Submitted:
13 February 2026
Posted:
14 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. The Emergence Hypothesis
3. Microbial Characteristics
3.1. Phenotypic
3.2. Genotypic
3.2. Diagnostics and Detection
4. Pathogenicity
4.1. Morphogenetic Switching
4.2. Colony Phenotypic Switching
4.3. Contextualizing The Damage Response Framework
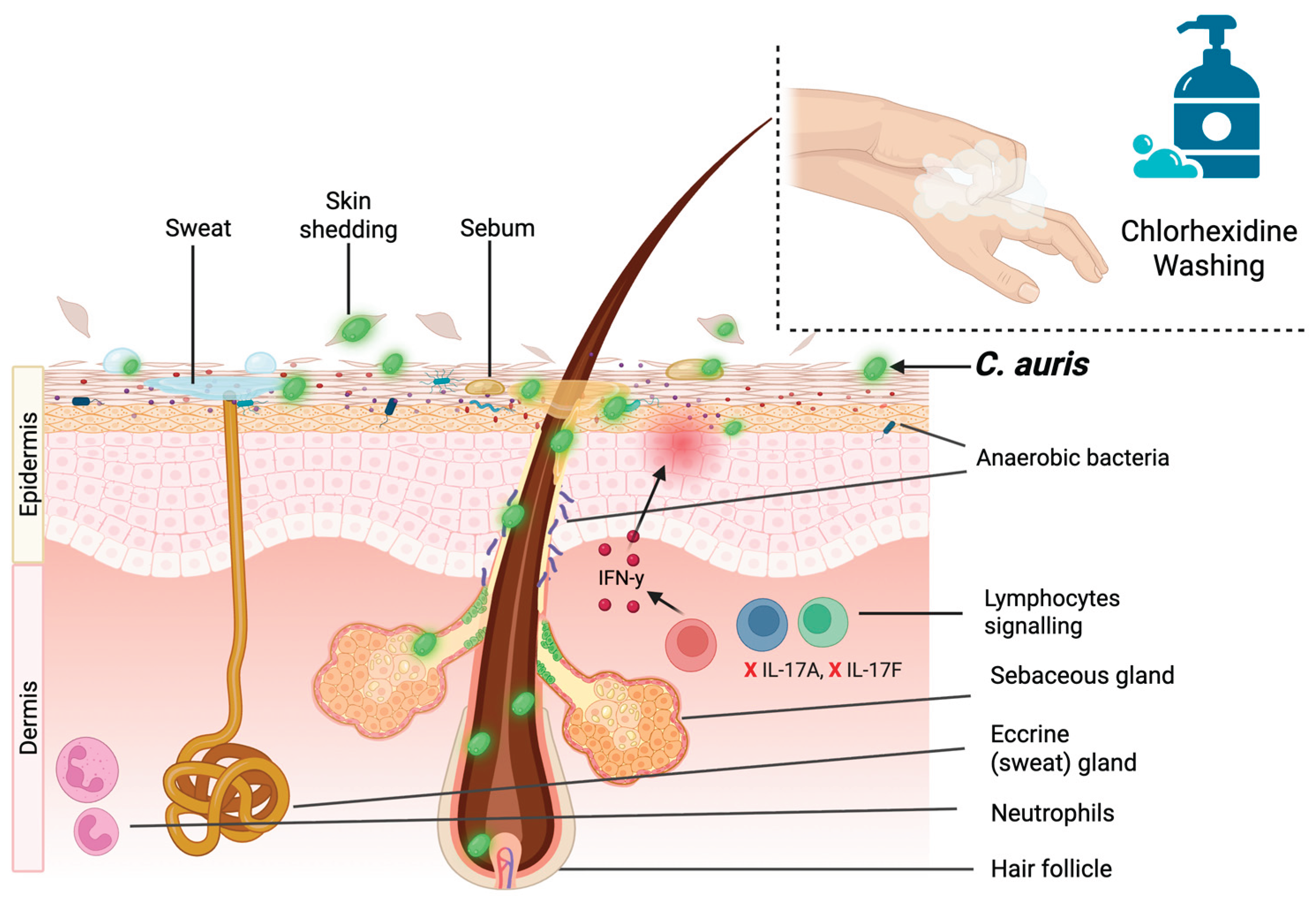
4.4. Skin Colonization and Persistence
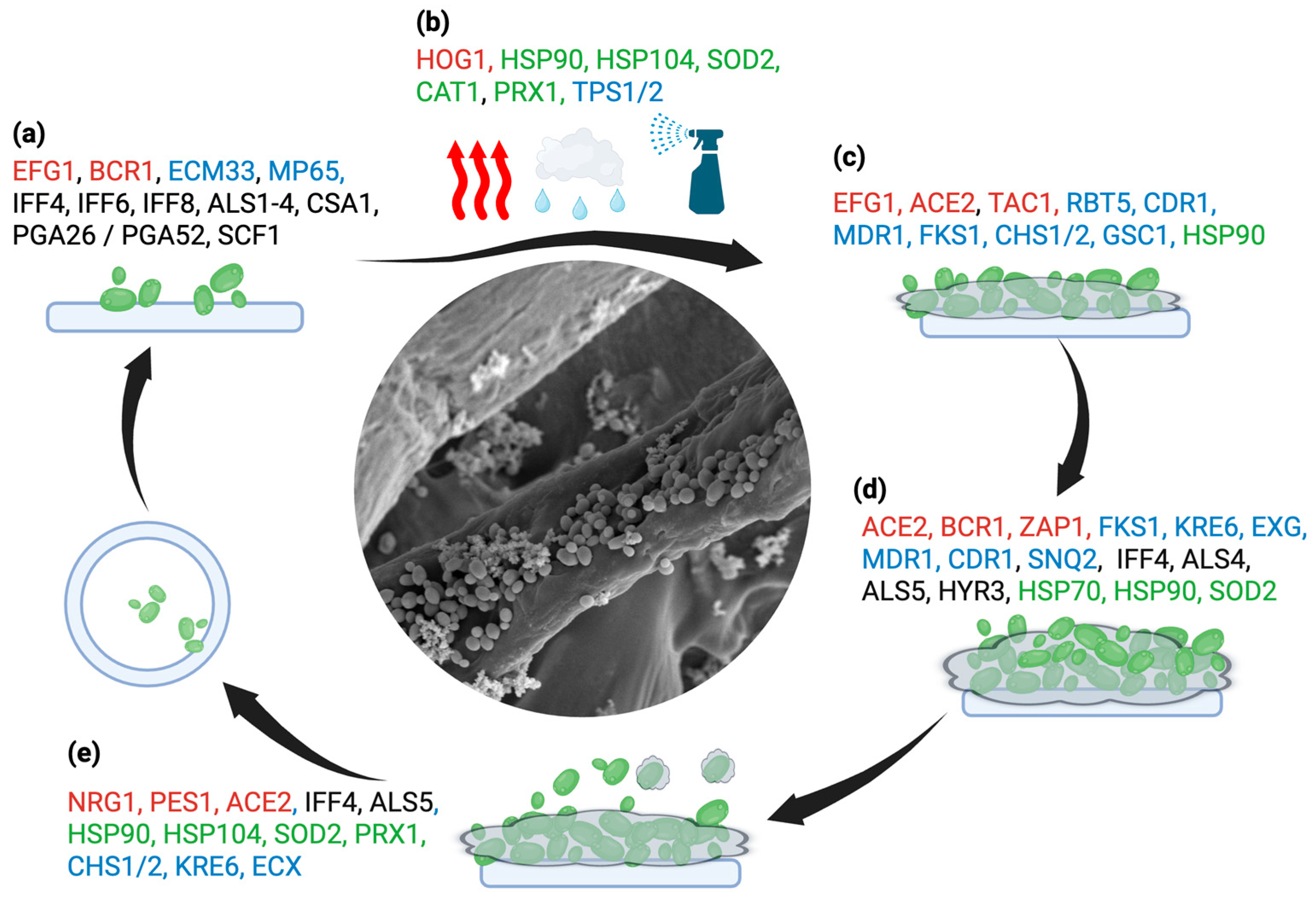
4.5. Biofilm Lifestyle: Impact on Pathogenesis and Clinical Repercussions
5. Antifungal Drug Resistance
5.1. Azoles
5.2. Polyenes
5.3. Echinocandins
6. Future Therapeutic Directions
| Fluconazole | ≥32 mg/mL |
| Amphotericin B | ≥2 mg/mL |
| Caspofungin | ≥2 mg/mL |
| Micafungin | ≥4 mg/mL |
| Anidulafungin | ≥4 mg/mL |
7. Conclusions and Outlook
Funding
Conflicts of Interest
References
- Rodrigues, M.L.; Nosanchuk, J.D. Fungal Diseases as Neglected Pathogens: A Wake-up Call to Public Health Officials. PLOS Neglected Tropical Diseases 2020, 14, e0007964. [CrossRef]
- Clinical Alert to U.S. Healthcare Facilities - June 2016 | Candida Auris | Fungal Diseases | CDC Available online: https://archive.cdc.gov/www_cdc_gov/fungal/candida-auris/candida-auris-alert.html (accessed on 25 November 2025).
- Denning, D.W. Global Incidence and Mortality of Severe Fungal Disease. The Lancet Infectious Diseases 2024, 24, e428–e438. [CrossRef]
