Submitted:
05 December 2025
Posted:
09 December 2025
You are already at the latest version
Abstract
Breast cancer remains a major global health challenge, strongly influenced by inequities in access to high-cost antibody-based diagnostic and therapeutic technologies. Because their manufacturing requires complex and expensive bioproduction systems, many regions, particularly in Latin America, face limited availability of tools essential for early detection and targeted treatment. This gap has highlighted the need for cost-effective and scalable theranostic alternatives, driving interest in aptamers, synthetic DNA or RNA oligomers selected through SELEX technology that fold into functional structures with high affinity, low immunogenicity, and reproducible chemical synthesis. Aptamers have expanded the landscape of diagnostic platforms by enabling sensitive aptasensors, liquid biopsy strategies, and nanomedicine-based imaging systems. They also contribute to targeted therapy by recognizing cancer biomarkers with precision and enabling controlled drug delivery. This review critically integrates advances in aptamer-based theranostics for breast cancer, examining their molecular mechanisms, including structural folding, selective ligand binding, and nanomaterial interfacing. We also discuss their application in extracellular vesicle capture, cancer stem cell detection, and therapeutic conjugates, emphasizing their advantages and limitations relative to antibody-based approaches. By consolidating current evidence, this work positions aptamers as emerging tools capable of democratizing precision oncology, particularly in regions where access to advanced technologies remains limited.
Keywords:
1. Introduction
1.1. Aptamers: Development and Advantages
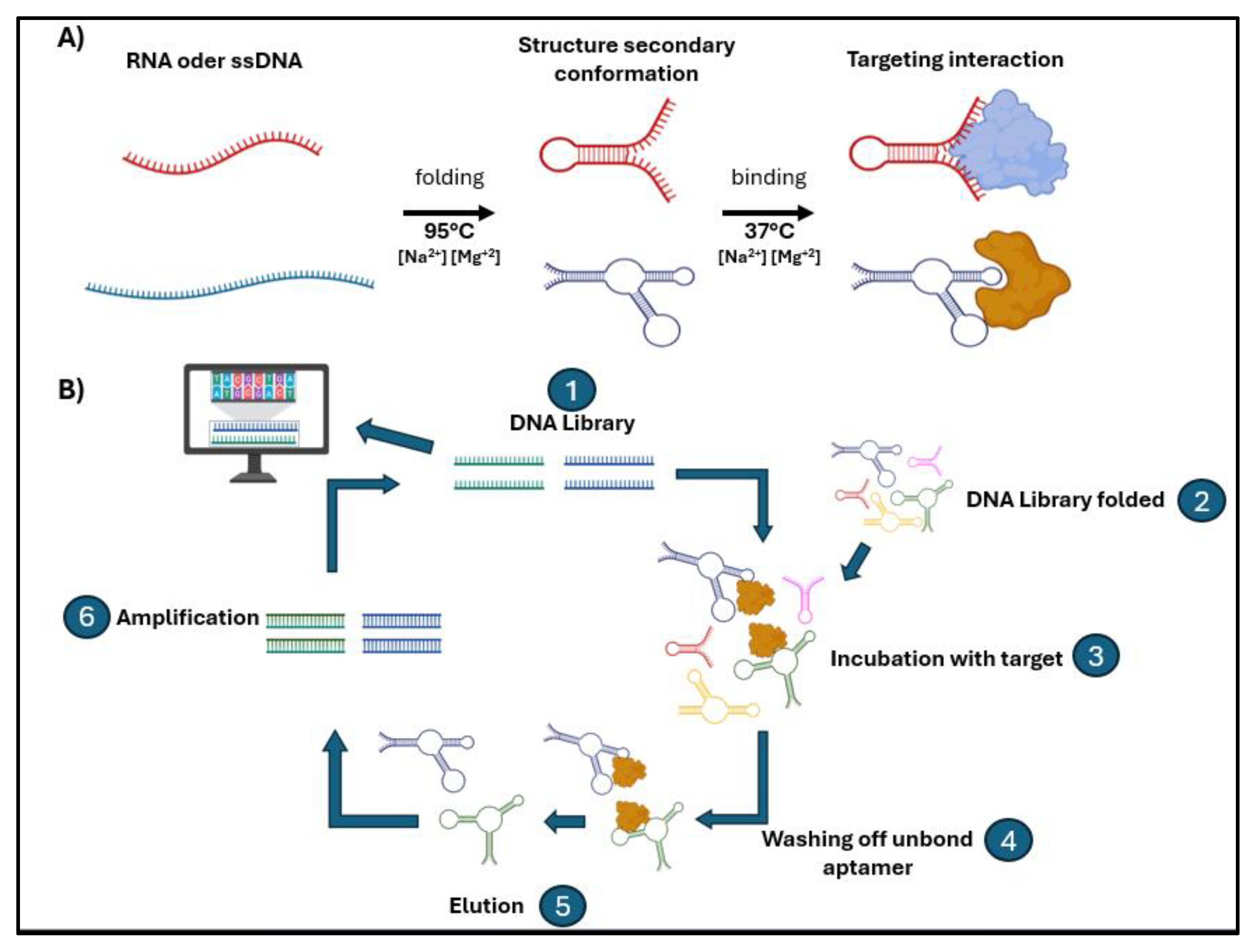
1.2. Advantage of Using Aptamers as Clinical Tools
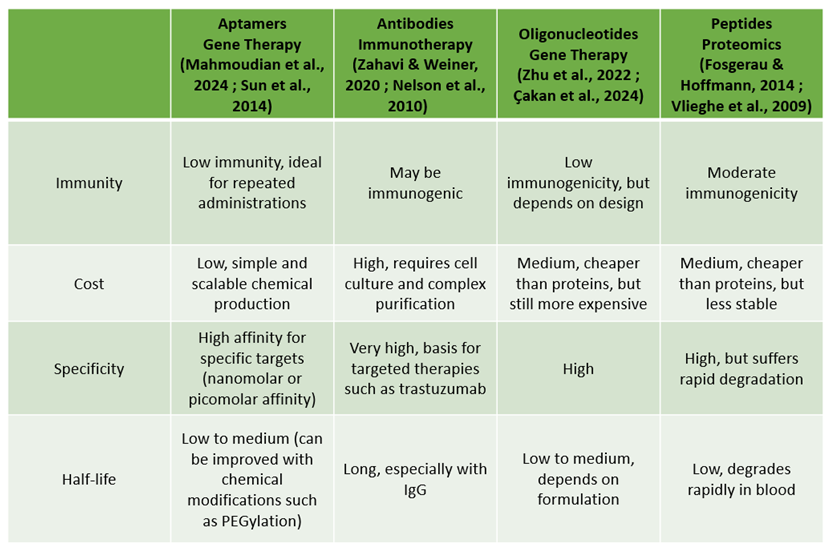 |
2. The Use of Aptamers in Breast Cancer Diagnostics
2.1. Aptamer Sensors in Solid Matrix
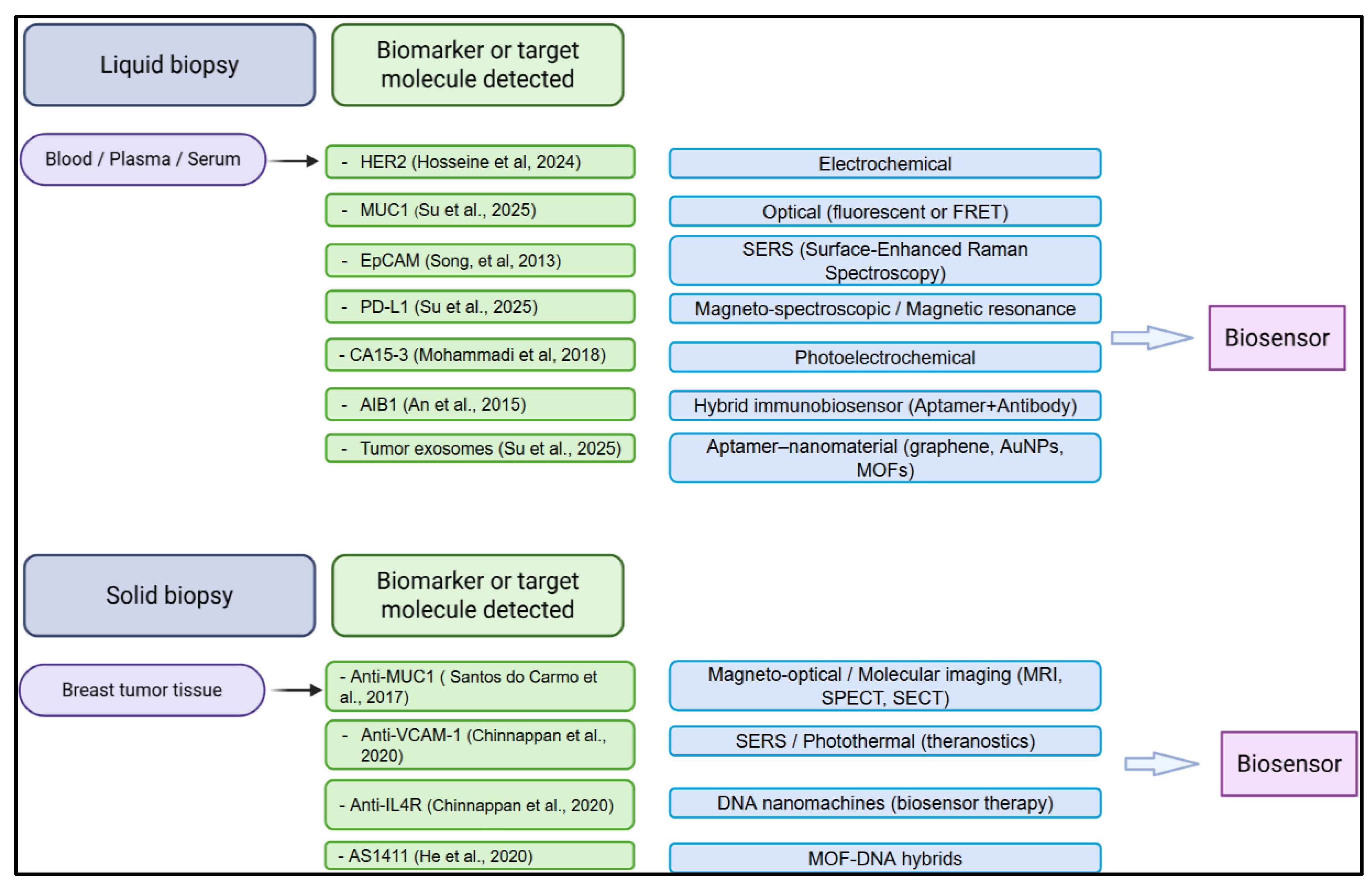
2.2. Aptasensor-Based Approaches for Liquid Biopsy and Early Detection of Breast Cancer
2.3. Aptamer Like a Target-Enhancer Imaging in Tumor
