Submitted:
02 February 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
Methods
2. Clinical and Therapeutic Challenges in CBF-AML
Limitations of Current Standard Therapies (7+3 Regimen, Consolidation)
3. Molecular Basis of Heterogeneity in CBF-AML
3.1. The t(8;21)(q22;q22) Translocation and RUNX1-RUNX1T1 Fusion Gene
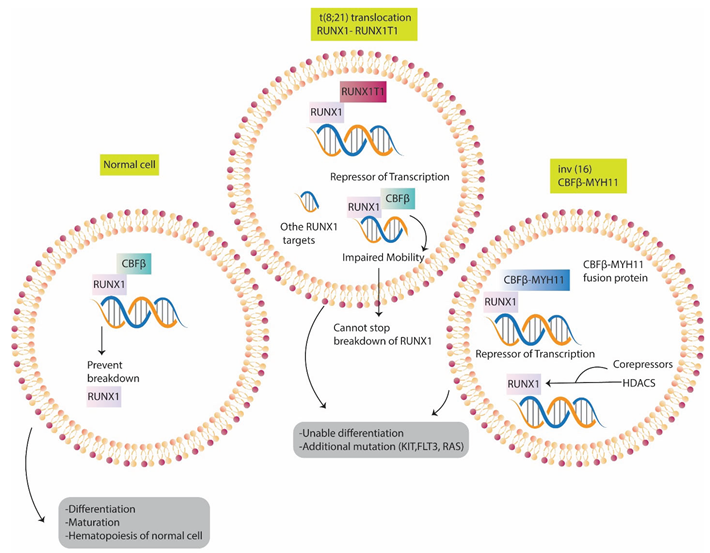
3.2. Clinical Phenotype and Treatment Sensitivity: Divergent Implications of CBF
Fusion Subtypes
3.3. Cooperation with Secondary Mutations and Therapeutic Challenges
3.4. FLT3 Mutations in CBF-AML: Infrequent but Clinically Relevant
3.5. KIT Mutations in CBF-AML: Subtype-Specific Risk Modifiers with Unresolved Therapeutic Implications
3.6. RAS Pathway Mutations in CBF-AML: Frequent but Largely Prognostically Neutral
3.7. ASXL1 and ASXL2 Mutations in CBF-AML: Frequency Does Not Equal Prognostic Impact
3.8. Cohesin Complex Alterations in CBF-AML: Markers of Clonal Evolution Rather than Immediate Therapeutic Targets
3.9. ZBTB7A Mutations in CBF-AML: Recurrent in t(8;21) but Clinically Unresolved
3.10. WT1 Alterations in CBF-AML: Adverse Marker or Surrogate of High-Risk Biology?
4. Molecular Risk Stratification and MRD-Guided Prognostic Modeling
5. Emerging Therapeutic Strategies Addressing Heterogeneity
5.1. Targeted Therapy in CBF-AML
5.2. KIT Inhibition
5.3. Exploratory Strategies: Menin Inhibition and Novel Combinations
5.4. Practice-Changing Targeted Therapy: Gemtuzumab Ozogamicin
5.5. Promising but Unproven Approaches: FLT3 Inhibition
5.6. Epigenetic Therapies in CBF-AML: Context-Dependent and Largely Investigational
5.7. Role of Hematopoietic Stem Cell Transplantation in High-Risk and Relapsed Patients
6. Conclusions
References
- Hasserjian RP. Acute myeloid leukemia: advances in diagnosis and classification. Int J Lab Hematol [Internet]. 2013 Jun [cited 2025 Oct 30];35(3):358–66. Available from: https://pubmed.ncbi.nlm.nih.gov/23590662/.
- Appelbaum FR. WHO, what, when, where, and why: New classification systems for acute myeloid leukemia and their impact on clinical practice. Best Pract Res Clin Haematol [Internet]. 2023 Dec 1 [cited 2025 Oct 30];36(4). Available from: https://pubmed.ncbi.nlm.nih.gov/38092471/.
- Orvain C, Bertoli S, Peterlin P, Desbrosses Y, Dumas PY, Iat A, et al. Molecular relapse after first-line intensive therapy in patients with CBF or NPM1-mutated acute myeloid leukemia – a FILO study. Leuk 2024 389 [Internet]. 2024 Jul 17 [cited 2025 Oct 30];38(9):1949–57. Available from: https://www.nature.com/articles/s41375-024-02335-2.
- Borthakur G, Kantarjian H. Core binding factor acute myelogenous leukemia-2021 treatment algorithm. Blood Cancer J 2021 116 [Internet]. 2021 Jun 16 [cited 2025 Oct 30];11(6):1–5. Available from: https://www.nature.com/articles/s41408-021-00503-6.
- Kundu M, Chen A, Anderson S, Kirby M, Xu LP, Castilla LH, et al. Role of Cbfb in hematopoiesis and perturbations resulting from expression of the leukemogenic fusion gene Cbfb-MYH11. Blood [Internet]. 2002 Oct 1 [cited 2025 Oct 30];100(7):2449–56. Available from: https://pubmed.ncbi.nlm.nih.gov/12239155/.
- Link KA, Chou FS, Mulloy JC. Core binding factor at the crossroads: Determining the fate of the HSC. J Cell Physiol. 2010 Jan;222(1):50–6.
- De Bruijn MFTR, Speck NA. Core-binding factors in hematopoiesis and immune function. Oncogene [Internet]. 2004 May 24 [cited 2025 Oct 30];23(24):4238–48. Available from: https://pubmed.ncbi.nlm.nih.gov/15156179/.
- Talami A, Bettelli F, Pioli V, Giusti D, Gilioli A, Colasante C, et al. How to Improve Prognostication in Acute Myeloid Leukemia with CBFB-MYH11 Fusion Transcript: Focus on the Role of Molecular Measurable Residual Disease (MRD) Monitoring. Biomed 2021, Vol 9, Page 953 [Internet]. 2021 Aug 3 [cited 2025 Oct 30];9(8):953. Available from: https://www.mdpi.com/2227-9059/9/8/953/htm.
- Khan M, Cortes J, Qiao W, Alzubaidi MA, Pierce SA, Ravandi F, et al. Outcomes of patients with relapsed core binding factor-positive acute myeloid leukemia. Clin Lymphoma Myeloma Leuk [Internet]. 2017 Jan 1 [cited 2025 Oct 30];18(1):e19. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC5861376/.
- George B, Yohannan B, Mohlere V, Gonzalez A. Therapy-related core binding factor acute myeloid leukemia. Int J Hematol Oncol [Internet]. 2023 Mar [cited 2025 Oct 31];12(1):IJH43. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC9979104/.
- Chiu M, Schimmer AD, Schuh AC, Bankar A, Richard-Carpentier G, Sibai H, et al. Genomic profiles and outcomes in de novo versus therapy-related core binding factor AML. Blood Cancer J 2024 141 [Internet]. 2024 Oct 31 [cited 2025 Oct 31];14(1):1–5. Available from: https://www.nature.com/articles/s41408-024-01166-9.
- Borthakur G, Kantarjian H. Core binding factor acute myelogenous leukemia-2021 treatment algorithm. Blood Cancer J 2021 116 [Internet]. 2021 Jun 16 [cited 2025 Oct 31];11(6):1–5. Available from: https://www.nature.com/articles/s41408-021-00503-6.
