Submitted:
03 February 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
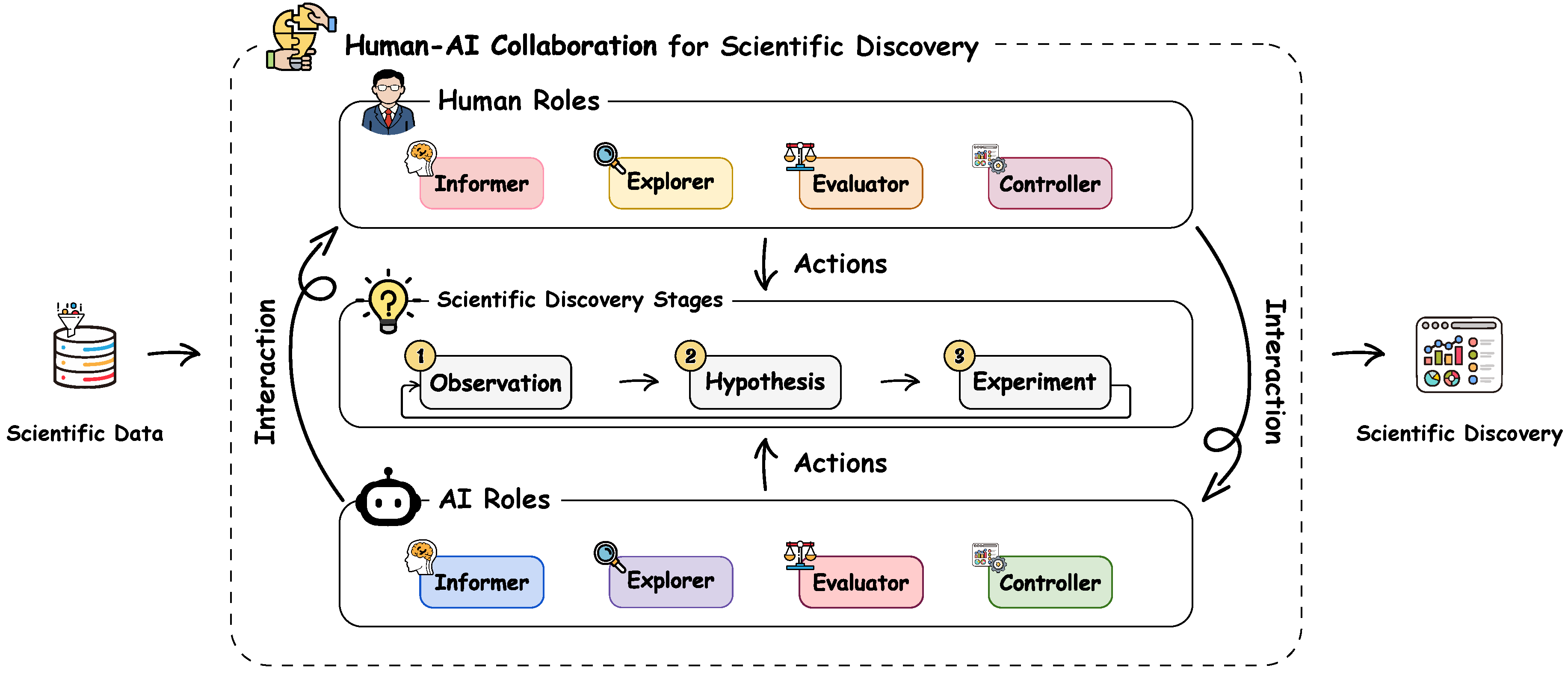
- We presented a comprehensive survey that systematically reviews human-AI collaboration specifically in scientific discovery.
- We introduced a novel taxonomy that defines the four roles of human and AI and characterized their distinct collaboration patterns across the three stages of scientific discovery.
- We identified critical challenges and future pathways for building human-AI partnerships in the scientific discovery process.
2. Methodology
2.1. Paper Collection
2.2. Paper Coding
3. Taxonomy
3.1. Roles of Human and AI
- ⋄Informer. The Informer synthesizes, distills, or articulates key information, insights, or constraints from raw data or intermediate analyses to guide the actions of other roles. For instance, the AI-Informer in THALIS extracts temporal patterns from longitudinal symptom records in cancer therapy [17]. This provides a summarized trajectory view for experts to analyze patient responses to treatment. Similarly, in ISHMAP for Mars rover operations [18], the human-Informer marks instrument states and operational events on the telemetry timeline. The AI uses these annotations to reduce false alarms during state changes and to highlight unexpected signals.