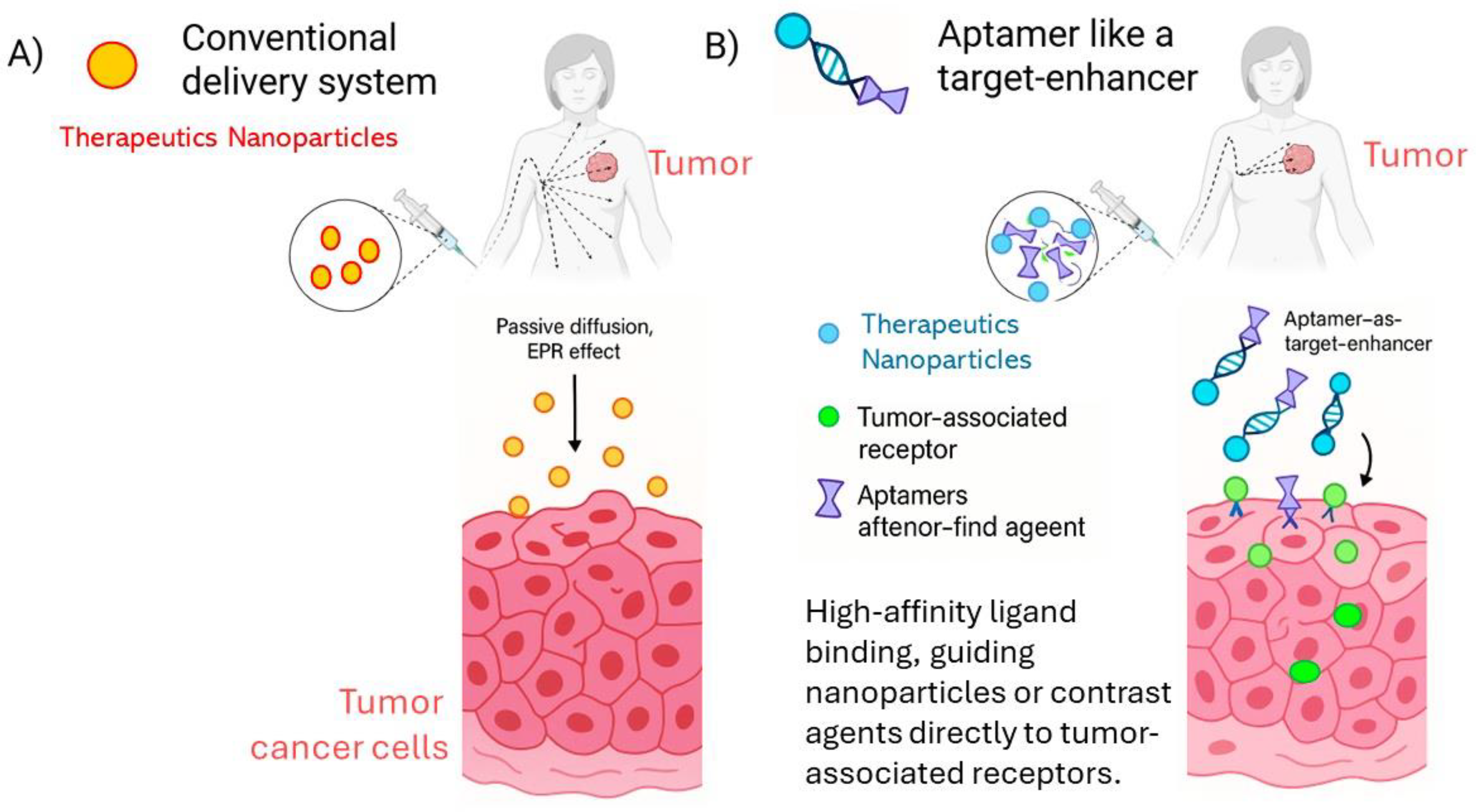
3. Aptamers Used in the Prognosis of Breast Cancer
3.1. Biomarkers for Quantifiable Prognosis
3.2. Aggressive Subpopulations and Cancer Stem Cells
3.3. Multiligand Signatures and Prediction of Clinical Outcomes
3.4. Oncogenes and Kinases with Prognostic Relevance
3.5. Tumor Microenvironment and Immunosuppression
4. Aptamers Used in Breast Cancer Therapy
4.1. Aptamers as Therapeutic Agents and Specific Target Modulators
4.2. Aptamers in Chemotherapeutic Drug Delivery Systems
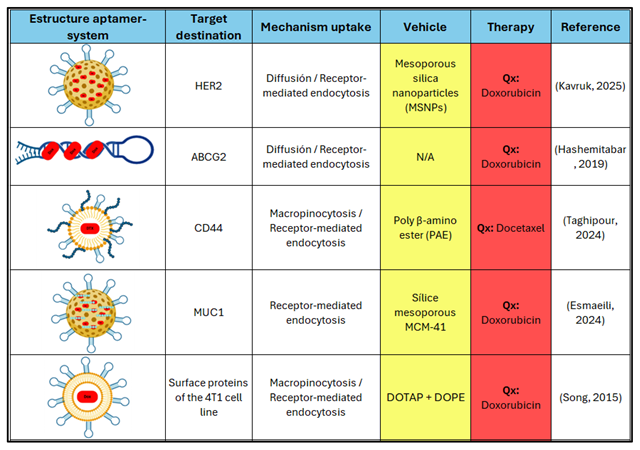 |
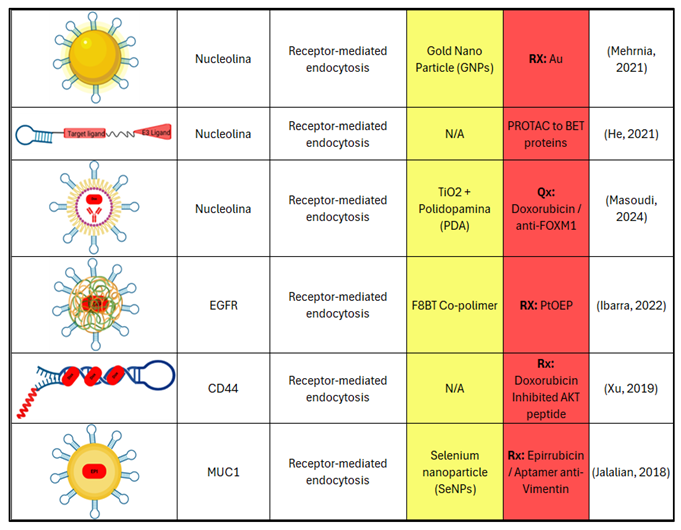 |
4.3. Combined Therapies with Chemotherapy, Radiotherapy, and Immunotherapy
4.4. Other Therapeutic Modalities
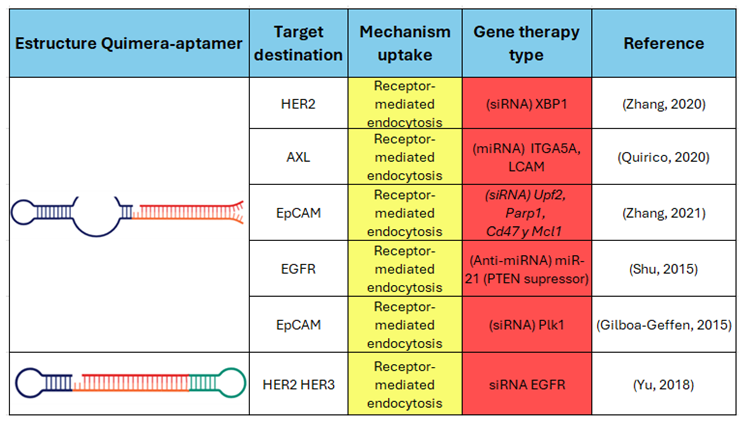 |
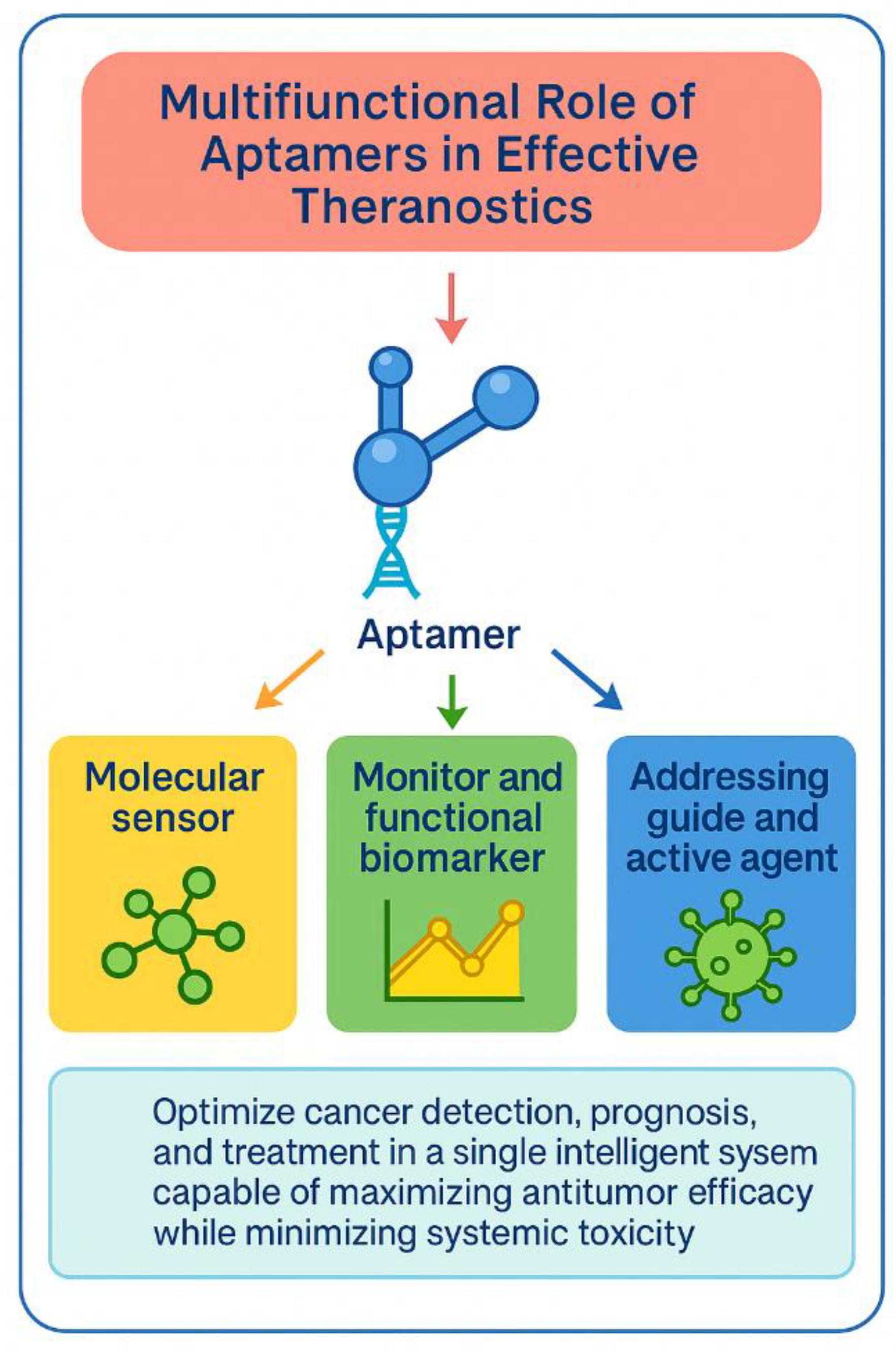
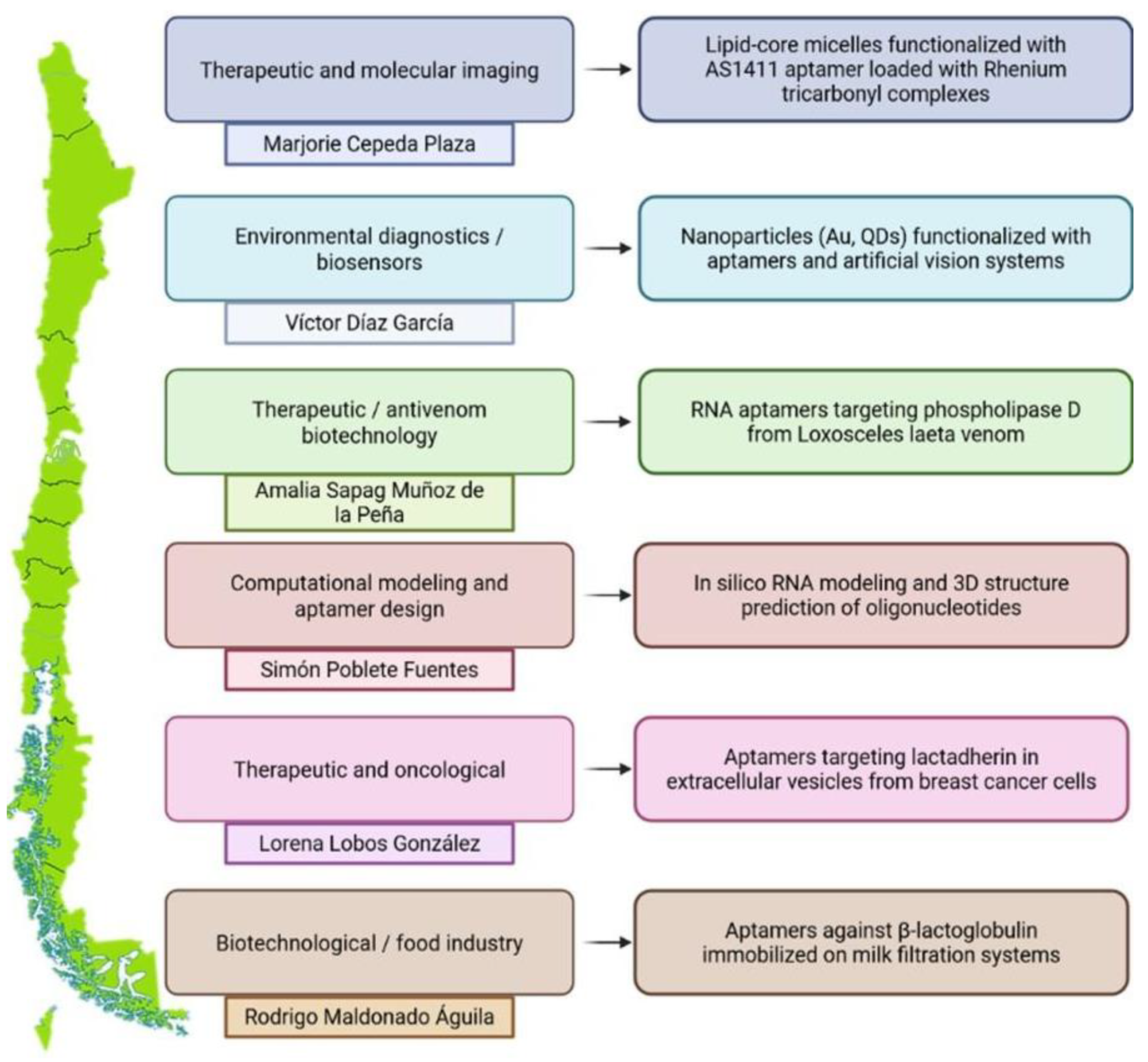
5. Perspectives and Challenges in Latin America
6. Conclusion
Abbreviations
| Breast cancer subtypes & clinical markers | |
| TNBC | Triple-Negative Breast Cancer |
| HER2 | Human Epidermal Growth Factor Receptor 2 |
| ER | Estrogen Receptor |
| PR | Progesterone Receptor |
| Liquid biopsy, vesicles & imaging | |
| CTC(s) | Circulating Tumor Cell(s) |
| EV(s) | Extracellular Vesicle(s) |
| SERS | Surface-Enhanced Raman Spectroscopy |
| SECT | Sequential Emission Computed Tomography |
| SPECT | Single Photon Emission Computed Tomography |
| MRI | Magnetic Resonance Imaging |
| Tumor biomarkers & receptors | |
| MUC1 | Mucin 1 |
| EpCAM | Epithelial Cell Adhesion Molecule |
| PD-L1 | Programmed Death-Ligand 1 |
| PDGFRβ | Platelet-Derived Growth Factor Receptor Beta |
| AIB1 | Amplified in Breast Cancer 1 |
| NCL | Nucleolin |
| OPN | Osteopontin |
| VRK1 | Vaccinia-Related Kinase 1 |
| MNK1b | MAP Kinase-Interacting Kinase 1b |
| Aptamer platforms & molecular tools | |
| SELEX | Systematic Evolution of Ligands by Exponential Enrichment |
| Cell-SELEX | Cell-based SELEX |
| Exo-SELEX | Exosome-guided SELEX |
| AS1411 | Nucleolin-Binding Aptamer |
| Q10 | Anti-metastatic Exo-SELEX Aptamer |
| Axl-148b | Anti-Axl Aptamer |
| Therapeutic systems & nanotechnology | |
| siRNA | Small Interfering RNA |
| PROTAC | Proteolysis-Targeting Chimera |