- Rogers HJ, Wang X, Xie Y, Davis AR, Thakral B, Wang SA, et al. Comparison of therapy-related and de novo core binding factor acute myeloid leukemia: A bone marrow pathology group study. Am J Hematol [Internet]. 2020 Jul 1 [cited 2025 Oct 31];95(7):799–808. Available from: /doi/pdf/10.1002/ajh.25814.
- Fareed S, Soliman DS, Al-Mashdali AF, Gameil A, Mulikandathil Y, Alshurafa A, et al. Acute Myeloid Leukemia with Core Binding Factor Rearrangements: A 10-Year Cancer Center Experience. Oncol [Internet]. 2025 [cited 2025 Oct 31]; Available from: https://dx.doi.org/10.1159/000544049.
- Shimony S, Stahl M, Stone RM. Acute Myeloid Leukemia: 2025 Update on Diagnosis, Risk-Stratification, and Management. Am J Hematol [Internet]. 2025 May 1 [cited 2025 Oct 31];100(5):860. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC11966364/.
- Begna KH, Xu X, Gangat N, Alkhateeb H, Patnaik MM, Al-Kali A, et al. Core-binding factor acute myeloid leukemia: long-term outcome of 70 patients uniformly treated with “7+3.” Blood Cancer J 2022 124 [Internet]. 2022 Apr 7 [cited 2025 Oct 31];12(4):1–4. Available from: https://www.nature.com/articles/s41408-022-00654-0.
- Ronnacker J, Muller PJ, Mikesch JH, Zukunft S, Weinbergerová B, Šrámek J, et al. Gemtuzumab ozogamicin in first-line treatment of CBF-AML: insights from a retrospective multi-center analysis. Leuk 2025 399 [Internet]. 2025 Jul 21 [cited 2025 Oct 31];39(9):2174–80. Available from: https://www.nature.com/articles/s41375-025-02700-9.
- Marcucci G, Geyer S, Laumann K, Zhao W, Bucci D, Uy GL, et al. Combination of dasatinib with chemotherapy in previously untreated core binding factor acute myeloid leukemia: CALGB 10801. Blood Adv [Internet]. 2020 Feb 25 [cited 2025 Oct 31];4(4):696–705. Available from: https://pubmed.ncbi.nlm.nih.gov/32092139/.
- Dasatinib Misses Survival End Points in Core-Binding Factor AML | CancerNetwork [Internet]. [cited 2025 Oct 31]. Available from: https://www.cancernetwork.com/view/dasatinib-misses-survival-end-points-in-core-binding-factor-aml?utm_source=chatgpt.com.
- Hills RK, Castaigne S, Appelbaum FR, Delaunay J, Petersdorf S, Othus M, et al. Addition of gemtuzumab ozogamicin to induction chemotherapy in adult patients with acute myeloid leukaemia: A meta-analysis of individual patient data from randomised controlled trials. Lancet Oncol [Internet]. 2014 Aug 1 [cited 2025 Oct 31];15(9):986–96. Available from: https://www.thelancet.com/action/showFullText?pii=S1470204514702815.
- Bruserud O, Reikvam H, Hatfield KJ, Kittang AO, Hovland R. Acute Myeloid Leukemia with the t(8;21) Translocation: Clinical Consequences and Biological Implications. J Biomed Biotechnol [Internet]. 2011 [cited 2025 Nov 8];2011:104631. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC3100545/.
- Al-Harbi S, Aljurf M, Mohty M, Almohareb F, Ahmed SOA. An update on the molecular pathogenesis and potential therapeutic targeting of AML with t(8;21)(q22;q22.1);RUNX1-RUNX1T1. Blood Adv [Internet]. 2020 Jan 14 [cited 2025 Oct 31];4(1):229–38. Available from: https://pubmed.ncbi.nlm.nih.gov/31935293/.
- Day RB, Hickman JA, Xu Z, Katerndahl CDS, Ferraro F, Ramakrishnan SM, et al. Proteogenomic analysis reveals cytoplasmic sequestration of RUNX1 by the acute myeloid leukemia–initiating CBFB::MYH11 oncofusion protein. J Clin Invest [Internet]. 2024 Feb 15 [cited 2025 Oct 31];134(4). Available from: https://www.researchgate.net/publication/376312045_Proteogenomic_analysis_reveals_cytoplasmic_sequestration_of_RUNX1_by_the_acute_myeloid_leukemia-initiating_CBFBMYH11_oncofusion_protein.
- Grinev V V., Barneh F, Ilyushonak IM, Nakjang S, Smink J, van Oort A, et al. RUNX1/RUNX1T1 mediates alternative splicing and reorganises the transcriptional landscape in leukemia. Nat Commun 2021 121 [Internet]. 2021 Jan 22 [cited 2025 Oct 31];12(1):1–16. Available from: https://www.nature.com/articles/s41467-020-20848-z.
- Grinev V V., Migas AA, Kirsanava AD, Mishkova OA, Siomava N, Ramanouskaya T V., et al. Decoding of exon splicing patterns in the human RUNX1-RUNX1T1 fusion gene. Int J Biochem Cell Biol [Internet]. 2015 Nov 9 [cited 2025 Oct 31];68:48–58. Available from: https://pubmed.ncbi.nlm.nih.gov/26320575/.
- Li X, Liu G. Acute Myeloid Leukemias with Variant RUNX1::RUNX1T1: Report of Three Cases. Hematopathology [Internet]. 2023 May 1 [cited 2025 Oct 31];8(1). Available from: http://journals.librarypublishing.arizona.edu/hemepath/article/id/5668/.
- Detection of inv(16) and t(16;16) by fluorescence in situ hybridization in acute myeloid leukemia M4Eo | Haematologica [Internet]. [cited 2025 Oct 31]. Available from: https://haematologica.org/article/view/1679.
- Mandoli A, Singh AA, Jansen PWTC, Wierenga ATJ, Riahi H, Franci G, et al. CBFB-MYH11/RUNX1 together with a compendium of hematopoietic regulators, chromatin modifiers and basal transcription factors occupies self-renewal genes in inv(16) acute myeloid leukemia. Leukemia [Internet]. 2014 [cited 2025 Oct 31];28(4):770–8. Available from: https://pubmed.ncbi.nlm.nih.gov/24002588/.
- Pulikkan JA, Castilla LH. Preleukemia and Leukemia-initiating cell activity in inv(16) acute myeloid Leukemia. Front Oncol [Internet]. 2018 Apr 26 [cited 2025 Oct 31];8(APR):324072. Available from: www.frontiersin.org.
- Hyde RK, Liu PP. RUNX1 repression-independent mechanisms of leukemogenesis by fusion genes CBFB-MYH11 and AML1-ETO (RUNX1-RUNX1T1). J Cell Biochem [Internet]. 2010 Aug 1 [cited 2025 Oct 31];110(5):1039–45. Available from: https://pubmed.ncbi.nlm.nih.gov/20589720/.
- Sinha C, Cunningham LC, Liu PP. Core Binding Factor Acute Myeloid Leukemia: New Prognostic Categories and Therapeutic Opportunities. Semin Hematol [Internet]. 2015 Jul 1 [cited 2025 Oct 31];52(3):215–22. Available from: https://pubmed.ncbi.nlm.nih.gov/26111469/.