- ⋄Explorer. The Explorer operates within the space of data patterns, hypotheses, or experimental designs to explore promising candidates or directions. Compared with Informer, whose output provides low-level data insights, the Explorer directly generates candidates tailored to the specific needs of each stage in the scientific discovery process. For instance, in the hAE interface [7], the AI-Explorer searches the parameter space to select the next experimental conditions for electron microscopy. ChemVA [19] enables the human-Explorer to interactively navigate a projected chemical space to identify molecular targets.
- ⋄Evaluator. Once artifacts are proposed, their scientific merit must be rigorously evaluated and even refined. The Evaluator assesses observed patterns, hypotheses, or experimental designs based on predefined criteria, evidence, and domain constraints, revising them as necessary to meet quality standards. For instance, in RetroLens [20], chemists serve as the human-Evaluator. Specifically, they can assess AI-predicted synthetic routes for chemical feasibility or refine the synthetic steps by themselves.
- ⋄Controller. The Controller oversees the scientific discovery workflow to ensure correct and constraint-compliant execution, intervening when necessary to adjust procedures and handle runtime exceptions. This role is central to BIA [21], where the AI-Controller orchestrates the execution of complex bioinformatics toolchains, dynamically handling errors and modifying the workflow logic to ensure successful completion.
3.2. Common Collaboration Patterns Within Each Stage of Scientific Discovery
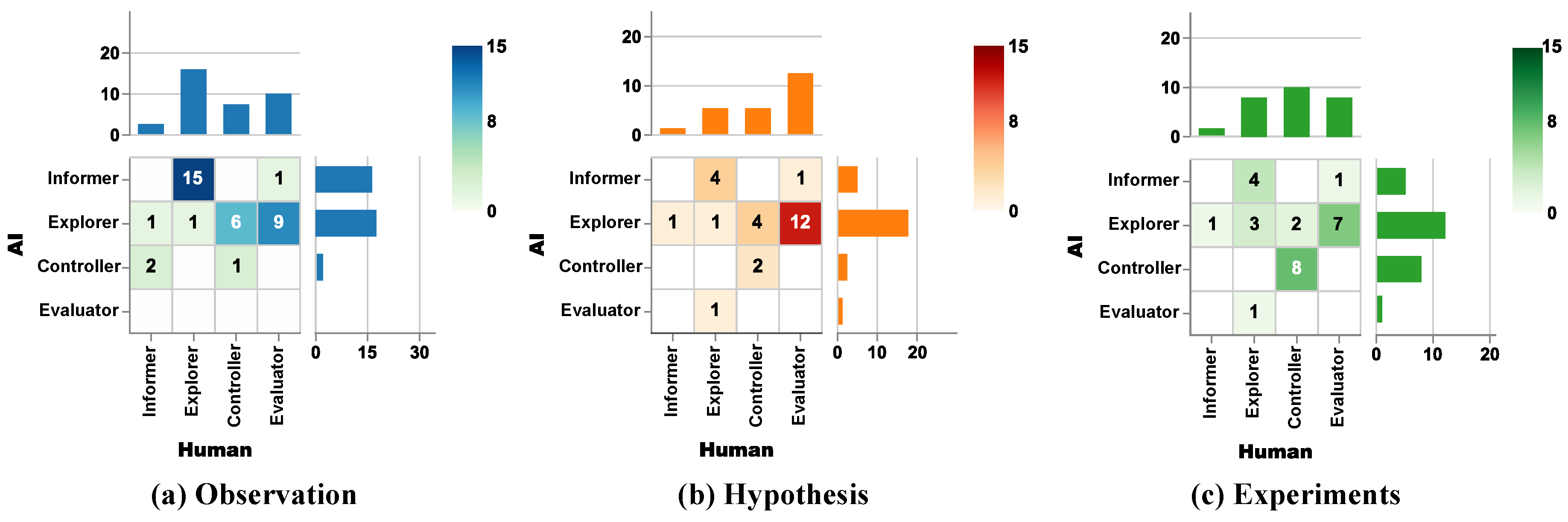
3.2.1. Observation Stage
3.2.2. Hypothesis Stage
3.2.3. Experiment Stage
3.2.4. Role Differences Across Three Stages
4. Discussion
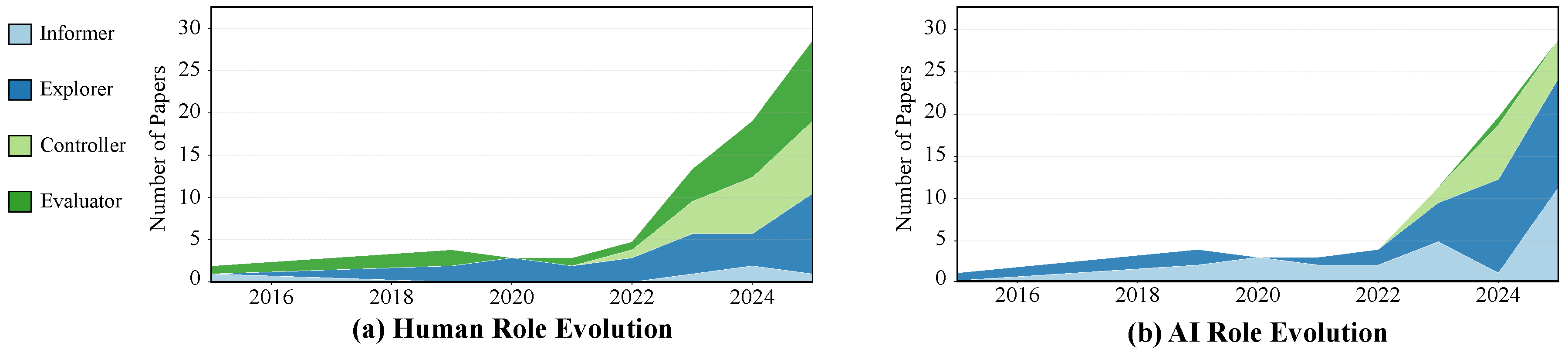
4.1. From Asymmetric Growth to Symbiotic Evolution
4.2. Generative Interfaces for Supporting Human Involvement
4.3. Adaptive Role Assignments Between Human and AI
4.4. Empowering Embodied AI in Scientific Experiments
4.5. Long-Term Implications for Leveraging AI in Scientific Discovery
5. Conclusion
6. Limitations
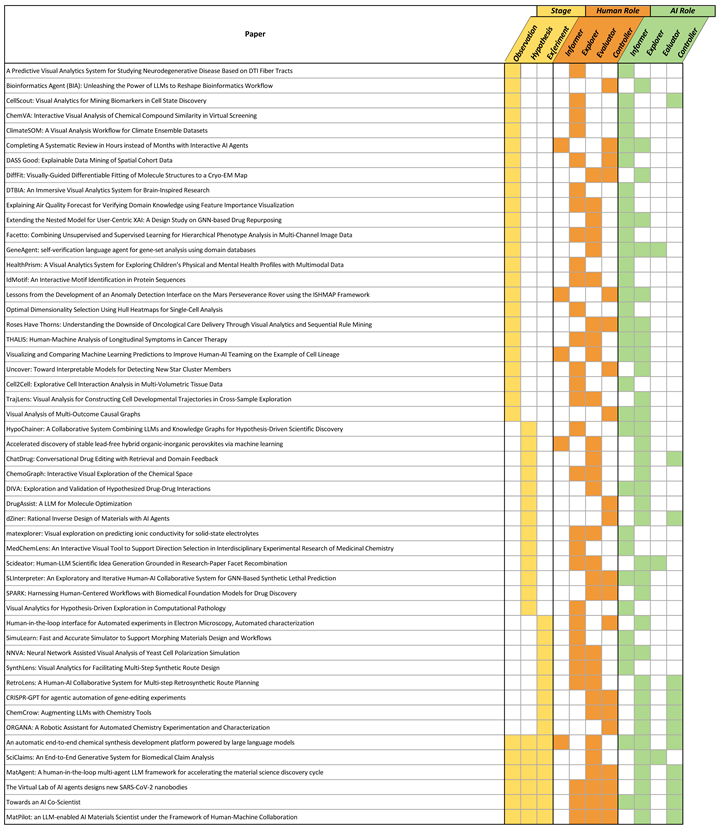
Appendix A. Coding Results
References
- Zhang, Y.; Chen, X.; Jin, B.; Wang, S.; Ji, S.; Wang, W.; Han, J. A Comprehensive Survey of Scientific Large Language Models and Their Applications in Scientific Discovery. In Proceedings of the Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing, 2024, pp. 8783–8817. [CrossRef]
- Tang, J.; Xia, L.; Li, Z.; Huang, C. AI-Researcher: Autonomous Scientific Innovation. In Proceedings of the The Thirty-ninth Annual Conference on Neural Information Processing Systems, 2025.
- Ye, F.; Zheng, Z.; Xue, D.; Shen, Y.; Wang, L.; Ma, Y.; Wang, Y.; Wang, X.; Zhou, X.; Gu, Q. ProteinBench: A Holistic Evaluation of Protein Foundation Models. In Proceedings of the International Conference on Representation Learning; Yue, Y.; Garg, A.; Peng, N.; Sha, F.; Yu, R., Eds., 2025, Vol. 2025, pp. 29857–29891.
