Submitted:
12 February 2026
Posted:
13 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
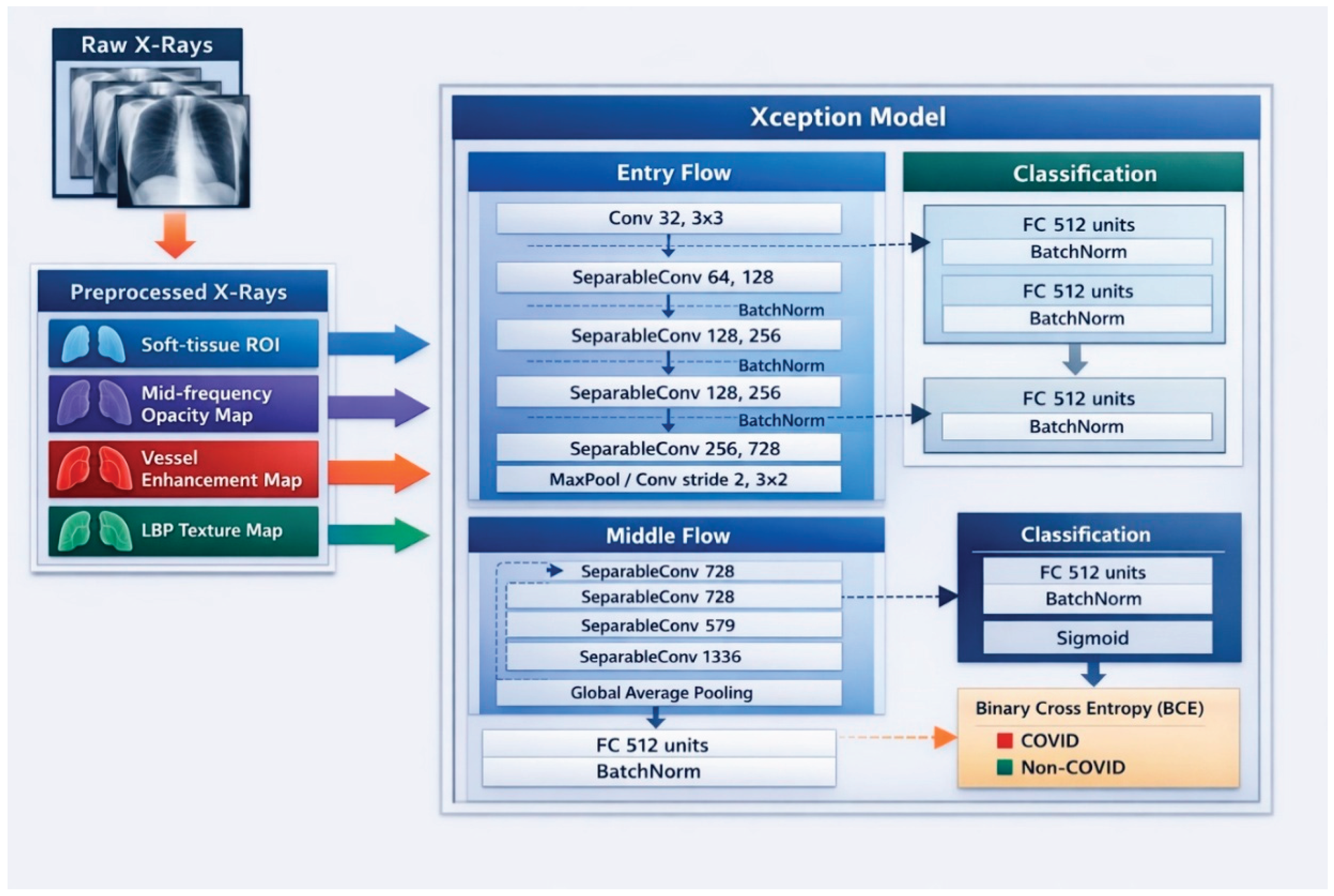
- • A novel four-channel input representation for thoracic chest X-ray analysis was proposed, integrating lung-region masking, frequency-domain enhancement, vesselness filtering, and texture-based features. While individual preprocessing and feature-enhancement techniques have been explored in prior chest radiograph studies, an identical combination of these anatomically and diagnostically motivated channels has not been identified in the reviewed literature. This novel four-channel combination provides complementary information beyond a single intensity channel, supporting improved sensitivity and anatomically aligned model attention when combined with explainable AI techniques.
- • A structured, explainable deep learning framework was developed for thoracic medical image analysis, integrating the proposed multi-channel input representation with a modified deep convolutional