Submitted:
05 May 2026
Posted:
07 May 2026
You are already at the latest version
Abstract
Keywords:
1.0. Introduction
2.0. Structure
3.0. Variants
3.1. Variants of Interest (VOI)
3.2. Variants of Concern (VOC)
3.3. Variants of High Consequence (VOHC)
3.4. Major Variants of Coronavirus
3.4.1. Alpha (B.1.1.7)
3.4.2. Beta (B.1.351)
3.4.3. Gamma (P.1)
3.4.4. Delta (B.1.617.2)
3.4.5. Omicron (B.1.1.529)
4.1. Mechanism of action
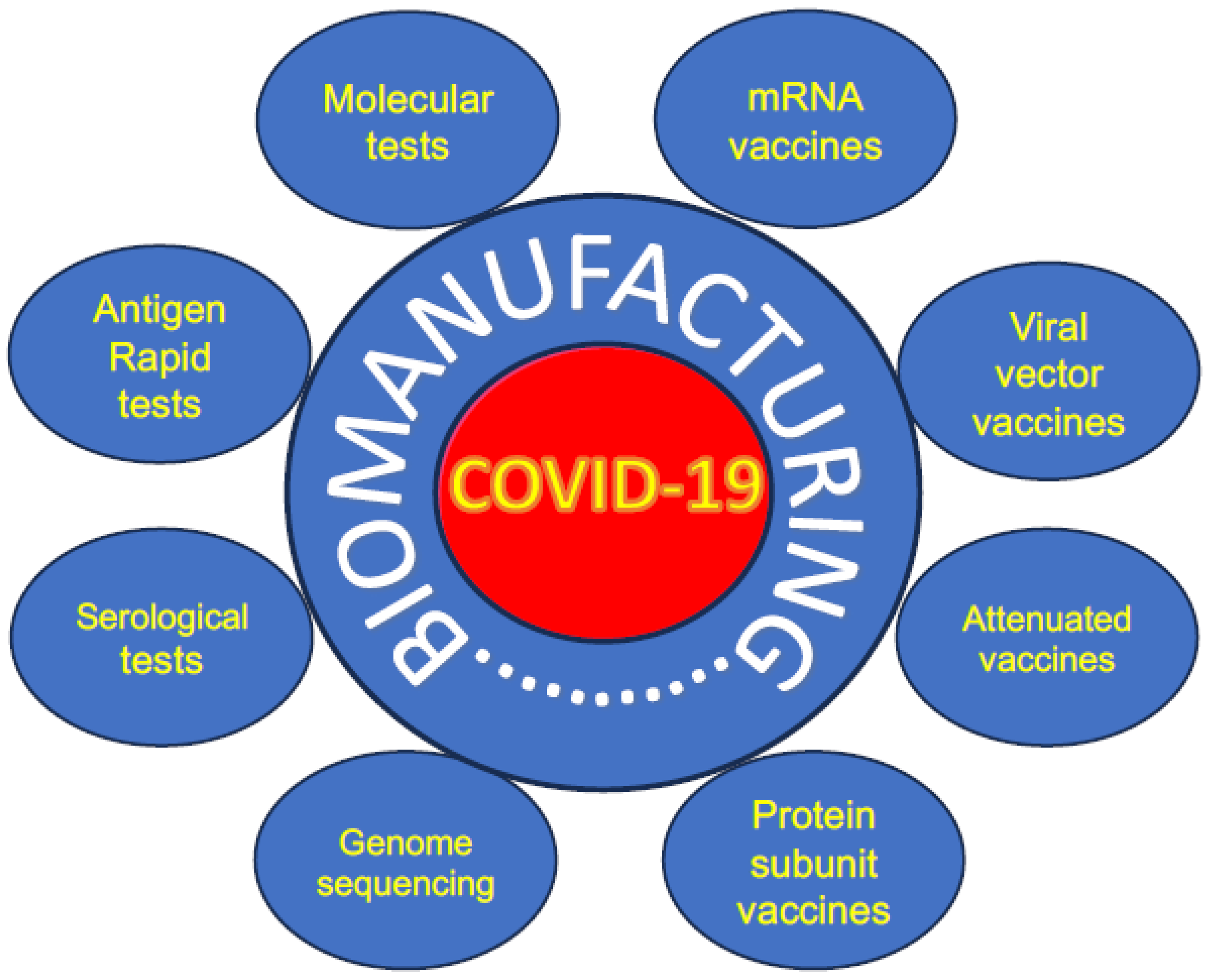
5.0. Response of Biomanufacturing Industry to COVID-19 Pandemic
5.1. SARS-CoV-2 Diagnostic Approaches
5.1.1. Polymerase Chain Reaction (PCR)
5.1.2. Antigen Rapid Diagnostic Test
5.1.3. Serology Test
5.1.4. Reverse Transcription Loop-Mediated Isothermal Amplification (RT-LAMP)
5.1.5. Next-Generation Sequencing
5.1.6. Nanotechnology Approaches in SARS-CoV-2 Diagnosis
6.0. Development of SARS-CoV-2 mRNA Vaccines
6.1. Vaccine Candidates
6.2. Design and Modifications to mRNA
6.3. Delivery of mRNA Vaccines
6.4. Next-Generation mRNA Vaccines
6.5. Advantages, Limitations and Caveats
6.5. Innovations in Large-Scale mRNA Vaccines and Diagnostics
6.5.1. Workforce and Expertise Gaps
6.6. Regulatory Affairs
7.0. Discussion
Author Contributions
Acknowledgements
Conflicts of interest
References
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–3. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. The Lancet 2020, 395, 565–74. [Google Scholar] [CrossRef]
- Felsenstein, S.; Herbert, J.A.; McNamara, P.S.; Hedrich, C.M. COVID-19: Immunology and treatment options. Clin. Immunol. 2020, 215, 108448. [Google Scholar] [CrossRef]
- Ghasemiyeh, P.; Borhani-Haghighi, A.; Karimzadeh, I.; Mohammadi-Samani, S.; Vazin, A.; Safari, A.; et al. Major Neurologic Adverse Drug Reactions, Potential Drug–Drug Interactions and Pharmacokinetic Aspects of Drugs Used in COVID-19 Patients with Stroke: A Narrative Review. Ther. Clin. Risk Manag 2020, Volume 16, 595–605. [Google Scholar] [CrossRef]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; et al. Remdesivir for the Treatment of Covid-19 - Final Report. N Engl. J. Med. 2020, 383, 1813–26. [Google Scholar] [CrossRef]
- Molnupiravir in unvaccinated patients with COVID-19. Drug Ther. Bull. 2022, 60, 35–35. [CrossRef] [PubMed]
- Vassilopoulos, A.; Mylonakis, E. patients with COVID-19 at risk for severe disease, nirmatrelvir + ritonavir reduced hospitalization or death. Ann. Intern Med. 2022, 175, JC63. [Google Scholar] [CrossRef] [PubMed]
- Uraki, R.; Korber, B.; Diamond, M.S.; Kawaoka, Y. SARS-CoV-2 variants: biology, pathogenicity, immunity and control. Nat. Rev. Microbiol. 2026, 24, 8–28. [Google Scholar] [CrossRef]
- Yadav, R.; Chaudhary, J.K.; Jain, N.; Chaudhary, P.K.; Khanra, S.; Dhamija, P.; et al. Role of Structural and Non-Structural Proteins and Therapeutic Targets of SARS-CoV-2 for COVID-19. Cells 2021, 10, 821. [Google Scholar] [CrossRef]
- Yan, W.; Zheng, Y.; Zeng, X.; He, B.; Cheng, W. Structural biology of SARS-CoV-2: open the door for novel therapies. Signal Transduct. Target Ther. 2022, 7, 26. [Google Scholar] [CrossRef]
- Malone, B.; Urakova, N.; Snijder, E.J.; Campbell, E.A. Structures and functions of coronavirus replication–transcription complexes and their relevance for SARS-CoV-2 drug design. Nat. Rev. Mol. Cell Biol. 2022, 23, 21–39. [Google Scholar] [CrossRef]
- Brian, D.A.; Baric, R.S. Coronavirus Genome Structure and Replication; 2005; pp. 1–30. [Google Scholar] [CrossRef]
- Jamison, D.A.; Anand Narayanan, S.; Trovão, N.S.; Guarnieri, J.W.; Topper, M.J.; Moraes-Vieira, P.M.; et al. A comprehensive SARS-CoV-2 and COVID-19 review, Part 1: Intracellular overdrive for SARS-CoV-2 infection. Eur. J. Hum. Genet. 2022, 30, 889–98. [Google Scholar] [CrossRef]
- Yan, W.; Zheng, Y.; Zeng, X.; He, B.; Cheng, W. Structural biology of SARS-CoV-2: open the door for novel therapies. Signal Transduct. Target Ther. 2022, 7, 26. [Google Scholar] [CrossRef]
- Wu, W.; Cheng, Y.; Zhou, H.; Sun, C.; Zhang, S. The SARS-CoV-2 nucleocapsid protein: its role in the viral life cycle, structure and functions, and use as a potential target in the development of vaccines and diagnostics. Virol. J. 2023, 20, 6. [Google Scholar] [CrossRef]
- Papageorgiou, A.C.; Mohsin, I. The SARS-CoV-2 Spike Glycoprotein as a Drug and Vaccine Target: Structural Insights into Its Complexes with ACE2 and Antibodies. Cells 2020, 9, 2343. [Google Scholar] [CrossRef] [PubMed]
- Peeling, R.W.; Heymann, D.L.; Teo, Y.-Y.; Garcia, P.J. Diagnostics for COVID-19: moving from pandemic response to control. The Lancet 2022, 399, 757–68. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef]
- Aleem, A.; Akbar Samad, A.B.; Vaqar, S. Emerging Variants of SARS-CoV-2 and Novel Therapeutics Against Coronavirus (COVID-19). 2024.
