Submitted:
12 February 2026
Posted:
13 February 2026
You are already at the latest version
Abstract
Myeloid/lymphoid neoplasms with tyrosine kinase rearrangements (MLN-TK) are rare clonal eosinophilias driven by PDGFRA, PDGFRB and other kinase fusions, highly sensitive to tyrosine kinase inhibitors. Their detection remains challenging, particularly for cryptic PDGFRA rearrangements. We performed a large multicenter real-world validation of the generic quantitative RT-PCR assay (gPDGFR), which detects 3′ PDGFRA/PDGFRB overexpression independently of fusion partner. A total of 231 consecutive patients with hypereosinophilia from 12 French centers were analyzed, and assay robustness was further assessed in an independent heterogeneous cohort of 102 TKI-treated patients. Twenty-two PDGFR-rearranged cases (14 PDGFRA-r, 8 PDGFRB-r) were identified. The assay demonstrated 100% sensitivity and 100% negative predictive value. For PDGFRA, positive predictive value and specificity reached 100%. In contrast, PDGFRB overexpression showed lower specificity due to borderline false-positive cases, underscoring the need for confirmatory testing. In selected patients, longitudinal gPDGFR kinetics paralleled fusion-specific RT-qPCR, supporting its use for molecular follow-up when dedicated assays are unavailable, although it does not provide quantitative measurable residual disease assessment. Overall, gPDGFR represents a robust, partner-independent first-line screening strategy that can be readily integrated into routine diagnostic workflows to enable timely identification of patients eligible for targeted therapy.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design and Population
