1. Introduction
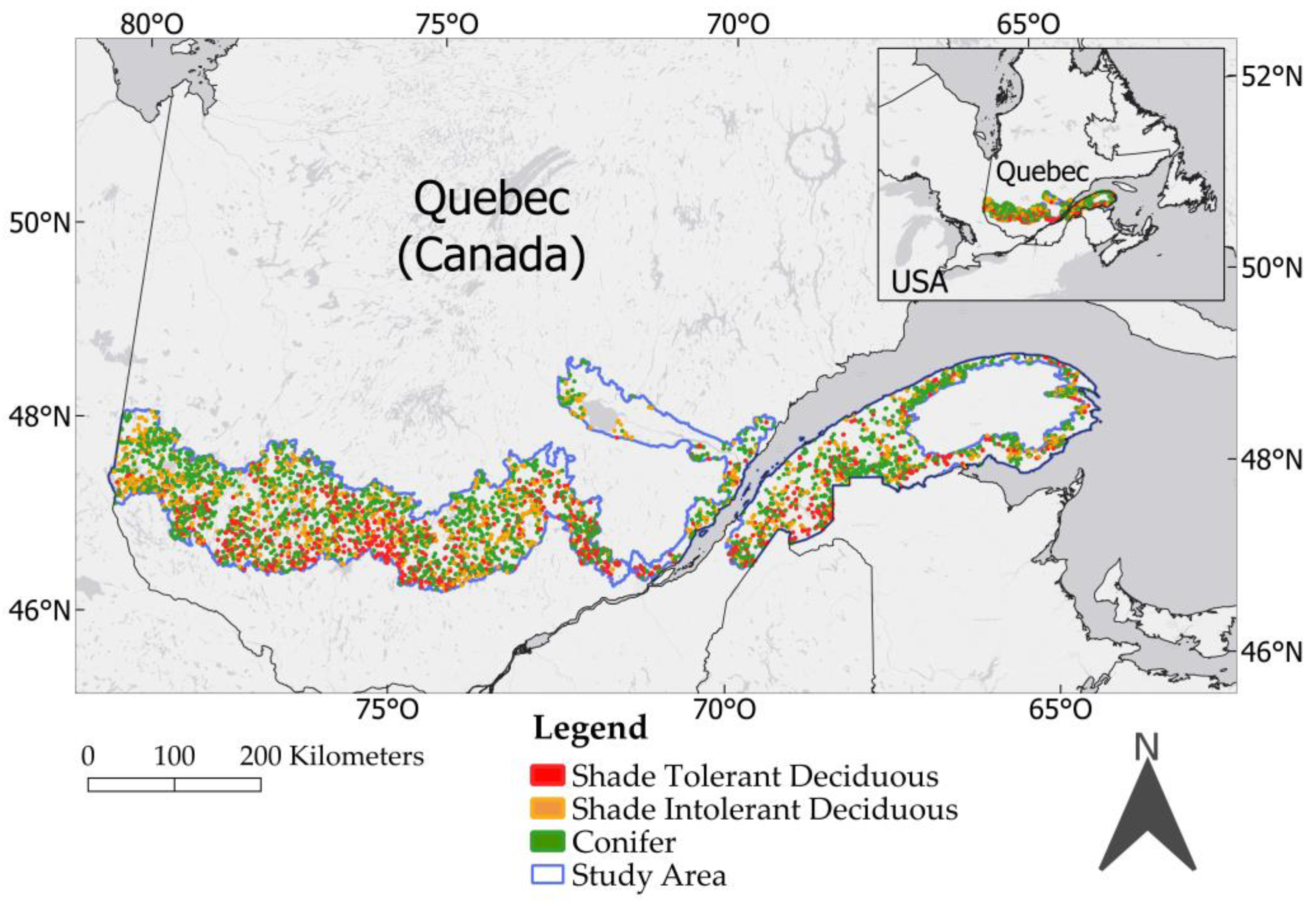
The forested area of Quebec covers nearly 905,800 km², which is more than the four countries with the most forest in Europe combined, excluding Russia (Sweden, Finland, Spain, France) [
1,
2]. In order to achieve sustainable forest management and effective operational planning in this vast territory, it is essential to have a reliable, complete and sufficiently rapid inventory technique for up-to-date monitoring. Stand basal area (BA) is an important variable for evaluating forest productivity [
3]. It represents the area covered by trees at breast height per hectare. It is highly correlated with volume and growth of forest stands [
4]. The classical method for determining stand BA uses ground sampling data and applies linear/nonlinear regressions models [
5,
6,
7,
8] or simultaneous equation methods [
9,
10] to extrapolate it to the entire stand. These approaches are becoming less common with the potential of new automated survey methods using technologies such as LiDAR (light detection and ranging) and photogrammetry [
11].
