Submitted:
11 February 2026
Posted:
12 February 2026
You are already at the latest version
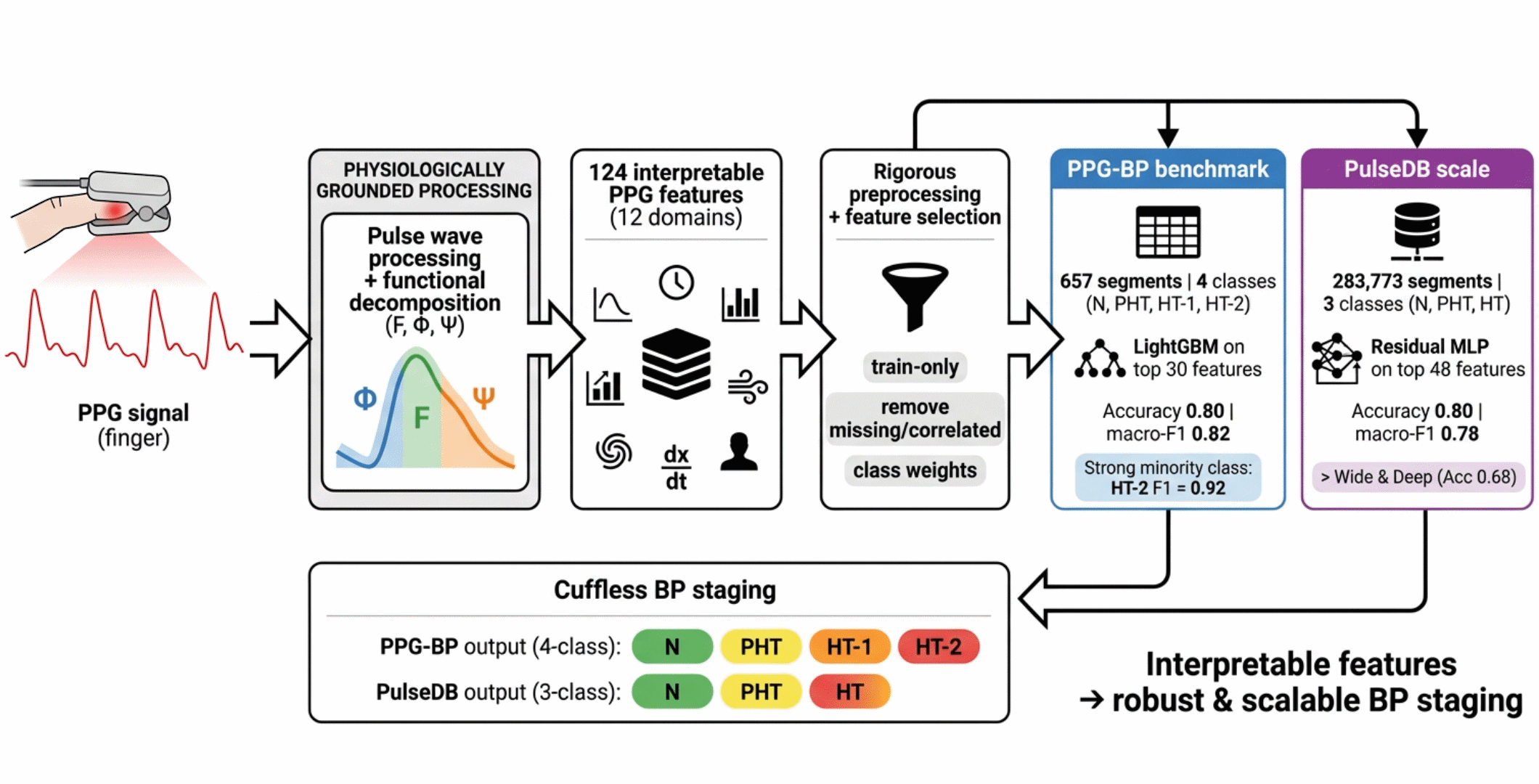
Abstract
Keywords:
1. Introduction
2. Materials and Methods
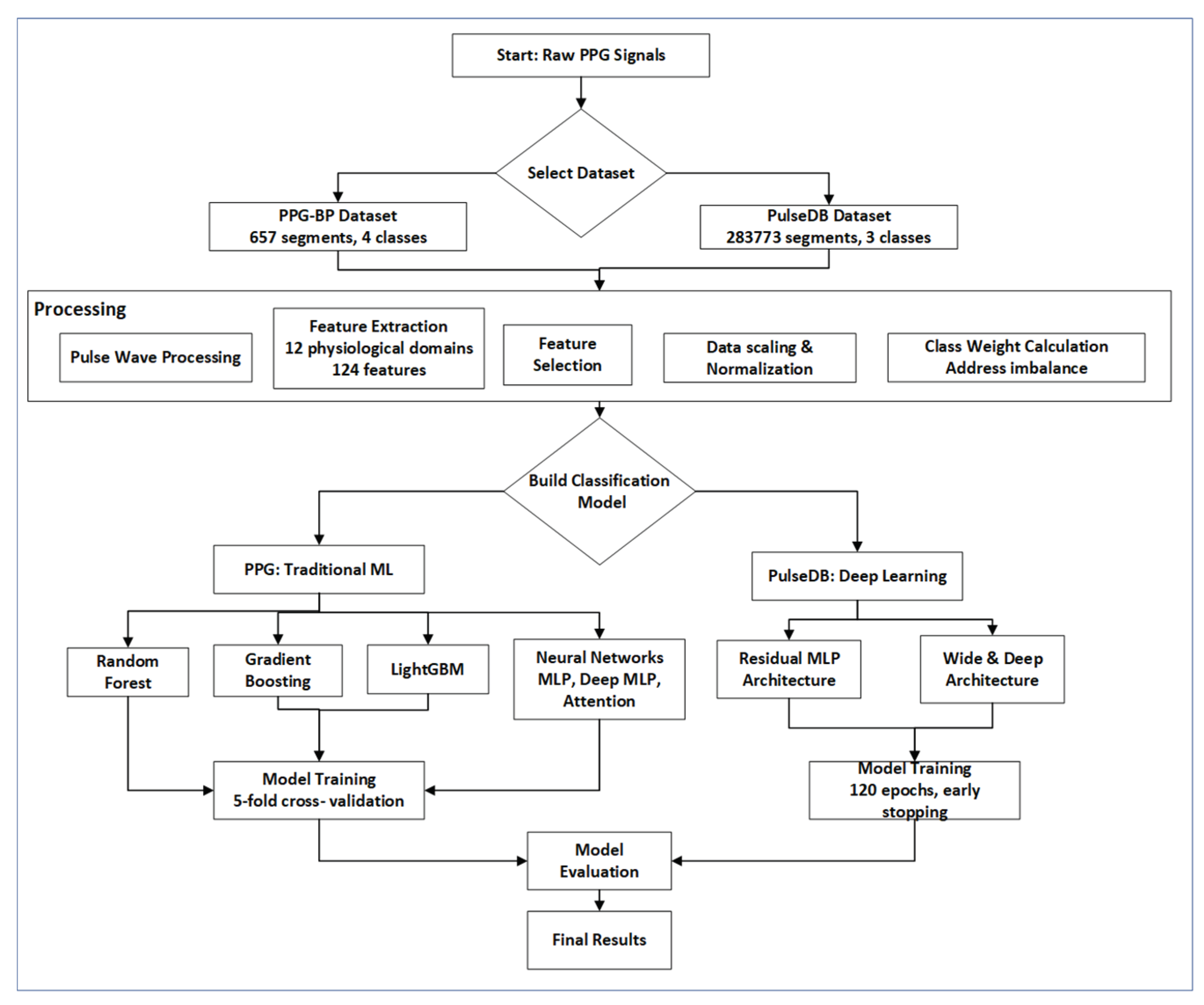
2.1. Datasets
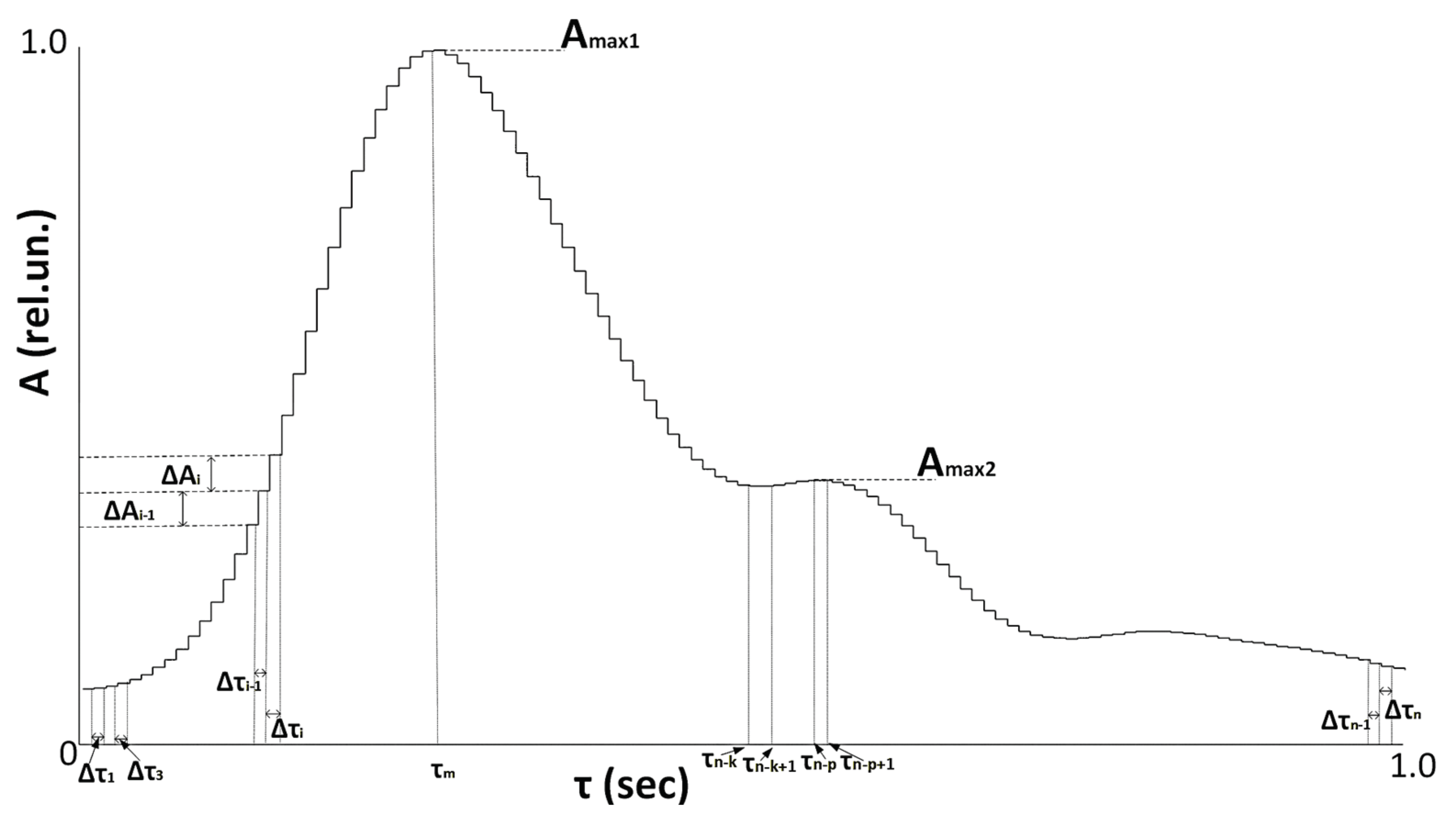
2.2. Pulse Wave Processing
2.3. Feature Extraction and Selection
- Basic morphology (26 features): These features capture fundamental PPG waveform shape characteristics related to cardiac ejection and vascular properties.
- Function-based (15 features): These features are derived from functional decomposition using basis functions (, , ) that represent distinct physiological components.
- Temporal-domain analysis (4 features): These features characterize timing relationships within and between cardiac cycles.
- Beat-to-beat analysis (7 features): These features quantify the stability and consistency of waveform characteristics across consecutive pulses.
- Velocity and acceleration analysis (11 features): These features analyze the first, second, and third derivatives of the PPG signal to characterize hemodynamic dynamics.
- Frequency-domain analysis (8 features): These features decompose the PPG signal into frequency components to capture periodic patterns and harmonics.
- Nonlinear dynamics (8 features): These features quantify irregularity, unpredictability, and self-similarity in PPG signals.
- Inter-beat comparison (7 features): These features quantify beat-to-beat similarity and variation in pulse-wave shape.