2.2. gPDGFR Assay
2.3. Definition of True Positive and True Negative Cases
3. Results
3.1. Diagnostic Performance
3.2. Clinical and Biological Characteristics
3.3. Treatment Response and Molecular Monitoring
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Appendix A
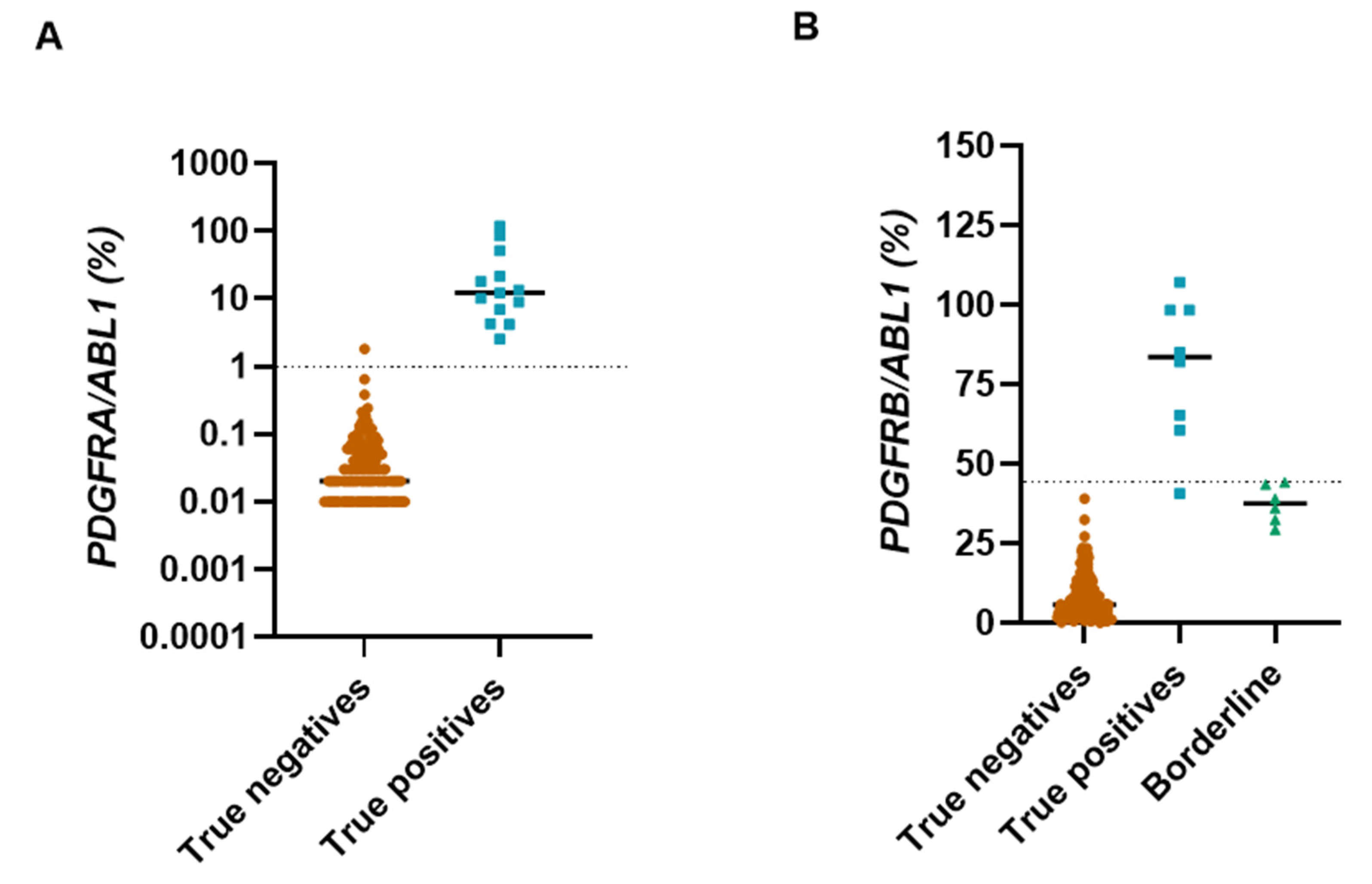
Appendix B
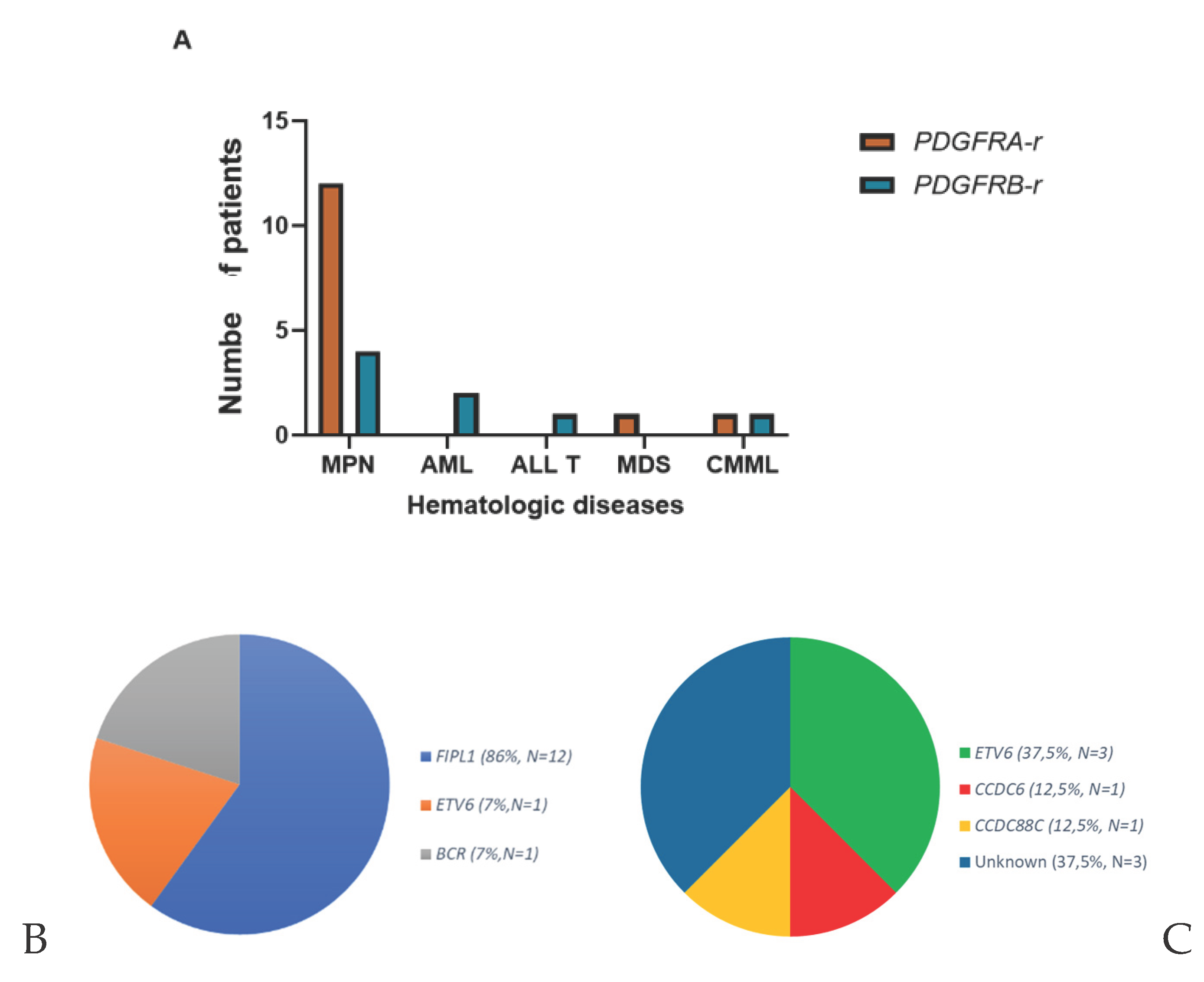
References
- Khoury, J.D.; Solary, E.; Abla, O.; Akkari, Y.; Alaggio, R.; Apperley, J.F.; Bejar, R.; Berti, E.; Busque, L.; Chan, J.K.C.; Chen, W.; Chen, X.; Chng, W.-J.; Choi, J.K.; Colmenero, I.; Coupland, S.E.; Cross, N.C.P.; De Jong, D.; Elghetany, M.T.; Takahashi, E.; Emile, J.-F.; Ferry, J.; Fogelstrand, L.; Fontenay, M.; Germing, U.; Gujral, S.; Haferlach, T.; Harrison, C.; Hodge, J.C.; Hu, S.; Jansen, J.H.; Kanagal-Shamanna, R.; Kantarjian, H.M.; Kratz, C.P.; Li, X.-Q.; Lim, M.S.; Loeb, K.; Loghavi, S.; Marcogliese, A.; Meshinchi, S.; Michaels, P.; Naresh, K.N.; Natkunam, Y.; Nejati, R.; Ott, G.; Padron, E.; Patel, K.P.; Patkar, N.; Picarsic, J.; Platzbecker, U.; Roberts, I.; Schuh, A.; Sewell, W.; Siebert, R.; Tembhare, P.; Tyner, J.; Verstovsek, S.; Wang, W.; Wood, B.; Xiao, W.; Yeung, C.; Hochhaus, A. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia 2022, 36, 1703–1719. [Google Scholar] [CrossRef]
