Submitted:
23 November 2025
Posted:
01 December 2025
You are already at the latest version
Abstract
Deep learning-based Medical Question Answering (MQA) systems are transforming access to healthcare information by enabling accurate and timely responses to complex queries. However, existing systems face challenges such as limited large-scale, high-quality medical datasets, inadequate contextual understanding, and difficulties in managing diverse medical terminologies This research proposes a novel closed-domain MQA system that addresses these limitations through innovative methodologies. The system employs BioBERT-based domain-specific embeddings trained on biomedical literature to accurately capture medical terminology, abbreviations, and contextual nuances. To model sequential dependencies in queries, Recurrent Neural Networks (RNNs) are integrated, enabling contextual interpretation across longer text sequences. Additionally, a question expansion mechanism utilizing medical dictionaries and ontologies like UMLS addresses synonymy, ambiguity, and terminological variations, ensuring that diverse medical expressions map to consistent, semantically relevant concepts for precise answer retrieval. Extensive evaluation using metrics such as F1-score, precision, recall, and exact match demonstrates the system’s superior performance compared to existing models. The key contributions include improved contextual understanding, better handling of medical terminology, and a scalable framework for future medical NLP applications. This system not only offers a reliable tool for healthcare professionals and patients but also advances the field of intelligent question answering by supporting evidence-based clinical decision-making.