The use of airborne laser scanning (ALS) for large-scale forest inventory has been used for several years and has shown that it can be the primary tool for evaluating stand dendrometric metrics [
12]. In Québec (Canada), the prediction of forest metrics, including BA, is done using the k-nearest neighbors method [
13] with data from an airborne LiDAR when available. This approach has limitations as it relies on ground-based data and photo-interpretation to generate and update its results. Annually, thousands of temporary and permanent samples are collected until the entire province is covered, temporary samples are taken once, while permanent samples are revisited to track changes over the years. It takes about 10 years to update the full inventory [
14] which makes it expensive and time-consuming to collect, it also requires significant labor in a labor shortage context [
15,
16]. From 2022, a tool called the LiDAR Dendrometric Map (LDM) was developed to predict forest metrics, derived from density and height of captured pulses combined with the effective ecoforest map data and a relative density index [
17]. This area-based approach (ABA) map is currently available only for bioclimatic domains dominated by conifers in northern Quebec. This approach has shown great potential since, without needing new costly ground sample plot acquisition programs, it’s possible to map forest metrics using ALS with a good level of accuracy in the boreal forest of Quebec, where the main species are coniferous or shade-intolerant deciduous [
17,
18].
The province now wants to extend the map to southern regions where shade-tolerant species are located. Before extending the ALS-based tool to southern forests, an essential step is to validate whether the tool will achieve the same accuracy in predicting basal area as it does in boreal forests. While ALS scanning approaches have demonstrated their efficiency in boreal forest with coniferous and shade-intolerant deciduous species [
19,
20,
21,
22], this has not been the case for shade-tolerant deciduous forests [
23,
24,
25]. Since ALS mainly gives information about crown heights and the relation between crown height and mean diameter at breast height (DBH) is weak in these forests, the accuracy tends to be lower [
23,
24]. To enhance forest metrics estimation from LiDAR, incorporating more canopy characteristics beyond vertical structure is necessary [
24,
26]. For the LDM, a relative density index (RDI) was generated using inventory data to predict crown closure, thereby strengthening the model. Data from the ecoforest map were also utilized, including dominant species, the proportion of merchantable conifer (FSPL), the standard version of the map, and two ecological land classifications [
17]. Given the unique approach used by the LDM for generating forest metrics, it is justified to test its effectiveness in shade-tolerant deciduous stands.
It is known that height-to-diameter relationships are related to growing environment and stand conditions [
27,
28], and the growth of shade-tolerant species is more influenced by abiotic factors than the growth of intolerant species [
29,
30]. Targeting improvements, adding data from outside the ecoforest map such as climate data, latitude and longitude to complement independent variables used by the LDM has not been tested. A tool called BIOSIM has been developed in Canada based on weather station data and provides a range of geospatial climate metrics that can be merged with the model [
31]. To add independent variables to a model, using a linear or non-linear parametric model is an option that is frequently used in forestry [
20,
24]. Another way is to use machine learning and train a new model with validation data [
32]. In forestry and biology, certain types of machine learning, such as neural networks, Maximum Likelihood and random forests are employed [
33,
34], but the use of ongoing advancements such as automated machine learning is still emerging [
35]. This approach allows one to test different types of machine learning or mix them to obtain the optimal solution for a specific situation [
36]. Although the use of this approach facilitates interaction compared to other methods of machine learning, certain criteria such as the proper selection of training and validation data, as well as suitable inputs, are crucial to maximize effectiveness [
37]. The large amount of inventory data in Quebec, both from permanent and temporary plots coinciding with LiDAR acquisition, provides an opportunity to test this approach [
38,
39,
40]. With a robust database and careful selection of inputs, the use of this modern method could lead to increased precision in forest types dominated by shade tolerant deciduous trees, aiming to achieve a level of accuracy similar to that found in boreal forests. Characterizing model error [
41] and calculating mean bias, precision and accuracy of a model’s predictions are essential steps of the quantitative validation process [
42] and are necessary to make a decision on the validity of a model [
43] and choose the best model.
The objectives of this study were as follows: (i) assess the accuracy of predictions for stand BA obtained with the LDM across three distinct stand types: shade-tolerant deciduous, shade-intolerant deciduous and conifers. (ii) identify and test independent variables that can increase the precision of BA predictions, including variables originally used for LDM production as well as additional geospatial and climatic factors. (iii) test model improvements by comparing linear, non-linear, and automated machine learning approaches to enhance the accuracy of LDM predictions.
3. Results
3.1. LiDAR Dendrometric Map Accuracy Assessment
For practical purposes (AAC calculations, harvest planning, etc.), forest inventory must provide useful information by species or groups of species.
Table 3 displays the average observed and predicted basal area for each of the three stand types and dominant species along with the computed statistics.
Bias indicates that on average, the LDM underestimated the basal area for coniferous stands while it overestimated it for the deciduous stands. Shade-tolerant deciduous stands showed slightly higher Bias (2.79 m²/ha) and MAE (6.01 m2/ha) than the other two stand types. According to RMSE, the LDM is slightly better at predicting BA for shade-intolerant deciduous than for the other stand types. R² and the r²ER showed that the independent variables in the LDM explained a greater proportion of the variability of the BA for the coniferous (53.88%) and shade-intolerant deciduous (46.78%) stands than for the shade-tolerant deciduous stands (26.58%).
