Submitted:
18 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Genomic Organization and Sequence Features of the miR-101 Family
1.2. Chronic Inflammation as a Driver of Malignant Transformation via miR-101a
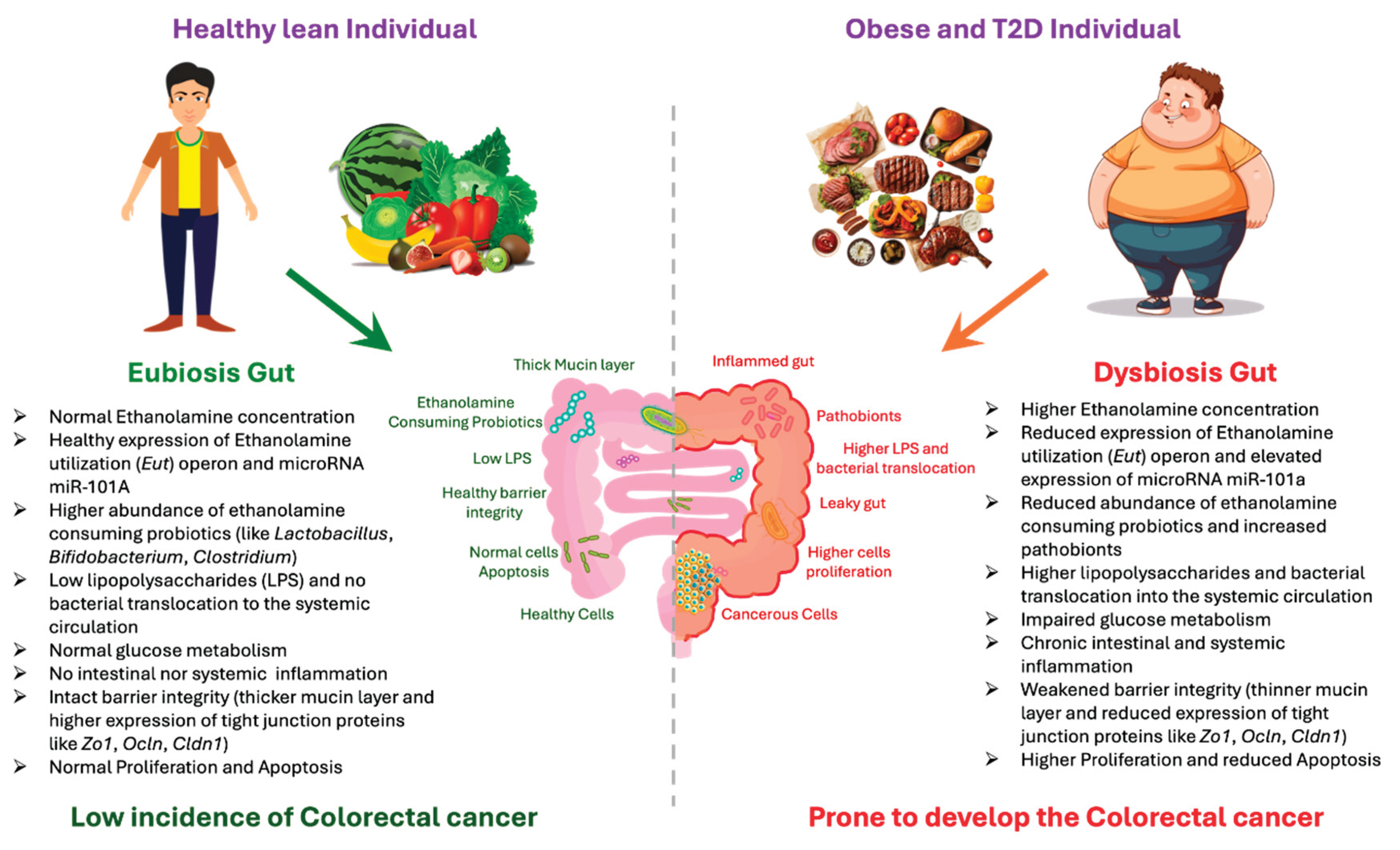
1.3. Obesity Associated Colorectal Cancer Development via miR-101a
2. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Akimoto, N. Rising incidence of early-onset colorectal cancer - a call to action. Nat Rev Clin Oncol 2021, 18(4), 230–243. [Google Scholar] [CrossRef] [PubMed]
- Mandic, M. Is the association of overweight and obesity with colorectal cancer underestimated? An umbrella review of systematic reviews and meta-analyses. Eur J Epidemiol 2023, 38(2), 135–144. [Google Scholar] [CrossRef]
- Arima, K. Western-Style Diet, pks Island-Carrying Escherichia coli, and Colorectal Cancer: Analyses From Two Large Prospective Cohort Studies. Gastroenterology 2022, 163(4), 862–874. [Google Scholar] [CrossRef] [PubMed]
- Lednovich, K.R. Intestinal FFA3 mediates obesogenic effects in mice on a Western diet. American Journal of Physiology-Endocrinology and Metabolism 2022, 323(3), E290–E306. [Google Scholar] [CrossRef] [PubMed]
- Lednovich, K.R. Intestinal FFA2 promotes obesity by altering food intake in Western diet-fed mice. Journal of Endocrinology 2024, 260(2), p. e230184. [Google Scholar] [CrossRef]
- He, L.; Hannon, G.J. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 2004, 5(7), 522–31. [Google Scholar] [CrossRef]
- Li, T. Translation suppresses exogenous target RNA-mediated microRNA decay. Nature Communications 2025, 16(1), p. 5257. [Google Scholar] [CrossRef]
- Xiao, C.; Rajewsky, K. MicroRNA Control in the Immune System: Basic Principles. Cell 2009, 136(1), 26–36. [Google Scholar] [CrossRef]
- Chaiwangyen, W. MicroRNA expression in response to environmental hazards: Implications for health. Ecotoxicology and Environmental Safety 2025, 300, 118420. [Google Scholar] [CrossRef]
- Singh, R. Dual Roles of miR-10a-5p and miR-10b-5p as Tumor Suppressors and Oncogenes in Diverse Cancers. Int J Mol Sci 2025, 26(1). [Google Scholar] [CrossRef]
- Otmani, K. The regulatory mechanisms of oncomiRs in cancer. Biomedicine & Pharmacotherapy 2024, 171, 116165. [Google Scholar] [CrossRef] [PubMed]
- Sareen, G. A new era of cancer immunotherapy: vaccines and miRNAs. Cancer Immunol Immunother 2025, 74(5), p. 163. [Google Scholar] [CrossRef]
- Chandramouli, A. MicroRNA-101 (miR-101) post-transcriptionally regulates the expression of EP4 receptor in colon cancers. Cancer Biology & Therapy 2012, 13(3), 175–183. [Google Scholar]
- Cohen, J.L. Amygdalar expression of the microRNA miR-101a and its target Ezh2 contribute to rodent anxiety-like behaviour. Eur J Neurosci 2017, 46(7), 2241–2252. [Google Scholar] [CrossRef]
- Dudakovic, A. MicroRNA-101a enhances trabecular bone accrual in male mice. Scientific Reports 2022, 12(1), p. 13361. [Google Scholar] [CrossRef]
- Liu, J.J. A novel AP-1/miR-101 regulatory feedback loop and its implication in the migration and invasion of hepatoma cells. Nucleic Acids Res 2014, 42(19), 12041–51. [Google Scholar] [CrossRef] [PubMed]
- de Sande, A.H. Cell-type-specific characterization of miRNA gene dynamics in immune cell subpopulations during aging and atherosclerosis disease development at single-cell resolution. bioRxiv 2023, 2023. 10.09.561173. [Google Scholar]
- Morita, M. Identification of microRNAs involved in acute rejection and spontaneous tolerance in murine hepatic allografts. Scientific Reports 2014, 4(1), p. 6649. [Google Scholar] [CrossRef]
- Wang, H. MiR-101 Targets the EZH2/Wnt/β-Catenin the Pathway to Promote the Osteogenic Differentiation of Human Bone Marrow-Derived Mesenchymal Stem Cells. Scientific Reports 2016, 6(1), p. 36988. [Google Scholar] [CrossRef] [PubMed]
- Vella, S. MicroRNA-101 is repressed by EZH2 and its restoration inhibits tumorigenic features in embryonal rhabdomyosarcoma. Clinical Epigenetics 2015, 7(1), p. 82. [Google Scholar] [CrossRef]
- Xu, B. Targeting EZH2 and PRC2 dependence as novel anticancer therapy. Experimental Hematology 2015, 43(8), 698–712. [Google Scholar] [CrossRef]
- Cao, Q. Coordinated Regulation of Polycomb Group Complexes through microRNAs in Cancer. Cancer Cell 2011, 20(2), 187–199. [Google Scholar] [CrossRef] [PubMed]
- Cao, P. MicroRNA-101 negatively regulates Ezh2 and its expression is modulated by androgen receptor and HIF-1alpha/HIF-1beta. Mol Cancer 2010, 9, 108. [Google Scholar] [CrossRef]
- Smits, M. miR-101 is down-regulated in glioblastoma resulting in EZH2-induced proliferation, migration, and angiogenesis. Oncotarget 2010, 1(8). [Google Scholar] [CrossRef]
- Hillyar, C.R.T. Complex Cross-Talk between EZH2 and Mirnas Confers Hallmark Characteristics and Shapes the Tumor Microenvironment. Epigenomics 2022, 14(11), 699–709. [Google Scholar] [CrossRef]