Keywords:
I. Introduction
A. Background
B. Research Motivation and Objectives
C. Research Contributions
II. Related Work
III. Methodology
A. Dataset Collection
- COVID-QA contains question-context-answer triples focused on COVID-19 topics like prevention and treatment. It tests the system’s ability to handle specialized medical vocabulary.
- MedQuAD includes a wide range of medically curated questions and answers from trusted sources like the U.S. National Library of Medicine, covering general medical topics.
| Dataset | Domain | Size | Purpose |
|---|---|---|---|
| COVID-QA | COVID-19 | 2,019 questions | Specialized COVID-19 queries |
| MedQuAD | General Medicine | 16,408 questions | Broad medical topics |
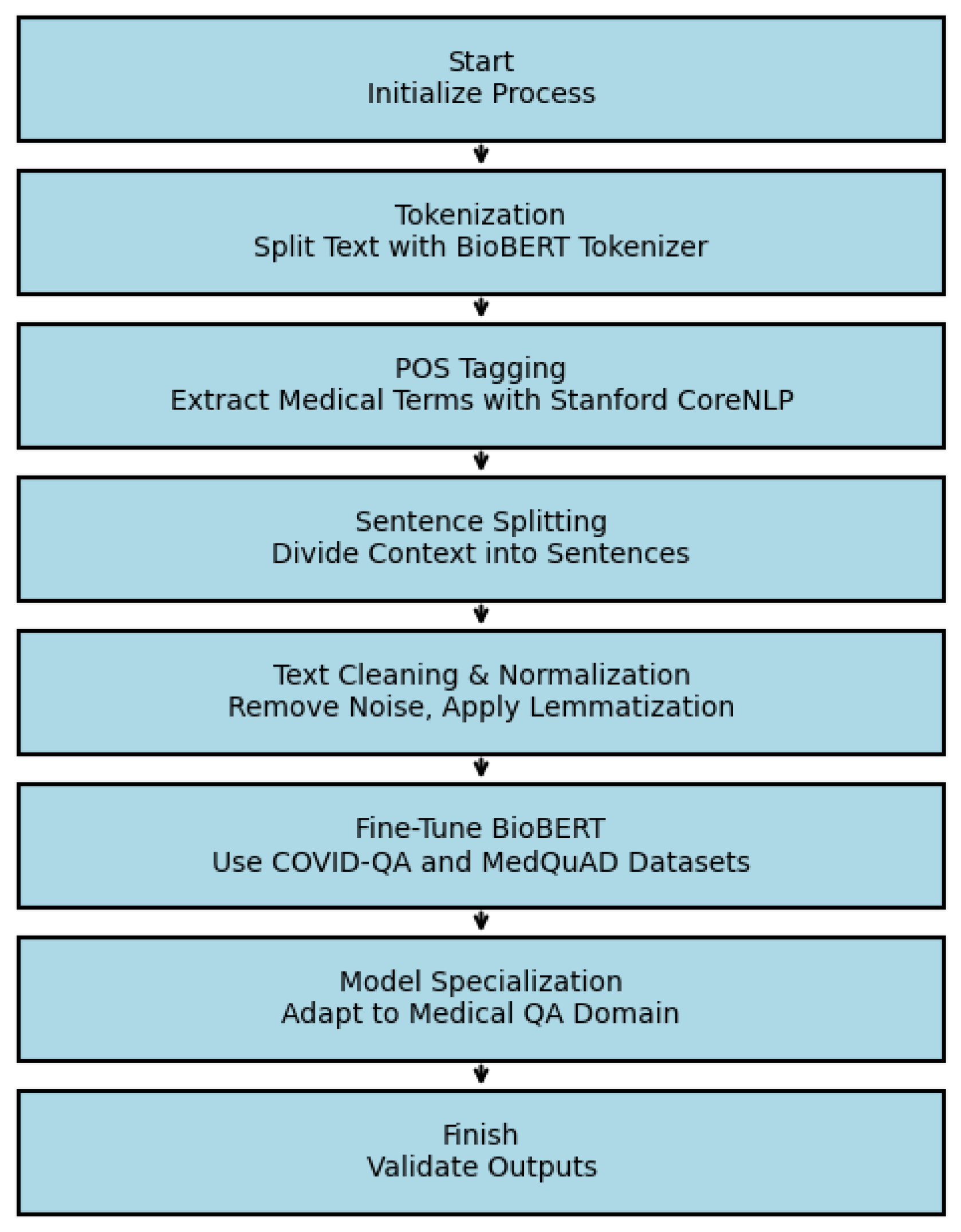
B. Data Preprocessing and Fine-Tuning
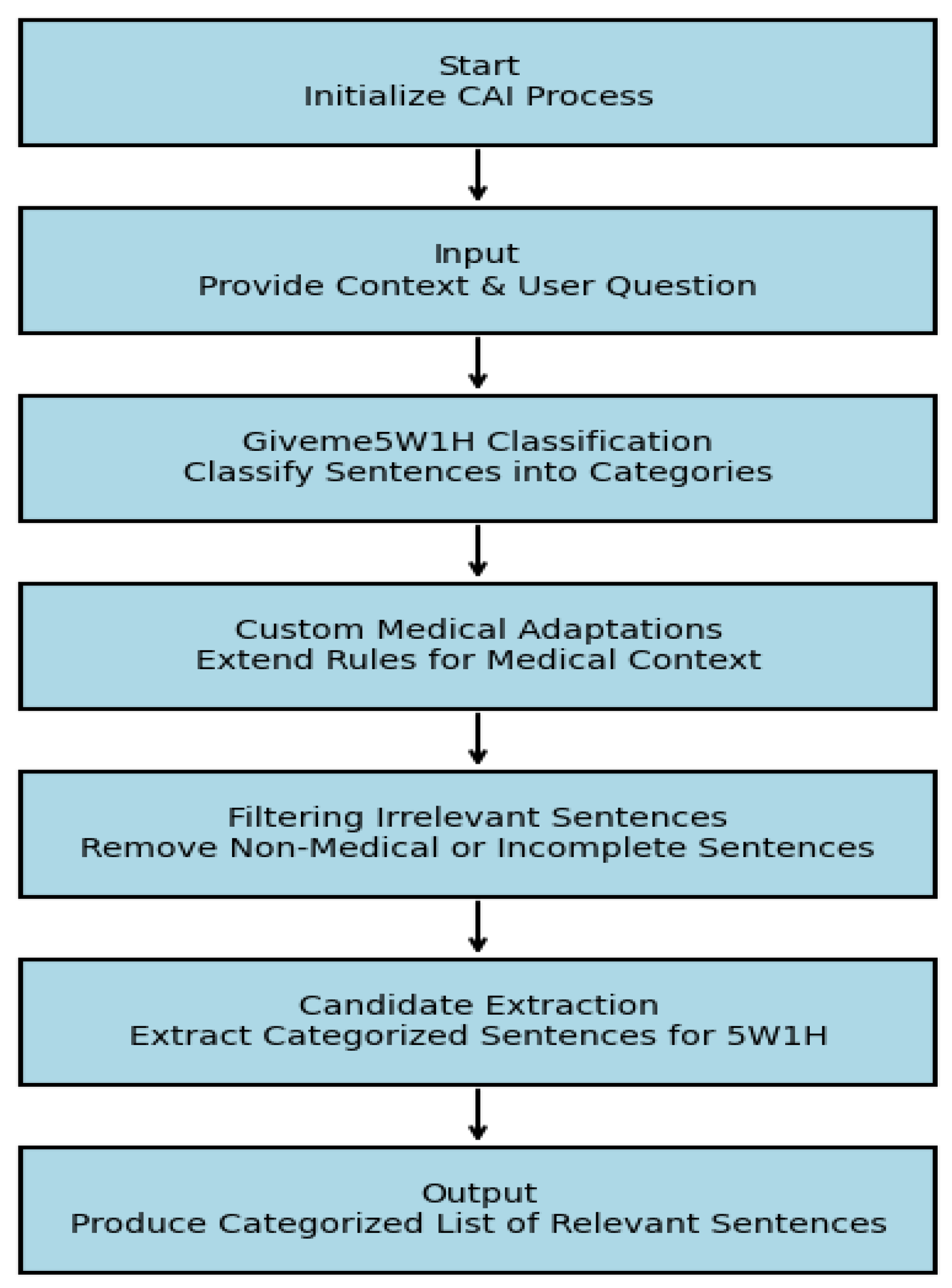