- Liu, F.; Hu, Z.-D.; Zhao, X.-M.; Zhao, W.-N.; Feng, Z.-X.; Yurkov, A.; Alwasel, S.; Boekhout, T.; Bensch, K.; Hui, F.-L.; et al. Phylogenomic Analysis of the Candida Auris- Candida Haemuli Clade and Related Taxa in the Metschnikowiaceae, and Proposal of Thirteen New Genera, Fifty-Five New Combinations and Nine New Species. Persoonia - Molecular Phylogeny and Evolution of Fungi 2024, 52, 22–43. [CrossRef]
- Du, H.; Bing, J.; Hu, T.; Ennis, C.L.; Nobile, C.J.; Huang, G. Candida Auris: Epidemiology, Biology, Antifungal Resistance, and Virulence. PLOS Pathogens 2020, 16, e1008921. [CrossRef]
- Chowdhary, A.; Jain, K.; Chauhan, N. Candida Auris Genetics and Emergence. Annual Review of Microbiology 2023, 77, 583–602. [CrossRef]
- Vazquez-Munoz, R.; Lopez, F.D.; Lopez-Ribot, J. Silver Nanoantibiotics Display Strong Antifungal Activity against the Emergent Multidrug-Resistant yeastCandida Aurisunder Both Planktonic and Biofilm Growing Conditions. 2020. [CrossRef]
- WHO Fungal Priority Pathogens List to Guide Research, Development and Public Health Action Available online: https://www.who.int/publications/i/item/9789240060241 (accessed on 25 November 2025).
- Casadevall, A.; Kontoyiannis, D.P.; Robert, V. On the Emergence of Candida Auris: Climate Change, Azoles, Swamps, and Birds. mBio 2019, 10. [CrossRef]
- Egger, N.B.; Hoenigl, M.; Sprute, R.; Arastehfar, A.; Cornely, O.A.; Krause, R.; Lass-Flörl, C.; Prattes, J. The Rise of Candida Auris: From Unique Traits to Co-Infection Potential. Microbial Cell 2022, 9, 141–144.
