Submitted:
11 February 2026
Posted:
12 February 2026
You are already at the latest version
Abstract
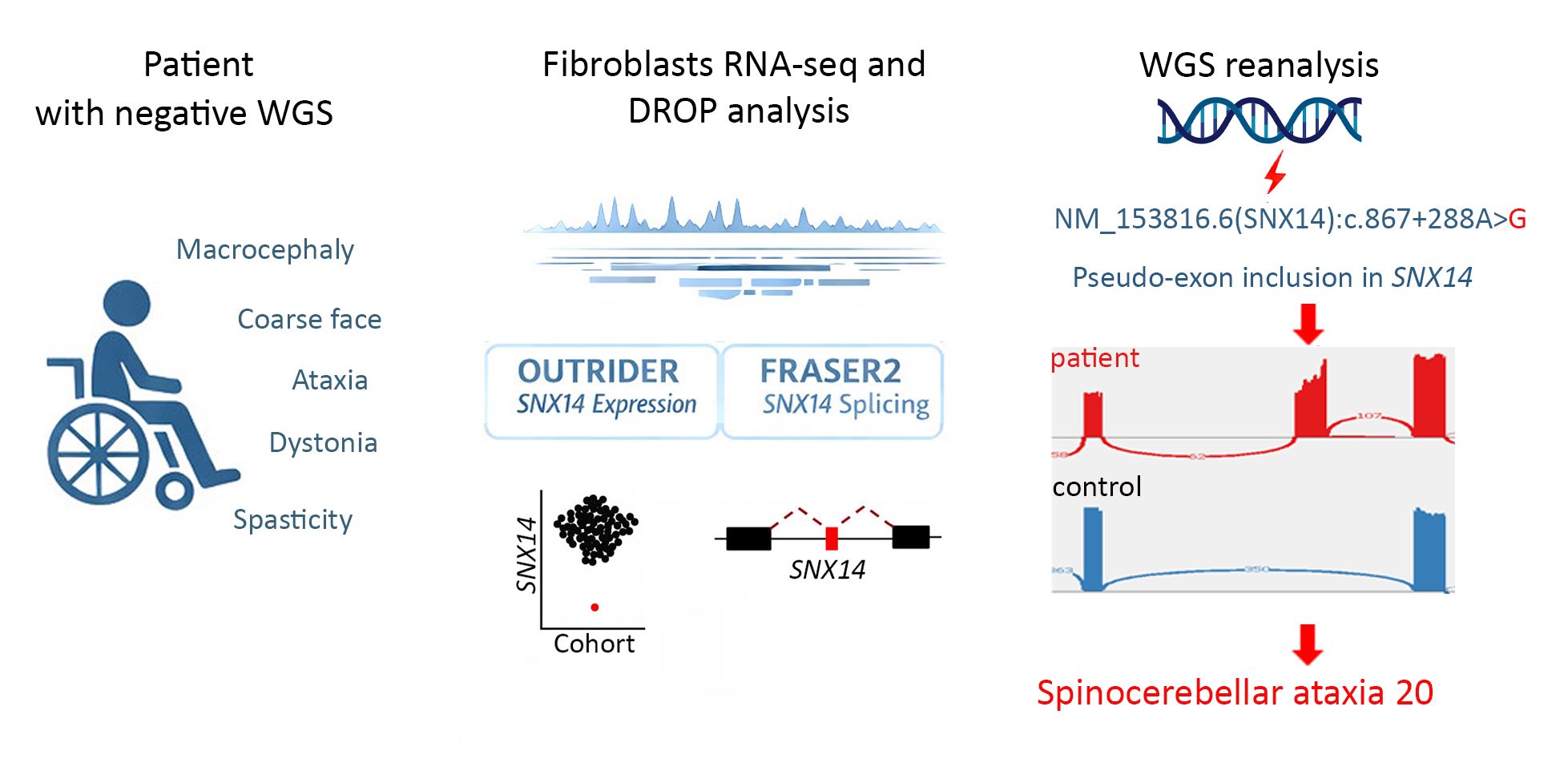
Keywords:
1. Introduction
2. Materials and Methods
2.1. Whole Genome Sequencing (WGS) and Data Analysis
2.2. RNA-seq and Data Analysis
3. Results
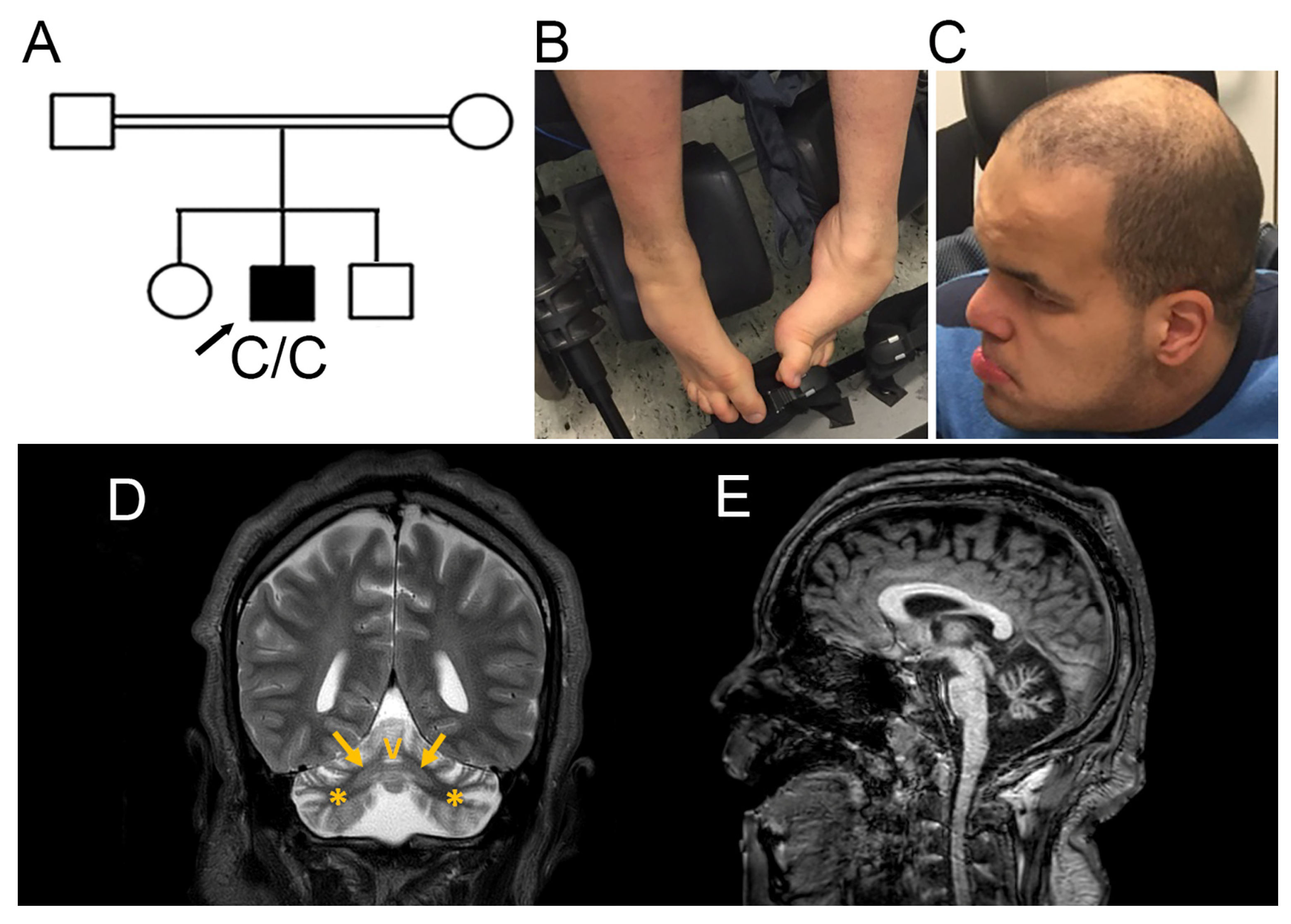
3.1. Clinical Presentation
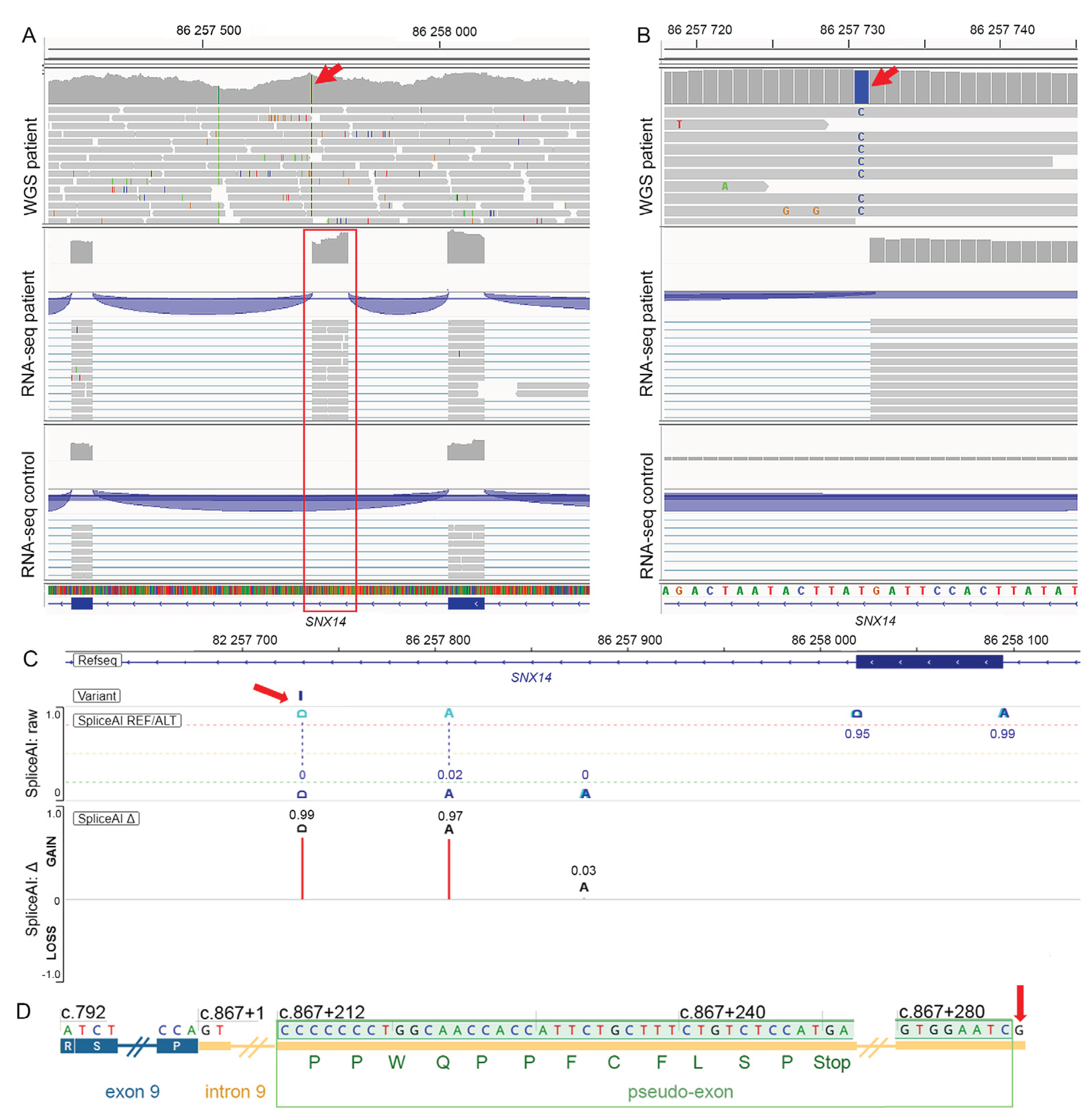
3.2. Variant Identification by DROP Pipeline on RNA-seq Data
3.3. Retrospective WGS Data Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Abbreviations
| ACMG/AMP | American College of Medical Genetics and Genomics/ Association for Molecular Pathology |
| CADD | Combined Annotation Dependent Depletion |
| DROP | Detection of RNA Outliers Pipeline |
| FDR | False discovery rate |
| IGV | Integrative Genomics Viewer |
| Indels | Small insertion/deletion |
| NAA | N-acetyl aspartate |
| NMD | Nonsense-mediated decay |
| RNA-seq | RNA-sequencing |
| SNVs | Single Nucleotide Variants |
| SNX14 | Sorting nexin 14 |
| SCAR20 | Spinocerebellar ataxia, autosomal recessive 20 |
| VUS | Variants of uncertain significance |
| WES | Whole exome sequencing |
| WGS | Whole genome sequencing |
References
- Pandey, R.; Brennan, N. F.; Trachana, K. A meta-analysis of diagnostic yield and clinical utility of genome and exome sequencing in pediatric rare and undiagnosed genetic diseases. Genet Med 2025, 27, pp. 101398. [Google Scholar] [CrossRef]
- Vaz-Drago, R.; Custodio, N.; Carmo-Fonseca, M. Deep intronic mutations and human disease. Hum Genet 2017, 136, 1093–1111. [Google Scholar] [CrossRef]
