Submitted:
13 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
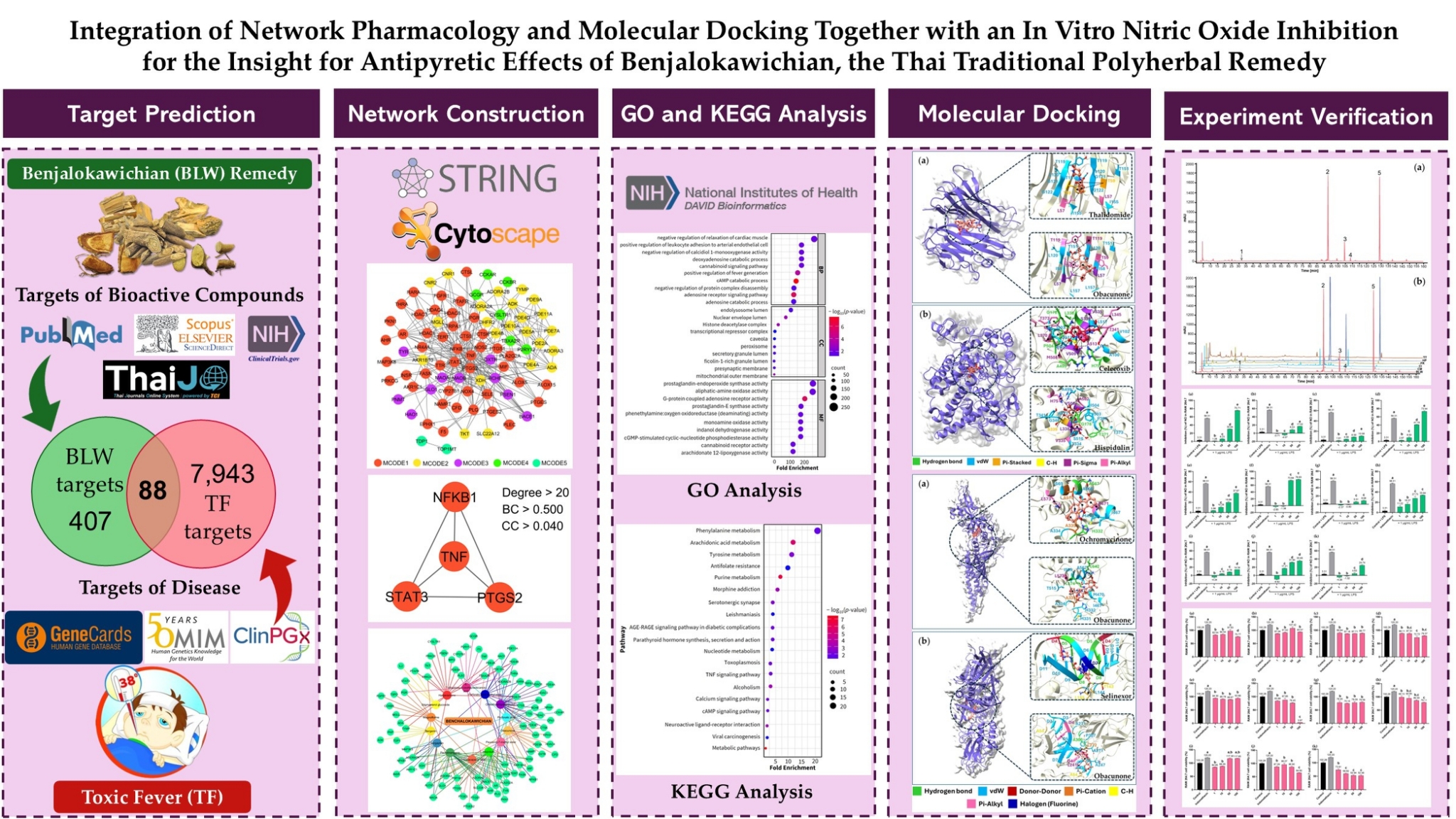
Keywords:
1. Introduction
2. Results
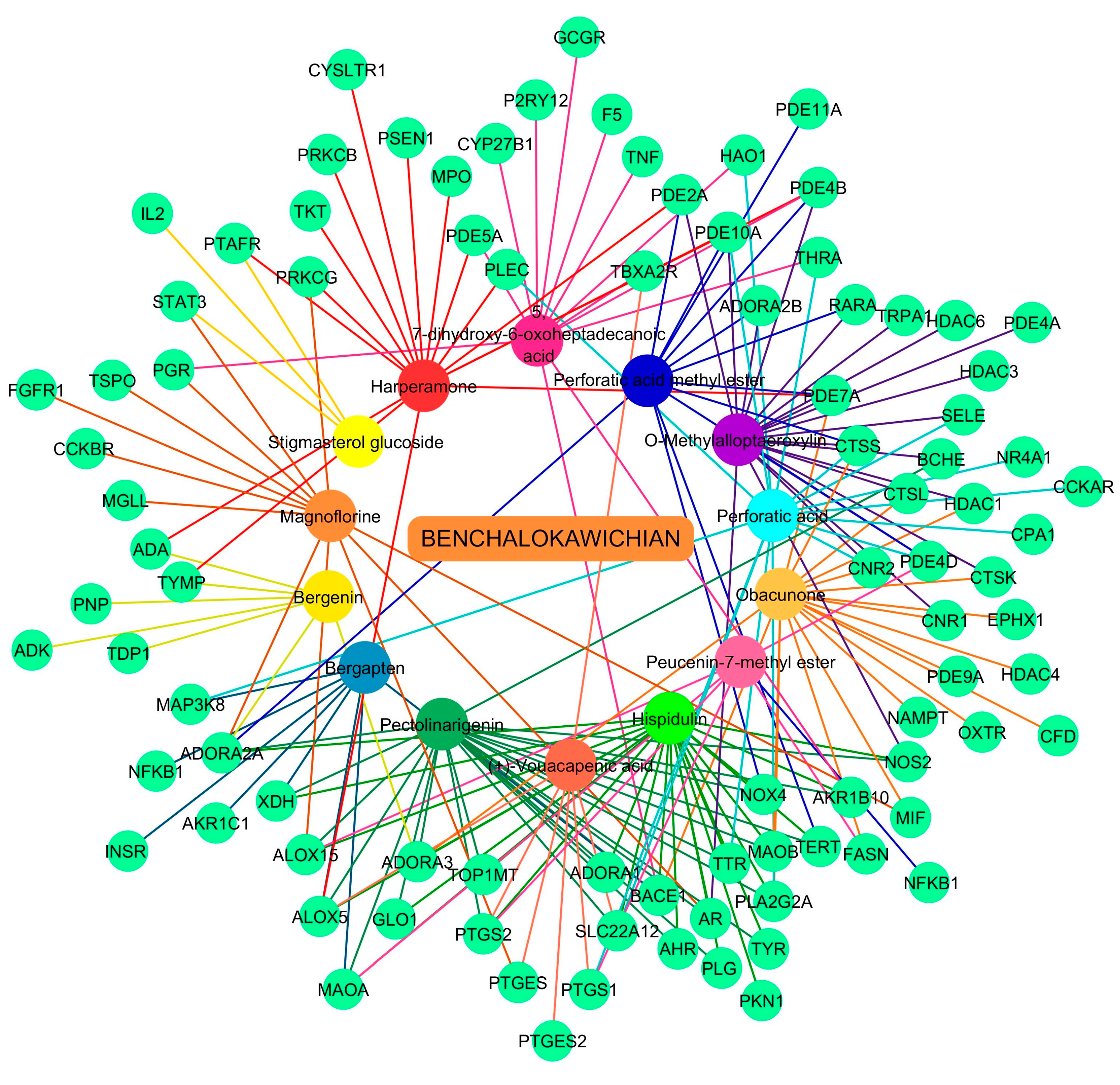
2.1. Screening of Bioactive Compounds and BLW-Related Targets Based on ADME
2.2. Obtention of Potential Therapeutic Targets of BLW for Treatment of TF
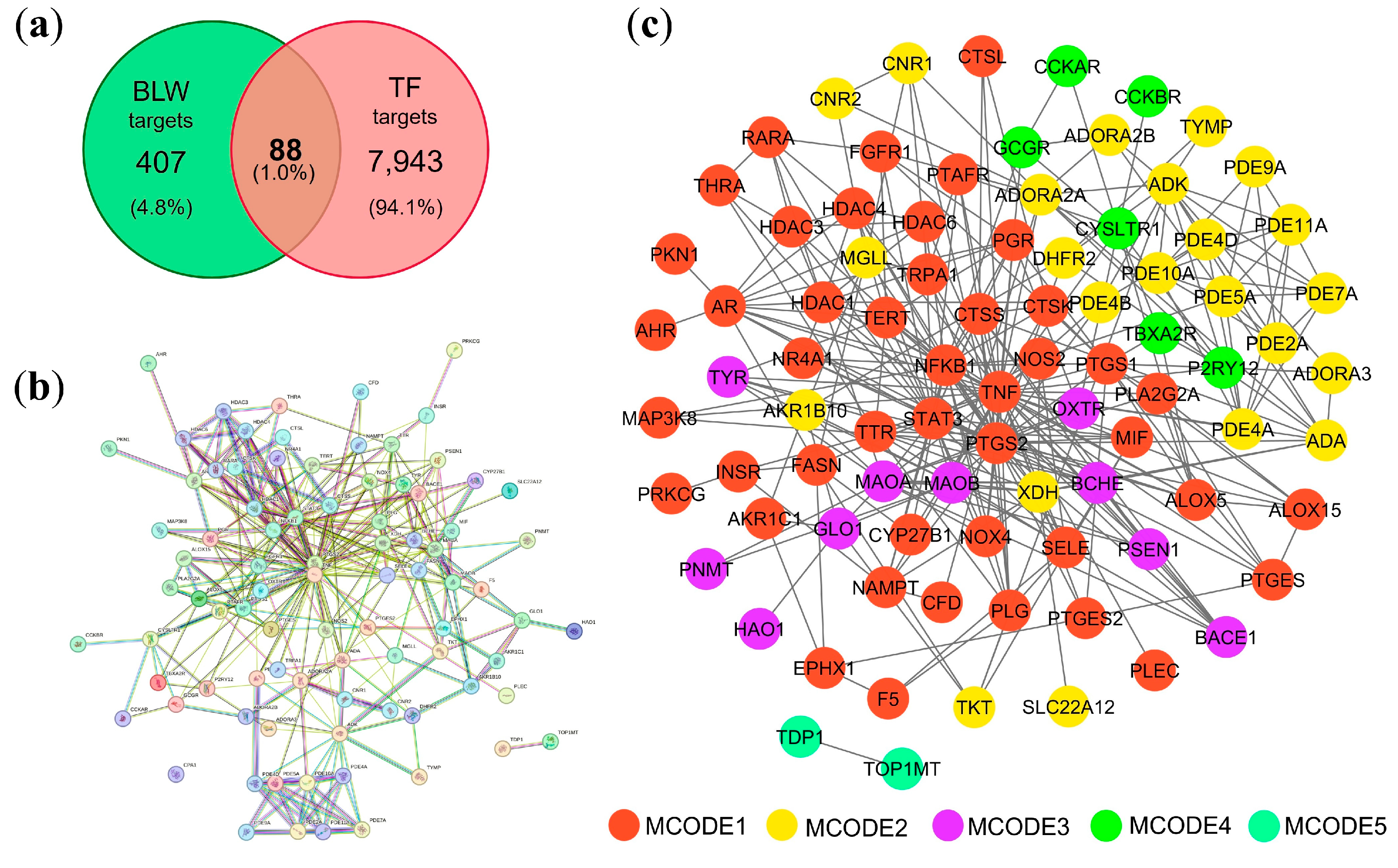
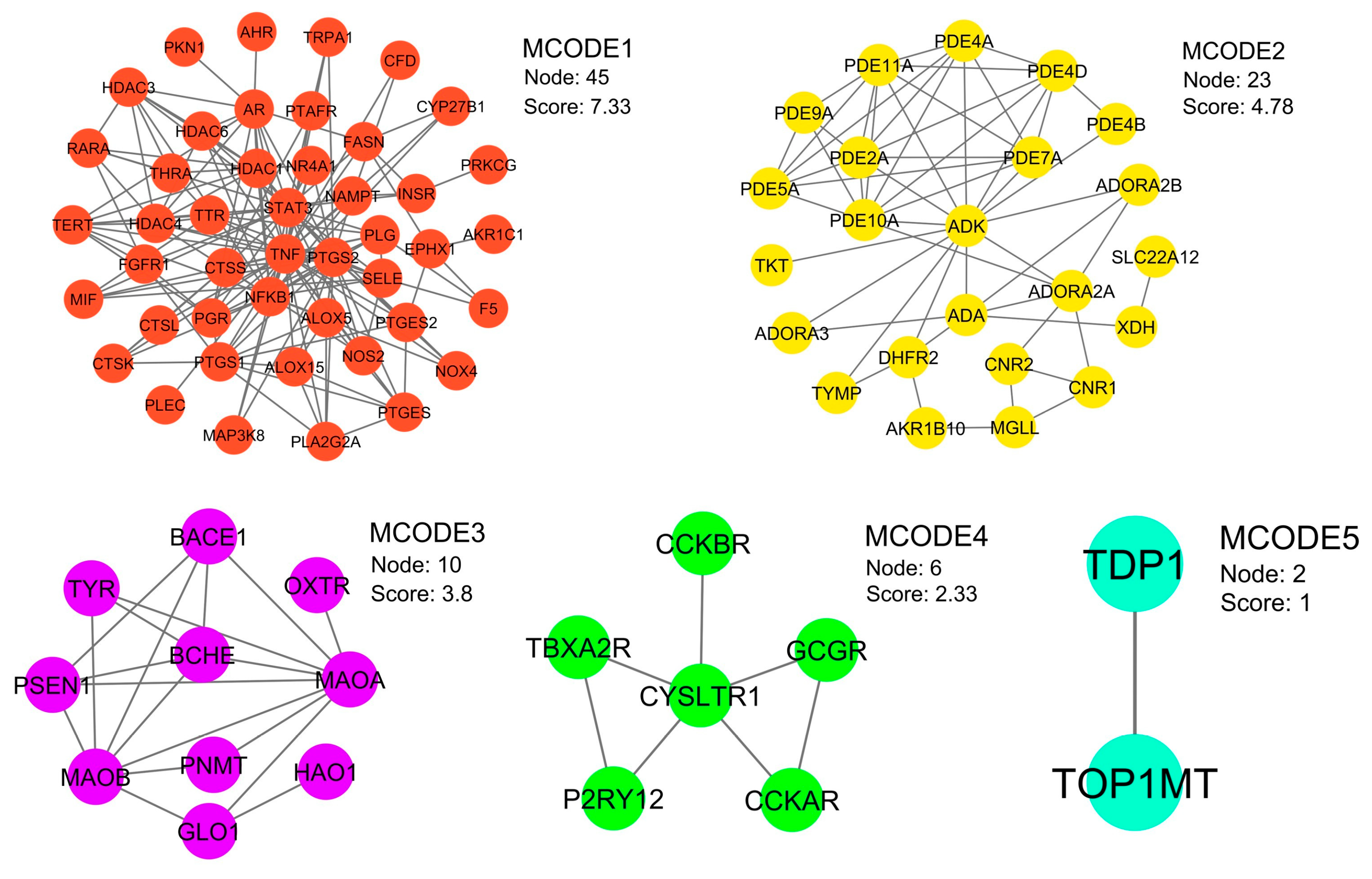
2.3. Protein–Protein Interaction (PPI) Network Visualization and Modular Analysis
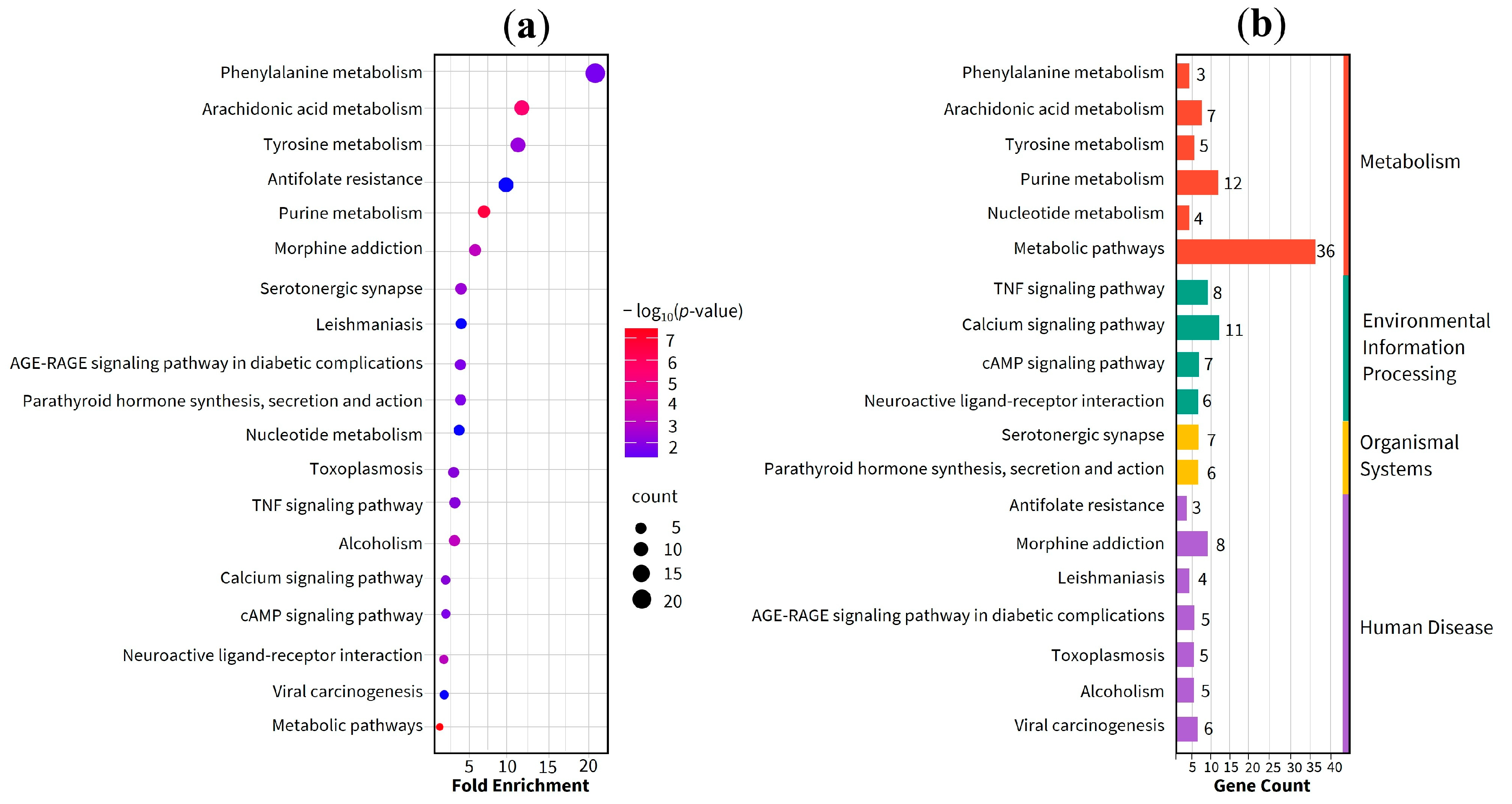
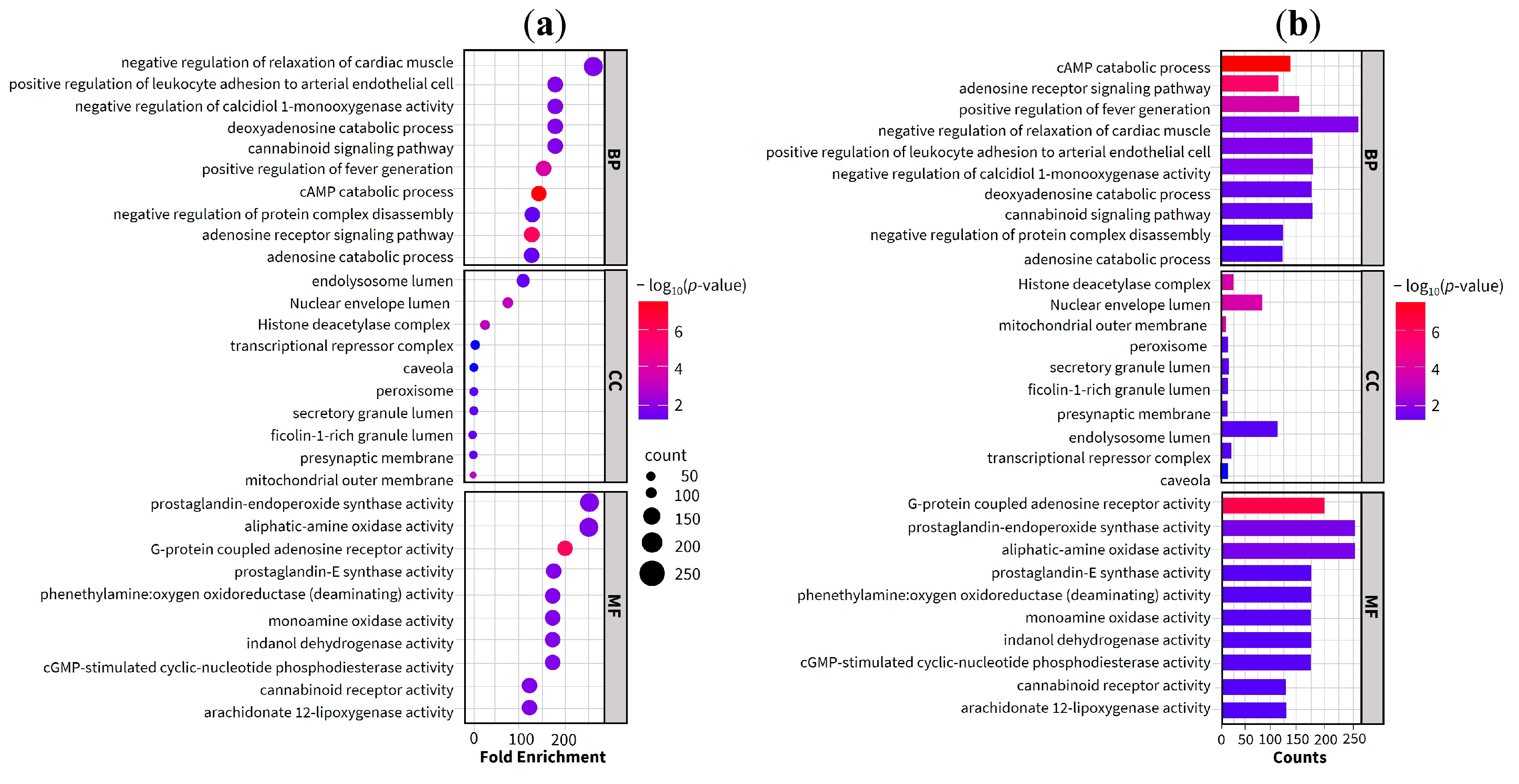
2.4. Gene Ontology (GO) and Enriched Pathway Analysis
2.5. Investigation of the Possible Therapeutic Targets of BLW for TF Treatment
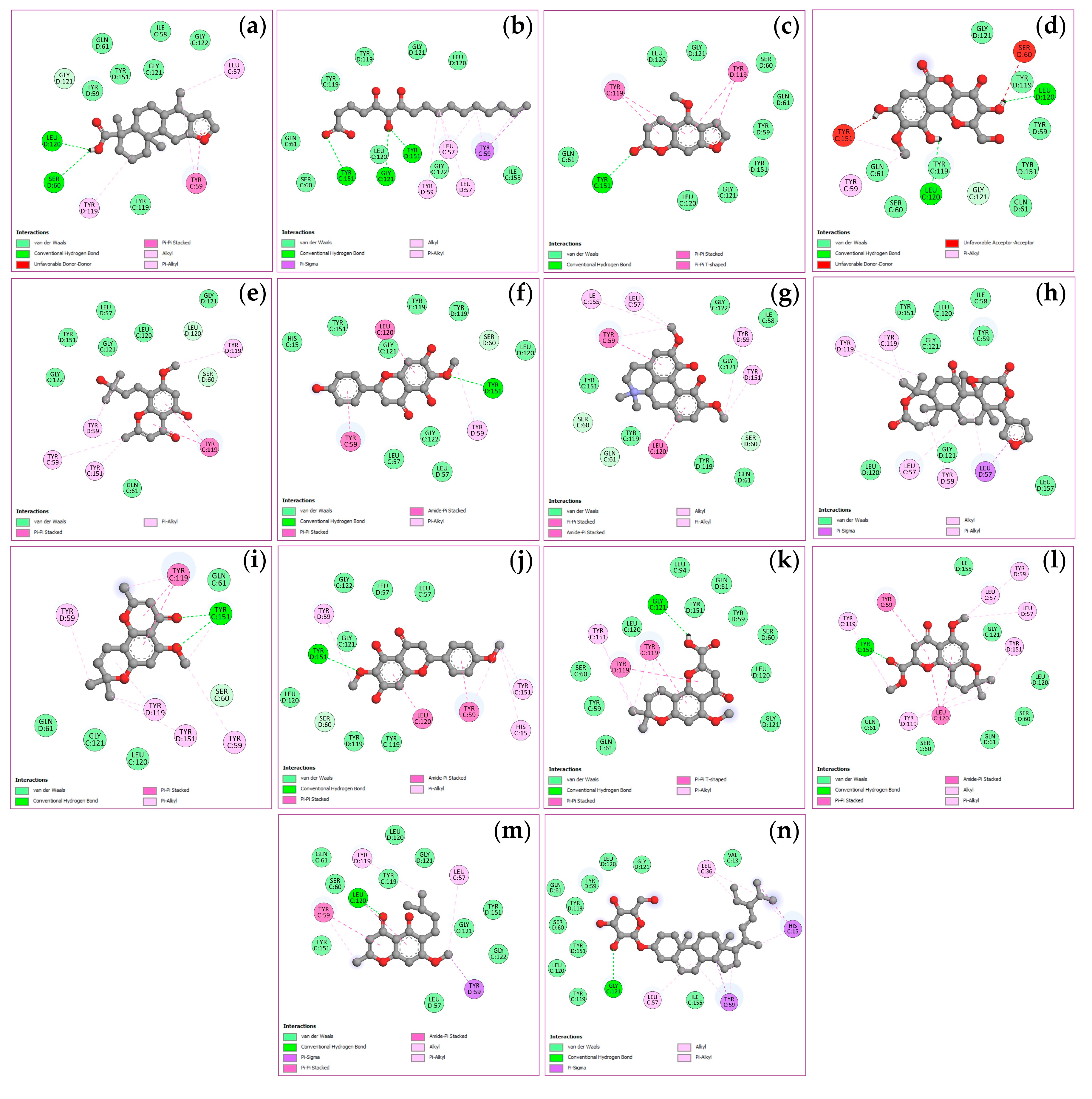
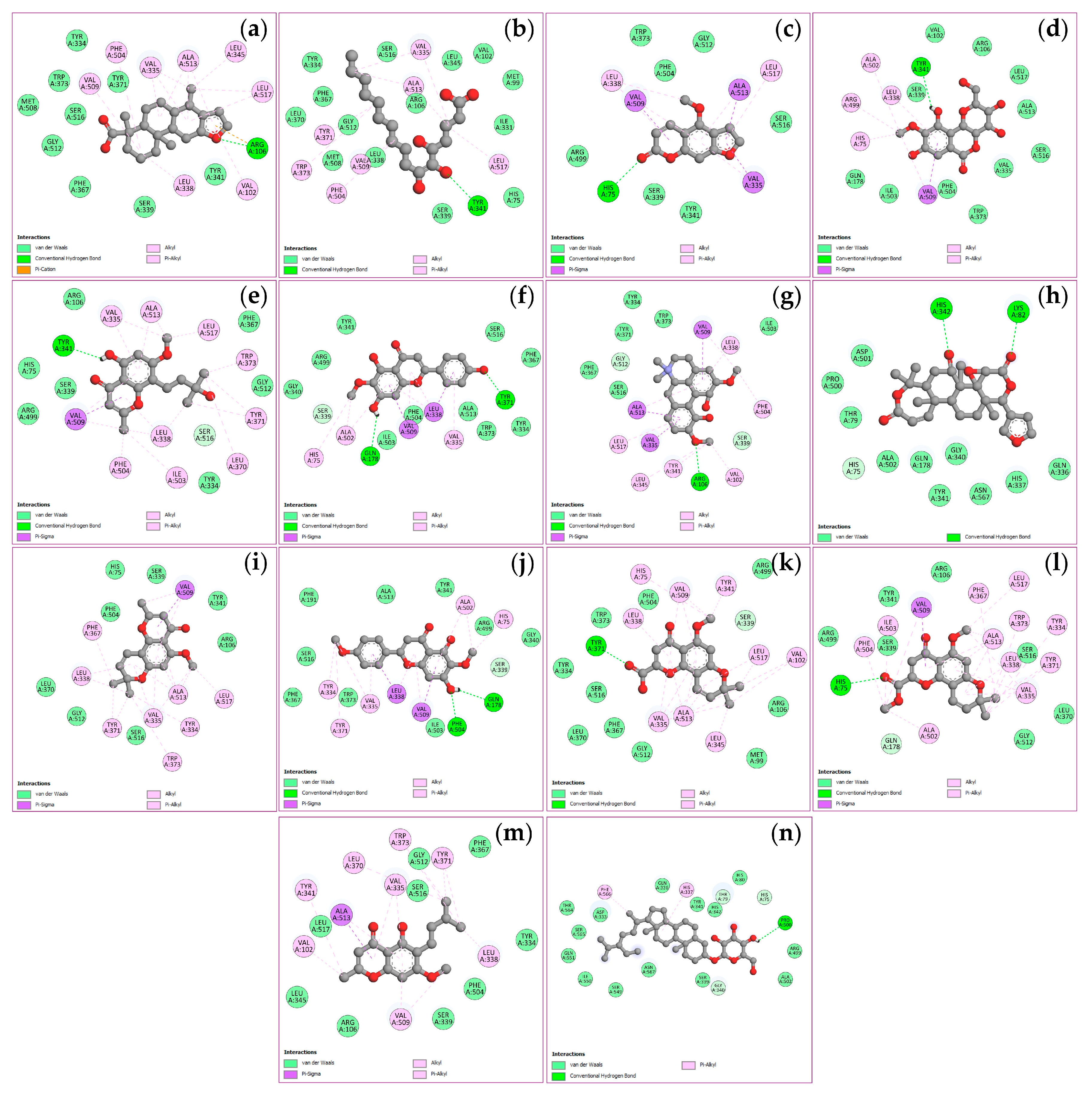
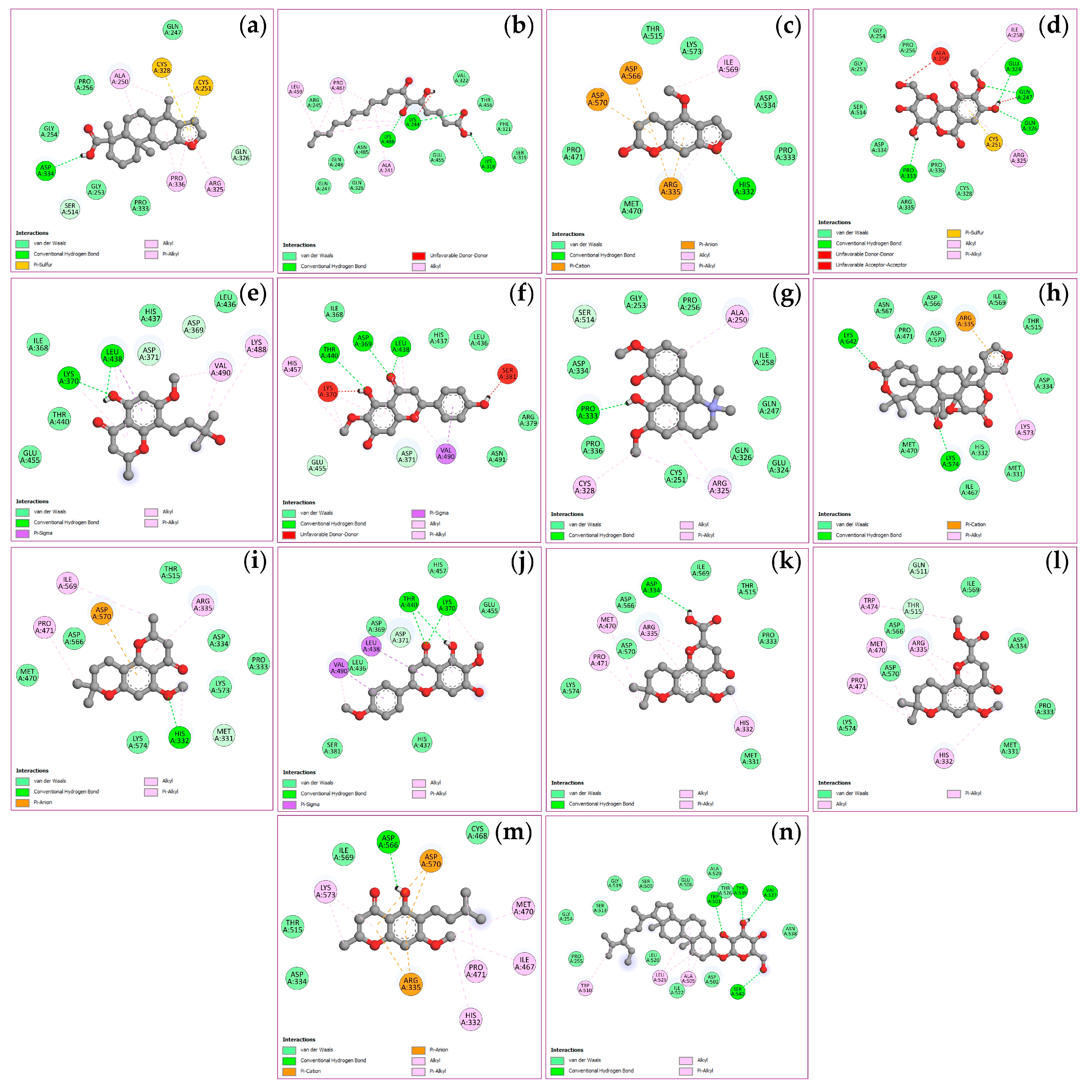
2.6. Molecular Docking of Key Targets
