Submitted:
19 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
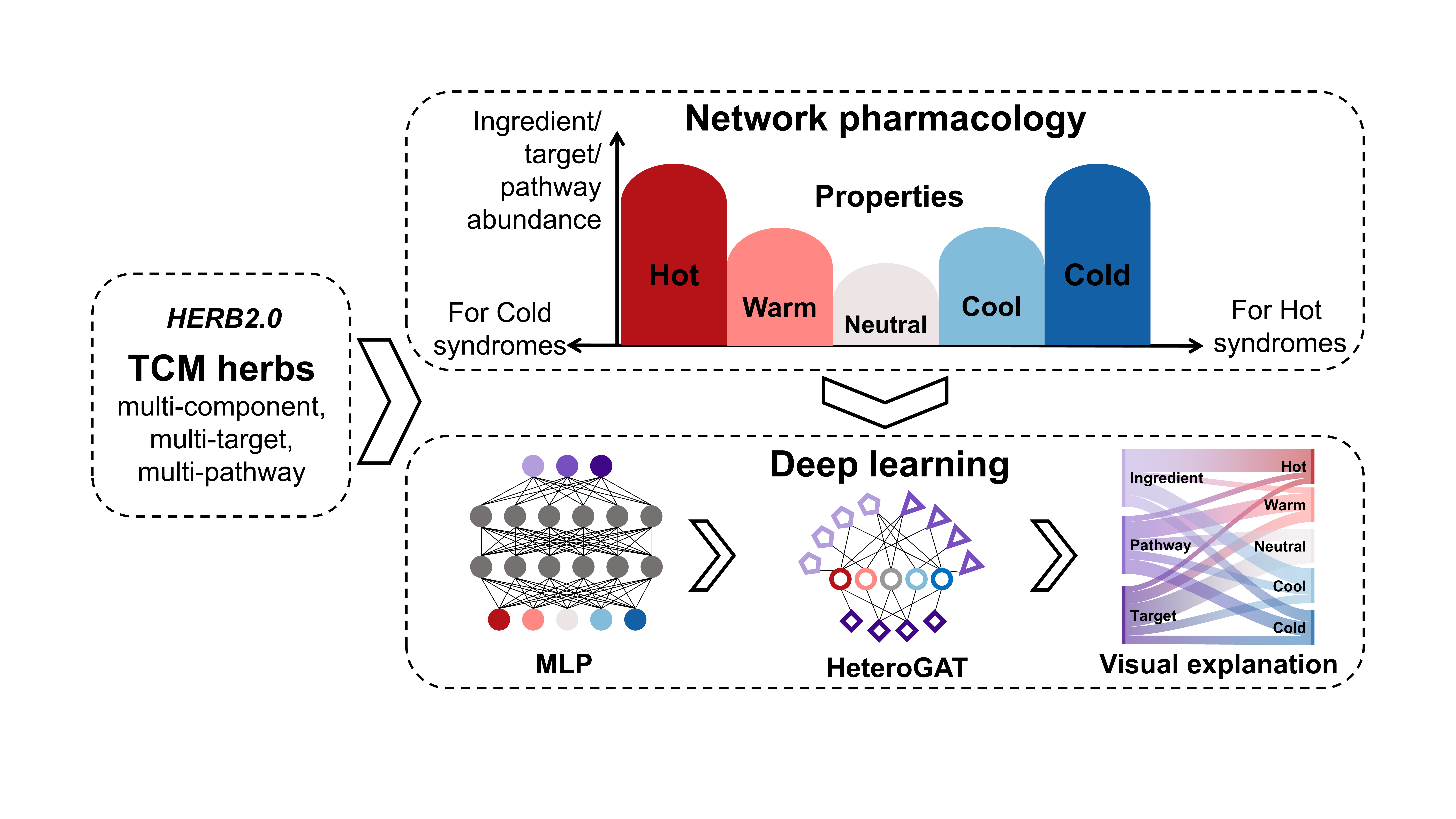
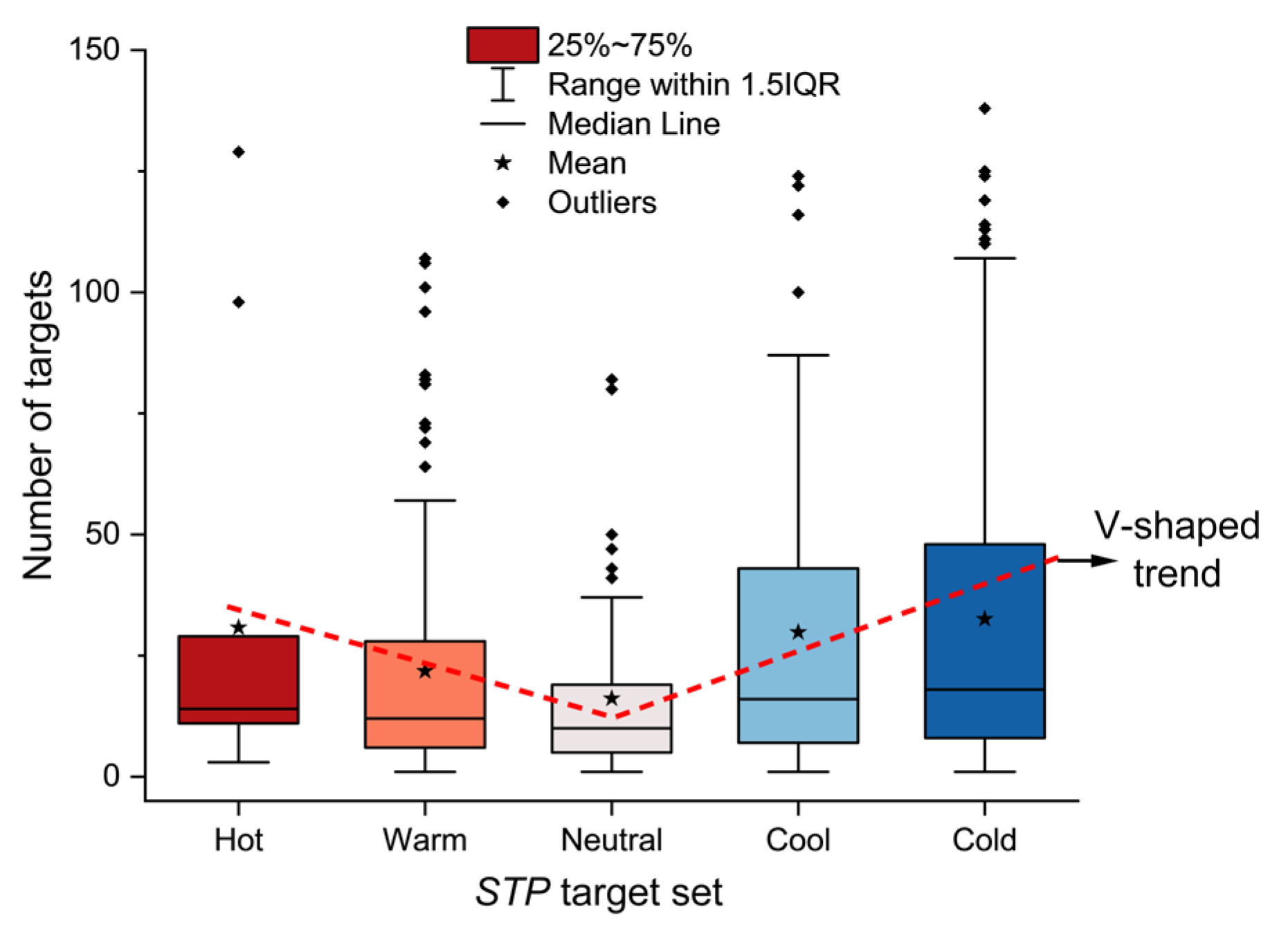
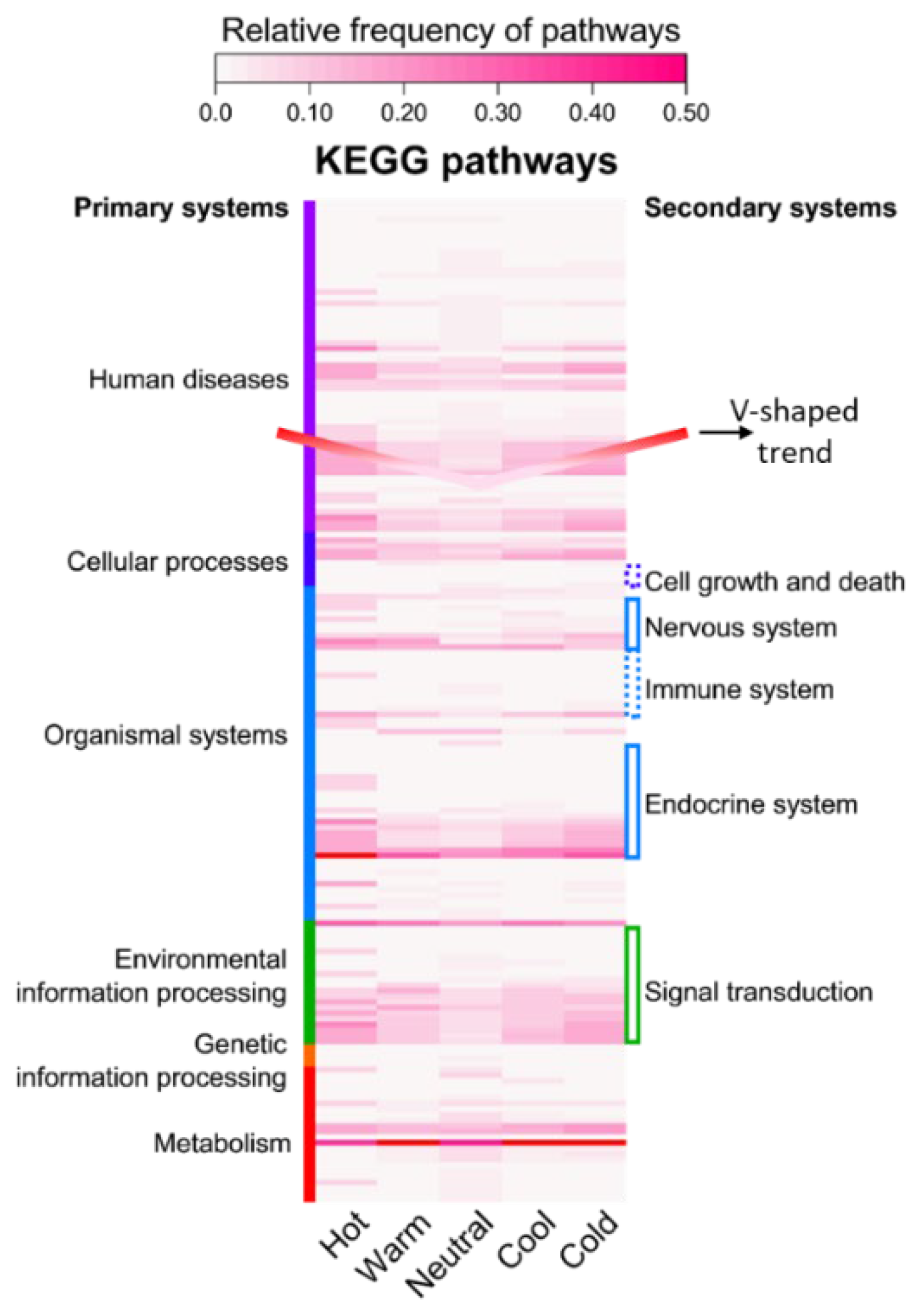
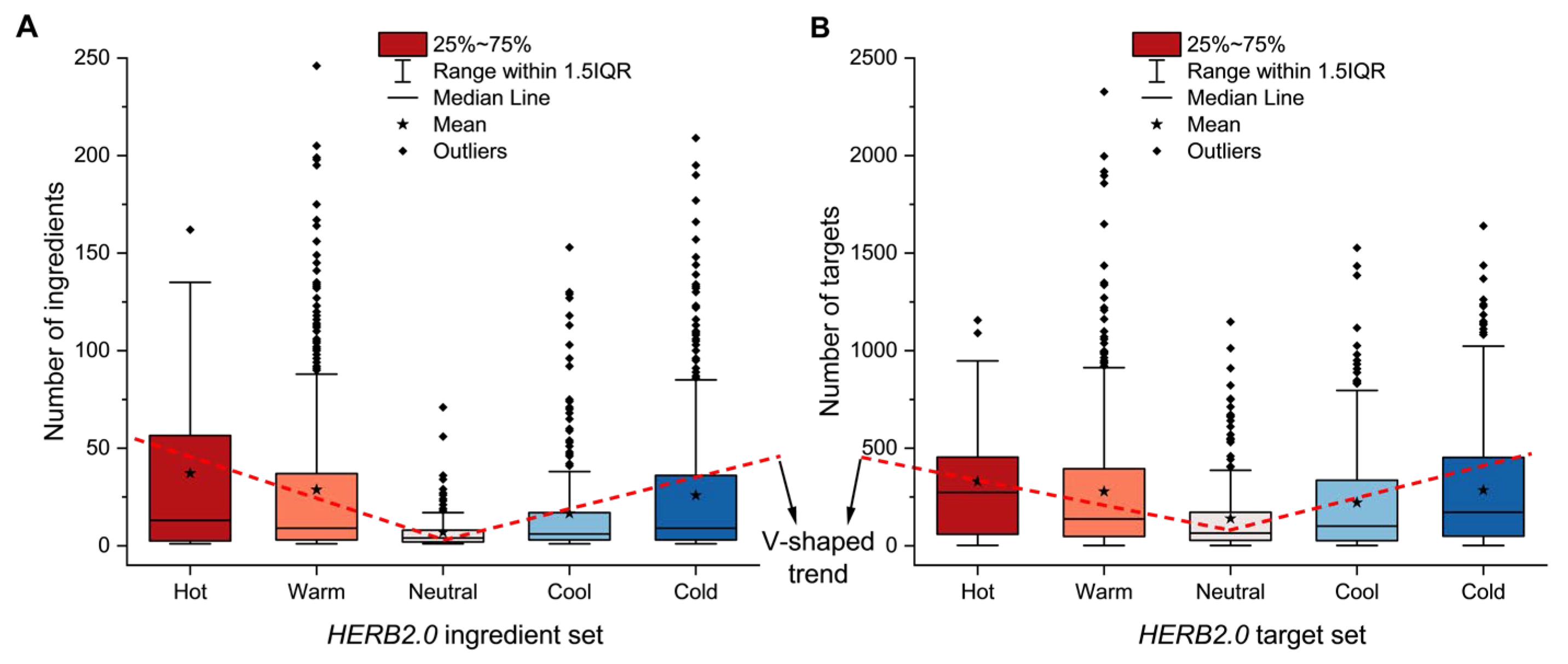
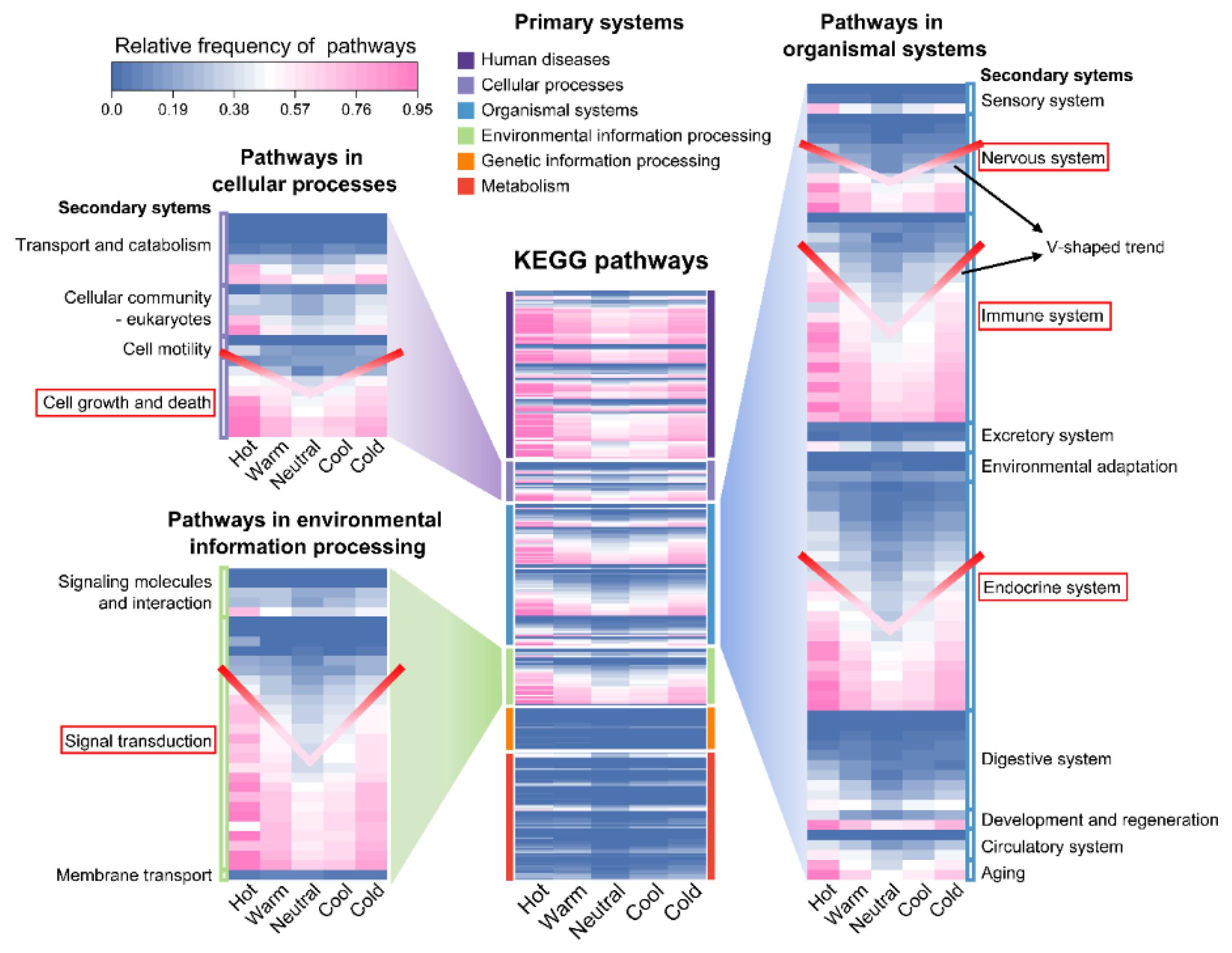
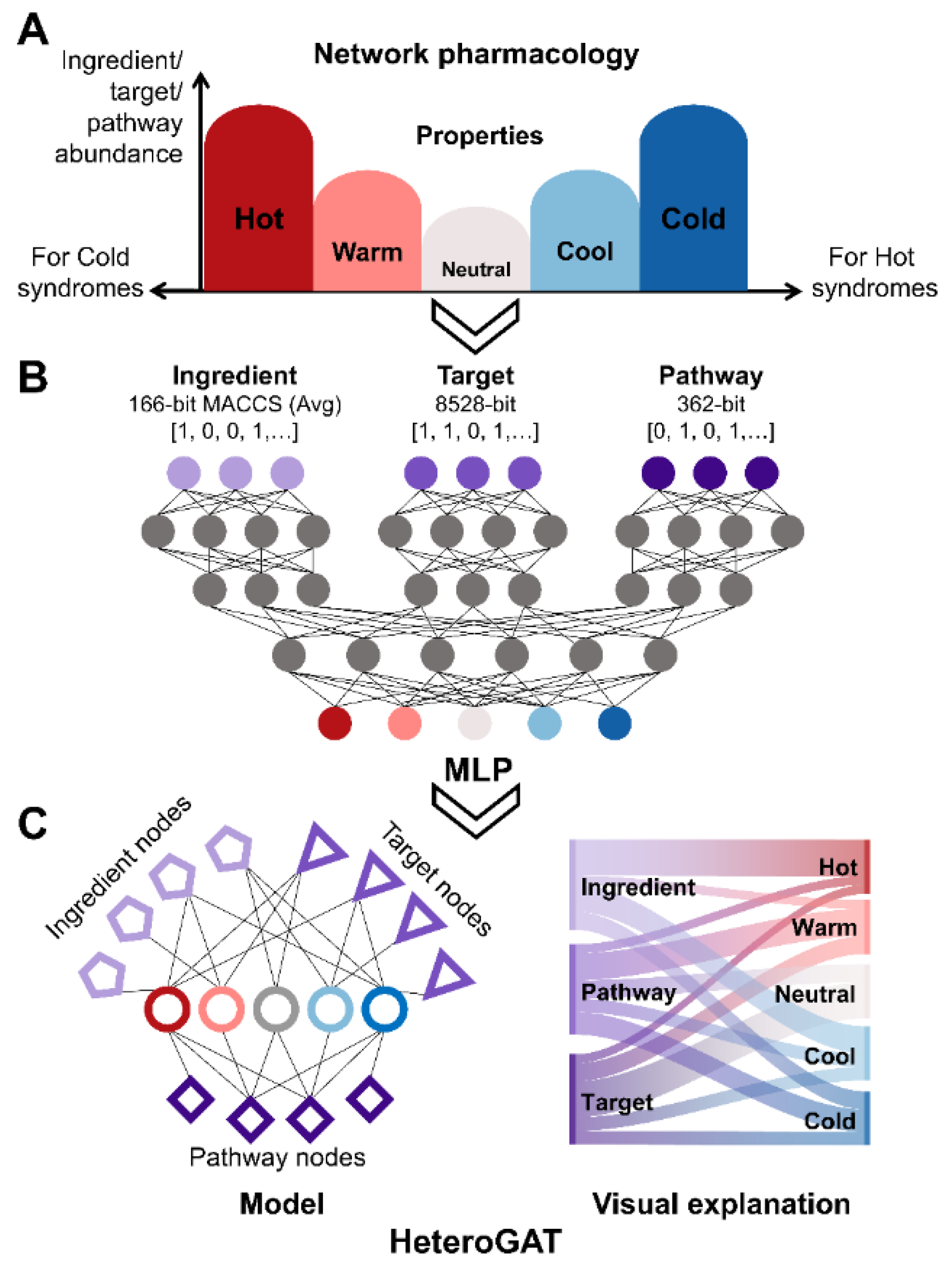
Extensive experimental data on Traditional Chinese Medicine are available in literature