- Park, E.; Pan, Z.; Zhang, Z. The Expanding Landscape of Alternative Splicing Variation in Human Populations. Am J Hum Genet 2018, 102, 11–26. [Google Scholar] [CrossRef]
- Lord, J.; Baralle, D. Splicing in the Diagnosis of Rare Disease: Advances and Challenges. Front Genet 2021, 12, pp. 689892. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Macakova, K.; Sinson, J. C. Clinical validation of RNA sequencing for Mendelian disorder diagnostics. Am J Hum Genet 2025, 112, 779–792. [Google Scholar] [CrossRef] [PubMed]
- Kernohan, K. D.; Boycott, K. M. The expanding diagnostic toolbox for rare genetic diseases. Nat Rev Genet 2024, 25, 401–415. [Google Scholar] [CrossRef] [PubMed]
- Yepez, V. A.; Mertes, C.; Muller, M. F. Detection of aberrant gene expression events in RNA sequencing data. Nat Protoc 2021, 16, 1276–1296. [Google Scholar] [CrossRef]
- Brechtmann, F.; Mertes, C.; Matuseviciute, A. OUTRIDER: A Statistical Method for Detecting Aberrantly Expressed Genes in RNA Sequencing Data. Am J Hum Genet 2018, 103, 907–917. [Google Scholar] [CrossRef]
- Scheller, I. F.; Lutz, K.; Mertes, C. Improved detection of aberrant splicing with FRASER 2.0 and the intron Jaccard index. Am J Hum Genet 2023, 110, 2056–2067. [Google Scholar] [CrossRef]
- Neethiraj, R.; Wm, J.; Jemt, A. nf-core/raredisease: 2.2.0 - Dogmatix. 2024, pp. [Google Scholar]
- Vasimuddin, M.; Misra, S.; Li, H. Efficient Architecture-Aware Acceleration of BWA-MEM for Multicore Systems. 2019 IEEE International Parallel and Distributed Processing Symposium (IPDPS), 2019. [Google Scholar]