- Scovino, A.M.; Dahab, E.C.; Vieira, G.F.; Freire-de-Lima, L.; Freire-de-Lima, C.G.; Morrot, A. SARS-CoV-2’s Variants of Concern: A Brief Characterization. Front Immunol. 2022, 13. [Google Scholar] [CrossRef]
- Ou, J.; Lan, W.; Wu, X.; Zhao, T.; Duan, B.; Yang, P.; et al. Tracking SARS-CoV-2 Omicron diverse spike gene mutations identifies multiple inter-variant recombination events. Signal Transduct. Target Ther. 2022, 7, 138. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.C.; Castro, J.; Lambrou, A.S.; Rose, E.B.; Cook, P.W.; Batra, D.; et al. Genomic Surveillance for SARS-CoV-2 Variants: Circulation of Omicron XBB and JN.1 Lineages — United States, May 2023–September 2024. MMWR Morb. Mortal. Wkly. Rep. 2024, 73, 938–45. [Google Scholar] [CrossRef] [PubMed]
- Salehi-Vaziri, M.; Fazlalipour, M.; Seyed Khorrami, S.M.; Azadmanesh, K.; Pouriayevali, M.H.; Jalali, T.; et al. The ins and outs of SARS-CoV-2 variants of concern (VOCs). Arch. Virol. 2022, 167, 327–44. [Google Scholar] [CrossRef] [PubMed]
- Davies, N.G.; Abbott, S.; Barnard, R.C.; Jarvis, C.I.; Kucharski, A.J.; Munday, J.D.; et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science (1979) 2021, 372. [Google Scholar] [CrossRef] [PubMed]
- Tegally, H.; Wilkinson, E.; Giovanetti, M.; Iranzadeh, A.; Fonseca, V.; Giandhari, J.; et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 2021, 592, 438–43. [Google Scholar] [CrossRef]
- Expression of concern on: Comparative genomics and characterization of SARS-CoV-2 P.1 (Gamma) variant of concern from Amazonas, Brazil. Front Med. 2024, 11. [CrossRef]
- Mlcochova, P.; Kemp, S.A.; Dhar, M.S.; Papa, G.; Meng, B.; Ferreira, I.A.T.M.; et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 2021, 599, 114–9. [Google Scholar] [CrossRef]
- Echaide, M.; Chocarro de Erauso, L.; Bocanegra, A.; Blanco, E.; Kochan, G.; Escors, D. mRNA Vaccines against SARS-CoV-2: Advantages and Caveats. Int. J. Mol. Sci. 2023, 24, 5944. [Google Scholar] [CrossRef]
- Willett, B.J.; Grove, J.; MacLean, O.A.; Wilkie, C.; De Lorenzo, G.; Furnon, W.; et al. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway. Nat. Microbiol. 2022, 7, 1161–79. [Google Scholar] [CrossRef] [PubMed]
- Wimalawansa, S.J. Unlocking insights: Navigating COVID-19 challenges and Emulating future pandemic Resilience strategies with strengthening natural immunity. Heliyon 2024, 10, e34691. [Google Scholar] [CrossRef]
- Mambelli, F.; de Araujo, A.C.V.S.C.; Farias, J.P.; de Andrade, K.Q.; Ferreira, L.C.S.; Minoprio, P.; et al. An Update on Anti-COVID-19 Vaccines and the Challenges to Protect Against New SARS-CoV-2 Variants. Pathogens 2025, 14, 23. [Google Scholar] [CrossRef]
- Jian, F.; Wang, J.; Yisimayi, A.; Song, W.; Xu, Y.; Chen, X.; et al. Evolving antibody response to SARS-CoV-2 antigenic shift from XBB to JN.1. Nature 2025, 637, 921–9. [Google Scholar] [CrossRef]
- Uraki, R.; Korber, B.; Diamond, M.S.; Kawaoka, Y. SARS-CoV-2 variants: biology, pathogenicity, immunity and control. Nat. Rev. Microbiol. 2026, 24, 8–28. [Google Scholar] [CrossRef]
- Nauthor Jaman, M.; Jaman, A.; Ahmmed, N.; Arbin, A.; khondaker, J.; et al. Emerging SARS-CoV-2 Omicron Subvariants in 2025: Clinical Impacts and Public Health Challenges. J. Biosci. Public Health 2025, 1, 1. [Google Scholar] [CrossRef]
- Umakanthan, S.; Chattu, V.K.; Ranade, A.V.; Das, D.; Basavarajegowda, A.; Bukelo, M. A rapid review of recent advances in diagnosis, treatment and vaccination for COVID-19. AIMS Public Health 2021, 8, 137–53. [Google Scholar] [CrossRef] [PubMed]
- Walls, A.C.; Park, Y.-J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e6. [Google Scholar] [CrossRef] [PubMed]
- Hartenian, E.; Nandakumar, D.; Lari, A.; Ly, M.; Tucker, J.M.; Glaunsinger, B.A. The molecular virology of coronaviruses. J. Biol. Chem. 2020, 295, 12910–34. [Google Scholar] [CrossRef]
- Snijder, E.J.; Limpens, R.W.A.L.; de Wilde, A.H.; de Jong, A.W.M.; Zevenhoven-Dobbe, J.C.; Maier, H.J.; et al. A unifying structural and functional model of the coronavirus replication organelle: Tracking down RNA synthesis. PLoS Biol. 2020, 18, e3000715. [Google Scholar] [CrossRef]
- Klein, S.; Cortese, M.; Winter, S.L.; Wachsmuth-Melm, M.; Neufeldt, C.J.; Cerikan, B.; et al. SARS-CoV-2 structure and replication characterized by in situ cryo-electron tomography. Nat. Commun. 2020, 11, 5885. [Google Scholar] [CrossRef]
- V’kovski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Coronavirus biology and replication: implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021, 19, 155–70. [Google Scholar] [CrossRef]
- Guan, W.; Ni, Z.; Hu, Y.; Liang, W.; Ou, C.; He, J.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–20. [Google Scholar] [CrossRef]
- Dutta, D.; Naiyer, S.; Mansuri, S.; Soni, N.; Singh, V.; Bhat, K.H.; et al. COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2. Diagnostics 2022, 12, 1503. [Google Scholar] [CrossRef]
- Drain, P.K. Rapid Diagnostic Testing for SARS-CoV-2. N. Engl. J. Med. 2022, 386, 264–72. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Wang, L.; Sakthivel, S.K.; Whitaker, B.; Murray, J.; Kamili, S.; et al. US CDC Real-Time Reverse Transcription PCR Panel for Detection of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2020, 26, 1654–65. [Google Scholar] [CrossRef]
- Cheng, M.P.; Papenburg, J.; Desjardins, M.; Kanjilal, S.; Quach, C.; Libman, M.; et al. Diagnostic Testing for Severe Acute Respiratory Syndrome–Related Coronavirus 2. Ann. Intern Med. 2020, 172, 726–34. [Google Scholar] [CrossRef] [PubMed]
- Dutta, D.; Naiyer, S.; Mansuri, S.; Soni, N.; Singh, V.; Bhat, K.H.; et al. COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2. Diagnostics 2022, 12, 1503. [Google Scholar] [CrossRef] [PubMed]
- Chu, V.T.; Schwartz, N.G.; Donnelly, M.A.P.; Chuey, M.R.; Soto, R.; Yousaf, A.R.; et al. Comparison of Home Antigen Testing With RT-PCR and Viral Culture During the Course of SARS-CoV-2 Infection. JAMA Intern Med. 2022, 182, 701. [Google Scholar] [CrossRef]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25. [Google Scholar] [CrossRef]
- Anantharajah, A.; Helaers, R.; Defour, J.-P.; Olive, N.; Kabera, F.; Croonen, L.; et al. How to choose the right real-time RT-PCR primer sets for the SARS-CoV-2 genome detection? J. Virol. Methods 2021, 295, 114197. [Google Scholar] [CrossRef]
- Liu, R.; Han, H.; Liu, F.; Lv, Z.; Wu, K.; Liu, Y.; et al. Positive rate of RT-PCR detection of SARS-CoV-2 infection in 4880 cases from one hospital in Wuhan, China, from Jan to Feb 2020. Clin. Chim. Acta 2020, 505, 172–5. [Google Scholar] [CrossRef]