| APC | Aptamer–PROTAC Conjugate |
| DOX | Doxorubicin |
| PTX | Paclitaxel |
| DOC | Docetaxel |
| PEG | Polyethylene Glycol |
| MOF | Metal–Organic Framework |
| ATP | Adenosine Triphosphate |
| NIR | Near-Infrared |
| Cell lines | |
| MCF-7 | Human Breast Adenocarcinoma Cells |
| SK-BR-3 | HER2-Positive Breast Cancer Cells |
| MDA-MB-231 | Triple-Negative Breast Cancer Cells |
| Nanomaterials | |
| Fe₃O₄ | Magnetite Nanoparticles |
| AuNPs | Gold Nanoparticles |
| Ag–Au | Silver–Gold Nanostructures |
| QDs | Quantum Dots |
| DNA-Au | DNA–Gold Nanomachine |
| Others | |
| miRNA | MicroRNA |
| TME | Tumor Microenvironment |
| EPR | Enhanced Permeability and Retention Effect |
References
- World Health Organization (WHO); World Health Organization. Breast Cancer: Key Facts – Updated 2024; Geneva, 2024. [Google Scholar]
- Sung, H.; et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians 2021, 71(3), 209–249. [Google Scholar] [CrossRef]
- Bianchini, G.; et al. Triple-negative breast cancer: Challenges and opportunities of a heterogeneous disease. Nature Reviews Clinical Oncology 2016, 13(11), 674–690. [Google Scholar] [CrossRef]
- Ellington, A. D.; Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346(6287), 818–822. [Google Scholar] [CrossRef]
- Zhou, J.; Rossi, J. Aptamers as targeted therapeutics: Current potential and challenges. Nature Reviews Drug Discovery 2017, 16(3), 181–202. [Google Scholar] [CrossRef] [PubMed]
- Gao, F.; et al. Recent advances in aptamer-based targeted drug delivery systems for cancer therapy. Frontiers in Bioengineering and Biotechnology 2022, 10, 972933. [Google Scholar] [CrossRef] [PubMed]
- Ni, X.; et al. Nucleic acid aptamers: Clinical applications and promising new horizons. Current Medicinal Chemistry 2011, 18(27), 4206–4214. [Google Scholar] [CrossRef]
- Jo, H.; Ban, C. Aptamer-nanoparticle complexes as powerful diagnostic and therapeutic tools. Experimental & Molecular Medicine 2016, 48(5), e230. [Google Scholar] [CrossRef]
- Liu, Z.; et al. Novel HER2 aptamer selectively delivers cytotoxic drug to HER2-positive breast cancer cells in vitro. Journal of Translational Medicine 2012, 10, 148. [Google Scholar] [CrossRef]
