Submitted:
21 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Screening with Molecular Docking
2.2. Molecular Dynamics Simulation
2.3. Materials and Reagents
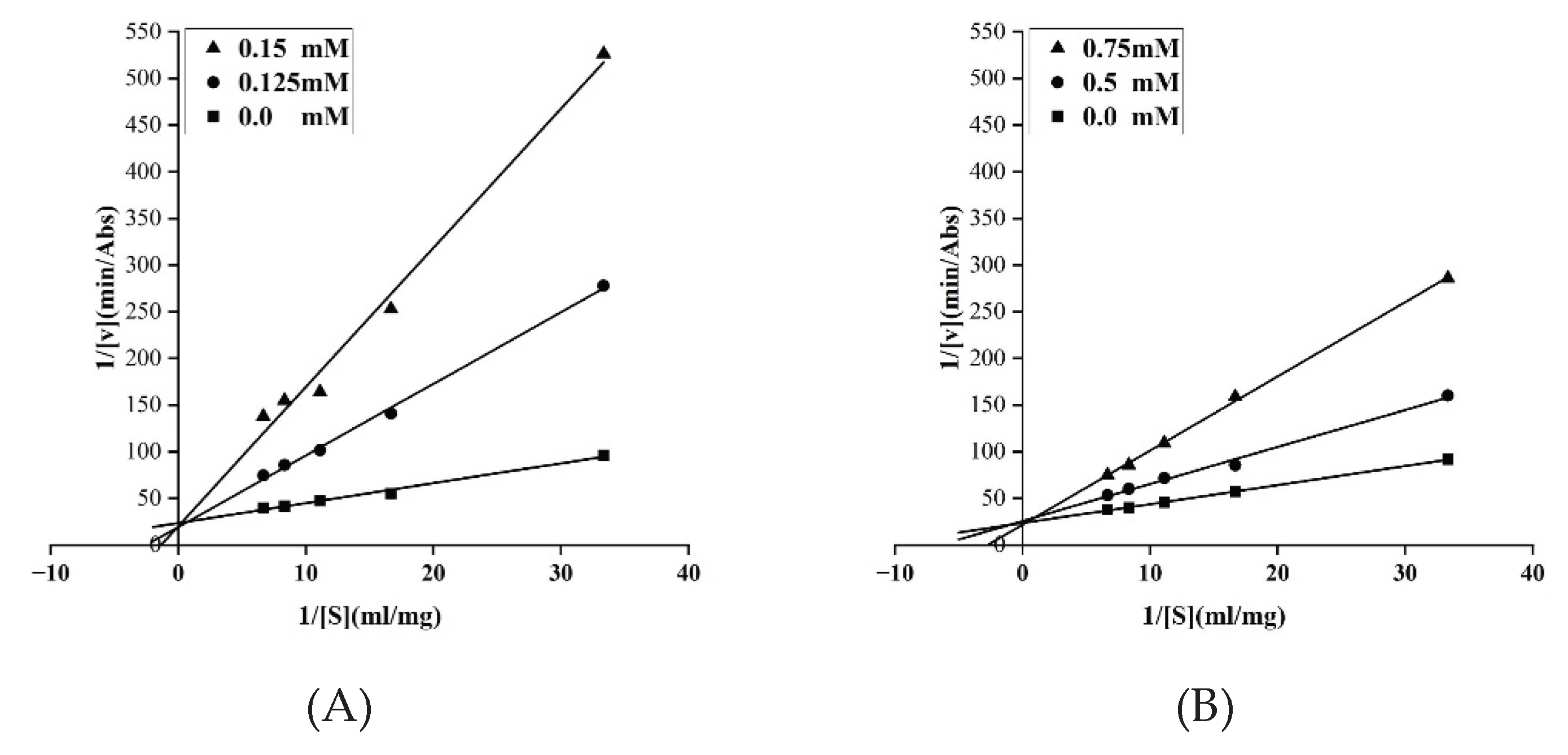
2.4. Inhibition Rate and Inhibition Type on Pancreatic Lipase
2.5. Statistic Analysis
3. Results
3.1. Molecular Docking
3.2. Molecular Dynamics Simulation
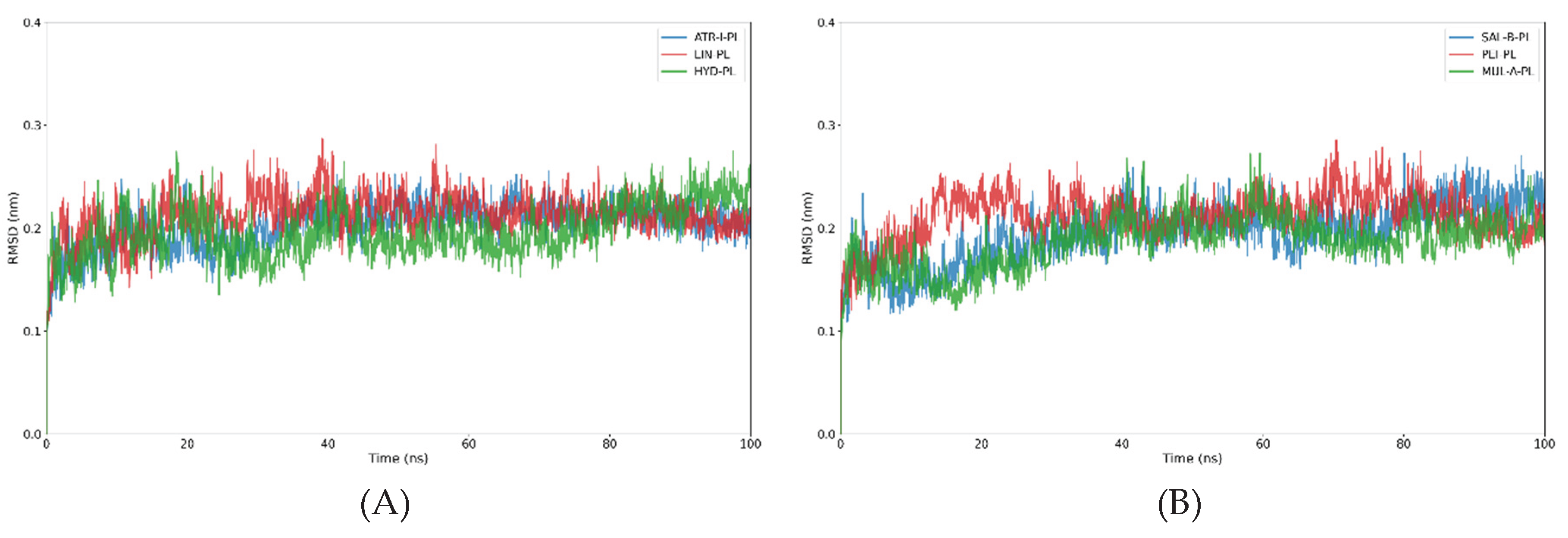
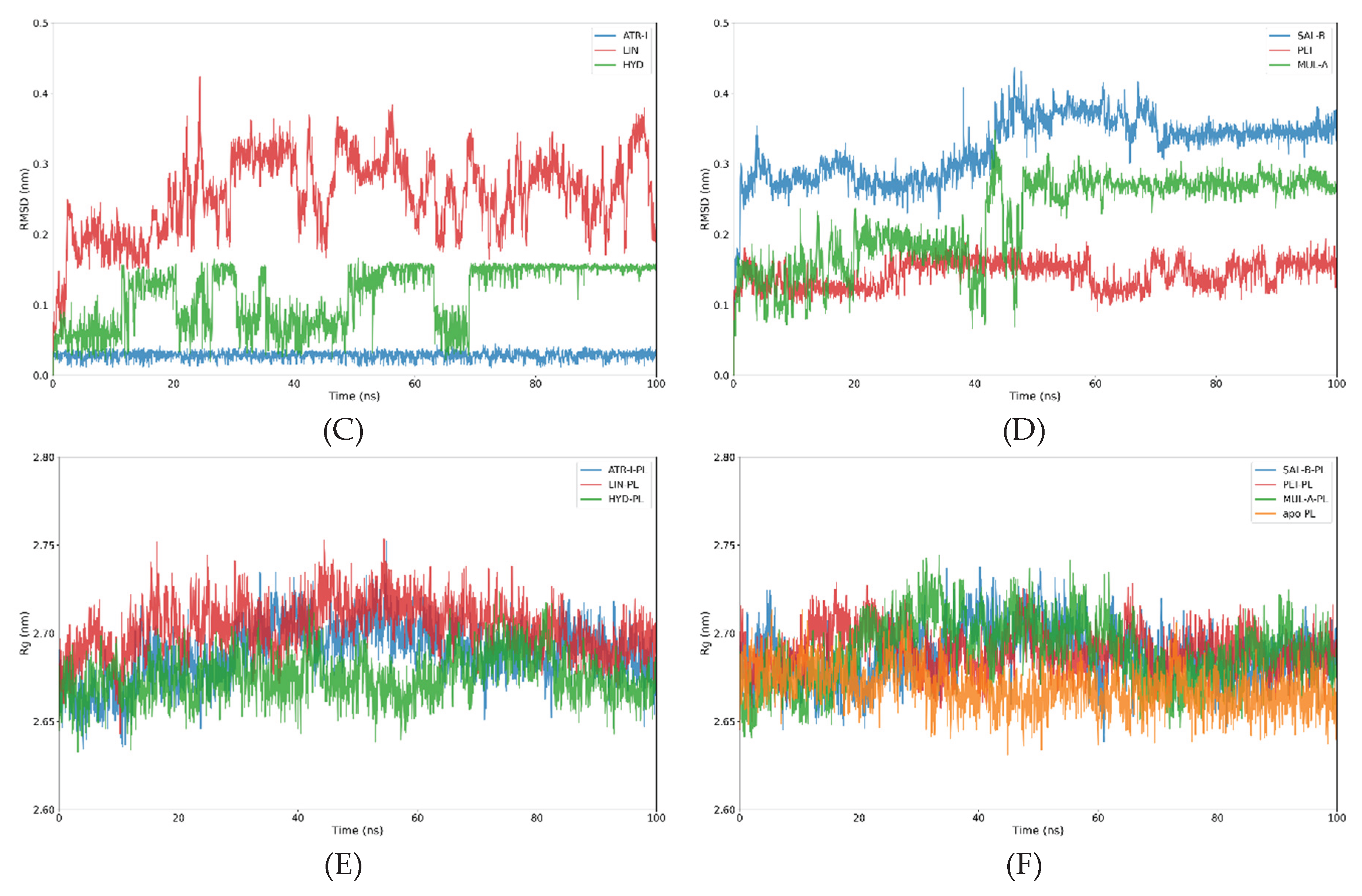
3.2.1. Analysis of the RMSD and Rg
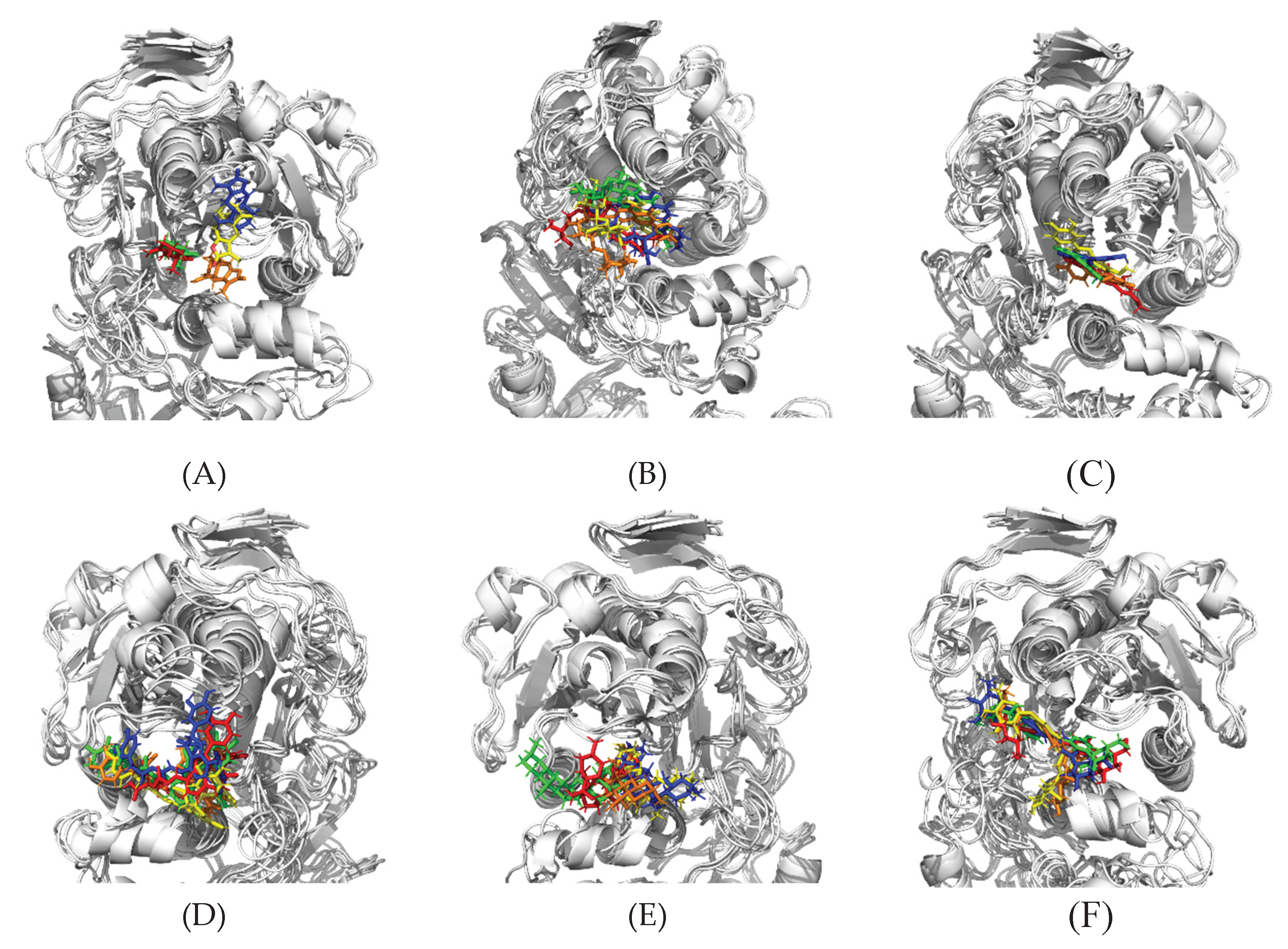
3.2.2. Binding Mode Variation During the Simulation
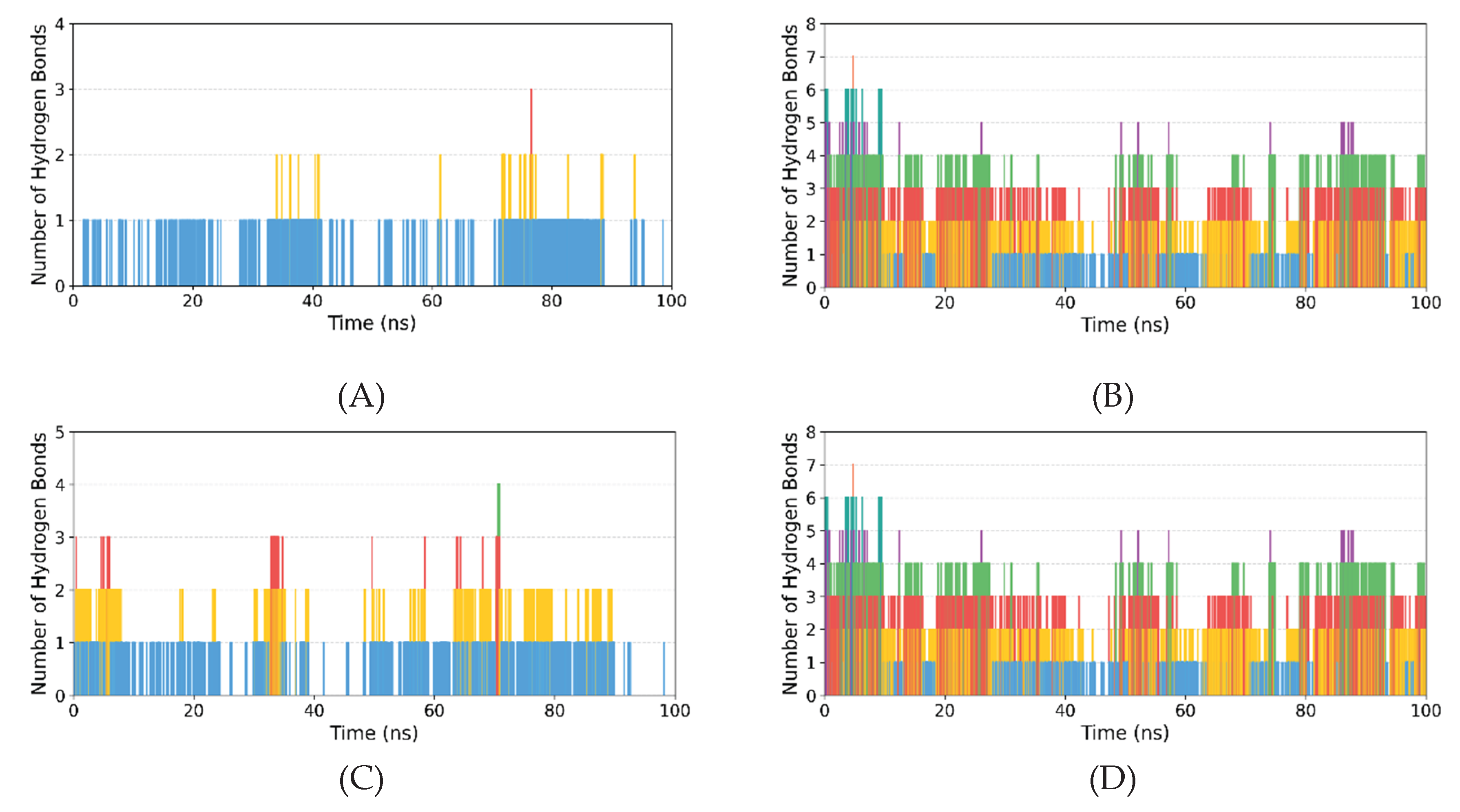
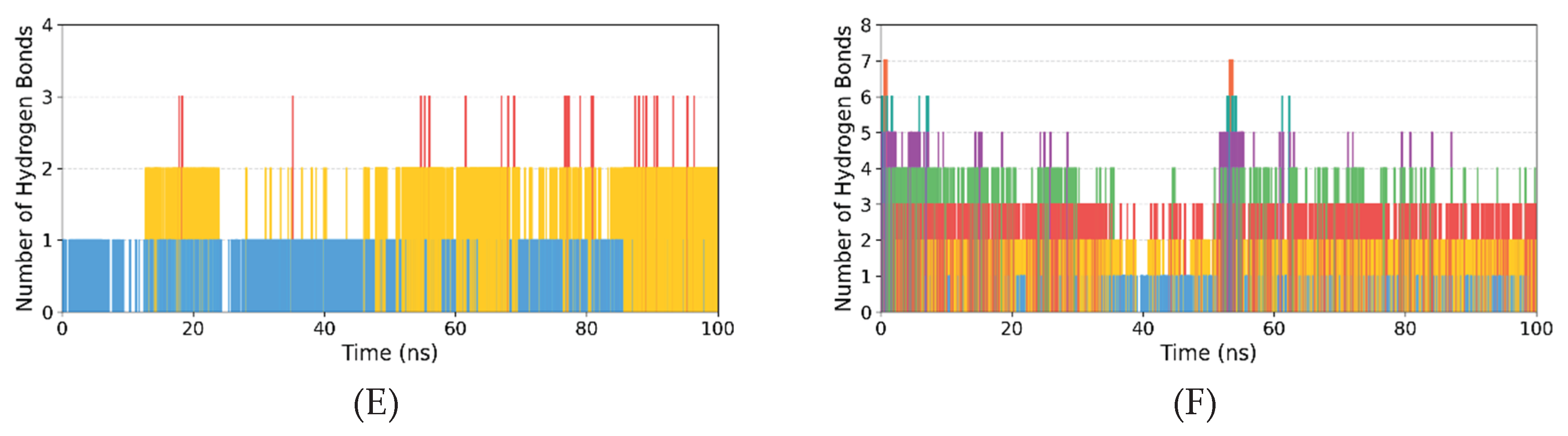
3.2.3. Hydrogen Bond Analysis
3.2.4. MM/PBSA Calculation
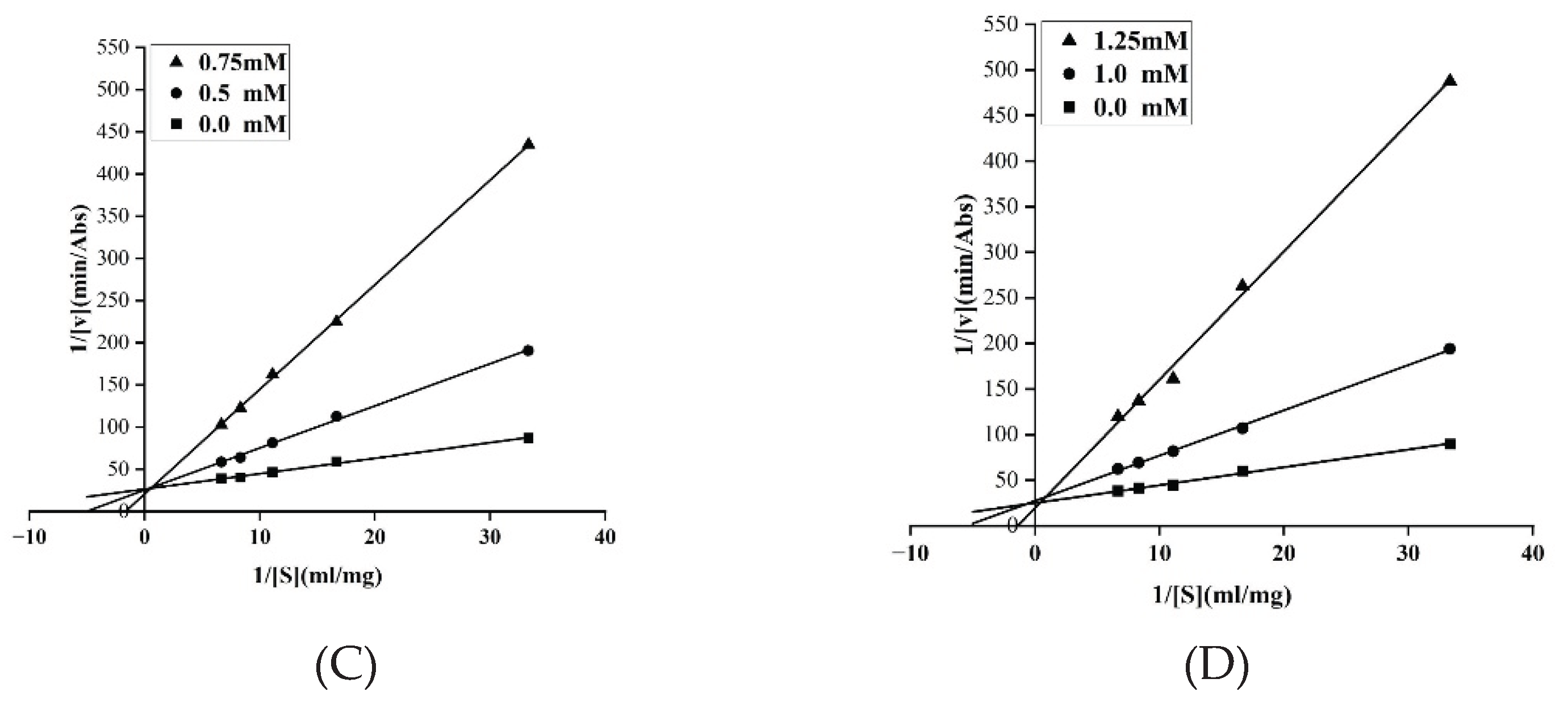
3.3. In Vitro Validation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PL | Pancreatic lipase |
| TCM | Traditional Chinese medicine |
| MD | Molecular dynamics |
| MM/PBSA | Mechanics/Poisson-Boltzmann surface area |
| RMSD | Root Mean Square Deviation |
| Rg | Radius of Gyration |
| pNPP | p-nitrophenyl palmitate |
| ATR-I | Atractylenolide I |
| LIN | Linarin |
| HYD | Hydroxygenkwanin |
| SAL-B | Salvianolic Acid B |
| PEI | Peiminine |