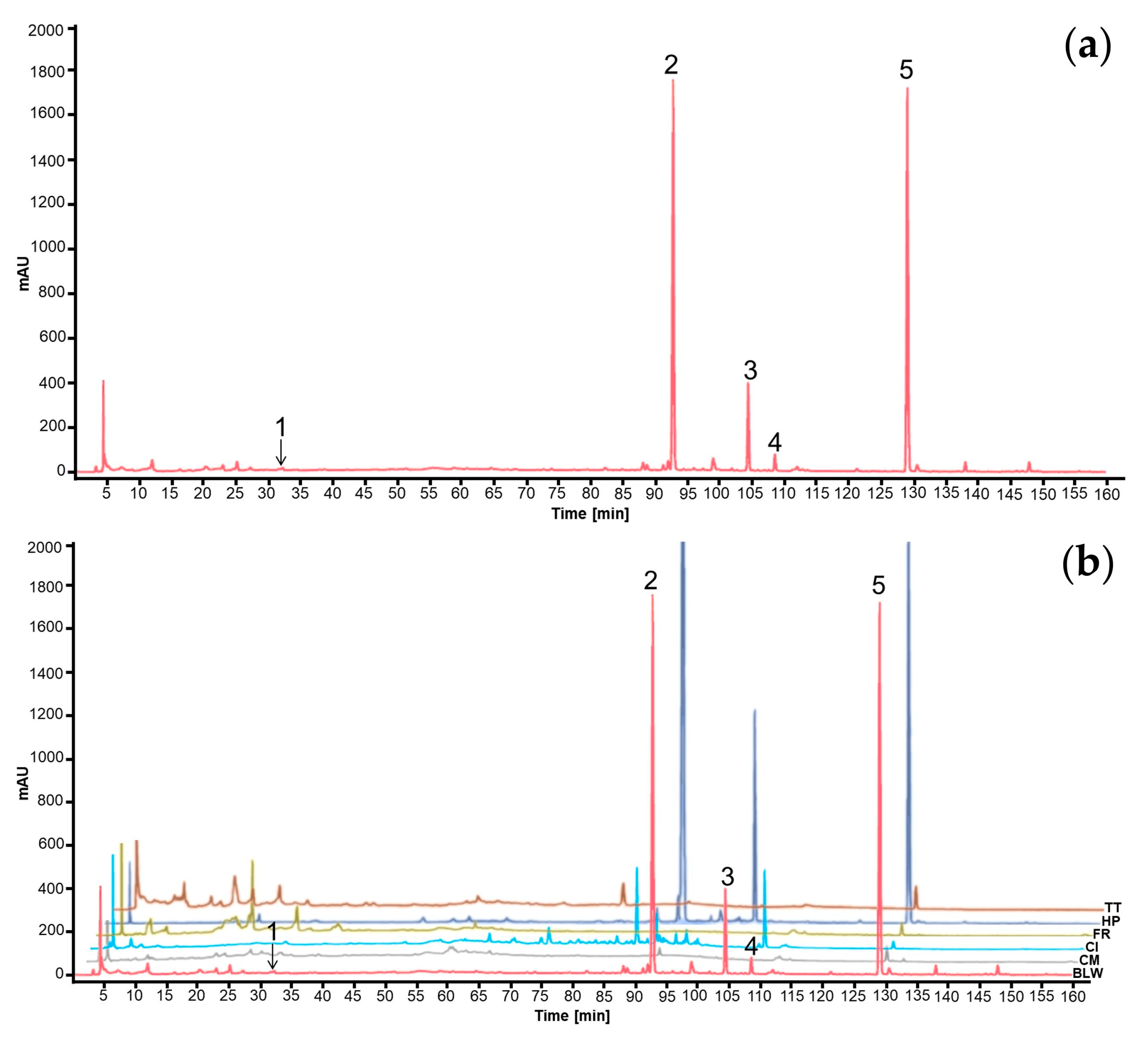
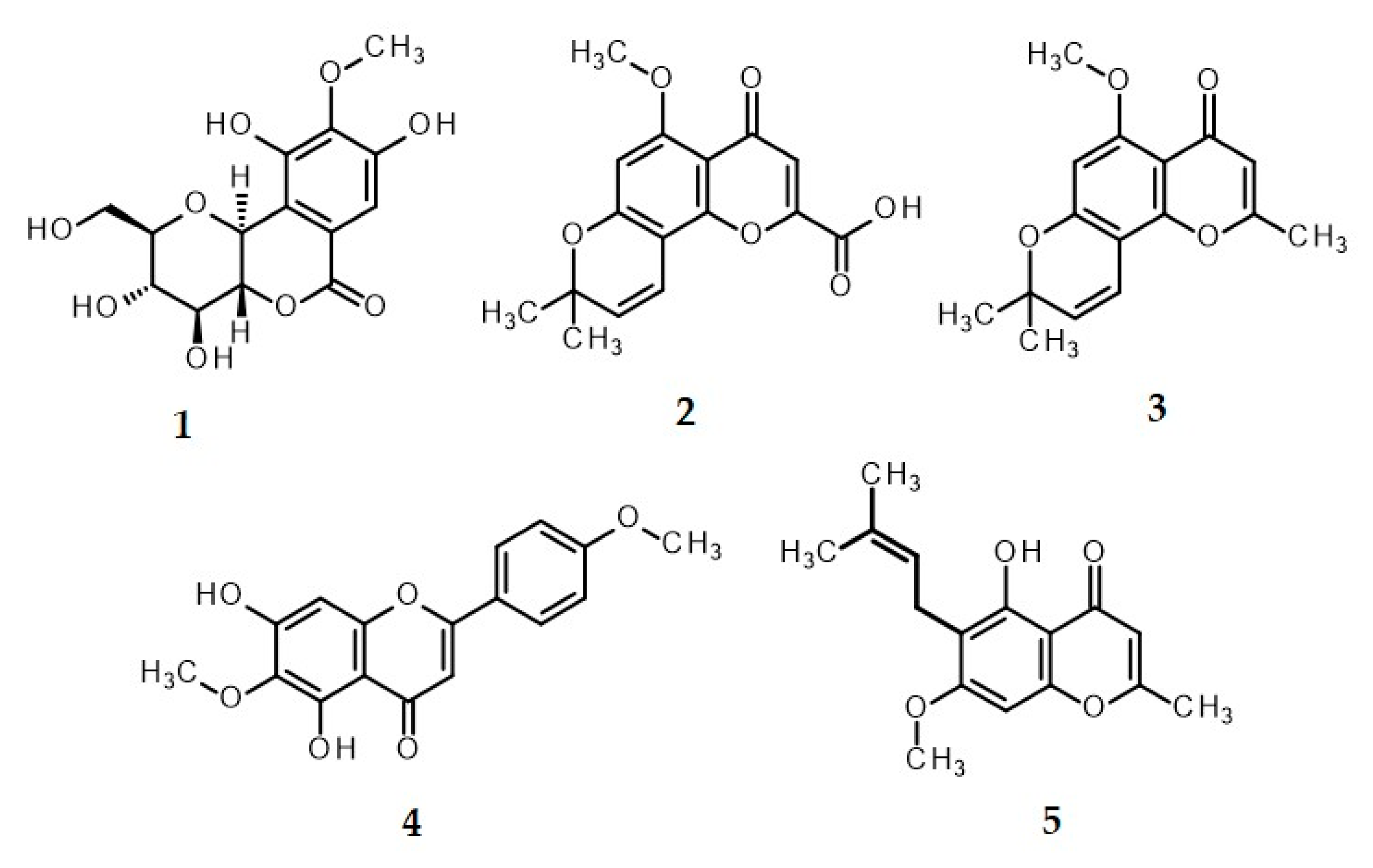
2.7. Characterization of the BLW Extract
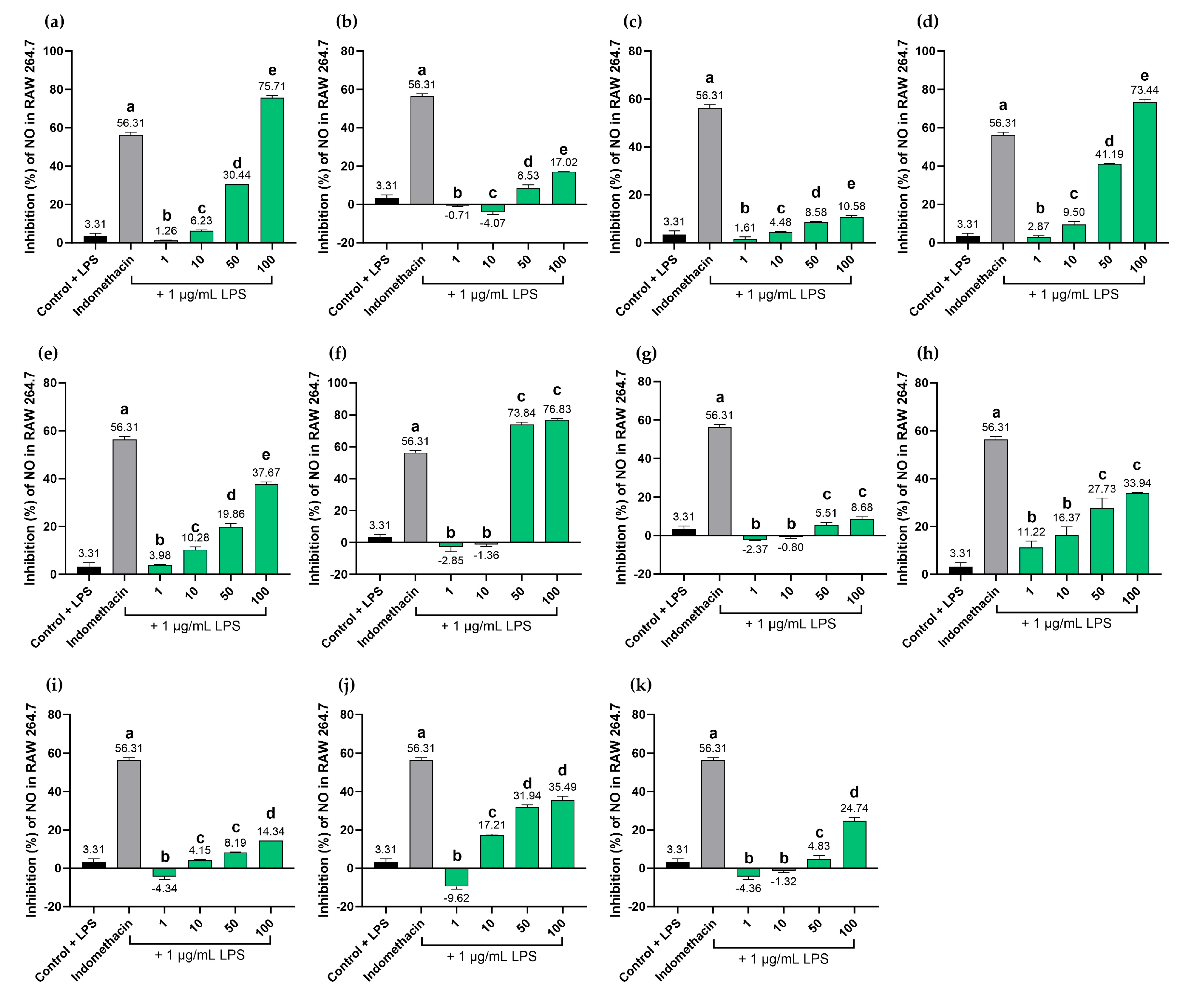
2.8. Effects of the BLW Extract and Biomarker Compounds on NO Production in LPS-Induced RAW264.7 Macrophages
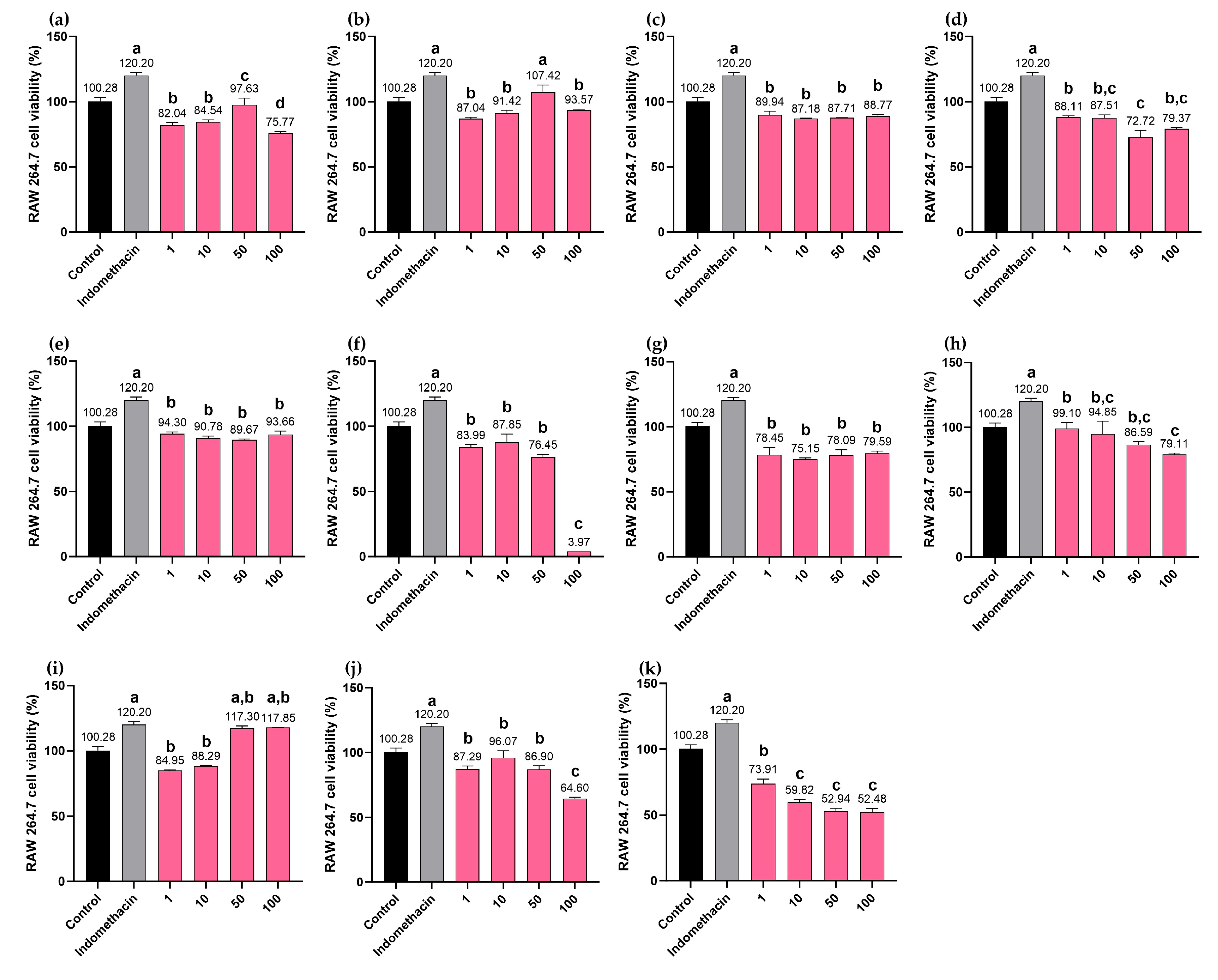
2.9. Cytotoxicity Effects of the BLW Extract and Biomarker Compounds on RAW264.7 Macrophages
3. Discussion
4. Materials and Methods
4.1. Screening for Potential Bioactive Compounds and BLW-Related Targets
4.2. Identification of TF-Related Targets
4.3. Protein-Protein Network Interaction and Modular Identification
4.4. Functional Enrichment and Pathway Analysis
4.5. Verification with Molecular Docking
4.5.1. Protein Structures and Modeling of Ligands Preparation
4.5.2. Protein-Ligand Docking
4.5.3. Docking Validation