- Kaur, M.D. Prabhjot; Khan, P.H.D. Wahab A. Myeloid/Lymphoid Neoplasms with Eosinophilia and Platelet Derived Growth Factor Receptor Alpha (PDGFRA) Rearrangement. In Exon Publications; 2022; pp. 129–146. [Google Scholar] [CrossRef]
- Erben, P.; Gosenca, D.; Muller, M.C.; Reinhard, J.; Score, J.; Del Valle, F.; Walz, C.; Mix, J.; Metzgeroth, G.; Ernst, T.; Haferlach, C.; Cross, N.C.P.; Hochhaus, A.; Reiter, A. Screening for diverse PDGFRA or PDGFRB fusion genes is facilitated by generic quantitative reverse transcriptase polymerase chain reaction analysis. Haematologica 2010, 95, 738–744. [Google Scholar] [CrossRef] [PubMed]
- Score, J.; Curtis, C.; Waghorn, K.; Stalder, M.; Jotterand, M.; Grand, F.H.; Cross, N.C.P. Identification of a novel imatinib responsive KIF5B-PDGFRA fusion gene following screening for PDGFRA overexpression in patients with hypereosinophilia. Leukemia 2006, 20, 827–832. [Google Scholar] [CrossRef] [PubMed]
- Ruminy, P.; Marchand, V.; Buchbinder, N.; Larson, T.; Joly, B.; Penther, D.; Lemasle, E.; Lepretre, S.; Angot, E.; Mareschal, S.; Viailly, P.-J.; Dubois, S.; Clatot, F.; Viennot, M.; Bohers, E.; Rizzo, D.; Cornic, M.; Bertrand, P.; Girod, C.; Camus, V.; Etancelin, P.; Buchonnet, G.; Schneider, P.; Picquenot, J.-M.; Vannier, J.-P.; Bastard, C.; Tilly, H.; Jardin, F. Multiplexed targeted sequencing of recurrent fusion genes in acute leukaemia. Leukemia 2016, 30, 757–760. [Google Scholar] [CrossRef] [PubMed]