- Döhner K, Döhner H. Molecular characterization of acute myeloid leukemia. Haematologica [Internet]. 2008 Jul 1 [cited 2025 Oct 31];93(7):976–82. Available from: https://haematologica.org/article/view/4939.
- Zhang W, Wang H, Zhang P, Li H, Ma X, Liu H. Rare type I CBFβ/MYH11 fusion transcript in primary acute myeloid leukemia with inv(16)(p13.1q22): a case report. Brazilian J Med Biol Res = Rev Bras Pesqui medicas e Biol [Internet]. 2021 [cited 2025 Oct 31];54(12). Available from: https://pubmed.ncbi.nlm.nih.gov/34730684/.
- Srinivasan S, Dhamne C, Patkar N, Chatterjee G, Moulik NR, Chichra A, et al. KIT exon 17 mutations are predictive of inferior outcome in pediatric acute myeloid leukemia with RUNX1::RUNX1T1. Pediatr Blood Cancer [Internet]. 2024 Feb 1 [cited 2025 Oct 31];71(2). Available from: https://pubmed.ncbi.nlm.nih.gov/38014874/.
- Yang M, Wang W, Zhang G, Qiu S, Liu B, Mi Y, et al. Clinical characteristics and therapeutic determinants of RUNX1::RUNX1T1 differ from those of CBFB::MYH11 acute myeloid leukemia. Haematologica [Internet]. 2025 Apr 30 [cited 2025 Oct 31];110(10):2281–92. Available from: https://haematologica.org/article/view/12048.
- Schuurhuis GJ, Heuser M, Freeman S, Buccisano F, Cloos J, Grimwade D, et al. Minimal/measurable residual disease in AML: a consensus document from the European LeukemiaNet MRD Working Party. [cited 2025 Oct 31]; Available from: www.bloodjournal.org.
- Yu S, Yang S, Hu L, Duan W, Zhao T, Qin Y, et al. Genetic abnormalities predict outcomes in patients with core binding factor acute myeloid leukemia. Ann Hematol [Internet]. 2025 Feb 1 [cited 2025 Oct 31];104(2):997–1006. Available from: https://pubmed.ncbi.nlm.nih.gov/39966122/.
- Ayatollahi H, Shajiei A, Sadeghian MH, Sheikhi M, Yazdandoust E, Ghazanfarpour M, et al. Prognostic Importance of C-KIT Mutations in Core Binding Factor Acute Myeloid Leukemia: A Systematic Review. Hematol Oncol Stem Cell Ther [Internet]. 2017 Mar 1 [cited 2025 Nov 4];10(1):1–7. Available from: https://pubmed.ncbi.nlm.nih.gov/27613372/.
- Paschka P, Marcucci G, Ruppert AS, Mrózek K, Chen H, Kittles RA, et al. Adverse Prognostic Significance of KIT Mutations in Adult Acute Myeloid Leukemia With inv(16) and t(8;21): A Cancer and Leukemia Group B Study. J Clin Oncol [Internet]. 2006 Aug 20 [cited 2025 Nov 5];24(24):3904–11. Available from: https://ascopubs.org/doi/10.1200/JCO.2006.06.9500.
- Qin YZ, Jiang Q, Wang Y, Jiang H, Xu LP, Zhao XS, et al. The impact of the combination of KIT mutation and minimal residual disease on outcome in t(8;21) acute myeloid leukemia. Blood Cancer J [Internet]. 2021 Apr 1 [cited 2025 Nov 7];11(4):67. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC8016839/.
- Quan X, Deng J. Core binding factor acute myeloid leukemia: Advances in the heterogeneity of KIT, FLT3, and RAS mutations (Review). Mol Clin Oncol [Internet]. 2020 [cited 2025 Nov 13];13(2):95. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC7366242/.
- Ramirez RMC, Ramirez RMC, Pachas MT de JC, Dongo RJZ. Prevalence and Prognosis of Secondary Genetic Aberrations Among Patients With Core Binding Factor Acute Myeloid Leukemia: A Mitelman Database Analysis. World J Oncol [Internet]. 2023 Nov 17 [cited 2025 Nov 13];14(6):488–98. Available from: https://www.wjon.org/index.php/wjon/article/view/1661.
- Goemans BF, Zwaan CM, Miller M, Zimmermann M, Harlow A, Meshinchi S, et al. Mutations in KIT and RAS are frequent events in pediatric core-binding factor acute myeloid leukemia. Leukemia [Internet]. 2005 [cited 2025 Nov 5];19(9):1536–42. Available from: https://pubmed.ncbi.nlm.nih.gov/16015387/.
- Maiti A, Naqvi K, Kadia TM, Borthakur G, Takahashi K, Bose P, et al. Phase II Trial of MEK Inhibitor Binimetinib (MEK162) in RAS-mutant Acute Myeloid Leukemia. Clin Lymphoma, Myeloma Leuk [Internet]. 2019 Mar 1 [cited 2025 Nov 4];19(3):142-148.e1. Available from: https://pubmed.ncbi.nlm.nih.gov/30635233/.
- Jahn N, Agrawal M, Bullinger L, Weber D, Corbacioglu A, Gaidzik VI, et al. Incidence and Prognostic Relevance of ASXL2 Mutations in Adult CBF-AML with t(8;21)(q22;q22): A Study of the German-Austrian AML Study Group (AMLSG). Blood. 2015 Dec 3;126(23):3818–3818.
- Medina EA, Delma CR, Yang FC. ASXL1/2 mutations and myeloid malignancies. J Hematol Oncol [Internet]. 2022;15(1):1–18. Available from: . [CrossRef]
- Hartmann L, Dutta S, Opatz S, Vosberg S, Reiter K, Leubolt G, et al. ZBTB7A mutations in acute myeloid leukaemia with t(8;21) translocation. Nat Commun [Internet]. 2016 Jun 2 [cited 2025 Nov 8];7:11733. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC4895769/.
- Kawashima N, Akashi A, Nagata Y, Kihara R, Ishikawa Y, Asou N, et al. Clinical significance of ASXL2 and ZBTB7A mutations and C-terminally truncated RUNX1-RUNX1T1 expression in AML patients with t(8;21) enrolled in the JALSG AML201 study. Ann Hematol [Internet]. 2019 Jan 30 [cited 2025 Nov 8];98(1):83–91. Available from: https://pubmed.ncbi.nlm.nih.gov/30251205/.
- Li S, Li N, Chen Y, Zheng Z, Guo Y. FLT3-TKD in the prognosis of patients with acute myeloid leukemia: A meta-analysis. Front Oncol. 2023 Feb 17;13:1086846.
- Guan W, Zhou L, Li Y, Yang E, Liu Y, Lv N, et al. Profiling of somatic mutations and fusion genes in acute myeloid leukemia patients with FLT3-ITD or FLT3-TKD mutation at diagnosis reveals distinct evolutionary patterns. Exp Hematol Oncol [Internet]. 2021 Dec 1 [cited 2025 Nov 2];10(1). Available from: https://pubmed.ncbi.nlm.nih.gov/33836835/.