- Tao, W.; et al. Polydopamine-based surface modification of novel nanoparticle-aptamer bioconjugates for in vivo breast cancer targeting and enhanced therapeutic effects. Theranostics 2016, 6(4), 470–484. [Google Scholar] [CrossRef]
- Wang, T.; et al. EpCAM Aptamer-mediated survivin silencing sensitized cancer stem cells to doxorubicin in a breast cancer model. Theranostics 2015, 5(12), 1456–1472. [Google Scholar]
- Hassan, E. M.; et al. Aptamers: Promising tools for the detection of circulating tumor cells. Nucleic Acid Therapeutics 2016, 26(6), 335–347. [Google Scholar] [CrossRef] [PubMed]
- Dzhumashev, D.; et al. Quantum dot-based screening identifies F3 peptide and reveals cell surface nucleolin as a therapeutic target for rhabdomyosarcoma. Cancers 2022, 14(20), 5048. [Google Scholar] [CrossRef]
- Chen, Z.; et al. Targeted immunotherapy of triple-negative breast cancer by aptamer-engineered NK cells. Biomaterials 2022, 280, 121259. [Google Scholar] [CrossRef]
- Camorani, S.; et al. Aptamer targeted therapy potentiates immune checkpoint blockade in triple-negative breast cancer. Journal of Experimental & Clinical Cancer Research 2020, 39(1), 180. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249(4968), 505–510. [Google Scholar] [CrossRef] [PubMed]
- Sefah, K.; et al. Development of DNA aptamers using Cell-SELEX. Nature Protocols 2010, 5(6), 1169–1185. [Google Scholar] [CrossRef]
- Maradani, B. S.; et al. Development and characterization of DNA aptamer against retinoblastoma by cell-SELEX and high-throughput sequencing. Scientific Reports 2022, 12, 20660. [Google Scholar] [CrossRef]
- Amero, P.; et al. Conversion of RNA aptamer into modified DNA aptamers provides for prolonged stability and enhanced antitumor activity. Journal of the American Chemical Society 2021, 143(20), 7655–7670. [Google Scholar] [CrossRef] [PubMed]
- Shraim, A. S.; et al. Therapeutic potential of aptamer–protein interactions. ACS Pharmacology & Translational Science 2022, 5(10), 989–1003. [Google Scholar] [CrossRef]
- Edwards, A. N.; et al. G-Quadruplex structure in the ATP-binding DNA aptamer strongly modulates ligand-binding activity. ACS Omega 2024, 9(2), 2105–2115. [Google Scholar] [CrossRef]