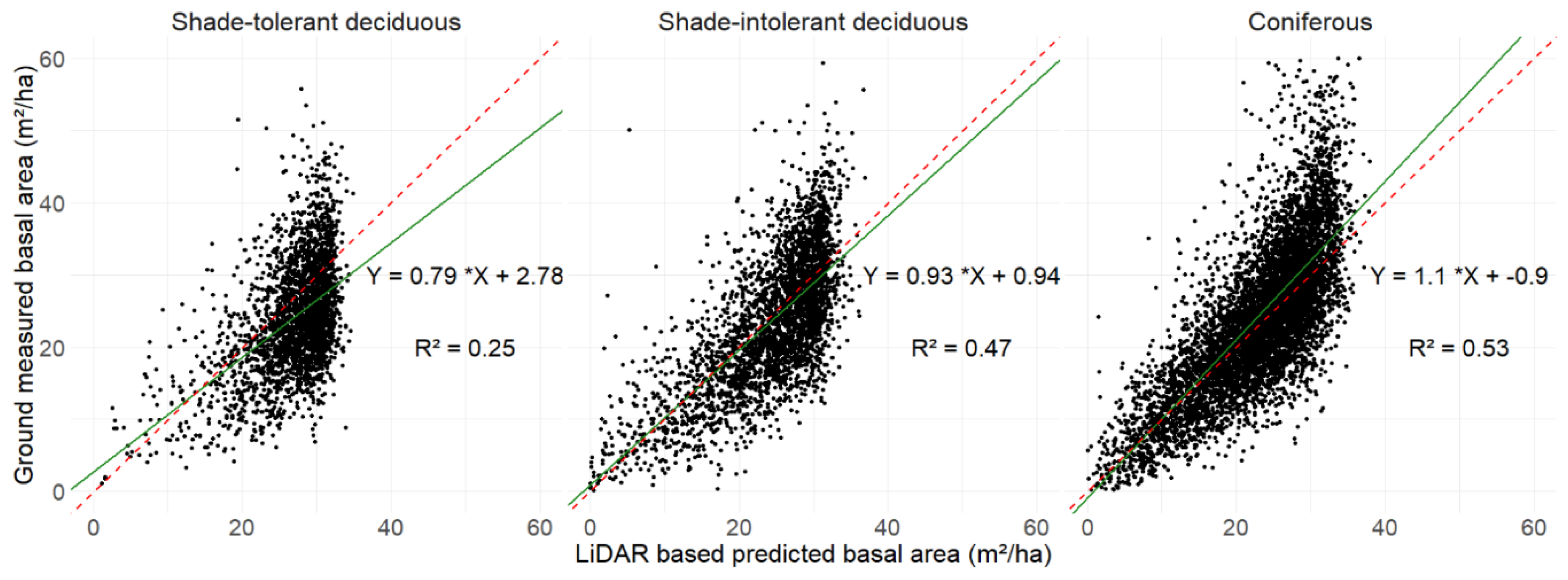
Figure 2 illustrates the relationship between ground-observed data and the LiDAR Dendrometric Map. The closer the line approaches a 1:1 ratio, the more accurate the results are.
Figure 2 illustrates that shade-tolerant deciduous stands exhibit a correlation deviating from the 1:1 line, as indicated by the equation Y = 0.79x + 2.78 (a). Conversely, shade-intolerant and coniferous stands demonstrate regressions closer to the 1:1 line, with equations of Y = 0.93x + 0.94 (b) and Y = 1.1x - 0.9 (c), respectively.
3.2. Independent Variables Analysis
Stepwise selection method