- Shen, Z.; Xiao, Y.; Kang, L.; Ma, W.; Shi, L.; Zhang, L.; et al. Genomic Diversity of Severe Acute Respiratory Syndrome–Coronavirus 2 in Patients With Coronavirus Disease 2019. Clin. Infect. Dis. 2020, 71, 713–20. [Google Scholar] [CrossRef]
- Perez-Romero, C.A.; Mendoza-Maldonado, L.; Tonda, A.; Coz, E.; Tabeling, P.; Vanhomwegen, J.; et al. An Innovative AI-based primer design tool for precise and accurate detection of SARS-CoV-2 variants of concern. Sci. Rep. 2023, 13, 15782. [Google Scholar] [CrossRef]
- Peeling, R.W.; Olliaro, P.L.; Boeras, D.I.; Fongwen, N. Scaling up COVID-19 rapid antigen tests: promises and challenges. Lancet Infect. Dis. 2021, 21, e290–5. [Google Scholar] [CrossRef]
- Brümmer, L.E.; Katzenschlager, S.; Gaeddert, M.; Erdmann, C.; Schmitz, S.; Bota, M.; et al. Accuracy of novel antigen rapid diagnostics for SARS-CoV-2: A living systematic review and meta-analysis. PLoS Med. 2021, 18, e1003735. [Google Scholar] [CrossRef]
- Schildgen, V.; Demuth, S.; Lüsebrink, J.; Schildgen, O. Limits and Opportunities of SARS-CoV-2 Antigen Rapid Tests: An Experienced-Based Perspective. Pathogens 2021, 10, 38. [Google Scholar] [CrossRef]
- Lange, S.J.; Kompaniyets, L.; Freedman, D.S.; Kraus, E.M.; Porter, R.; Blanck, H.M.; et al. Longitudinal Trends in Body Mass Index Before and During the COVID-19 Pandemic Among Persons Aged 2–19 Years — United States, 2018–2020. MMWR Morb. Mortal. Wkly. Rep. 2021, 70, 1278–83. [Google Scholar] [CrossRef] [PubMed]
- Kohmer, N.; Toptan, T.; Pallas, C.; Karaca, O.; Pfeiffer, A.; Westhaus, S.; et al. The Comparative Clinical Performance of Four SARS-CoV-2 Rapid Antigen Tests and Their Correlation to Infectivity In Vitro. J. Clin. Med. 2021, 10, 328. [Google Scholar] [CrossRef] [PubMed]
- Patriquin, G.; LeBlanc, J.J.; Williams, C.; Hatchette, T.F.; Ross, J.; Barrett, L.; et al. Comparison between Nasal and Nasopharyngeal Swabs for SARS-CoV-2 Rapid Antigen Detection in an Asymptomatic Population, and Direct Confirmation by RT-PCR from the Residual Buffer. Microbiol. Spectr. 2022, 10. [Google Scholar] [CrossRef]
- Dinnes, J.; Berhane, S.; Walsh, J.; Reidy, P.; Doherty, A.; Hillier, B.; et al. Rapid, point-of-care antigen tests for diagnosis of SARS-CoV-2 infection. Cochrane Database Syst. Rev. 2025, 2025. [Google Scholar] [CrossRef]
- Arshadi, M.; Fardsanei, F.; Deihim, B.; Farshadzadeh, Z.; Nikkhahi, F.; Khalili, F.; et al. Diagnostic Accuracy of Rapid Antigen Tests for COVID-19 Detection: A Systematic Review With Meta-analysis. Front Med. 2022, 9. [Google Scholar] [CrossRef]
- Baro, B.; Rodo, P.; Ouchi, D.; Bordoy, A.E.; Saya Amaro, E.N.; Salsench, S.V.; et al. Performance characteristics of five antigen-detecting rapid diagnostic test (Ag-RDT) for SARS-CoV-2 asymptomatic infection: a head-to-head benchmark comparison. J. Infect. 2021, 82, 269–75. [Google Scholar] [CrossRef] [PubMed]
- Kim, A.E.; Bennett, J.C.; Luiten, K.; O’Hanlon, J.A.; Wolf, C.R.; Magedson, A.; et al. Comparative Diagnostic Utility of SARS-CoV-2 Rapid Antigen and Molecular Testing in a Community Setting. J. Infect. Dis. 2024, 230, 363–73. [Google Scholar] [CrossRef]
- Wagenhäuser, I.; Knies, K.; Pscheidl, T.; Eisenmann, M.; Flemming, S.; Petri, N.; et al. SARS-CoV-2 antigen rapid detection tests: test performance during the COVID-19 pandemic and the impact of COVID-19 vaccination. EBioMedicine 2024, 109, 105394. [Google Scholar] [CrossRef] [PubMed]
- Derin, D.Ç.; Gültekin, E. Design of a lateral flow assay targeting the conserved NIID_2019-nCoV_N gene region for molecular viral diagnosis. Braz. J. Med. Biol. Res. 2025, 58. [Google Scholar] [CrossRef]
- Kumar, A.; Tripathi, P.; Kumar, P.; Shekhar, R.; Pathak, R. From Detection to Protection: Antibodies and Their Crucial Role in Diagnosing and Combatting SARS-CoV-2. Vaccines 2024, 12, 459. [Google Scholar] [CrossRef]
- Kevadiya, B.D.; Machhi, J.; Herskovitz, J.; Oleynikov, M.D.; Blomberg, W.R.; Bajwa, N.; et al. Diagnostics for SARS-CoV-2 infections. Nat. Mater. 2021, 20, 593–605. [Google Scholar] [CrossRef]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.C.; et al. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822–35. [Google Scholar] [CrossRef] [PubMed]
- Long, Q.-X.; Tang, X.-J.; Shi, Q.-L.; Li, Q.; Deng, H.-J.; Yuan, J.; et al. Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat. Med. 2020, 26, 1200–4. [Google Scholar] [CrossRef]
- Thapa, S.; Singh, K.R.; Verma, R.; Singh, J.; Singh, R.P. State-of-the-Art Smart and Intelligent Nanobiosensors for SARS-CoV-2 Diagnosis. Biosensors 2022, 12, 637. [Google Scholar] [CrossRef]
- Choi, G.; Moehling, T.J.; Meagher, R.J. Advances in RT-LAMP for COVID-19 testing and diagnosis. Expert Rev. Mol. Diagn. 2023, 23, 9–28. [Google Scholar] [CrossRef]
- Fernandes, R.S.; de Oliveira Silva, J.; Gomes, K.B.; Azevedo, R.B.; Townsend, D.M.; de Paula Sabino, A.; et al. Recent advances in point of care testing for COVID-19 detection. Biomed. Pharmacother. 2022, 153, 113538. [Google Scholar] [CrossRef]
- Lim, J.; Koprowski, K.; Stavins, R.; Xuan, N.; Hoang, T.-H.; Baek, J.; et al. Point-of-Care Multiplex Detection of Respiratory Viruses. ACS Sens. 2024, 9, 4058–68. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, T.; Aonuma, H.; Natsuhara, D.; Hoshina, T.; Ote, M.; Saiki, E.; et al. Development of a colorimetric RT-LAMP-based microfluidic device for SARS-CoV-2 detection. Trop. Med. Health 2026, 54, 25. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, N.N.T.; McCarthy, C.; Lantigua, D.; Camci-Unal, G. Development of Diagnostic Tests for Detection of SARS-CoV-2. Diagnostics 2020, 10, 905. [Google Scholar] [CrossRef] [PubMed]
- Behjati, S.; Tarpey, P.S. What is next generation sequencing? Arch. Dis. Child. Educ. Pract. Ed. 2013, 98, 236–8. [Google Scholar] [CrossRef]
- Shu, Y.; McCauley, J. GISAID: Global initiative on sharing all influenza data – from vision to reality. Eurosurveillance 2017, 22. [Google Scholar] [CrossRef]
- Hatcher, E.L.; Zhdanov, S.A.; Bao, Y.; Blinkova, O.; Nawrocki, E.P.; Ostapchuck, Y.; et al. Virus Variation Resource – improved response to emergent viral outbreaks. Nucleic Acids Res. 2017, 45, D482–90. [Google Scholar] [CrossRef]
- Zhao, W.-M.; Song, S.-H.; Chen, M.-L.; Zou, D.; Ma, L.-N.; Ma, Y.-K.; et al. The 2019 novel coronavirus resource. Yi Chuan 2020, 42, 212–21. [Google Scholar] [CrossRef]
- Zhang, Y.; Zmasek, C.; Sun, G.; Larsen, C.N.; Scheuermann, R.H. Hepatitis C Virus Database and Bioinformatics Analysis Tools in the Virus Pathogen Resource (ViPR); 2019; pp. 47–69. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–33. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Chen, X.; Hu, T.; Li, J.; Song, H.; Liu, Y.; et al. A Novel Bat Coronavirus Closely Related to SARS-CoV-2 Contains Natural Insertions at the S1/S2 Cleavage Site of the Spike Protein. Curr. Biol. 2020, 30, 2196–2203.e3. [Google Scholar] [CrossRef]