and databases. However, many studies focus on specific diseases or pathways with small sample sizes. As a result, the fundamental pharmacological basis underlying TCM herb properties remains insufficiently elucidated. Based on the concept of the multi-component, multi-target, multi-pathway network of TCM, a data-driven strategy was developed for the profiling of TCM herb properties through network pharmacology and deep learning, facilitating the exploration of the scientific evidence underlying TCM herb properties. Large-scale ingredient and target data of TCM herbs were curated from the HERB2.0 database. KEGG pathway enrichment was conducted for each herb with relative frequency profiling of distinct property groups. Deep learning models were developed and optimized for classification with visual explanation. As a result, high-relative frequency pathways were highly concentrated in five systems (endocrine, immune, nervous, signal transduction, cell growth and death) of KEGG. Herbs with distinct properties exhibited a V-shaped trend (Hot>Warm>Neutral<Cool<Cold) in terms of the abundance of ingredients, targets and high-frequency pathways. The HeteroGAT model improved classification accuracy and provided visual explanations at the ingredient–target–pathway level. We demonstrated a viable strategy to profile TCM property classification from a holistic perspective on ingredients, targets, and pathways, which could help elucidate the scientific basis of TCM properties. However, further advances in model refinement and data matrices are required to enhance the effectiveness of this strategy.
