Submitted:
26 January 2026
Posted:
26 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
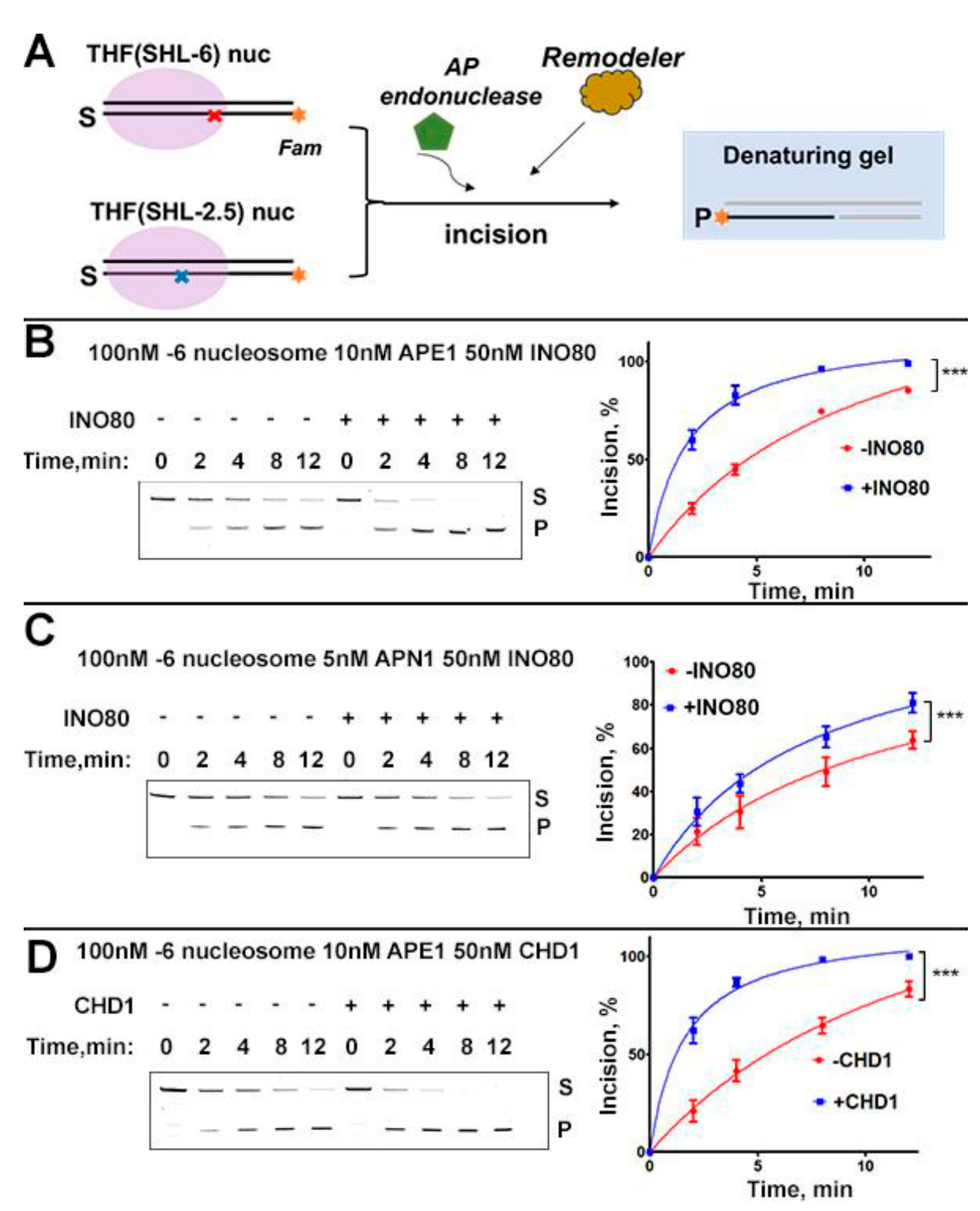
2.1. INO80 Stimulation of AP-Site Incision on Nucleosome Is Conserved in Yeast
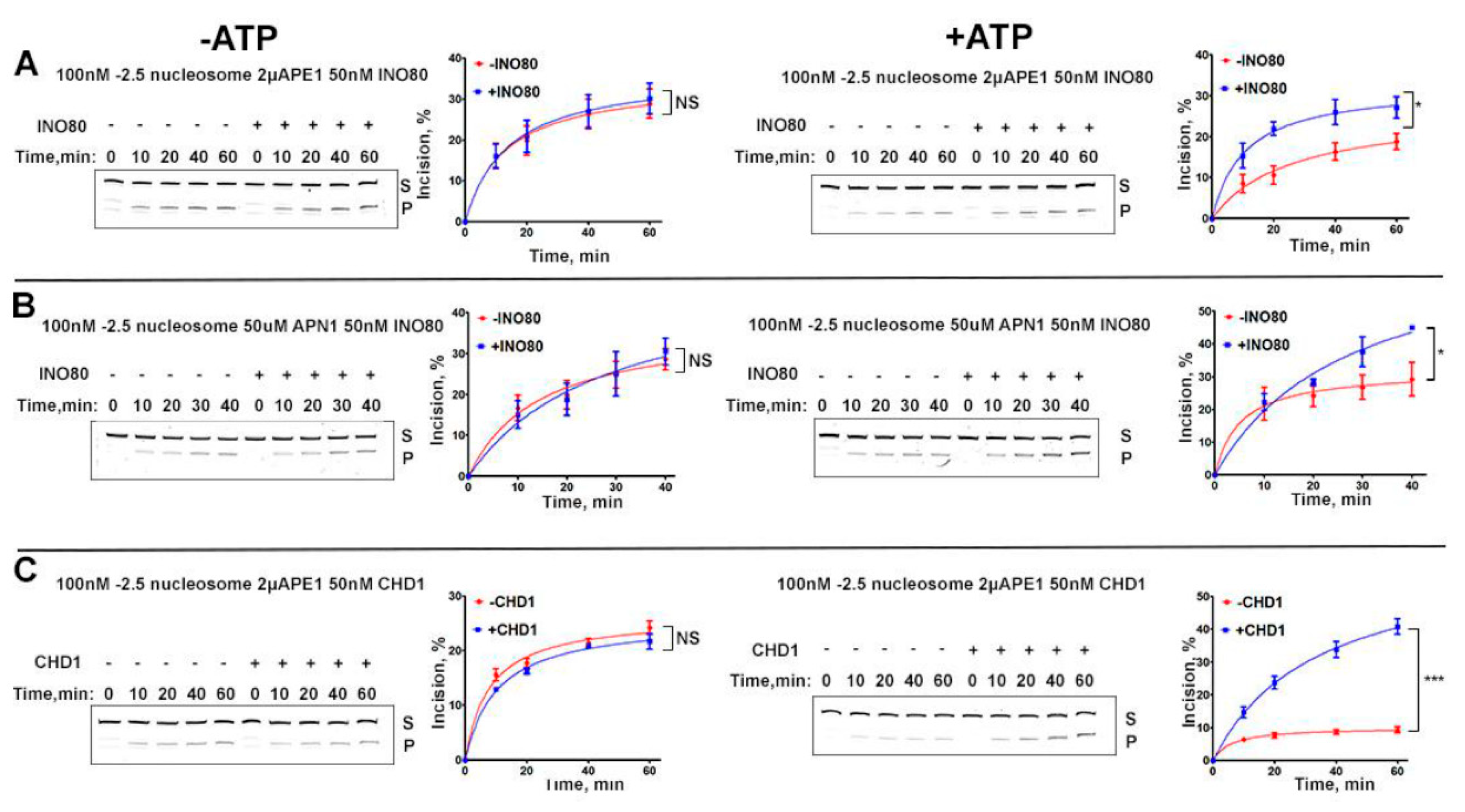
2.2. Efficient Incision of Internal Nucleosome AP-Site Requires Chromatin Remodeler and ATP
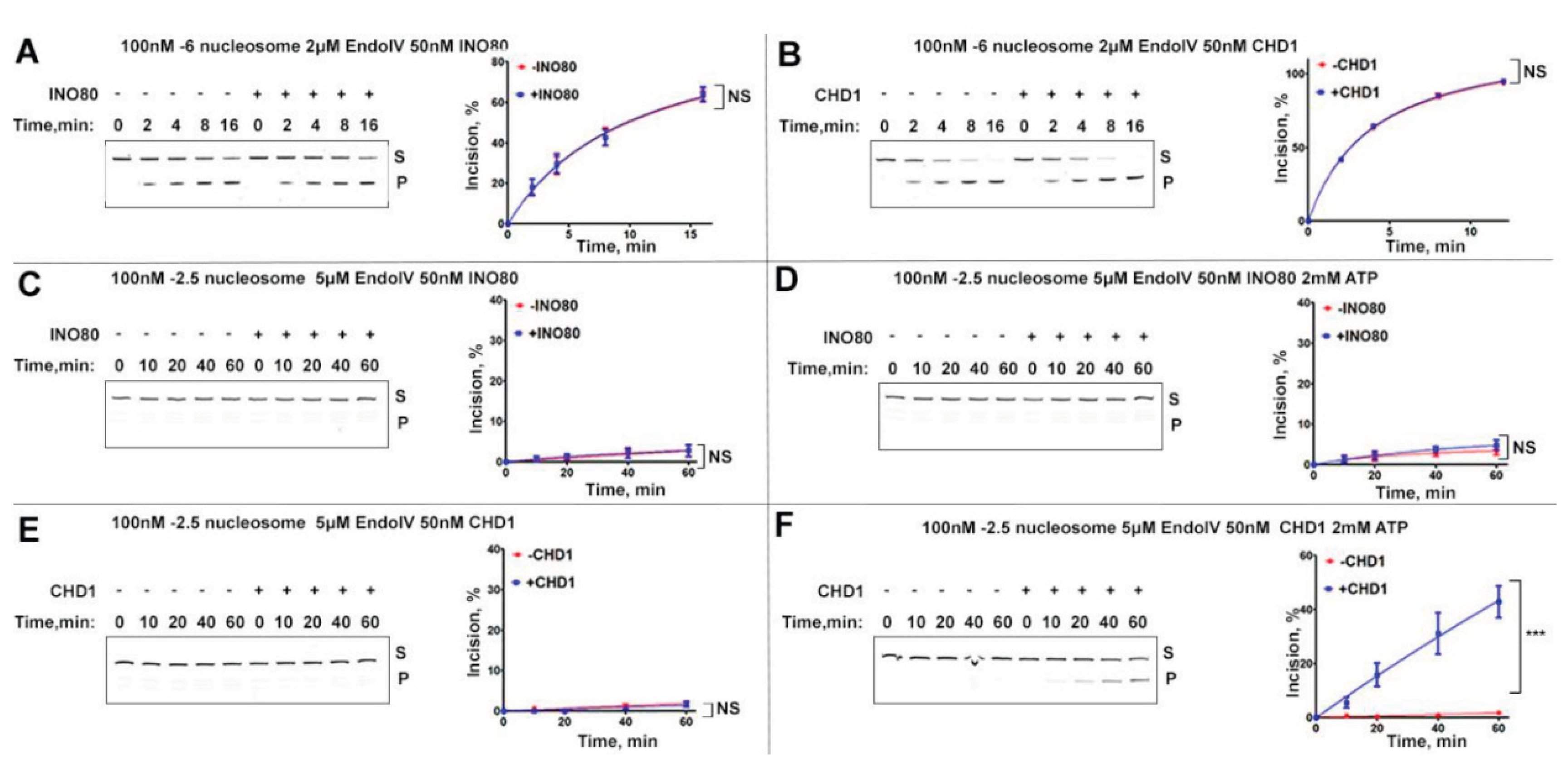
2.3. Endo IV Is Ineffective in Incising Internal AP-Sites on Nucleosomes Without Chromatin Remodelers
3. Discussion
4. Materials and Methods
4.1. Expression of Recombinant Chromatin Remodelers
4.2. Histone Purification and Octamer Reconstitution
4.3. APN1 Purification
4.4. APE1 Purification
4.5. DNA
4.5.1. Purification of 0N80 DNA
4.5.2. Separation of 0N80Double Strand DNA into Its Single Strands
4.5.3. Fluorescent Damaged DNA Preparation
4.5.4. Nucleosome Reconstitutions
4.6. AP-Endonuclease Incision Assay
4.7. Nucleosome Sliding Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- David, S.S.; O'Shea, V.L.; Kundu, S. Base-excision repair of oxidative DNA damage. Nature 2007, 447, 941–950. [Google Scholar] [CrossRef]
- Girard, P.M.; Guibourt, N.; Boiteux, S. The Ogg1 protein of Saccharomyces cerevisiae: A 7,8-dihydro-8-oxoguanine DNA glycosylase/AP lyase whose lysine 241 is a critical residue for catalytic activity. Nucleic Acids Res 1997, 25, 3204–3211. [Google Scholar] [CrossRef]