- Chen, X.; Kang, Y.; Luo, J.; Pang, K.; Xu, X.; Wu, J.; et al. Next-Generation Sequencing Reveals the Progression of COVID-19. Front Cell Infect. Microbiol. 2021, 11. [Google Scholar] [CrossRef]
- Ramuta, M.D.; Newman, C.M.; Brakefield, S.F.; Stauss, M.R.; Wiseman, R.W.; Kita-Yarbro, A.; et al. SARS-CoV-2 and other respiratory pathogens are detected in continuous air samples from congregate settings. Nat. Commun. 2022, 13, 4717. [Google Scholar] [CrossRef]
- Minor, N.R.; Ramuta, M.D.; Stauss, M.R.; Harwood, O.E.; Brakefield, S.F.; Alberts, A.; et al. Metagenomic sequencing detects human respiratory and enteric viruses in air samples collected from congregate settings. Sci. Rep. 2023, 13, 21398. [Google Scholar] [CrossRef]
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–2. [Google Scholar] [CrossRef] [PubMed]
- Yoo, H.M.; Kim, I.-H.; Kim, S. Nucleic Acid Testing of SARS-CoV-2. Int. J. Mol. Sci. 2021, 22, 6150. [Google Scholar] [CrossRef] [PubMed]
- Campos, G.S.; Sardi, S.I.; Falcao, M.B.; Belitardo, E.M.M.A.; Rocha, D.J.P.G.; Rolo, C.A.; et al. Ion torrent-based nasopharyngeal swab metatranscriptomics in COVID-19. J. Virol. Methods 2020, 282, 113888. [Google Scholar] [CrossRef]
- Paden, C.R.; Tao, Y.; Queen, K.; Zhang, J.; Li, Y.; Uehara, A.; et al. Rapid, Sensitive, Full-Genome Sequencing of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2020, 26, 2401–5. [Google Scholar] [CrossRef] [PubMed]
- Pillay, S.; Giandhari, J.; Tegally, H.; Wilkinson, E.; Chimukangara, B.; Lessells, R.; et al. Whole Genome Sequencing of SARS-CoV-2: Adapting Illumina Protocols for Quick and Accurate Outbreak Investigation during a Pandemic. Genes 2020, 11, 949. [Google Scholar] [CrossRef]
- Mercatelli, D.; Holding, A.N.; Giorgi, F.M. Web tools to fight pandemics: the COVID-19 experience. Brief. Bioinform. 2021, 22, 690–700. [Google Scholar] [CrossRef]
- Hu, T.; Li, J.; Zhou, H.; Li, C.; Holmes, E.C.; Shi, W. Bioinformatics resources for SARS-CoV-2 discovery and surveillance. Brief. Bioinform. 2021, 22, 631–41. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, I.; Yusuf, K.; Roy, B.C.; Stubbs, J.; Anant, S.; Attard, T.M.; et al. Dietary Interventions Ameliorate Infectious Colitis by Restoring the Microbiome and Promoting Stem Cell Proliferation in Mice. Int. J. Mol. Sci. 2021, 23, 339. [Google Scholar] [CrossRef]
- Ahmed, I.; Roy, B.C.; Raach, R.-M.T.; Owens, S.M.; Xia, L.; Anant, S.; et al. Enteric infection coupled with chronic Notch pathway inhibition alters colonic mucus composition leading to dysbiosis, barrier disruption and colitis. PLoS ONE 2018, 13, e0206701. [Google Scholar] [CrossRef]
- Coşgun, Y.; Yalçın, S.; Dedeoğlu, E.; Ünal, G.; Kopp, K.; Musul, B.; et al. Enhancing public health surveillance: a comparative study of platform-specific and hybrid assembly approaches in SARS-CoV-2 genome sequencing. Microb. Genom. 2025, 11. [Google Scholar] [CrossRef]
- Hirschhorn, J.W.; Babady, N.E.; Bateman, A.; Blankenship, H.M.; Bard, J.D.; Florek, K.; et al. Considerations for Severe Acute Respiratory Syndrome Coronavirus 2 Genomic Surveillance. J. Mol. Diagn. 2025, 27, 12–24. [Google Scholar] [CrossRef]
- Haleem, A.; Javaid, M.; Singh, R.P.; Rab, S.; Suman, R. Applications of nanotechnology in medical field: a brief review. Glob. Health J. 2023, 7, 70–7. [Google Scholar] [CrossRef]
- Singh, P.; Singh, D.; Sa, P.; Mohapatra, P.; Khuntia, A.; K Sahoo, S. Insights From Nanotechnology in COVID-19: Prevention, Detection, Therapy and Immunomodulation. Nanomedicine 2021, 16, 1219–35. [Google Scholar] [CrossRef]
- Ayan, S.; Aranci-Ciftci, K.; Ciftci, F.; Ustundag, C.B. Nanotechnology and COVID-19: Prevention, diagnosis, vaccine, and treatment strategies. Front Mater. 2023, 9. [Google Scholar] [CrossRef]
- Sharma, A.; Kontodimas, K.; Bosmann, M. Nanomedicine: A Diagnostic and Therapeutic Approach to COVID-19. Front Med. 2021, 8. [Google Scholar] [CrossRef]
- Valenzuela-Fernández, A.; Cabrera-Rodriguez, R.; Ciuffreda, L.; Perez-Yanes, S.; Estevez-Herrera, J.; González-Montelongo, R.; et al. Nanomaterials to combat SARS-CoV-2: Strategies to prevent, diagnose and treat COVID-19. Front Bioeng. Biotechnol. 2022, 10. [Google Scholar] [CrossRef] [PubMed]
- Alhalaili, B.; Popescu, I.N.; Kamoun, O.; Alzubi, F.; Alawadhia, S.; Vidu, R. Nanobiosensors for the Detection of Novel Coronavirus 2019-nCoV and Other Pandemic/Epidemic Respiratory Viruses: A Review. Sensors 2020, 20, 6591. [Google Scholar] [CrossRef] [PubMed]
- Valerio, T.L.; Anastácio, R.; da Silva, S.S.; de Oliveira, C.C.; Vidotti, M. An overview of electrochemical biosensors used for COVID-19 detection. Anal. Methods 2024, 16, 2164–76. [Google Scholar] [CrossRef]
- Tekin, Y.S.; Kul, S.M.; Sagdic, O.; Rodthongkum, N.; Geiss, B.; Ozer, T. Optical biosensors for diagnosis of COVID-19: nanomaterial-enabled particle strategies for post pandemic era. Microchim. Acta 2024, 191, 320. [Google Scholar] [CrossRef]
- Harun-Ur-Rashid, M.; Foyez, T.; Jahan, I.; Pal, K.; Bin, Imran A. Rapid diagnosis of COVID-19 via nano-biosensor-implemented biomedical utilization: a systematic review. RSC Adv. 2022, 12, 9445–65. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–79. [Google Scholar] [CrossRef]
- Martinon, F.; Krishnan, S.; Lenzen, G.; Magné, R.; Gomard, E.; Guillet, J.; et al. Induction of virus-specific cytotoxic T lymphocytes in vivo by liposome-entrapped mRNA. Eur. J. Immunol. 1993, 23, 1719–22. [Google Scholar] [CrossRef] [PubMed]
- BRENNER, S.; JACOB, F.; MESELSON, M. An Unstable Intermediate Carrying Information from Genes to Ribosomes for Protein Synthesis. Nature 1961, 190, 576–81. [Google Scholar] [CrossRef] [PubMed]
- Lockard, R.E.; Lingrel, J.B. The synthesis of mouse hemoglobin chains in a rabbit reticulocyte cell-free system programmed with mouse reticulocyte 9S RNA. Biochem Biophys. Res. Commun. 1969, 37, 204–12. [Google Scholar] [CrossRef]
- Krieg, P.A.; Melton, D.A. [25] In vitro RNA synthesis with SP6 RNA polymerase; 1987; pp. 397–415. [Google Scholar] [CrossRef]
- Krieg, P.A.; Melton, D.A. Functional messenger RNAs are produced by SP6 in vitro transcription of cloned cDNAs. Nucleic Acids Res. 1984, 12, 7057–70. [Google Scholar] [CrossRef]
- Malone, R.W.; Felgner, P.L.; Verma, I.M. Cationic liposome-mediated RNA transfection. Proc. Natl. Acad. Sci. 1989, 86, 6077–81. [Google Scholar] [CrossRef]
- Wolff, J.A.; Budker, V. The Mechanism of Naked DNA Uptake and Expression; 2005; pp. 1–20. [Google Scholar] [CrossRef]
