Submitted:
18 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- We introduce an ensemble of neural networks that leverages heterogeneous feature representations for training, improving robustness and generalization.
- The proposed approach is built upon a well-established, for this topic, neural network architecture, extending it through the exploration of different network hyperparameters and pooling strategies to construct the ensemble.
- We provide a publicly available implementation of the proposed system to ensure reproducibility https://github.com/LorisNanni/Ensemble-Deep-Learning-Models-on-Raw-DNA-Sequences-for-Viral-Genome-Identification-in-Human-Samples.
- Extensive experimental evaluations demonstrate that the proposed method achieves state-of-the-art performance when compared to existing approaches on the same dataset and under the same evaluation protocol.
2. Materials and Methods
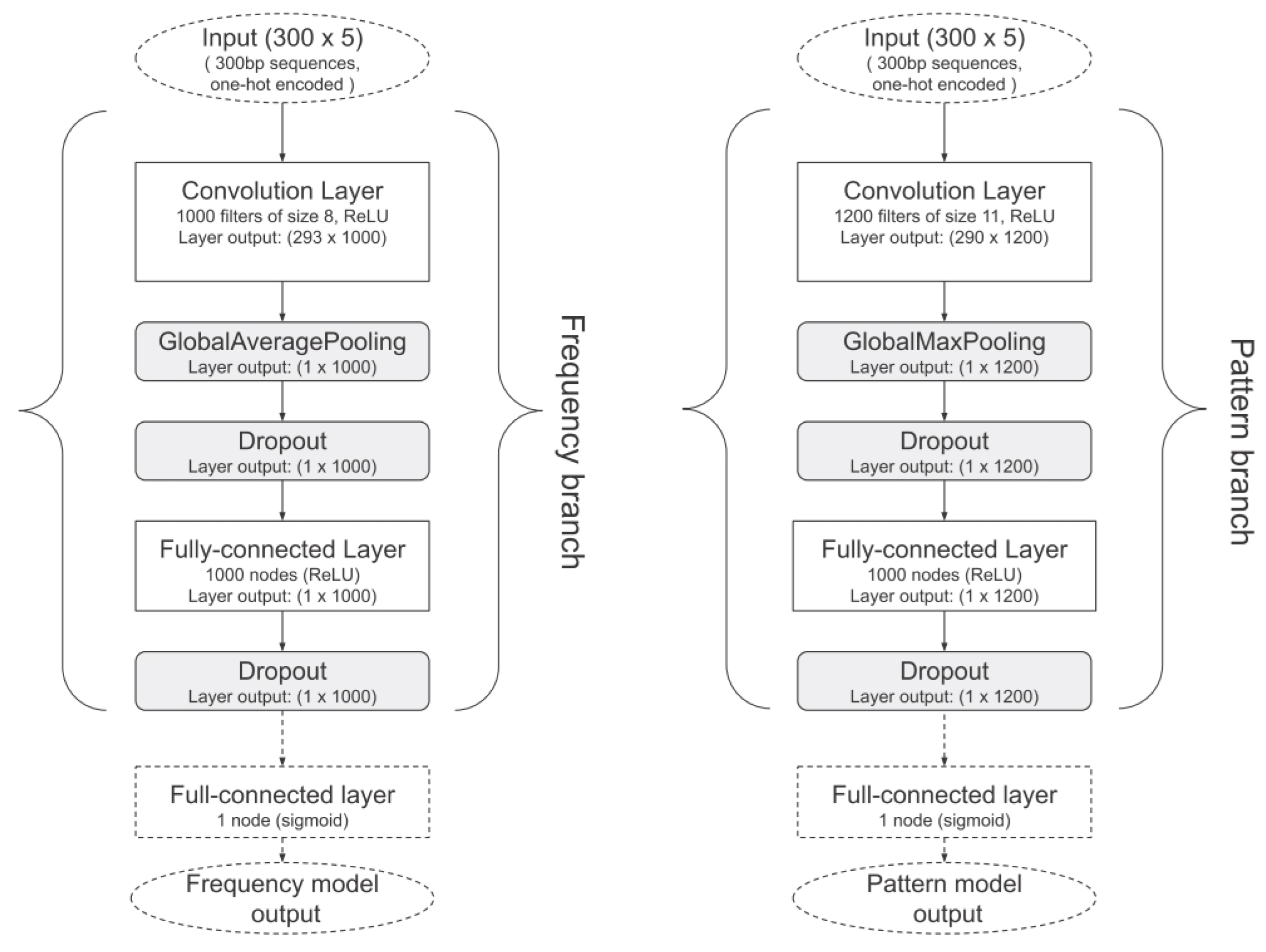
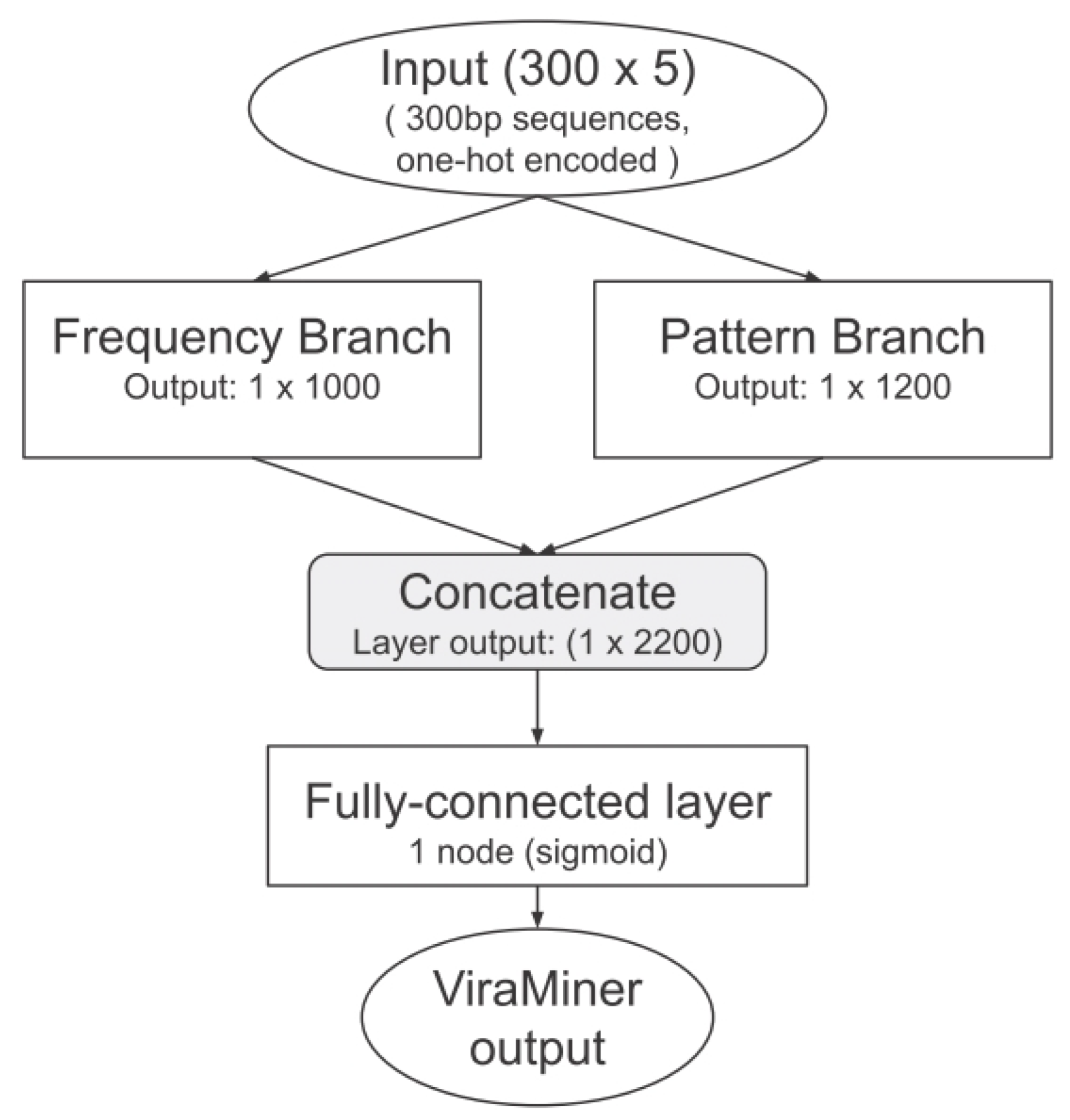
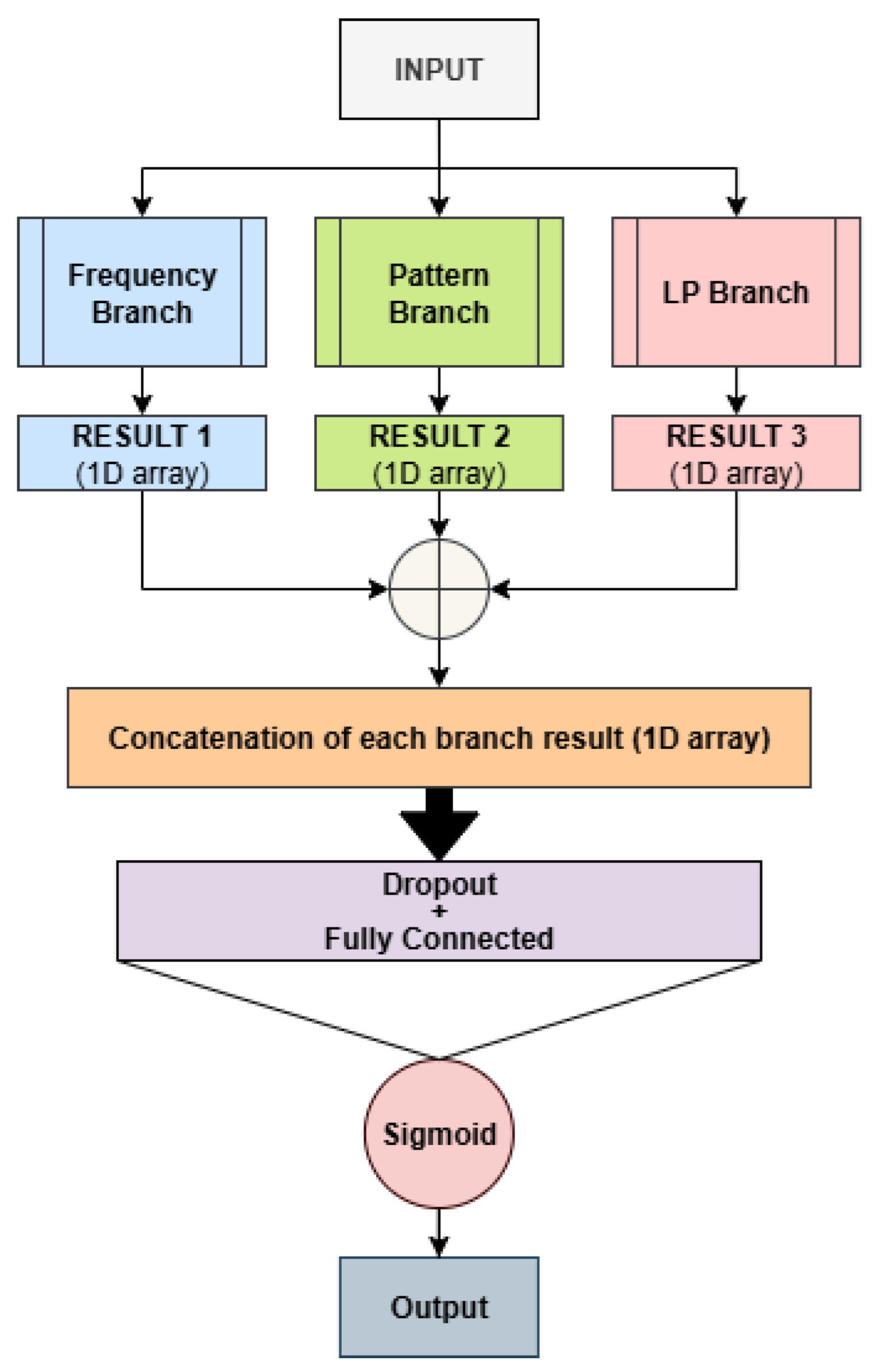
- Bf (VM)
- Bf
- B1
- Bp (VM)
- Bp
- Mf+p+1
- Mf+1
- Mp+1
- Mf+p
- Original ViraMiner architecture.