- Zhang, Y.; Wei, Z.; Yuan, Y.; Li, C.; Huang, W. EquiPocket: an E(3)-Equivariant Geometric Graph Neural Network for Ligand Binding Site Prediction. In Proceedings of the Proceedings of the 41st International Conference on Machine Learning; Salakhutdinov, R.; Kolter, Z.; Heller, K.; Weller, A.; Oliver, N.; Scarlett, J.; Berkenkamp, F., Eds., 2024, Vol. 235, pp. 60021–60039.
- Ma, Y.; Gou, Z.; Hao, J.; Xu, R.; Wang, S.; Pan, L.; Yang, Y.; Cao, Y.; Sun, A. SciAgent: Tool-augmented Language Models for Scientific Reasoning. In Proceedings of the Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing; Al-Onaizan, Y.; Bansal, M.; Chen, Y.N., Eds., 2024, pp. 15701–15736. [CrossRef]
- Yao, S.; Zhao, J.; Yu, D.; Du, N.; Shafran, I.; Narasimhan, K.; Cao, Y. ReAct: Synergizing Reasoning and Acting in Language Models. In Proceedings of the International Conference on Learning Representations, 2023.
- Pratiush, U.; Duscher, G.; Kalinin, S. Human-in-the-loop interface for Automated experiments in Electron Microscopy, Automated characterization. In Proceedings of the AI for Accelerated Materials Design - NeurIPS 2024, 2024.
- Jansen, P.; Tafjord, O.; Radensky, M.; Siangliulue, P.; Hope, T.; Dalvi Mishra, B.; Majumder, B.P.; Weld, D.S.; Clark, P. CodeScientist: End-to-End Semi-Automated Scientific Discovery with Code-based Experimentation. 2025, pp. 13370–13467. [CrossRef]
- Zheng, T.; Deng, Z.; Tsang, H.T.; Wang, W.; Bai, J.; Wang, Z.; Song, Y. From Automation to Autonomy: A Survey on Large Language Models in Scientific Discovery. 2025, pp. 17744–17761. [CrossRef]
- Reddy, C.K.; Shojaee, P. Towards scientific discovery with generative AI: progress, opportunities, and challenges. In Proceedings of the Proceedings of the Thirty-Ninth AAAI Conference on Artificial Intelligence. AAAI Press, 2025. [CrossRef]
- Huang, C.; Deng, Y.; Lei, W.; Lv, J.; Chua, T.S.; Huang, J. How to Enable Effective Cooperation Between Humans and NLP Models: A Survey of Principles, Formalizations, and Beyond. In Proceedings of the Proceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers); Che, W.; Nabende, J.; Shutova, E.; Pilehvar, M.T., Eds., 2025, pp. 466–488. [CrossRef]
- Rajashekar, N.C.; Shin, Y.E.; Pu, Y.; Chung, S.; You, K.; Giuffre, M.; Chan, C.E.; Saarinen, T.; Hsiao, A.; Sekhon, J.; et al. Human-Algorithmic Interaction Using a Large Language Model-Augmented Artificial Intelligence Clinical Decision Support System. In Proceedings of the Proceedings of the 2024 CHI Conference on Human Factors in Computing Systems, 2024, CHI ’24. [CrossRef]
- Li, H.; Wang, Y.; Qu, H. Where Are We So Far? Understanding Data Storytelling Tools from the Perspective of Human-AI Collaboration. In Proceedings of the Proceedings of the 2024 CHI Conference on Human Factors in Computing Systems, 2024. [CrossRef]
- Mohanty, V.; Lim, J.; Luther, K. What Lies Beneath? Exploring the Impact of Underlying AI Model Updates in AI-Infused Systems. In Proceedings of the Proceedings of the 2025 CHI Conference on Human Factors in Computing Systems, 2025, CHI ’25. [CrossRef]
- Wang, H.; Fu, T.; Du, Y.; Gao, W.; Huang, K.; Liu, Z.; Chandak, P.; Liu, S.; Van Katwyk, P.; Deac, A.; et al. Scientific discovery in the age of artificial intelligence. Nature 2023, 620, 47–60. [CrossRef]
- Wang, X.; Huey, S.L.; Sheng, R.; Mehta, S.; Wang, F. SciDaSynth: Interactive Structured Data Extraction From Scientific Literature With Large Language Model. Campbell Systematic Reviews 2025, 21, e70073. [CrossRef]