- Jirikowski, G.F.; Sanna, P.P.; Maciejewski-Lenoir, D.; Bloom, F.E. Reversal of Diabetes Insipidus in Brattleboro Rats: Intrahypothalamic Injection of Vasopressin mRNA. Science (1979) 1992, 255, 996–8. [Google Scholar] [CrossRef]
- Hassett, K.J.; Benenato, K.E.; Jacquinet, E.; Lee, A.; Woods, A.; Yuzhakov, O.; et al. Optimization of Lipid Nanoparticles for Intramuscular Administration of mRNA Vaccines. Mol. Ther. Nucleic Acids 2019, 15, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Conry, R.M.; LoBuglio, A.F.; Loechel, F.; Moore, S.E.; Sumerel, L.A.; Barlow, D.L.; et al. A carcinoembryonic antigen polynucleotide vaccine has in vivo antitumor activity. Gene Ther. 1995, 2, 59–65. [Google Scholar]
- Karikó, K.; Ni, H.; Capodici, J.; Lamphier, M.; Weissman, D. mRNA Is an Endogenous Ligand for Toll-like Receptor 3. J. Biol. Chem. 2004, 279, 12542–50. [Google Scholar] [CrossRef]
- Andries, O.; Mc Cafferty, S.; De Smedt, S.C.; Weiss, R.; Sanders, N.N.; Kitada, T. N1-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. J. Control. Release 2015, 217, 337–44. [Google Scholar] [CrossRef] [PubMed]
- Karikó, K.; Muramatsu, H.; Ludwig, J.; Weissman, D. Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res. 2011, 39, e142–e142. [Google Scholar] [CrossRef]
- Karikó, K.; Buckstein, M.; Ni, H.; Weissman, D. Suppression of RNA Recognition by Toll-like Receptors: The Impact of Nucleoside Modification and the Evolutionary Origin of RNA. Immunity 2005, 23, 165–75. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-Z.; Holmes, E.C. A Genomic Perspective on the Origin and Emergence of SARS-CoV-2. Cell 2020, 181, 223–7. [Google Scholar] [CrossRef]
- Freed, N.E.; Vlková, M.; Faisal, M.B.; Silander, O.K. Rapid and inexpensive whole-genome sequencing of SARS-CoV-2 using 1200 bp tiled amplicons and Oxford Nanopore Rapid Barcoding. Biol. Methods Protoc. 2020, 5. [Google Scholar] [CrossRef] [PubMed]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science (1979) 2020, 367, 1260–3. [Google Scholar] [CrossRef] [PubMed]
- Corbett, K.S.; Edwards, D.; Leist, S.R.; Abiona, O.M.; Boyoglu-Barnum, S.; Gillespie, R.A.; et al. SARS-CoV-2 mRNA Vaccine Development Enabled by Prototype Pathogen Preparedness 2020. [CrossRef]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; et al. Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine. N. Engl. J. Med. 2020, 383, 2603–15. [Google Scholar] [CrossRef]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–16. [Google Scholar] [CrossRef]
- Sadoff, J.; Gray, G.; Vandebosch, A.; Cárdenas, V.; Shukarev, G.; Grinsztejn, B.; et al. Safety and Efficacy of Single-Dose Ad26.COV2.S Vaccine against Covid-19. N. Engl. J. Med. 2021, 384, 2187–201. [Google Scholar] [CrossRef]
- Yin, Z.; Chen, J.-L.; Lu, Y.; Wang, B.; Godfrey, L.; Mentzer, A.J.; et al. Evaluation of T cell responses to naturally processed variant SARS-CoV-2 spike antigens in individuals following infection or vaccination. Cell Rep. 2023, 42, 112470. [Google Scholar] [CrossRef]
- Tamandjou Tchuem, C.R.; Auvigne, V.; Vaux, S.; Montagnat, C.; Paireau, J.; Monnier Besnard, S.; et al. Vaccine effectiveness and duration of protection of COVID-19 mRNA vaccines against Delta and Omicron BA.1 symptomatic and severe COVID-19 outcomes in adults aged 50 years and over in France. Vaccine 2023, 41, 2280–8. [Google Scholar] [CrossRef]
- Powell, A.A.; Kirsebom, F.; Stowe, J.; Ramsay, M.E.; Lopez-Bernal, J.; Andrews, N.; et al. Protection against symptomatic infection with delta (B.1.617.2) and omicron (B.1.1.529) BA.1 and BA.2 SARS-CoV-2 variants after previous infection and vaccination in adolescents in England, August, 2021–March, 2022: a national, observational, test-negative, case-control study. Lancet Infect. Dis. 2023, 23, 435–44. [Google Scholar] [CrossRef] [PubMed]
- Abu-Raddad, L.J.; Chemaitelly, H.; Ayoub, H.H.; AlMukdad, S.; Yassine, H.M.; Al-Khatib, H.A.; et al. Effect of mRNA Vaccine Boosters against SARS-CoV-2 Omicron Infection in Qatar. N. Engl. J. Med. 2022, 386, 1804–16. [Google Scholar] [CrossRef] [PubMed]
- Heinz, F.X.; Stiasny, K. Profiles of current COVID-19 vaccines. Wien. Klin. Wochenschr. 2021, 133, 271–83. [Google Scholar] [CrossRef]
- Ada, G.L. The ideal vaccine. World J. Microbiol. Biotechnol. 1991, 7, 105–9. [Google Scholar] [CrossRef] [PubMed]
- Florindo, H.F.; Kleiner, R.; Vaskovich-Koubi, D.; Acúrcio, R.C.; Carreira, B.; Yeini, E.; et al. Immune-mediated approaches against COVID-19. Nat. Nanotechnol. 2020, 15, 630–45. [Google Scholar] [CrossRef]
- Yadav, T.; Srivastava, N.; Mishra, G.; Dhama, K.; Kumar, S.; Puri, B.; et al. Recombinant vaccines for COVID-19. Hum. Vaccin Immunother. 2020, 16, 2905–12. [Google Scholar] [CrossRef]
- Jimenez-Guardeño, J.M.; Regla-Nava, J.A.; Nieto-Torres, J.L.; DeDiego, M.L.; Castaño-Rodriguez, C.; Fernandez-Delgado, R.; et al. Identification of the Mechanisms Causing Reversion to Virulence in an Attenuated SARS-CoV for the Design of a Genetically Stable Vaccine. PLoS Pathog. 2015, 11, e1005215. [Google Scholar] [CrossRef]
- Dong, Y.; Dai, T.; Wei, Y.; Zhang, L.; Zheng, M.; Zhou, F. A systematic review of SARS-CoV-2 vaccine candidates. Signal Transduct. Target Ther. 2020, 5, 237. [Google Scholar] [CrossRef]
- Dunkle, L.M.; Kotloff, K.L.; Gay, C.L.; Áñez, G.; Adelglass, J.M.; Barrat Hernández, A.Q.; et al. Efficacy and Safety of NVX-CoV2373 in Adults in the United States and Mexico. N Engl. J. Med. 2022, 386, 531–43. [Google Scholar] [CrossRef]
- Hotez, P.J.; Bottazzi, M.E. Whole Inactivated Virus and Protein-Based COVID-19 Vaccines. Annu Rev. Med. 2022, 73, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Wilson-Welder, J.H.; Torres, M.P.; Kipper, M.J.; Mallapragada, S.K.; Wannemuehler, M.J.; Narasimhan, B. Vaccine adjuvants: Current challenges and future approaches. J. Pharm. Sci. 2009, 98, 1278–316. [Google Scholar] [CrossRef] [PubMed]
- Heath, P.T.; Galiza, E.P.; Baxter, D.N.; Boffito, M.; Browne, D.; Burns, F.; et al. Safety and Efficacy of NVX-CoV2373 Covid-19 Vaccine. N. Engl. J. Med. 2021, 385, 1172–83. [Google Scholar] [CrossRef]
- Keech, C.; Albert, G.; Cho, I.; Robertson, A.; Reed, P.; Neal, S.; et al. Phase 1–2 Trial of a SARS-CoV-2 Recombinant Spike Protein Nanoparticle Vaccine. N. Engl. J. Med. 2020, 383, 2320–32. [Google Scholar] [CrossRef] [PubMed]
- McConeghy, K.W.; Davidson, H.E.; Canaday, D.H.; Han, L.; Hayes, K.; Baier, R.R.; et al. Recombinant vs Egg-Based Quadrivalent Influenza Vaccination for Nursing Home Residents. JAMA Netw. Open 2025, 8, e2452677. [Google Scholar] [CrossRef]