- Vascular compliance (7 features): These features estimate arterial stiffness using pulse-wave-velocity proxies and reflection characteristics.
- Composite clinical indices (5 features): These features combine multiple signal characteristics into clinically interpretable indices.
- Engineered features (22 features): These features combine signal-derived characteristics with demographic variables using domain knowledge.
- Demographics and heart rate (4 features): These features include age, sex, BMI, and heart rate, which provide context for interpretation and normalization.
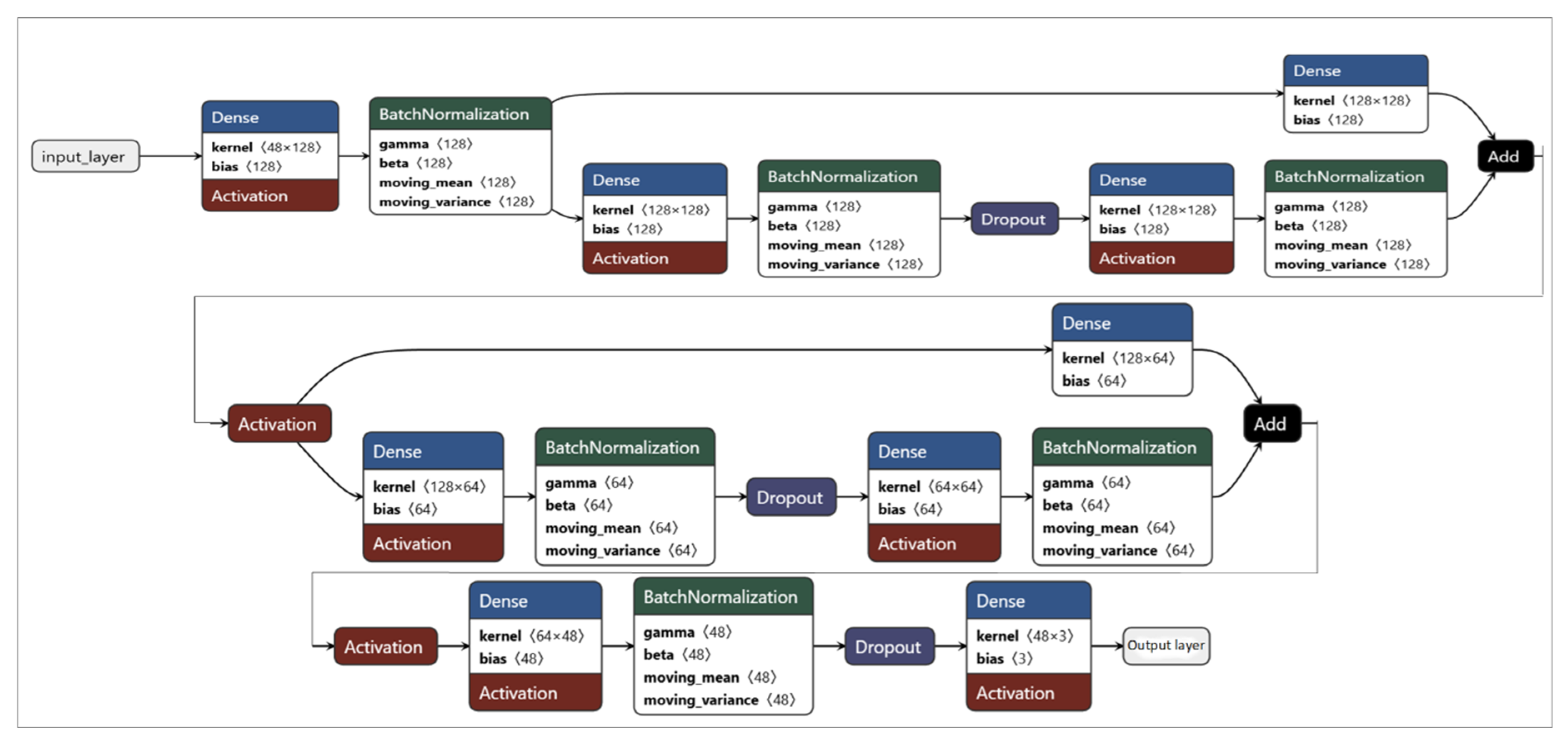
2.4. Classification of BP Stages
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| PPG | Photoplethysmography |
| SBP | Systolic blood pressure |
| DBP | Diastolic blood pressure |
| CNN | Convolutional neural network |
| GRU | Gated recurrent unit |
| SVM | Support vector machine |
| LDA | Linear discriminant analysis |
| LSVM | Linear Support Vector Machine |
| LSTM | Long Short-Term Memory |
| BMI | Body mass index |
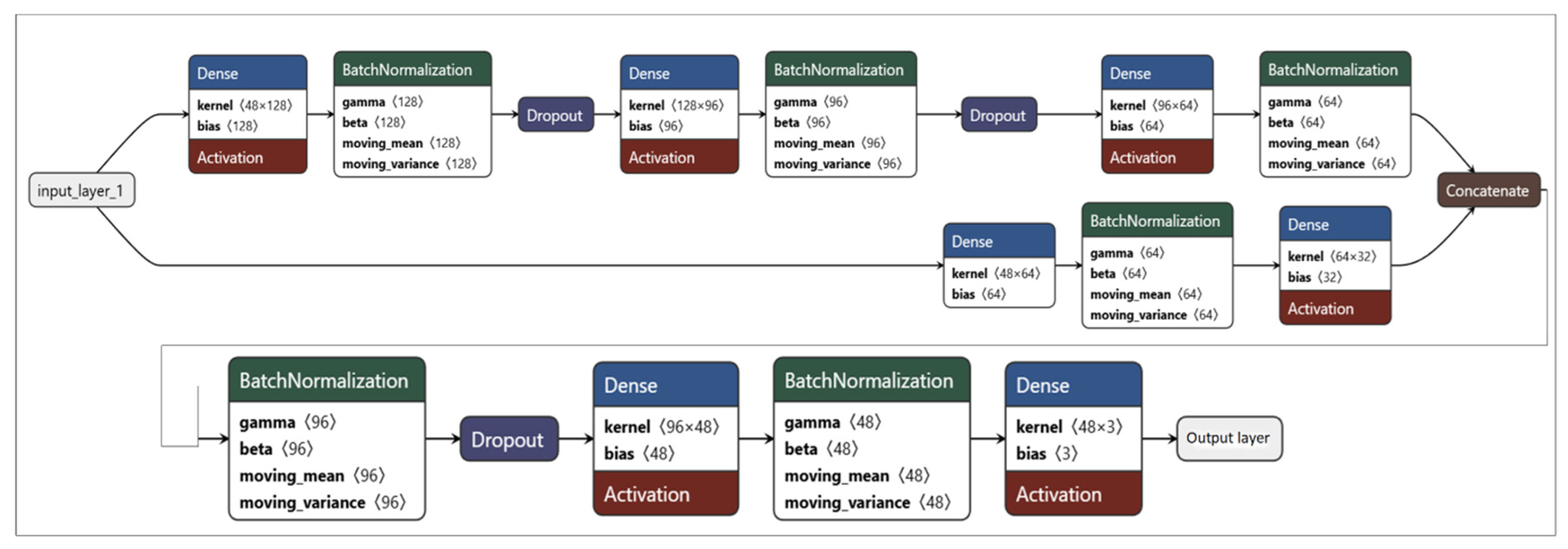
| MLP | Multi-Layer Perceptron |
| PHT | Prehypertension |
| HT | Hypertension |
References
- World Health Organization. Cardiovascular Diseases (CVDs). Available online: https://www.who.int/health-topics/cardiovascular-diseases#tab=tab_1 (accessed on February 2026).