- Sun, H.; et al. Oligonucleotide aptamers: New tools for targeted cancer therapy. Molecular Therapy – Nucleic Acids 2014, 3, e182. [Google Scholar] [CrossRef] [PubMed]
- Mahmoudian, F.; et al. Aptamers as an approach to targeted cancer therapy. Cancer Cell International 2024, 24(1), 108. [Google Scholar] [CrossRef]
- Zahavi, D.; Weiner, L. Monoclonal antibodies in cancer therapy. Antibodies 2020, 9(3), 34. [Google Scholar] [CrossRef]
- Nelson, A. L.; et al. Development trends for human monoclonal antibody therapeutics. Nature Reviews Drug Discovery 2010, 9(10), 767–774. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; et al. RNA-based therapeutics: An overview and prospectus. Cell Death and Disease 2022, 13(7), 644. [Google Scholar] [CrossRef] [PubMed]
- Çakan, E.; et al. Therapeutic antisense oligonucleotides in oncology: From bench to bedside. Cancers 2024, 16(17), 2940. [Google Scholar] [CrossRef] [PubMed]
- Fosgerau, K.; Hoffmann, T. Peptide therapeutics: Current status and future directions. Drug Discovery Today 2014, 20(1), 122–128. [Google Scholar] [CrossRef]
- Vlieghe, P.; et al. Synthetic therapeutic peptides: Science and market. Drug Discovery Today 2009, 15(1-2), 40–56. [Google Scholar] [CrossRef]
- Domsicova, M.; et al. New insights into aptamers: An alternative to antibodies in the detection of molecular biomarkers. International Journal of Molecular Sciences 2024, 25(13), 6833. [Google Scholar] [CrossRef]
- Agnello, L.; et al. Aptamers and antibodies: Rivals or allies in cancer targeted therapy? Exploration of Targeted Anti-Tumor Therapy 2021, 2(1), 107–121. [Google Scholar] [CrossRef]
- Hosseine, M.; et al. Label-free electrochemical biosensor based on green-synthesized reduced graphene oxide/Fe3O4/nafion/polyaniline for ultrasensitive detection of SKBR3 cell line of HER2 breast cancer biomarker. Scientific Reports 2024, 14, 11928. [Google Scholar] [CrossRef]
- An, Y.; et al. Selection and application of DNA aptamer against oncogene amplified in breast cancer 1. Journal of Molecular Evolution 2015, 81(5-6), 179–185. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; et al. Selection of DNA aptamers against epithelial cell adhesion molecule for cancer cell imaging and circulating tumor cell capture. Analytical Chemistry 2013, 85(8), 4141–4149. [Google Scholar] [CrossRef]