C. Candidate Answer Identification (CAI) Module
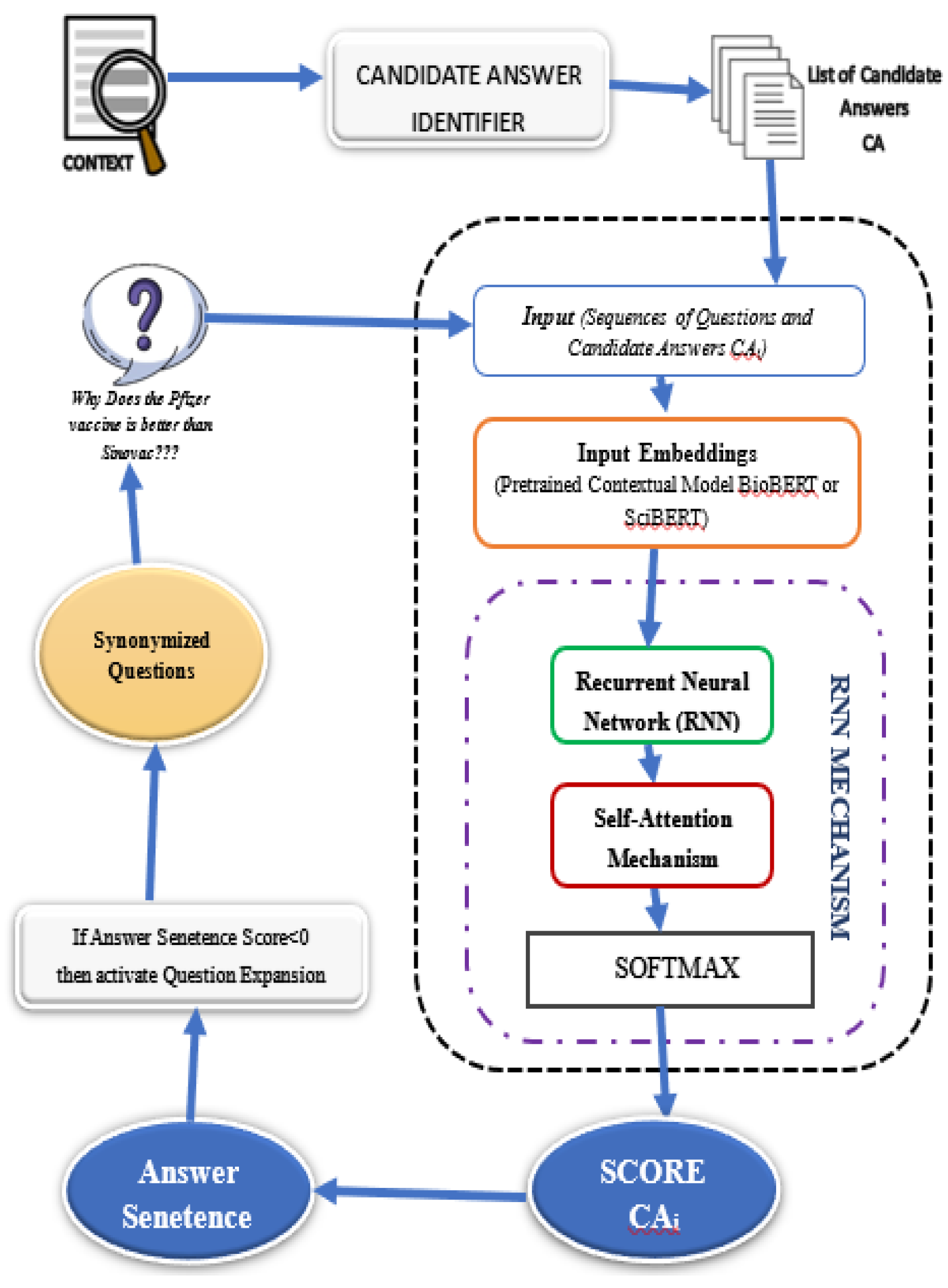
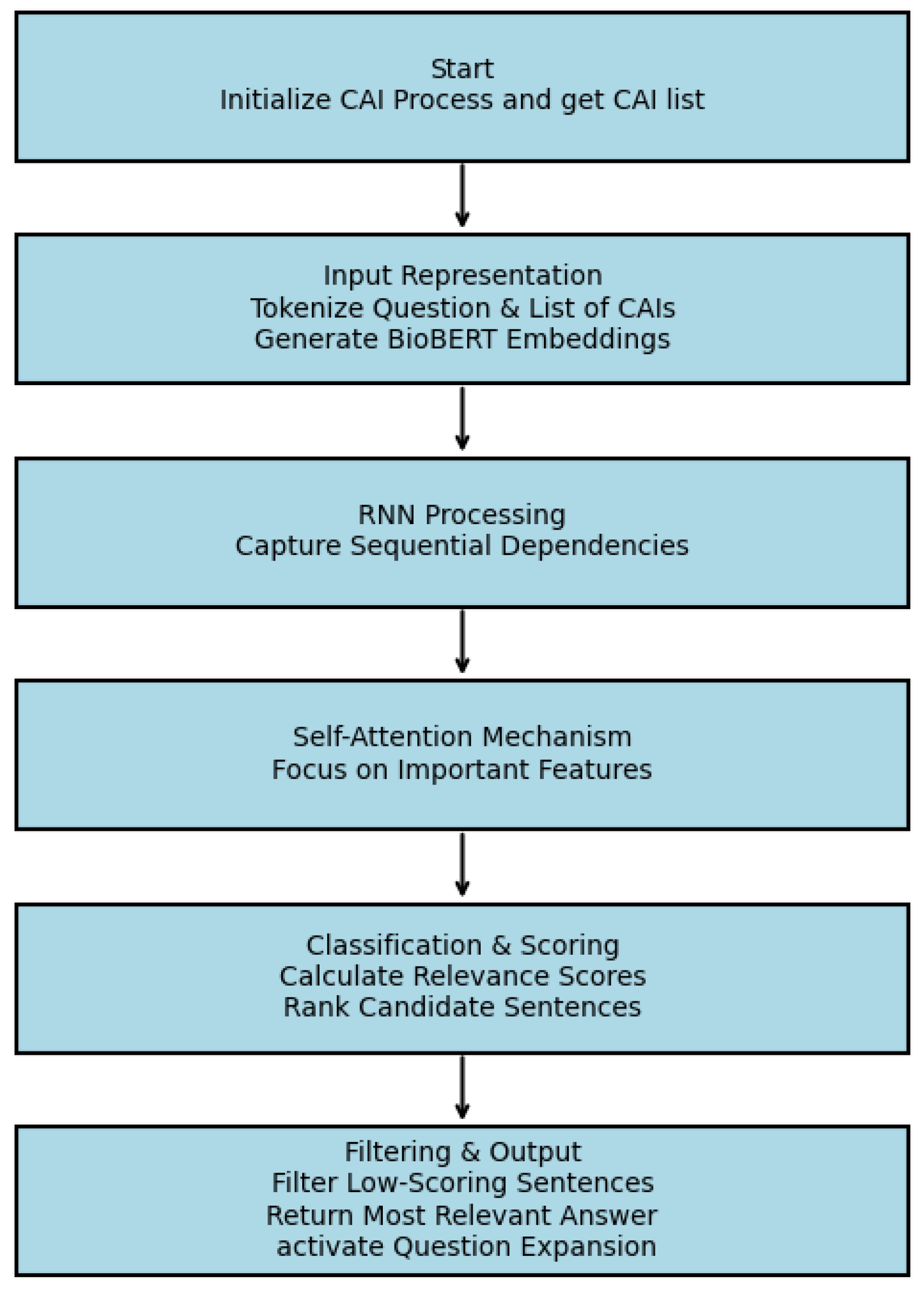
D. RNN-Attention based Answer Selector
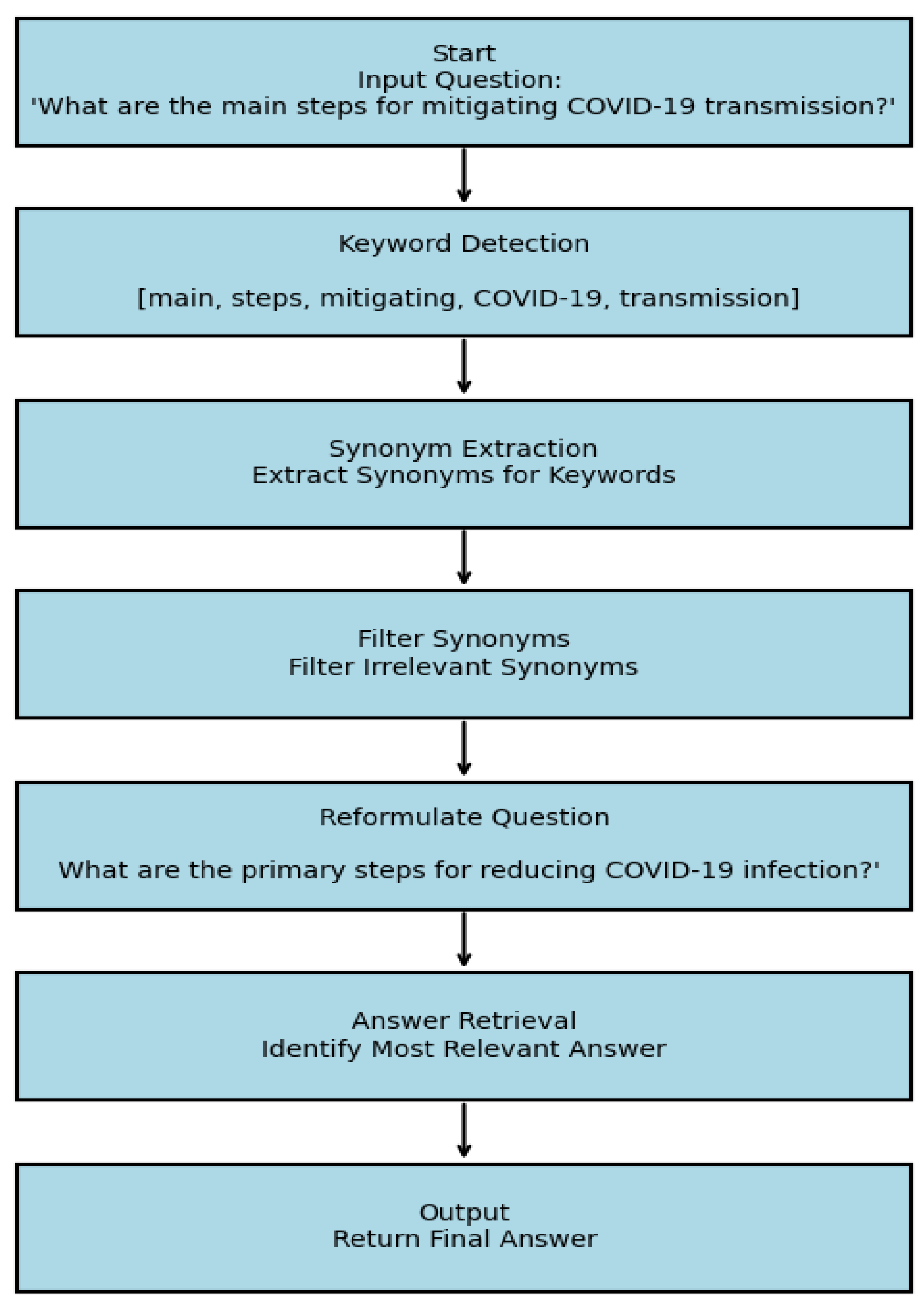
E. Question Expansion (QE) Module
IV. Experiment Setup
V. Results & Discussion
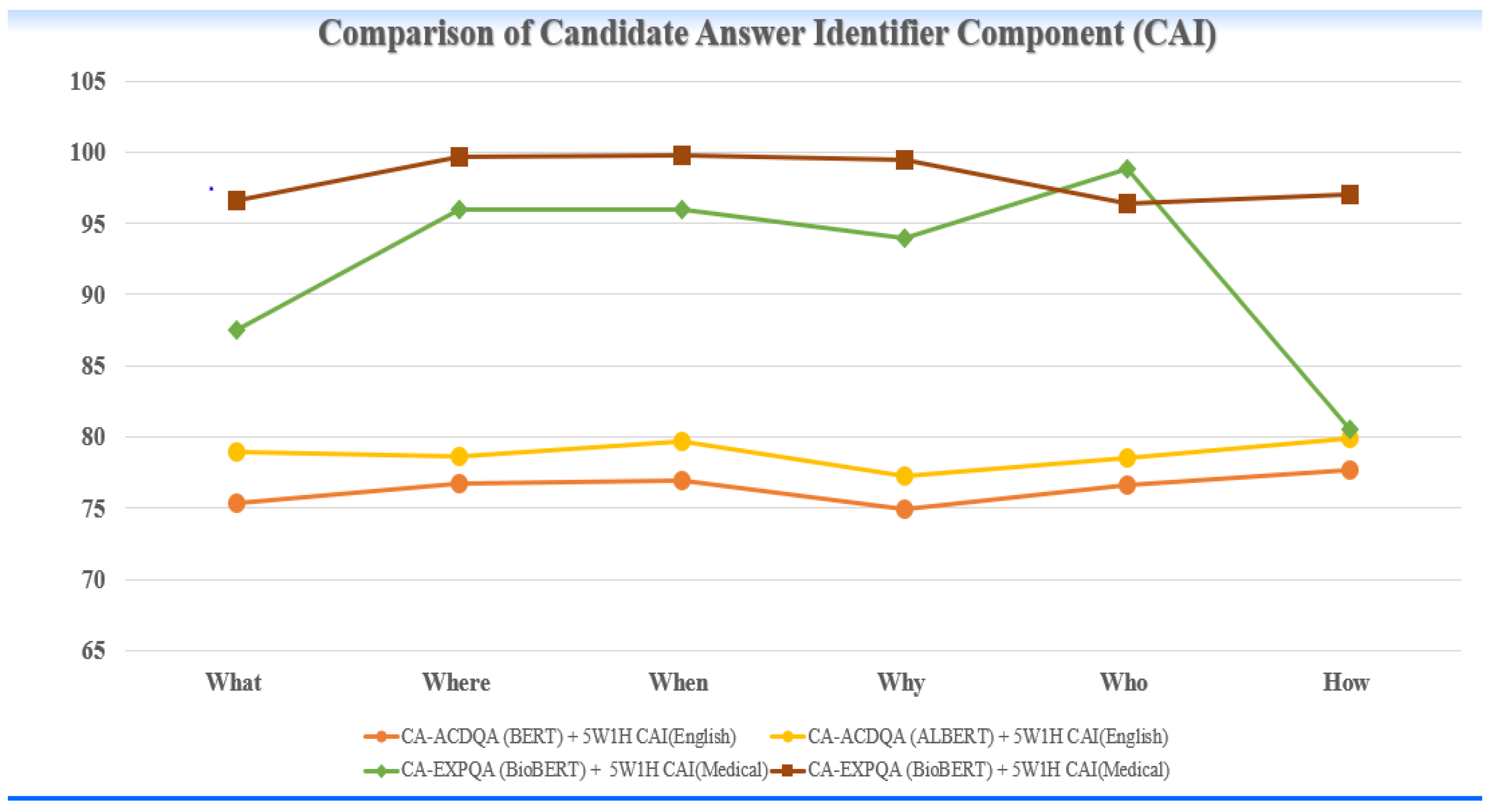
A. Results of Candidate Answer Identifier Module:
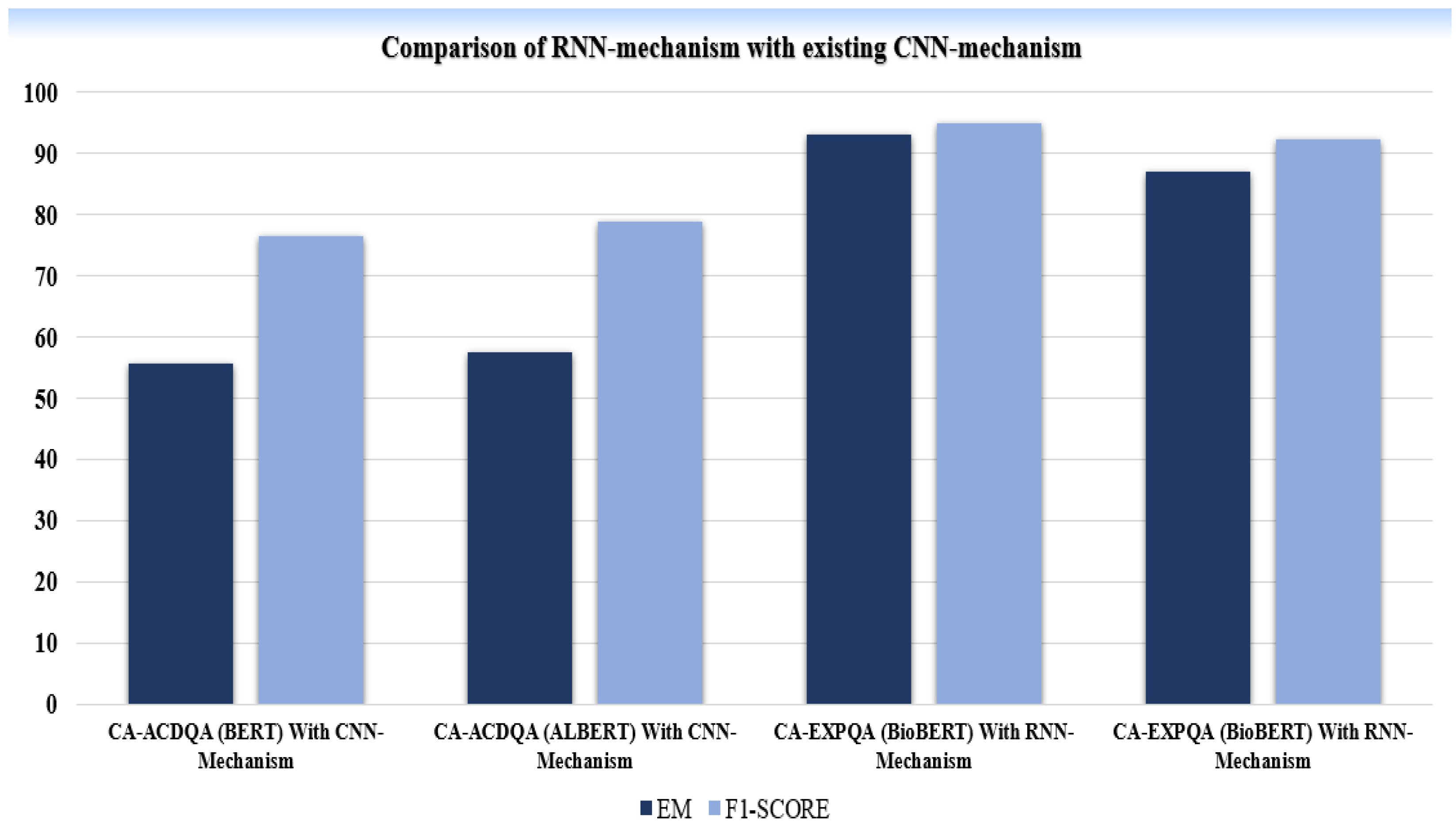
B. Results of RNN-Attention Module:
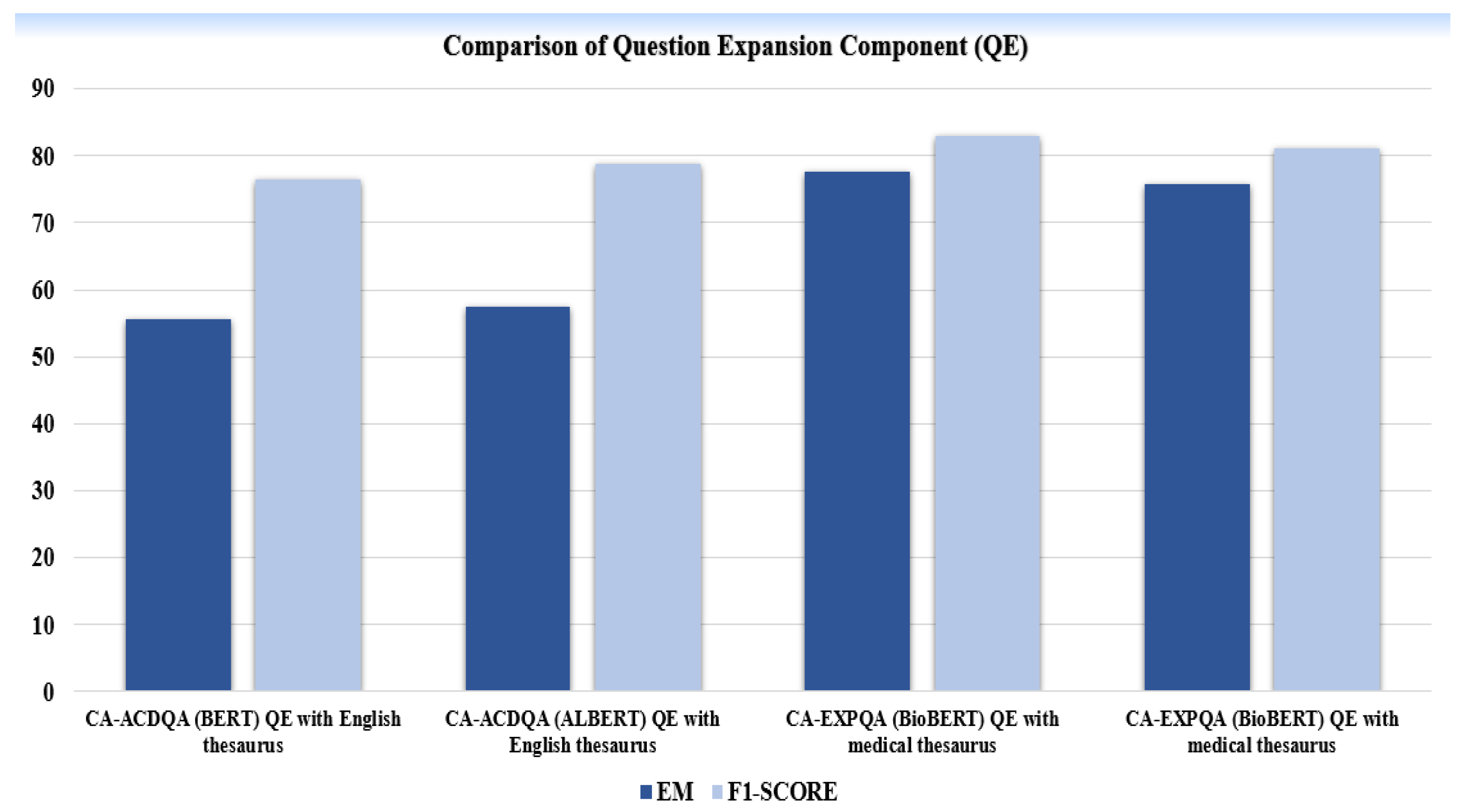
C. Results of Question Expansion Module:
VI. Conclusion & Future Work
References
- Saeed, N., Ashraf, H., & Jhanjhi, N. Z. (2023). Deep Learning-Based Question Answering System (Survey). Preprints, 2023121739.
- Mutabazi, E., Ni, J., Tang, G., & Cao, W. (2021). A Review on Medical Textual Question Answering Systems Based on Deep Learning Approaches. Applied Sciences, 11(12), 25456. [CrossRef]