- Gómez-Acebo, I. Role of circulating MicroRNAs in prostate cancer diagnosis and risk stratification in the MCC Spain study. Sci Rep 2025, 15(1), p. 17517. [Google Scholar] [CrossRef]
- Huang, X. SChLAP1 regulates the metastasis and apoptosis of prostate cancer partly via miR-101. Transl Androl Urol 2025, 14(6), 1782–1796. [Google Scholar] [CrossRef]
- Liu, N. MiR-101: An Important Regulator of Gene Expression and Tumor Ecosystem. Cancers (Basel) 2022, 14(23). [Google Scholar] [CrossRef] [PubMed]
- Pastena, P. Unraveling Biomarker Signatures in Triple-Negative Breast Cancer: A Systematic Review for Targeted Approaches. International Journal of Molecular Sciences 2024, 25(5), 2559. [Google Scholar] [CrossRef]
- Tessitore, A. MicroRNAs in the DNA Damage/Repair Network and Cancer. In Int J Genomics; 2014; p. 820248. [Google Scholar]
- Schnekenburger, M.; Diederich, M. Epigenetics Offer New Horizons for Colorectal Cancer Prevention. Curr Colorectal Cancer Rep 2012, 8(1), 66–81. [Google Scholar] [PubMed]
- Tâlvan, C.-D. Exploring miRNA Profiles in Colon Cancer: A Focus on miR101-3p, miR106a-5p, and miR326. Cancers 2024, 16(12), 2285. [Google Scholar] [CrossRef]
- Mishra, S.P. A mechanism by which gut microbiota elevates permeability and inflammation in obese/diabetic mice and human gut. Gut 2023, 72(10), 1848–1865. [Google Scholar] [CrossRef]
- Tarek, G. MiR-101-3p and miR-106b-5p roles in EMT pathway: prognostic and therapeutic insights for luminal breast cancer. Journal of the Egyptian National Cancer Institute 2025, 37(1), p. 43. [Google Scholar] [CrossRef]
- Wu, H.; Ballantyne, C.M. Metabolic Inflammation and Insulin Resistance in Obesity. Circ Res 2020, 126(11), 1549–1564. [Google Scholar] [CrossRef] [PubMed]
- Jia, X. Impact of Gut Microbiota and Microbiota-Related Metabolites on Hyperlipidemia. Front Cell Infect Microbiol 2021, 11, 634780. [Google Scholar] [CrossRef] [PubMed]
- Semo, D.; Reinecke, H.; Godfrey, R. Gut microbiome regulates inflammation and insulin resistance: a novel therapeutic target to improve insulin sensitivity. Signal Transduction and Targeted Therapy 2024, 9(1), p. 35. [Google Scholar] [CrossRef] [PubMed]
- Yarahmadi, A. Understanding the complex function of gut microbiota: its impact on the pathogenesis of obesity and beyond: a comprehensive review. Diabetology & Metabolic Syndrome 2024, 16(1), 308. [Google Scholar] [CrossRef]
- Upadhyay, G. Targeted and untargeted lipidomics with integration of liver dynamics and microbiome after dietary reversal of obesogenic diet targeting inflammation-resolution signaling in aging mice. Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids 2024, 1869(8), 159542. [Google Scholar] [CrossRef]
- Nagpal, R.; Mishra, S.P.; Yadav, H. Unique Gut Microbiome Signatures Depict Diet-Versus Genetically Induced Obesity in Mice. International Journal of Molecular Sciences 2020, 21(10), 3434. [Google Scholar] [CrossRef]
- Glaros, S.B. Systemic and gut microbiome changes with metformin and liraglutide in youth-onset type 2 diabetes: the MIGHTY study. Gut Microbes 2025, 17(1), p. 2558071. [Google Scholar] [CrossRef]
- Dixon, S.A. The effects of prebiotics on gastrointestinal side effects of metformin in youth: A pilot randomized control trial in youth-onset type 2 diabetes. In Frontiers in Endocrinology; 2023; pp. 14–2023. [Google Scholar]
- Soundararajan, R. Integration of lipidomics with targeted, single cell, and spatial transcriptomics defines an unresolved pro-inflammatory state in colon cancer. Gut 2025, 74(4), 586–602. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J. Ethanolamine Metabolism in the Mammalian Gastrointestinal Tract: Mechanisms, Patterns, and Importance. Curr Mol Med 2017, 17(2), 92–99. [Google Scholar] [CrossRef]
- Bourdeau-Julien, I. The diet rapidly and differentially affects the gut microbiota and host lipid mediators in a healthy population. Microbiome 2023, 11(1), p. 26. [Google Scholar] [CrossRef]
- Tâlvan, C.-D. The Impact of miRNA Expression on Colon Cancer Severity, Invasiveness, and Localization. Cancers 2025, 17(7), 1091. [Google Scholar] [CrossRef]
- Rahimi-Farsi, N. Comprehensive systems biology analysis of microRNA-101-3p regulatory network identifies crucial genes and pathways in hepatocellular carcinoma. Journal of Genetic Engineering and Biotechnology 2025, 23(1), p. 100471. [Google Scholar] [CrossRef]
- Mature hsa-miR-101-3p. 2025. Available online: https://mirbase.org/mature/MIMAT0000099?utm_source=chatgpt.com.
- Zeng, X. CEBPα/miR-101b-3p promotes meningoencephalitis in mice infected with Angiostrongylus cantonensis by promoting microglial pyroptosis. Cell Communication and Signaling 2023, 21(1), p. 31. [Google Scholar] [CrossRef]
- MIR101-1 microRNA 101-1 [ Homo sapiens (human) ]. 2025. Available online: https://www.ncbi.nlm.nih.gov/gene?Cmd=DetailsSearch&Db=gene&Term=406893.
- RefSeq Gene MIR101-2. 2025.
- Perez, G. The UCSC Genome Browser database: 2025 update. Nucleic Acids Res 2025, 53(D1), D1243–d1249. [Google Scholar] [CrossRef]
- Mir101a Gene Detail. 2025.