A total of 25 variables were retained, including localization data, species composition percentages, and meteorological variables. The selected variables are indicated with the index 1 in
Table 2.
Boruta method
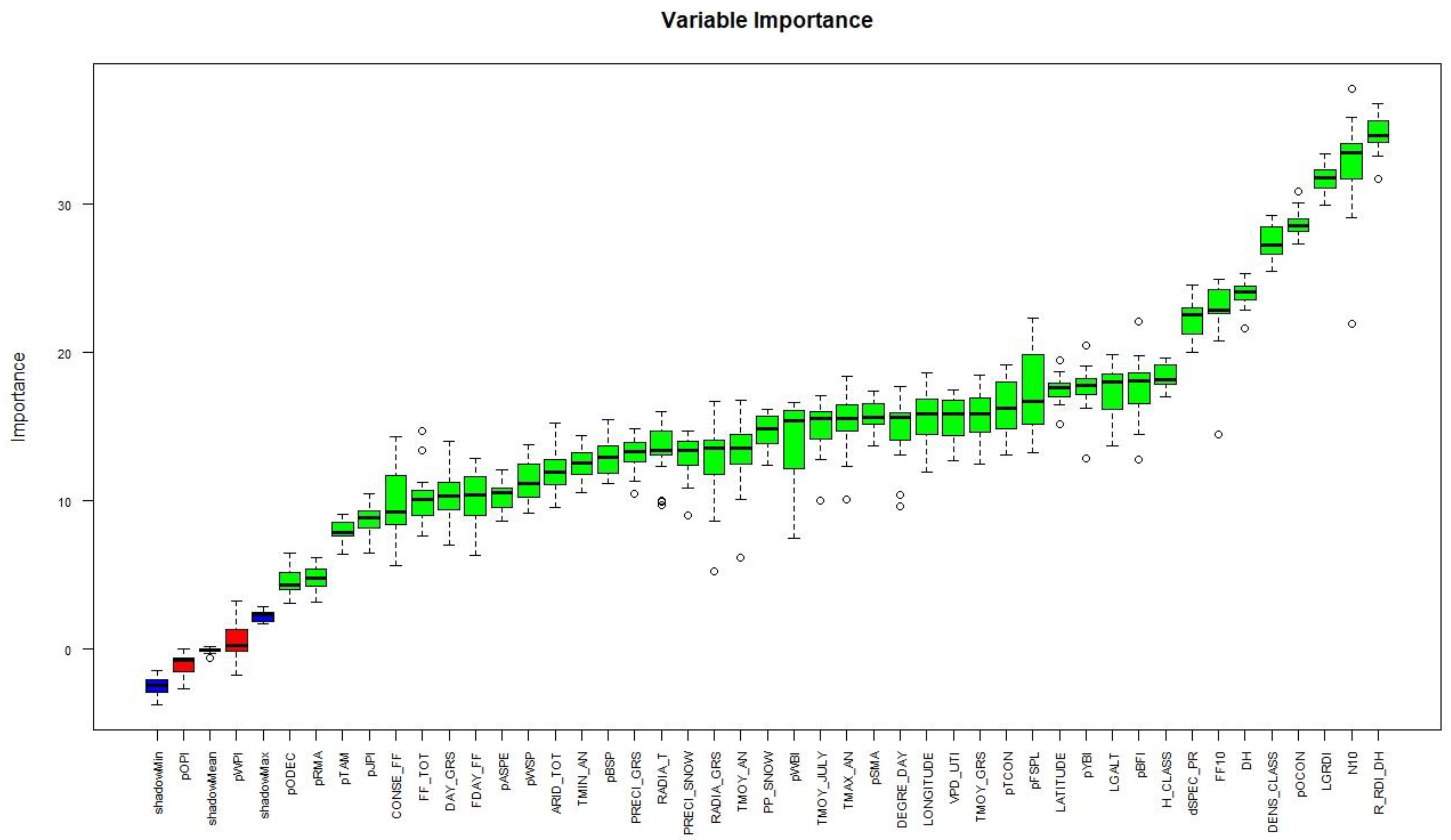
Figure 3 depicts the Boruta variable importance chart. Blue boxes represent the maximum, mean, and minimum Z score of a shadow attribute. Green and red boxplots represent Z score of confirmed and rejected variables respectively.
Out of the 44 variables, two were rejected (pOPI and pWPI) and 42 were confirmed.
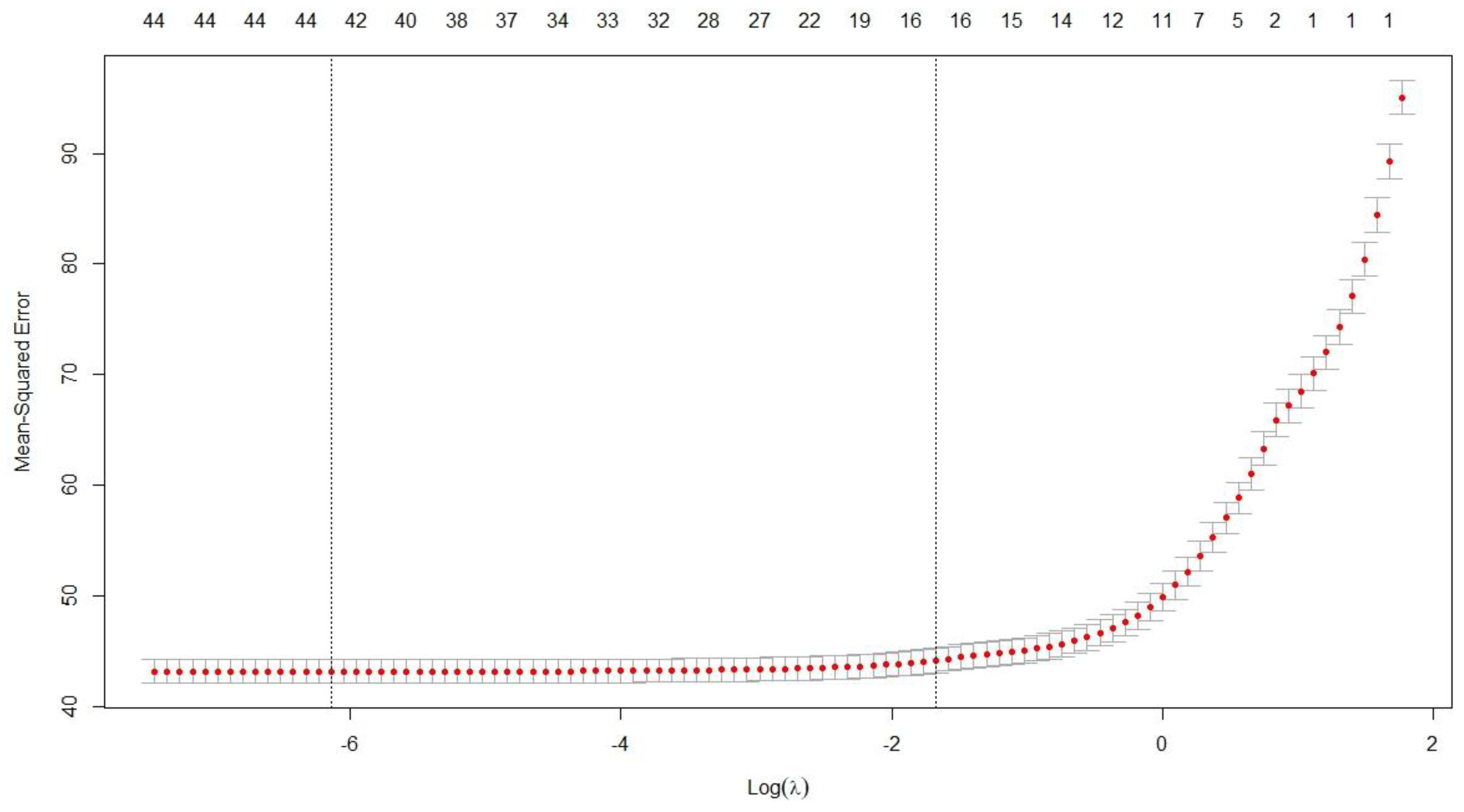
Figure 4 and
Table 4 illustrate the results obtained from the Lasso method.