- Kia, M. A., Garifullina, A., Kern, M., Chamberlain, J., & Jameel, S. (2022). Adaptable Closed-Domain Question Answering Using Contextualized CNN-Attention Models and Question Expansion. IEEE Access. [CrossRef]
- Jili Qian et al., “A Liver Cancer Question-Answering System Based on Next-Generation Intelligence and the Large Model Med-PaLM 2,” International Journal of Computer Science and Information Technology, 2024. [CrossRef]
- Jafar A. Alzubi et al., “COBERT: COVID-19 Question Answering System Using BERT,” Arabian Journal for Science and Engineering, 2023. [CrossRef]
- Husamelddin A.M.N Balla et al., “Arabic Medical Community Question Answering Using ON-LSTM and CNN,” ICMLC, 2022.
- Dimitris Pappas et al., “Data Augmentation for Biomedical Factoid Question Answering,” 2022.
- Jimenez Eladio and Hao Wu, “emrQA-msquad: A Medical Dataset Structured with the SQuAD V2.0 Framework,” Beijing Institute of Technology, 2024.
- Juraj Vladika and Florian Matthes, “Improving Health Question Answering with Reliable and Time-Aware Evidence Retrieval,” Technical University of Munich, 2024.
- Rui Yang et al., “KG-Rank: Enhancing Large Language Models for Medical QA with Knowledge Graphs and Ranking Techniques,” 2024.
- Chengyu Huang and Feng Yao, “Research on Question-Answering System of Children’s Disease Based on ALBERT,” ISAIMS, 2023.
- Yu-Hsuan Chang et al., “Interactive Healthcare Robot Using Attention-Based Question-Answer Retrieval and Medical Entity Extraction Models,” IEEE Journal of Biomedical and Health Informatics, 2023. [CrossRef]
- Shrutikirti Singh and Seba Susan, “Healthcare Question-Answering System: Trends and Perspectives,” 2023.
- Prateek Chhikara et al., “Privacy-Aware Question-Answering System for Online Mental Health Risk Assessment,” University of Southern California, 2023.
- Yun Zhao et al., “JMS-QA: A Joint Hierarchical Architecture for Mental Health Question Answering,” IEEE/ACM Transactions on Audio, Speech, and Language Processing, 2024. [CrossRef]
- Baivab Das and S. Jaya Nirmala, “Improving Healthcare Question Answering System by Identifying Suitable Answers,” 2022 IEEE MysuruCon.
- Gezheng Xu et al., “External Features-Enriched Model for Biomedical Question Answering,” BMC Bioinformatics, 2021. [CrossRef]
- Gregory Kell et al., “Question Answering Systems for Health Professionals at the Point of Care—A Systematic Review,” Journal of the American Medical Informatics Association, 2024. [CrossRef]
- Prateek Chhikara et al., “Privacy Aware Question-Answering System for Online Mental Health Risk Assessment,” 2023.