- World Health Organization. Hypertension. Available online: https://www.who.int/news-room/fact-sheets/detail/hypertension (accessed on 1 February 2026).
- Haugg, F.; Elgendi, M.; Menon, C. Assessment of blood pressure using only a smartphone and machine learning techniques: a systematic review. Front. Cardiovasc. Med. 2022, 9, 894224. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Yang, C.; Yang, S.; Kwong, C.F.; et al. Photoplethysmography-based non-invasive blood pressure monitoring via ensemble model and imbalanced dataset processing. Phys. Eng. Sci. Med. 2024, 47, 1307–1321. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Yin, S.; Zhang, X.; et al. Blood pressure stratification using photoplethysmography and light gradient boosting machine. Front. Physiol. 2023, 14, 1072273. [Google Scholar] [CrossRef] [PubMed]
- Allen, J. Photoplethysmography and its application in clinical physiological measurement. Physiol. Meas. 2007, 28, R1–R39. [Google Scholar] [CrossRef]
- Elgendi, M. On the analysis of fingertip photoplethysmogram signals. Curr. Cardiol. Rev. 2012, 8, 14–25. [Google Scholar] [CrossRef]
- Charlton, P.H.; Paliakaitė, B.; Pilt, K.; et al. Assessing hemodynamics from the photoplethysmogram to gain insights into vascular age: a review from VascAgeNet. Am. J. Physiol. Heart Circ. Physiol. 2022, 322, H493–H522. [Google Scholar] [CrossRef]
- Mejía-Mejía, E.; Allen, J.; Budidha, K.; El Hajj, C.; Kyriacou, P.; Charlton, P. Photoplethysmography signal processing and synthesis. In Photoplethysmography; Elsevier, 2022; pp. 69–146. [Google Scholar] [CrossRef]
- Elgendi, M.; Jost, E.; Alian, A.; et al. Photoplethysmography Features Correlated with Blood Pressure Changes. Diagnostics 2024, 14, 2309. [Google Scholar] [CrossRef]
- Yen, C.-T.; Chang, S.-N.; Liao, C.-H. Deep learning algorithm evaluation of hypertension classification in less photoplethysmography signals conditions. Meas. Control 2021, 54, 439–445. [Google Scholar] [CrossRef]
- Nasir, N.; Sameer, M.; Barneih, F.; Alshaltone, O. Deep Learning Classification of Photoplethysmogram Signal for Hypertension Levels. arXiv 2024, arXiv:2405.14556. [Google Scholar] [CrossRef]
- Tjahjadi, H.; Ramli, K. Noninvasive Blood Pressure Classification Based on Photoplethysmography Using K-Nearest Neighbors Algorithm: A Feasibility Study. Information 2020, 11, 93. [Google Scholar] [CrossRef]
- Abdullah, S.; Kristoffersson, A. Machine learning approaches for cardiovascular hypertension stage estimation using photoplethysmography and clinical features. Front. Cardiovasc. Med. 2023, 10, 1285066. [Google Scholar] [CrossRef] [PubMed]