2.1. Dataset
3. Results
-
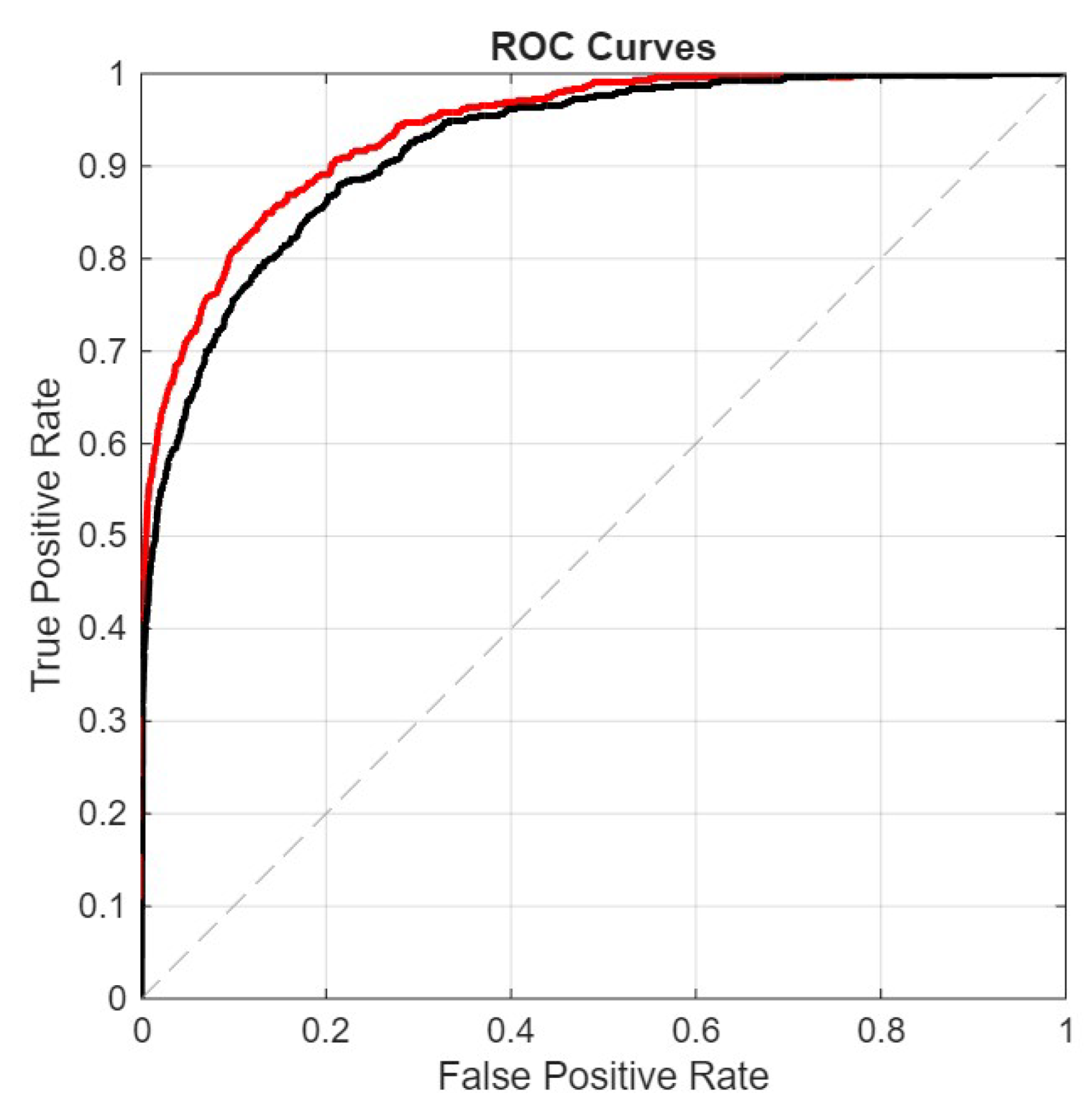
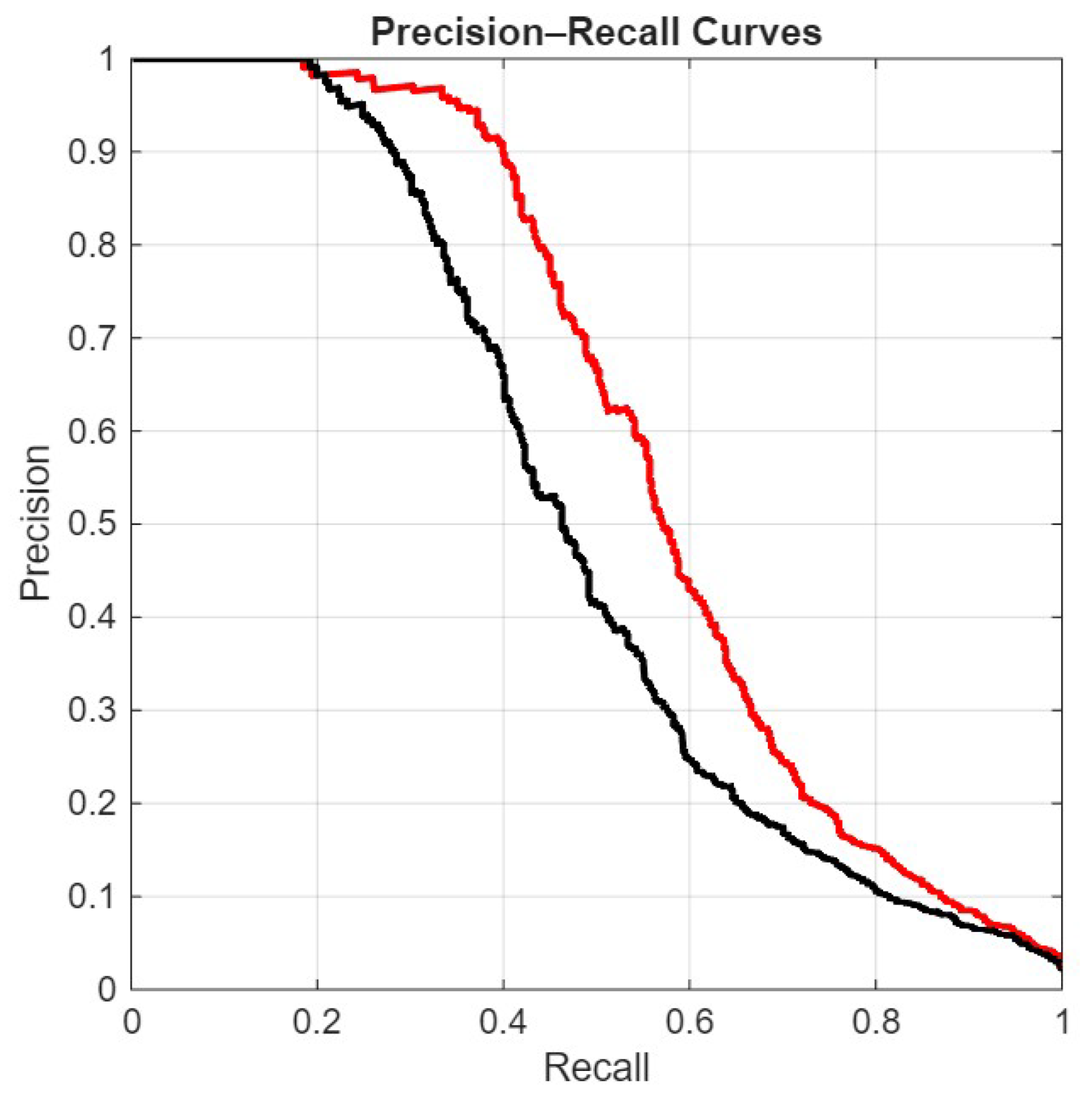
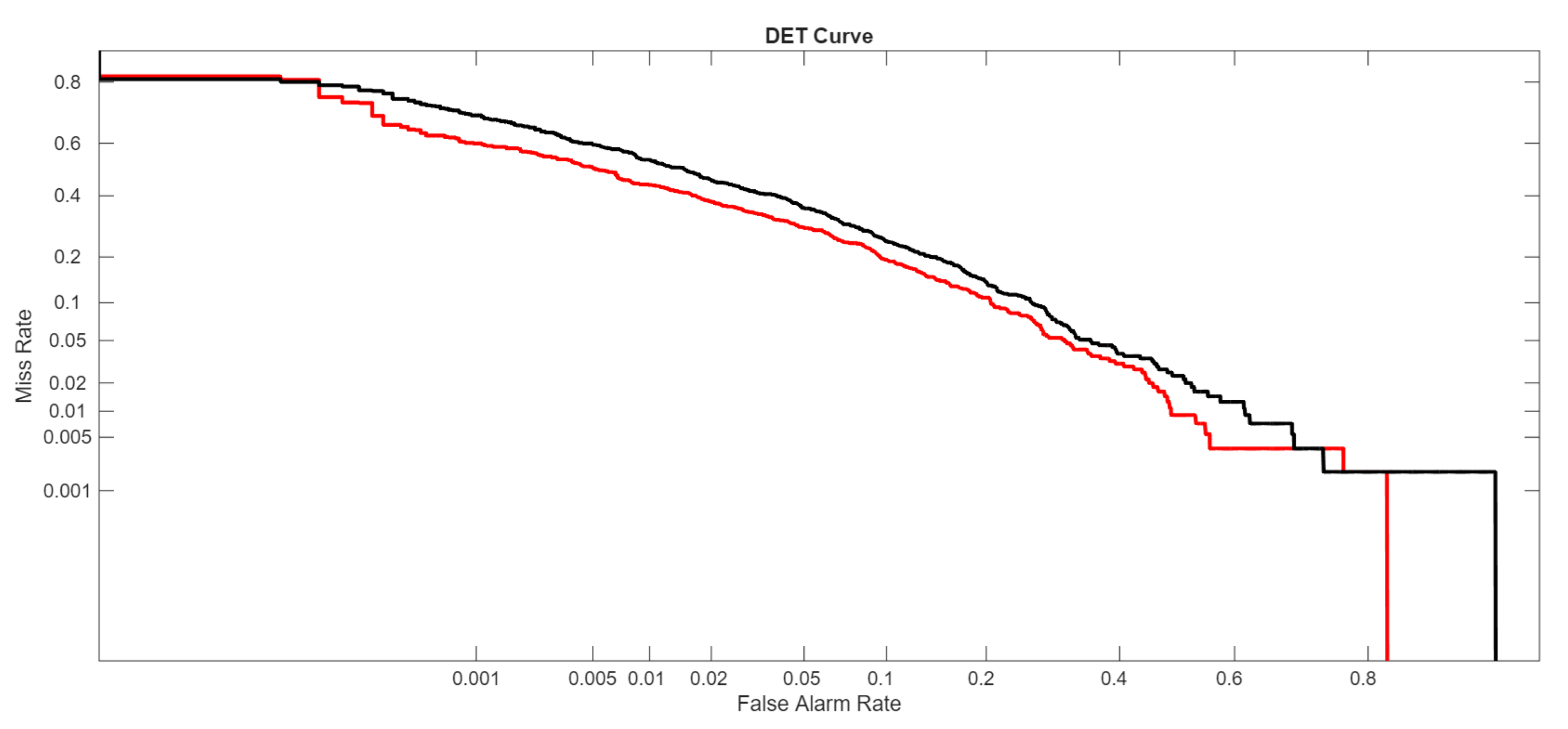
The ROC curve plots the True Positive Rate (TPR) against the False Positive Rate (FPR):The AUROC is the area under this curve:
- FullE, the ensemble proposed here;
- Mf+p+1, our best stand-alone net;
- Viraminer, the original method proposed in [9];
- [2], the comparison of our method with this paper is very interesting, not only because it uses the same dataset and the same data split, but also because this method is based on self-attention and convolutional operations on nucleic acid sequences, leveraging two prominent deep learning strategies commonly used in computer vision and natural language processing.
4. Conclusion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Slack, F.J.; Chinnaiyan, A.M. The Role of Non-coding RNAs in Oncology. Cell 2019, 179, 1033–1055. [Google Scholar] [CrossRef]
- He, S.; Gao, B.; Sabnis, R.; Sun, Q. Nucleic Transformer: Classifying DNA Sequences with Self-Attention and Convolutions. ACS Synthetic Biology 2023, 12, 3205–3214. [Google Scholar] [CrossRef] [PubMed]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate Structure Prediction