- Floricel, C.; Nipu, N.; Biggs, M.; Wentzel, A.; Canahuate, G.; Van Dijk, L.; Mohamed, A.; Fuller, C.; Marai, G. THALIS: Human-Machine Analysis of Longitudinal Symptoms in Cancer Therapy. IEEE Transactions on Visualization and Computer Graphics 2022, 28, 151–161. [CrossRef]
- Wright, A.P.; Nemere, P.; Galvin, A.; Chau, D.H.; Davidoff, S. Lessons from the Development of an Anomaly Detection Interface on the Mars Perseverance Rover using the ISHMAP Framework. In Proceedings of the Proceedings of the 28th International Conference on Intelligent User Interfaces, 2023, p. 91–105. [CrossRef]
- Sabando, M.V.; Ulbrich, P.; Selzer, M.; Byška, J.; Mičan, J.; Ponzoni, I.; Soto, A.J.; Ganuza, M.L.; Kozlíková, B. ChemVA: Interactive Visual Analysis of Chemical Compound Similarity in Virtual Screening. IEEE Transactions on Visualization and Computer Graphics 2021, 27, 891–901. [CrossRef]
- Shi, C.; Hu, Y.; Wang, S.; Ma, S.; Zheng, C.; Ma, X.; Luo, Q. RetroLens: A Human-AI Collaborative System for Multi-step Retrosynthetic Route Planning. 2023. [CrossRef]
- Xin, Q.; Kong, Q.; Ji, H.; Shen, Y.; Liu, Y.; Sun, Y.; Zhang, Z.; Li, Z.; Xia, X.; Deng, B.; et al. BioInformatics Agent (BIA): Unleashing the Power of Large Language Models to Reshape Bioinformatics Workflow. bioRxiv 2024, [https://www.biorxiv.org/content/early/2024/05/22/2024.05.22.595240.full.pdf]. [CrossRef]
- Kawakami, Y.; Cayan, D.; Liu, D.; Ma, K.L. ClimateSOM: a Visual Analysis Workflow for Climate Ensemble Datasets. IEEE Transactions on Visualization and Computer Graphics 2025, pp. 1–11. [CrossRef]
- Jeong, H.; Jeong, H.o.; Lee, S.; Jeong, W.K. Optimal Dimensionality Selection Using Hull Heatmaps for Single-Cell Analysis. Computer Graphics Forum 2025, 44, e70151. [CrossRef]
- Mörth, E.; Sidak, K.; Maliga, Z.; Möller, T.; Gehlenborg, N.; Sorger, P.; Pfister, H.; Beyer, J.; Krüger, R. Cell2Cell: Explorative Cell Interaction Analysis in Multi-Volumetric Tissue Data. IEEE Transactions on Visualization and Computer Graphics 2025, 31, 569–579. [CrossRef]
- Jiang, Z.; Chen, H.; Zhou, R.; Deng, J.; Zhang, X.; Zhao, R.; Xie, C.; Wang, Y.; Ngai, E.C. HealthPrism: A Visual Analytics System for Exploring Children’s Physical and Mental Health Profiles with Multimodal Data . IEEE Transactions on Visualization & Computer Graphics 2024, 30, 1205–1215. [CrossRef]
- Yao, J.H.; Li, M.; Liu, J.; Li, Y.; Feng, J.; Han, J.; Zheng, Q.; Feng, J.; Chen, S. DTBIA: An Immersive Visual Analytics System for Brain-Inspired Research. IEEE Transactions on Visualization and Computer Graphics 2025, 31, 3796–3808. [CrossRef]
- Wang, Q.; Ruan, S.; Sheng, R.; Wang, Y.; Zhu, M.; Qu, H. TrajLens: Visual Analysis for Constructing Cell Developmental Trajectories in Cross-Sample Exploration. IEEE Transactions on Visualization and Computer Graphics 2025, pp. 1–11. [CrossRef]
- Sheng, R.; Zang, Z.; Wang, J.; Luo, Y.; Chen, Z.; Zhou, Y.; Ruan, S.; Qu, H. CellScout: Visual Analytics for Mining Biomarkers in Cell State Discovery. IEEE Transactions on Visualization and Computer Graphics 2025, pp. 1–16. [CrossRef]
- Park, J.H.; Prasad, V.; Newsom, S.; Najar, F.; Rajan, R. IdMotif: An Interactive Motif Identification in Protein Sequences. IEEE Computer Graphics and Applications 2024, 44, 114–125. [CrossRef]
- Palaniyappan Velumani, R.; Xia, M.; Han, J.; Wang, C.; LAU, A.K.; Qu, H. AQX: Explaining Air Quality Forecast for Verifying Domain Knowledge using Feature Importance Visualization. In Proceedings of the Proceedings of the 27th International Conference on Intelligent User Interfaces, 2022, p. 720–733. [CrossRef]