The two vertical lines correspond to λ_min, which minimizes the cross-validated mean squared error (MSE) and retains more predictors, and λ_1se, within one standard error of the minimum, producing a more parsimonious model with fewer variables. According to these values, the optimal number of variables falls within the range of 16 to 43.
The method utilized ranks variables based on their coefficients, indicating the change in the response variable for a one-unit increase in the corresponding predictor variable, while holding all other variables constant. Because variables were set on the same scale, the absolute values of the coefficients directly correlate with their importance in the model.
Table 4 shows the variable rank by coefficient value.
Table 4.
Variables Best 25 Selected by Lasso Regression Ranked by Absolute Coefficient Value.
Table 4.
Variables Best 25 Selected by Lasso Regression Ranked by Absolute Coefficient Value.
| Rank |
Variable |
Coefficient |
Abs (Coefficient) |
| 1 |
R_RDI_DH |
1037.24 |
1037.24 |
| 2 |
FF10 |
15.93 |
15.93 |
| 3 |
LGALT |
-2.60 |
2.60 |
| 4 |
TMOY_GRS |
-2.30 |
2.30 |
| 5 |
TMAX_AN |
1.70 |
1.70 |
| 6 |
LATITUDE |
1.01 |
1.01 |
| 7 |
LGRDI |
-0.92 |
0.92 |
| 8 |
H_CLASS |
-0.63 |
0.63 |
| 9 |
ARID_TOT |
-0.63 |
0.63 |
| 10 |
TMOY_JULY |
-0.38 |
0.38 |
| 11 |
TMIN_AN |
-0.32 |
0.32 |
| 12 |
pOCON |
0.24 |
0.24 |
| 13 |
DH |
-0.12 |
0.12 |
| 14 |
LONGITUDE |
0.12 |
0.12 |
| 15 |
dSPEC_PR |
0.11 |
0.11 |
| 16 |
FF_TOT |
-0.10 |
0.10 |
| 17 |
DAY_GRS |
0,09 |
0,09 |
| 18 |
pWSP |
0,07 |
0,07 |
| 19 |
CONSE_FF |
0,06 |
0,06 |
| 20 |
pBFI |
0,05 |
0,05 |
| 21 |
pWBI |
0,04 |
0,04 |
| 22 |
pBSP |
0,04 |
0,04 |
| 23 |
pASPE |
0,03 |
0,03 |
| 24 |
PRECI_GRS |
-0,03 |
0,03 |
| 25 |
FDAY_FF |
-0,03 |
0,03 |
Table 4 presents the ranking of the 25 most influential variables identified by the Lasso regression; beyond this point, coefficient magnitudes gradually decreased from PP_SNOW (0.03) to N10 (-0.0003). The only variable with a coefficient of 0 is
TMOY_AN, which means it was not considered to have any influence on the model and was excluded from the lasso selection.
Because each feature selection method produced a different number of selected variables, subsets were created to allow a fair comparison by using the same number of variables across methods. Since the stepwise method does not provide a variable ranking, subsets were generated only for Boruta and Lasso. Three rankings were considered: the 16 best variables according to Lasso λ_1se (within one standard error of the minimum), the 25 best variables corresponding to the number selected by the stepwise method, and the corresponding number of variables selected by each method. This procedure resulted in seven subsets, with the selected variables for each method presented in
Table 2 along with their respective indices.
1 Stepwise method, 25 selected variables
2 Boruta method, best 16 variables
3 Boruta method, best 25 variables
4 Boruta method, 42 selected variables
5 Lasso method, λ_1se 16 variables
6Lasso method, best 25 variables
7 Lasso method, λ_min 43 variables
3.3. Model Improvement with Three Approaches
3.3.1. Linear Parametric, Non-Linear Parametric and H20 AutoML Models
For comparison, all new modeling approaches were developed using the same set of variables selected through the stepwise backward selection method (Leboeuf et al., 2022).