- Hassija, V.; Chamola, V.; Mahapatra, A.; et al. Interpreting Black-Box Models: A Review on Explainable Artificial Intelligence. Cogn. Comput. 2024, 16, 45–74. [Google Scholar] [CrossRef]
- Wu, J.; Liang, H.; Ding, C.; Huang, X.; Huang, J.; Peng, Q. Improving the Accuracy in Classification of Blood Pressure from Photoplethysmography Using Continuous Wavelet Transform and Deep Learning. Int. J. Hypertens. 2021, 2021, 9938584. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Ríos, E.; Montesinos, L.; Alfaro-Ponce, M. A machine learning approach for hypertension detection based on photoplethysmography and clinical data. Comput. Biol. Med. 2022, 145, 105479. [Google Scholar] [CrossRef]
- Fahoum, A.A.; Al Omari, A.; Al Omari, G.; Zyout, A. Development of a novel light-sensitive PPG model using PPG scalograms and PPG-NET learning for non-invasive hypertension monitoring. Heliyon 2024, 10, e39745. [Google Scholar] [CrossRef]
- Promi, R.T.H.; Nazri, R.A.; Salim, M.S.; Raju, S.M.T.U. A Deep Learning Approach for Non-Invasive Hypertension Classification from PPG Signal. In *Proceedings of the 2023 International Conference on Next-Generation Computing, IoT and Machine Learning (NCIM), Gazipur, Bangladesh, 24–25 November 2023; pp. 1–5. [Google Scholar] [CrossRef]
- Frederick, G.; Yaswant, T.; Brintha, A. PPG Signals for Hypertension Diagnosis: A Novel Method using Deep Learning Models. arXiv 2023, arXiv:2304.06952. [Google Scholar] [CrossRef]
- Nour, M.; Polat, K. Automatic Classification of Hypertension Types Based on Personal Features by Machine Learning Algorithms. Math. Probl. Eng. 2020, 28, 1–13. [Google Scholar] [CrossRef]
- Pankaj, P.; Kumar, A.; Kumar, M.; et al. Optimized deep neural network models for blood pressure classification using Fourier analysis-based time–frequency spectrogram of photoplethysmography signal. Biomed. Eng. Lett. 2023, 13, 739–750. [Google Scholar] [CrossRef]
- Tanc, Y.H.; Ozturk, M. Hypertension Classification Using PPG Signals. In Proceedings of the 2022 Medical Technologies Congress (TIPTEKNO), Antalya, Turkey, 31 October–2 November 2022; pp. 1–4. [Google Scholar] [CrossRef]
- Liang, Y.; Chen, Z.; Liu, G.; Elgendi, M. A new, short-recorded photoplethysmogram dataset for blood pressure monitoring in China. Sci. Data 2018, 5, 180020. [Google Scholar] [CrossRef]
- Wang, W.; Mohseni, P.; Kilgore, K.L.; Najafizadeh, L. PulseDB: A large, cleaned dataset based on MIMIC-III and VitalDB for benchmarking cuff-less blood pressure estimation methods. Front. Digit. Health 2023, 4, 1090854. [Google Scholar] [CrossRef]
