- Blaak, H.; de Kruijf, P.; Hamidjaja, R. A.; van Hoek, A. H. A. M.; de Roda Husman, A. M.; Schets, F. M. Prevalence and Characteristics of ESBL-Producing E. Coli in Dutch Recreational Waters Influenced by Wastewater Treatment Plants. Veterinary Microbiology 2014, 171(3), 448–459. [Google Scholar] [CrossRef] [PubMed]
- van Heijnsbergen, E.; Niebaum, G.; Lämmchen, V.; Borneman, A.; Hernández Leal, L.; Klasmeier, J.; Schmitt, H. (Antibiotic-Resistant) E. Coli in the Dutch–German Vecht Catchment─Monitoring and Modeling. Environ. Sci. Technol. 2022, 56(21), 15064–15073. [Google Scholar] [CrossRef] [PubMed]
- Jäger, T.; Hembach, N.; Elpers, C.; Wieland, A.; Alexander, J.; Hiller, C.; Krauter, G.; Schwartz, T. Reduction of Antibiotic Resistant Bacteria During Conventional and Advanced Wastewater Treatment, and the Disseminated Loads Released to the Environment. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Bivins, A.; North, D.; Wu, Z.; Shaffer, M.; Ahmed, W.; Bibby, K. Within-and between-Day Variability of SARS-CoV-2 RNA in Municipal Wastewater during Periods of Varying COVID-19 Prevalence and Positivity. Acs Es&T Water 2021, 1(9), 2097–2108. [Google Scholar]
- Murray, G. E.; Tobin, R. S.; Junkins, B.; Kushner, D. J. Effect of Chlorination on Antibiotic Resistance Profiles of Sewage-Related Bacteria. Applied and Environmental Microbiology 1984, 48(1), 73–77. [Google Scholar] [CrossRef]
- Garcia, C. R.; Norfolk, W. A.; Howard, A. K.; Glatter, A. L.; Beaudry, M. S.; Mallis, N. A.; Welton, M.; Glenn, T. C.; Lipp, E. K.; Ottesen, E. A. Long-Term Gut Colonization with ESBL-Producing Escherichia Coli in Participants without Known Risk Factors from the Southeastern United States. medRxiv 2024, 2024.02.03.24302254. [Google Scholar] [CrossRef]
- Keenum, I.; Calarco, J.; Majeed, H.; Hager-Soto, E. E.; Bott, C.; Garner, E.; Harwood, V. J.; Pruden, A. To What Extent Do Water Reuse Treatments Reduce Antibiotic Resistance Indicators? A Comparison of Two Full-Scale Systems. Water Research 2024, 254, 121425. [Google Scholar] [CrossRef]
- Craddock, H. A.; Chattopadhyay, S.; Rjoub, Y.; Rosen, D.; Greif, J.; Lipchin, C.; Mongodin, E. F.; Sapkota, A. R. Antibiotic-Resistant Escherichia Coli and Klebsiella Spp. in Greywater Reuse Systems and Pond Water Used for Agricultural Irrigation in the West Bank, Palestinian Territories. Environmental Research 2020, 188, 109777. [Google Scholar] [CrossRef]
- Diriba, K.; Awulachew, E.; Gemede, A.; Anja, A. The Magnitude of Extended-Spectrum Beta-Lactamase- Producing Enterobacteriaceae from Clinical Samples in Ethiopia: A Systematic Review and Meta-Analysis. Access Microbiol 2021, 3(3), 000195. [Google Scholar] [CrossRef]
- Woerther, P.-L.; Burdet, C.; Chachaty, E.; Andremont, A. Trends in Human Fecal Carriage of Extended-Spectrum β-Lactamases in the Community: Toward the Globalization of CTX-M. Clinical Microbiology Reviews 2013, 26(4), 744–758. [Google Scholar] [CrossRef]
- Ruppé, E. Lessons from a Global Antimicrobial Resistance Surveillance Network. Bull World Health Organ 2023, 101(10), 672–678. [Google Scholar] [CrossRef]
- Milenkov, M.; Proux, C.; Rasolofoarison, T. L.; Rakotomalala, F. A.; Rasoanandrasana, S.; Rahajamanana, V. L.; Rafalimanana, C.; Ravaoarisaina, Z.; Ramahatafandry, I. T.; Westeel, E.; Petitjean, M.; Berti, V.; Marin, J.; Mullaert, J.; Han, L.; Clermont, O.; Raskine, L.; Endtz, H.; Andremont, A.; Denamur, E.; Komurian-Pradel, F.; Samison, L. H.; Armand-Lefevre, L. Implementation of the WHO Tricycle Protocol for Surveillance of Extended-Spectrum β-Lactamase Producing Escherichia Coli in Humans, Chickens, and the Environment in Madagascar: A Prospective Genomic Epidemiology Study. The Lancet Microbe 2024, 5(8), 100850. [Google Scholar] [CrossRef] [PubMed]