- Kayser S, Kramer M, Martínez-Cuadrón D, Grenet J, Metzeler KH, Sustkova Z, et al. Characteristics and outcome of patients with core-binding factor acute myeloid leukemia and FLT3-ITD: results from an international collaborative study. Haematologica [Internet]. 2022 Apr 1 [cited 2025 Nov 2];107(4):836–43. Available from: https://haematologica.org/article/view/haematol.2021.278645.
- Cortes JE, Khaled S, Martinelli G, Perl AE, Ganguly S, Russell N, et al. Quizartinib versus salvage chemotherapy in relapsed or refractory FLT3-ITD acute myeloid leukaemia (QuANTUM-R): a multicentre, randomised, controlled, open-label, phase 3 trial. Lancet Oncol [Internet]. 2019 Jul 1 [cited 2025 Nov 2];20(7):984–97. Available from: https://pubmed.ncbi.nlm.nih.gov/31175001/.
- Bazinet A, Bataller A, Kadia T, Daver N, Short NJ, Yilmaz M, et al. A retrospective study of outcomes across time and treatment regimens in newly diagnosed, FMS-like tyrosine kinase 3 (FLT3)-mutated acute myeloid leukemia. Cancer [Internet]. 2025 Mar 15 [cited 2025 Nov 2];131(6):e35813. Available from: /doi/pdf/10.1002/cncr.35813.
- Travaglini S, Gurnari C, Ottone T, Voso MT. Advances in the pathogenesis of FLT3 -mutated acute myeloid leukemia and targeted treatments. Curr Opin Oncol [Internet]. 2024 Nov 1 [cited 2025 Nov 6];36(6):569–76. Available from: https://journals.lww.com/co-oncology/fulltext/2024/11000/advances_in_the_pathogenesis_of_flt3_mutated_acute.15.aspx.
- Pommert L, Tarlock K. The evolution of targeted therapy in pediatric AML: gemtuzumab ozogamicin, FLT3/IDH/BCL2 inhibitors, and other therapies. Hematology [Internet]. 2022 Dec 9 [cited 2025 Nov 9];2022(1):603–10. Available from: https://dx.doi.org/10.1182/hematology.2022000358.
- Mrózek K, Bloomfield CD. Clinical Significance of the Most Common Chromosome Translocations in Adult Acute Myeloid Leukemia. J Natl Cancer Inst Monogr [Internet]. 2008 [cited 2025 Nov 4];(39). Available from: https://academic.oup.com/jncimono/article/2008/39/52/953332.
- Marcucci G, Geyer S, Laumann K, Zhao W, Bucci D, Uy GL, et al. Combination of dasatinib with chemotherapy in previously untreated core binding factor acute myeloid leukemia: CALGB 10801. Blood Adv [Internet]. 2020 Feb 25 [cited 2025 Nov 4];4(4):696. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC7042984/.
- Boissel N, Renneville A, Leguay T, Lefebvre PC, Recher C, Lecerf T, et al. Dasatinib in high-risk core binding factor acute myeloid leukemia in first complete remission: a French Acute Myeloid Leukemia Intergroup trial. Haematologica [Internet]. 2015 [cited 2025 Nov 4];100(6):780–5. Available from: https://pubmed.ncbi.nlm.nih.gov/25715404/.
- Hu Z, Tang X, Chen F, Li T, Liu Y, Zhou G, et al. Molecular genetics profiling of core-binding factor acute myeloid leukemia in pediatrics. Ther Adv Hematol [Internet]. 2025 Jan 1 [cited 2025 Nov 4];16. Available from: https://pubmed.ncbi.nlm.nih.gov/40290757/.
- Kantarjian HM, DiNardo CD, Kadia TM, Daver NG, Altman JK, Stein EM, et al. Acute myeloid leukemia management and research in 2025. CA Cancer J Clin [Internet]. 2025 Jan 1 [cited 2025 Nov 4];75(1):46–67. Available from: /doi/pdf/10.3322/caac.21873.
- (PDF) Rapid mutational analysis of N-ras proto-oncogene in hematologic malignancies: A study of 77 Greek patients [Internet]. [cited 2025 Nov 4]. Available from: https://www.researchgate.net/publication/11812910_Rapid_mutational_analysis_of_N-ras_proto-oncogene_in_hematologic_malignancies_A_study_of_77_Greek_patients.
- Alawieh D, Cysique-Foinlan L, Willekens C, Renneville A. RAS mutations in myeloid malignancies: revisiting old questions with novel insights and therapeutic perspectives. Blood Cancer J [Internet]. 2024 Dec 1 [cited 2025 Nov 4];14(1). Available from: https://pubmed.ncbi.nlm.nih.gov/38658558/.
- Desikan SP, Ravandi F, Pemmaraju N, Konopleva M, Loghavi S, Jabbour EJ, et al. A Phase II Study of Azacitidine, Venetoclax, and Trametinib in Relapsed or Refractory Acute Myeloid Leukemia Harboring RAS Pathway-Activating Mutations. Acta Haematol [Internet]. 2022 Sep 1 [cited 2025 Nov 4];145(5):529–36. Available from: https://pubmed.ncbi.nlm.nih.gov/35717939/.
- José-Enériz ES, Gimenez-Camino N, Agirre X, Prosper F. HDAC Inhibitors in Acute Myeloid Leukemia. Cancers (Basel) [Internet]. 2019 Nov 1 [cited 2025 Nov 6];11(11):1794. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC6896008/.
- Conneely SE, Quezada A, Kurtz KJ, Zhang N, Fuente JD La, Mercer N, et al. Cohesin haploinsufficiency is tolerated in Cbfb::MYH11-driven murine acute myeloid leukemia. Exp Hematol [Internet]. 2025 Oct [cited 2025 Nov 8];105287. Available from: https://pubmed.ncbi.nlm.nih.gov/41173205/.
- Martín-Izquierdo M, Abáigar M, Hernández-Sánchez JM, Tamborero D, López-Cadenas F, Ramos F, et al. Co-occurrence of cohesin complex and Ras signaling mutations during progression from myelodysplastic syndromes to secondary acute myeloid leukemia. Haematologica [Internet]. 2021 Aug 1 [cited 2025 Nov 8];106(8):2215–23. Available from: https://haematologica.org/article/view/9805.
- Tothova Z, Valton AL, Gorelov RA, Vallurupalli M, Krill-Burger JM, Holmes A, et al. Cohesin mutations alter DNA damage repair and chromatin structure and create therapeutic vulnerabilities in MDS/AML. JCI Insight [Internet]. 2021 Feb 8 [cited 2025 Nov 8];6(3). Available from: . [CrossRef]
- Redondo Monte E, Wilding A, Leubolt G, Kerbs P, Bagnoli JW, Hartmann L, et al. ZBTB7A prevents RUNX1-RUNX1T1-dependent clonal expansion of human hematopoietic stem and progenitor cells. Oncogene [Internet]. 2020 Apr 9 [cited 2025 Nov 8];39(15):3195. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC7142018/.