- Matsumura, T.; Takano, T.; Takahashi, Y. Immune responses related to the immunogenicity and reactogenicity of COVID-19 mRNA vaccines. Int. Immunol. 2023, 35, 213–20. [Google Scholar] [CrossRef]
- Duan, L.; Zheng, Q.; Zhang, H.; Niu, Y.; Lou, Y.; Wang, H. The SARS-CoV-2 Spike Glycoprotein Biosynthesis, Structure, Function, and Antigenicity: Implications for the Design of Spike-Based Vaccine Immunogens. Front Immunol. 2020, 11. [Google Scholar] [CrossRef]
- Li, K.; Huang, B.; Wu, M.; Zhong, A.; Li, L.; Cai, Y.; et al. Dynamic changes in anti-SARS-CoV-2 antibodies during SARS-CoV-2 infection and recovery from COVID-19. Nat. Commun. 2020, 11, 6044. [Google Scholar] [CrossRef]
- Huang, Y.; Yang, C.; Xu, X.; Xu, W.; Liu, S. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol. Sin. 2020, 41, 1141–9. [Google Scholar] [CrossRef]
- Okuyama, R. mRNA and Adenoviral Vector Vaccine Platforms Utilized in COVID-19 Vaccines: Technologies, Ecosystem, and Future Directions. Vaccines 2023, 11, 1737. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–79. [Google Scholar] [CrossRef]
- Kim, S.C.; Sekhon, S.S.; Shin, W.-R.; Ahn, G.; Cho, B.-K.; Ahn, J.-Y.; et al. Modifications of mRNA vaccine structural elements for improving mRNA stability and translation efficiency. Mol. Cell Toxicol. 2022, 18, 1–8. [Google Scholar] [CrossRef]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; et al. Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine. N. Engl. J. Med. 2020, 383, 2603–15. [Google Scholar] [CrossRef]
- Walsh, E.E.; Pérez Marc, G.; Zareba, A.M.; Falsey, A.R.; Jiang, Q.; Patton, M.; et al. Efficacy and Safety of a Bivalent RSV Prefusion F Vaccine in Older Adults. N. Engl. J. Med. 2023, 388, 1465–77. [Google Scholar] [CrossRef] [PubMed]
- Mugridge, J.S.; Coller, J.; Gross, J.D. Structural and molecular mechanisms for the control of eukaryotic 5′–3′ mRNA decay. Nat. Struct. Mol. Biol. 2018, 25, 1077–85. [Google Scholar] [CrossRef]
- Chaudhary, N.; Weissman, D.; Whitehead, K.A. mRNA vaccines for infectious diseases: principles, delivery and clinical translation. Nat. Rev. Drug Discov. 2021, 20, 817–38. [Google Scholar] [CrossRef]
- Chatterjee, S.; Pal, J.K. Role of 5′- and 3′-untranslated regions of mRNAs in human diseases. Biol. Cell 2009, 101, 251–62. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, J.; Schmeing, T.M.; Sonenberg, N. The multifaceted eukaryotic cap structure. WIREs RNA 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Tuller, T.; Zur, H. Multiple roles of the coding sequence 5′ end in gene expression regulation. Nucleic Acids Res. 2015, 43, 13–28. [Google Scholar] [CrossRef]
- Courel, M.; Clément, Y.; Bossevain, C.; Foretek, D.; Vidal Cruchez, O.; Yi, Z.; et al. GC content shapes mRNA storage and decay in human cells. Elife 2019, 8. [Google Scholar] [CrossRef]
- Kudla, G.; Lipinski, L.; Caffin, F.; Helwak, A.; Zylicz, M. High Guanine and Cytosine Content Increases mRNA Levels in Mammalian Cells. PLoS Biol. 2006, 4, e180. [Google Scholar] [CrossRef]
- Silva, J.; Fernandes, R.; Romão, L. Translational Regulation by Upstream Open Reading Frames and Human Diseases; 2019; pp. 99–116. [Google Scholar] [CrossRef]
- Eckmann, C.R.; Rammelt, C.; Wahle, E. Control of poly(A) tail length. WIREs RNA 2011, 2, 348–61. [Google Scholar] [CrossRef]
- Biziaev, N.; Shuvalov, A.; Salman, A.; Egorova, T.; Shuvalova, E.; Alkalaeva, E. The impact of mRNA poly(A) tail length on eukaryotic translation stages. Nucleic Acids Res. 2024, 52, 7792–808. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–79. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, S.; Satapathy, S.R.; Dutta, T. Delivery Strategies for mRNA Vaccines. Pharm. Med. 2022, 36, 11–20. [Google Scholar] [CrossRef]
- Chen, S.; Huang, X.; Xue, Y.; Álvarez-Benedicto, E.; Shi, Y.; Chen, W.; et al. Nanotechnology-based mRNA vaccines. Nat. Rev. Methods Prim. 2023, 3, 63. [Google Scholar] [CrossRef] [PubMed]
- Kiaie, S.H.; Majidi Zolbanin, N.; Ahmadi, A.; Bagherifar, R.; Valizadeh, H.; Kashanchi, F.; et al. Recent advances in mRNA-LNP therapeutics: immunological and pharmacological aspects. J. Nanobiotechnology 2022, 20, 276. [Google Scholar] [CrossRef] [PubMed]
- Heshmati, N.; Chakka, L.R.J.; Zhang, Y.; Maniruzzaman, M. Fabrication of mRNA encapsulated lipid nanoparticles using state of the art SMART-MaGIC technology and transfection in vitro. Sci. Rep. 2024, 14, 22714. [Google Scholar] [CrossRef]
- Kiaie, S.H.; Majidi Zolbanin, N.; Ahmadi, A.; Bagherifar, R.; Valizadeh, H.; Kashanchi, F.; et al. Recent advances in mRNA-LNP therapeutics: immunological and pharmacological aspects. J. Nanobiotechnology 2022, 20, 276. [Google Scholar] [CrossRef]
- Kowalski, P.S.; Rudra, A.; Miao, L.; Anderson, D.G. Delivering the Messenger: Advances in Technologies for Therapeutic mRNA Delivery. Mol. Ther. 2019, 27, 710–28. [Google Scholar] [CrossRef]
- Jackson, L.A.; Anderson, E.J.; Rouphael, N.G.; Roberts, P.C.; Makhene, M.; Coler, R.N.; et al. An mRNA Vaccine against SARS-CoV-2 — Preliminary Report. N. Engl. J. Med. 2020, 383, 1920–31. [Google Scholar] [CrossRef]
- Verbeke, R.; Lentacker, I.; De Smedt, S.C.; Dewitte, H. The dawn of mRNA vaccines: The COVID-19 case. J. Control. Release 2021, 333, 511–20. [Google Scholar] [CrossRef]
- Hou, X.; Zaks, T.; Langer, R.; Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 2021, 6, 1078–94. [Google Scholar] [CrossRef]
- Li, X.; Qi, J.; Wang, J.; Hu, W.; Zhou, W.; Wang, Y.; et al. Nanoparticle technology for mRNA: Delivery strategy, clinical application and developmental landscape. Theranostics 2024, 14, 738–60. [Google Scholar] [CrossRef]
- Choi, J.; Kim, S.-H.; Park, S.; Jang, H.M.; Lee, Y.J.; Kim, H.-J.; et al. Robust neutralization of emerging JN.1 subvariants by the updated JN.1 vaccine: rationale for catch-up booster recommendations in high-risk individuals. Front Immunol. 2026, 16. [Google Scholar] [CrossRef] [PubMed]
- Levine, K.S.; Blanc, R.; Wang, Q.; Malca, H.; Sekulovich, R.; Jin, H.; et al. Self-amplifying COVID-19 mRNA vaccination induces longitudinally enhanced antibody function in a Phase 3 trial. npj Vaccines 2026. [Google Scholar] [CrossRef]
- Okada, Y.; Kumagai, Y.; Okura, I.; Otsuki, M.; Ishida, N.; Iwama, Y.; et al. Immunogenicity of a booster dose of a bivalent (Asp614Gly and omicron BA.4/5 variant) self-amplifying mRNA SARS-CoV-2 booster vaccine versus the BNT162b2 omicron BA.4/5 mRNA vaccine: a randomised phase 3 trial. Lancet Infect. Dis. 2025, 25, 290–300. [Google Scholar] [CrossRef] [PubMed]