Linear Model
Equation (7) displays the results of the optimized linear model.
Where
BAcorrected is the prediction of the model,
factor(dSPEC_PR) represents a coefficient B assigned to each species, which is only applied when that species is the dominant one in the stand. Other predictors are also present in the model but are not explicitly displayed in the formula due to their lower relative importance
. Results are shown in
Table 5.
Non-Linear Model
Out of all the non-linear models tested, the quadratic regression model produced the best results. Equation (8) displays the optimized quadratic base on the same approach as the linear model.
Where
BAcorrected is the prediction of the model,
factor(dSPEC_PR) represents a coefficient B assigned to each species, which is only applied when that species is the dominant one in the stand. Other predictors are also present in the model but are not explicitly displayed in the formula due to their lower relative importance
. Results are shown in
Table 5.
H2O AutoML Model
The H2O AutoML function does not provide a detailed description of the best resulting model because its primary goal is to automate the model selection and tuning process rather than to generate interpretable model outputs. The best-performing model often corresponds to a Stacked Ensemble, which combines multiple base learners (e.g., GBM, DRF, DNN, GLM) to maximize predictive accuracy. This ensemble model integrates the predictions of several algorithms through a meta-learner, making the internal structure complex and not easily summarized in a single set of equations or coefficients.
The leaderboard indicates that the Stack Ensemble Model achieved the highest performance among all AutoML candidates, followed by GBM, and outperformed DRF, DNN, and GLM. Results for the best model are presented in
Table 5 under H20 AutoML.
For all three modeling approaches, the best performance for shade-tolerant species was achieved by training models exclusively on data from shade-tolerant deciduous stands, using the same input variables.
3.3.2. Comparing Models Performance: New Approaches vs. Initial Model
Table 5 shows that the H2O AutoML method achieved the best overall performance across both groups. For all commercial species, r²
ER increased from 45.59 % (LDM) to 59.75 %, with MAE reduced from 5.47 m²/ha to 4.78 m²/ha and bias remained close to zero (-0.05). For shade-tolerant species, r²
ER improved from 13.37 % to 31.01 %, MAE decreased from 6.03 m²/ha to 5.13 m²/ha, and bias was also minimized from 2.70 m²/ha to 0.01 m²/ha. Linear and quadratic models also provided considerable improvements, with the linear model performing slightly better for both all commercial species and shade-tolerant species.
In order to further improve model performance, subsequent evaluations were conducted using the other variable subsets with the H20 AutoML, the best-performing modeling approach.
3.3.3. Testing the variable selection impacts on the best performing model (H20 AutoML)
To select the inputs for automatic machine learning, a comparison between metrics selection approaches was conducted and is presented in
Table 6.
Based on the results presented in
Table 6, using only 16 input variables is insufficient to maximize the explained variance, while increasing the number of inputs beyond 25 does not lead to a significant improvement in predictive accuracy. No significant differences in performance were observed between the feature selection methods. The stepwise method with 25 selected variables was therefore chosen to generate the final model. This configuration provides performance comparable to both the 25-variable Boruta subset and the 25-variable Lasso subset, with the added advantage that all 25 inputs were retained by the stepwise method, whereas Boruta and Lasso retain 42 and 43 variables, respectively, without providing additional predictive gain.