| MUL-A | Mulberroside A |
References
- Ji, T.; Li, Y.; Ma, L. Sarcopenic obesity: An emerging public health problem. Aging Dis. 2022, 13, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Li, H. Obesity: Epidemiology, pathophysiology, and therapeutics. Front. Endocrinol. 2021, 12, 706978. [Google Scholar] [CrossRef] [PubMed]
- Swinburn, B.A.; Sacks, G.; Hall, K.D.; McPherson, K.; Finegood, D.T.; Moodie, M.L.; Gortmaker, S.L. The global obesity pandemic: shaped by global drivers and local environments. Lancet 2011, 378(9793), 804–814. [Google Scholar] [CrossRef]
- Knowler, W.C.; Barrett-Connor, E.; Fowler, S.E.; Hamman, R.F.; Lachin, J.M.; Walker, E.A.; Nathan, D.M. Diabetes Prevention Program Research Group. Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N. Engl. J. Med. 2002, 346, 393–403. [Google Scholar] [PubMed]
- Lopez-Jimenez, F.; Almahmeed, W.; Bays, H.; Cuevas, A.; Di Angelantonio, E.; le Roux, C.W.; Sattar, N.; Sun, M.C.; Wittert, G.; Pinto, F.J.; Wilding, J.P.H. Obesity and cardiovascular disease: Mechanistic insights and management strategies. Eur. J. Prev. Cardiol. 2022, 29, 2218–2237. [Google Scholar] [CrossRef]