- Satoh, K.; Makimura, K.; Hasumi, Y.; Nishiyama, Y.; Uchida, K.; Yamaguchi, H. Candida Aurissp. Nov., a Novel Ascomycetous Yeast Isolated from the External Ear Canal of an Inpatient in a Japanese Hospital. Microbiology and Immunology 2009, 53, 41–44. [CrossRef]
- Jones, C.R.; Neill, C.; Borman, A.M.; Budd, E.L.; Cummins, M.; Fry, C.; Guy, R.L.; Jeffery, K.; Johnson, E.M.; Manuel, R.; et al. The Laboratory Investigation, Management, and Infection Prevention and Control of Candida Auris: A Narrative Review to Inform the 2024 National Guidance Update in England: This Article Is Part of the Candida Collection. Journal of Medical Microbiology 2024, 73. [CrossRef]
- Salmanton-García, J.; Nóbrega De Almeida, J.; Colombo, A.L. Candidozyma Auris (Formerly Candida Auris): Resistant, Long Lasting, and Everywhere. Clinical Microbiology and Infection 2026, S1198743X25006330. [CrossRef]
- Casadevall, A.; Kontoyiannis, D.P.; Robert, V. Environmental Candida Auris and the Global Warming Emergence Hypothesis. mBio 2021, 12. [CrossRef]
- Santana, D.J.; Anku, J.A.E.; Zhao, G.; Zarnowski, R.; Johnson, C.J.; Hautau, H.; Visser, N.D.; Ibrahim, A.S.; Andes, D.; Nett, J.E.; et al. A Candida Auris–Specific Adhesin, Scf1, Governs Surface Association, Colonization, and Virulence. Science 2023, 381, 1461–1467. [CrossRef]
- Horton, M.V.; Johnson, C.J.; Kernien, J.F.; Patel, T.D.; Lam, B.C.; Cheong, J.Z.A.; Meudt, J.J.; Shanmuganayagam, D.; Kalan, L.R.; Nett, J.E. Candida Auris Forms High-Burden Biofilms in Skin Niche Conditions and on Porcine Skin. mSphere 2020, 5, e00910-19. [CrossRef]
- Day, A.M.; McNiff, M.M.; Da Silva Dantas, A.; Gow, N.A.R.; Quinn, J. Hog1 Regulates Stress Tolerance and Virulence in the Emerging Fungal Pathogen Candida Auris. mSphere 2018, 3, e00506-18. [CrossRef]
- Arora, P.; Singh, P.; Wang, Y.; Yadav, A.; Pawar, K.; Singh, A.; Padmavati, G.; Xu, J.; Chowdhary, A. Environmental Isolation of Candida Auris from the Coastal Wetlands of Andaman Islands, India. mBio 2021, 12. [CrossRef]
- Escandón, P. Novel Environmental Niches for Candida Auris: Isolation from a Coastal Habitat in Colombia. Journal of Fungi 2022, 8, 748. [CrossRef]
- Silva, I.; Miranda, I.M.; Costa-de-Oliveira, S. Potential Environmental Reservoirs of Candida Auris: A Systematic Review. Journal of Fungi 2024, 10, 336. [CrossRef]
- Akinbobola, A.; Kean, R.; Quilliam, R.S. Plastic Pollution as a Novel Reservoir for the Environmental Survival of the Drug Resistant Fungal Pathogen Candida Auris. Marine Pollution Bulletin 2024, 198, 115841. [CrossRef]
- Nwaubani, D.A.; Baral, R.; Solomon, T.; Idris, O.; Sherchan, S.P. Wastewater Surveillance of Candida Auris in Baltimore. International Journal of Hygiene and Environmental Health 2025, 263, 114486. [CrossRef]
- Kim, S.H.; Iyer, K.R.; Pardeshi, L.; Muñoz, J.F.; Robbins, N.; Cuomo, C.A.; Wong, K.H.; Cowen, L.E. Erratum for Kim et al., “Genetic Analysis of Candida Auris Implicates Hsp90 in Morphogenesis and Azole Tolerance and Cdr1 in Azole Resistance.” mBio 2019, 10. [CrossRef]
- Sudbery, P.; Gow, N.; Berman, J. The Distinct Morphogenic States of Candida Albicans. Trends in Microbiology 2004, 12, 317–324. [CrossRef]
- Bravo Ruiz, G.; Ross, Z.K.; Gow, N.A.R.; Lorenz, A. Pseudohyphal Growth of the Emerging Pathogen Candida Auris Is Triggered by Genotoxic Stress through the S Phase Checkpoint. mSphere 2020, 5. [CrossRef]
- Yue, H.; Bing, J.; Zheng, Q.; Zhang, Y.; Hu, T.; Du, H.; Wang, H.; Huang, G. Filamentation in Candida Auris , an Emerging Fungal Pathogen of Humans: Passage through the Mammalian Body Induces a Heritable Phenotypic Switch. Emerging Microbes & Infections 2018, 7, 1–13. [CrossRef]
- Larkin, E.; Hager, C.; Chandra, J.; Mukherjee, P.K.; Retuerto, M.; Salem, I.; Long, L.; Isham, N.; Kovanda, L.; Borroto-Esoda, K.; et al. The Emerging Pathogen Candida Auris: Growth Phenotype, Virulence Factors, Activity of Antifungals, and Effect of SCY-078, a Novel Glucan Synthesis Inhibitor, on Growth Morphology and Biofilm Formation. Antimicrob Agents Chemother 2017, 61, e02396-16. [CrossRef]
- Rossato, L.; Colombo, A.L. Candida Auris: What Have We Learned About Its Mechanisms of Pathogenicity? Frontiers in Microbiology 2018, 9. [CrossRef]
- Ramage, G.; Sherry, L.; Kean, R. Microbe Profile: Candidozyma Auris: An Emergent and Resilient Yeast and New Antifungal Strategies: This Article Is Part of the Microbe Profiles Collection. Microbiology 2026, 172. [CrossRef]
- Chybowska, A.D.; Childers, D.S.; Farrer, R.A. Nine Things Genomics Can Tell Us About Candida Auris. Frontiers in Genetics 2020, 11. [CrossRef]
- Santos, M.A.S.; Gomes, A.C.; Santos, M.C.; Carreto, L.C.; Moura, G.R. The Genetic Code of the Fungal CTG Clade. Comptes Rendus. Biologies 2011, 334, 607–611. [CrossRef]
- Lockhart, S.R.; Etienne, K.A.; Vallabhaneni, S.; Farooqi, J.; Chowdhary, A.; Govender, N.P.; Colombo, A.L.; Calvo, B.; Cuomo, C.A.; Desjardins, C.A.; et al. Simultaneous Emergence of Multidrug-Resistant Candida Auris on 3 Continents Confirmed by Whole-Genome Sequencing and Epidemiological Analyses. Clin Infect Dis 2017, 64, 134–140. [CrossRef]
- Sharma, M.; Chakrabarti, A. On the Origin of Candida Auris: Ancestor, Environmental Stresses, and Antiseptics. mBio 2020, 11. [CrossRef]