4.6. Preparation of BLW Extract
4.7. Chemicals and Reagents
4.8. HPLC Analysis
4.9. Nitric Oxide (NO) Inhibitory Activity and Cytotoxicity of the BLW Extract and Biomarkers on RAW 264.7 Macrophages
4.10. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BLW | Benjalokawichian |
| BP | Biological Process |
| CC | Cellular Component |
| GI | Gastrointestinal |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| MCODE | Molecular Complex Detection |
| MF | Molecular Function |
| NFKB1 | Nuclear factor-kappa-B p105 subunit |
| PPI | Protein-Protein Interaction |
| PTGS2 | Prostaglandin-Endoperoxide Synthase 2 |
| STAT3 | Signal Transducer and Activator of Transcription 3 |
| STRING | Search Tool for the Retrieval of Interacting Genes |
| TF | Toxic Fever |
| TNF | Tumor Necrosis Factor-α |
| TTM | Thai Traditional Medicine |
References
- Lai, J.; Wu, H.; Qin, A. Cytokines in febrile diseases. J. Interferon Cytokine Res. 2021, 41, 1–11. [Google Scholar] [CrossRef]
- Ma, L.L.; Liu, H.M.; Luo, C.H.; He, Y.N.; Wang, F.; Huang, H.Z.; Han, L.; Yang, M.; Xu, R.C.; Zhang, D.K. Fever and antipyretic supported by traditional Chinese medicine: A multi-pathway regulation. Front. Pharmacol. 2021, 12, 583279. [Google Scholar] [CrossRef]
- Palo, T.; Thaworn, A.; Charoenkij, P.; Thamsermsang, O.; Chotewuttakorn, S.; Tripatara, P.; Laohapand, T.; Akarasereenont, P. The effects of Thai herbal Ha-Rak formula on COX isoform expression in human umbilical vein endothelial cells induced by IL-1β. Evid. Based Complement Alternat. Med. 2017, 2017, 9383272. [Google Scholar] [CrossRef]
- Wang, B.; Wu, L.; Chen, J.; Dong, L.; Chen, C.; Wen, Z.; Hu, J.; Fleming, I.; Wang, D.W. Metabolism pathways of arachidonic acids: Mechanisms and potential therapeutic targets. Sig. Transduct. Target. Ther. 2021, 6, 94. [Google Scholar] [CrossRef] [PubMed]
- Tredicine, M.; Mucci, M.; Recchiuti, A.; Mattoscio, D. Immunoregulatory mechanisms of the arachidonic acid pathway in cancer. FEBS Lett. 2025, 599, 927–951. [Google Scholar] [CrossRef] [PubMed]
- Subin, P.; Sabuhom, P.; Naladta, A.; Luecha, P.; Nualkaew, S.; Nualkaew, N. An evaluation of the anti-inflammatory effects of a Thai traditional polyherbal recipe TPDM6315 in LPS-induced RAW264.7 macrophages and TNF-induced 3T3-L1 adipocytes. Curr. Issues Mol. Biol. 2023, 45, 4891–4907. [Google Scholar] [CrossRef] [PubMed]
- Balli, S.; Shumway, K.R.; Sharan, S. Physiology, fever. In StatPearls [Internet]; Treasure Island (FL), Ed.; StatPearls: Florida, USA, 2023; Available online: https://www.ncbi.nlm.nih.gov/books/NBK562334/ (accessed on 12 August 2024).