- Subramaniyan, V.; Hanim, Y.U. Role of pancreatic lipase inhibition in obesity treatment: mechanisms and challenges towards current insights and future directions. Int J Obes. 2025, 49, 492–506. [Google Scholar] [CrossRef]
- McClendon, K.S.; Riche, D.M.; Uwaifo, G.I. Orlistat: Current status in clinical therapeutics. Expert Opin. Drug Saf. 2009, 8, 727–744. [Google Scholar] [CrossRef]
- Fan, Q.; Xu, F.; Liang, B.; Zou, X. The anti-obesity effect of traditional Chinese medicine on lipid metabolism. Front. Pharmacol. 2021, 12, 696603. [Google Scholar] [CrossRef]
- Li, C.; Zhang, H.; Li, X. The mechanism of traditional Chinese medicine for the treatment of obesity. Diabetes Metab. Syndr. Obes. 2020, 13, 3371–3381. [Google Scholar] [CrossRef]
- Kontoyianni, M. Docking and virtual screening in drug discovery. Methods Mol. Biol. 2017, 1647, 255–266. [Google Scholar]
- Pantsar, T.; Poso, A. Binding Affinity via Docking: Fact and Fiction. Molecules 2018, 23(8), 1899. [Google Scholar] [CrossRef] [PubMed]
- Menchon, G.; Maveyraud, L.; Czaplicki, G. Molecular dynamics as a tool for virtual ligand screening. Methods Mol. Biol. 2018, 1762, 145–178. [Google Scholar] [PubMed]
- Genheden, S.; Ryde, U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin Drug Discov. 2015, 10(5), 449–61. [Google Scholar] [CrossRef] [PubMed]