- Ahmad, S.; Alfouzan, W. Candida Auris: Epidemiology, Diagnosis, Pathogenesis, Antifungal Susceptibility, and Infection Control Measures to Combat the Spread of Infections in Healthcare Facilities. Microorganisms 2021, 9, 807. [CrossRef]
- Chow, N.A.; de Groot, T.; Badali, H.; Abastabar, M.; Chiller, T.M.; Meis, J.F. Potential Fifth Clade ofCandida Auris,Iran, 2018. Emerging Infectious Diseases 2019, 25, 1780–1781. [CrossRef]
- Khan, T.; Faysal, N.I.; Hossain, M.M.; Mah-E-Muneer, S.; Haider, A.; Moon, S.B.; Sen, D.; Ahmed, D.; Parnell, L.A.; Jubair, M.; et al. Emergence of the Novel Sixth Candida Auris Clade VI in Bangladesh. Microbiology Spectrum 2024, 12. [CrossRef]
- Lee, W.G.; Shin, J.H.; Uh, Y.; Kang, M.G.; Kim, S.H.; Park, K.H.; Jang, H.-C. First Three Reported Cases of Nosocomial Fungemia Caused by Candida Auris. Journal of Clinical Microbiology 2011, 49, 3139–3142. [CrossRef]
- Chow, N.A.; Muñoz, J.F.; Gade, L.; Berkow, E.L.; Li, X.; Welsh, R.M.; Forsberg, K.; Lockhart, S.R.; Adam, R.; Alanio, A.; et al. Tracing the Evolutionary History and Global Expansion of Candida Auris Using Population Genomic Analyses. mBio 2020, 11. [CrossRef]
- Chowdhary, A.; Sharma, C.; Meis, J.F. Candida Auris: A Rapidly Emerging Cause of Hospital-Acquired Multidrug-Resistant Fungal Infections Globally. PLoS Pathog 2017, 13, e1006290. [CrossRef]
- Lockhart, S.R.; Lyman, M.M.; Sexton, D.J. Tools for Detecting a “Superbug”: Updates on Candida Auris Testing. Journal of Clinical Microbiology 2022, 60. [CrossRef]
- Kordalewska, M.; Perlin, D.S. Molecular Diagnostics in the Times of Surveillance for Candida Auris. Journal of Fungi 2019, 5, 77. [CrossRef]
- Arastehfar, A.; Daneshnia, F.; Kord, M.; Roudbary, M.; Zarrinfar, H.; Fang, W.; Hashemi, S.J.; Najafzadeh, M.J.; Khodavaisy, S.; Pan, W.; et al. Comparison of 21-Plex PCR and API 20C AUX, MALDI-TOF MS, and rDNA Sequencing for a Wide Range of Clinically Isolated Yeast Species: Improved Identification by Combining 21-Plex PCR and API 20C AUX as an Alternative Strategy for Developing Countries. Frontiers in Cellular and Infection Microbiology 2019, 9. [CrossRef]
- Pfaller, M.A.; Diekema, D.J.; Turnidge, J.D.; Castanheira, M.; Jones, R.N. Twenty Years of the SENTRY Antifungal Surveillance Program: Results for Candida Species From 1997–2016. Open Forum Infectious Diseases 2019, 6, S79–S94. [CrossRef]
- Graf, B.; Adam, T.; Zill, E.; Göbel, U.B. Evaluation of the VITEK 2 System for Rapid Identification of Yeasts and Yeast-Like Organisms. Journal of Clinical Microbiology 2000, 38, 1782–1785. [CrossRef]
- Ramani, R.; Gromadzki, S.; Pincus, D.H.; Salkin, I.F.; Chaturvedi, V. Efficacy of API 20C and ID 32C Systems for Identification of Common and Rare Clinical Yeast Isolates. Journal of Clinical Microbiology 1998, 36, 3396–3398. [CrossRef]
- Hinrikson, H.P.; Hurst, S.F.; Lott, T.J.; Warnock, D.W.; Morrison, C.J. Assessment of Ribosomal Large-Subunit D1-D2, Internal Transcribed Spacer 1, and Internal Transcribed Spacer 2 Regions as Targets for Molecular Identification of Medically Important Aspergillus Species. Journal of Clinical Microbiology 2005, 43, 2092–2103. [CrossRef]
- Leaw, S.N.; Chang, H.C.; Sun, H.F.; Barton, R.; Bouchara, J.-P.; Chang, T.C. Identification of Medically Important Yeast Species by Sequence Analysis of the Internal Transcribed Spacer Regions. Journal of Clinical Microbiology 2006, 44, 693–699. [CrossRef]
- Pezzotti, G.; Kobara, M.; Nakaya, T.; Imamura, H.; Fujii, T.; Miyamoto, N.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Ohgitani, E.; et al. Raman Metabolomics of Candida Auris Clades: Profiling and Barcode Identification. IJMS 2022, 23, 11736. [CrossRef]
- Xue, J.; Yue, H.; Lu, W.; Li, Y.; Huang, G.; Fu, Y.V. Application of Raman Spectroscopy and Machine Learning for Candida Auris Identification and Characterization. Appl Environ Microbiol 2024, 90, e01025-24. [CrossRef]
- Bradley, S.F. Candida Auris Infection. JAMA 2019, 322, 1526. [CrossRef]
- Baker, A.D.; Gold, J.A.W.; Forsberg, K.; Jones, S.; Lyman, M.M. Progression from Candida Auris Colonization Screening to Clinical Case Status, United States, 2016–2023. Emerging Infectious Diseases 2025, 31. [CrossRef]
- Ramage, G.; Kean, R.; MacKay, W.G.; Vaccaro, A.; Lopez-Ribot, J.L.; Howe, M.H. Candidozyma Auris: Implications for Infection Prevention and Control in the Management of This Resilient Biofilm Former. In Fungal Biofilms; Springer Nature Switzerland, 2025; pp. 165–192 ISBN 978-3-031-93315-8.
- Ware, A.; Johnston, W.; Delaney, C.; Butcher, M.C.; Ramage, G.; Price, L.; Butcher, J.; Kean, R. Dry Surface Biofilm Formation by Candida Auris Facilitates Persistence and Tolerance to Sodium Hypochlorite. APMIS 2025, 133, e70022. [CrossRef]
- Spampinato, C.; Leonardi, D. CandidaInfections, Causes, Targets, and Resistance Mechanisms: Traditional and Alternative Antifungal Agents. BioMed Research International 2013, 2013, 1–13. [CrossRef]
- CDC Data and Statistics on Candidemia Available online: https://www.cdc.gov/candidiasis/data-research/facts-stats/index.html (accessed on 25 November 2025).
- Wall, G.; Montelongo-Jauregui, D.; Vidal Bonifacio, B.; Lopez-Ribot, J.L.; Uppuluri, P. Candida Albicans Biofilm Growth and Dispersal: Contributions to Pathogenesis. Current Opinion in Microbiology 2019, 52, 1–6. [CrossRef]
- CDC Data and Statistics on Candidemia Available online: https://www.cdc.gov/candidiasis/data-research/facts-stats/index.html (accessed on 25 November 2025).