- Rohmer, J.; Couteau-Chardon, A.; Trichereau, J.; Panel, K.; Gesquiere, C.; Ben Abdelali, R.; Bidet, A.; Bladé, J.-S.; Cayuela, J.-M.; Cony-Makhoul, P.; Cottin, V.; Delabesse, E.; Ebbo, M.; Fain, O.; Flandrin, P.; Galicier, L.; Godon, C.; Grardel, N.; Guffroy, A.; Hamidou, M.; Hunault, M.; Lengline, E.; Lhomme, F.; Lhermitte, L.; Machelart, I.; Mauvieux, L.; Mohr, C.; Mozicconacci, M.-J.; Naguib, D.; Nicolini, F.E.; Rey, J.; Rousselot, P.; Tavitian, S.; Terriou, L.; Lefèvre, G.; Preudhomme, C.; Kahn, J.-E.; Groh, M. CEREO and GBMHM collaborators, Epidemiology, clinical picture and long-term outcomes of FIP1L1-PDGFRA-positive myeloid neoplasm with eosinophilia: Data from 151 patients. Am J Hematol 2020, 95, 1314–1323. [Google Scholar] [CrossRef] [PubMed]
- Ogbogu, P.U.; Bochner, B.S.; Butterfield, J.H.; Gleich, G.J.; Huss-Marp, J.; Kahn, J.E.; Leiferman, K.M.; Nutman, T.B.; Pfab, F.; Ring, J.; Rothenberg, M.E.; Roufosse, F.; Sajous, M.-H.; Sheikh, J.; Simon, D.; Simon, H.-U.; Stein, M.L.; Wardlaw, A.; Weller, P.F.; Klion, A.D. Hypereosinophilic syndromes: a multicenter, retrospective analysis of clinical characteristics and response to therapy. J Allergy Clin Immunol 2009, 124, 1319–25.e3. [Google Scholar] [CrossRef] [PubMed]
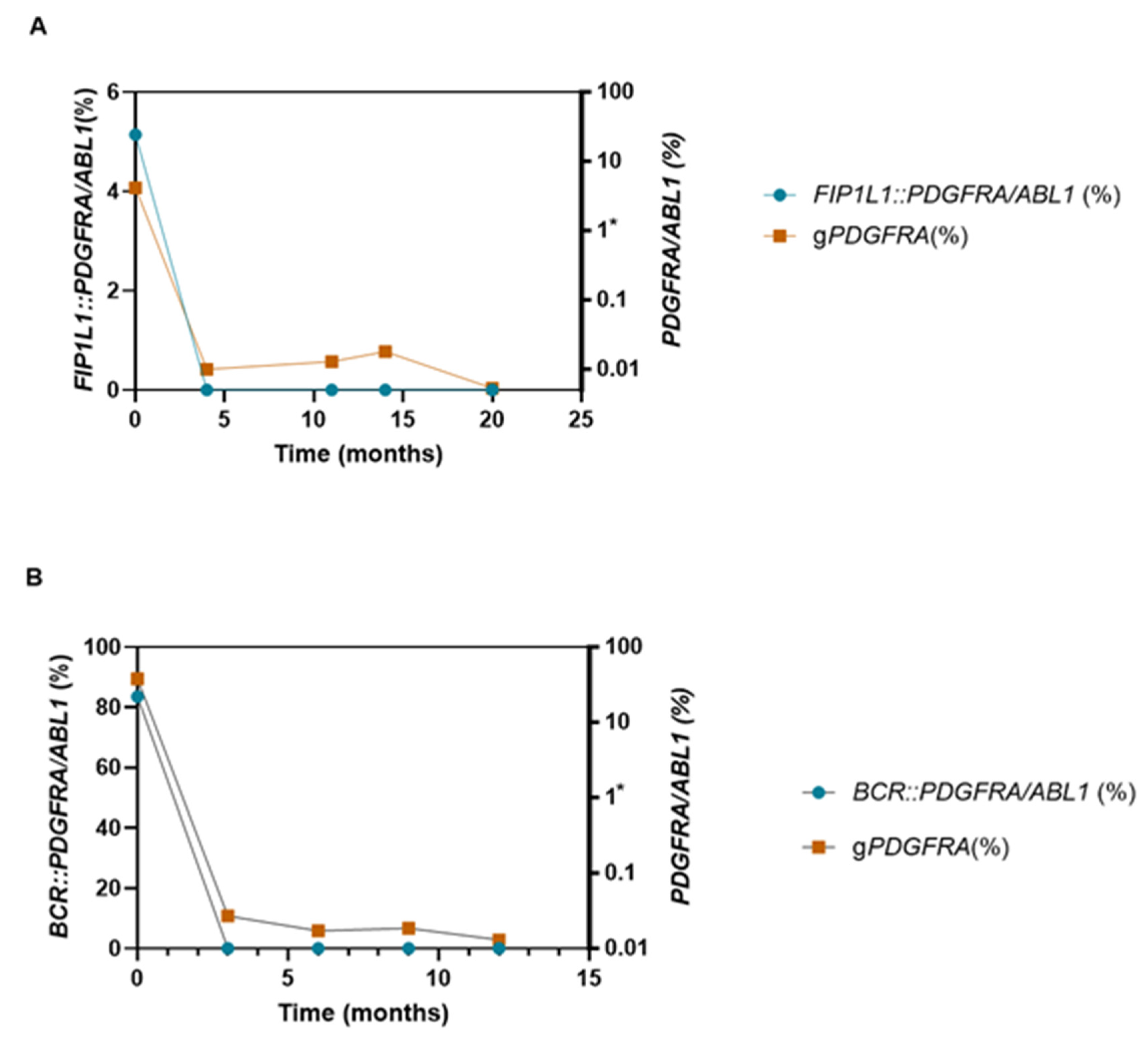
| UPN | Sex | Age (years) | Hematologic diseases | Rearrangement | % overexpression | Eosinophils (10⁹/L) | Tryptase (µg/L) | Viamin B12 (pg/mL) | Bone marrow | Karyotype | FISH | RT-PCR or RT-MLPA | TKI treatment | Time to remission |