- Msokar, S.; Davydov, R.V.; Mazing, M.S.; Isakova, D.D. New opportunities for studying the oxygen saturation of blood hemoglobin in capillaries and tissues. St. Petersburg Polytech. Univ. J. Phys. Math. 2023, 16, 328–332. [Google Scholar] [CrossRef]
- Davydov, R.; Zaitceva, A.; Davydov, V.; Isakova, D.; Mazing, M. New Methodology of Human Health Express Diagnostics Based on Pulse Wave Measurements and Occlusion Test. J. Pers. Med. 2023, 13, 443. [Google Scholar] [CrossRef]
- Su, Y.; Li, Y.; Zhang, S.; Wang, H. Linear and Nonlinear Analysis of Photoplethysmogram Signals and Electrodermal Activity to Recognize Three Different Levels of Human Stress. Int. J. Adv. Comput. Sci. Appl. 2023, 14. [Google Scholar] [CrossRef]
- Dehkordi, P.; Garde, A.; Karlen, W.; Petersen, C.L.; Ansermino, J.M.; Dumont, G.A. Detrended fluctuation analysis of photoplethysmogram pulse rate intervals in sleep disordered breathing. In Proceedings of the 2014 IEEE Healthcare Innovation Conference (HIC), Seattle, WA, USA, 8–10 October 2014; pp. 323–326. [Google Scholar] [CrossRef]
- van Roon, A.M.; Span, M.M.; Lefrandt, J.D.; Riese, H. Overview of Mathematical Relations Between Poincaré Plot Measures and Time and Frequency Domain Measures of Heart Rate Variability. Entropy 2025, 27, 861. [Google Scholar] [CrossRef]
- Alotaiby, T.N.; Alshebeili, S.A.; Alotibi, G.; Alotaibi, G.N. Recurrence Quantification Analysis for PPG/ECG-Based Subject Authentication. In Proceedings of the 2022 4th International Conference on Data Intelligence and Security (ICDIS), Shenzhen, China, 24–26 August 2022; pp. 288–291. [Google Scholar] [CrossRef]
- Chen, D.; Liu, Z.; Ma, X.; Hua, D. Selecting genes by test statistics. J. Biomed. Biotechnol. 2005, 132–138. [Google Scholar] [CrossRef]
- Msokar, S. Classification-BP-stages. GitHub. 2026. Available online: https://github.com/souhair/Classification-BP-stages.
- Chobanian, A.V.; Bakris, G.L.; Black, H.R.; Cushman, W.C.; Green, L.A.; Izzo, J.L.; Jones, D.W.; Materson, B.J.; Oparil, S.; Wright, J.T.; et al. The seventh report of the joint national committee on prevention, detection, evaluation, and treatment of high blood pressure; the JNC 7 report. JAMA 2003, 289, 2560–2572. [Google Scholar] [CrossRef]
| Study | Study focus | Dataset | Methods | Results | Limitations |
|---|---|---|---|---|---|
| Martínez-Ríos et al. (2022) [17] | Hypertension detection using raw PPG and clinical data | PPG-BP | (SVM) with early/late fusion | 71.42% accuracy, 0.76 F1-score for prehypertension | Binary classification (Normal vs prehypertension) |
| Al Fahoum et al. (2024) [18] | Lightweight model for wearable deployment using raw PPG | PPG-BP | Custom PPG-NET (depthwise separable convolutions) | PPG-NET model achieving 100% across all metrics | Implausible performance, small dataset, requires validation, potential data leakage |