- Bing, J.; Guan, Z.; Zheng, T.; Zhang, Z.; Fan, S.; Ennis, C.L.; Nobile, C.J.; Huang, G. Clinical Isolates of Candida Auris with Enhanced Adherence and Biofilm Formation Due to Genomic Amplification of ALS4. PLoS Pathog 2023, 19, e1011239. [CrossRef]
- Borman, A.M.; Szekely, A.; Johnson, E.M. Comparative Pathogenicity of United Kingdom Isolates of the Emerging Pathogen Candida Auris and Other Key Pathogenic Candida Species. mSphere 2016, 1, e00189-16. [CrossRef]
- Brown, J.L.; Delaney, C.; Short, B.; Butcher, M.C.; McKloud, E.; Williams, C.; Kean, R.; Ramage, G. Candida Auris Phenotypic Heterogeneity Determines Pathogenicity In Vitro. mSphere 2020, 5. [CrossRef]
- Ben-Ami, R.; Berman, J.; Novikov, A.; Bash, E.; Shachor-Meyouhas, Y.; Zakin, S.; Maor, Y.; Tarabia, J.; Schechner, V.; Adler, A.; et al. Multidrug-Resistant Candida Haemulonii and C. Auris , Tel Aviv, Israel. Emerging Infectious Diseases 2017, 23. [CrossRef]
- Nielsen, K.; Heitman, J. Sex and Virulence of Human Pathogenic Fungi. In Fungal Genomics; Elsevier, 2007; pp. 143–173.
- Kvaal, C.; Lachke, S.A.; Srikantha, T.; Daniels, K.; McCoy, J.; Soll, D.R. Misexpression of the Opaque-Phase-Specific Gene PEP1 ( SAP1 ) in the White Phase of Candida Albicans Confers Increased Virulence in a Mouse Model of Cutaneous Infection. Infection and Immunity 1999, 67, 6652–6662. [CrossRef]
- Kvaal, C.A.; Srikantha, T.; Soll, D.R. Misexpression of the White-Phase-Specific Gene WH11 in the Opaque Phase of Candida Albicans Affects Switching and Virulence. Infection and Immunity 1997, 65, 4468–4475. [CrossRef]
- Bentz, M.L.; Sexton, D.J.; Welsh, R.M.; Litvintseva, A.P. Phenotypic Switching in Newly Emerged Multidrug-Resistant Pathogen Candida Auris. Medical Mycology 2018, 57, 636–638. [CrossRef]
- Mulet Bayona, J.V.; Salvador García, C.; Tormo Palop, N.; Gimeno Cardona, C. Evaluation of a Novel Chromogenic Medium for Candida Spp. Identification and Comparison with CHROMagarTM Candida for the Detection of Candida Auris in Surveillance Samples. Diagnostic Microbiology and Infectious Disease 2020, 98, 115168. [CrossRef]
- Fan, S.; Li, C.; Bing, J.; Huang, G.; Du, H. Discovery of the Diploid Form of the Emerging Fungal Pathogen Candida Auris. ACS Infectious Diseases 2020, 6, 2641–2646. [CrossRef]
- Muñoz, J.F.; Gade, L.; Chow, N.A.; Loparev, V.N.; Juieng, P.; Berkow, E.L.; Farrer, R.A.; Litvintseva, A.P.; Cuomo, C.A. Genomic Insights into Multidrug-Resistance, Mating and Virulence in Candida Auris and Related Emerging Species. Nature Communications 2018, 9. [CrossRef]
- Casadevall, A.; Pirofski, L. The Damage-Response Framework of Microbial Pathogenesis. Nature Reviews Microbiology 2003, 1, 17–24. [CrossRef]
- Tharp, B.; Zheng, R.; Bryak, G.; Litvintseva, A.P.; Hayden, M.K.; Chowdhary, A.; Thangamani, S. Role of Microbiota in the Skin Colonization of Candida Auris. mSphere 2023, 8, e00623-22. [CrossRef]
- Swaney, M.H.; Nelsen, A.; Sandstrom, S.; Kalan, L.R. Sweat and Sebum Preferences of the Human Skin Microbiota. Microbiol Spectr 2026, 11, e04180-22. [CrossRef]
- Sexton, D.J.; Bentz, M.L.; Welsh, R.M.; Derado, G.; Furin, W.; Rose, L.J.; Noble-Wang, J.; Pacilli, M.; McPherson, T.D.; Black, S.; et al. Positive Correlation Between Candida Auris Skin-Colonization Burden and Environmental Contamination at a Ventilator-Capable Skilled Nursing Facility in Chicago. Clin Infect Dis 2021, 73, 1142–1148. [CrossRef]
- Kim, H.Y.; Nguyen, T.A.; Kidd, S.; Chambers, J.; Alastruey-Izquierdo, A.; Shin, J.-H.; Dao, A.; Forastiero, A.; Wahyuningsih, R.; Chakrabarti, A.; et al. Candida Auris —a Systematic Review to Inform the World Health Organization Fungal Priority Pathogens List. Medical Mycology 2024, 62, myae042. [CrossRef]
- Bruno, M.; Kersten, S.; Bain, J.M.; Jaeger, M.; Rosati, D.; Kruppa, M.D.; Lowman, D.W.; Rice, P.J.; Graves, B.; Ma, Z.; et al. Transcriptional and Functional Insights into the Host Immune Response against the Emerging Fungal Pathogen Candida Auris. Nature Microbiology 2020, 5, 1516–1531. [CrossRef]
- Johnson, C.J.; Davis, J.M.; Huttenlocher, A.; Kernien, J.F.; Nett, J.E. H. mBio 2018, 9, e01403-18. [CrossRef]
- Biswal, M.; Rudramurthy, S.M.; Jain, N.; Shamanth, A.S.; Sharma, D.; Jain, K.; Yaddanapudi, L.N.; Chakrabarti, A. Controlling a Possible Outbreak of Candida Auris Infection: Lessons Learnt from Multiple Interventions. Journal of Hospital Infection 2017, 97, 363–370. [CrossRef]
- Proctor, D.M.; Drummond, R.A.; Lionakis, M.S.; Segre, J.A. One Population, Multiple Lifestyles: Commensalism and Pathogenesis in the Human Mycobiome. Cell Host & Microbe 2023, 31, 539–553. [CrossRef]