| P1 | M | 49 | MPN | FIP1L1::PDGFRA | 6,85 | 4,4 | unk | 1902 | HE > 20% | Normal | n.t | FIP1L1::PDGFRA | Imatinib 100 mg/d | unk |
| P2 | M | 85 | MPN | FIP1L1::PDGFRA | 10,1 | 6,42 | 51,9 | 890 | unk | n.t | n.t | FIP1L1::PDGFRA | Imatinib 100 mg/d | unk |
| P3 | M | 84 | MPN | FIP1L1::PDGFRA | 4,23 | 3,25 | n.t | 367 | unk | Normal | n.t | FIP1L1::PDGFRA | Imatinib 100 mg/d | 3 months |
| P4 | M | 42 | MPN | FIP1L1::PDGFRA | 4,16 | 2,24 | 22 | 2160 | Mild HE | Normal | CHIC2 deletion | FIP1L1::PDGFRA | Imatinib 100 mg/d | unk |
| P5 | M | 60 | MPN | FIP1L1::PDGFRA | 116,64 | 10,44 | n.t | n.t | unk | Normal | n.t | FIP1L1::PDGFRA | unk | unk |
| P6 | M | 58 | MPN | FIP1L1::PDGFRA | 51,37 | 21 | Normal | 2500 | HE | Normal | CHIC2 deletion | FIP1L1::PDGFRA | Imatinib 100 mg/d | unk |
| P7 | M | 54 | MPN | FIP1L1::PDGFRA | 17,93 | 4,4 | unk | 1902 | unk | n.t | n.t | FIP1L1::PDGFRA | Imatinib 100 mg/d | unk |
| P8 | M | 75 | CMML | FIP1L1::PDGFRA | 13,29 | 14,5 | n.t | 8500 | unk | Normal | CHIC2 deletion | FIP1L1::PDGFRA | Imatinib 100 mg/d | unk |
| P9 | M | 27 | MPN | FIP1L1::PDGFRA | 2,53 | 88,13 | 38 | n.t | unsuitable | n.t | n.t | FIP1L1::PDGFRA | Imatinib 100 mg/d | unk |
| P10 | F | 77 | MPN | FIP1L1::PDGFRA | 9,81 | 6,22 | unk | unk | HE=15% | Normal | CHIC2 deletion | FIP1L1::PDGFRA | Imatinib 100 mg/d | 3 months |
| P11 | F | 48 | MPN | FIP1L1::PDGFRA | 12,17 | 1,43 | unk | unk | HE=13% | Normal | CHIC2 deletion | FIP1L1::PDGFRA | « Watch and see » | unk |
| P12 | M | 76 | MPN | FIP1L1::PDGFRA | 10,78 | unk | unk | unk | unk | unk | unk | FIP1L1::PDGFRA | unk | unk |