of Biomolecular Interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Do, D.T.; Le, T.Q.T.; Le, N.Q.K. Using Deep Neural Networks and Biological Subwords to Detect Protein S-sulfenylation Sites. Briefings in Bioinformatics 2020, 22, bbaa128. [Google Scholar] [CrossRef] [PubMed]
- Wylie, K.M.; Mihindukulasuriya, K.A.; Sodergren, E.; Weinstock, G.M.; Storch, G.A. Sequence analysis of the human virome in febrile and afebrile children. PLoS ONE 2012, 7, e27735. [Google Scholar] [CrossRef] [PubMed]
- Cesanelli, F.; Scarvaglieri, I.; De Francesco, M.A.; Alberti, M.; Salvi, M.; Tiecco, G.; Castelli, F.; Quiros-Roldan, E. The Human Virome in Health and Its Remodeling During HIV Infection and Antiretroviral Therapy: A Narrative Review. Microorganisms 2026, 14. [Google Scholar] [CrossRef]
- Zhang, D.; Cao, Y.; Dai, B.; Zhang, T.; Jin, X.; Lan, Q.; Qian, C.; He, Y.; Jiang, Y. The virome composition of respiratory tract changes in school-aged children with Mycoplasma pneumoniae infection. Virology Journal 2025, 22, 10. [Google Scholar] [CrossRef] [PubMed]
- Mercalli, A.; Lampasona, V.; Klingel, K.; Albarello, L.; Lombardoni, C.; Ekstrom, J.; et al. No evidence of enteroviruses in the intestine of patients with type 1 diabetes. Diabetologia 2012, 55, 2479–2488. [Google Scholar] [CrossRef]