- Poplin, R.; Chang, P. C.; Alexander, D. A universal SNP and small-indel variant caller using deep neural networks. Nature Biotechnology 2018, 36, 983–987. [Google Scholar] [CrossRef]
- Yun, T.; Li, H.; Chang, P. C. Accurate, scalable cohort variant calls using DeepVariant and GLnexus. Bioinformatics 2021, 36, 5582–5589. [Google Scholar] [CrossRef]
- McLaren, W.; Gil, L.; Hunt, S. E. The Ensembl Variant Effect Predictor. Genome Biol 2016, 17, pp. 122. [Google Scholar] [CrossRef]
- Rentzsch, P.; Schubach, M.; Shendure, J. CADD-Splice-improving genome-wide variant effect prediction using deep learning-derived splice scores. Genome Med 2021, 13, pp. 31. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, B. S.; Layer, R. M.; Quinlan, A. R. Vcfanno: fast, flexible annotation of genetic variants. Genome Biol 2016, 17, pp. 118. [Google Scholar] [CrossRef] [PubMed]
- Danecek, P.; Bonfield, J. K.; Liddle, J. Twelve years of SAMtools and BCFtools. Gigascience 2021, 10, pp. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Francioli, L. C.; Goodrich, J. K. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 2024, 625, 92–100. [Google Scholar] [CrossRef]
- Landrum, M. J.; Lee, J. M.; Benson, M. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res 2016, 44, D862–868. [Google Scholar] [CrossRef]
- Jaganathan, K.; Kyriazopoulou Panagiotopoulou, S.; McRae, J. F. Predicting Splicing from Primary Sequence with Deep Learning. Cell 2019, 176, 535–548 e524. [Google Scholar] [CrossRef]