- Bergeron, G.; Bloch, D.; Murray, K.; Kratz, M.; Parton, H.; Ackelsberg, J.; Antwi, M.; Del Rosso, P.; Dorsinville, M.; Kubinson, H.; et al. Candida Auris Colonization After Discharge to a Community Setting: New York City, 2017–2019. Open Forum Infect Dis 2020, 8, ofaa620. [CrossRef]
- Huang, X.; Hurabielle, C.; Drummond, R.A.; Bouladoux, N.; Desai, J.V.; Sim, C.K.; Belkaid, Y.; Lionakis, M.S.; Segre, J.A. Murine Model of Colonization with Fungal Pathogen Candida Auris to Explore Skin Tropism, Host Risk Factors and Therapeutic Strategies. Cell Host & Microbe 2021, 29, 210-221.e6. [CrossRef]
- Eyre, D.W.; Sheppard, A.E.; Madder, H.; Moir, I.; Moroney, R.; Quan, T.P.; Griffiths, D.; George, S.; Butcher, L.; Morgan, M.; et al. A Candida Auris Outbreak and Its Control in an Intensive Care Setting. New England Journal of Medicine 2018, 379, 1322–1331. [CrossRef]
- Moore, G.; Schelenz, S.; Borman, A.M.; Johnson, E.M.; Brown, C.S. Yeasticidal Activity of Chemical Disinfectants and Antiseptics against Candida Auris. Journal of Hospital Infection 2017, 97, 371–375. [CrossRef]
- Johnson, C.J.; Eix, E.F.; Lam, B.C.; Wartman, K.M.; Meudt, J.J.; Shanmuganayagam, D.; Nett, J.E. Augmenting the Activity of Chlorhexidine for Decolonization of Candida Auris from Porcine Skin. J Fungi (Basel) 2021, 7, 804. [CrossRef]
- Santana, D.J.; Anku, J.A.E.; Zhao, G.; Zarnowski, R.; Johnson, C.J.; Hautau, H.; Visser, N.D.; Ibrahim, A.S.; Andes, D.; Nett, J.E.; et al. A Candida Auris-Specific Adhesin, SCF1, Governs Surface Association, Colonization, and Virulence. Science 2023, 381, 1461–1467. [CrossRef]
- Wang, T.W.; Sofras, D.; Montelongo-Jauregui, D.; Paiva, T.O.; Carolus, H.; Dufrêne, Y.F.; Alfaifi, A.A.; McCracken, C.; Bruno, V.M.; Van Dijck, P.; et al. Functional Redundancy in Candida Auris Cell Surface Adhesins Crucial for Cell-Cell Interaction and Aggregation. Nat Commun 2024, 15, 9212. [CrossRef]
- Zhao, G.; Lyu, J.; Veniaminova, N.A.; Zarnowski, R.; Mattos, E.; Johnson, C.J.; Quintanilla, D.; Hautau, H.; Hold, L.A.; Xu, B.; et al. Adhesin Als4112 Promotes Candida Auris Skin Colonization through Interactions with Keratinocytes and Extracellular Matrix Proteins. Nat Commun 2025, 16, 5673. [CrossRef]
- Wang, T.W.; Sofras, D.; Montelongo-Jauregui, D.; Paiva, T.O.; Carolus, H.; Dufrêne, Y.F.; Alfaifi, A.A.; McCracken, C.; Bruno, V.M.; Van Dijck, P.; et al. Functional Redundancy in Candida Auris Cell Surface Adhesins Crucial for Cell-Cell Interaction and Aggregation. Nat Commun 2024, 15, 9212. [CrossRef]
- Shivarathri, R.; Chauhan, M.; Datta, A.; Das, D.; Karuli, A.; Jenull, S.; Kuchler, K.; Thangamani, S.; Chowdhary, A.; Desai, J.V.; et al. The Candida Auris Hog1 MAP Kinase Is Essential for the Colonization of Murine Skin and Intradermal Persistence. bioRxiv 2024, 2024.03.18.585572. [CrossRef]
- Huang, X.; Welsh, R.M.; Deming, C.; Proctor, D.M.; Thomas, P.J.; Gussin, G.M.; Huang, S.S.; Kong, H.H.; Bentz, M.L.; Vallabhaneni, S.; et al. Skin Metagenomic Sequence Analysis of Early Candida Auris Outbreaks in U.S. Nursing Homes. mSphere 2026, 6, e00287-21. [CrossRef]
- Kean, R.; McKloud, E.; Townsend, E.M.; Sherry, L.; Delaney, C.; Jones, B.L.; Williams, C.; Ramage, G. The Comparative Efficacy of Antiseptics against Candida Auris Biofilms. International Journal of Antimicrobial Agents 2018, 52, 673–677. [CrossRef]
- Cerbu, D.; Seiser, S.; Phan-Canh, T.; Moser, D.; Freystätter, C.; Matiasek, J.; Kuchler, K.; Elbe-Bürger, A. Octenidine Effectively Reduces Candida Auris Colonisation on Human Skin. Sci Rep 2025, 15, 27034. [CrossRef]
- Wu, W.-G.; Luk, K.S.; Hung, M.-F.; Tsang, W.-Y.; Lee, K.-P.; Lam, B.H.-S.; Cheng, K.-L.; Cheung, W.-S.; Tang, H.-L.; To, W.-K. Antifungal Efficacy of Natural Antiseptic Products against Candida Auris. Med Mycol 2024, 62. [CrossRef]
- Baldim, I.; Paziani, M.H.; Grizante Barião, P.H.; Kress, M.R. von Z.; Oliveira, W.P. Nanostructured Lipid Carriers Loaded with Lippia Sidoides Essential Oil as a Strategy to Combat the Multidrug-Resistant Candida Auris. Pharmaceutics 2022, 14, 180. [CrossRef]
- Jaromin, A.; Zarnowski, R.; Markowski, A.; Zagórska, A.; Johnson, C.J.; Etezadi, H.; Kihara, S.; Mota-Santiago, P.; Nett, J.E.; Boyd, B.J.; et al. Liposomal Formulation of a New Antifungal Hybrid Compound Provides Protection against Candida Auris in the Ex Vivo Skin Colonization Model. Antimicrob Agents Chemother 2026, 68, e00955-23. [CrossRef]
- Eix, E.F.; Johnson, C.J.; Wartman, K.M.; Kernien, J.F.; Meudt, J.J.; Shanmuganayagam, D.; Gibson, A.L.F.; Nett, J.E. Ex Vivo Human and Porcine Skin Effectively Model Candida Auris Colonization, Differentiating Robust and Poor Fungal Colonizers. J Infect Dis 2022, 225, 1791–1795. [CrossRef]