- Cao, W. miRNASNP-v4: a comprehensive database for miRNA-related SNPs across 17 species. Nucleic Acids Research 2024, 53(D1), D1066–D1074. [Google Scholar] [CrossRef]
- SiamiGorji, S. Effects of microRNAs polymorphism in cancer progression. Med J Islam Repub Iran 2020, 34, 3. [Google Scholar] [CrossRef] [PubMed]
- Chen, J. Genetic variations in the flanking regions of miR-101-2 are associated with increased risk of breast cancer. PLoS One 2014, 9(1), e86319. [Google Scholar] [CrossRef] [PubMed]
- Omariba, G. Genome-Wide Analysis of MicroRNA-related Single Nucleotide Polymorphisms (SNPs) in Mouse Genome. Sci Rep 2020, 10(1), p. 5789. [Google Scholar] [CrossRef]
- Miao, L. MicroRNA-101 polymorphisms and risk of head and neck squamous cell carcinoma in a Chinese population. Tumor Biology 2016, 37(3), 4169–4174. [Google Scholar] [CrossRef]
- Cao, P. MicroRNA-101 negatively regulates Ezh2 and its expression is modulated by androgen receptor and HIF-1α/HIF-1β. Molecular Cancer 2010, 9(1), p. 108. [Google Scholar] [CrossRef] [PubMed]
- Su, H. MicroRNA-101, down-regulated in hepatocellular carcinoma, promotes apoptosis and suppresses tumorigenicity. Cancer Res 2009, 69(3), 1135–42. [Google Scholar] [CrossRef]
- Lippi, G. MicroRNA-101 Regulates Multiple Developmental Programs to Constrain Excitation in Adult Neural Networks. Neuron 2016, 92(6), 1337–1351. [Google Scholar] [CrossRef]
- Kavakiotis, I. DIANA-miTED: a microRNA tissue expression database. Nucleic Acids Res 2022, 50(D1), D1055–d1061. [Google Scholar] [CrossRef]
- Panwar, B.; Omenn, G.S.; Guan, Y. miRmine: a database of human miRNA expression profiles. Bioinformatics 2017, 33(10), 1554–1560. [Google Scholar] [CrossRef]
- Su, H. MicroRNA-101, Down-regulated in Hepatocellular Carcinoma, Promotes Apoptosis and Suppresses Tumorigenicity. Cancer Research 2009, 69(3), 1135–1142. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T. A microRNA, miR-101a, controls mammary gland development by regulating cyclooxygenase-2 expression. Differentiation 2009, 77(2), 181–187. [Google Scholar] [CrossRef] [PubMed]
- Konno, Y. MicroRNA-101 targets EZH2, MCL-1 and FOS to suppress proliferation, invasion and stem cell-like phenotype of aggressive endometrial cancer cells. Oncotarget 2014, 5(15), 6049–62. [Google Scholar] [CrossRef]
- Wang, C.Z. MiR-101: a potential therapeutic target of cancers. Am J Transl Res 2018, 10(11), 3310–3321. [Google Scholar]
- Ochs, M.J.; Steinhilber, D.; Suess, B. MicroRNA Involved in Inflammation: Control of Eicosanoid Pathway. Front Pharmacol 2011, 2, 39. [Google Scholar] [CrossRef] [PubMed]
- Li, M. MicroRNA-101 is a potential prognostic indicator of laryngeal squamous cell carcinoma and modulates CDK8. Journal of Translational Medicine 2015, 13(1), p. 271. [Google Scholar] [CrossRef]
- Mishra, R.; Singh, S.K. HIV-1 Tat C Modulates Expression of miRNA-101 to Suppress VE-Cadherin in Human Brain Microvascular Endothelial Cells. The Journal of Neuroscience 2013, 33(14), 5992–6000. [Google Scholar] [CrossRef] [PubMed]
- Shukla, A. Hypoxia-Induced miR-101 Impairs Endothelial Barrier Integrity Through Altering VE-Cadherin and Claudin-5. Mol Neurobiol 2024, 61(3), 1807–1817. [Google Scholar] [CrossRef]
- Friedman, J.M. The Putative Tumor Suppressor microRNA-101 Modulates the Cancer Epigenome by Repressing the Polycomb Group Protein EZH2. Cancer Research 2009, 69(6), 2623–2629. [Google Scholar] [CrossRef]
- Camfield, S. Secrets of DNA-PKcs beyond DNA repair. npj Precision Oncology 2024, 8(1), 154. [Google Scholar] [CrossRef]
- Yan, D. Targeting DNA-PKcs and ATM with miR-101 sensitizes tumors to radiation. PLoS One 2010, 5(7), e11397. [Google Scholar] [CrossRef]
- Huang, Z.; Wu, X.; Li, J. miR-101 suppresses colon cancer cell migration through the regulation of EZH2. Rev Esp Enferm Dig 2021, 113(4), 255–260. [Google Scholar] [CrossRef]
- Man, X. DNMT3A and DNMT3B in Breast Tumorigenesis and Potential Therapy. Front Cell Dev Biol 2022, 10, 916725. [Google Scholar] [CrossRef] [PubMed]
- Ma, L. The Mechanism of DNA Methylation and miRNA in Breast Cancer. International Journal of Molecular Sciences 2023, 24(11), 9360. [Google Scholar] [CrossRef]
- Szczepanek, J.; Tretyn, A. MicroRNA-Mediated Regulation of Histone-Modifying Enzymes in Cancer: Mechanisms and Therapeutic Implications. Biomolecules 2023, 13(11). [Google Scholar] [CrossRef]
- Varambally, S. Genomic Loss of microRNA-101 Leads to Overexpression of Histone Methyltransferase EZH2 in Cancer. Science 2008, 322(5908), 1695–1699. [Google Scholar] [CrossRef]
- Zhu, L. MiR-101 inhibits cell proliferation and invasion of pancreatic cancer through targeting STMN1. Cancer Biomark 2018, 23(2), 301–309. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y. miR-101 inhibits autophagy and enhances cisplatin-induced apoptosis in hepatocellular carcinoma cells. Oncol Rep 2013, 29(5), 2019–24. [Google Scholar] [CrossRef]
- Sun, Q. MiR-101 sensitizes human nasopharyngeal carcinoma cells to radiation by targeting stathmin 1. Mol Med Rep 2015, 11(5), 3330–6. [Google Scholar] [CrossRef]
- Tonetti, F.R.; Eguileor, A.; Llorente, C. Goblet cells: guardians of gut immunity and their role in gastrointestinal diseases. eGastroenterology 2024, 2(3). [Google Scholar] [CrossRef] [PubMed]
- Stanforth, K.J. The MUC2 Gene Product: Polymerisation and Post-Secretory Organisation—Current Models. Polymers 2024, 16(12), 1663. [Google Scholar] [CrossRef]
- Schulz-Heddergott, R. Therapeutic Ablation of Gain-of-Function Mutant p53 in Colorectal Cancer Inhibits Stat3-Mediated Tumor Growth and Invasion. Cancer Cell 2018, 34(2), 298–314.e7. [Google Scholar] [CrossRef] [PubMed]
- Yu, H. Revisiting STAT3 signalling in cancer: new and unexpected biological functions. Nat Rev Cancer 2014, 14(11), 736–46. [Google Scholar] [CrossRef]
- Heichler, C. STAT3 activation through IL-6/IL-11 in cancer-associated fibroblasts promotes colorectal tumour development and correlates with poor prognosis. Gut 2020, 69(7), 1269–1282. [Google Scholar] [CrossRef]
- Patel, D.; Witt, S.N. Ethanolamine and Phosphatidylethanolamine: Partners in Health and Disease. In Oxid Med Cell Longev; 2017; p. 4829180. [Google Scholar]
- Sogame, R. Dietary Ethanolamine Plasmalogen from Ascidian Alleviates Chronic Hepatic Injury in Mice Treated with Continuous Acetaminophen. Applied Sciences 2025, 15(11), 5968. [Google Scholar] [CrossRef]
- Zhou, J. Ethanolamine enhances intestinal functions by altering gut microbiome and mucosal anti-stress capacity in weaned rats. British Journal of Nutrition 2018, 120(3), 241–249. [Google Scholar] [CrossRef] [PubMed]
- Everard, A. Intestinal epithelial N-acylphosphatidylethanolamine phospholipase D links dietary fat to metabolic adaptations in obesity and steatosis. Nat Commun 2019, 10(1), p. 457. [Google Scholar] [CrossRef] [PubMed]
- Elmihi, K.A. Female ethanolamine phosphate phospholyase knockout mice resisted high-fat diet-induced obesity with attenuated hepatic cholesterol deposition. Am J Physiol Gastrointest Liver Physiol 2025, 329(3), G390–g402. [Google Scholar] [CrossRef]
- Shi, Z. Dynamic regulation of gut microbiota in high-fat or normal-fed mice by dietary phosphatidylcholine and phosphatidylethanolamine. Food Bioscience 2025. 71, 107314. [Google Scholar] [CrossRef]
- Barnes, A.J. Ethanolamine metabolism through two genetically distinct loci enables Klebsiella pneumoniae to bypass nutritional competition in the gut. PLoS Pathog 2024, 20(5), e1012189. [Google Scholar] [CrossRef]