- Shimizu, M. Clinical features of cytokine storm syndrome. Adv. Exp. Med. Biol. 2024, 1448, 33–42. [Google Scholar]
- Traditional Thai Medicine Rehabilitation Foundation, Ayurveda College (Jivaka Kumar Bhaccha). Original Thai traditional medical textbook (Paetsart Sonkau): Conservation edition, 3rd ed.; Printing Press of Chulalongkorn University: Bangkok, Thailand, 2015; pp. 345–356. [Google Scholar]
- Chaloemram, C. Scientific-based explanation of Thai traditional medicine theory for Thai traditional herbal remedy (A case study of Mo-Ha-Rak formula). Doctoral dissertation, Mahasarakham University, Maha Sarakham, Thailand, January 2025. [Google Scholar]
- National Drug Committee. List of herbal medicine products A.D.; Agricultural Co-operative Federation of Thailand, Ltd. : Nonthaburi, Thailand, 2006. [Google Scholar]
- Suttana, W.; Singharachai, C.; Charoensup, R.; Rujanapun, N.; Suya, C. Antiproliferative and apoptosis-inducing activities of Benchalokawichian remedy against doxorubicin-sensitive and -resistant erythromyelogenous leukemic cells. CMU J. Nat. Sci. 2021, 20, e2021056. [Google Scholar] [CrossRef]
- Herbal Products Division, Food and Drug Administration. National list of essential herbal medicines 2023; Minnie Group Co., Ltd.: Pathum Thani, Thailand, 2023; pp. 73–74. [Google Scholar]
- Wang, X-H.; Li, Y-H.; Zhang, J-C.; Li, Z.; Liu, G-X.; Zhang, T.; Zhang, M-Y. Analysis of the effect of Bupleurum on fever in Xiaochaihu Decoction based on network pharmacology. J. Hainan Med. Univ. 2021, 27, 55–59. [Google Scholar]
- Pei, K.; Wang, Y.; Guo, W.; Lin, H.; Lin, Z.; Lv, G. Antipyretic mechanism of Bai Hu Tang on LPS-induced fever in rat: A network pharmacology and metabolomics analysis. Pharmaceuticals 2025, 18, 610. [Google Scholar] [CrossRef]
- Zhang, H.; Ge, S.; Diao, F.; Song, W.; Zhang, Y.; Zhuang, P.; Zhang, Y. Network pharmacology integrated with experimental verification reveals the antipyretic characteristics and mechanism of Zi Xue powder. Pharm. Biol. 2023, 61, 1512–1524. [Google Scholar] [CrossRef]
- Zhou, W.; Zhang, H.; Wang, X.; Kang, J.; Guo, W.; Zhou, L.; Liu, H.; Wang, M.; Jia, R.; Du, X.; Wang, W.; Zhang, B.; Li, S. Network pharmacology to unveil the mechanism of Moluodan in the treatment of chronic atrophic gastritis. Phytomedicine 2022, 95, 153837. [Google Scholar] [CrossRef]
- Chen, W.; Li, Y.; Zhang, C.; Zhou, H.; Ma, J.; Vaishnani, D.K.; Zeng, B.; Yu, J.; Mao, H.; Zheng, J. Multi-omics and experimental validation reveal anti-HCC mechanisms of Tibetan Liwei Muxiang pill and quercetin. Pharmaceuticals 2025, 18, 900. [Google Scholar] [CrossRef] [PubMed]
- Chaloemram, C.; Rattarom, R.; Kijjoa, A.; Nualkaew, S. Integration of network pharmacology, molecular docking and in vitro nitric oxide inhibition assay to explore the mechanism of action of Thai traditional polyherbal remedy, Mo-Ha-Rak, in the treatment of prolonged fever. Pharmaceuticals 2025, 18, 1541. [Google Scholar] [CrossRef]
- Patel, V.R.; Saini, S.; Dwivedi, J.; Gupta, A.K.; Shrivastava, A.K.; Misra, A. Exploring the concept and scope of polyherbal formulations: A comprehensive review. Int. J. Herb. Med. 2025, 13, 9–16. [Google Scholar] [CrossRef]
- Dubey, S.; Dixit, A.K. Preclinical evidence of polyherbal formulations on wound healing: A systematic review on research trends and perspectives. J. Ayurveda Integr. Med. 2023, 14, 100688. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Qi, X.; Yang, C.; Wang, S.; Jiang, J.; Wang, L.; Song, J.; Yu, B.; Sun, M. Network pharmacology, molecular docking, and in vitro experiments reveal the role and mechanism of Tanshinone IIA in colorectal cancer treatment through the PI3K/AKT Pathway. Drug Des. Devel. Ther. 2025, 19, 2959–2977. [Google Scholar] [CrossRef] [PubMed]
- Nadjar, A.; Bluthé, RM.; May, M.J.; Dantzer, R.; Parnet, P. Inactivation of the cerebral NFκB pathway inhibits interleukin-1β-induced sickness behavior and c-Fos expression in various brain nuclei. Neuropsychopharmacol 2005, 30, 1492–1499. [Google Scholar] [CrossRef]
- Juckmeta, T. Biological activities of the ethanolic extracts from Pikut Benjalokawichian (Ha-Rak) and its isolate compounds. Master’s Thesis, Thammasat University, Bangkok, Thailand, 2011. [Google Scholar]
- Feng, Y.; Bhandari, R.; Li, C.; Shu, P.; Shaikh, I.I. Pectolinarigenin suppresses LPS-induced inflammatory response in macrophages and attenuates DSS-induced colitis by modulating the NF-κB/Nrf2 signaling pathway. Inflammation 2022, 45, 2529–2543. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.S.; Yang, X.; Zhang, W.X.; Zhou, Y.; Wei, T.T.; Cui, N.; Du, J.; Liu, W.; Lu, Q.B. Metabolic alterations in urine among the patients with severe fever with thrombocytopenia syndrome. Virol. J. 2024, 21, 11. [Google Scholar] [CrossRef]
- Luporini, R.L.; Pott-Junior, H.; Di Medeiros Leal, M.C.B.; Castro, A.; Ferreira, A.G.; Cominetti; de Freitas Anibal, M.R.F. Phenylalanine and COVID-19: Tracking disease severity markers. Int. Immunopharmacol. 2021, 101, 108313. [Google Scholar] [CrossRef]
- Kaewnoi, A.; Duanyai, S.; Vallisuta, O.; Yahuafai, J. Comparison of in vitro anti-inflammatory activity of extracts from original Ya-Ha-Rak and adapted formula. J. Basic. App. Pharmacol. 2024, 4, O98–O107. [Google Scholar]
- Booranasubkajorn, S.; Kanlaya, H.; Huabprasert, S.; Lumlerdkij, N.; Akarasereenont, P.; Tripatara, P. The effect of Thai herbal Ha-Rak formula (HRF) on LPS-induced systemic inflammation in Wistar rats. Siriraj Med. J. 2017, 69, 356–362. [Google Scholar]
- Konsue, A.; Sattayasai, J.; Puapairoj, P.; Picheansoonthon, C. Antipyretic effects of Bencha-Loga-Wichien herbal drug in rats. Thai J. Pharmacol. 2008, 29, 79–82. [Google Scholar]
- Jongchanapong, A.; Singharachai, C.; Palanuvej, C.; Ruangrungsi, N.; Towiwat, P. Antipyretic and antinociceptive effects of Ben-Cha-Lo-Ka-Wi-Chian remedy. J. Health Res. 2010, 24, 15–22. [Google Scholar]