- Vigeland, M. D.; Gjotterud, K. S.; Selmer, K. K. FILTUS: a desktop GUI for fast and efficient detection of disease-causing variants, including a novel autozygosity detector. Bioinformatics 2016, 32, 1592–1594. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Thorvaldsdottir, H.; Robinson, J. T.; Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 2013, 14, 178–192. [Google Scholar] [CrossRef]
- Walker, L. C.; Hoya, M.; Wiggins, G. A. R. Using the ACMG/AMP framework to capture evidence related to predicted and observed impact on splicing: Recommendations from the ClinGen SVI Splicing Subgroup. Am J Hum Genet 2023, 110, 1046–1067. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015, 17, 405–424. [Google Scholar] [CrossRef]
- Akizu, N.; Cantagrel, V.; Zaki, M. S. Biallelic mutations in SNX14 cause a syndromic form of cerebellar atrophy and lysosome-autophagosome dysfunction. Nat Genet 2015, 47, 528–534. [Google Scholar] [CrossRef]
- Jazayeri, R.; Hu, H.; Fattahi, Z. Exome Sequencing and Linkage Analysis Identified Novel Candidate Genes in Recessive Intellectual Disability Associated with Ataxia. Arch Iran Med 2015, 18, 670–682. [Google Scholar]
- Karaca, E.; Harel, T.; Pehlivan, D. Genes that Affect Brain Structure and Function Identified by Rare Variant Analyses of Mendelian Neurologic Disease. Neuron 2015, 88, 499–513. [Google Scholar] [CrossRef] [PubMed]
- Maia, N.; Soares, G.; Silva, C. Two Compound Heterozygous Variants in SNX14 Cause Stereotypies and Dystonia in Autosomal Recessive Spinocerebellar Ataxia 20. Front Genet 2020, 11, pp. 1038. [Google Scholar] [CrossRef] [PubMed]
- Raslan, I. R.; Silva, T. Y. T.; Kok, F. Clinical and Genetic Characterization of a Cohort of Brazilian Patients With Congenital Ataxia. Neurol Genet 2024, 10, pp. e200153. [Google Scholar] [CrossRef] [PubMed]