- Jacobs, A.L.; Schar, P. DNA glycosylases: In DNA repair and beyond. Chromosoma 2012, 121, 1–20. [Google Scholar] [CrossRef] [PubMed]
- van der Kemp, P.A.; Thomas, D.; Barbey, R.; de Oliveira, R.; Boiteux, S. Cloning and expression in Escherichia coli of the OGG1 gene of Saccharomyces cerevisiae, which codes for a DNA glycosylase that excises 7,8-dihydro-8-oxoguanine and 2,6-diamino-4-hydroxy-5-N-methylformamidopyrimidine. Proc Natl Acad Sci U S A 1996, 93, 5197–5202. [Google Scholar] [CrossRef]
- Xanthoudakis, S.; Smeyne, R.J.; Wallace, J.D.; Curran, T. The redox/DNA repair protein, Ref-1, is essential for early embryonic development in mice. Proc Natl Acad Sci U S A 1996, 93, 8919–8923. [Google Scholar] [CrossRef] [PubMed]
- Sudhakar, J.; Khetan, V.; Madhusudan, S.; Krishnakumar, S. Dysregulation of human apurinic/apyrimidinic endonuclease 1 (APE1) expression in advanced retinoblastoma. Br J Ophthalmol 2014, 98, 402–407. [Google Scholar] [CrossRef]
- Dumitrache, L.C.; Shimada, M.; Downing, S.M.; Kwak, Y.D.; Li, Y.; Illuzzi, J.L.; Russell, H.R.; Wilson, D.M., 3rd; McKinnon, P.J. Apurinic endonuclease-1 preserves neural genome integrity to maintain homeostasis and thermoregulation and prevent brain tumors. Proc Natl Acad Sci U S A 2018, 115, E12285–E12294. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Zhao, J.; Talluri, S.; Buon, L.; Mu, S.; Potluri, L.B.; Liao, C.; Shi, J.; Chakraborty, C.; Gonzalez, G.B.; et al. Elevated APE1 Dysregulates Homologous Recombination and Cell Cycle Driving Genomic Evolution, Tumorigenesis, and Chemoresistance in Esophageal Adenocarcinoma. Gastroenterology 2023, 165, 357–373. [Google Scholar] [CrossRef]
- Bhakat, K.K.; Mantha, A.K.; Mitra, S. Transcriptional regulatory functions of mammalian AP-endonuclease (APE1/Ref-1), an essential multifunctional protein. Antioxid Redox Signal 2009, 11, 621–638. [Google Scholar] [CrossRef]
- Antoniali, G.; Dalla, E.; Mangiapane, G.; Zhao, X.; Jing, X.; Cheng, Y.; De Sanctis, V.; Ayyildiz, D.; Piazza, S.; Li, M.; et al. APE1 controls DICER1 expression in NSCLC through miR-33a and miR-130b. Cell Mol Life Sci 2022, 79, 446. [Google Scholar] [CrossRef]
- Antoniali, G.; Serra, F.; Lirussi, L.; Tanaka, M.; D'Ambrosio, C.; Zhang, S.; Radovic, S.; Dalla, E.; Ciani, Y.; Scaloni, A.; et al. Mammalian APE1 controls miRNA processing and its interactome is linked to cancer RNA metabolism. Nat Commun 2017, 8, 797. [Google Scholar] [CrossRef]
- Codrich, M.; Degrassi, M.; Malfatti, M.C.; Antoniali, G.; Gorassini, A.; Ayyildiz, D.; De Marco, R.; Verardo, G.; Tell, G.; et al. APE1 interacts with the nuclear exosome complex protein MTR4 and is involved in cisplatin- and 5-fluorouracil-induced RNA damage response. FEBS J 2023, 290, 1740–1764. [Google Scholar] [CrossRef] [PubMed]