- Dankwa, B.; Broni, E.; Enninful, K.S.; Kwofie, S.K.; Wilson, M.D. Consensus docking and MM-PBSA computations identify putative furin protease inhibitors for developing potential therapeutics against COVID-19. Struct. Chem. 2022, 33, 2221–2241. [Google Scholar] [CrossRef]
- Al-Nema, M.; Gaurav, A.; Lee, V.S. Docking based screening and molecular dynamics simulations to identify potential selective PDE4B inhibitor. Heliyon. 2020, 6(9), e04856. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Nguyen, P.T.V.; Huynh, H.A.; Truong, D.V.; Tran, T.D.; Vo, C.T. Exploring Aurone Derivatives as Potential Human Pancreatic Lipase Inhibitors through Molecular Docking and Molecular Dynamics Simulations. Molecules 2020, 25(20), 4657. [Google Scholar] [CrossRef]
- Vulichi, S.R.; Runthala, A.; Rachamreddy, S.K.; Yaramanedi, R.S.P.; Sahoo, P.S.; Burra, P.V.L.S.; Kaur, N.; Akkiraju, S.; Kanala, S.R.; Chippada, A.R.; Murthy, S.D.S. Appraisal of pancreatic lipase inhibitory potential of Ziziphus oenoplia (L.)Mill. leaves by in vitro and in Silico Approaches. ACS Omega. 2023, 8, 16630–16646. [Google Scholar] [CrossRef]