| P13 | F | 78 | MPN | BCR::PDGFRA | 83,52 | 3,11 | unk | unk | unk | t(4;22) | unk | BCR ::PDGFRA | Imatinib 400 mg/d | 6 months |
| P14 | F | 91 | MDS-IB2 | ETV6::PDGFRA | 21,4 | 0,03 | unk | unk | MDS-IB2 | t(4;12) | ETV6::PDGFRA rearrangement | n.t | Imatinib 100 mg/d | unk |
| P15 | M | 57 | AML | ETV6::PDGFB | 98,27 | 0,6 | n.t | n.t | AML | t(5;12) | ETV6::PDGFRB rearrangement | n.t | Imatinib 400 mg/d | unk |
| P16 | M | 77 | MPN | ETV6::PDGFRB | 60,5 | 50,68 | n.t | n.t | HE | t(5;12) | ETV6::PDGFRB rearrangement | Negative | Ruxolitinib | unk |
| P17 | M | 58 | AML | ETV6 ::PDGFRB | 98,36 | unk | n.t | n.t | unk | unk | n.t | ETV6 ::PDGFRB | Imatinib 400 mg/d | 1 months |
| P18 | M | 77 | CMML | CCDC88C::PDGFRB | 85,09 | 2,19 | n.t | n.t | CMML | t(5;14) | PDGFRB rearrangement | n.t | Imatinib 100 mg/d | unk |
| P19 | M | 70 | CML | CCDC6::PDGFRB | 40,7 | 1,9 | n.t | n.t | MPN | t(5;10) | CCDC6::PDGFRB rearrangement | n.t | Imatinib 200 mg/d | unk |
| P20 | M | 93 | MPN | PDGFRB unknown partner | 82 | unk | n.t | n.t | unk | unk | PDGFRB rearrangement | n.t | unk | unk |
| P21 | M | 65 | MPN | PDGFRB unknown partner |
107 | 16,28 | n.t | >1476 | unk | n.t | PDGFRB rearrangement | n.t | Imatinib 100 mg/d | unk |
| P22 | M | 25 | ALL T | PDGFRB unknown partner | 65,37 | 1,3 | n.t | n.t | ALL | t(5;12) | PDGFRB rearrangement | Negative | Imatinib 500 mg/d | unk |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