neural network architecture. The framework was designed to balance diagnostic performance and interpretability, addressing the limitations of conventional single-channel pipelines and black-box deep learning models commonly reported in medical imaging literature.
- • A novel approach to interpreting and communicating model attention was introduced by combining quantitative spatial attention analysis with rule-based natural-language explanations. Rather than presenting saliency maps solely as visual artefacts, this study quantified the distribution of model attention inside and outside lung regions and translated these measurements into concise, human-readable explanations. This structured explanation strategy improved the accessibility and interpretability of explainable AI outputs for non-technical users, including clinicians.
- • The proposed framework enhanced clinical relevance and trustworthiness of AI-assisted diagnosis, by ensuring that model attention was anatomically meaningful and aligned with lung regions of interest. This design supported transparent decision-making and addressed key ethical, regulatory, and usability concerns associated with the deployment of deep learning models in real-world clinical settings.
- • The study provided practical insights into the integration of explainable AI within medical imaging workflows, demonstrating how anatomically guided preprocessing, multi-channel learning, and explainability mechanisms can be combined into a cohesive and computationally feasible diagnostic system.
2. Literature Review
2.1. Deep Learning for Chest X-Ray Classification
2.2. Explainable Artificial Intelligence in Medical Imaging
2.3. Addressing Gaps and Advancing Knowledge
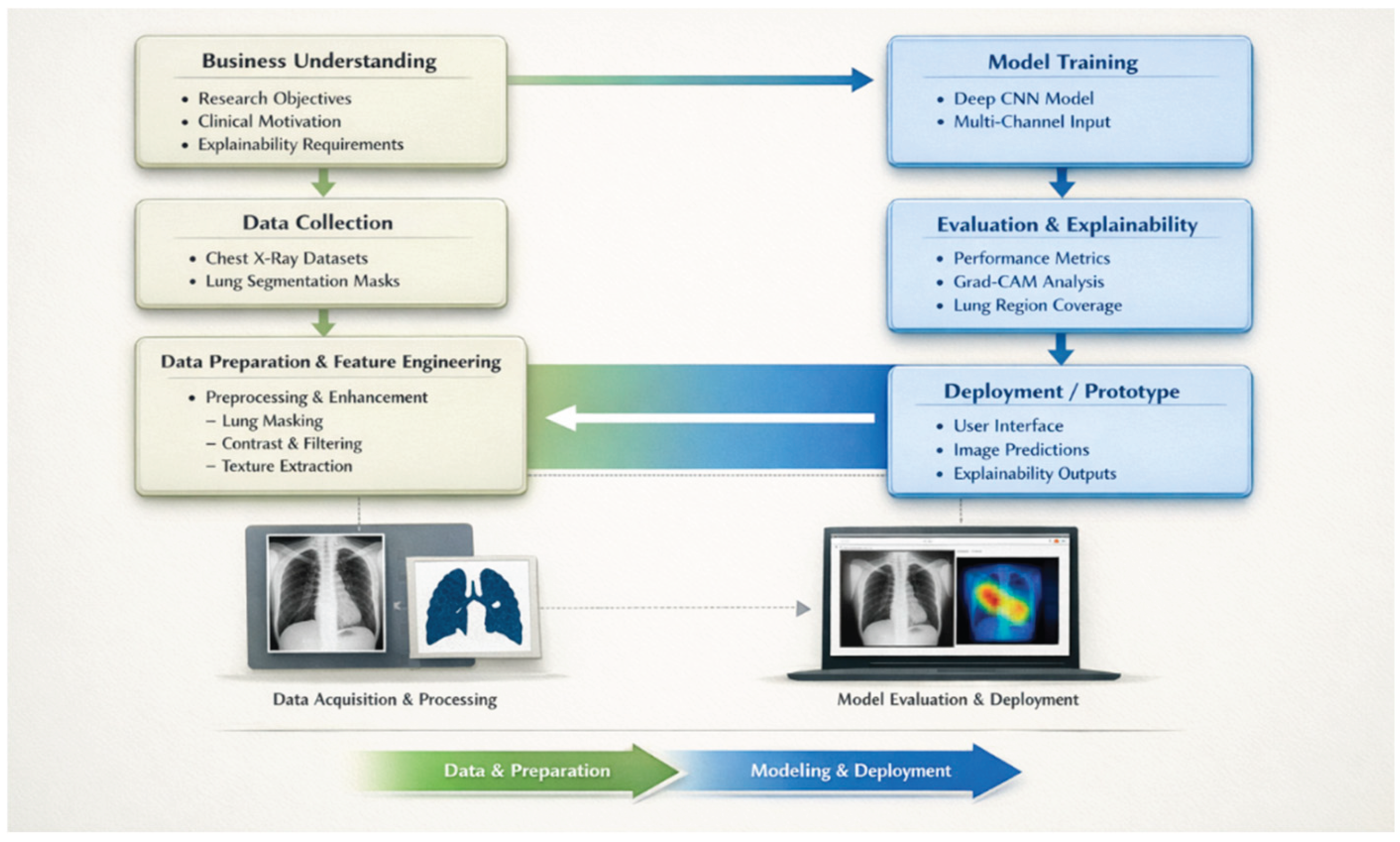
3. Proposed Methodology
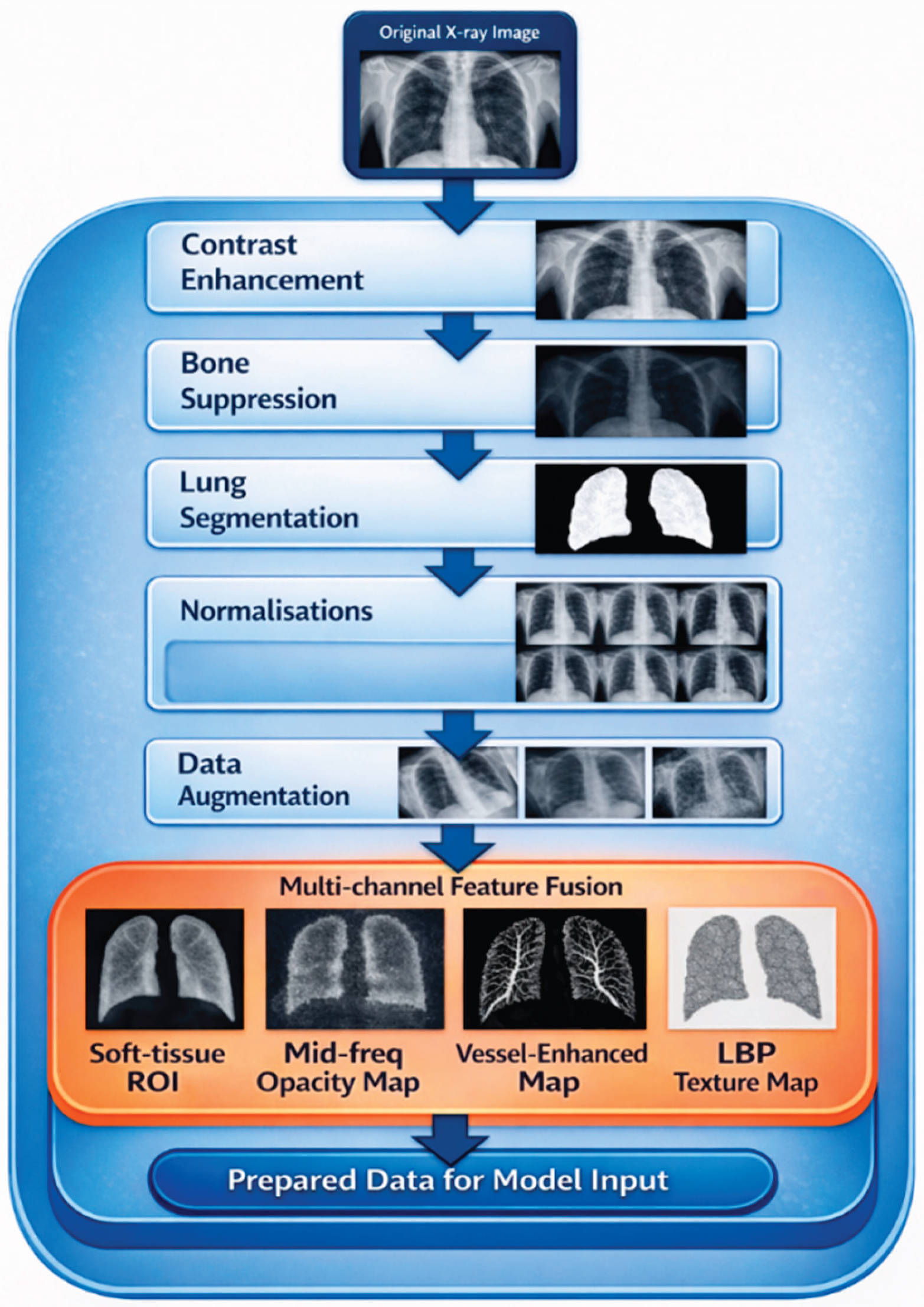
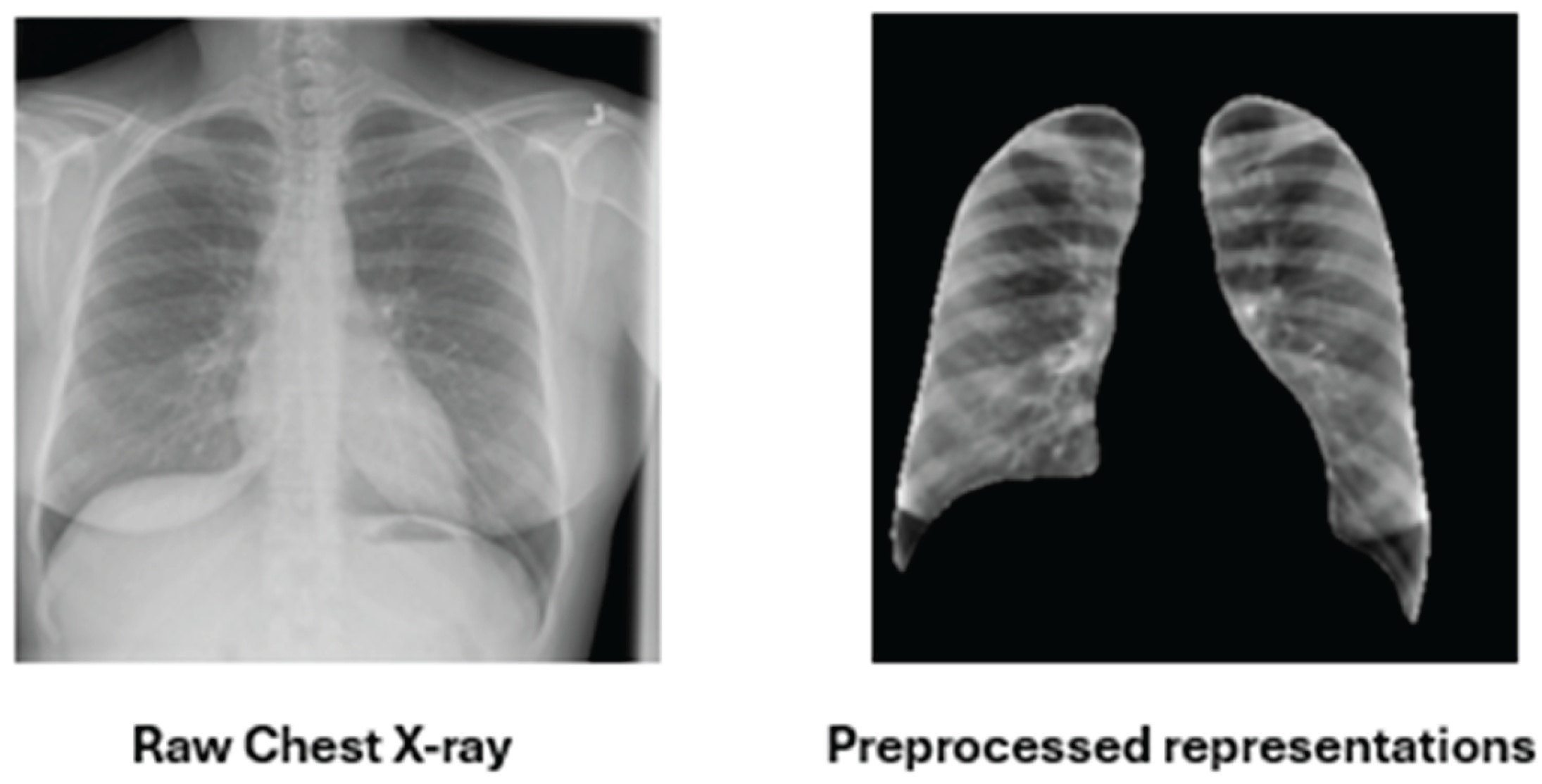
3.1. Image Preprocessing and Normalisation
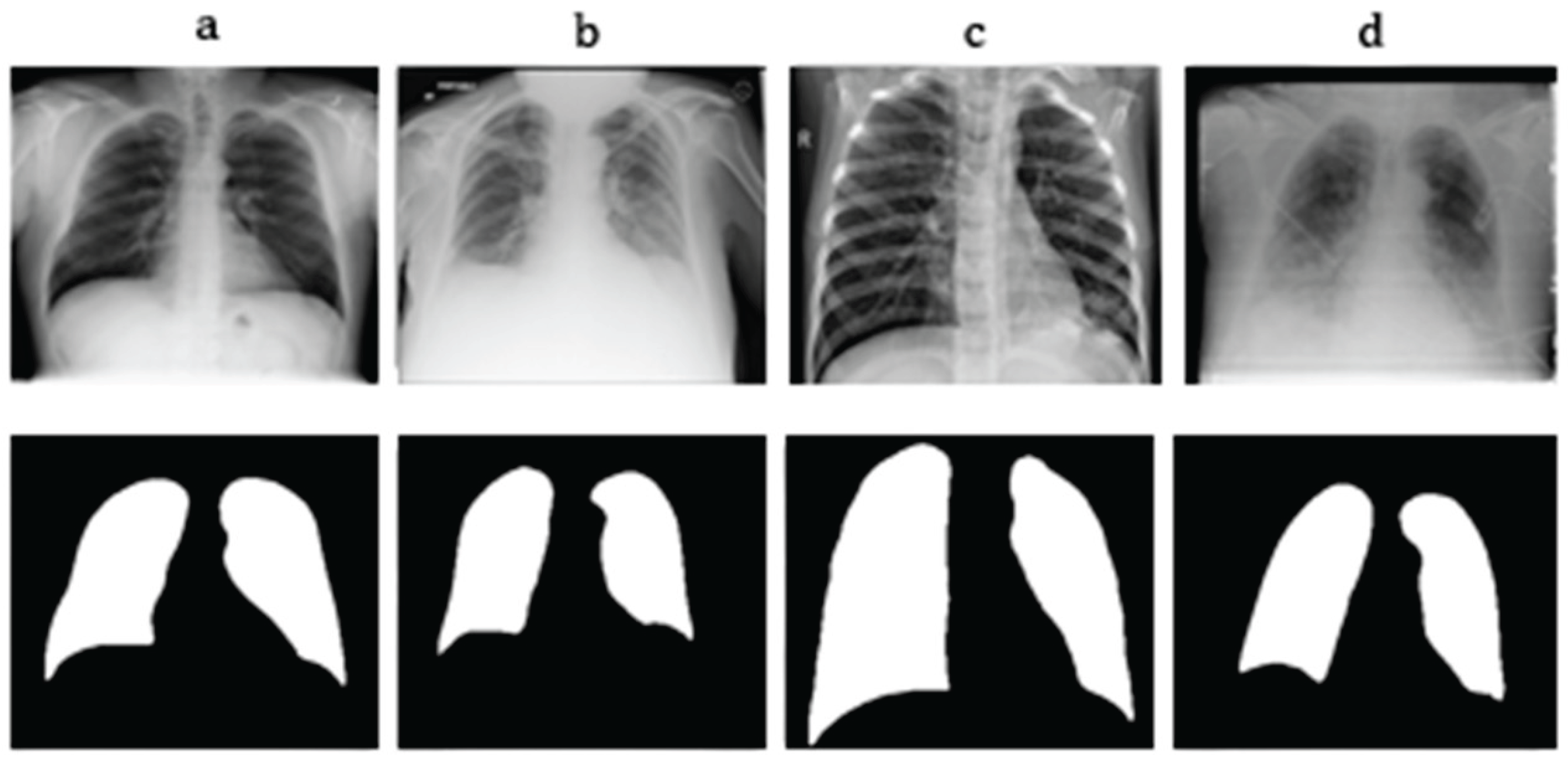
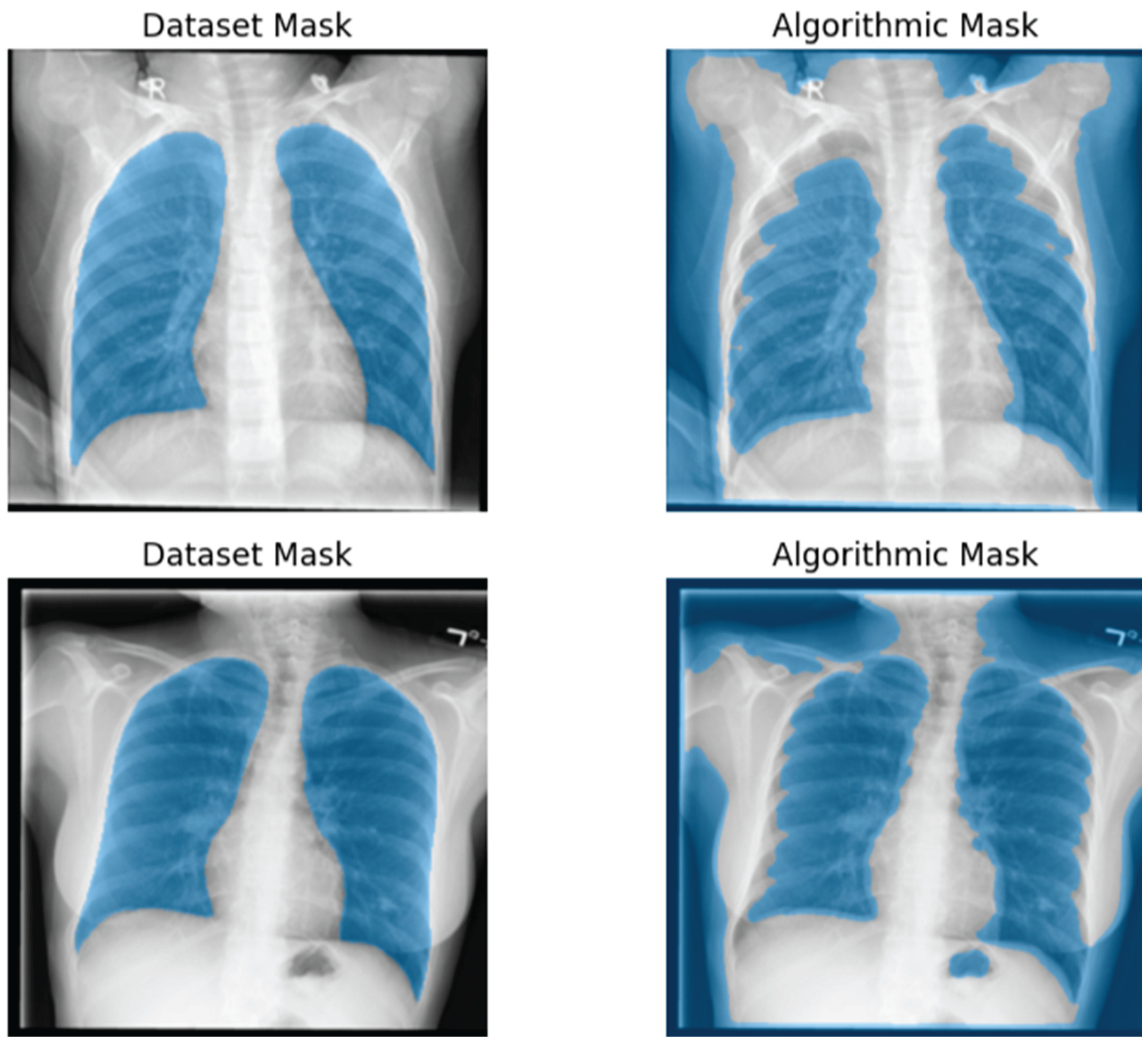
3.1.1. Pulmonary Region of Interest (ROI) Extraction
- I.