- Rowley, C.A.; Anderson, C.J.; Kendall, M.M. Ethanolamine Influences Human Commensal Escherichia coli Growth, Gene Expression, and Competition with Enterohemorrhagic E. coli O157:H7. mBio 2018, 9(5). [Google Scholar] [CrossRef]
- Thiennimitr, P. Intestinal inflammation allows Salmonella to use ethanolamine to compete with the microbiota. Proc Natl Acad Sci U S A 2011, 108(42), 17480–5. [Google Scholar] [CrossRef]
- Ormsby, M.J. Inflammation associated ethanolamine facilitates infection by Crohn’s disease-linked adherent-invasive Escherichia coli. EBioMedicine 2019, 43, 325–332. [Google Scholar] [CrossRef]
- Pacheco, A.R.; Sperandio, V. Enteric Pathogens Exploit the Microbiota-generated Nutritional Environment of the Gut. Microbiology Spectrum 2015, 3(3). [Google Scholar] [CrossRef]
- Lengfelder, I. Complex Bacterial Consortia Reprogram the Colitogenic Activity of Enterococcus faecalis in a Gnotobiotic Mouse Model of Chronic, Immune-Mediated Colitis. Front Immunol 2019, 10, 1420. [Google Scholar] [CrossRef]
- Garsin, D.A. Ethanolamine utilization in bacterial pathogens: roles and regulation. Nature Reviews Microbiology 2010, 8(4), 290–295. [Google Scholar] [CrossRef]
- Anderson, C.J. Ethanolamine Signaling Promotes Salmonella Niche Recognition and Adaptation during Infection. PLoS Pathog 2015, 11(11), e1005278. [Google Scholar]
- Fang, H. Postbiotic Impact on Host Metabolism and Immunity Provides Therapeutic Potential in Metabolic Disease. Endocrine Reviews 2024, 46(1), 60–79. [Google Scholar] [CrossRef]
- Breton, J.; Galmiche, M.; Déchelotte, P. Dysbiotic Gut Bacteria in Obesity: An Overview of the Metabolic Mechanisms and Therapeutic Perspectives of Next-Generation Probiotics. Microorganisms 2022, 10(2). [Google Scholar] [CrossRef]
- Zeng, M.Y.; Inohara, N.; Nuñez, G. Mechanisms of inflammation-driven bacterial dysbiosis in the gut. Mucosal Immunology 2017, 10(1), 18–26. [Google Scholar] [CrossRef]
- Mishra, S.P. New Horizons in Microbiota and Metabolic Health Research. The Journal of Clinical Endocrinology & Metabolism 2020, 106(2), e1052–e1059. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S. A Cascade of Microbiota-Leaky Gut-Inflammation- Is it a Key Player in Metabolic Disorders? Current Obesity Reports 2025, 14(1), p. 32. [Google Scholar] [CrossRef] [PubMed]
- Okumura, R.; Takeda, K. The role of the mucosal barrier system in maintaining gut symbiosis to prevent intestinal inflammation. Semin Immunopathol 2024, 47(1), p. 2. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X. Mucosal immune response in biology, disease prevention and treatment. Signal Transduction and Targeted Therapy 2025, 10(1), p. 7. [Google Scholar] [CrossRef]
- Song, C. Intestinal mucus components and secretion mechanisms: what we do and do not know. Experimental & Molecular Medicine 2023, 55(4), 681–691. [Google Scholar] [CrossRef]
- Mishra, S.P. Abnormalities in microbiota/butyrate/FFAR3 signaling in aging gut impair brain function. JCI Insight 2024, 9(3). [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.E.V.; Larsson, J.M.H.; Hansson, G.C. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host–microbial interactions. Proceedings of the National Academy of Sciences 2011, 108 (supplement_1), 4659–4665. [Google Scholar] [CrossRef]
- McKenna, L.B. MicroRNAs Control Intestinal Epithelial Differentiation, Architecture, and Barrier Function. Gastroenterology 2010, 139(5), 1654–1664.e1. [Google Scholar] [CrossRef]
- Bergstrom, K.S. Muc2 protects against lethal infectious colitis by disassociating pathogenic and commensal bacteria from the colonic mucosa. PLoS Pathog 2010, 6(5), e1000902. [Google Scholar] [CrossRef]
- Johansson, M.E.V. Bacteria penetrate the normally impenetrable inner colon mucus layer in both murine colitis models and patients with ulcerative colitis. Gut 2014, 63(2), 281–291. [Google Scholar] [CrossRef] [PubMed]
- van der Post, S. Structural weakening of the colonic mucus barrier is an early event in ulcerative colitis pathogenesis. Gut 2019, 68(12), 2142–2151. [Google Scholar] [CrossRef]
- Jiang, Z. Proteus mirabilis exacerbates ulcerative colitis by inhibiting mucin production. Front Microbiol 2025, 16, 1556953. [Google Scholar] [CrossRef]
- Guagliano, G. The hidden piece in inflammatory bowel diseases: The significance of intestinal mucus in mediating the bacteria-cells crosstalk. Materials Today Bio 2025. 33, 101937. [Google Scholar] [CrossRef] [PubMed]
- Genua, F. The Role of Gut Barrier Dysfunction and Microbiome Dysbiosis in Colorectal Cancer Development. Front Oncol 2021, 11, 626349. [Google Scholar] [CrossRef] [PubMed]
- Neurath, M.F.; Artis, D.; Becker, C. The intestinal barrier: a pivotal role in health, inflammation, and cancer. Lancet Gastroenterol Hepatol 2025, 10(6), 573–592. [Google Scholar] [CrossRef]
- Kearns, R. Gut modulation to regulate NF-κB in colorectal and gastric cancer therapy and inflammation. Cancer Immunol Immunother 2025, 74(8), p. 264. [Google Scholar] [CrossRef]
- Lei, L. Increased intestinal permeability and lipopolysaccharide contribute to swainsonine-induced systemic inflammation. Ecotoxicology and Environmental Safety 2024, 284, 116912. [Google Scholar] [CrossRef]
- He, R. Dysbiosis and extraintestinal cancers. Journal of Experimental & Clinical Cancer Research 2025, 44(1), p. 44. [Google Scholar] [CrossRef]
- Matar, A. Intestinal Barrier Impairment, Preservation, and Repair: An Update. Nutrients 2024, 16(20), 3494. [Google Scholar] [CrossRef]
- Neurath, M.F.; Artis, D.; Becker, C. The intestinal barrier: a pivotal role in health, inflammation, and cancer. The Lancet Gastroenterology & Hepatology 2025, 10(6), 573–592. [Google Scholar] [CrossRef] [PubMed]
- Dmytriv, T.R.; Storey, K.B.; Lushchak, V.I. Intestinal barrier permeability: the influence of gut microbiota, nutrition, and exercise. Front Physiol 2024, 15, 1380713. [Google Scholar] [CrossRef]
- Mazaheri-Tehrani, S. Serum lipopolysaccharide binding protein (LBP) and metabolic syndrome: a systematic review and meta-analysis. Diabetology & Metabolic Syndrome 2025, 17(1), p. 268. [Google Scholar] [CrossRef]
- Mazaheri-Tehrani, S. Serum lipopolysaccharide binding protein (LBP) and metabolic syndrome: a systematic review and meta-analysis. Diabetol Metab Syndr 2025, 17(1), p. 268. [Google Scholar] [CrossRef] [PubMed]
- Starbæk, S.M.R. Meta-inflammation and endotoxemia in a highly translational porcine model of diet-induced obesity. Lab Animal 2025. [Google Scholar] [CrossRef] [PubMed]
- Hu, J. Brd4 modulates metabolic endotoxemia-induced inflammation by regulating colonic macrophage infiltration in high-fat diet-fed mice. Communications Biology 2024, 7(1), p. 1708. [Google Scholar] [CrossRef]
- Dimeji, I.Y.; Ayodeji, A.S. Pharmacological modulation of the gut microbiota and endotoxemia: A next-generation approach to treating metabolic syndrome. ASPET Discovery 2025. 1, 100010. [Google Scholar] [CrossRef]
- Jacobson, R. Collagenolytic Enterococcus faecalis induces DDR1 signaling, proliferation and altered immune infiltrate in colorectal peritoneal metastases. Surgery Open Science 2025. 28, 65–72. [Google Scholar] [CrossRef]
- Fang, J. Slimy partners: the mucus barrier and gut microbiome in ulcerative colitis. Experimental & Molecular Medicine 2021, 53(5), 772–787. [Google Scholar] [CrossRef]
- Rath, E.; Haller, D. Intestinal epithelial cell metabolism at the interface of microbial dysbiosis and tissue injury. Mucosal Immunology 2022, 15(4), 595–604. [Google Scholar] [CrossRef]
- Mishra, P. From Dysbiosis to Diagnosis: The Role of Gut Microbiota in Breast Cancer Etiology and Management. Applied Microbiology 2025, 5(4), 134. [Google Scholar] [CrossRef]