| Abdullah & Kristoffersson (2023) [14] | Hypertension staging with APG features (using raw PPG) | PPG-BP | Decision Tree, Linear Discriminant Analysis (LDA), (Linear Support Vector Machine) LSVM | DT model achieved training accuracy of 100% during cross-validation and accuracy of 96.87% on the test dataset | The authors used systolic BP (SBP) as an input feature, which introduces data leakage and likely overfitting. |
| Promi et al. (2023) [19] | Hybrid spatiotemporal model (using raw PPG and clinical data) | PPG-BP | CNN-GRU hybrid with 10-fold CV | 85.00% accuracy, 0.8608 F1-score | The paper does not explore the individual contributions of the CNN and GRU components, making it unclear whether both are necessary for performance gains. Although 10-fold cross-validation is used, the training and validation loss/accuracy curves (Figure 4 [19]) suggest some overfitting, as validation loss plateaus early while training loss continues to decrease. |
| Yen et al. (2021) [11] | Hybrid models for imbalanced data (using raw PPG) | PPG-BP | ResNetCNN-BiLSTM, Xception-BiLSTM | 76% accuracy 0.45 recall, and a precision of 0.48 for Xception BiLSTM | Poor precision, recall and F1-score, especially for minority classes. |
| Nasir et al. (2024) [12] | Ensemble and stacked models | PPG-BP | CNN, LSTM, BiLSTM with stacking | 71.9% accuracy (LSTM-CNN-RF), 100% with ensembles | Binary classification, potential data leakage, F1-score: 0.66 and recall: 0.60 for LSTM-CNN+SVM |
| Frederick et al. (2023) [20] | Pooling layer optimization (raw PPG) | PPG-BP | AlexNet, ResNet-50, VGG-16, AvgPool_VGG-16 | 80% accuracy with AvgPool_VGG-16 | Severe class imbalance, poor minority class performance (F1-score: 0.75 for HT-1 and 0.10 for HT-2) |
| Tjahjadi & Ramli (2020) [13] | Raw signal classification | PPG-BP | K-Nearest Neighbors (KNN) | 86.7% accuracy, 100% F1 for some binary tasks | Binary classification. Data leakage risk, overfitting with K=1, excessive accuracy for binary tasks. Requires validation. |
| Nour & Polat (2020) [21] | Hypertension type classification | PPG-BP | C4.5 Decision Tree, Random Forest, LDA, LSVM | 99.5% accuracy with tree-based models | Authors used SBP & DBP as input features of DL models and that led to data leakage. |
| Pankaj et al. (2023) [22] | BP classification using Fourier-based spectrograms (PPG raw) | MIMIC-III + Figshare databases | DenseNet-201, AlexNet, GoogLeNet (transfer learning) | 96.51% accuracy with DenseNet-201 across 3 classes of BP stages | High computational cost |