- Vascotto, C.; Fantini, D.; Romanello, M.; Cesaratto, L.; Deganuto, M.; Leonardi, A.; Radicella, J.P.; Kelley, M.R.; D'Ambrosio, C.; Scaloni, A.; et al. APE1/Ref-1 interacts with NPM1 within nucleoli and plays a role in the rRNA quality control process. Mol Cell Biol 2009, 29, 1834–1854. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Wang, X.; Bao, S.; Guo, R.; Johnson, D.G.; Shen, X.; Li, L. INO80 chromatin remodeling complex promotes the removal of UV lesions by the nucleotide excision repair pathway. Proc Natl Acad Sci U S A 2010, 107, 17274–17279. [Google Scholar] [CrossRef] [PubMed]
- Kothandapani, A.; Gopalakrishnan, K.; Kahali, B.; Reisman, D.; Patrick, S.M. Downregulation of SWI/SNF chromatin remodeling factor subunits modulates cisplatin cytotoxicity. Exp Cell Res 2012, 318, 1973–1986. [Google Scholar] [CrossRef]
- Hara, R.; Sancar, A. The SWI/SNF chromatin-remodeling factor stimulates repair by human excision nuclease in the mononucleosome core particle. Mol Cell Biol 2002, 22, 6779–6787. [Google Scholar] [CrossRef]
- Ruthemann, P.; Balbo Pogliano, C.; Codilupi, T.; Garajova, Z.; Naegeli, H. Chromatin remodeler CHD1 promotes XPC-to-TFIIH handover of nucleosomal UV lesions in nucleotide excision repair. EMBO J 2017, 36, 3372–3386. [Google Scholar] [CrossRef]
- Sokolova, V.; Lee, G.; Mullins, A.; Mody, P.; Watanabe, S.; Tan, D. DNA-translocation-independent role of INO80 remodeler in DNA damage repairs. J Biol Chem 2023, 299, 105245. [Google Scholar] [CrossRef]