- Prabhakar, L.; Dicky John, D.G.; Singh, S.R.; Murali, A. Computational analysis of marine algal compounds for obesity management against pancreatic lipase. J. Biomol. Struct. Dyn. 2023, 41, 4863–4872. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Ru, J.; Li, P.; Wang, J.; Zhou, W.; Li, B.; Huang, C.; Li, P.; Guo, Z.; Tao, W.; Yang, Y.; Xu, X.; Li, Y.; Wang, Y.; Yang, L. TCMSP: A database of systems pharmacology for drug discovery from herbal medicines. J. Cheminform. 2014, 6, 13. [Google Scholar] [CrossRef] [PubMed]
- Páll, S.; Zhmurov, A.; Bauer, P.; Abraham, M.; Lundborg, M.; Gray, A.; Hess, B.; Lindahl, E. Heterogeneous parallelization and acceleration of molecular dynamics simulations in GROMACS. J. Chem. Phys. 2020, 153, 134110. [Google Scholar] [CrossRef] [PubMed]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory. Comput. 2015, 11(8), 3696–3713. [Google Scholar] [CrossRef] [PubMed]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; et al. Gaussian 09, Revision A.02; Gaussian, Inc.: Wallingford CT, USA, 2016. [Google Scholar]
- Case, DA; Aktulga, HM; Belfon, K; Cerutti, DS; Cisneros, GA; et al. AmberTools. J Chem Inf Model. 2023, 63(20), 6183–6191. [Google Scholar] [CrossRef]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25(9), 1157–74. [Google Scholar] [CrossRef]