- Mukherjee, T. The NF-κB signaling system in the immunopathogenesis of inflammatory bowel disease. Science Signaling 2024, 17(818), p. eadh1641. [Google Scholar] [CrossRef]
- Chen, L. Interactions between toll-like receptors signaling pathway and gut microbiota in host homeostasis. Immun Inflamm Dis 2024, 12(7), p. e1356. [Google Scholar] [CrossRef]
- Clarke, Thomas B. Invasive Bacterial Pathogens Exploit TLR-Mediated Downregulation of Tight Junction Components to Facilitate Translocation across the Epithelium. Cell Host & Microbe 2011, 9(5), 404–414. [Google Scholar] [CrossRef] [PubMed]
- Garavaglia, B. The role of gut microbiota, immune system, and autophagy in the pathogenesis of inflammatory bowel disease: Molecular mechanisms and therapeutic approaches. Aspects of Molecular Medicine 2024. 4, 100056. [Google Scholar] [CrossRef]
- Robles-Vera, I. Microbiota translocation following intestinal barrier disruption promotes Mincle-mediated training of myeloid progenitors in the bone marrow. Immunity 2025. 58, 2, 381–396.e9. [Google Scholar] [CrossRef]
- Lei, W. Gut microbiota-driven neuroinflammation in Alzheimer’s disease: from mechanisms to therapeutic opportunities. Front Immunol 2025, 16, 1582119. [Google Scholar] [CrossRef]
- Desind, S.Z. Long noncoding RNAs and their complex role in shaping and regulating arachidonic acid metabolism: Learning to love the (not-really) junk. Wiley Interdiscip Rev RNA 2023, p. e1828. [Google Scholar] [CrossRef]
- Hao, Y. Enforced expression of miR-101 inhibits prostate cancer cell growth by modulating the COX-2 pathway in vivo. Cancer Prev Res (Phila) 2011, 4(7), 1073–83. [Google Scholar] [CrossRef]
- Che, L. MicroRNA-101 inhibits cadmium-induced angiogenesis by targeting cyclooxygenase-2 in primary human umbilical vein endothelial cells. Biochemical Pharmacology 2021, 189, 114192. [Google Scholar] [CrossRef]
- Liu, N. MiR-101: An Important Regulator of Gene Expression and Tumor Ecosystem. Cancers 2022, 14(23), 5861. [Google Scholar] [CrossRef] [PubMed]
- Chandimali, N. Free radicals and their impact on health and antioxidant defenses: a review. Cell Death Discovery 2025, 11(1), p. 19. [Google Scholar] [CrossRef]
- Morris, G. Redox regulation of the immune response. Cellular & Molecular Immunology 2022, 19(10), 1079–1101. [Google Scholar] [CrossRef] [PubMed]
- Manoharan, R.R. ROS signaling in innate immunity via oxidative protein modifications. Front Immunol 2024, 15, 1359600. [Google Scholar] [CrossRef] [PubMed]
- Li, Q. Signaling pathways involved in colorectal cancer: pathogenesis and targeted therapy. Signal Transduction and Targeted Therapy 2024, 9(1), p. 266. [Google Scholar] [CrossRef]
- Shahgoli, V.K. Inflammatory bowel disease, colitis, and cancer: unmasking the chronic inflammation link. Int J Colorectal Dis 2024, 39(1), p. 173. [Google Scholar] [CrossRef]
- Farmanbar, A.; Kneller, R.; Firouzi, S. Mutational signatures reveal mutual exclusivity of homologous recombination and mismatch repair deficiencies in colorectal and stomach tumors. Scientific Data 2023, 10(1), p. 423. [Google Scholar] [CrossRef]
- Thatikonda, V. Comprehensive analysis of mutational signatures reveals distinct patterns and molecular processes across 27 pediatric cancers. Nature Cancer 2023, 4(2), 276–289. [Google Scholar] [CrossRef]
- Grzelakowska, A.; Kalyanaraman, B.; Zielonka, J. Small molecule probes for peroxynitrite detection. Redox Biochemistry and Chemistry 2024, 10, 100034. [Google Scholar] [CrossRef] [PubMed]
- Seiwert, N. Heme oxygenase 1 protects human colonocytes against ROS formation, oxidative DNA damage and cytotoxicity induced by heme iron, but not inorganic iron. Cell Death & Disease 2020, 11(9), p. 787. [Google Scholar] [CrossRef] [PubMed]
- Irrazabal, T. Limiting oxidative DNA damage reduces microbe-induced colitis-associated colorectal cancer. Nature Communications 2020, 11(1), p. 1802. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y. Inflammation in cancer: therapeutic opportunities from new insights. Molecular Cancer 2025, 24(1), p. 51. [Google Scholar] [CrossRef]
- Wen, Y. Chronic inflammation, cancer development and immunotherapy. Front Pharmacol 2022, 13, 1040163. [Google Scholar] [CrossRef]
- Tripathi, S.; Sharma, Y.; Kumar, D. Unveiling the link between chronic inflammation and cancer. Metabolism Open 2025, 25, 100347. [Google Scholar] [CrossRef]
- Zhao, H. Inflammation and tumor progression: signaling pathways and targeted intervention. Signal Transduction and Targeted Therapy 2021, 6(1), 263. [Google Scholar] [CrossRef]
- Greten, F.R.; Grivennikov, S.I. Inflammation and Cancer: Triggers, Mechanisms, and Consequences. Immunity 2019, 51(1), 27–41. [Google Scholar] [CrossRef]
- Chi, X. T-cell exhaustion and stemness in antitumor immunity: Characteristics, mechanisms, and implications. Front Immunol 2023, 14, 1104771. [Google Scholar] [CrossRef]
- Voshagh, Q. Investigating the association between the tissue expression of miRNA-101, JAK2/STAT3 with TNF-α, IL-6, IL-1β, and IL-10 cytokines in the ulcerative colitis patients. Immun Inflamm Dis 2024, 12(3), p. e1224. [Google Scholar] [CrossRef]
- Hu, Y.; Dong, Z.; Liu, K. Unraveling the complexity of STAT3 in cancer: molecular understanding and drug discovery. Journal of Experimental & Clinical Cancer Research 2024, 43(1), p. 23. [Google Scholar] [CrossRef]
- Rahbar Farzam, O. Interplay of miRNAs and lncRNAs in STAT3 signaling pathway in colorectal cancer progression. Cancer Cell International 2024, 24(1), p. 16. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Augenlicht, L.H. Intestinal stem cells: guardians of homeostasis in health and aging amid environmental challenges. Experimental & Molecular Medicine 2024, 56(3), 495–500. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z. Oxidative stress caused by 3-monochloro-1,2-propanediol provokes intestinal stem cell hyperproliferation and the protective role of quercetin. Ecotoxicology and Environmental Safety 2025. 291, 117851. [Google Scholar] [CrossRef]
- Good, H.J. Prostaglandin E(2) and Akt Promote Stemness in Apc Mutant Dclk1+ Cells to Give Rise to Colitis-associated Cancer. Cell Mol Gastroenterol Hepatol 2025, 19(6), p. 101469. [Google Scholar] [CrossRef]
- Cineus, R. The IL-22–oncostatin M axis promotes intestinal inflammation and tumorigenesis. Nature Immunology 2025, 26(6), 837–853. [Google Scholar] [CrossRef] [PubMed]
- Morrison, H.A. NF-κB Inducing Kinase Attenuates Colorectal Cancer by Regulating Noncanonical NF-κB Mediated Colonic Epithelial Cell Regeneration. Cellular and Molecular Gastroenterology and Hepatology 2024, 18(3). [Google Scholar] [CrossRef]
- Song, P. Wnt/β-catenin signaling pathway in carcinogenesis and cancer therapy. Journal of Hematology & Oncology 2024, 17(1), p. 46. [Google Scholar]
- Deng, B. COX2 Enhances Neovascularization of Inflammatory Tenocytes Through the HIF-1α/VEGFA/PDGFB Pathway. Front Cell Dev Biol 2021, 9, 670406. [Google Scholar] [CrossRef]
- Geindreau, M.; Bruchard, M.; Vegran, F. Role of Cytokines and Chemokines in Angiogenesis in a Tumor Context. Cancers (Basel) 2022, 14(10). [Google Scholar] [CrossRef]
- Jacobson, R.A. Enterococcus faecalis exploits the human fibrinolytic system to drive excess collagenolysis: implications in gut healing and identification of druggable targets. Am J Physiol Gastrointest Liver Physiol 2020, 318(1), G1–g9. [Google Scholar] [CrossRef]
- Strillacci, A. MiR-101 downregulation is involved in cyclooxygenase-2 overexpression in human colon cancer cells. Exp Cell Res 2009, 315(8), 1439–47. [Google Scholar] [CrossRef]