- Puspandari, N.; Sunarno, S.; Febrianti, T.; Febriyana, D.; Saraswati, R. D.; Rooslamiati, I.; Amalia, N.; Nursofiah, S.; Hartoyo, Y.; Herna, H.; Mursinah, M.; Muna, F.; Aini, N.; Risniati, Y.; Dhewantara, P. W.; Allamanda, P.; Wicaksana, D. N.; Sukoco, R.; Efadeswarni; Nelwan, E. J.; Cahyarini; Haryanto, B.; Sihombing, B.; Soares Magalhães, R. J.; Kakkar, M.; Setiawaty, V.; Matheu, J. Extended Spectrum Beta-Lactamase-Producing Escherichia Coli Surveillance in the Human, Food Chain, and Environment Sectors: Tricycle Project (Pilot) in Indonesia. One Health 2021, 13, 100331. [Google Scholar] [CrossRef] [PubMed]
- Appling, K. C.; Sobsey, M. D.; Durso, L. M.; Fisher, M. B. Environmental Monitoring of Antimicrobial Resistant Bacteria in North Carolina Water and Wastewater Using the WHO Tricycle Protocol in Combination with Membrane Filtration and Compartment Bag Test Methods for Detecting and Quantifying ESBL E. Coli. PLOS Water 2023, 2(9), e0000117. [Google Scholar] [CrossRef]
- Heida, A.; Hamilton, M. T.; Gambino, J.; Sanderson, K.; Schoen, M. E.; Jahne, M. A.; Garland, J.; Ramirez, L.; Quon, H.; Lopatkin, A. J.; Hamilton, K. A. Population Ecology-Quantitative Microbial Risk Assessment (QMRA) Model for Antibiotic-Resistant and Susceptible E. Coli in Recreational Water. Environ. Sci. Technol. 2025, 59(9), 4266–4281. [Google Scholar] [CrossRef]
- Jørgensen, S. B.; Søraas, A. V.; Arnesen, L. S.; Leegaard, T. M.; Sundsfjord, A.; Jenum, P. A. A Comparison of Extended Spectrum β-Lactamase Producing Escherichia Coli from Clinical, Recreational Water and Wastewater Samples Associated in Time and Location. PLOS ONE 2017, 12(10), e0186576. [Google Scholar] [CrossRef]
- Carroll, S.; Hargreaves, M.; Goonetilleke, A. Sourcing Faecal Pollution from Onsite Wastewater Treatment Systems in Surface Waters Using Antibiotic Resistance Analysis*. J. Appl. Microbiol. 2005, 99(3), 471–482. [Google Scholar] [CrossRef]
- Lee, S.; Suits, M.; Wituszynski, D.; Winston, R.; Martin, J.; Lee, J. Residential Urban Stormwater Runoff: A Comprehensive Profile of Microbiome and Antibiotic Resistance. Science of The Total Environment 2020, 723, 138033. [Google Scholar] [CrossRef]
- Herrig, I.; Fleischmann, S.; Regnery, J.; Wesp, J.; Reifferscheid, G.; Manz, W. Prevalence and Seasonal Dynamics of blaCTX-M Antibiotic Resistance Genes and Fecal Indicator Organisms in the Lower Lahn River, Germany. PLOS ONE 2020, 15(4), e0232289. [Google Scholar] [CrossRef]
- Kalanxhi, E.; Laxminarayan, R. Climate Change and Antimicrobial Resistance. Nat Rev Microbiol 2026, 1–12. [Google Scholar] [CrossRef]
- MacFadden, D. R.; McGough, S. F.; Fisman, D.; Santillana, M.; Brownstein, J. S. Antibiotic Resistance Increases with Local Temperature. Nature Clim Change 2018, 8(6), 510–514. [Google Scholar] [CrossRef] [PubMed]
- Louisiana Department of Health. The Louisiana Antibiogram: Louisiana Antibiotic Resistance 2016. 2016. Available online: https://ldh.la.gov/assets/oph/Center-PHCH/Center-CH/infectious-epi/AntibioticSensitivity/LAAntibiogram/LouisianaAntibioticResistance2016_FINAL.pdf (accessed on 2025-01-10).