- Lung Region Isolation
- i.
- Segmentation Phase: Otsu’s Thresholding
- ii.
- Formula for Between-Class Variance:where:
- iii.
- Binary Mask Result:where denotes the image intensity at pixel location .
- iv.
- Geometric Phase: Connected Components
- v.
- Assuming the lungs correspond to the two largest contiguous dark regions, the mask was filtered by retaining the two largest components:
- vi.
- Formula for Component Area:
- vii.
- Validation Phase: Area Fraction Heuristic
- viii.
- Decision Rule
- II.
- Soft-Tissue Enhancement: CLAHE and Bone Suppression
- is the normalised histogram
- is the number of grayscale levels
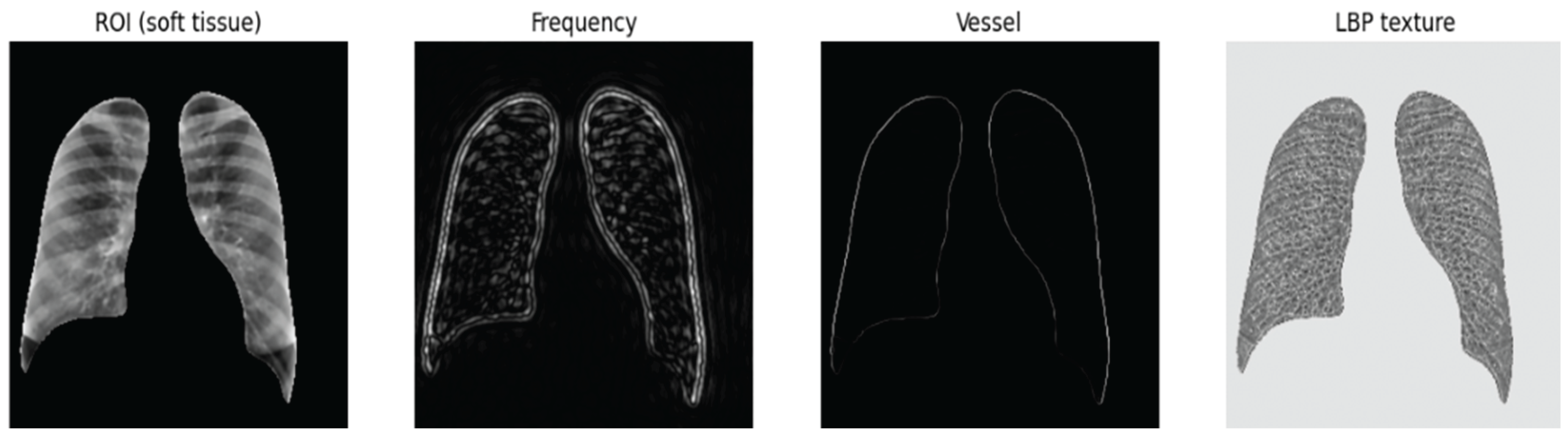
3.1.2. Multi-Channel Feature Construction
- Mid-frequency opacity mapping (Fourier band-pass filtering)
- a)
- 2D DFT:
- b)
- Distance to Center:
- c)
- Filter Mask (Ideal Band-Pass):
- d)
- Filtered Transform:
- e)
- Inverse DFT:
- II.
- Vessel enhancement using the Frangi filter
- III.
- Texture encoding using Local Binary Patterns (LBP)
3.2. Normalisation, Class Balancing, and Augmentation
3.3. Model Architecture
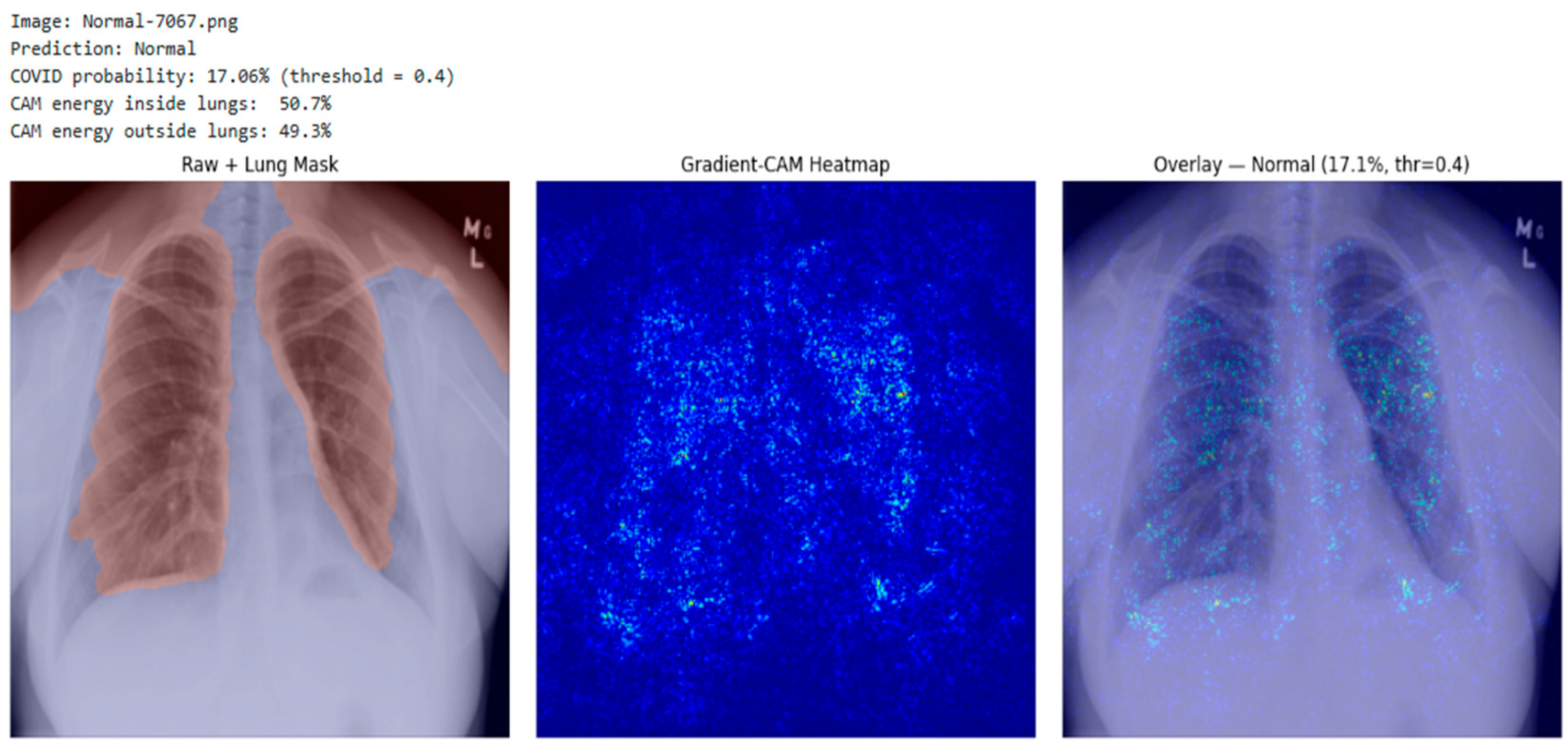
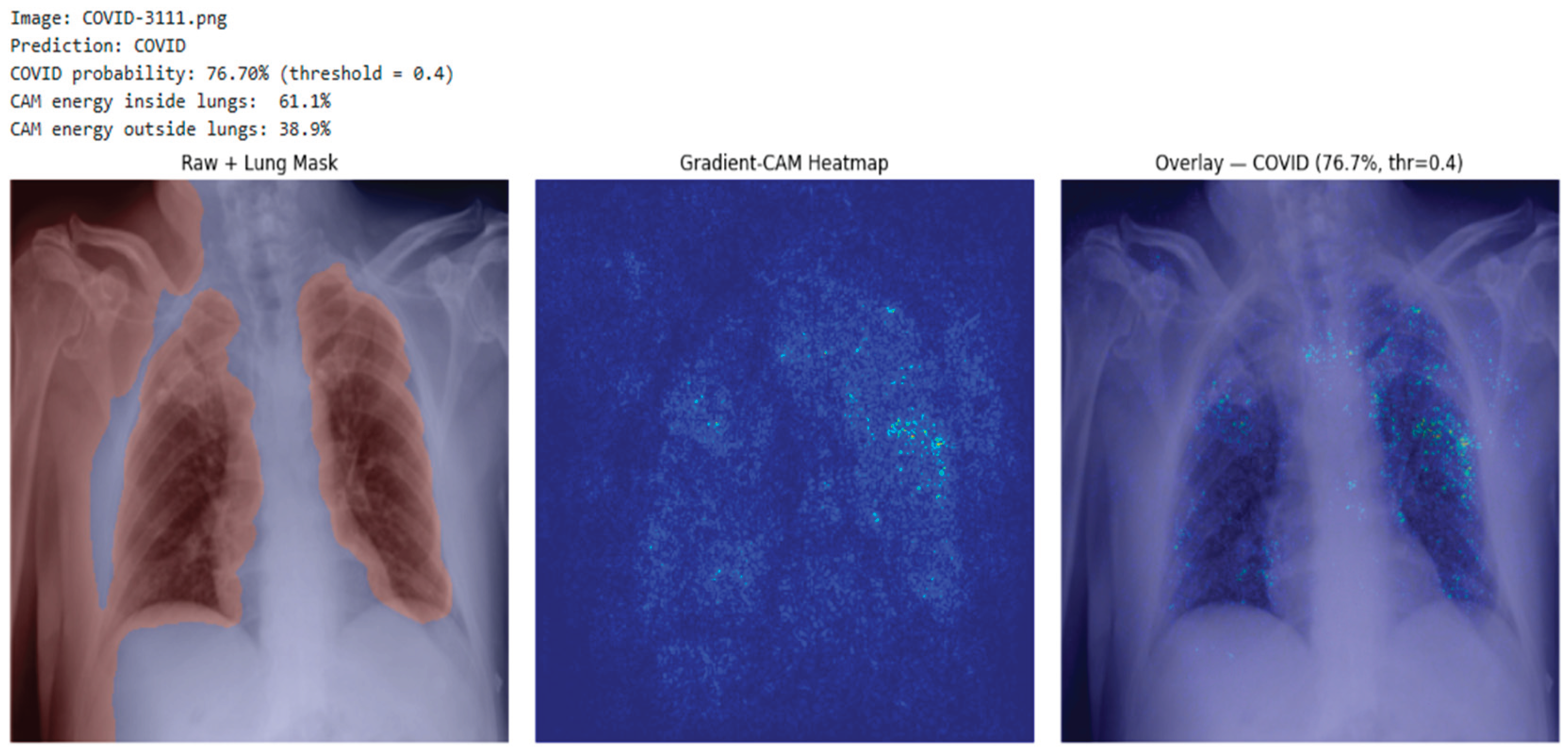
3.4. Explainability Integration
3.5. Quantitative Classification Performance
- (a)
- CAM Energy
4. Experimental Results and Analysis
4.1. Implementation Setup
4.2. Model Training Configuration
4.3. Quantitative Classification Performance
4.4. Quantitative Classification Performance
- (a)
- CAM Energy
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mercaldo, F.; Belfiore, M.P.; Reginelli, A.; Brunese, L.; Santone, A. Coronavirus covid-19 detection by means of explainable deep learning. Scientific Reports 2023, 13, 462. [Google Scholar] [CrossRef]
- Chadaga, K.; Prabhu, S.; Sampathila, N.; Chadaga, R.; Umakanth, S.; Bhat, D.; GS, S.K. Explainable artificial intelligence approaches for COVID-19 prognosis prediction using clinical markers. Scientific Reports 2024, 14, 1783. [Google Scholar] [CrossRef]
- Pham, N.T.; Ko, J.; Shah, M.; Rakkiyappan, R.; Woo, H.G.; Manavalan, B. Leveraging deep transfer learning and explainable AI for accurate COVID-19 diagnosis: Insights from a multi-national chest CT scan study. Comput. Biol. Med. 2025, 185, 109461. [Google Scholar] [CrossRef] [PubMed]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef] [PubMed]
- Esteva, A.; Robicquet, A.; Ramsundar, B.; Kuleshov, V.; DePristo, M.; Chou, K.; Cui, C.; Corrado, G.; Thrun, S.; Dean, J. A guide to deep learning in healthcare. Nat. Med. 2019, 25, 24–29. [Google Scholar] [CrossRef]
- Shobayo, O.; Saatchi, R. Developments in Deep Learning Artificial Neural Network Techniques for Medical Image Analysis and Interpretation. Diagnostics 2025, 15, 1072. [Google Scholar] [CrossRef] [PubMed]
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017, 542, 115–118. [Google Scholar] [CrossRef]
- Rajpurkar, P.; Irvin, J.; Zhu, K.; et al. CheXNet: Radiologist-level pneumonia detection on chest X-rays with deep learning. arXiv 2017, arXiv:1711.05225. [Google Scholar]
- El-Magd, L.M.A.; Dahy, G.; Farrag, T.A.; Darwish, A.; Hassnien, A.E. An interpretable deep learning based approach for chronic obstructive pulmonary disease using explainable artificial intelligence. International Journal of Information Technology 2025, 17, 4077–4092. [Google Scholar] [CrossRef]