Keywords:
1. Introduction
2. Results and Discussions
2.1. Overview of Data Characteristics
2.2. Property Profiling Through Network Pharmacology
2.3. Establishment of Classification Models of TCM Herb Properties by Deep Learning
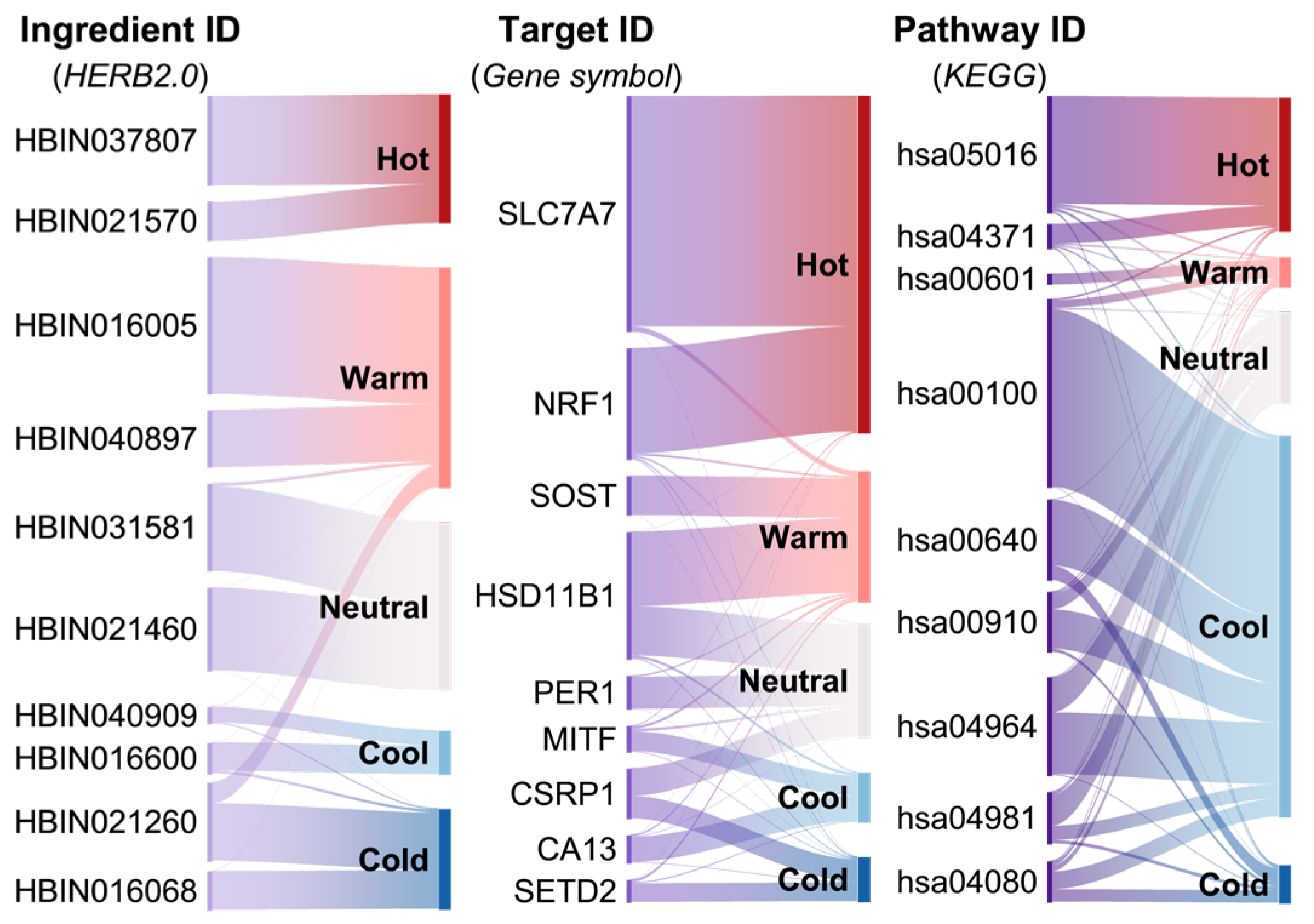
2.4. Property Profiling Through HeteroGAT
3. Materials and Methods
3.1. Data Acquisition
3.2. KEGG Pathway Enrichment Analysis
3.3. Statistical Analysis of Network Pharmacology
3.4. Classification Models of TCM Herbs Properties by MLP
3.5. Classification Models of TCM Herbs Properties by HeteroGAT
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| AUC | Averaged area under the receiver operating characteristic curve |
| HeteroGAT | Heterogeneous Graph Attention Network |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| MLP | Multilayer Perceptron |
| SMILES | Simplified Molecular Input Line Entry System |
| STP | SwissTargetPrediction |
| TCM | Traditional Chinese Medicine |
Appendix A
References
- Chen, P.; Wang, B.-Y.; Zhang, P.; Li, S. Cold and Hot Syndromes in Traditional Chinese Medicine: Insights from the Perspective of Immunometabolic Homeostasis. World Journal of Traditional Chinese Medicine 2024, 10, 434–442. [Google Scholar] [CrossRef]
- Wang, B.-Y.; Chen, P.; Zhang, P.; Li, S. Biological Mechanism of Traditional Chinese Medicine Formula and Herbs in Treating Diseases from the Perspective of Cold and Hot. World Journal of Traditional Chinese Medicine 2024, 10, 274–283. [Google Scholar] [CrossRef]
- Liu, J.; Feng, W.; Peng, C. A Song of Ice and Fire: Cold and Hot Properties of Traditional Chinese Medicines. Front Pharmacol 2020, 11, 598744. [Google Scholar] [CrossRef] [PubMed]
- Eigenschink, M.; Dearing, L.; Dablander, T.E.; Maier, J.; Sitte, H.H. A critical examination of the main premises of Traditional Chinese Medicine. Wien Klin Wochenschr 2020, 132, 260–273. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Liu, Z.; Liao, J.; Chen, Q.; Lu, X.; Fan, X. Network pharmacology approaches for research of Traditional Chinese Medicines. Chin J Nat Med 2023, 21, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhang, B. Traditional Chinese medicine network pharmacology: theory, methodology and application. Chinese Journal of Natural Medicines 2014, 11, 110–120. [Google Scholar] [CrossRef]