| Wu et al. (2021) [16] | CWT optimization for BP classification (PPG raw) | MIMIC-III (311,000 segments) | Custom PPG-NET (depthwise separable convolutions) | 90% accuracy with cgau1 wavelet, 2.4s segments | Binary classification, lacks clinical granularity |
| Hu et al. (2023) [5] | Automated feature extraction with LightGBM | MIMIC-III (121 records) | LightGBM with Optuna optimization | F1-scores: 0.90-0.97 across tasks | Binary classification. The feature set contains 794 features, which exceeds the number of training records. |
| Tanc & Ozturk (2022) [23] | SST-based classification | MIMIC-II (150 records) | GoogLeNet (transfer learning) | 0.968 F1-score | Binary classification oversimplification, potential data leakage. |
| Systolic Blood Pressure (mmHg) | Diastolic Blood Pressure (mmHg) | |
|---|---|---|
| Normal (N) | < 120, and | < 80 |
| Prehypertension (PHT) | 120-139, or | 80-89 |
| Hypertension Stage 1 (HT-1) | 140-159, or | 90-99 |
| Hypertension Stage 2 (HT-2) | ≥160, or | ≥100 |
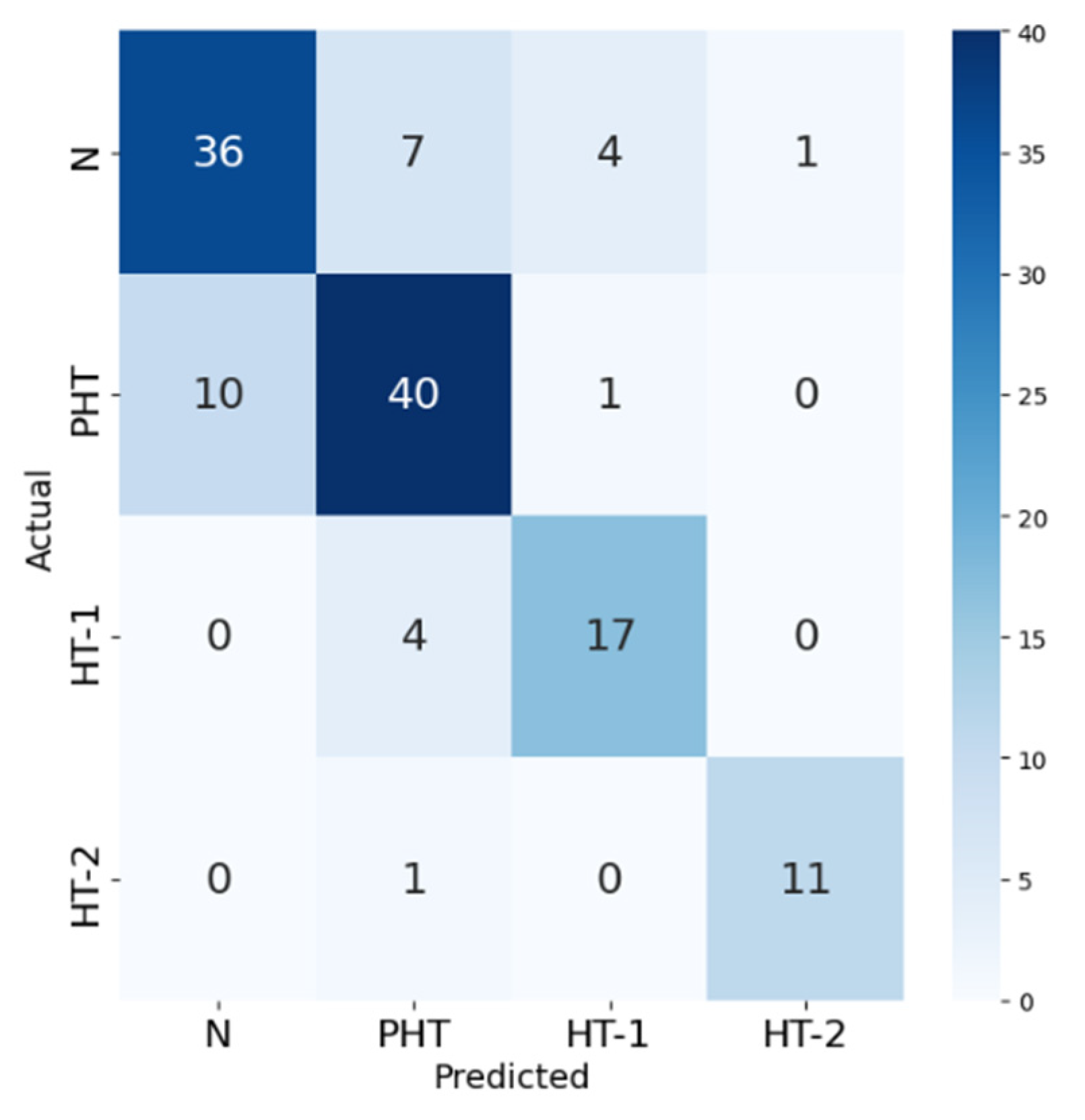
| precision | recall | F1-score | support | |
|---|---|---|---|---|
| Normal (N) | 0.78 | 0.75 | 0.77 | 48 |
| Prehypertension (PHT) | 0.77 | 0.78 | 0.78 | 51 |
| Hypertension Stage 1 (HT-1) | 0.77 | 0.81 | 0.79 | 21 |
| Hypertension Stage 2 (HT-2) | 0.92 | 0.92 | 0.92 | 12 |
| Accuracy | 0.79 | 132 | ||
| Macro avg | 0.81 | 0.82 | 0.81 | 132 |
| Weighted avg | 0.79 | 0.79 | 0.79 | 132 |
| Models | Macro-F1 |
|---|---|
| LightGBM | 0.813 |
| Gradient Boosting | 0.766 |
| Ensemble model1 | 0.751 |
| Random Forest | 0.723 |
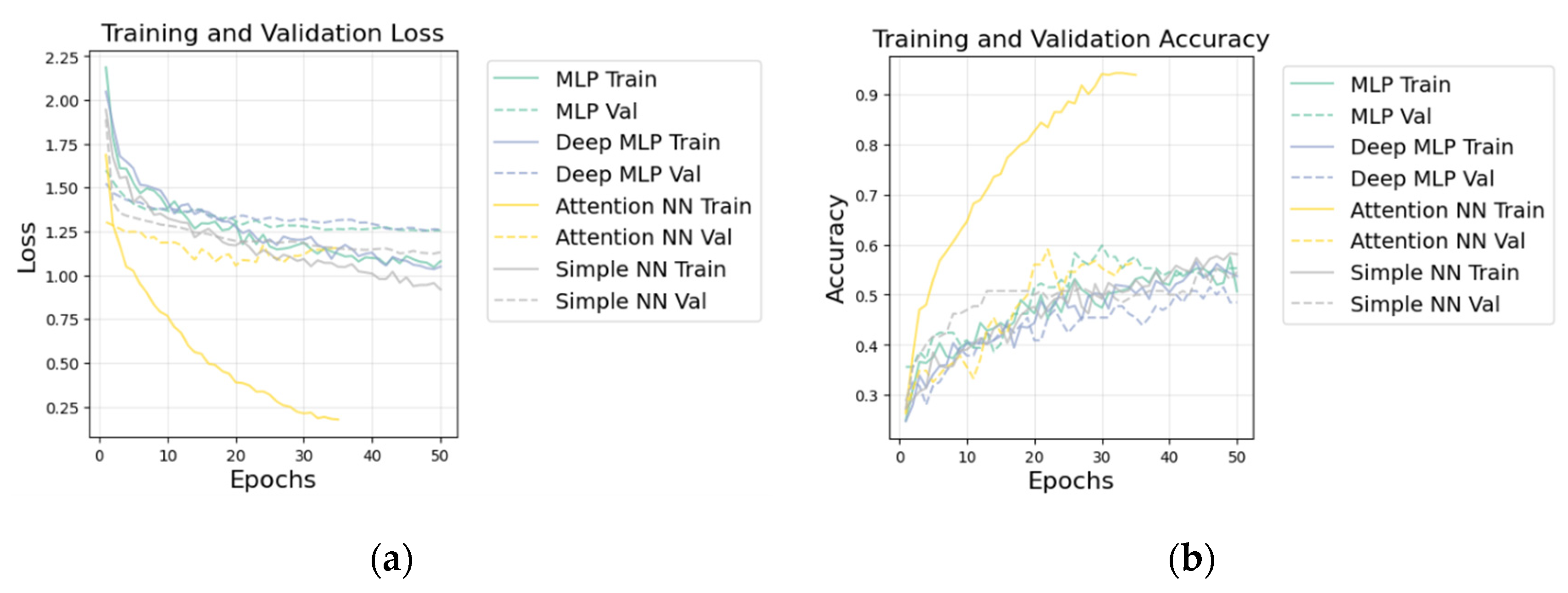
| Deep MLP | 0.563 |
| Simple NN | 0.528 |
| MLP | 0.462 |
| Attention NN | 0.324 |
| LightGBM model | F1-score |
|---|---|
| Normal (N) | 0.766 |
| Prehypertension (PHT) | 0.777 |
| Hypertension Stage 1 (HT-1) | 0.791 |
| Hypertension Stage 2 (HT-2) | 0.917 |
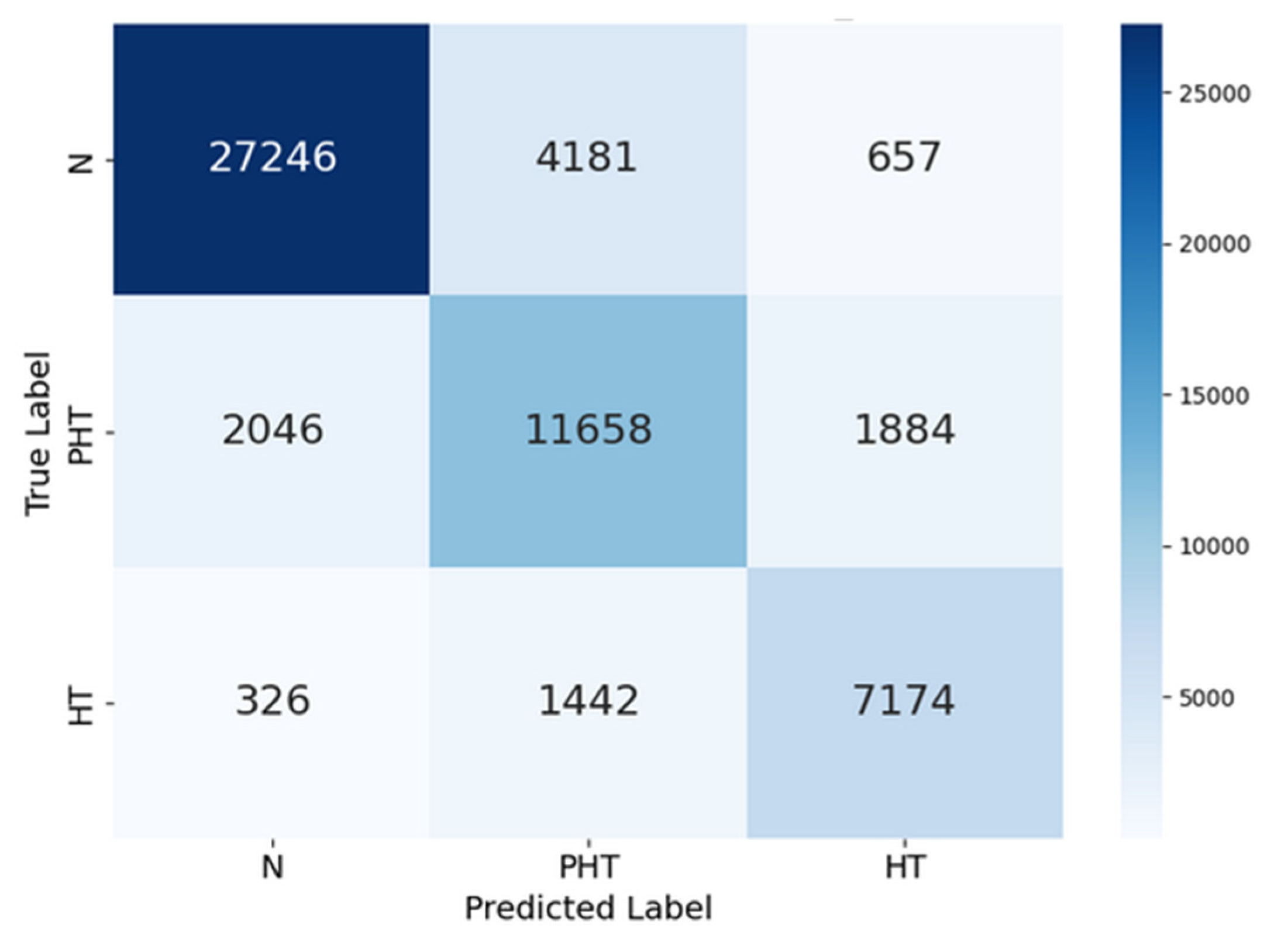
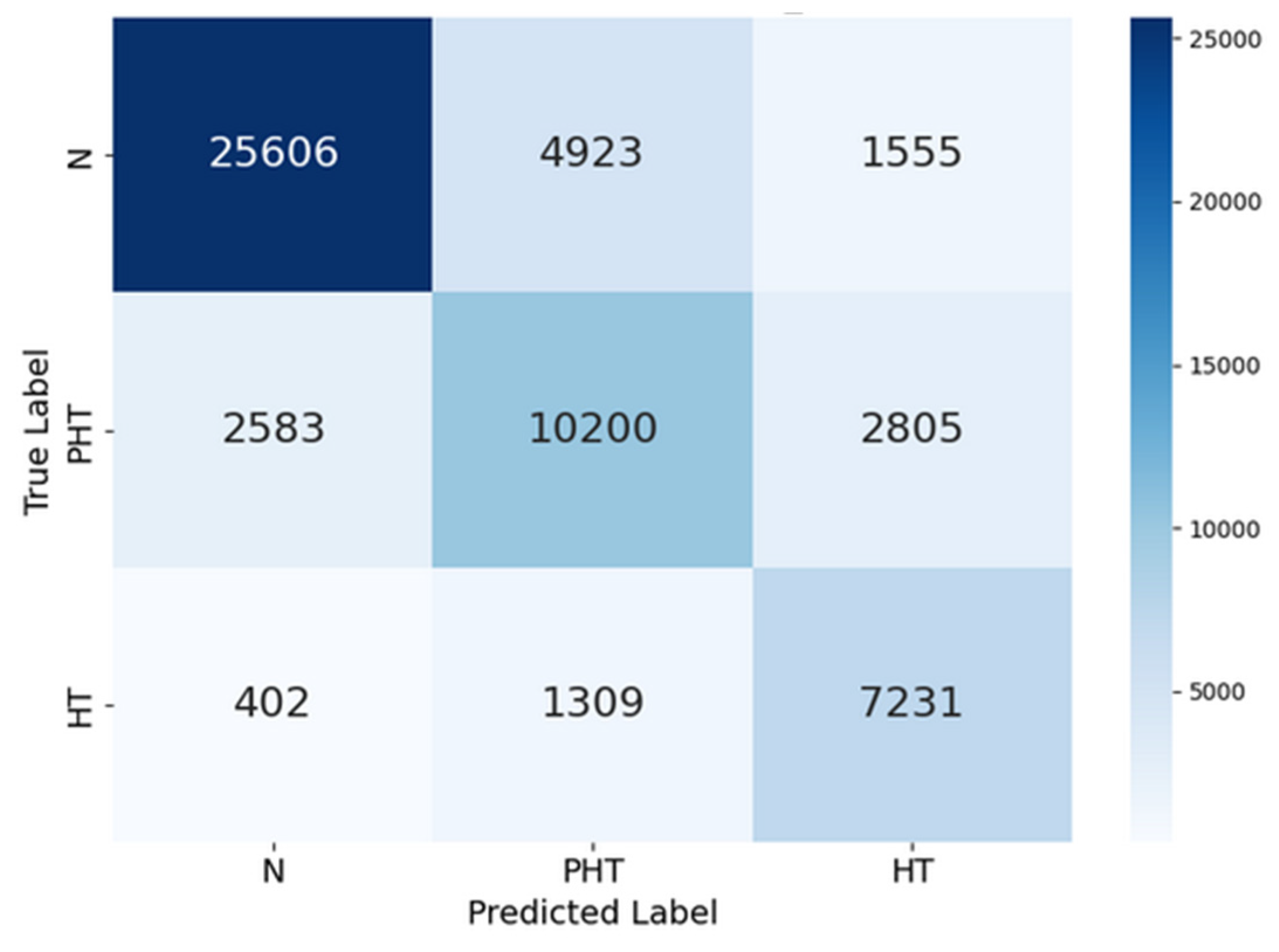
| precision | recall | F1-score | support | |
|---|---|---|---|---|
| Normal (N) | 0.92 | 0.85 | 0.88 | 32084 |
| Prehypertension (PHT) | 0.67 | 0.75 | 0.71 | 15588 |
| Hypertension (HT) | 0.74 | 0.80 | 0.77 | 8942 |
| Accuracy | 0.81 | 56614 | ||
| Macro avg | 0.78 | 0.80 | 0.79 | 56614 |
| Weighted avg | 0.82 | 0.81 | 0.82 | 56614 |
| precision | recall | F1-score | support | |
|---|---|---|---|---|
| Normal (N) | 0.90 | 0.80 | 0.84 | 32084 |
| Prehypertension (PHT) | 0.62 | 0.65 | 0.64 | 15588 |
| Hypertension (HT) | 0.62 | 0.81 | 0.70 | 8942 |
| Accuracy | 0.76 | 56614 | ||
| Macro avg | 0.71 | 0.75 | 0.73 | 56614 |
| Weighted avg | 0.78 | 0.76 | 0.76 | 56614 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