- Adadi, A.; Berrada, M. Peeking inside the black box: A survey on explainable artificial intelligence (XAI). IEEE Access 2018, 6, 52138–52160. [Google Scholar] [CrossRef]
- Solayman, S.; Aumi, S.A.; Mery, C.S.; Mubassir, M.; Khan, R. Automatic COVID-19 prediction using explainable machine learning techniques. International Journal of Cognitive Computing in Engineering 2023, 4, 36–46. [Google Scholar] [CrossRef]
- Wachter, S.; Mittelstadt, B.; Floridi, L. Why a right to explanation of automated decision-making does not exist in the GDPR. Int. Data Priv. Law 2017, 7, 76–99. [Google Scholar] [CrossRef]
- Singh, J.; Sillerud, B.; Yednock, J.; Larson, C.; Steffen, A.; Singh, A. Healthcare leaders’ attitudes and perceptions on the use of artificial intelligence and artificial intelligence enabled tools in healthcare settings. Journal of Medical Artificial Intelligence 2025, 8, 41. [Google Scholar] [CrossRef]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual explanations from deep networks via gradient-based localization. Proc. IEEE ICCV 2017, 618–626. [Google Scholar]
- Gulum, M.A.; Trombley, C.M.; Kantardzic, M. Explainable deep learning for medical image analysis: A survey. J. Imaging 2021, 7, 102. [Google Scholar]
- Samek, W.; Montavon, G.; Vedaldi, A.; Hansen, L.K.; Müller, K. Explainable AI: interpreting, explaining and visualizing deep learning; Springer Nature, 2019; Vol. 11700. [Google Scholar]
- Holzinger, A.; Langs, G.; Denk, H.; Zatloukal, K.; Müller, H. Causability and explainability of artificial intelligence in medicine. WIREs Data Min. Knowl. Discov. 2019, 9, e1312. [Google Scholar] [CrossRef]
- Slack, D.; Hilgard, A.; Jia, E.; Singh, S.; Lakkaraju, H. Fooling LIME and SHAP: Adversarial attacks on post hoc explanation methods. Proc. AAAI Conf. Artif. Intell. 2020, 34, 180–187. [Google Scholar]
- Nneji, G.U.; Cai, J.; Deng, J.; Su, S. Multi-channel deep learning framework for chest X-ray classification. Comput. Biol. Med. 2022, 145, 105410. [Google Scholar]
- Çallı, E.; Sogancioglu, E.; van Ginneken, B.; van Leeuwen, K.G.; Murphy, K. Deep learning for chest X-ray analysis: A survey. Med. Image Anal. 2021, 72, 102125. [Google Scholar] [CrossRef]
- Ait Nasser, A.; Jilbab, A.; Bourouhou, A. Deep learning for chest X-ray analysis: A survey. Artif. Intell. Rev. 2023, 56, 1243–1290. [Google Scholar]
- Arun, N.; Gaw, N.; Singh, P.; Chang, K.; Aggarwal, M.; Chen, B.; Hoebel, K.; Gupta, S.; Patel, J.; Gidwani, M.; et al. Assessing the trustworthiness of saliency maps for localizing abnormalities in medical imaging. Radiol. Artif. Intell. 2021, 3, e200267. [Google Scholar] [CrossRef]
- Singh, P.; Gaw, N.; Chang, K.; et al. Are saliency maps trustworthy? A large-scale evaluation in medical imaging. Med. Image Anal. 2022, 75, 102327. [Google Scholar]
- Adebayo, J.; Gilmer, J.; Muelly, M.; Goodfellow, I.; Hardt, M.; Kim, B. Sanity checks for saliency maps. Advances in neural information processing systems 2018, 31. [Google Scholar]
- Tjoa, E.; Guan, C. A survey on explainable artificial intelligence (XAI): Toward medical XAI. IEEE Trans. Neural Netw. Learn. Syst. 2020, 32, 4793–4813. [Google Scholar] [CrossRef]