- Xu, C.; Neuroth, T.; Fujiwara, T.; Liang, R.; Ma, K.L. A Predictive Visual Analytics System for Studying Neurodegenerative Disease Based on DTI Fiber Tracts. IEEE Transactions on Visualization and Computer Graphics 2023, 29, 2020–2035. [CrossRef]
- Krueger, R.; Beyer, J.; Jang, W.D.; Kim, N.W.; Sokolov, A.; Sorger, P.K.; Pfister, H. Facetto: Combining Unsupervised and Supervised Learning for Hierarchical Phenotype Analysis in Multi-Channel Image Data. IEEE Transactions on Visualization and Computer Graphics 2020, 26, 227–237. [CrossRef]
- Wang, Q.; Huang, K.; Chandak, P.; Zitnik, M.; Gehlenborg, N. Extending the Nested Model for User-Centric XAI: A Design Study on GNN-based Drug Repurposing. IEEE Transactions on Visualization and Computer Graphics 2023, 29, 1266–1276. [CrossRef]
- Luo, D.; Alsuwaykit, Z.; Khan, D.; Strnad, O.; Isenberg, T.; Viola, I. DiffFit: Visually-Guided Differentiable Fitting of Molecule Structures to a Cryo-EM Map. IEEE Transactions on Visualization and Computer Graphics 2025, 31, 558–568. [CrossRef]
- Floricel, C.; Wentzel, A.; Mohamed, A.; Fuller, C.; Canahuate, G.; Marai, G. Roses Have Thorns: Understanding the Downside of Oncological Care Delivery Through Visual Analytics and Sequential Rule Mining. IEEE Transactions on Visualization and Computer Graphics 2024, 30, 1227–1237. [CrossRef]
- Hong, J.; Maciejewski, R.; Trubuil, A.; Isenberg, T. Visualizing and Comparing Machine Learning Predictions to Improve Human-AI Teaming on the Example of Cell Lineage. IEEE Transactions on Visualization and Computer Graphics 2024, 30, 1956–1969. [CrossRef]
- Wang, Z.; Jin, Q.; Wei, C.H.; Tian, S.; Lai, P.T.; Zhu, Q.; Day, C.P.; Ross, C.; Leaman, R.; Lu, Z. GeneAgent: Self-Verification Language Agent for Gene-Set Analysis Using Domain Databases. Nature Methods 2025, 22, 1677–1685. [CrossRef]
- Qiu, R.; Chen, S.; Su, Y.; Yen, P.Y.; Shen, H.W. Completing A Systematic Review in Hours instead of Months with Interactive AI Agents. In Proceedings of the Proceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers); Che, W.; Nabende, J.; Shutova, E.; Pilehvar, M.T., Eds., 2025, pp. 31559–31593. [CrossRef]
- Ratzenböck, S.; Obermüller, V.; Möller, T.; Alves, J.; Bomze, I.M. Uncover: Toward Interpretable Models for Detecting New Star Cluster Members. IEEE Transactions on Visualization and Computer Graphics 2023, 29, 3855–3872. [CrossRef]
- Wentzel, A.; Floricel, C.; Canahuate, G.; Naser, M.A.; Mohamed, A.S.; Fuller, C.D.; van Dijk, L.; Marai, G.E. DASS Good: Explainable Data Mining of Spatial Cohort Data. Computer Graphics Forum 2023, 42, 283–295. [CrossRef]
- Fan, M.; Yu, J.; Weiskopf, D.; Cao, N.; Wang, H.Y.; Zhou, L. Visual Analysis of Multi-Outcome Causal Graphs. IEEE Transactions on Visualization and Computer Graphics 2025, 31, 656–666. [CrossRef]
- Liu, S.; Wang, J.; Yang, Y.; Wang, C.; Liu, L.; Guo, H.; Xiao, C. Conversational Drug Editing Using Retrieval and Domain Feedback. In Proceedings of the The Twelfth International Conference on Learning Representations (ICLR), 2024.
- Lu, S.; Zhou, Q.; Ouyang, Y.; Guo, Y.; Li, Q.; Wang, J. Accelerated Discovery of Stable Lead-Free Hybrid Organic-Inorganic Perovskites via Machine Learning. Nature Communications 2018, 9, 3405. [CrossRef]
- Ni, Z.; Li, Y.; Hu, K.; Han, K.; Xu, M.; Chen, X.; Liu, F.; Ye, Y.; Bai, S. MatPilot: An LLM-Enabled AI Materials Scientist Under the Framework of Human-Machine Collaboration. arXiv preprint arXiv:2411.08063 2024.