- Valdés-Tresanco, M.S.; Valdés-Tresanco, M.E.; Valiente, P.A.; Moreno, E. gmx_MMPBSA: A new tool to perform end-state free energy calculations with GROMACS. J. Chem. Theory Comput. 2021, 17, 6281–6291. [Google Scholar] [CrossRef]
- Glisan, S.L.; Grove, K.A.; Yennawar, N.H.; Lambert, J.D. Inhibition of pancreatic lipase by black tea theaflavins: Comparative enzymology and in silico modeling studies. Food. Chem. 2017, 216, 296–300. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; Vanderplas, J.; Passos, A.; Cournapeau, D.; Brucher, M.; Perrot, M.; Duchesnay, E. Scikit-learn: Machine Learning in Python. J. Mach. Learn Res. 2011, 12, 2825–2830. [Google Scholar]
- Winkler, F.K.; D'Arcy, A.; Hunziker, W. Structure of human pancreatic lipase. Nature 1990, 343, 771–774. [Google Scholar] [CrossRef]
- Chen, T.; Tong, M.; Ma, K. Synthesis of Oleanolic Acid Derivatives and Their Inhibitory Effects on Pancreatic Lipase. Nat. Prod. Res. Dev. 2016, 28, 11–16. [Google Scholar]
- Hou, T.; et al. Assessing the performance of MM/PBSA and MM/GBSA methods. 2. Entropy calculations and their influence on the prediction of binding affinities. J Chem Theory Comput. 2011, 7, 27–39. [Google Scholar]