- Wani, N.A.; Kumar, R.; Bedi, J. DeepXplainer: An interpretable deep learning based approach for lung cancer detection using explainable artificial intelligence. Comput. Methods Programs Biomed. 2024, 243, 107879. [Google Scholar] [CrossRef]
- Wang, X.; Peng, Y.; Lu, L.; Lu, Z.; Bagheri, M.; Summers, R.M. ChestX-ray8: Hospital-scale chest X-ray database and benchmarks on weakly supervised classification and localization of common thorax diseases. Proc. IEEE CVPR 2017, 2097–2106. [Google Scholar]
- Irvin, J.; Rajpurkar, P.; Ko, M.; Yu, Y.; Ciurea-Ilcus, S.; Chute, C.; et al. CheXpert: A large chest radiograph dataset with uncertainty labels and expert comparison. Proc. AAAI Conf. Artif. Intell. 2019, 33, 590–597. [Google Scholar] [CrossRef]
- Tang, Y.; Tang, Y.; Peng, Y.; Yan, K.; Bagheri, M.; Redd, B.A.; Brandon, C.J.; Lu, Z.; Han, M.; Xiao, J. Automated abnormality classification of chest radiographs using deep convolutional neural networks. NPJ digital medicine 2020, 3, 70. [Google Scholar] [CrossRef]
- Wang, L.; Lin, Z.Q.; Wong, A. COVID-Net: A tailored deep convolutional neural network design for detection of COVID-19 cases from chest X-ray images. Sci. Rep. 2020, 10, 19549. [Google Scholar] [CrossRef] [PubMed]
- Narin, A.; Kaya, C.; Pamuk, Z. Automatic detection of coronavirus disease (COVID-19) using X-ray images and deep convolutional neural networks. Pattern Anal. Appl. 2021, 24, 1207–1220. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J.P.; Hashir, M.; Brooks, R.; Bertrand, H. On the limits of cross-domain generalization in automated X-ray prediction. Proc. Med. Imaging Deep Learn. (MIDL) 2020, 136–155. [Google Scholar]
- Maguolo, G.; Nanni, L. A critic evaluation of methods for COVID-19 automatic detection from X-ray images. Inf. Fusion 2021, 76, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, H.; Liu, Q. Lung segmentation improves explainability of deep learning models for chest X-ray classification. Comput. Methods Programs Biomed. 2021, 207, 106173. [Google Scholar]
- Babu, P.A.; Rai, A.K.; Ramesh, J.V.N.; Nithyasri, A.; Sangeetha, S.; Kshirsagar, P.R.; Rajendran, A.; Rajaram, A.; Dilipkumar, S. An explainable deep learning approach for oral cancer detection. Journal of Electrical Engineering & Technology 2024, 19, 1837–1848. [Google Scholar]
- Lundberg, S.M.; Lee, S.I. A unified approach to interpreting model predictions. Adv. Neural Inf. Process. Syst. 2017, 30, 4765–4774. [Google Scholar]
- Aasem, M.; Javed Iqbal, M. Toward explainable AI in radiology: Ensemble-CAM for effective thoracic disease localization in chest X-ray images using weak supervised learning. Front. Big Data 2024, 7, 1366415. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Li, O.; Tao, D.; Barnett, A.; Rudin, C.; Su, J.K. This looks like that: deep learning for interpretable image recognition. Advances in neural information processing systems 2019, 32, 8930–8941. [Google Scholar]
- Koh, P.W.; Nguyen, T.; Tang, Y.S.; Mussmann, S.; Pierson, E.; Kim, B.; Liang, P. In In Concept bottleneck models; International conference on machine learning; PMLR: 2020; 5338–5348.