- Brandt, P.; Mirhakkak, M.H.; Wagner, L.; Driesch, D.; Möslinger, A.; Fänder, P.; Schäuble, S.; Panagiotou, G.; Vylkova, S. High-Throughput Profiling of Candida Auris Isolates Reveals Clade-Specific Metabolic Differences. Microbiol Spectr 2026, 11, e00498-23. [CrossRef]
- Nicklas, J.P.; Deming, C.; Lee-Lin, S.; Conlan, S.; Shen, Z.; Blaustein, R.A.; Segre, J.A. Candida Auris Metabolism and Growth Preferences in Physiologically Relevant Skin-like Conditions. bioRxiv 2025, 2025.05.18.654780. [CrossRef]
- Fanning, S.; Mitchell, A.P. Fungal Biofilms. PLoS Pathogens 2012, 8, e1002585. [CrossRef]
- Ramage, G.; Kean, R.; Rautemaa-Richardson, R.; Williams, C.; Lopez-Ribot, J.L. Fungal Biofilms in Human Health and Disease. Nat Rev Microbiol 2025. [CrossRef]
- Sherry, L.; Ramage, G.; Kean, R.; Borman, A.; Johnson, E.M.; Richardson, M.D.; Rautemaa-Richardson, R. Biofilm-Forming Capability of Highly Virulent, Multidrug-Resistant Candida Auris. Emerg Infect Dis 2017, 23, 328–331. [CrossRef]
- Wang, X.; Bing, J.; Zheng, Q.; Zhang, F.; Liu, J.; Yue, H.; Tao, L.; Du, H.; Wang, Y.; Wang, H.; et al. The First Isolate of Candida Auris in China: Clinical and Biological Aspects. Emerging Microbes & Infections 2018, 7, 1–9. [CrossRef]
- Vila, T.; Montelongo-Jauregui, D.; Ahmed, H.; Puthran, T.; Sultan, A.S.; Jabra-Rizk, M.A. Comparative Evaluations of the Pathogenesis of Candida Auris Phenotypes and Candida Albicans Using Clinically Relevant Murine Models of Infections. mSphere 2020, 5, 10.1128/msphere.00760-20. [CrossRef]
- Pathirana, R.U.; Friedman, J.; Norris, H.L.; Salvatori, O.; McCall, A.D.; Kay, J.; Edgerton, M. Fluconazole-Resistant Candida Auris Is Susceptible to Salivary Histatin 5 Killing and to Intrinsic Host Defenses. Antimicrobial Agents and Chemotherapy 2018, 62. [CrossRef]
- Kean, R.; Delaney, C.; Sherry, L.; Borman, A.; Johnson, E.M.; Richardson, M.D.; Rautemaa-Richardson, R.; Williams, C.; Ramage, G. Transcriptome Assembly and Profiling of Candida Auris Reveals Novel Insights into Biofilm-Mediated Resistance. mSphere 2018, 3, e00334-18. [CrossRef]
- Frías-De-León, M.G.; Hernández-Castro, R.; Vite-Garín, T.; Arenas, R.; Bonifaz, A.; Castañón-Olivares, L.; Acosta-Altamirano, G.; Martínez-Herrera, E. Antifungal Resistance in Candida Auris: Molecular Determinants. Antibiotics 2020, 9, 568. [CrossRef]
- Lockhart, S.R. Candida Auris and Multidrug Resistance: Defining the New Normal. Fungal Genetics and Biology 2019, 131, 103243. [CrossRef]
- Chaabane, F.; Graf, A.; Jequier, L.; Coste, A.T. Review on Antifungal Resistance Mechanisms in the Emerging Pathogen Candida Auris. Frontiers in Microbiology 2019, 10. [CrossRef]
- Healey, K.R.; Kordalewska, M.; Jiménez Ortigosa, C.; Singh, A.; Berrío, I.; Chowdhary, A.; Perlin, D.S. Limited ERG11 Mutations Identified in Isolates of Candida Auris Directly Contribute to Reduced Azole Susceptibility. Antimicrobial Agents and Chemotherapy 2018, 62. [CrossRef]
- Rybak, J.M.; Doorley, L.A.; Nishimoto, A.T.; Barker, K.S.; Palmer, G.E.; Rogers, P.D. Abrogation of Triazole Resistance upon Deletion of CDR1 in a Clinical Isolate of Candida Auris. Antimicrobial Agents and Chemotherapy 2019, 63. [CrossRef]
- Barker, K.S.; Santana, D.J.; Zhang, Q.; Peters, T.L.; Rybak, J.M.; Morschhäuser, J.; Cuomo, C.A.; Rogers, P.D. Mutations in TAC1B Drive Increased CDR1 and MDR1 Expression and Azole Resistance in Candida Auris. Antimicrobial Agents and Chemotherapy 2025, 69. [CrossRef]
- Fisher, M.C.; Alastruey-Izquierdo, A.; Berman, J.; Bicanic, T.; Bignell, E.M.; Bowyer, P.; Bromley, M.; Brüggemann, R.; Garber, G.; Cornely, O.A.; et al. Tackling the Emerging Threat of Antifungal Resistance to Human Health. Nature Reviews Microbiology 2022, 20, 557–571. [CrossRef]
- Nett, J.E.; Andes, D.R. Antifungal Agents. Infectious Disease Clinics of North America 2016, 30, 51–83. [CrossRef]
- Chen, J.; Tian, S.; Han, X.; Chu, Y.; Wang, Q.; Zhou, B.; Shang, H. Is the Superbug Fungus Really so Scary? A Systematic Review and Meta-Analysis of Global Epidemiology and Mortality of Candida Auris. BMC Infect Dis 2020, 20, 827. [CrossRef]
- Siopi, M.; Peroukidou, I.; Beredaki, M.-I.; Spruijtenburg, B.; De Groot, T.; Meis, J.F.; Vrioni, G.; Tsakris, A.; Pournaras, S.; Meletiadis, J. Overestimation of Amphotericin B Resistance in Candida Auris with Sensititre YeastOne Antifungal Susceptibility Testing: A Need for Adjustment for Correct Interpretation. Microbiol Spectr 2023, 11, e04431-22. [CrossRef]
- Chen, S.C.A.; Sorrell, T.C. Antifungal Agents. Medical Journal of Australia 2007, 187, 404–409. [CrossRef]