- Vincente, A.T.Z.B.J.D.R. ViraMiner: Deep learning on raw DNA sequences for identifying viral genomes in human samples. PLOS ONE 2019, 1–17. [Google Scholar]
- Meiring, T.L.; Salimo, A.T.; Coetzee, B.; Maree, H.J.; Moodley, J.; Hitzeroth, I. Next-generation sequencing of cervical DNA detects human papillomavirus types not detected by commercial kits. Virology Journal 2012, 9, 164. [Google Scholar] [CrossRef]
- Mistry, J.; Finn, R.D.; Eddy, S.R.; Bateman, A.; Punta, M. Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Research 2013, 41, e121. [Google Scholar] [CrossRef]
- Randhawa, G.S.; Soltysiak, M.P.M.; El Roz, H.; de Souza, C.P.E.; Hill, K.A.; Kari, L. Machine learning using intrinsic genomic signatures for rapid classification of novel pathogens: COVID-19 case study. PLOS ONE 2020, 15, e0232391. [Google Scholar] [CrossRef]
- Alipanahi, B.; Delong, A.; Weirauch, M.T.; Frey, B.J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nature Biotechnology 2015, 33, 831–838. [Google Scholar] [CrossRef]
- Kelley, D.R.; Snoek, J.; Rinn, J.L. Basset: learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Research 2016, 26, 990–999. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Troyanskaya, O.G. Predicting effects of noncoding variants with deep learning–based sequence model. Nature Methods 2015, 12, 931. [Google Scholar] [CrossRef]
- Angermueller, C.; Pärnamaa, T.; Parts, L.; Stegle, O. Deep learning for computational biology. Molecular Systems Biology 2016, 12, 878. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. In Proceedings of the Proceedings of the 3rd International Conference on Learning Representations (ICLR), San Diego, CA, USA, 2015. [Google Scholar]
- Radenović, F.; Tolias, G.; Chum, O. Fine-Tuning CNN Image Retrieval with No Human Annotation. IEEE Transactions on Pattern Analysis and Machine Intelligence;arXiv 2019, arXiv:1711.0251241, 1655–1668. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Nowicki, M.; Bzhalava, D.; Bała, P. Massively Parallel Implementation of Sequence Alignment with Basic Local Alignment Search Tool Using Parallel Computing in Java Library. Journal of Computational Biology 2018, 25, 871–881. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Ahlgren, N.A.; Lu, Y.Y.; Fuhrman, J.A.; Sun, F. VirFinder: a novel k-mer based tool for identifying viral sequences from assembled metagenomic data. Microbiome 2017, 5, 69. [Google Scholar] [CrossRef] [PubMed]