- Sait, H.; Moirangthem, A.; Agrawal, V. Autosomal recessive spinocerebellar ataxia-20 due to a novel SNX14 variant in an Indian girl. Am J Med Genet A 2022, 188, 1909–1914. [Google Scholar] [CrossRef]
- Shao, Y.; Yang, S.; Li, J. Compound heterozygous mutation of the SNX14 gene causes autosomal recessive spinocerebellar ataxia 20. Front Genet 2024, 15, pp. 1379366. [Google Scholar] [CrossRef]
- Shukla, A.; Upadhyai, P.; Shah, J. Autosomal recessive spinocerebellar ataxia 20: Report of a new patient and review of literature. Eur J Med Genet 2017, 60, 118–123. [Google Scholar] [CrossRef]
- Sousa, S. B.; Ramos, F.; Garcia, P. Intellectual disability, coarse face, relative macrocephaly, and cerebellar hypotrophy in two sisters. Am J Med Genet A 2014, 164A, 10–14. [Google Scholar] [CrossRef]
- Thomas, A. C.; Williams, H.; Seto-Salvia, N. Mutations in SNX14 cause a distinctive autosomal-recessive cerebellar ataxia and intellectual disability syndrome. Am J Hum Genet 2014, 95, 611–621. [Google Scholar] [CrossRef]
- Trujillano, D.; Bertoli-Avella, A. M.; Kumar Kandaswamy, K. Clinical exome sequencing: results from 2819 samples reflecting 1000 families. Eur J Hum Genet 2017, 25, 176–182. [Google Scholar] [CrossRef]
- Al Shamsi, B.; Al Maimani, A.; Al Hanaie, M. Exploring the Genetic Variations Underlying SNX14-Linked Autosomal Recessive Spinocerebellar Ataxia Type 20: A Case Series of 17 Patients From a Single Center in the Omani Population and Review of Literature. Am J Med Genet A 2025, pp. e70001. [CrossRef] [PubMed]