- Li, R.; Ma, T.; Gu, J.; Liang, X.; Li, S. Imbalanced network biomarkers for traditional Chinese medicine Syndrome in gastritis patients. Sci Rep 2013, 3, 1543. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Fu, X.; Wang, Z. Multisolvent Similarity Measure of Chinese Herbal Medicine Ingredients for Cold-Hot Nature Identification. J Chem Inf Model 2019, 59, 5065–5073. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Mervin, L.H.; Li, X.; Yu, H.; Li, J.; Mohamad Zobir, S.Z.; Zoufir, A.; Zhou, Y.; Song, Y.; Wang, Z.; et al. Toward Understanding the Cold, Hot, and Neutral Nature of Chinese Medicines Using in Silico Mode-of-Action Analysis. J Chem Inf Model 2017, 57, 468–483. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Xu, B. New insights into molecular mechanisms of "Cold or Hot" nature of food: When East meets West. Food Res Int 2021, 144, 110361. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Li, Y.; Cheng, H.; Li, G.; Liang, Z. Definition of the molecular bases of cold and hot properties of traditional Chinese medicine through machine learning. Pharmacological Research - Modern Chinese Medicine 2022, 4. [Google Scholar] [CrossRef]
- Yang, M.; Liu, W. Classification of cold and hot medicinal properties of Chinese herbal medicines based on graph convolutional network. Digital Chinese Medicine 2024, 7, 356–364. [Google Scholar] [CrossRef]
- Li, Y.; Li, R.; Ouyang, Z.; Li, S. Herb Network Analysis for a Famous TCM Doctor's Prescriptions on Treatment of Rheumatoid Arthritis. Evid Based Complement Alternat Med 2015, 2015, 451319. [Google Scholar] [CrossRef] [PubMed]
- Luo, C.; Yu, H.; Yang, T.; Bai, C.; He, B.; Yan, Y.; Liu, T.; Wang, J.; Gu, X. Data Mining and Systematic Pharmacology to Reveal the Mechanisms of Traditional Chinese Medicine in Recurrent Respiratory Tract Infections' Treatment. Evid Based Complement Alternat Med 2020, 2020, 8979713. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Jafari, M.; Tang, Y.; Tang, J. Predicting Meridian in Chinese traditional medicine using machine learning approaches. PLoS Comput Biol 2019, 15, e1007249. [Google Scholar] [CrossRef] [PubMed]
- Gao, K.; Liu, L.; Lei, S.; Li, Z.; Huo, P.; Wang, Z.; Dong, L.; Deng, W.; Bu, D.; Zeng, X.; et al. HERB 2.0: an updated database integrating clinical and experimental evidence for traditional Chinese medicine. Nucleic Acids Research 2024, 53, D1404–D1414. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Scientific Reports 2017, 7, 42717. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Research 2019, 47, W357–W364. [Google Scholar] [CrossRef]