- Kale, B.; Clyde, A.; Sun, M.; Ramanathan, A.; Stevens, R.; Papka, M.E. ChemoGraph: Interactive Visual Exploration of the Chemical Space. 2023, Vol. 42, pp. 13–24. [CrossRef]
- Swanson, K.; Wu, W.; Bulaong, N.L.; Pak, J.E.; Zou, J. The Virtual Lab of AI agents designs new SARS-CoV-2 nanobodies. Nature 2025, 646, 716–723. [CrossRef]
- Wang, Q.; Schaeffer, R.; Khani, F.; Ilievski, F.; et al. Scideator: Human-LLM Scientific Idea Generation Grounded in Research-Paper Facet Recombination. Transactions on Machine Learning Research 2025.
- Kakar, T.; Qin, X.; Rundensteiner, E.A.; Harrison, L.; Sahoo, S.K.; De, S. DIVA: Exploration and Validation of Hypothesized Drug-Drug Interactions. In Proceedings of the Computer Graphics Forum, 2019, Vol. 38, pp. 95–106. [CrossRef]
- Ortega, R.; Gomez-Perez, J.M. SciClaims: An End-to-End Generative System for Biomedical Claim Analysis. In Proceedings of the Proceedings of the 2025 Conference on Empirical Methods in Natural Language Processing: System Demonstrations; Habernal, I.; Schulam, P.; Tiedemann, J., Eds., 2025, pp. 141–154. [CrossRef]
- Ansari, M.; Watchorn, J.; Brown, C.E.; Brown, J.S. dZiner: Rational Inverse Design of Materials with AI Agents. In Proceedings of the AI for Accelerated Materials Design - NeurIPS 2024, 2024.
- Ye, G.; Cai, X.; Lai, H.; Wang, X.; Huang, J.; Wang, L.; Liu, W.; Zeng, X. DrugAssist: a large language model for molecule optimization. Briefings in Bioinformatics 2025, 26, bbae693. [CrossRef]
- Kwon, B.C.; Rabinovici-Cohen, S.; Moturi, B.; Mwaura, R.; Wahome, K.; Njeru, O.; Shinyenyi, M.; Wanjiru, C.; Remy, S.; Ogallo, W.; et al. SPARK: harnessing human-centered workflows with biomedical foundation models for drug discovery. 2024, IJCAI ’24. [CrossRef]
- Jiang, H.; Shi, S.; Zhang, S.; Zheng, J.; Li, Q. SLInterpreter: An Exploratory and Iterative Human-AI Collaborative System for GNN-Based Synthetic Lethal Prediction. IEEE Transactions on Visualization and Computer Graphics 2025, 31, 919–929. [CrossRef]
- Jiang, H.; Shi, S.; Yao, Y.; Jiang, C.; Li, Q. HypoChainer: a Collaborative System Combining LLMs and Knowledge Graphs for Hypothesis-Driven Scientific Discovery. IEEE Transactions on Visualization and Computer Graphics 2025, pp. 1–11. [CrossRef]
- Shi, C.; Nie, F.; Hu, Y.; Xu, Y.; Chen, L.; Ma, X.; Luo, Q. MedChemLens: An Interactive Visual Tool to Support Direction Selection in Interdisciplinary Experimental Research of Medicinal Chemistry. IEEE Transactions on Visualization and Computer Graphics 2023, 29, 63–73. [CrossRef]
- Corvò, A.; Caballero, H.S.G.; Westenberg, M.A.; van Driel, M.A.; van Wijk, J.J. Visual Analytics for Hypothesis-Driven Exploration in Computational Pathology. IEEE Transactions on Visualization and Computer Graphics 2021, 27, 3851–3866. [CrossRef]
- Pu, J.; Shao, H.; Gao, B.; Zhu, Z.; Zhu, Y.; Rao, Y.; Xiang, Y. matExplorer: Visual Exploration on Predicting Ionic Conductivity for Solid-state Electrolytes. IEEE Transactions on Visualization and Computer Graphics 2022, 28, 65–75. [CrossRef]
- Darvish, K.; Skreta, M.; Zhao, Y.; Yoshikawa, N.; Som, S.; Bogdanovic, M.; Cao, Y.; Hao, H.; Xu, H.; Aspuru-Guzik, A.; et al. ORGANA: A robotic assistant for automated chemistry experimentation and characterization. Matter 2025, 8, 101897. [CrossRef]
- Bran, A.M.; Cox, S.; Schilter, O.; Baldassari, C.; White, A.D.; Schwaller, P. Augmenting large language models with chemistry tools. Nature Machine Intelligence 2024, 6, 525–535. [CrossRef]
- Boiko, D.A.; MacKnight, R.; Kline, B.; Gomes, G. Autonomous chemical research with large language models. Nature 2023, 624, 570–578. [CrossRef]
- Qu, Y.; Huang, K.; Yin, M.; Zhan, K.; Liu, D.; Yin, D.; Cousins, H.C.; Johnson, W.A.; Wang, X.; Shah, M.; et al. CRISPR-GPT for agentic automation of gene-editing experiments. Nature Biomedical Engineering 2025. [CrossRef]
- Bazgir, A.; chandra Praneeth Madugula, R.; Zhang, Y. MatAgent: A human-in-the-loop multi-agent LLM framework for accelerating the material science discovery cycle. In Proceedings of the AI for Accelerated Materials Design - ICLR 2025, 2025.