- Kongsted, J.; Ryde, U. An improved method to predict the entropy term with the MM/PBSA approach. J Comput-Aided Mol Des. 2009, 23, 63–71. [Google Scholar] [CrossRef]
- Cardullo, N.; Muccilli, V.; Pulvirenti, L.; Tringali, C. Natural Isoflavones and Semisynthetic Derivatives as Pancreatic Lipase Inhibitors. J. Nat. Prod. 2021, 84, 654–665. [Google Scholar] [CrossRef]
- Li, M.; Chen, Y.T.; Ruan, J.C.; Wang, W.J.; Chen, J.G.; Zhang, Q.F. Structure-activity relationship of dietary flavonoids on pancreatic lipase. Curr. Res. Food. Sci. 2023, 6, 100424. [Google Scholar] [CrossRef]
- Tran, T.H.; Mai, T.T.; Ho, T.T.; Le, T.N.; Cao, T.C.; Thai, K.M.; Tran, T.S. Inhibition of pancreatic lipase by flavonoid derivatives: in vitro and in silico investigations. Adv Pharmacol Pharm Sci. 2024, 6655996. [Google Scholar]
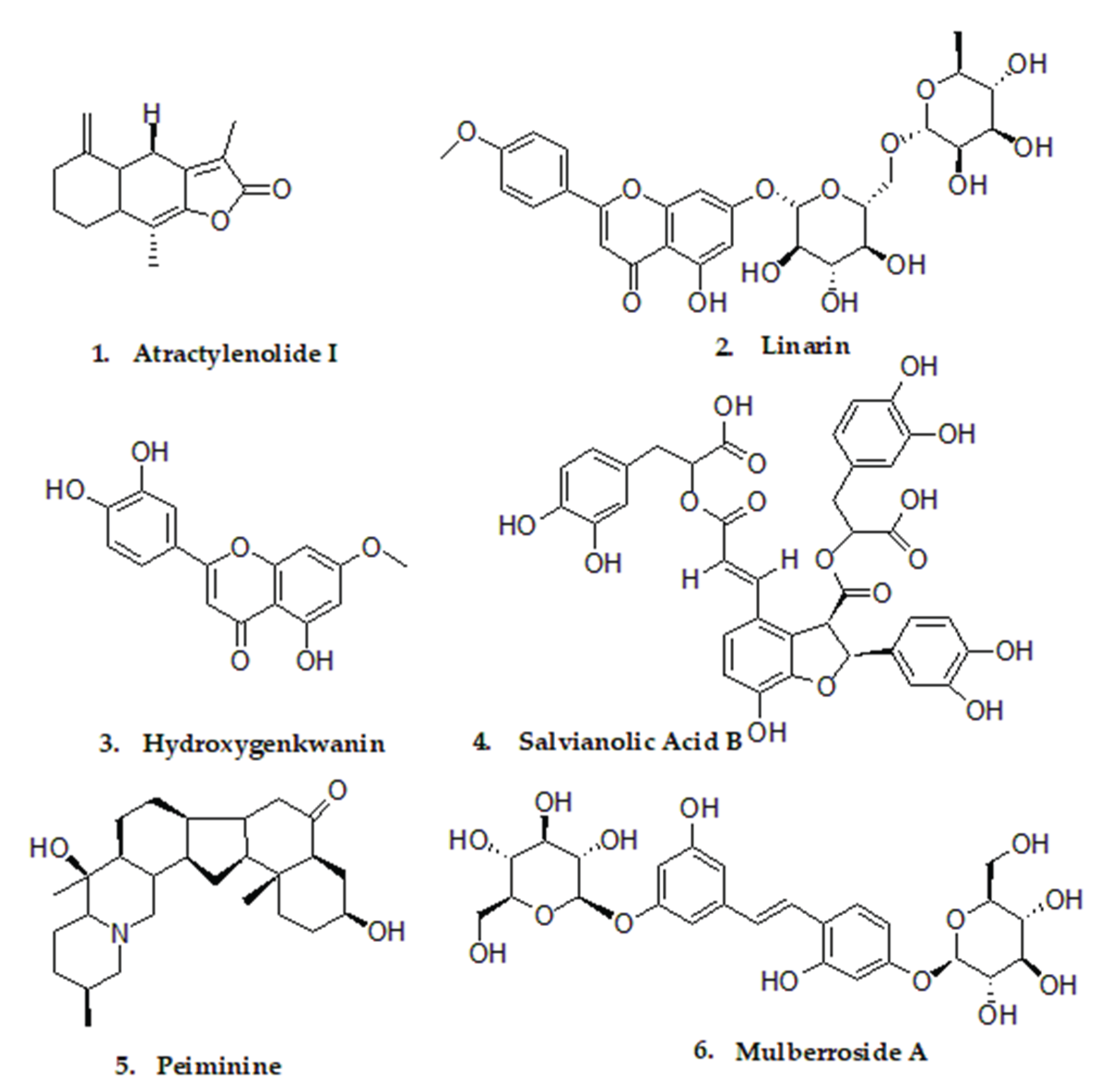
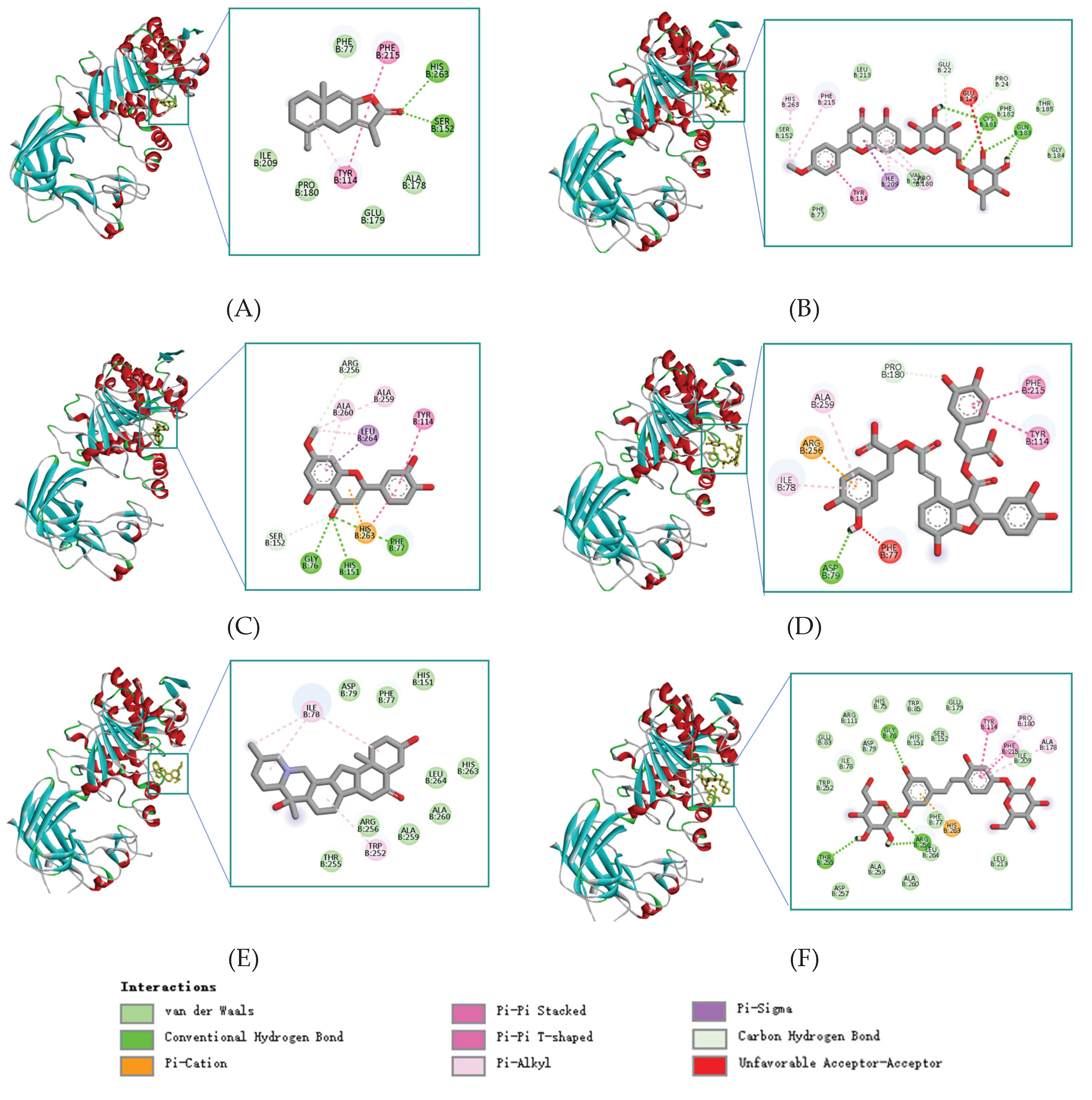
| Mol_ID | Mol_Name | Score | Hydrogen Bonds | Hydrophobic Interactions |
|---|---|---|---|---|
| MOL000043 | Atractylenolide I (ATR-I) |
−9.0 | Ser152,His263 | Phe77,Tyr114,Ala178,Phe215 |
| MOL001790 | Linarin (LIN) | −9.3 | Cys181,Glu183 | Phe182,Thr185,Val210,Leu213 |