- Luo, X.; Yu, Z.; Yue, B.; Ren, J.; Zhang, J.; Mani, S.; Wang, Z.; Dou, W. Obacunone reduces inflammatory signalling and tumour occurrence in mice with chronic inflammation-induced colorectal cancer. Pharm. Biol. 2020, 58, 886–897. [Google Scholar] [CrossRef]
- Choodej, S.; Sommit, D.; Pudhom, K. Rearranged limonoids and chromones from Harrisonia perforata and their anti-inflammatory activity. Bioorg. Med. Chem. Lett. 2013, 23, 3896–3900. [Google Scholar] [CrossRef]
- Tangsongcharoen, T.; Issaravanich, S.; Palanuvej, C.; Ruangrungsi, N. Quantitative analysis of hispidulin content in Clerodendrum petasites roots distributed in Thailand. Pharmacog. J. 2019, 11, 1093–1099. [Google Scholar] [CrossRef]
- Yu, CI.; Cheng, C.I.; Kang, Y.F.; Chang, P.C.; Lin, I.P.; Kuo, Y.H.; Jhou, A.J.; Lin, M.Y.; Chen, C.Y.; Lee, C.H. Hispidulin inhibits neuroinflammation in lipopolysaccharide-activated BV2 microglia and attenuates the activation of Akt, NF-κB, and STAT3 pathway. Neurotox. Res. 2020, 38, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Jain, R.; Rawat, S.; Jain, S.C. Phytochemicals and antioxidant evaluation of Ficus racemosa root bark. J. Pharm. Res. 2013, 6, 615–619. [Google Scholar] [CrossRef]
- Boonsong, N.; Preeprame, S.; Putalun, W. Quantitative determination of bergenin in callus, twig and root of Ficus racemosa L. extract by high performance liquid chromatography. KKU Res. J. (GS) 2022, 22, 87–98. [Google Scholar]
- Stitmannaitham, M. Isolation and structural determination of compounds from roots of Harrisonia perforata Merr. Master’s Thesis, Chulalongkorn University, Bangkok, Thailand, 1992. [Google Scholar]
- Thadaniti, S.; Archakunakorn, W.; Tuntiwachwuttikul, P.; Bremner, J.B. Chromones from Harrisonia perforata (Blanco.) Merr. J. Sci. Soc. Thailand 1994, 20, 183–187. [Google Scholar] [CrossRef]
- Chunthorng-Orn, J.; Pipatrattanaseree, W.; Juckmeta, T.; Dechayont, B.; Phuaklee, P.; Itharat, A. Quality evaluation and pectolinarigenin contents analysis of Harak remedy in Thailand. J. Health Sci. Altern. Med. 2019, 1, 25–33. [Google Scholar]
- Sakpakdeejaroen, I.; Juckmeta, T.; Itharat, A. Development and validation of RP-HPLC method to determine anti-allergic compound in Thai traditional remedy called Benjalokawichien. J. Med. Assoc. Thai. 2014, 97, S76–S80. [Google Scholar]
- Gao, X.J.; Guo, M.Y.; Zhang, Z.C.; Wang, T.C.; Cao, Y.G.; Zhang, N.S. Bergenin plays an anti-inflammatory role via the modulation of MAPK and NF-κB signaling pathways in a mouse model of LPS-induced mastitis. Inflammation 2015, 38, 1142–1150. [Google Scholar] [CrossRef] [PubMed]
- Alanazi, S.T.; Salama, S.A.; Althobaiti, M.M.; Alotaibi, R.A.; AlAbdullatif, A.A.; Musa, A.; Harisa, G.I. Alleviation of copper-induced hepatotoxicity by bergenin: Diminution of oxidative stress, inflammation, and apoptosis via targeting SIRT1/FOXO3a/NF-κB axes and p38 MAPK signaling. Biol. Trace. Elem. Res. 2025, 203, 3195–3207. [Google Scholar] [CrossRef]
- Juckmeta, T.; Pipatrattanaseree, W.; Jaidee, W.; Dechayont, B.; Chunthorng-Orn, J.; Andersen, R.J.; Itharat, A. Cytotoxicity to five cancer cell lines of the respiratory tract system and anti-inflammatory activity of Thai traditional remedy. Nat. Prod. Commun. 2019, 14, 1–6. [Google Scholar] [CrossRef]
- Nutmakul, T.; Pattanapanyasat, K.; Soonthornchareonnon, N.; Shiomi, K.; Mori, M.; Prathanturarug, S. Antiplasmodial activities of a Thai traditional antipyretic formulation, Bencha-Loga-Wichian: A comparative study between the roots and their substitutes, the stems. J. Ethnopharmacol. 2016, 193, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Chuchote, C.; Somwong, P. Similarity analysis of the chromatographic fingerprints of Thai herbal Ya-Ha-Rak remedy using HPLC. Interprof. J. Health Sci. 2019, 17, 55–63. [Google Scholar]
- Somwong, P.; Chuchote, C. Determination of lupeol, a cytotoxic compound against SW620 cells in the extracts of Ha-Rak recipe. Pharmacogn J. 2021, 13, 133–138. [Google Scholar] [CrossRef]
- Ryszkiewicz, P.; Schlicker, E.; Malinowska, B. Is inducible nitric oxide synthase (iNOS) promising as a new target against pulmonary hypertension? Antioxidants 2025, 14, 377. [Google Scholar] [CrossRef] [PubMed]
- Soufli, I.; Toumi, R.; Rafa, H.; Touil-Boukoffa, C. Overview of cytokines and nitric oxide involvement in immuno-pathogenesis of inflammatory bowel diseases. World J. Gastrointest. Pharmacol. Ther. 2016, 7, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Pal, P.P.; Begum, A.S.; Basha, S.A.; Araya, H.; Fujimoto, Y. New natural pro-inflammatory cytokines (TNF-α, IL-6 and IL-1β) and iNOS inhibitors identified from Penicillium polonicum through in vitro and in vivo studies. Int. Immunopharmacol. 2023, 117, 109940. [Google Scholar]
- Pratama, R.R.; Sari, R.A.; Sholikhah, I.; Mansor, H.; Chang, H-I.; Sukardiman; Widyowati, R. Inhibition of nitric oxide production in RAW 264.7 cells and cytokines IL-1β in osteoarthritis rat models of 70% ethanol extract of Arcangelisia flava (L.) merr stems. Heliyon 2024, 10, e35730. [Google Scholar] [CrossRef] [PubMed]
- Juckmeta, T.; Itharat, A. Anti-inflammatory and antioxidant activities of Thai traditional remedy called “Ya-ha-rak”. J. Health Res. 2012, 26, 205–210. [Google Scholar]
- Chandranipapongse, W.; Palo, T.; Chotewuttakorn, S.; Tripatara, P.; Booranasubkajorn, S.; Laohapand, T.; Akarasereenont, P. Study the effect of an antipyretic drug, Thai herbal Ha-Rak formula on platelet aggregation in healthy Thai volunteers: A randomized, placebo-controlled trial. Siriraj Med. J. 2017, 69, 283–289. [Google Scholar]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; Wang, J.; Yu, B.; Zhang, J.; Bryant, S.H. PubChem substance and compound databases. Nucleic Acids Res. 2016, 44, D1202–D1213. [Google Scholar] [CrossRef]
- Gfeller, D.; Grosdidier, A.; Wirth, W.; Daina, A.; Michielin, O.; Zoete, V. Swisstarget prediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res. 2014, 42, W32–W38. [Google Scholar] [CrossRef]