- Gottweis, J.; Weng, W.H.; Daryin, A.; Tu, T.; Palepu, A.; Sirkovic, P.; Myaskovsky, A.; Weissenberger, F.; Rong, K.; Tanno, R.; et al. Towards an AI Co-Scientist. arXiv preprint arXiv:2502.18864 2025.
- Yang, H.; Qian, K.; Liu, H.; Yu, Y.; Gu, J.; McGehee, M.; Zhang, Y.J.; Yao, L. SimuLearn: Fast and Accurate Simulator to Support Morphing Materials Design and Workflows. In Proceedings of the Proceedings of the 33rd Annual ACM Symposium on User Interface Software and Technology, 2020, UIST ’20, p. 71–84. [CrossRef]
- Hazarika, S.; Li, H.; Wang, K.C.; Shen, H.W.; Chou, C.S. NNVA: Neural Network Assisted Visual Analysis of Yeast Cell Polarization Simulation. IEEE Transactions on Visualization and Computer Graphics 2020, 26, 34–44. [CrossRef]
- Wang, Q.; Sheng, R.; Ruan, S.; Jin, X.; Shi, C.; Zhu, M. SynthLens: Visual Analytics for Facilitating Multi-Step Synthetic Route Design. IEEE Transactions on Visualization and Computer Graphics 2025, 31, 7647–7660. [CrossRef]
- Xie, Q.; Li, Q.; Yu, Z.; Zhang, Y.; Zhang, Y.; Yang, L. An Empirical Analysis of Uncertainty in Large Language Model Evaluations. In Proceedings of the The Thirteenth International Conference on Learning Representations, 2025.
- Heo, J.; Xiong, M.; Heinze-Deml, C.; Narain, J. Do LLMs estimate uncertainty well in instruction-following? In Proceedings of the The Thirteenth International Conference on Learning Representations, 2025.
- Chen, Y.; Zhang, C.; Luo, D.; D’Haro, L.F.; Tan, R.; Li, H. Unveiling the Achilles’ Heel of NLG Evaluators: A Unified Adversarial Framework Driven by Large Language Models. In Proceedings of the Findings of the Association for Computational Linguistics: ACL 2024, 2024, pp. 1359–1375. [CrossRef]
- Rezaei, M.; Fu, Y.; Cuvin, P.; Ziems, C.; Zhang, Y.; Zhu, H.; Yang, D. EgoNormia: Benchmarking Physical-Social Norm Understanding. In Proceedings of the Findings of the Association for Computational Linguistics: ACL 2025, 2025, pp. 19256–19283. [CrossRef]
- Chen, J.; Zhang, Y.; Zhang, Y.; Shao, Y.; Yang, D. Generative Interfaces for Language Models. arXiv preprint arXiv:2508.19227 2025.
- Sheng, R.; Wang, X.; Wang, J.; Jin, X.; Sheng, Z.; Xu, Z.; Rajendran, S.; Qu, H.; Wang, F. TrialCompass: Visual Analytics for Enhancing the Eligibility Criteria Design of Clinical Trials. IEEE Transactions on Visualization and Computer Graphics 2025, pp. 1–11. [CrossRef]
- Luro, S.; Potvin-Trottier, L.; Okumus, B.; Paulsson, J. Isolating live cells after high-throughput, long-term time-lapse microscopy. Nature Methods 2020, 17, 93–100. [CrossRef]
- Wright, P.M.; Seiple, I.B.; Myers, A.G. The Evolving Role of Chemical Synthesis in Antibacterial Drug Discovery. Angewandte Chemie International Edition 2014, 53, 8840–8869. [CrossRef]
- Schmidgall, S.; Su, Y.; Wang, Z.; Sun, X.; Wu, J.; Yu, X.; Liu, J.; Moor, M.; Liu, Z.; Barsoum, E. Agent Laboratory: Using LLM Agents as Research Assistants. In Proceedings of the Findings of the Association for Computational Linguistics: EMNLP 2025; Christodoulopoulos, C.; Chakraborty, T.; Rose, C.; Peng, V., Eds., 2025, pp. 5977–6043. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