- Park SH, Lee HJ, Kim IS, Kang JE, Lee EY, Kim HJ, et al. Incidences and Prognostic Impact of c-KIT, WT1, CEBPA, and CBL Mutations, and Mutations Associated With Epigenetic Modification in Core Binding Factor Acute Myeloid Leukemia: A Multicenter Study in a Korean Population. Ann Lab Med [Internet]. 2015 May 1 [cited 2025 Nov 8];35(3):288–97. Available from: https://pubmed.ncbi.nlm.nih.gov/25932436/.
- Kreutmair S, Pfeifer D, Waterhouse M, Takács F, Graessel L, Döhner K, et al. First-in-human study of WT1 recombinant protein vaccination in elderly patients with AML in remission: a single-center experience. Cancer Immunol Immunother [Internet]. 2022 Dec 1 [cited 2025 Nov 8];71(12):2913–28. Available from: https://pubmed.ncbi.nlm.nih.gov/35476127/.
- Liu Yin JA, O’Brien MA, Hills RK, Daly SB, Wheatley K, Burnett AK. Minimal residual disease monitoring by quantitative RT-PCR in core binding factor AML allows risk stratification and predicts relapse: results of the United Kingdom MRC AML-15 trial. Blood [Internet]. 2012 Oct 4 [cited 2025 Nov 5];120(14):2826–35. Available from: https://dx.doi.org/10.1182/blood-2012-06-435669.
- Yoon JH, Kim H, Shin SH, Jeon YW, Kim JH, Lee SE, et al. Molecular and Cytogenetic Risk Stratification For Core-Binding Factor-Positive Adult AML With Analysis Of Post-Remission Treatment Outcomes Including Transplantation. Blood. 2013 Nov 15;122(21):1301–1301.
- Tarlock K, Gerbing RB, Ries RE, Smith JL, Leonti A, Huang BJ, et al. Prognostic impact of cooccurring mutations in FLT3-ITD pediatric acute myeloid leukemia. Blood Adv [Internet]. 2024 May 14 [cited 2025 Nov 7];8(9):2094–103. Available from: https://dx.doi.org/10.1182/bloodadvances.2023011980.
- Fan J, Gao L, Chen J, Hu S. Influence of KIT mutations on prognosis of pediatric patients with core-binding factor acute myeloid leukemia: a systematic review and meta-analysis. Transl Pediatr [Internet]. 2020 Dec 1 [cited 2025 Nov 4];9(6):72633–733. Available from: https://tp.amegroups.org/article/view/56363/html.
- Voso MT, Ottone T, Lavorgna S, Venditti A, Maurillo L, Lo-Coco F, et al. MRD in AML: The Role of New Techniques. Front Oncol [Internet]. 2019 Jul 23 [cited 2025 Nov 7];9:655. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC6664148/.
- Cilloni D, Renneville A, Hermitte F, Hills RK, Daly S, Jovanovic J V., et al. Real-time quantitative polymerase chain reaction detection of minimal residual disease by standardized WT1 assay to enhance risk stratification in acute myeloid leukemia: A European LeukemiaNet Study. J Clin Oncol [Internet]. 2009 Nov 1 [cited 2025 Nov 7];27(31):5195–201. Available from: https://ascopubs.org/doi/10.1200/JCO.2009.22.4865.
- Ivey A, Hills RK, Simpson MA, Jovanovic J V., Gilkes A, Grech A, et al. Assessment of Minimal Residual Disease in Standard-Risk AML. N Engl J Med [Internet]. 2016 Feb 4 [cited 2025 Nov 7];374(5):422–33. Available from: https://www.nejm.org/doi/full/10.1056/NEJMoa1507471.
- Schlenk RF, Kayser S, Bullinger L, Kobbe G, Casper J, Ringhoffer M, et al. Differential impact of allelic ratio and insertion site in FLT3-ITD-positive AML with respect to allogeneic transplantation. Blood [Internet]. 2014 Sep 30 [cited 2025 Nov 7];124(23):3441–9. Available from: https://pubmed.ncbi.nlm.nih.gov/25270908/.
- Li Y, Solis-Ruiz J, Yang F, Long N, Tong CH, Lacbawan FL, et al. NGS-defined measurable residual disease (MRD) after initial chemotherapy as a prognostic biomarker for acute myeloid leukemia. Blood Cancer J 2023 131 [Internet]. 2023 Apr 24 [cited 2025 Nov 7];13(1):1–9. Available from: https://www.nature.com/articles/s41408-023-00833-7.
- Svaton M, Skotnicova A, Reznickova L, Rennerova A, Valova T, Kotrova M, et al. NGS-Based MRD Quantitation: An Alternative to qPCR Validated on a Large Consecutive Cohort of Children with ALL. Blood. 2021 Nov 23;138(Supplement 1):1314.
- Yao Q, Bai Y, Kumar S, Au E, Orfao A, Chim CS. Minimal Residual Disease Detection by Next-Generation Sequencing in Multiple Myeloma: A Comparison With Real-Time Quantitative PCR. Front Oncol. 2021 Jan 29;10:611021.
- Puckrin R, Atenafu EG, Claudio JO, Chan S, Gupta V, Maze D, et al. Measurable residual disease monitoring provides insufficient lead-time to prevent morphological relapse in the majority of patients with core-binding factor acute myeloid leukemia. Haematologica [Internet]. 2020 Jan 1 [cited 2025 Nov 7];106(1):56. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC7776265/.
- Puckrin R, Atenafu EG, Claudio JO, Chan S, Gupta V, Maze D, et al. Measurable residual disease monitoring provides insufficient lead-time to prevent morphologic relapse in the majority of patients with core-binding factor acute myeloid leukemia. Haematologica [Internet]. 2021 Jan 1 [cited 2025 Nov 7];106(1):56–63. Available from: https://haematologica.org/article/view/9590.
- Heuser M, Freeman SD, Ossenkoppele GJ, Buccisano F, Hourigan CS, Ngai LL, et al. 2021 Update on MRD in acute myeloid leukemia: a consensus document from the European LeukemiaNet MRD Working Party. Blood [Internet]. 2021 Dec 30 [cited 2025 Nov 13];138(26):2753–67. Available from: https://dx.doi.org/10.1182/blood.2021013626.
- Onecha E, Linares M, Rapado I, Ruiz-Heredia Y, Martinez-Sanchez P, Cedena T, et al. A novel deep targeted sequencing method for minimal residual disease monitoring in acute myeloid leukemia. Haematologica [Internet]. 2018 Jan 31 [cited 2025 Nov 13];104(2):288. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC6355493/.
- Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, et al. Genomic Classification and Prognosis in Acute Myeloid Leukemia. N Engl J Med [Internet]. 2016 Jun 9 [cited 2025 Nov 5];374(23):2209–21. Available from: https://www.nejm.org/doi/full/10.1056/NEJMoa1516192.
- Cioccio J, Claxton D. Therapy of acute myeloid leukemia: therapeutic targeting of tyrosine kinases. Expert Opin Investig Drugs [Internet]. 2019 Apr 3 [cited 2025 Nov 5];28(4):337–49. Available from: https://pubmed.ncbi.nlm.nih.gov/30775933/.
- Arock M, Wedeh G, Hoermann G, Bibi S, Akin C, Peter B, et al. Preclinical human models and emerging therapeutics for advanced systemic mastocytosis. Haematologica [Internet]. 2018 Oct 31 [cited 2025 Nov 5];103(11):1760. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC6278969/.