- Schaefer, J.S. MicroRNA signatures differentiate Crohn’s disease from ulcerative colitis. BMC Immunology 2015, 16(1), 5. [Google Scholar] [CrossRef] [PubMed]
- Chandramouli, A. MicroRNA-101 (miR-101) post-transcriptionally regulates the expression of EP4 receptor in colon cancers. Cancer Biol Ther 2012, 13(3), 175–83. [Google Scholar] [CrossRef] [PubMed]
- Umansky, V. The Role of Myeloid-Derived Suppressor Cells (MDSC) in Cancer Progression. Vaccines (Basel) 2016, 4(4). [Google Scholar] [CrossRef]
- Liu, J. [miR-101 inhibits the proliferation and migration of breast cancer cells via downregulating the expression of DNA methyltransferase 3a]. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 2016, 32(3), 299–303. [Google Scholar]
- Wang, H. MiR-101 Targets the EZH2/Wnt/β-Catenin the Pathway to Promote the Osteogenic Differentiation of Human Bone Marrow-Derived Mesenchymal Stem Cells. Sci Rep 2016, 6, 36988. [Google Scholar] [CrossRef]
- Vella, S. MicroRNA-101 is repressed by EZH2 and its restoration inhibits tumorigenic features in embryonal rhabdomyosarcoma. Clin Epigenetics 2015, 7(1), p. 82. [Google Scholar] [CrossRef]
- Smits, M. miR-101 is down-regulated in glioblastoma resulting in EZH2-induced proliferation, migration, and angiogenesis. Oncotarget 2010, 1(8), 710–20. [Google Scholar] [CrossRef]
- Friedman, J.M. The putative tumor suppressor microRNA-101 modulates the cancer epigenome by repressing the polycomb group protein EZH2. Cancer Res 2009, 69(6), 2623–9. [Google Scholar] [CrossRef]
- Varambally, S. Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science 2008, 322(5908), 1695–9. [Google Scholar] [CrossRef]
- Chakrabarty, A. MicroRNA regulation of cyclooxygenase-2 during embryo implantation. Proc Natl Acad Sci U S A 2007, 104(38), 15144–9. [Google Scholar] [CrossRef]
- Tanaka, T. A microRNA, miR-101a, controls mammary gland development by regulating cyclooxygenase-2 expression. Differentiation 2009, 77(2), 181–7. [Google Scholar] [CrossRef]
- Liu, Y. MicroRNA-101 inhibits angiogenesis via COX-2 in endometrial carcinoma. Mol Cell Biochem 2018, 448(1-2), 61–69. [Google Scholar] [CrossRef] [PubMed]
- Che, L. MicroRNA-101 inhibits cadmium-induced angiogenesis by targeting cyclooxygenase-2 in primary human umbilical vein endothelial cells. Biochem Pharmacol 2021, 189, 114192. [Google Scholar] [CrossRef] [PubMed]
- Harati, R. Loss of miR-101-3p Promotes Transmigration of Metastatic Breast Cancer Cells through the Brain Endothelium by Inducing COX-2/MMP1 Signaling. Pharmaceuticals (Basel) 2020, 13(7). [Google Scholar] [CrossRef] [PubMed]
- Cai, Y. Eicosapentaenoic acid’s metabolism of 15-LOX-1 promotes the expression of miR-101 thus inhibits Cox2 pathway in colon cancer. Onco Targets Ther 2020, 13, 5605–5616. [Google Scholar] [CrossRef]
- Fan, M.J. Long non-coding RNA SPRY4-IT1 promotes epithelial-mesenchymal transition of cervical cancer by regulating the miR-101-3p/ZEB1 axis. Biosci Rep 2019, 39(6). [Google Scholar] [CrossRef]
- Xiong, W.C. Interplay between long noncoding RNA ZEB1-AS1 and miR-101/ZEB1 axis regulates proliferation and migration of colorectal cancer cells. Am J Transl Res 2018, 10(2), 605–617. [Google Scholar]
- Liang, H. LncRNA PTAR promotes EMT and invasion-metastasis in serous ovarian cancer by competitively binding miR-101-3p to regulate ZEB1 expression. Mol Cancer 2018, 17(1), p. 119. [Google Scholar] [CrossRef]
- Yin, L.; Wang, Y. Extracellular vesicles derived from M2-polarized tumor-associated macrophages promote immune escape in ovarian cancer through NEAT1/miR-101-3p/ZEB1/PD-L1 axis. Cancer Immunol Immunother 2023, 72(3), 743–758. [Google Scholar] [CrossRef]
- Shahverdi, M. Knockdown of Myeloid Cell Leukemia-1 by MicroRNA-101 Increases Sensitivity of A549 Lung Cancer Cells to Etoposide. Iran J Med Sci 2021, 46(4), 298–307. [Google Scholar]
- Wang, H. MALAT1/miR-101-3p/MCL1 axis mediates cisplatin resistance in lung cancer. Oncotarget 2018, 9(7), 7501–7512. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Li, Z. Overexpression of miR-101 promotes TRAIL-induced mitochondrial apoptosis in papillary thyroid carcinoma by targeting c-met and MCL-1. Oncotarget 2017, 8(65), 108665–108675. [Google Scholar] [CrossRef] [PubMed]
- Shen, W. MicroRNA-101-5p inhibits the growth and metastasis of cervical cancer cell by inhibiting CXCL6. Eur Rev Med Pharmacol Sci 2019, 23(5), 1957–1968. [Google Scholar] [PubMed]
- Chen, Q. miRNA-101-5p inhibits the growth and aggressiveness of NSCLC cells through targeting CXCL6. Onco Targets Ther 2019, 12, 835–848. [Google Scholar] [CrossRef]
- Chen, F. MicroRNA-101 Targets CXCL12-Mediated Akt and Snail Signaling Pathways to Inhibit Cellular Proliferation and Invasion in Papillary Thyroid Carcinoma. Oncol Res 2019, 27(6), 691–701. [Google Scholar] [CrossRef]
- Chen, H. Circular RNA hsa_circ_0007364 increases cervical cancer progression through activating methionine adenosyltransferase II alpha (MAT2A) expression by restraining microRNA-101-5p. Bioengineered 2020, 11(1), 1269–1279. [Google Scholar] [CrossRef]
- Wang, H.; Xiao, R.; Yang, B. MiR-101-3p Suppresses Progression of Cervical Squamous Cell Carcinoma by Targeting and Down-Regulating KPNA2. Technol Cancer Res Treat 2021, 20, p. 15330338211055948. [Google Scholar] [CrossRef]
- Li, Z. miR-101-3p sensitizes non-small cell lung cancer cells to irradiation. Open Med (Wars) 2020, 15(1), 413–423. [Google Scholar] [CrossRef]
- Chen, M. Long Noncoding RNA FAM201A Mediates the Radiosensitivity of Esophageal Squamous Cell Cancer by Regulating ATM and mTOR Expression via miR-101. Front Genet 2018, 9, 611. [Google Scholar] [CrossRef]
- Shao, N.; Song, L.; Sun, X. Exosomal circ_PIP5K1A regulates the progression of non-small cell lung cancer and cisplatin sensitivity by miR-101/ABCC1 axis. Mol Cell Biochem 2021, 476(6), 2253–2267. [Google Scholar] [CrossRef]
- Xie, F.W.; Liu, J.C. LncRNA SNHG12 regulates the miR-101-3p/CUL4B axis to mediate the proliferation, migration and invasion of non-small cell lung cancer. Kaohsiung J Med Sci 2021, 37(8), 664–674. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H. Circular RNA ZFR accelerates non-small cell lung cancer progression by acting as a miR-101-3p sponge to enhance CUL4B expression. Artif Cells Nanomed Biotechnol 2019, 47(1), 3410–3416. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z. Inhibition of MicroRNA miR-101-3p on prostate cancer progression by regulating Cullin 4B (CUL4B) and PI3K/AKT/mTOR signaling pathways. Bioengineered 2021, 12(1), 4719–4735. [Google Scholar] [CrossRef] [PubMed]
- Li, K. Long non-coding RNA SNHG6 promotes the growth and invasion of non-small cell lung cancer by downregulating miR-101-3p. Thorac Cancer 2020, 11(5), 1180–1190. [Google Scholar] [CrossRef] [PubMed]
- Ding, C. Exosomal Circ-MEMO1 Promotes the Progression and Aerobic Glycolysis of Non-small Cell Lung Cancer Through Targeting MiR-101-3p/KRAS Axis. Front Genet 2020, 11, 962. [Google Scholar] [CrossRef] [PubMed]
- Li, L. MiR-101-3p inhibits EMT to attenuate proliferation and metastasis in glioblastoma by targeting TRIM44. J Neurooncol 2019, 141(1), 19–30. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.H. Long noncoding RNA CRNDE promotes proliferation, migration and invasion in prostate cancer through miR-101/Rap1A. Neoplasma 2020, 67(3), 584–594. [Google Scholar] [CrossRef]
- Zhou, Z. MicroRNA-101 suppresses colorectal cancer progression by negative regulation of Rap1b. Oncol Lett 2020, 20(3), 2225–2231. [Google Scholar] [CrossRef]
- Jin, Q. MicroRNA-101-3p inhibits proliferation in retinoblastoma cells by targeting EZH2 and HDAC9. Exp Ther Med 2018, 16(3), 1663–1670. [Google Scholar] [CrossRef]