- Lin, T.-Y.; Goyal, P.; Girshick, R.; He, K.; Dollár, P. Focal loss for dense object detection. In Proceedings of the IEEE international conference on computer vision; 2017; pp. 2980–2988. [Google Scholar]
- Wang, X.; Ji, H.; Shi, C.; Wang, B.; Ye, Y.; Cui, P.; Yu, P.S. Heterogeneous Graph Attention Network. In Proceedings of the The World Wide Web Conference, San Francisco, CA, USA, 2019; pp. 2022–2032. [Google Scholar]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-cam: Visual explanations from deep networks via gradient-based localization. In Proceedings of the Proceedings of the IEEE international conference on computer vision, 2017; pp. 618–626. [Google Scholar]
| Properties | Number of herbs |
|---|---|
| Hot | 36 |
| Warm | 530 |
| Neutral | 256 |
| Cool | 255 |
| Cold | 504 |
| Feature combinations | MLP | HeteroGAT | ||
|---|---|---|---|---|
| BalAcc | Macro AUC | BalAcc | Macro AUC | |
| I | 0.3632 ± 0.0310 | 0.6616 ± 0.0272 | 0.4638 ± 0.0300 | 0.6983 ± 0.0203 |
| T | 0.3658 ± 0.0285 | 0.6483 ± 0.0219 | 0.3575 ± 0.0375 | 0.6264 ± 0.0219 |
| P | 0.3257 ± 0.0300 | 0.6294 ± 0.0240 | 0.3286 ± 0.0347 | 0.5849 ± 0.0241 |
| I-T | 0.3646 ± 0.0303 | 0.6540 ± 0.0245 | 0.4594 ± 0.0323 | 0.6954 ± 0.0214 |
| I-P | 0.3497 ± 0.0321 | 0.6522 ± 0.0221 | 0.4617 ± 0.0343 | 0.7035 ± 0.0233 |
| I-T-P | 0.3652 ± 0.0294 | 0.6527 ± 0.0254 | 0.4586 ± 0.0305 | 0.7016 ± 0.0209 |
| Models | Statistics of recall per class | ||||
|---|---|---|---|---|---|
| Hot | Warm | Neutral | Cool | Cold | |
| MLP | 0.31 ± 0.14 | 0.52 ± 0.06 | 0.27 ± 0.09 | 0.17 ± 0.08 | 0.54 ± 0.10 |
| HeteroGAT | 0.66 ± 0.14 | 0.49 ± 0.08 | 0.35 ± 0.09 | 0.36 ± 0.12 | 0.45 ± 0.13 |
| Resources | Tasks | Features | Models | Performance |
|---|---|---|---|---|
|
TCMID (583 herbs) |
Meridian binary classification |
Ingredient (4922) | RF, SVM, kNN [15] | BalAcc: 0.67 |
|
TCMSP, ETCM (393 herbs) |
Property binary classification |
Ingredient (12793) | RF, SVM, GNB [11] | AUC: 0.82 |
|
TCMID (459 herbs) |
Property binary classification |
Ingredient (8075) | GCN [12] | Acc: 0.84 F1: 0.85 |
|
HERB2.0 (1581herbs) |
Property five-class classification |
Ingredient (18410) Target (8528) Pathway (362) |
HeteroGAT | BalAcc: 0.46 Macro AUC: 0.70 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