3.3.4. Testing Best Model with best variable selection
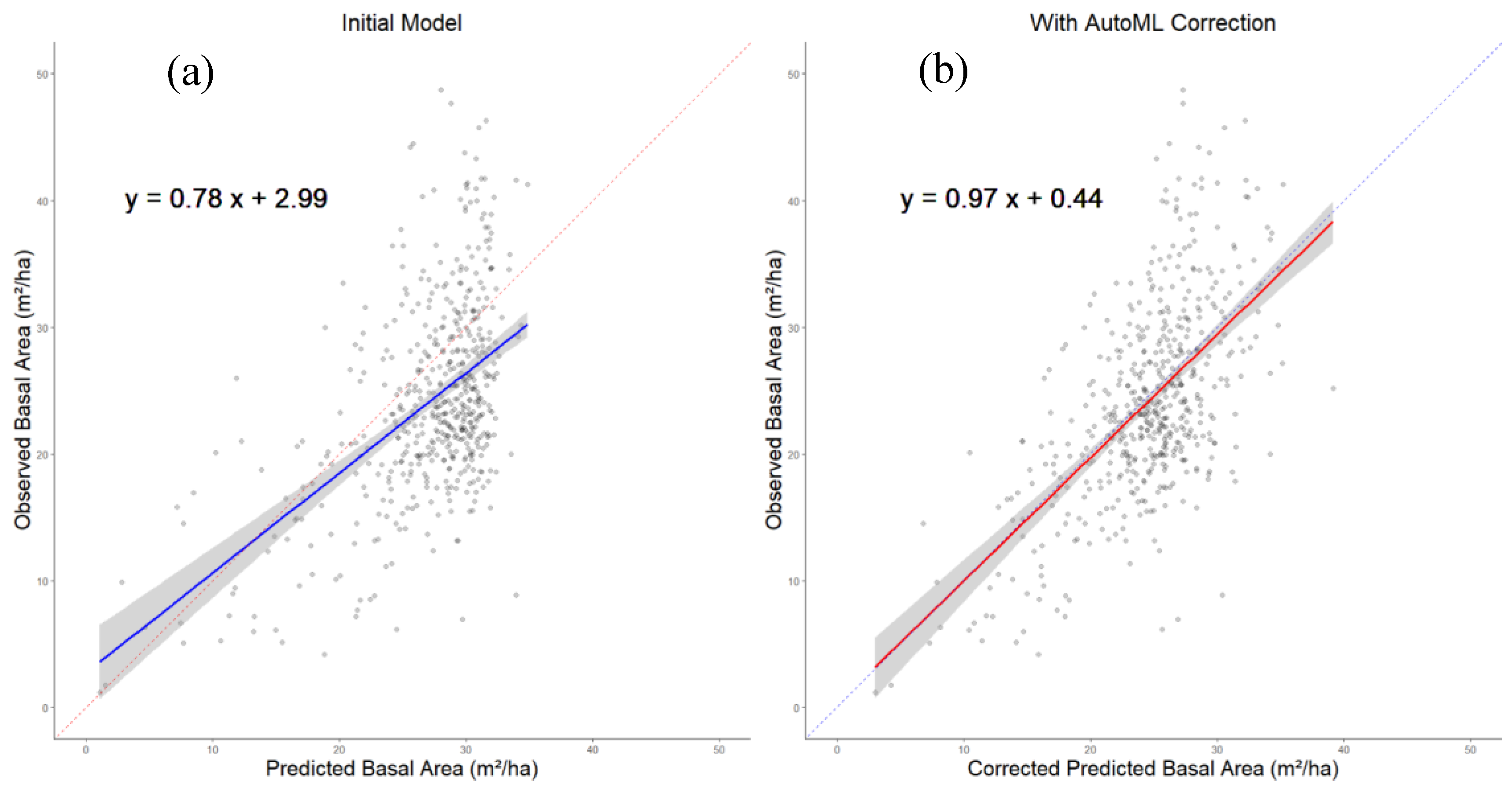
A graphical representation illustrating the correction's impact using AutoML on the distribution of data, comparing observed and predicted basal area in shade-tolerant deciduous, is depicted in
Figure 5.
We observe that the results are centered on the 1:1 line in the Auto ML corrected model; however, the data points are still widely scattered around this line, highlighting the lack of accuracy.
Results for each stand type show the improvement from the initial model to the best model (
Table 7). The test dataset is used for the comparison, which represents 20% of the total data, ensuring trustworthy results by using data that was not part of the model training.
Improvements were observed across all categories, with the most significant corrections seen in shade-tolerant deciduous stands in terms of bias, MAE and r²ER. Coniferous species received the most significant correction in terms of RMSE and R². Despite the improvements, shade-tolerant deciduous stands still exhibit a lower performance indicator than the other stands in the initial model.
4. Discussion
This project has provided essential confirmation that ALS-based approaches, such as the LDM developed by the Quebec government's forest inventory program, still require refinement before they can be extended to temperate forests with complex stands, like those in the southern regions of Quebec. In particular, shade-tolerant deciduous stands have been identified as the main challenge requiring further research and development. The incorporation of geolocalized data independent of ground samples, together with the original metrics of the LDM, combined with the use of automated machine learning, has led to notable improvements. Although these enhancements have not yet yielded competitive precision for shade-tolerant deciduous stands, the approach developed in this project opens a new avenue for generating data that can be integrated with future advancements in the field. The approach adopted in this project lays the foundation for leveraging automated machine learning to fully exploit the potential of large datasets and incorporate new inputs derived from recent advances in forest remote sensing.
When assessing the LDM model's precision in predicting plot basal area across different stand types, it yielded R² values of 53.4% in coniferous stands, 45.2% in shade-intolerant deciduous stands, and only 24.6% in shade-tolerant deciduous stands. The low R² in the latter indicates a weak correlation between the predicted and observed basal area. Although shade-tolerant deciduous stands have the lowest R², the RMSE is higher in coniferous stands, this suggests that while the model captures overall trends better in conifers (higher R²), individual plot predictions for conifers are more variable, leading to larger absolute errors.
Further analysis revealed that dominant species composition significantly affects BA prediction accuracy, consistent with previous studies [
23,
24,
25]. Stands dominated by sugar maple and yellow birch showed particularly poor performance, with R² values of 22.6% and 24.3%, and overestimations of 4.4 m²/ha and 2.4 m²/ha, respectively. The r²
ER of -17.7% for the sugar maple indicates that the LiDAR model performs worse than a trivial baseline, due to systematic overestimation of BA. This result highlights a clear performance issue of the model in this forest type. Stands dominated by white cedar also performed poorly, with an R² of 27.1% and an underestimation of 6.6 m²/ha. This bias contributes to the higher RMSE observed in conifer stands. These species-specific biases highlight the importance of considering species composition in the model.