- Albanese, C. M.; et al. A genome-inspired, reverse selection approach to aptamer discovery. Talanta 2018, 177, 150–156. [Google Scholar] [CrossRef]
- Chinnappan, R.; et al. Anti-VCAM-1 and anti-IL4Rα aptamer-conjugated super paramagnetic iron oxide nanoparticles for enhanced breast cancer diagnosis and therapy. Molecules 2020, 25(15), 3437. [Google Scholar] [CrossRef]
- Wu, P.; et al. Aptamer-guided silver-gold bimetallic nanostructures with highly active surface-enhanced Raman scattering for specific detection and near-infrared photothermal therapy of human breast cancer cells. Analytical Chemistry 2012, 84(18), 7692–7699. [Google Scholar] [CrossRef]
- He, Y.; et al. Iron(II) phthalocyanine loaded and AS1411 aptamer targeting nanoparticles: A nanocomplex for dual modal imaging and photothermal therapy of breast cancer. International Journal of Nanomedicine 2020, 15, 5927–5949. [Google Scholar] [CrossRef]
- Yu, S.; et al. Endogenous mRNA triggered DNA-Au nanomachine for in situ imaging and targeted multimodal synergistic cancer therapy. Angewandte Chemie 2021, 60(11), 5948–5958. [Google Scholar] [CrossRef]
- Luo, J.; et al. Aptamer-based photoelectrochemical assay for the determination of MCF-7. Mikrochimica Acta 2020, 187(5), 257. [Google Scholar] [CrossRef]
- Santos do Carmo, F.; et al. Anti-MUC1 nano-aptamers for triple-negative breast cancer imaging by single-photon emission computed tomography in inducted animals: initial considerations. International Journal of Nanomedicine 2017, 12, 53–60. [Google Scholar] [CrossRef] [PubMed]
- He, S.; et al. Aptamer-PROTAC conjugates (APCs) for tumor-specific targeting in breast cancer. Angewandte Chemie International Edition 2021, 60(43), 23299–23305. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; et al. Discovery of a novel DNA aptamer for impeding tumor metastasis by blocking the functional activity of target protein on exosome. Chemical Engineering Journal 2025, 522, 167253. [Google Scholar] [CrossRef]