- Sultana, S.; Hossain, A.A.; Alam, J. COVID-19 detection from optimized features of breathing audio signals using explainable ensemble machine learning. Results in Control and Optimization 2025, 18, 100538. [Google Scholar] [CrossRef]
- Pino, C.; Carrara, F.; Bellio, M.; Neri, E. Influence of explainable AI on trust in medical image analysis. Artif. Intell. Med. 2021, 115, 102079. [Google Scholar]
- Mohammed, M.A.; Abdulkareem, K.H.; Garcia-Zapirain, B.; Mostafa, S.A.; Maashi, M.S. A comprehensive investigation of deep learning-based COVID-19 detection from chest X-ray images. Comput. Biol. Med. 2022, 136, 104730. [Google Scholar]
- Suzuki, K.; Abe, H.; MacMahon, H.; Doi, K. Image-processing technique for suppressing ribs in chest radiographs by means of massive training artificial neural network. IEEE Trans. Med. Imaging 2006, 25, 406–416. [Google Scholar] [CrossRef]
- Harrison, A.P.; Xu, Z.; George, K.; Lu, L.; Summers, R.M.; Mollura, D.J. Progressive and multi-path holistic lung segmentation from chest X-ray images. Med. Image Anal. 2021, 67, 101840. [Google Scholar]
- Saha, M.; Chakraborty, C.; Racoceanu, D. Efficient deep learning model for explainable COVID-19 detection from chest X-ray images. Diagnostics. 2021, 11, 1772. [Google Scholar]
- Rahman, T.; Chowdhury, M.E.H.; Khandakar, A.; et al. COVID-19 radiography database. arXiv. 2020, arXiv:2005.06794. [Google Scholar]
- Jacobi, A.; Chung, M.; Bernheim, A.; Eber, C. Portable chest X-ray in coronavirus disease-19 (COVID-19): A pictorial review. Clin. Imaging. 2020, 64, 35–42. [Google Scholar] [CrossRef]
- Pisano, E.D.; Zong, S.; Hemminger, B.M.; et al. Contrast limited adaptive histogram equalization image processing to improve the detection of simulated spiculations in dense mammograms. J. Digit. Imaging 1998, 11, 193–200. [Google Scholar] [CrossRef]
- Pizer, S.M.; Amburn, E.P.; Austin, J.D.; et al. Adaptive histogram equalization and its variations. Comput. Vis. Graph. Image Process. 1990, 39, 355–368. [Google Scholar] [CrossRef]
- Chung, A.G.; Shafiee, M.J.; Khalvati, F.; Haider, M.A.; Wong, A. Radiomics-based multi-scale texture analysis for infectious lung disease characterization. Comput. Biol. Med. 2021, 133, 104372. [Google Scholar]
- Frangi, A.F.; Niessen, W.J.; Vincken, K.L.; Viergever, M.A. Multiscale vessel enhancement filtering. Lect. Notes Comput. Sci. (MICCAI) 1998, 130–137. [Google Scholar]