- Cao, K. miR-101 Inhibiting Cell Proliferation, Migration and Invasion in Hepatocellular Carcinoma through Downregulating Girdin. Mol Cells 2016, 39(2), 96–102. [Google Scholar] [CrossRef]
- Liu, Z. MicroRNA-101 suppresses migration and invasion via targeting vascular endothelial growth factor-C in hepatocellular carcinoma cells. Oncol Lett 2016, 11(1), 433–438. [Google Scholar] [CrossRef]
- Liu, P. LncRNA-MALAT1 mediates cisplatin resistance via miR-101-3p/VEGF-C pathway in bladder cancer. Acta Biochim Biophys Sin (Shanghai) 2019, 51(11), 1148–1157. [Google Scholar] [CrossRef]
- Calastri, M.C.J. Investigating VEGF. miR-145-3p, and miR-101-3p Expression in Patients with Cholangiocarcinoma. Asian Pac J Cancer Prev 2022, 23(7), 2233–2241. [Google Scholar] [CrossRef]
- Guo, X. Cancer-Associated Fibroblasts Promote Migration and Invasion of Non-Small Cell Lung Cancer Cells via miR-101-3p Mediated VEGFA Secretion and AKT/eNOS Pathway. Front Cell Dev Biol 2021, 9, 764151. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J. miR-101 inhibits cholangiocarcinoma angiogenesis through targeting vascular endothelial growth factor (VEGF). Am J Pathol 2013, 182(5), 1629–39. [Google Scholar] [CrossRef]
- Du, Y.L. LncRNA XIST Promotes Migration and Invasion of Papillary Thyroid Cancer Cell by Modulating MiR-101-3p/CLDN1 Axis. Biochem Genet 2021, 59(2), 437–452. [Google Scholar] [CrossRef] [PubMed]
- Si, W. The coordination between ZNF217 and LSD1 contributes to hepatocellular carcinoma progress and is negatively regulated by miR-101. Exp Cell Res 2019, 379(1), 1–10. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y. MicroRNA-101-3p suppresses proliferation and migration in hepatocellular carcinoma by targeting the HGF/c-Met pathway. Invest New Drugs 2020, 38(1), 60–69. [Google Scholar] [CrossRef]
- Meng, X. Influence of miR-101 on proliferation of liver cancer cells through the MAPK/ERK signaling pathway. Oncol Lett 2020, 19(2), 1310–1316. [Google Scholar] [CrossRef]
- Saika, R. MicroRNA-101a regulates microglial morphology and inflammation. Journal of Neuroinflammation 2017, 14(1), p. 109. [Google Scholar] [CrossRef]
- Wang, L. IL-1β-Mediated Repression of microRNA-101 Is Crucial for Inflammation-Promoted Lung Tumorigenesis. Cancer Research 2014, 74(17), 4720–4730. [Google Scholar] [CrossRef]
- Ma, S. A novel regulatory loop miR-101/ANXA2/EGR1 mediates malignant characteristics of liver cancer stem cells. Carcinogenesis 2021, 42(1), 93–104. [Google Scholar] [CrossRef]
- Wu, F. MicroRNA-101-3p regulates gastric cancer cell proliferation, invasion and apoptosis by targeting PIM 1 expression. Cell Mol Biol (Noisy-le-grand) 2019, 65(7), 118–122. [Google Scholar] [CrossRef]
- Chen, L. MicroRNA-101 inhibits cell migration and invasion in bladder cancer via targeting FZD4. Exp Ther Med 2019, 17(2), 1476–1485. [Google Scholar] [CrossRef] [PubMed]
- Tao, L. HIPK3 Inhibition by Exosomal hsa-miR-101-3p Is Related to Metabolic Reprogramming in Colorectal Cancer. Front Oncol 2021, 11, 758336. [Google Scholar] [CrossRef]
- Wu, H.B. CircAPLP2 regulates the proliferation and metastasis of colorectal cancer by targeting miR-101-3p to activate the Notch signalling pathway. Am J Transl Res 2020, 12(6), 2554–2569. [Google Scholar]
- Yang, Q.; Yu, W.; Han, X. Overexpression of microRNA-101 causes anti-tumor effects by targeting CREB1 in colon cancer. Mol Med Rep 2019, 19(4), 3159–3167. [Google Scholar] [CrossRef] [PubMed]
- Wang, H. miR-101-3p and miR-199b-5p promote cell apoptosis in oral cancer by targeting BICC1. Mol Cell Probes 2020, 52, 101567. [Google Scholar] [CrossRef] [PubMed]
- Wang, L. miR-101 promotes breast cancer cell apoptosis by targeting Janus kinase 2. Cell Physiol Biochem 2014, 34(2), 413–22. [Google Scholar] [CrossRef] [PubMed]
| Gene/Protein | Function | Ref. |
| E-cadherin | Overexpressed miR-101 suppresses DNMT3A, thereby restoring E-cadherin and inhibiting proliferation and migration in MDA-MB-231 breast cancer cells | [177] |
| EZH2 | Overexpression of miR-101a-3p in the amygdala increases anxiety-like behavior in rats by repressing Ezh2 | [14] |
| miR-101 promotes osteogenic differentiation of hBMSCs by targeting EZH2 and activating the Wnt/β-catenin pathway | [178] | |
| miR-101 is downregulated in eRMS, inversely correlated with EZH2, and its re-expression suppresses EZH2, reducing migration, clonogenicity, and tumorigenic potential of eRMS cells | [179] | |
| miR-101 suppresses Ezh2, reducing invasion and migration of prostate cancer cells, with its expression further modulated by androgen signaling and HIF-1α/β induction | [23] | |
| miR-101 downregulation drives EZH2 overexpression in GBM, and inhibiting EZH2 suppresses tumor growth, invasion, and angiogenesis both in vitro and in vivo | [180] | |
| Re-expression of miR-101 suppresses proliferation, invasion, and self-renewal of aggressive endometrial cancer cells by directly targeting EZH2, MCL-1, and FOS | [66] | |
| miR-101 is downregulated in bladder TCC, and its restoration suppresses proliferation by directly repressing EZH2, highlighting its role as a tumor suppressor. | [181] | |
| miR-101 is downregulated in CRC, and its restoration suppresses EZH2-driven migration of colorectal cancer cells. | [75] | |
| Genomic loss of miR-101 in prostate cancer reduces its repression of EZH2, leading to EZH2 overexpression and epigenetic dysregulation that drives tumor progression. | [182] | |
| PTGS2/COX2 | miR-101a directly repress COX-2 translation, with precursor overexpression reducing and antisense inhibition restoring COX-2 reporter and protein levels | [183] |
| miR-101a upregulation during mammary gland development suppresses COX-2, thereby inhibiting proliferation and modulating differentiation of mammary epithelial cells. | [184] | |
| miR-101 is downregulated in endometrial cancer, and its restoration suppresses angiogenesis and tumor growth partly through COX-2 regulation. | [185] | |
| Exogenous miR-101 directly targets COX-2, suppressing prostate cancer cell proliferation and reducing tumor growth in vitro and in vivo. | [142] | |
| miR-101 directly inhibits COX-2 translation, and its downregulation correlates with COX-2 overexpression in colorectal cancer cells and patient tissues | [173] | |
| CdCl₂ induces ER stress–driven COX-2/VEGF upregulation causing abnormal angiogenesis and cytotoxicity, which can be rescued by miR-101, siPTGS2, or COX-2 inhibition | [186] | |
| miR-101-3p suppresses breast cancer cell transmigration across the brain endothelium by downregulating COX-2/MMP1 signaling, thereby preserving junctional integrity and reducing brain metastasis potential | [187] | |
| EPA upregulates miR-101 via the 15-LOX-1 pathway, leading to COX-2 suppression and inhibiting colon cancer progression. | [188] | |
| ZEB1 | miR-101a directly targets and suppresses ZEB1, thereby inhibiting epithelial–mesenchymal transition (EMT), reducing invasion and migration, and functioning as a tumor-suppressive regulator. | [189] |
| ZEB1-AS1 promotes CRC proliferation and migration by sponging miR-101 to upregulate ZEB1, while miR-101 restoration or ZEB1-AS1 depletion suppresses these effects. | [190] | |
| miR-101 directly suppresses ZEB1 expression, thereby inhibiting epithelial-to-mesenchymal transition (EMT), reducing cell migration, and limiting metastatic potential. | [191] | |
| M2-TAM–derived EVs deliver NEAT1, which sponges miR-101-3p to upregulate ZEB1/PD-L1, thereby promoting ovarian cancer growth and inducing CD8⁺ T-cell apoptosis. | [192] | |