| MOL005530 | Hydroxygenkwanin (HYD) |
−9.3 | Gly76,Phe77,His151 | Arg256,Tyr114,Ala260,Leu264 |
| MOL007074 | Salvianolic Acid B (SAL-B) | −9.1 | Asp79 | Pro180,Ile78,Tyr114,Phe215 |
| MOL009593 | Peiminine (PEI) | −9.4 | None | Phe77,Ile78,His151,Trp252, Thr255,Arg256,Ala259,Leu264 |
| MOL012733 | Mulberroside A (MUL-A) | −9.9 | Gly76,Thr255,Arg256 | Phe77,ILE78,Tyr114,Pro180, Ile209,Phe215,Ala259,Leu264 |
| Compound | ΔEVDW | ΔEELE | ΔEPB | ΔENPOLAR | ΔGGAS | ΔGSOLV | ΔGTOTAL |
|---|---|---|---|---|---|---|---|
| ATR-I | -17.08±0.51 | -5.35±0.50 | 11.09±0.60 | -2.11±0.04 | -22.43±0.90 | 8.99±0.56 | -13.44±0.45 |
| LIN | -26.47±0.72 | -43.49±1.61 | 53.71±1.39 | -3.29±0.05 | -69.96±1.62 | 50.42±1.37 | -19.54±0.43 |
| HYD | -35.35±0.39 | -6.88±0.60 | 29.99±0.93 | -3.34±0.02 | -42.23±0.82 | 26.65±0.92 | -15.58±0.85 |
| SAL-B | -28.27±0.47 | -63.91±1.09 | 75.13±0.83 | -4.33±0.03 | -92.18±0.88 | 70.80±0.82 | -21.38±0.40 |
| PEI | -38.97±0.29 | -13.26±0.65 | 34.85±0.50 | -3.87±0.04 | -52.23±0.62 | 30.99±0.50 | -21.24±0.39 |
| MUL-A | -48.58±0.43 | -28.76±0.76 | 68.96±1.07 | -4.95±0.03 | -77.34±0.84 | 64.01±1.06 | -13.33±0.58 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