- Kim, J. H.; et al. DNA aptamers against Vaccinia-Related Kinase 1 block proliferation in MCF7 breast cancer cells. Biochemical Pharmacology 2020, 175, 113862. [Google Scholar] [CrossRef]
- Pichiorri, F.; et al. In vivo NCL targeting affects breast cancer aggressiveness through miRNA regulation. The Journal of Experimental Medicine 2013, 210(5), 951–968. [Google Scholar] [CrossRef]
- Song, X.; et al. Targeted delivery of doxorubicin to breast cancer cells by aptamer functionalized DOTAP/DOPE liposomes. Oncology Reports 2015, 34(4), 1953–1960. [Google Scholar] [CrossRef] [PubMed]
- Esmaeili, Y.; et al. Smart co-delivery of plasmid DNA and doxorubicin using MCM-chitosan-PEG polymerization functionalized with MUC-1 aptamer against breast cancer. Biomedicine & Pharmacotherapy 2024, 173, 116465. [Google Scholar] [CrossRef]
- Taghipour, Y. D.; et al. Enhanced docetaxel therapeutic effect using dual targeted SRL-2 and TA1 aptamer conjugated micelles in inhibition Balb/c mice breast cancer model. Scientific Reports 2024, 14, 24603. [Google Scholar] [CrossRef] [PubMed]
- Alibolandi, M.; et al. MUC1 aptamer-conjugated mesoporous silica nanoparticles effectively target breast cancer cells. Molecular Pharmaceutics 2015, 12(8), 2576–2587. [Google Scholar] [CrossRef]
- Liu, Z.; et al. PEGylated anti-MUC1 aptamer–doxorubicin complex for targeted drug delivery to MCF7 breast cancer cells. European Journal of Pharmaceutical Sciences 2012, 47(1), 256–263. [Google Scholar] [CrossRef]
- Camorani, S.; et al. Aptamer-targeted therapy potentiates immune checkpoint blockade in triple-negative breast cancer. Molecular Therapy – Nucleic Acids 2020, 22, 231–241. [Google Scholar] [CrossRef]
- Yu, J.; et al. Targeting EGFR/HER2/HER3 with a three-in-one aptamer–siRNA chimera confers superior activity against HER2+ breast cancer. Molecular Therapy – Nucleic Acids 2018, 13, 511–523. [Google Scholar] [CrossRef]
- Li, J.; et al. Co-delivery of paclitaxel and PLK1-targeted siRNA using aptamer-functionalized cationic liposome for synergistic anti-breast cancer effects in vivo. Journal of Nanobiotechnology 2020, 18, 123. [Google Scholar] [CrossRef]