- Ayala, R.; Willhoft, O.; Aramayo, R.J.; Wilkinson, M.; McCormack, E.A.; Ocloo, L.; Wigley, D.B.; Zhang, X. Structure and regulation of the human INO80-nucleosome complex. Nature 2018, 556, 391–395. [Google Scholar] [CrossRef]
- Eustermann, S.; Schall, K.; Kostrewa, D.; Lakomek, K.; Strauss, M.; Moldt, M.; Hopfner, K.P. Structural basis for ATP-dependent chromatin remodelling by the INO80 complex. Nature 2018, 556, 386–390. [Google Scholar] [CrossRef]
- Torigoe, S.E.; Patel, A.; Khuong, M.T.; Bowman, G.D.; Kadonaga, J.T. ATP-dependent chromatin assembly is functionally distinct from chromatin remodeling. Elife 2013, 2, e00863. [Google Scholar] [CrossRef] [PubMed]
- Dyer, P.N.; Edayathumangalam, R.S.; White, C.L.; Bao, Y.; Chakravarthy, S.; Muthurajan, U.M.; Luger, K. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol 2004, 375, 23–44. [Google Scholar]
- Lowary, P.T.; Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J Mol Biol 1998, 276, 19–42. [Google Scholar] [CrossRef]
- Duan, M.R.; Smerdon, M.J. UV damage in DNA promotes nucleosome unwrapping. J Biol Chem 2010, 285, 26295–26303. [Google Scholar] [CrossRef] [PubMed]
- Westman, E.; Eriksson, S.; Laas, T.; Pernemalm, P.A.; Skold, S.E. Separation of DNA restriction fragments by ion-exchange chromatography on FPLC columns Mono P and Mono Q. Anal Biochem 1987, 166, 158–171. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