- Gow, N.A.R.; Latge, J.-P.; Munro, C.A. The Fungal Cell Wall: Structure, Biosynthesis, and Function. Microbiology Spectrum 2017, 5. [CrossRef]
- Kordalewska, M.; Lee, A.; Park, S.; Berrio, I.; Chowdhary, A.; Zhao, Y.; Perlin, D.S. Understanding Echinocandin Resistance in the Emerging Pathogen Candida Auris. Antimicrobial Agents and Chemotherapy 2018, 62. [CrossRef]
- Ramos, L.S.; Barbosa, P.F.; Lorentino, C.M.A.; Lima, J.C.; Braga, A.L.; Lima, R.V.; Giovanini, L.; Casemiro, A.L.; Siqueira, N.L.M.; Costa, S.C.; et al. The Multidrug-Resistant Candida Auris, Candida Haemulonii Complex and Phylogenetic Related Species: Insights into Antifungal Resistance Mechanisms. Current Research in Microbial Sciences 2025, 8, 100354. [CrossRef]
- Misas, E.; Parnell, L.A.; Rajeev, M.; López, L.F.; Santos, A.R.D.; Mudge, Z.B.; Gade, L.; Forsberg, K.; Lyman, M.; Sexton, D.J.; et al. A Benchmark Dataset for Validating FKS1 Mutations in Candida Auris. Microbiol Spectr 2025, 13, e03147-24. [CrossRef]
- Cao, C.; Wang, Y.; Husain, S.; Soteropoulos, P.; Xue, C. A Mechanosensitive Channel Governs Lipid Flippase-Mediated Echinocandin Resistance in Cryptococcus Neoformans. mBio 2019, 10. [CrossRef]
- Arendrup, M.C.; Lockhart, S.R.; Wiederhold, N. Candida Auris MIC Testing by EUCAST and Clinical and Laboratory Standards Institute Broth Microdilution, and Gradient Diffusion Strips; to Be or Not to Be Amphotericin B Resistant? Clinical Microbiology and Infection 2025, 31, 108–112. [CrossRef]
- Castanheira, M.; Deshpande, L.M.; Rhomberg, P.R.; Carvalhaes, C.G. Recent Increase in Candida Auris Frequency in the SENTRY Surveillance Program: Antifungal Activity and Genotypic Characterization. Antimicrob Agents Chemother 2024, 68, e00570-24. [CrossRef]
- Candida Auris: Methods and Protocols; Lorenz, A., Ed.; Methods in Molecular Biology; 1st ed. 2022.; Springer US: New York, NY, 2022; ISBN 978-1-0716-2417-3.
- Vatanshenassan, M.; Boekhout, T.; Lass-Flörl, C.; Lackner, M.; Schubert, S.; Kostrzewa, M.; Sparbier, K. Proof of Concept for MBT ASTRA, a Rapid Matrix-Assisted Laser Desorption Ionization–Time of Flight Mass Spectrometry (MALDI-TOF MS)-Based Method To Detect Caspofungin Resistance in Candida Albicans and Candida Glabrata. Journal of Clinical Microbiology 2018, 56. [CrossRef]
- Di Luca, M.; Koliszak, A.; Karbysheva, S.; Chowdhary, A.; Meis, J.; Trampuz, A. Thermogenic Characterization and Antifungal Susceptibility of Candida Auris by Microcalorimetry. Journal of Fungi 2019, 5, 103. [CrossRef]
- CDC Antifungal Susceptibility Testing for C. auris Available online: https://www.cdc.gov/candida-auris/hcp/laboratories/antifungal-susceptibility-testing.html (accessed on 25 November 2025).
- CDC Antifungal Susceptibility Testing for C. auris Available online: https://www.cdc.gov/candida-auris/hcp/laboratories/antifungal-susceptibility-testing.html (accessed on 25 November 2025).
- Szymański, P.; Markowicz, M.; Mikiciuk-Olasik, E. Adaptation of High-Throughput Screening in Drug Discovery—Toxicological Screening Tests. International Journal of Molecular Sciences 2011, 13, 427–452. [CrossRef]
- Pierce, C.G.; Uppuluri, P.; Tristan, A.R.; Wormley, F.L.; Mowat, E.; Ramage, G.; Lopez-Ribot, J.L. A Simple and Reproducible 96-Well Plate-Based Method for the Formation of Fungal Biofilms and Its Application to Antifungal Susceptibility Testing. Nat Protoc 2008, 3, 1494–1500. [CrossRef]
- Ajetunmobi, O.H.; Wall, G.; Bonifacio, B.V.; Montelongo-Jauregui, D.; Lopez-Ribot, J.L. A 384-Well Microtiter Plate Model for Candida Biofilm Formation and Its Application to High-Throughput Screening. In Antifungal Drug Resistance; Springer US, 2023; pp. 53–64 ISBN 978-1-0716-3155-3.
- Singh, U.; Akhtar, S.; Mishra, A.; Sarkar, D. A Novel Screening Method Based on Menadione Mediated Rapid Reduction of Tetrazolium Salt for Testing of Anti-Mycobacterial Agents. Journal of Microbiological Methods 2011, 84, 202–207. [CrossRef]
- Pushpakom, S.; Iorio, F.; Eyers, P.A.; Escott, K.J.; Hopper, S.; Wells, A.; Doig, A.; Guilliams, T.; Latimer, J.; McNamee, C.; et al. Drug Repurposing: Progress, Challenges and Recommendations. Nature Reviews Drug Discovery 2018, 18, 41–58. [CrossRef]
- Shaban, S.; Patel, M.; Ahmad, A. Improved Efficacy of Antifungal Drugs in Combination with Monoterpene Phenols against Candida Auris. Scientific Reports 2020, 10. [CrossRef]
- Lionakis, M.S.; Drummond, R.A.; Hohl, T.M. Immune Responses to Human Fungal Pathogens and Therapeutic Prospects. Nature Reviews Immunology 2023, 23, 433–452. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