- Levchenko, O.; Filatova, A.; Mishina, I. Homozygous deep intronic variant in SNX14 cause autosomal recessive Spinocerebellar ataxia 20: a case report. Front Genet 2023, 14, pp. 1197681. [Google Scholar] [CrossRef]
- Zhang, H.; Hong, Y.; Yang, W. SNX14 deficiency-induced defective axonal mitochondrial transport in Purkinje cells underlies cerebellar ataxia and can be reversed by valproate. Natl Sci Rev 2021, 8, pp. nwab024. [Google Scholar] [CrossRef] [PubMed]
- Bryant, D.; Liu, Y.; Datta, S. SNX14 mutations affect endoplasmic reticulum-associated neutral lipid metabolism in autosomal recessive spinocerebellar ataxia 20. Hum Mol Genet 2018, 27, 1927–1940. [Google Scholar] [CrossRef]
- Ebrahimi-Fakhari, D. Congenital Disorders of Autophagy: What a Pediatric Neurologist Should Know. Neuropediatrics 2018, 49, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Ebrahimi-Fakhari, D.; Saffari, A.; Wahlster, L. Congenital disorders of autophagy: an emerging novel class of inborn errors of neuro-metabolism. Brain 2016, 139, 317–337. [Google Scholar] [CrossRef] [PubMed]
- Russell, S. R.; Drack, A. V.; Cideciyan, A. V. Intravitreal antisense oligonucleotide sepofarsen in Leber congenital amaurosis type 10: a phase 1b/2 trial. Nat Med 2022, 28, 1014–1021. [Google Scholar] [CrossRef] [PubMed]
- Lauffer, M. C.; van Roon-Mom, W.; Aartsma-Rus, A. Possibilities and limitations of antisense oligonucleotide therapies for the treatment of monogenic disorders. Commun Med (Lond) 2024, 4, pp. 6. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