- Pruden, A.; Vikesland, P. J.; Davis, B. C.; de Roda Husman, A. M. Seizing the Moment: Now Is the Time for Integrated Global Surveillance of Antimicrobial Resistance in Wastewater Environments. Current Opinion in Microbiology 2021, 64, 91–99. [Google Scholar] [CrossRef] [PubMed]
- Chau, K. K.; Barker, L.; Budgell, E. P.; Vihta, K. D.; Sims, N.; Kasprzyk-Hordern, B.; Harriss, E.; Crook, D. W.; Read, D. S.; Walker, A. S.; Stoesser, N. Systematic Review of Wastewater Surveillance of Antimicrobial Resistance in Human Populations. Environment International 2022, 162, 107171. [Google Scholar] [CrossRef]
- Pärnänen, K. M. M.; Narciso-da-Rocha, C.; Kneis, D.; Berendonk, T. U.; Cacace, D.; Do, T. T.; Elpers, C.; Fatta-Kassinos, D.; Henriques, I.; Jaeger, T.; Karkman, A.; Martinez, J. L.; Michael, S. G.; Michael-Kordatou, I.; O’Sullivan, K.; Rodriguez-Mozaz, S.; Schwartz, T.; Sheng, H.; Sørum, H.; Stedtfeld, R. D.; Tiedje, J. M.; Giustina, S. V. D.; Walsh, F.; Vaz-Moreira, I.; Virta, M.; Manaia, C. M. Antibiotic Resistance in European Wastewater Treatment Plants Mirrors the Pattern of Clinical Antibiotic Resistance Prevalence. Science Advances 2019, 5(3), eaau9124. [Google Scholar] [CrossRef] [PubMed]
- Yasmin, S.; Karim, A.-M.; Lee, S.-H.; Zahra, R. Temporal Variation of Meropenem Resistance in E. Coli Isolated from Sewage Water in Islamabad, Pakistan. Antibiotics (Basel) 2022, 11(5), 635. [Google Scholar] [CrossRef]
- Loumame, E. hassan; Tounsi, A.; Amir, S.; Soraa, N.; Ouazzani, N. Microbial Resistance to Carbapenems in Effluents from Gynaecological, Paediatric and Surgical Hospital Units. Antibiotics 2022, 11(8). [Google Scholar] [CrossRef] [PubMed]
- Ogura, Y.; Ueda, T.; Nukazawa, K.; Hiroki, H.; Xie, H.; Arimizu, Y.; Hayashi, T.; Suzuki, Y. The Level of Antimicrobial Resistance of Sewage Isolates Is Higher than That of River Isolates in Different Escherichia Coli Lineages. Sci Rep 2020, 10(1), 17880. [Google Scholar] [CrossRef]
- Gschwind, R.; Bonnet, M.; Abramova, A.; Jarquín-Díaz, V. H.; Wenne, M.; Löber, U.; Godron, N.; Kampouris, I. D.; Tskhay, F.; Nahid, F.; Debroucker, C.; Bui-Hai, M.; El Aiba, I.; Klümper, U.; Berendonk, T. U.; Forslund-Startceva, S. K.; Zahra, R.; Bengtsson-Palme, J.; Ruppé, E. Cefiderocol Resistance Genes Identified in Environmental Samples Using Functional Metagenomics. ISME J 2026, 20(1), wrag010. [Google Scholar] [CrossRef]
| River |
3GCR E. coli by TBX-CRO (log10 CFU/100 mL) |
3GCR E. coli by TBX-CTX (log10 CFU/100 mL) |
E. coli by TBX (log10 CFU/100 mL) |
| Natalbany (NR) | LC (< 0): 125 RC (> 2.176): 0 UC: 18 µ = -1.77 σ = 1.54 |
LC (< 0): 55 RC (> 2.176): 0 UC: 17 µ = -0.89 σ = 1.24 |
LC (< 0) = 17 RC (> 2.176) = 25 UC: 96 µ = 1.54 σ = 0.98 |
| Yellow Water (YW) | LC (< 0): 80 RC (> 2.176): 0 UC: 62 µ = -0.21 σ = 1.05 |
LC (< 0): 37 RC (> 2.176): 2 UC: 33 µ = -0.08 σ = 1.10 |
LC (< 0): 9 RC (> 2.176): 73 UC: 55 µ = 2.39 σ = 1.14 |
| NR vs. YW | p = 4.85 x 10-8 | p = 0.0089 | p = 1.91 x 10-8 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