- Yokoyama A, Somervaille TCP, Smith KS, Rozenblatt-Rosen O, Meyerson M, Cleary ML. The menin tumor suppressor protein is an essential oncogenic cofactor for MLL-associated leukemogenesis. Cell [Internet]. 2005 Oct 21 [cited 2025 Nov 5];123(2):207–18. Available from: https://pubmed.ncbi.nlm.nih.gov/16239140/.
- Krivtsov A V., Evans K, Gadrey JY, Eschle BK, Hatton C, Uckelmann HJ, et al. A Menin-MLL Inhibitor Induces Specific Chromatin Changes and Eradicates Disease in Models of MLL-Rearranged Leukemia. Cancer Cell [Internet]. 2019 Dec 9 [cited 2025 Nov 5];36(6):660-673.e11. Available from: https://pubmed.ncbi.nlm.nih.gov/31821784/.
- Jurcic JG. What happened to anti-CD33 therapy for acute myeloid Leukemia? Curr Hematol Malig Rep [Internet]. 2012 Mar 23 [cited 2025 Nov 5];7(1):65–73. Available from: https://link.springer.com/article/10.1007/s11899-011-0103-0.
- Swaminathan M, Cortes JE. Update on the role of gemtuzumab-ozogamicin in the treatment of acute myeloid leukemia. Ther Adv Hematol [Internet]. 2023 Jan 1 [cited 2025 Nov 5];14. Available from: https://journals.sagepub.com/doi/full/10.1177/20406207231154708.
- Ronnacker J, Muller PJ, Mikesch JH, Zukunft S, Weinbergerová B, Šrámek J, et al. Gemtuzumab ozogamicin in first-line treatment of CBF-AML: insights from a retrospective multi-center analysis. Leuk 2025 399 [Internet]. 2025 Jul 21 [cited 2025 Nov 5];39(9):2174–80. Available from: https://www.nature.com/articles/s41375-025-02700-9.
- Khan N, Hills RK, Virgo P, Couzens S, Clark N, Gilkes A, et al. Expression of CD33 is a predictive factor for effect of gemtuzumab ozogamicin at different doses in adult acute myeloid leukaemia. Leuk 2017 315 [Internet]. 2016 Oct 31 [cited 2025 Nov 5];31(5):1059–68. Available from: https://www.nature.com/articles/leu2016309.
- Hills RK, Castaigne S, Appelbaum FR, Delaunay J, Petersdorf S, Othus M, et al. Addition of gemtuzumab ozogamicin to induction chemotherapy in adult patients with acute myeloid leukaemia: A meta-analysis of individual patient data from randomised controlled trials. Lancet Oncol [Internet]. 2014 Aug 1 [cited 2025 Nov 5];15(9):986–96. Available from: https://www.thelancet.com/action/showFullText?pii=S1470204514702815.
- Castaigne S, Pautas C, Terré C, Raffoux E, Bordessoule D, Bastie JN, et al. Effect of gemtuzumab ozogamicin on survival of adult patients with de-novo acute myeloid leukaemia (ALFA-0701): A randomised, open-label, phase 3 study. Lancet [Internet]. 2012 Apr 21 [cited 2025 Nov 5];379(9825):1508–16. Available from: https://www.thelancet.com/action/showFullText?pii=S0140673612604851.
- Amadori S, Venditti A, Voso MT, Annino L, De Fabritiis P, Alimena G, et al. Gemtuzumab ozogamicin versus best supportive care in older patients with newly diagnosed acute myeloid leukemia unsuitable for intensive chemotherapy: Results of the randomized phase III EORTC-GIMEMA AML-19 Trial. J Clin Oncol [Internet]. 2016 Mar 20 [cited 2025 Nov 5];34(9):972–9. Available from: https://ascopubs.org/doi/10.1200/JCO.2015.64.0060.
- Kiyoi H. FLT3 INHIBITORS: RECENT ADVANCES AND PROBLEMS FOR CLINICAL APPLICATION. Nagoya J Med Sci [Internet]. 2015 [cited 2025 Nov 5];77(1–2):7. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC4361503/.
- Shah V, Shah A, Jain P, Panchal V, Fernandes BCA, Gangasani S, et al. AML-1203: Efficacy of Gilteritinib Versus Salvage Chemotherapy in Relapsed or Refractory FLT3-Mutated Acute Myeloid Leukemia: A Meta-Analysis. Clin Lymphoma Myeloma Leuk. 2025 Sep 1;25:S492–3.
- Pratz KW, Cherry M, Altman JK, Cooper BW, Podoltsev NA, Cruz JC, et al. Gilteritinib in Combination with Induction and Consolidation Chemotherapy and as Maintenance Therapy: A Phase IB Study in Patients with Newly Diagnosed AML. J Clin Oncol [Internet]. 2023 Sep 10 [cited 2025 Nov 6];41(26):4236–46. Available from: https://ascopubs.org/doi/10.1200/JCO.22.02721.
- Wang ES, Goldberg AD, Tallman M, Walter RB, Karanes C, Sandhu K, et al. Crenolanib and Intensive Chemotherapy in Adults With Newly Diagnosed FLT3-Mutated AML. J Clin Oncol [Internet]. 2024 May 20 [cited 2025 Nov 6];42(15):1776–87. Available from: https://ascopubs.org/doi/pdf/10.1200/JCO.23.01061.
- Garciaz S, Hospital MA. FMS-Like Tyrosine Kinase 3 Inhibitors in the Treatment of Acute Myeloid Leukemia: An Update on the Emerging Evidence and Safety Profile. Onco Targets Ther [Internet]. 2023 [cited 2025 Nov 6];16:31. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC9869913/.
- Bazarbachi A, Labopin M, Battipaglia G, Djabali A, Passweg J, Socié G, et al. Sorafenib improves survival of FLT3-mutated acute myeloid leukemia in relapse after allogeneic stem cell transplantation: a report of the EBMT Acute Leukemia Working Party. Haematologica [Internet]. 2019 Sep 1 [cited 2025 Nov 6];104(9):e398–401. Available from: https://haematologica.org/article/view/9059.
- Morin S, Giannotti F, Mamez AC, Pradier A, Masouridi-Levrat S, Simonetta F, et al. Real-world experience of sorafenib maintenance after allogeneic hematopoietic stem cell transplantation for FLT3-ITD AML reveals high rates of toxicity-related treatment interruption. Front Oncol [Internet]. 2023 [cited 2025 Nov 6];13:1095870. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC10050716/.
- Yamaura T, Nakatani T, Uda K, Ogura H, Shin W, Kurokawa N, et al. A novel irreversible FLT3 inhibitor, FF-10101, shows excellent efficacy against AML cells with FLT3 mutations. Blood [Internet]. 2018 Jan 25 [cited 2025 Nov 6];131(4):426–38. Available from: https://dx.doi.org/10.1182/blood-2017-05-786657.
- Ferng TT, Terada D, Ando M, Tarver TC, Chaudhary F, Lin KC, et al. The irreversible FLT3 inhibitor FF-10101 is active against a diversity of FLT3 inhibitor resistance mechanisms. Mol Cancer Ther [Internet]. 2022 May 1 [cited 2025 Nov 6];21(5):844. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC9081245/.