- Mehrnia, M.; et al. AS1411 aptamer-targeted gold nanoclusters enhance the efficacy of radiation therapy in breast tumor-bearing mice. Nanomedicine 2021, 16(18), 1501–1516. [Google Scholar] [CrossRef]
- Wei, X.; et al. PD-L1 aptamer-functionalized degradable hafnium oxide nanoparticles for near infrared-II diagnostic imaging and radiosensitization. Advanced Science 2023, 10(5), 2204568. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; et al. Targeted immunotherapy of triple-negative breast cancer by aptamer-engineered NK cells. Nature Biomedical Engineering 2022, 6(2), 170–183. [Google Scholar] [CrossRef]
- Yang, L.; et al. Photothermal therapeutic response of cancer cells to aptamer-gold nanoparticle-hybridized graphene oxide under NIR illumination. ACS Applied Materials & Interfaces 2015, 7(9), 5097–5106. [Google Scholar] [CrossRef]
- Ibarra, L. E.; et al. Selective photo-assisted eradication of triple-negative breast cancer cells through aptamer decoration of doped conjugated polymer nanoparticles. Pharmaceutics 2022, 14(3), 626. [Google Scholar] [CrossRef]
- Wang, Y.; et al. pH- and miRNA-responsive DNA-tetrahedra/metal–organic framework conjugates: Functional sense-and-treat carriers. ACS Nano 2021, 15(4), 7896–7909. [Google Scholar] [CrossRef]
- Zhang, Y.; et al. ATP/pH dual responsive nanoparticle with d-[des-Arg10] Kallidin mediated efficient in vivo targeting drug delivery. Journal of Nanobiotechnology 2022, 20, 102. [Google Scholar] [CrossRef]
- Nano(bio)sensores electrouímicos y plasmónicos para la cuantificación de biomarcadores de relevancia clínica - CONICET. Available online: https://bicyt.conicet.gov.ar/fichas/produccion/11834746.
- López Santini, B. P. Estudio in silico de la interacción entre el aptámero anti-MUC1 y el epítope de la Mucina 1 (Tesis de Maestría). Centro de Investigación Científica y de Educación Superior de Ensenada (CICESE); 2018; Available online: https://cicese.repositorioinstitucional.mx/jspui/bitstream/1007/2081/1/tesis_L%C3%B3pez_Santini_Brianda_Paola_07_mayo_2018B.pdf.
- Salgado, H.; et al. RegulonDB v12.0: a comprehensive resource of transcriptional regulation in E. coli K-12. Nucleic Acids Research 2023, 52(D1), D255–D264. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