- Safran, M.; Dalah, I.; Alexander, J.; Rosen, N.; Stein, T.I.; Shmoish, M.; Nativ, N.; Bahir, I.; Doniger, T.; Krug, H.; Sirota-Madi, A.; Olender, T.; Golan, Y.; Stelzer, G.; Harel, A.; Lancet, D. GeneCards version 3: The human gene integrator. Database (Oxford) 2010, 2010, baq020. [Google Scholar] [CrossRef]
- Amberger, J.S.; Bocchini, C.A.; Schiettecatte, F.; Scott, A.F.; Hamosh, A. OMIM.org: Online mendelian inheritance in man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 2015, 43, D789–D798. [Google Scholar] [CrossRef]
- Whirl-Carrillo, M.; McDonagh, E.M.; Hebert, J.M.; Gong, L.; Sangkuhl, K.; Thorn, C.F.; Altman, R.B.; Klein, T.E. An evidence-based framework for evaluating pharmacogenomics knowledge for personalized medicine. Clin. Pharm. Therap. 2021, 92, 414–417. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; Jensen, E.J.; von Mering, C. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022, 50, W216–W221. [Google Scholar] [CrossRef]
- Salman, H.A.; Yaakop, A.S.; Aladaileh, S.; Mustafa, M.; Gharaibeh, M.; Kahar, U.M. Inhibitory effects of Ephedra alte on IL-6, hybrid TLR4, TNF-α, IL-1β, and extracted TLR4 receptors: In silico molecular docking. Heliyon 2023, 9, e12730. [Google Scholar] [CrossRef]
- Shrivastava, N.; Joshi, J.; Sehgal, N.; Kumar, I.P. Cyclooxygenase-2 identified as a potential target for novel radiomodulator scopolamine methyl bromide: An in silico study. Inform. Med. Unlocked 2017, 9, 18–25. [Google Scholar] [CrossRef]
- Liu, W.; Chu, Z.; Yang, C.; Yang, Y.; Wu, H.; Sun, J. Discovery of potent STAT3 inhibitors using structure-based virtual screening, molecular dynamic simulation, and biological evaluation. Front. Oncol. 2023, 13, 1287797. [Google Scholar] [CrossRef]
- Piccagli, L.; Fabbri, E.; Borgatti, M.; Bezzerri, V.; Mancini, I.; Nicolis, E.; Dechecchi, M.C.; Lampronti, I.; Cabrini, G.; Gambari, R. Docking of molecules identified in bioactive medicinal plants extracts into the p50 NF-kappaB transcription factor: Correlation with inhibition of NF-kappaB/DNA interactions and inhibitory effects on IL-8 gene expression. BMC Struct. Biol. 2008, 8, 38. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Shivaleela, B.; Srushti, S.C.; Shreedevi, S.J.; Babu, R.L. Thalidomide-based inhibitor for TNF-α: Designing and in silico evaluation. Futur. J. Pharm. Sci. 2022, 8, 5. [Google Scholar] [CrossRef]
- Antoniou, K.; Malamas, M.; Drosos, A.A. Clinical pharmacology of celecoxib, a COX-2 selective inhibitor. Expert Opin. Pharmacother. 2007, 8, 1719–1732. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, K.; Takaishi, M.; Nakajima, K.; Ikeda, M.; Kanda, T.; Tarutani, M.; Iiyama, T.; Asao, N.; DiGiovanni, J.; Sano, S. STAT3 as a therapeutic target for the treatment of psoriasis: A clinical feasibility study with STA-21, a STAT3 inhibitor. J. Invest. Dermatol. 2011, 131, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Kashyap, T.; Argueta, C.; Aboukameel, A.; Unger, T.J.; Klebanov, B.; Mohammad, R.M.; Muqbil, I.; Azmi, A.S.; Drolen, C.; Senapedis, W.; Lee, M.; Kauffman, M.; Shacham, S.; Landesman, Y. Selinexor, a selective inhibitor of nuclear export (SINE) compound, acts through NF-κB deactivation and combines with proteasome inhibitors to synergistically induce tumor cell death. Oncotarget 2016, 7, 78883–78895. [Google Scholar] [CrossRef]
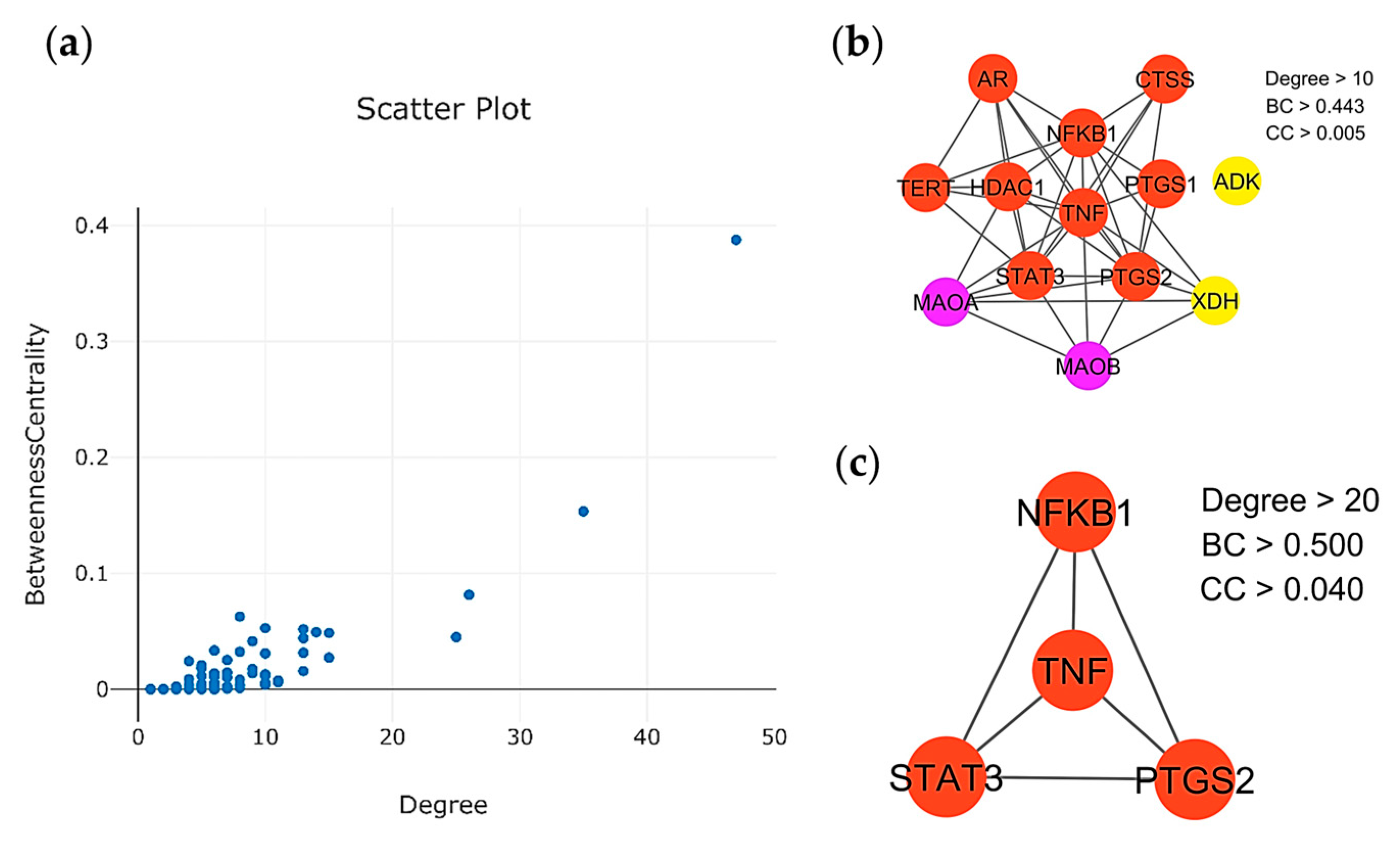

| Target | Degree | Betweenness centrality | Closeness centrality | Type |
|---|---|---|---|---|
| TNF | 47 | 0.669354 | 0.387608 | Cytokines |
| PTGS2 | 35 | 0.588652 | 0.153506 | Oxidoreductase |
| STAT3 | 26 | 0.532051 | 0.081417 | Transcription |
| NFKB1 | 25 | 0.515527 | 0.044900 | Transcription |
| MCODE | Pathway | Description | Fold enrichment |
|---|---|---|---|
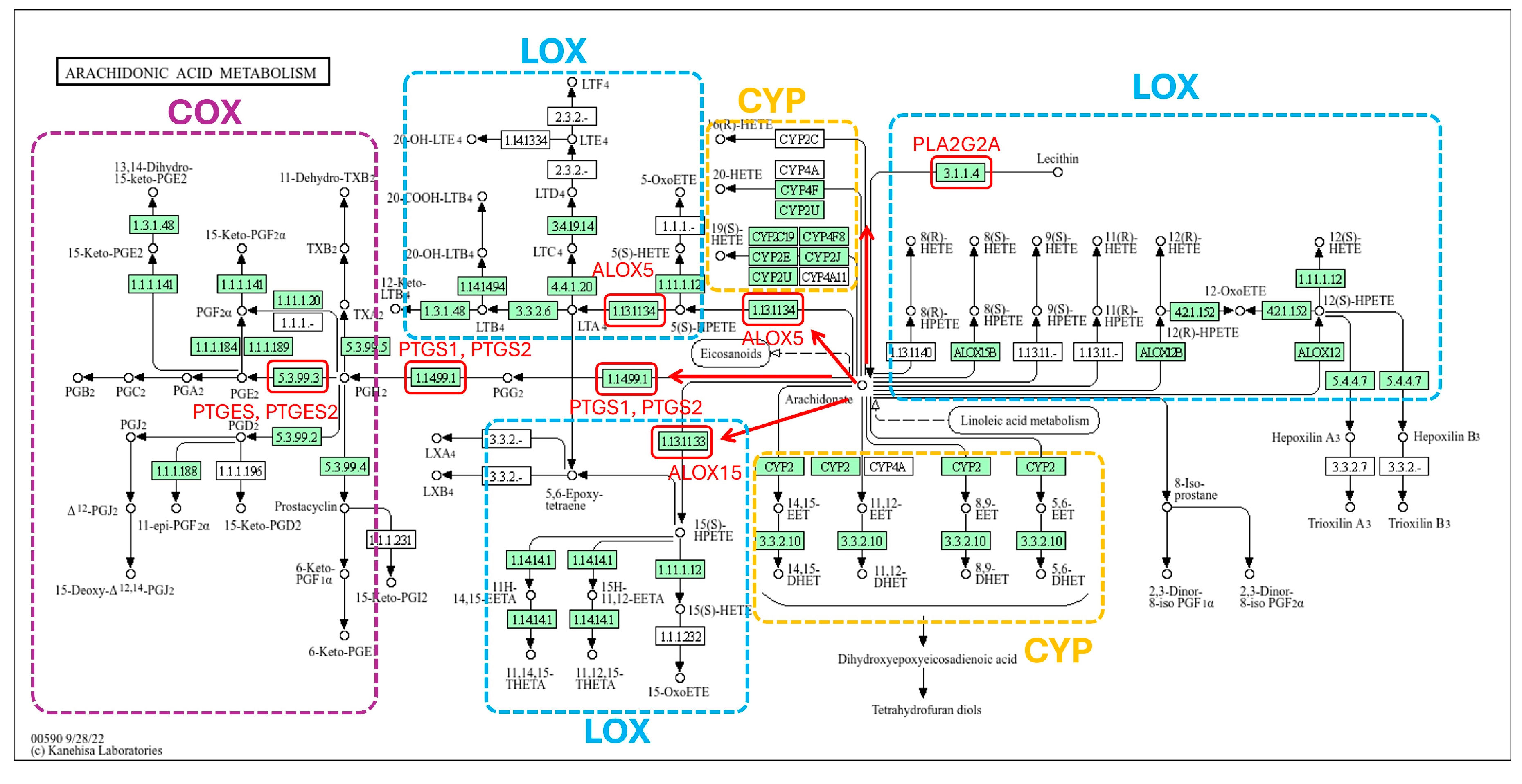
| MCODE1 | hsa00590 | Arachidonic acid metabolism | 22.6 |
| MCODE1 | hsa05143 | African trypanosomiasis | 16.0 |
| MCODE1 | hsa04913 | Ovarian steroidogenesis | 11.6 |
| MCODE2 | hsa00230 | Purine metabolism | 36.9 |
| MCODE2 | hsa05032 | Morphine addiction | 30.3 |
| MCODE2 | hsa01232 | Nucleotide metabolism | 18.5 |
| MCODE3 | hsa00360 | Phenylalanine metabolism | 120.3 |
| MCODE3 | hsa00350 | Tyrosine metabolism | 106.9 |
| MCODE3 | hsa00340 | Histidine metabolism | 87.5 |
| MCODE4 | hsa04020 | Calcium signaling pathway | 22.8 |
| MCODE4 | hsa04080 | Neuroactive ligand-receptor interaction | 19.7 |
| Compounds | Binding energy (ΔGbind, kcal/mol) | |||
|---|---|---|---|---|
| TNF (PDB: 2AZ5) |
PTGS2 (PDB: 3LN1) |
STAT3 (PDB: 6NJS) |
NFKB1 (PDB: 1NFK) |
|
| Bioactive compounds | ||||
| (+)-Vouacapenic acid | -8.9 | -7.8 | -7.5 | -8.1 |
| 5, 7-Dihydroxy-6-oxoheptade canoic acid | -5.1 | -6.5 | -4.5 | -5.1 |
| Bergapten | -6.5 | -8.1 | -6.0 | -7.6 |
| Bergenin | -7.3 | -7.2 | -7.1 | -8.6 |
| Harperamone | -6.6 | -8.0 | -5.9 | -8.0 |
| Hispidulin | -7.4 | -9.4 | -7.7 | -9.3 |
| Magnoflorine | -7.9 | -8.8 | -7.2 | -9.5 |
| Obacunone | -9.1 | -8.8 | -9.3 | -11.1 |
| O-Methylalloptaeroxylin | -7.4 | -8.9 | -6.9 | -8.1 |
| Pectolinarigenin | -7.5 | -8.6 | -7.1 | -9.6 |
| Perforatic acid | -7.5 | -8.9 | -7.2 | -8.5 |
| Perforatic acid methyl ester | -7.4 | -9.3 | -7.4 | -7.9 |
| Peucenin-7-methyl ether | -6.5 | -8.3 | -6.3 | -7.9 |
| Stigmasterol glucoside | -8.9 | -8.5 | -7.7 | -8.5 |
| Standard drug | ||||
| Thalidomide | -7.4 | - | - | - |
| Celecoxib | - | -12.1 | - | - |
| Ochromycinone | - | - | -8.3 | - |
| Selinexor | - | - | - | -10.2 |
| Marker compounds | Content of biomarkers (mg/g extract) | |||||
|---|---|---|---|---|---|---|
| BLW | CM | CI | FR | HP | TT | |
| Bergenin (1) | 0.22 ± 0.00a,* | ND | ND | 5.45 ± 0.01** | ND | ND |
| Perforatic acid (2) | 80.89 ± 0.00b,* | ND | ND | ND | 288.39 ± 0.00a,** | ND |
| O-Methyllaloptaeroxyrin (3) | 53.29 ± 0.02c,* | ND | ND | ND | 216.46 ± 0.03b,** | ND |
| Pectolinarigenin (4) | 2.50 ± 0.00d,* | ND | 3.76 ± 0.00* | ND | ND | ND |
| Peucenin-7-methyl ether (5) | 35.03 ± 0.00e | ND | ND | ND | 73.51 ± 0.00c,** | ND |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