- Beghini A, Beghini A. Core Binding Factor Leukemia: Chromatin Remodeling Moves Towards Oncogenic Transcription. Cancers 2019, Vol 11, [Internet]. 2019 Dec 7 [cited 2025 Nov 6];11(12). Available from: https://www.mdpi.com/2072-6694/11/12/1973.
- Zhang J, Gao X, Yu L. Roles of Histone Deacetylases in Acute Myeloid Leukemia With Fusion Proteins. Front Oncol [Internet]. 2021 Sep 1 [cited 2025 Nov 6];11:741746. Available from: www.frontiersin.org.
- Yu X, Li H, Hu P, Qing Y, Wang X, Zhu M, et al. Natural HDAC-1/8 inhibitor baicalein exerts therapeutic effect in CBF-AML. Clin Transl Med [Internet]. 2020 Aug 1 [cited 2025 Nov 6];10(4):e154. Available from: https://onlinelibrary.wiley.com/doi/full/10.1002/ctm2.154.
- Kuykendall A, Duployez N, Boissel N, Lancet JE, Welch JS. Acute Myeloid Leukemia: The Good, the Bad, and the Ugly. Am Soc Clin Oncol Educ B [Internet]. 2018 May [cited 2025 Nov 6];(38):555–73. Available from: https://ascopubs.org/doi/10.1200/EDBK_199519.
- Vanegas YM, Badar T. Clinical Utility of Azacitidine in the Management of Acute Myeloid Leukemia: Update on Patient Selection and Reported Outcomes. Cancer Manag Res [Internet]. 2022 Dec 1 [cited 2025 Nov 6];14:3527. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC9793740/.
- Gabellier L, Peterlin P, Thepot S, Hicheri Y, Paul F, Gallego-Hernanz MP, et al. Hypomethylating agent monotherapy in core binding factor acute myeloid leukemia: a French multicentric retrospective study. Ann Hematol [Internet]. 2024 Mar 1 [cited 2025 Nov 6];103(3):759. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC10867066/.
- Zhang K, Zhang X, Xu Y, Xue S, Qiu H, Tang X, et al. Efficacy of venetoclax combined with hypomethylating agents in young, and unfit patients with newly diagnosed core binding factor acute myeloid leukemia. Blood Cancer J 2023 131 [Internet]. 2023 Oct 11 [cited 2025 Nov 6];13(1):1–4. Available from: https://www.nature.com/articles/s41408-023-00928-1.
- Santini V, Lübbert M, Wierzbowska A, Ossenkoppele GJ. The Clinical Value of Decitabine Monotherapy in Patients with Acute Myeloid Leukemia. Adv Ther [Internet]. 2021 Apr 1 [cited 2025 Nov 6];39(4):1474. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC8989816/.
- Bhattacharya S, Banerjee A, Dolai TK, Sadhukhan SK, Med IJ, Oncol P. Role of Maintenance Therapy with Decitabine in Acute Myeloid Leukemia in First Remission: A Prospective Interventional Study from a Tertiary Care Center in Eastern India. 2025 [cited 2025 Nov 6]; Available from: .
- Arslan S, Zhang J, Dhakal P, Moran JA, Naidoo N, Lombardi Story J, et al. Outcomes of Therapy with Venetoclax Combined with Hypomethylating Agents in Favorable-Risk Acute Myeloid Leukemia (AML). Blood [Internet]. 2020 Nov 5 [cited 2025 Nov 6];136(Supplement 1):41–2. Available from: https://dx.doi.org/10.1182/blood-2020-142780.
- He S, Li Y, Shi X, Wang L, Cai D, Zhou J, et al. DNA methylation landscape reveals LIN7A as a decitabine-responsive marker in patients with t(8;21) acute myeloid leukemia. Clin Epigenetics [Internet]. 2023 Dec 1 [cited 2025 Nov 6];15(1):37. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC9983225/.
- Imai Y, Maru Y, Tanaka J. Action mechanisms of histone deacetylase inhibitors in the treatment of hematological malignancies. Cancer Sci [Internet]. 2016 Nov 1 [cited 2025 Nov 6];107(11):1543. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC5132279/.
- Salmon JM, Bots M, Vidacs E, Stanley KL, Atadja P, Zuber J, et al. Combining the differentiating effect of panobinostat with the apoptotic effect of arsenic trioxide leads to significant survival benefit in a model of t(8;21) acute myeloid leukemia. Clin Epigenetics [Internet]. 2015 Jan 22 [cited 2025 Nov 7];7(1):2. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC4308003/.
- Bots M, Verbrugge I, Martin BP, Salmon JM, Ghisi M, Baker A, et al. Differentiation therapy for the treatment of t(8;21) acute myeloid leukemia using histone deacetylase inhibitors. Blood [Internet]. 2014 Feb 27 [cited 2025 Nov 7];123(9):1341. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC3938147/.
- Garcia-Manero G, Podoltsev NA, Othus M, Pagel JM, Radich JP, Fang M, et al. Standard versus high-dose cytarabine with or without vorinostat for AML. Leukemia [Internet]. 2023 [cited 2025 Nov 7];38(1):58. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC11729399/.
- Silva G, Cardoso BA, Belo H, Almeida AM. Vorinostat Induces Apoptosis and Differentiation in Myeloid Malignancies: Genetic and Molecular Mechanisms. PLoS One [Internet]. 2013 Jan 8 [cited 2025 Nov 7];8(1):e53766. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC3540071/.
- Collados-Ros A, Muro M, Legaz I. Gemtuzumab Ozogamicin in Acute Myeloid Leukemia: Efficacy, Toxicity, and Resistance Mechanisms—A Systematic Review. Biomedicines [Internet]. 2024 Jan 1 [cited 2025 Nov 13];12(1):208. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC10813452/.
- Travaglini S, Gurnari C, Ottone T, Voso MT. Advances in the pathogenesis of FLT3 -mutated acute myeloid leukemia and targeted treatments. Curr Opin Oncol [Internet]. 2024 Nov 1 [cited 2025 Nov 13];36(6):569–76. Available from: https://journals.lww.com/co-oncology/fulltext/2024/11000/advances_in_the_pathogenesis_of_flt3_mutated_acute.15.aspx.
- José-Enériz ES, Gimenez-Camino N, Agirre X, Prosper F, José-Enériz ES, Gimenez-Camino N, et al. HDAC Inhibitors in Acute Myeloid Leukemia. Cancers 2019, Vol 11, [Internet]. 2019 Nov 14 [cited 2025 Nov 13];11(11). Available from: https://www.mdpi.com/2072-6694/11/11/1794.
- Wang T, Chen S, Chen J, Liu T, Zhang T, Qiu H, et al. Allogeneic Hematopoietic Stem Cell Transplantation Improved Survival for Adult Core Binding Factor Acute Myelogenous Leukemia Patients with Intermediate- and Adverse-Risk Genetics in the 2017 European LeukemiaNet. Transplant Cell Ther [Internet]. 2021 Feb 1 [cited 2025 Nov 7];27(2):173.e1-173.e9. Available from: https://pubmed.ncbi.nlm.nih.gov/33830030/.