- Bzhalava, Z.; Tampuu, A.; Bała, P.; Vicente, R.; Dillner, J. Machine Learning for detection of viral sequences in human metagenomic datasets. BMC Bioinformatics 2018, 19, 336. [Google Scholar] [CrossRef]
- Bzhalava, Z.; Tampuu, A.; Bała, P.; Vicente, R.; Dillner, J. Machine Learning for detection of viral sequences in human metagenomic datasets. BMC Bioinformatics 2018, 19, 336. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Li, L.; Myers, J.R.; Marth, G.T. ART: a next-generation sequencing read simulator. Bioinformatics 2011, 28, 593–594. [Google Scholar] [CrossRef] [PubMed]
- Zabihi, S.; Hashemi, S.; Mansoori, E. EDEN: multiscale expected density of nucleotide encoding for enhanced DNA sequence classification with hybrid deep learning. BMC Bioinformatics 2026, 27, 40. [Google Scholar] [CrossRef] [PubMed]
| Hyperparameter | Tested Configurations |
|---|---|
| Learning rate | 0.1, 0.01, 0.001. |
| Optimizer | Adam |
| Epochs | 30 |
| Batch size | 128 |
| Loss function | Binary Cross-Entropy |
| Dropout | 0.1, 0.2, 0.5. |
| Number of filters | 1000, 1200, 1500 |
| Kernel size | from 5 to 18 |
| Norm type (only LP) | from 1 to 10 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