| MCL-1/2 | miR-101 suppresses Mcl-1, inhibits A549 lung cancer cell growth, and synergistically enhances sensitivity to etoposide by promoting apoptosis. | [193] |
| miR-101 suppresses MCL-1 by targeting its 3′-UTR, thereby promoting apoptosis and enhancing chemosensitivity. | [194] | |
| Overexpression of miR-101 sensitizes papillary thyroid carcinoma cells to TRAIL-induced apoptosis by targeting c-Met and MCL-1 and inhibiting the PI3K/AKT pathway. | [195] | |
| CXCL6 | Overexpression of miR-101-5p suppresses cervical cancer proliferation, migration, invasion, and tumor growth by directly targeting CXCL6 | [196] |
| Overexpression of miR-101-5p suppresses NSCLC growth, invasion, and metastasis by directly targeting CXCL6. | [197] | |
| CXCL12 | miR-101 is downregulated in PTC and suppresses proliferation, survival, migration, and invasion by directly targeting CXCL12 and inhibiting its downstream Akt/EMT signaling | [198] |
| MAT2A | hsa_circ_0007364 drives cervical cancer progression by sponging miR-101-5p, thereby relieving suppression of MAT2A and enhancing tumor cell proliferation, invasion, and growth. | [199] |
| KPNA2 | miR-101-3p is downregulated in cervical squamous cell carcinoma, and its overexpression suppresses tumor cell growth by directly targeting KPNA2. | [200] |
| mTOR | miR-101-3p enhances radiosensitivity of NSCLC by inhibiting the mTOR signaling pathway, reducing survival and promoting apoptosis in irradiated cells. | [201] |
| lncRNA FAM201A decreases radiosensitivity in ESCC by suppressing miR-101a, which in turn upregulates mTOR (and ATM) signaling. | [202] | |
| ABCC1 | Exosomal circ_PIP5K1A promotes NSCLC progression and cisplatin resistance by sponging miR-101a, which normally suppresses ABCC1 expression to inhibit tumor growth and enhance chemosensitivity. | [203] |
| CUL4B | miR-101a directly targets CUL4B, and its suppression by lncRNA SNHG12 enhances CUL4B expression, thereby promoting proliferation, migration, and invasion in non-small cell lung cancer. | [204] |
| circZFR promotes NSCLC progression by sponging miR-101a-3p, thereby relieving its repression of CUL4B and enhancing proliferation, migration, and invasion | [205] | |
| miR-101a directly targets CUL4B, and its overexpression suppresses prostate cancer cell proliferation, migration, and invasion while promoting apoptosis by inhibiting the PI3K/AKT/mTOR pathway | [206] | |
| CDYL | SNHG6 promotes NSCLC growth and invasion by suppressing miR-101a-3p, thereby relieving its inhibition on CDYL. | [207] |
| KRAS | miR-101a directly targets the KRAS 3′UTR, and its repression by circ-MEMO1 leads to KRAS upregulation, thereby promoting proliferation, glycolysis, and tumor growth in NSCLC. | [208] |
| TRIM44 | miR-101-3p directly targets TRIM44, suppressing EMT and thereby reducing proliferation, migration, and invasion of glioblastoma cells. | [209] |
| RAP1A/B | miR-101a suppresses Rap1A expression in prostate cancer, an effect antagonized by lncRNA CRNDE, thereby restraining proliferation, migration, and invasion. | [210] |
| miR-101a suppresses colorectal cancer progression by directly targeting Rap1b, forming a negative feedback loop that inhibits proliferation, migration, and invasion. | [211] | |
| HDAC9 | miR-101a-3p suppresses retinoblastoma cell proliferation by directly targeting HDAC9, and restoring HDAC9 reverses this anti-proliferative effect. | [212] |
| Girdin | miR-101a acts as a tumor suppressor in HCC by directly targeting Girdin, thereby inhibiting cell proliferation, migration, and invasion. | [213] |
| VEGF-C | miR-101a suppresses HCC cell migration and invasion by directly targeting VEGF-C, highlighting its role in restraining cytoskeletal remodeling and girding during metastasis. | [214] |
| miR-101a suppresses VEGF-C, and its inhibition by MALAT1 promotes cisplatin resistance in bladder cancer cells. | [215] | |
| In cholangiocarcinoma, miR-101a is underexpressed, correlating with VEGF overexpression, suggesting its loss may promote tumor progression and reduced survival. | [216] | |
| miR-101a-3p suppresses VEGFA in cancer-associated fibroblasts, thereby reducing CAF-driven EMT, invasion, and metastasis in lung cancer. | [217] | |
| miR-101a suppresses cholangiocarcinoma growth by directly targeting VEGF and indirectly repressing its transcription via COX-2 inhibition, thereby blocking angiogenesis. | [218] | |
| ZO-1 | miR-101a-3p, upregulated by ethanolamine accumulation in obesity, destabilizes ZO-1 mRNA, thereby weakening intestinal barriers and promoting gut permeability, inflammation, and metabolic dysfunction. | [33] |
| CLDN1 | miR-101a suppresses CLDN1 expression to inhibit papillary thyroid carcinoma cell migration and invasion, while XIST promotes metastasis by sponging miR-101a and restoring CLDN1. | [219] |
| ZNF217 | miR-101 suppresses ZNF217 in hepatocellular carcinoma, thereby restoring CDH1 expression and inhibiting proliferation, EMT, and invasion. | [220] |
| c-Met pathway | miR-101a sensitizes papillary thyroid carcinoma cells to TRAIL by targeting c-Met (and MCL-1), thereby inhibiting PI3K/AKT signaling and reducing apoptosis resistance. | [195] |
| c-Met pathway MAPK/ERK pathway |
miR-101a suppresses tumor growth and progression by downregulating pro-angiogenic signaling, such as VEGF-related pathways, through targeting upstream regulators like the HGF/c-MET axis. | [221] |
| miR-101a is downregulated in liver cancer, and its overexpression suppresses proliferation by targeting EZH2 and inhibiting the MAPK/ERK signaling pathway | [222] | |
| miR-101a, enriched in the brain, promotes differentiation of bone marrow cells into microglia-like cells by enhancing inflammatory responsiveness and downregulating MAPK phosphatase-1. | [223] | |
| Lin28B | In NSCLC, IL-1β suppresses miR-101, leading to Lin28B upregulation and enhanced proliferation and migration, an effect reversible by COX-2 inhibition. | [224] |
| ANXA2 | miR-101a suppresses ANXA2, thereby inhibiting ERK signaling, reducing LCSC proliferation and metastasis, and forming a regulatory miR-101/ANXA2/EGR1 loop in liver cancer. | [225] |
| SRF | miR-101a suppresses gastric adenocarcinoma cell proliferation and invasion by directly targeting SRF, thereby downregulating HOTAIR transcription. | [225] |
| PIM 1 | miR-101a suppresses PIM1 expression in gastric cancer cells, thereby inhibiting proliferation and invasion while promoting apoptosis. | [226] |
| FZD4 | miR-101a is downregulated in bladder cancer, and its restoration suppresses migration and invasion by directly targeting FZD4. | [227] |
| STMN1 | miR-101a suppresses pancreatic cancer cell proliferation and invasion by directly targeting and downregulating STMN1. | [80] |
| HIPK3 | miR-101a-3p acts as an oncomiR in colorectal cancer by targeting HIPK3, thereby enhancing cell growth, migration, glycolysis, and reducing chemosensitivity. | [228] |
| Notch1 | miR-101a suppresses NOTCH1 signaling, while CircAPLP2 promotes colorectal cancer proliferation and metastasis by sponging miR-101a and thereby reactivating the Notch pathway. | [229] |
| CREB1 | miR-101a is downregulated in colon cancer, and its overexpression suppresses proliferation and migration by directly targeting CREB1. | [230] |
| BICC1 | miR-101a is downregulated in oral cancer, and its restoration suppresses BICC1, thereby reducing cell viability and promoting apoptosis. | [231] |
| Jak2 | miR-101a suppresses proliferation and induces apoptosis in breast cancer cells by directly targeting Jak2. | [232] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