- Carotti, M.; Salaffi, F.; Sarzi-Puttini, P.; Agostini, A.; Borgheresi, A.; Minorati, D.; Galli, M.; Giovagnoni, A. Chest CT features of coronavirus disease 2019 (COVID-19) pneumonia: Key points for radiologists. Radiol. Med. 2020, 125, 636–646. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, S.; Dong, D.; Tian, J.; Zhou, X. Vascular feature learning for pulmonary disease diagnosis. Med. Image Anal. 2019, 58, 101541. [Google Scholar]
- Ojala, T.; Pietikäinen, M.; Mäenpää, T. Multiresolution gray-scale and rotation invariant texture classification with local binary patterns. IEEE Trans. Pattern Anal. Mach. Intell. 2002, 24, 971–987. [Google Scholar] [CrossRef]
- Zhou, S.K.; Greenspan, H.; Davatzikos, C.; Duncan, J.S.; van Ginneken, B.; Madabhushi, A.; Prince, J.L.; Rueckert, D.; Summers, R.M. A review of deep learning in medical imaging: Imaging traits, technology trends, case studies with progress highlights, and future promises. Proc. IEEE 2021, 109, 820–838. [Google Scholar] [CrossRef] [PubMed]
- Chicco, D.; Jurman, G. The advantages of the Matthews correlation coefficient over F1 score and accuracy in binary classification evaluation. BMC Genomics 2020, 21, 6. [Google Scholar] [CrossRef] [PubMed]
| Diagnostic Class | Number of Images | Description |
|---|---|---|
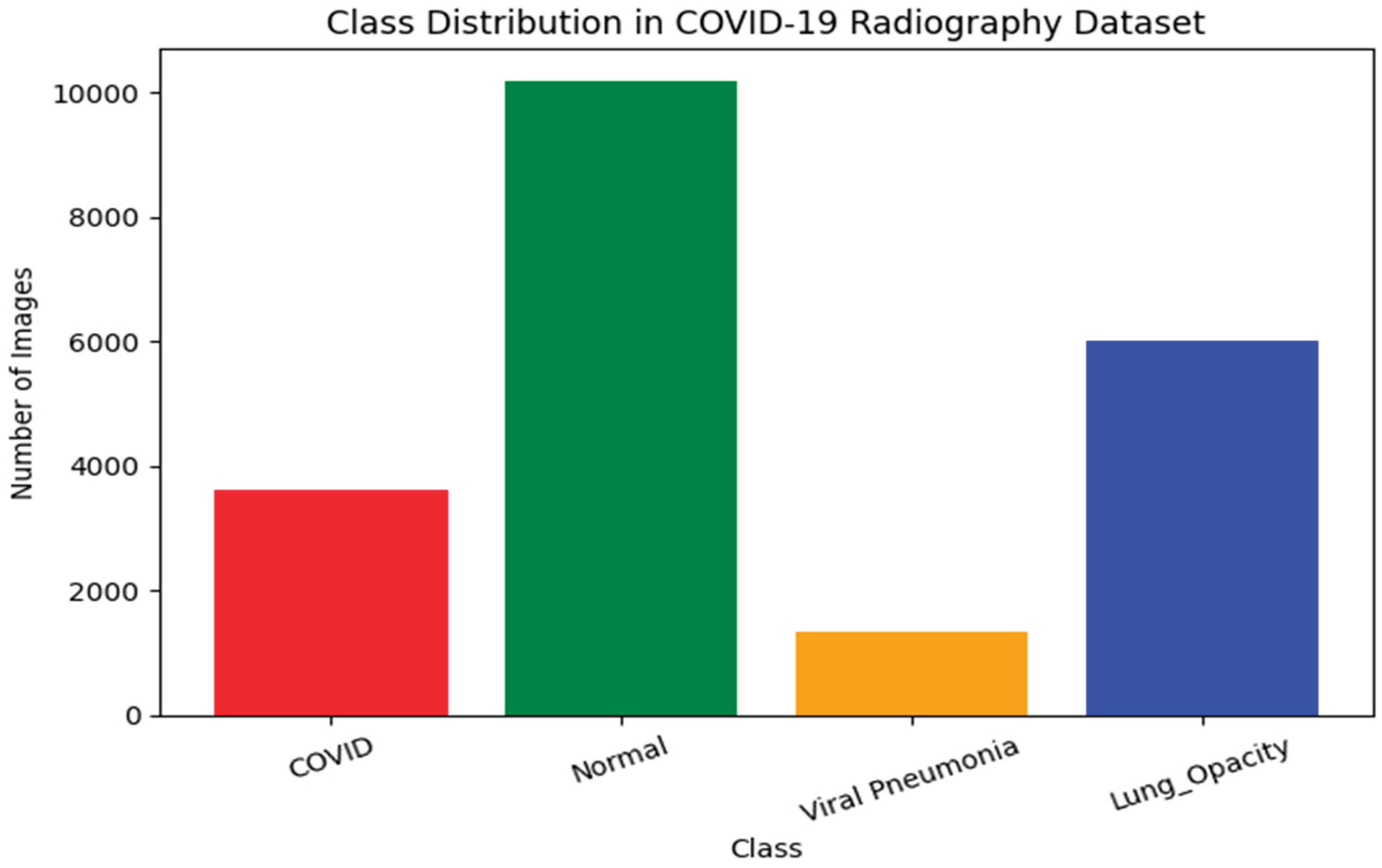
| COVID-19 | 3,616 | Confirmed COVID-19 radiographs sourced from curated public repositories. |
| Normal | 10,192 | Chest radiographs with clear lung fields and no radiographic abnormalities. |
| Lung Opacity | 6,012 | Images showing non-COVID pulmonary opacities caused by various conditions. |
| Viral Pneumonia | 1,345 | Radiographs depicting viral pneumonia distinct from COVID-19. |
| Threshold | Accuracy | Precision | Recall | F1 | MCC |
|---|---|---|---|---|---|
| 0.05 | 0.70 | 0.37 | 0.99 | 0.53 | 0.48 |
| 0.10 | 0.82 | 0.49 | 0.99 | 0.65 | 0.61 |
| 0.15 | 0.88 | 0.58 | 0.98 | 0.73 | 0.69 |
| 0.20 | 0.91 | 0.67 | 0.97 | 0.80 | 0.76 |
| 0.25 | 0.93 | 0.71 | 0.95 | 0.81 | 0.78 |
| 0.30 | 0.94 | 0.76 | 0.93 | 0.84 | 0.81 |
| 0.35 | 0.95 | 0.81 | 0.91 | 0.86 | 0.83 |
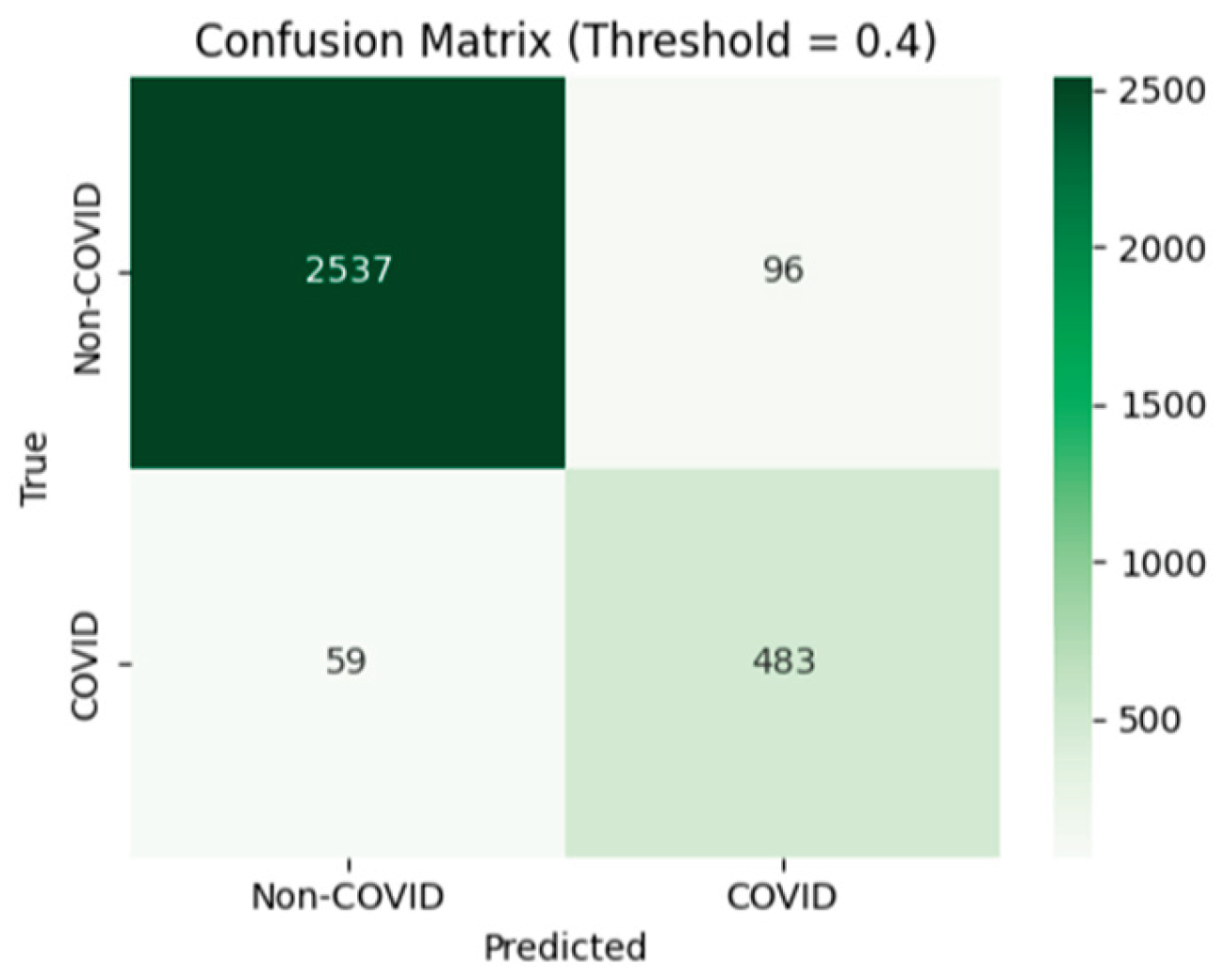
| 0.40 | 0.95 | 0.83 | 0.89 | 0.86 | 0.83 |
| 0.45 | 0.95 | 0.85 | 0.87 | 0.86 | 0.83 |
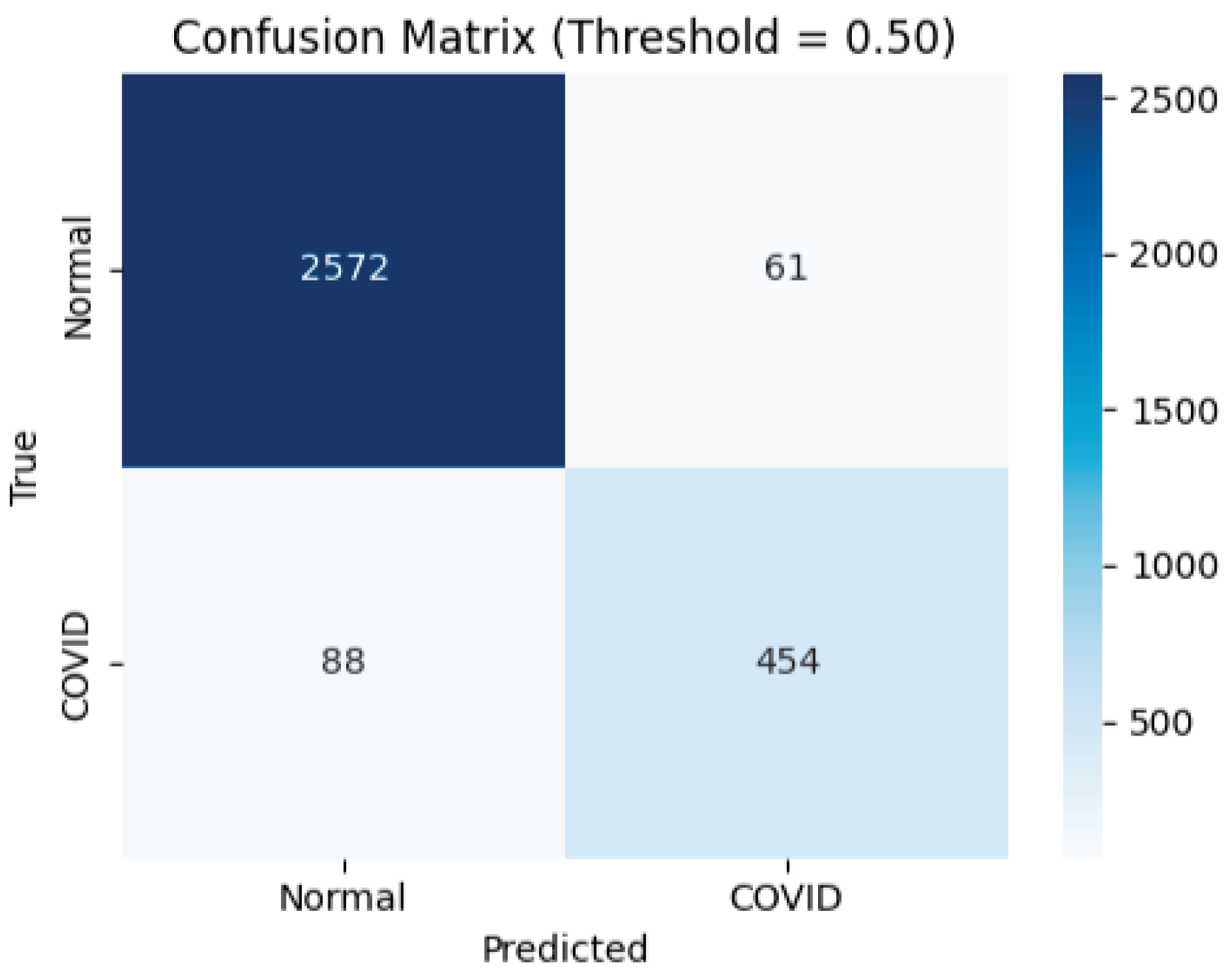
| 0.50 | 0.95 | 0.88 | 0.84 | 0.86 | 0.83 |
| 0.55 | 0.95 | 0.90 | 0.82 | 0.86 | 0.83 |
| 0.60 | 0.95 | 0.93 | 0.78 | 0.85 | 0.82 |
| 0.65 | 0.95 | 0.94 | 0.74 | 0.83 | 0.80 |
| 0.70 | 0.94 | 0.96 | 0.69 | 0.80 | 0.78 |
| 0.75 | 0.94 | 0.98 | 0.65 | 0.78 | 0.77 |
| 0.80 | 0.92 | 0.98 | 0.57 | 0.72 | 0.71 |
| 0.85 | 0.91 | 0.99 | 0.48 | 0.65 | 0.66 |
| 0.90 | 0.89 | 1.00 | 0.37 | 0.54 | 0.57 |
| 0.95 | 0.86 | 0.99 | 0.20 | 0.34 | 0.41 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