- Hồ, N.T.; Hughes, S.G.; Ta, V.T.; Phan, L.T.; Đỗ, Q.; Nguyễn, T.V.; et al. Safety, immunogenicity and efficacy of the self-amplifying mRNA ARCT-154 COVID-19 vaccine: pooled phase 1, 2, 3a and 3b randomized, controlled trials. Nat. Commun. 2024, 15, 4081. [Google Scholar] [CrossRef] [PubMed]
- Longet, S.; Paul, S. Current status of intranasal and inhaled COVID-19 vaccines. npj Vaccines 2026. [Google Scholar] [CrossRef]
- Plett, P.C. [Peter Plett and other discoverers of cowpox vaccination before Edward Jenner]. Sudhoffs Arch. 2006, 90, 219–32. [Google Scholar]
- Jenner, E. Dr. Jenner, on the Vaccine Inoculation. Med. Phys. J. 1800, 3, 502–3. [Google Scholar] [PubMed]
- Maruggi, G.; Zhang, C.; Li, J.; Ulmer, J.B.; Yu, D. mRNA as a Transformative Technology for Vaccine Development to Control Infectious Diseases. Mol. Ther. 2019, 27, 757–72. [Google Scholar] [CrossRef]
- Krammer, F. SARS-CoV-2 vaccines in development. Nature 2020, 586, 516–27. [Google Scholar] [CrossRef]
- Ruiz-Fresneda, M.A.; Ruiz-Pérez, R.; Ruiz-Fresneda, C.; Jiménez-Contreras, E. Differences in Global Scientific Production Between New mRNA and Conventional Vaccines Against COVID-19. Environ. Sci. Pollut. Res. 2022, 29, 57054–66. [Google Scholar] [CrossRef] [PubMed]
- Pollard, A.J.; Bijker, E.M. A guide to vaccinology: from basic principles to new developments. Nat. Rev. Immunol. 2021, 21, 83–100. [Google Scholar] [CrossRef]
- Wang, Y.-S.; Kumari, M.; Chen, G.-H.; Hong, M.-H.; Yuan, J.P.-Y.; Tsai, J.-L.; et al. mRNA-based vaccines and therapeutics: an in-depth survey of current and upcoming clinical applications. J. BioMed Sci. 2023, 30, 84. [Google Scholar] [CrossRef] [PubMed]
- Awakoaiye, B.; Li, S.; Sanchez, S.; Dangi, T.; Irani, N.; Arroyo, L.; et al. Comparative analysis of adenovirus, mRNA, and protein vaccines reveals context-dependent immunogenicity and efficacy. JCI Insight 2025, 10. [Google Scholar] [CrossRef]
- Liu, Y.; Ye, Q. Safety and Efficacy of the Common Vaccines against COVID-19. Vaccines 2022, 10, 513. [Google Scholar] [CrossRef]
- Mostafavi, F.; Bahardoust, M.; Sera, F.; Amirabadizadeh, A.; Allahyari, S.; Ssentongod, P.; et al. COVID-19 Vaccine Effectiveness of Booster Doses Against Delta and Omicron Variants Over Follow-up Times Using Longitudinal Meta-analysis. J. Res. Health Sci. 2024, 24, e00626. [Google Scholar] [CrossRef]
- Echaide, M.; Chocarro de Erauso, L.; Bocanegra, A.; Blanco, E.; Kochan, G.; Escors, D. mRNA Vaccines against SARS-CoV-2: Advantages and Caveats. Int. J. Mol. Sci. 2023, 24, 5944. [Google Scholar] [CrossRef]
- Patone, M.; Mei, X.W.; Handunnetthi, L.; Dixon, S.; Zaccardi, F.; Shankar-Hari, M.; et al. Risk of Myocarditis After Sequential Doses of COVID-19 Vaccine and SARS-CoV-2 Infection by Age and Sex. Circulation 2022, 146, 743–54. [Google Scholar] [CrossRef]
- Xu, S.; Yang, K.; Li, R.; Zhang, L. mRNA Vaccine Era—Mechanisms, Drug Platform and Clinical Prospection. Int. J. Mol. Sci. 2020, 21, 6582. [Google Scholar] [CrossRef]
- Echaide, M.; Chocarro de Erauso, L.; Bocanegra, A.; Blanco, E.; Kochan, G.; Escors, D. mRNA Vaccines against SARS-CoV-2: Advantages and Caveats. Int. J. Mol. Sci. 2023, 24, 5944. [Google Scholar] [CrossRef]
- Wang, J.; Ding, Y.; Chong, K.; Cui, M.; Cao, Z.; Tang, C.; et al. Recent Advances in Lipid Nanoparticles and Their Safety Concerns for mRNA Delivery. Vaccines 2024, 12, 1148. [Google Scholar] [CrossRef]
- Webb, C.; Ip, S.; Bathula, N.V.; Popova, P.; Soriano, S.K.V.; Ly, H.H.; et al. Current Status and Future Perspectives on MRNA Drug Manufacturing. Mol. Pharm. 2022, 19, 1047–58. [Google Scholar] [CrossRef] [PubMed]
- SeyedAlinaghi, S.; Karimi, A.; Pashaei, Z.; Afzalian, A.; Mirzapour, P.; Ghorbanzadeh, K.; et al. Safety and Adverse Events Related to COVID-19 mRNA Vaccines; a Systematic Review. Arch. Acad. Emerg. Med. 2022, 10, e41. [Google Scholar] [CrossRef] [PubMed]
- Feddema, J.J.; Fernald, K.D.S.; Schikan, H.G.C.P.; van de Burgwal, L.H.M. Upscaling vaccine manufacturing capacity - key bottlenecks and lessons learned. Vaccine 2023, 41, 4359–68. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, S.M.; Rosen, A.M.; Edwards, D.; Bolio, A.; Larson, H.J.; Servin, M.; et al. Opportunities and challenges to implementing mRNA-based vaccines and medicines: lessons from COVID-19. Front Public Health 2024, 12. [Google Scholar] [CrossRef]
- Seo, S.H.; Song, M.K. Advancements and challenges in next-generation mRNA vaccine manufacturing systems. Clin. Exp. Vaccine Res. 2025, 14, 299. [Google Scholar] [CrossRef]
- Hengelbrock, A.; Schmidt, A.; Helgers, H.; Vetter, F.L.; Strube, J. Scalable mRNA Machine for Regulatory Approval of Variable Scale between 1000 Clinical Doses to 10 Million Manufacturing Scale Doses. Processes 2023, 11, 745. [Google Scholar] [CrossRef]
- Shepherd, S.J.; Han, X.; Mukalel, A.J.; El-Mayta, R.; Thatte, A.S.; Wu, J.; et al. Throughput-scalable manufacturing of SARS-CoV-2 mRNA lipid nanoparticle vaccines. Proc. Natl. Acad. Sci. 2023, 120. [Google Scholar] [CrossRef]
- Whitley, J.; Zwolinski, C.; Denis, C.; Maughan, M.; Hayles, L.; Clarke, D.; et al. Development of mRNA manufacturing for vaccines and therapeutics: mRNA platform requirements and development of a scalable production process to support early phase clinical trials. Transl. Res. 2022, 242, 38–55. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Basu, M.; Chakraborty, D.; Ghosh, P.; Ghosh, M.K. Unraveling COVID-19 Diagnostics: A Roadmap for Future Pandemic. Nat. Cell Sci. 2024, 000, 000–000. [Google Scholar] [CrossRef]
- Chisholm, O.; Critchley, H. Future directions in regulatory affairs. Front Med. 2023, 9. [Google Scholar] [CrossRef]
- Ostad Ali Dehaghi, R.; Khadem Broojerdi, A.; Magdy, A.; Valentin, M.; Dahlan, J.; Malik, O.; et al. Strengthening Vaccine Regulation: Insights from COVID-19 Vaccines, Best Practices, and Lessons for Future Public Health Emergencies. Vaccines 2025, 13, 638. [Google Scholar] [CrossRef]
- Quinn, S.C.; Jamison, A.M.; Freimuth, V. Communicating Effectively About Emergency Use Authorization and Vaccines in the COVID-19 Pandemic. Am. J. Public Health 2021, 111, 355–8. [Google Scholar] [CrossRef] [PubMed]
- Xu, A.; Xie, L.; Zhong, L.; Zhao, X.; Wan, Z.; Wang, J.-H.; et al. Development of a multiplex real-time RT-PCR assay for simultaneous detection of 18 respiratory viruses. Front Cell Infect. Microbiol. 2026, 15. [Google Scholar] [CrossRef] [PubMed]
- Erlichster, M.; Chana, G.; Zantomio, D.; Goudey, B.; Skafidas, E. Pan-Family Assays for Rapid Viral Screening: Reducing Delays in Public Health Responses During Pandemics. Clin. Infect. Dis. 2021, 73, e3047–52. [Google Scholar] [CrossRef]
- Li, X.; Guo, J.; Yang, H.; Wu, Y.; Xie, Z.; Li, D. A CRISPR-assisted passive microfluidic chip for rapid, visual detection of multiple respiratory viruses. Sci. Rep. 2025, 16, 2033. [Google Scholar] [CrossRef]