- Halaburda K, Labopin M, Mailhol A, Socié G, Craddock C, Aljurf M, et al. Allogeneic stem cell transplantation in second complete remission for core binding factor acute myeloid leukemia: a study from the Acute Leukemia Working Party of the European Society for Blood and Marrow Transplantation. Haematologica [Internet]. 2020 Jun 1 [cited 2025 Nov 7];105(6):1723–30. Available from: https://haematologica.org/article/view/9449.
- Kawashima N, Ishikawa Y, Atsuta Y, Sugiura I, Sawa M, Dobashi N, et al. Prospective Evaluation of Prognostic Relevance of KIT Mutations in Core-Binding Factor Acute Myeloid Leukemia: Results from the JALSG CBF-AML209-KIT Study. Blood [Internet]. 2018 Nov 29 [cited 2025 Nov 7];132(Supplement 1):438–438. Available from: https://dx.doi.org/10.1182/blood-2018-99-113931.
- Duan W, Liu X, JIA J, Wang J, Gong L, Jiang Q, et al. Both the Subtypes of Kit Mutation and Minimal Residual Disease Are Associated with Prognosis in Core Banding Factor Acute Myeloid Leukemia. Blood [Internet]. 2020 Nov 5 [cited 2025 Nov 7];136(Supplement 1):4–5. Available from: https://dx.doi.org/10.1182/blood-2020-141905.
- Jahn N, Terzer T, Sträng E, Dolnik A, Cocciardi S, Panina E, et al. Genomic heterogeneity in core-binding factor acute myeloid leukemia and its clinical implication. Blood Adv [Internet]. 2020 Dec 22 [cited 2025 Nov 2];4(24):6342. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC7757000/.
- Quan X, Deng J. Core binding factor acute myeloid leukemia: Advances in the heterogeneity of KIT, FLT3, and RAS mutations (Review). Mol Clin Oncol [Internet]. 2020 [cited 2025 Nov 2];13(2):95. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC7366242/.
- Cornelissen JJ, Blaise D. Hematopoietic stem cell transplantation for patients with AML in first complete remission. Blood [Internet]. 2016 Jan 7 [cited 2025 Nov 7];127(1):62–70. Available from: https://pubmed.ncbi.nlm.nih.gov/26660427/.
- Al Hamed R, Labopin M, Wu D, Gedde-Dahl T, Aljurf M, Forcade E, et al. Allogeneic stem cell transplantation in de novo core-binding factor acute myeloid leukemia in first complete remission: data from the EBMT. Bone Marrow Transplant [Internet]. 2024 Oct 1 [cited 2025 Nov 7];59(10):1458–65. Available from: https://pubmed.ncbi.nlm.nih.gov/39095548/.
| Gene/Mutation Class | Prevalence (Overall CBF-AML) | Prognostic Impact in t(8;21) (RUNX1::RUNX1T1) | Prognostic Impact in inv(16) (CBFB::MYH11) | Key References |
| KIT (Exon 17 D816/N822) | 20%–45% (Higher in t(8;21)) | Highly Adverse (Increased relapse risk, shorter EFS/OS). Guide for Allo-HSCT in CR1. | Less Consistent/Less Significant Adverse Impact | [34,38,39,40] |
| FLT3-ITD/TKD | Detected in a subset; 4%-11% (Less common than NK-AML) | Adverse (Independent factor for poorer survival/relapse) | Adverse (Predictive of short progression-free survival (PFS)) | [41] |
| RAS (NRAS/KRAS) | Frequent (Overall 15-20%); Highest in inv(16) (up to 36%) | Prognosis varies; potentially favorable when isolated (no RTK mutations) | Higher prevalence; outcome influenced by co-mutations/allelic ratio | [42,43,44] |
| ASXL2 | Frequent in t(8;21) (22%-23%) | No statistically significant independent adverse prognosis demonstrated to date | Less Common; Role not defined | [45,46] |
| ZBTB7A | Primarily in t(8;21) (up to 23%) | Implicated in leukemogenesis; clinical utility/prognosis not established | Rare/Absent | [47,48] |
| Methodology | Primary Target(s) | Typical Sensitivity (LOD) | Key Advantages | Limitations in CBF-AML | References |
| RT-qPCR (Real-Time Quantitative PCR) | Fusion transcripts (RUNX1::RUNX1T1, CBFB::MYH11) | High (10-4 to 10-6) | Highly specific, standardized, and cost-effective; established prognostic marker. | Limited to tracking only the fusion transcript; fails to capture clonal evolution of secondary mutations. | [71,84] |
| NGS (Next-Generation Sequencing) | Co-occurring somatic mutations (e.g., KIT, FLT3, RAS, cohesin) & Fusion transcripts (DNA-based) | Moderate to High (10-4 to 10-5 for SNVs/Indels) | Comprehensive monitoring of multiple mutations and clonal evolution; detects preleukemic persistence. | Lower sensitivity than optimized qPCR for fusion transcripts; lack of standardized thresholds; complex bioinformatics. | [85] |
| Flow Cytometry (MFC) (Multi-parameter Flow Cytometry) | Aberrant immunophenotypes (e.g., CD19 co-expression in t(8;21)) | Moderate 10-3 to 10-4) | Rapid, can be applied to many AML subtypes, widely available. | Requires sufficient aberrant markers; interpretation can be challenging in monocytic subtypes (inv(16) M4Eo). | [85] |
| Agent Class (Example Agent) | Mechanism of Action | Relevance/Role in CBF-AML | Clinical Status/Key Findings | Key References |
| Anti-CD33 ADC (Gemtuzumab Ozogamicin - GO) | Monoclonal antibody-drug conjugate; targeted delivery of calicheamicin toxin to CD33+ cells upon internalization. | Frontline therapy; high CD33 expression and low Multi-Drug Resistance (MDR) in CBF-AML. | Significantly improves OS/EFS, reducing relapse; established standard of care in favorable-risk AML. | [123] |
| FLT3 Inhibitors (Midostaurin, Gilteritinib) | Blocks ligand-independent activation of FLT3 Receptor Tyrosine Kinase (RTK). | Targets adverse FLT3-ITD/TKD mutations associated with poor outcomes. | Used widely in FLT3-mutated AML; specific benefit in CBF-AML requires focused investigation due to mixed results. | [51,98,100,124] |
| KIT Inhibitors (Dasatinib) | Blocks KIT RTK constitutive signaling. | Targets adverse KIT mutations, particularly in the t(8;21) subtype. | Phase III studies failed to show clear EFS/OS benefit; therapeutic challenge remains despite prognostic significance. | [19,57,87] |
| DNMT Inhibitors (Azacitidine, Decitabine) | Restores gene expression by inhibiting DNA Methyltransferases (e.g., LIN7A reactivation). | Used in unfit/older patients and maintenance therapy. Potential for synergy (e.g., with Venetoclax). | AZA/VEN response superior in inv(16) vs. t(8;21); Decitabine promising for maintenance, guided by molecular markers. | [113,114,117] |
| HDAC Inhibitors (Panobinostat, Vorinostat) | Targets Histone Deacetylases, restoring acetylation and gene expression; promotes fusion protein degradation. | Potential for combination synergy with IC or DNMTi; robust preclinical activity in t(8;21) models. | Limited clinical monotherapy efficacy; require randomized trials in combination regimens tailored to specific CBF subtypes. | [118,120,122,125] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