Given that other studies have reported R² values exceeding 90% for basal area estimation [
19,
20], we examined the relationship between ground observations and predictions graphically to identify potential weaknesses in the model. The model appears resistant to predicting values above 35 m²/ha, suggesting a saturation effect driven by model constraints that limit overestimating basal area at the pixel level. While this limitation may be beneficial for large-scale estimations, it restricts accuracy at the plot level, suggesting that the 5m × 5m data provided by the LDM is not well-suited for fine-scale predictions where higher variability in BA is typically observed.
To complement the species information already considered in the LDM, an alternative metric - the proportion of basal area covered by species- was tested alongside LDM inputs, geospatial climate variables, and location coordinates. Six variables were consistently identified as most important across all input selection methods: R_RDI_DH, H_CLASS, dSPEC_PR and pOCON, TMOY_GRS and LATITUDE. Among these, R_RDI_DH and H_CLASS confirmed their strong relationship with BA, whereas species-related metrics such as dSPEC_PR and pOCON emphasize the role of species composition and help explain the poor performance of the LDM in cedar-dominated stands [
67]. Specifically, the former captures species-specific height-to-diameter ratios, while the latter reflects structural divergence in stands such as cedar. Climatic and geographic variables (TMOY_GRS, LATITUDE) further demonstrate how environmental conditions shape forest structure. Conversely, the rejection of redundant predictors such as TMOY_AN illustrates the model’s ability to prioritize the most informative variables.
Integrating these variables into the three modeling approaches improved the results. For shade-tolerant species, the initial R² of 26.6% increased to 31.2% (AutoML), 31.0% (linear regression), and 30.6% (quadratic regression). For all commercial species, the initial R² of 45.6% improved to 59.8%, 55.9%, and 54.6% under AutoML, linear regression, and quadratic regression, respectively. Given its superior performance, the AutoML modeling approach was selected for the subsequent analyses. The results showed that using 16 input variables was insufficient to achieve optimal accuracy, whereas increasing the number of inputs beyond 25 did not yield substantial gains. No significant difference was observed between the model selection methods. In the comparison between AutoML (25 splitwise-selected variables) and the LDM model (
Figure 5), the elimination of the prediction plateau at 35 m²/ha in the AutoML outputs suggests that this approach better captures stand dynamics and reduces systematic bias. The closer alignment of predicted versus observed values with the 1:1 line further indicates an improved capacity to generalize across conditions. However, despite the gains observed in shade-tolerant deciduous stands, model precision remains limited compared to the performance levels achieved in coniferous and shade-intolerant deciduous stands, underscoring the persistent challenges of modeling heterogeneous forest structures.
This research supports the MNRF’s efforts to identify optimal LiDAR-based approaches for southern Quebec’s forest inventory. It confirms that the LDM requires further refinement to handle structurally irregular, hardwood-dominated forests. The improvements made here offer guidance for future aerial LiDAR applications in complex forest environments.
To enhance BA prediction in shade-tolerant deciduous stands, new structural variables must be explored. Satellite imagery has shown mixed results [
68,
69,
70], and canopy complexity remains a challenge [
71]. The height-BA relationship may be insufficient, prompting investigation into variables like slope angle, stem size distribution [
72], and vertical foliage profiles [
73]. Promising LiDAR-derived metrics include mean height filtered by intensity thresholds and leaf area density variation [
24,
74]. With ongoing advances, species-level identification via aerial LiDAR may soon be feasible [
75,
76], offering transformative inputs for models highly sensitive to dominant species.
This study faced several challenges that may have influenced model performance and interpretation. A key limitation was the need to obtain a large volume of observed data corresponding to MNRF validation plots, where GPS imprecision and human interpretation bias may have introduced spatial and semantic errors. Despite the careful selection process of ground sample plots, it was not possible to verify whether unrecorded disturbances (e.g., harvesting, windthrow, etc.) occurred between the field data collection period and LiDAR acquisition, potentially degrading the quality of the training dataset. Moreover, some variables used in the model were designed for broad-scale applications and lacked the spatial specificity required for fine-scale plot-level predictions. For instance, tree species composition percentages are currently derived from photo-interpretation over sectors spanning several hectares. This coarse resolution may introduce discrepancies when applied to 400 m² plots, where species composition can vary significantly. The available LiDAR dataset had already been transformed into predefined summary metrics, with a single value per plot. This constrained the capacity to extract new structural information directly from the point cloud. As a result, model improvements had to rely on external inputs independent of LiDAR, such as climate variables or geolocation. Incorporating custom LiDAR-derived variables—tailored to local canopy structure and vertical heterogeneity—could enhance the model’s sensitivity to subtle structural variations. Another limitation stems from the use of automated machine learning (AutoML), particularly the Stacked Ensemble Model, which does not provide interpretable model parameters. While feature importance can be assessed, the internal logic of the ensemble remains opaque. This reinforces the importance of careful input selection, as model transparency is limited. Additionally, a common issue in machine learning regression models was observed: low values were generally overestimated, whereas high values were underestimated [
77]. To address this, the Regression of Observed on Estimated values (ROE) method was applied to correct the bias [
78], but the improvements were marginal. This suggests that further investigation is needed to determine whether H2O's AutoML framework exhibits structural bias in extreme values.
In summary, the study has advanced our understanding of the link between stand type and ALS-based model accuracy and highlighted potential improvements.