- Awakoaiye, B.; Li, S.; Sanchez, S.; Dangi, T.; Irani, N.; Arroyo, L.; et al. Comparative analysis of adenovirus, mRNA, and protein vaccines reveals context-dependent immunogenicity and efficacy. JCI Insight 2025, 10. [Google Scholar] [CrossRef]
- Bouazzaoui, A.; Abdellatif, A.A.H.; Al-Allaf, F.A.; Bogari, N.M.; Al-Dehlawi, S.; Qari, S.H. Strategies for Vaccination: Conventional Vaccine Approaches Versus New-Generation Strategies in Combination with Adjuvants. Pharmaceutics 2021, 13, 140. [Google Scholar] [CrossRef]
- Gote, V.; Bolla, P.K.; Kommineni, N.; Butreddy, A.; Nukala, P.K.; Palakurthi, S.S.; et al. A Comprehensive Review of mRNA Vaccines. Int. J. Mol. Sci. 2023, 24, 2700. [Google Scholar] [CrossRef]
- Hirschhorn, J.W.; Babady, N.E.; Bateman, A.; Blankenship, H.M.; Bard, J.D.; Florek, K.; et al. Considerations for Severe Acute Respiratory Syndrome Coronavirus 2 Genomic Surveillance: A Joint Consensus Recommendation of the Association for Molecular Pathology and Association of Public Health Laboratories. J. Mol. Diagn. 2025, 27, 12–24. [Google Scholar] [CrossRef]
- Bosch, I.; de Puig, H.; Hiley, M.; Carré-Camps, M.; Perdomo-Celis, F.; Narváez, C.F.; et al. Rapid antigen tests for dengue virus serotypes and Zika virus in patient serum. Sci. Transl. Med. 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Shaffaf, T.; Ghafar-Zadeh, E. COVID-19 Diagnostic Strategies Part II: Protein-Based Technologies. Bioengineering 2021, 8, 54. [Google Scholar] [CrossRef] [PubMed]
- Aoki, M.N.; de Oliveira Coelho, B.; Góes, L.G.B.; Minoprio, P.; Durigon, E.L.; Morello, L.G.; et al. Colorimetric RT-LAMP SARS-CoV-2 diagnostic sensitivity relies on color interpretation and viral load. Sci. Rep. 2021, 11, 9026. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Coulter, F.J.; Yang, M.; Smith, J.L.; Tafesse, F.G.; Messer, W.B.; et al. A lyophilized colorimetric RT-LAMP test kit for rapid, low-cost, at-home molecular testing of SARS-CoV-2 and other pathogens. Sci. Rep. 2022, 12, 7043. [Google Scholar] [CrossRef]
- Atmar, R.L.; Ramani, S. Immunologic Detection and Characterization. In Viral Infections of Humans; Springer US: New York, NY, 2022; pp. 1–30. [Google Scholar] [CrossRef]
- Najafabadi, Z.Y.; Fanuel, S.; Falak, R.; Kaboli, S.; Kardar, G.A. The Trend of CRISPR-Based Technologies in COVID-19 Disease: Beyond Genome Editing. Mol. Biotechnol. 2023, 65, 146–61. [Google Scholar] [CrossRef]
- Li, X.; Qi, J.; Wang, J.; Hu, W.; Zhou, W.; Wang, Y.; et al. Nanoparticle technology for mRNA: Delivery strategy, clinical application and developmental landscape. Theranostics 2024, 14, 738–60. [Google Scholar] [CrossRef]
- Jones, I.; Roy, P. Sputnik V COVID-19 vaccine candidate appears safe and effective. The Lancet 2021, 397, 642–3. [Google Scholar] [CrossRef]
- Pramod, S.; Govindan, D.; Ramasubramani, P.; Kar, S.S.; Aggarwal, R.; Manoharan, N.; et al. Effectiveness of Covishield vaccine in preventing Covid-19 – A test-negative case-control study. Vaccine 2022, 40, 3294–7. [Google Scholar] [CrossRef]
| Name of company | Country | Product | Function | References |
|---|---|---|---|---|
| GenScript USA Inc. | USA | Rapid antigen test | Gold-coated antibodies produce colorimetric signal on paper in presence of target serum | [212,213] |
| Abbot Diagnostics | USA | RT-LAMP | Assay amplifies target regions of the SARS-CoV-2 RdRp and N gene with the same fluorophore within a single well. | [214,215] |
| Bio-Rad Laboratories DRG Diagnostics GmbH Euroimmun US Inc. |
USA | ELISA | Viral antigens are immobilized on plate to catch antibodies from patient’s antibodies | [213,216] |
| Mammoth Biosciences/ Sherlock Biosciences | USA | CRISPR | Rapid detection of SARS-CoV-2 | [217] |
| Identify Sensors | USA | Check 4 | Rapid detection of SARS-CoV-2 |
www.statnano.com |
| NanoEnTek | South Korea | Frend COVID-19 Ag | Rapid detection of SARS-CoV-2 | |
| Zepto Life Technology | USA | COVID-19 Test | Rapid detection of SARS-CoV-2 |
|
| Mologic Ltd | UK |
COVIS-19 test | Rapid detection of SARS-CoV-2 | |
| Lucence Diagnostics Pte Ltd. | Singapore | SAFER Sample Kit | Rapid detection of SARS-CoV-2 | |
| Graphene Leaders Canada | Canada | GLCM Biosensor test | Rapid detection of SARS-CoV-2 | |
| Nanoshel | India | Hand Sanitizer | Anti-viral | |
| Curran Biotech | USA | Filter | Capture coating | |
| Medicevo Corporation | India | Face mask with graphene | Virus control and personal protection | |
| Flextrapower | USA | Graphene mask | Virus control and personal protection |
|
| Integricote | USA | Respiratory mask | Virus control and personal protection | |
| Master Dynamic Limited | China | Diamond face mask |
Virus removal | |
| Yamashin Filter Corp | Japan | Nanofiber mask |
Virus removal | |
| MVX Prime Ltd. | UK | Nano mask | Anti-viral Anti-bacterial |
| Name of the company | Name of the vaccine | Adult dosage (www.who.int) |
Ingredients |
|---|---|---|---|
| Biontech/Pfizer (USA) |
BNT162b2 | 2 doses of 30µg each Intramuscularly |
Nucleoside-modified mRNA encoding the viral spike (S) glycoprotein of SARS-CoV-2 [150], DSPC, cholesterol, ALC-0315, ALC-0159 [218] |
| Moderna (USA) |
mRNA-1273 | 2 doses of 50µg each Intramuscularly |
Nucleoside-modified mRNA encoding the viral spike (S) glycoprotein , DSPC, cholesterol, SM102, DMG-PEG2000 [218] |
| Novavax (India) |
NVX-CoV2337 | 2 doses of 0.5 ml each Intramuscularly |
SARS-CoV-2 recombinant spike protein [140] |
| J&J/Janssen (USA) |
Ad26.CoV2.S | 1 dose of 0.5 ml each Intramuscularly |
Recombinant, replication-incompetent Ad26 vector, encoding a stabilized variant of the SARS-CoV-2 Spike (S) protein. [126] |
| Sinovac (China) |
Coronavac | 2 doses of 0.5 ml each Intramuscularly |
Aluminum hydroxide–adjuvanted, inactivated whole-virus vaccine [138] |
| Gamaleya (Russia) |
Sputnik V | 2 doses of 0.5 ml each Intramuscularly |
Recombinant, replication-incompetent Ad26 vector, encoding a stabilized variant of the SARS-CoV-2 Spike (S) protein. [219] |
| Bharat Biotech (India) |
Covaxin | 2 doses of 0.5 ml each Intramuscularly |
Alhydroxiquim-II–adjuvanted, inactivated whole-virus vaccine [138] |
| Astrazenica (UK) |
Covishield | 2 doses of 0.5 ml each Intramuscularly |
Recombinant, replication-incompetent ChAdOx1 vector, encoding a stabilized variant of the SARS-CoV-2 Spike (S) protein. [220] |
| Sinopharm (China) |
BBIBP-CorV | 2 doses of 0.5 ml each Intramuscularly |
Aluminum hydroxide–adjuvanted, inactivated whole-virus vaccine [138] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