- Singh, S., & Susan, S. (2023). Healthcare Question–Answering System: Trends and Perspectives. Springer.
- Xu, G., Rong, W., Wang, Y., Ouyang, Y., & Xiong, Z. (2021). External Features Enriched Model for Biomedical Question Answering. BMC Bioinformatics. [CrossRef]
- Yagnik, N., Jhaveri, J., Sharma, V., & Pila, G. (2024). MedLM: Exploring Language Models for Medical Question Answering Systems. arXiv.
- Das, B., & Nirmala, S. J. (2022). Improving Healthcare Question Answering System by Identifying Suitable Answers. IEEE MysuruCon.
- Pergola, G., Kochkina, E., Gui, L., Liakata, M., & He, Y. (2021). Boosting Low-Resource Biomedical QA via Entity-Aware Masking Strategies. arXiv.
- Peng, K., Yin, C., Rong, W., Lin, C., Zhou, D., & Xiong, Z. (2022). Named Entity Aware Transfer Learning for Biomedical Factoid Question Answering. IEEE/ACM Transactions on Computational Biology and Bioinformatics. [CrossRef]
- Pappas, D., Malakasiotis, P., & Androutsopoulos, I. (2022). Data Augmentation for Biomedical Factoid Question Answering.
| MODEL | DATASET | EVALUATION METRICES | |
|---|---|---|---|
| EM | F1-SCORE | ||
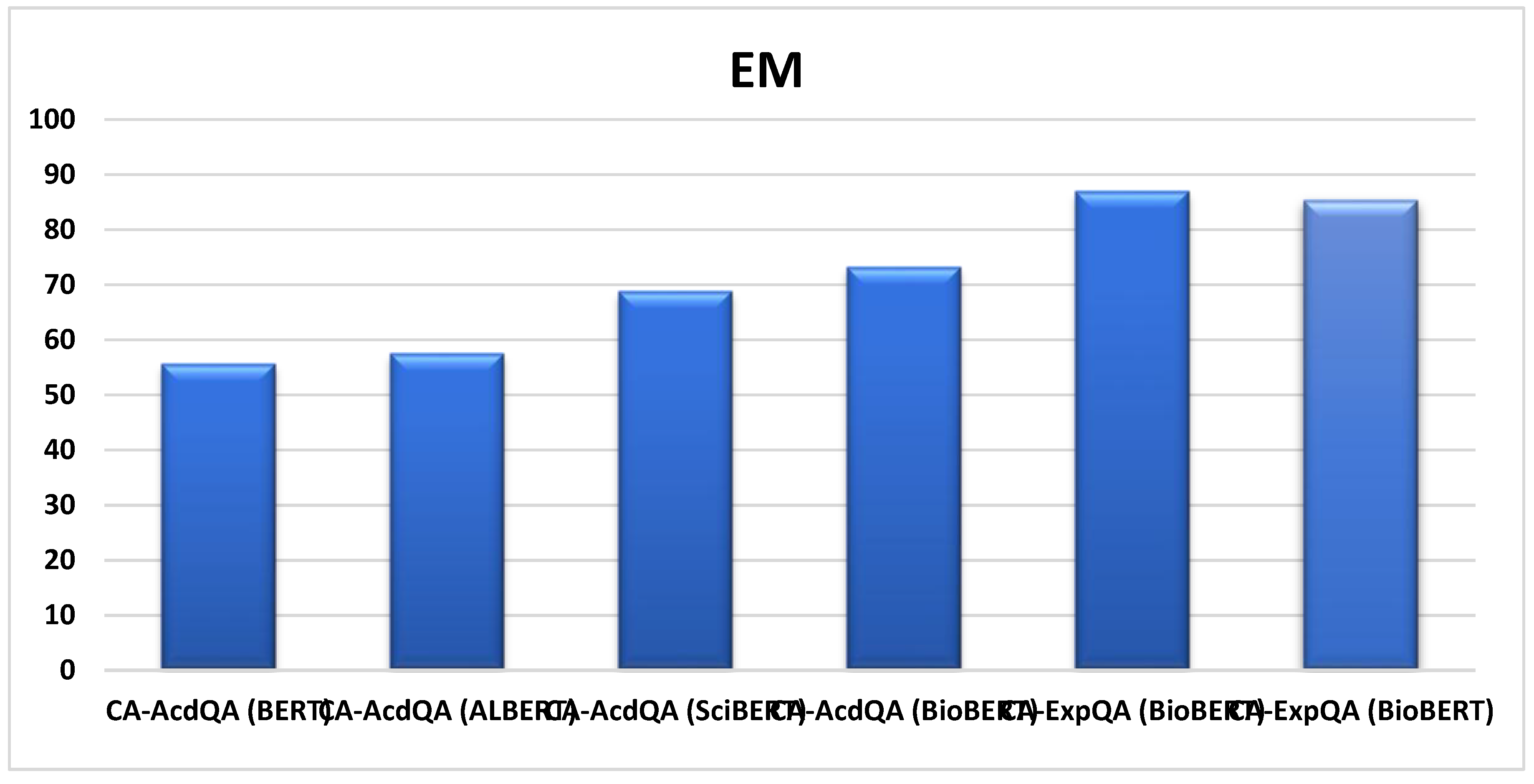
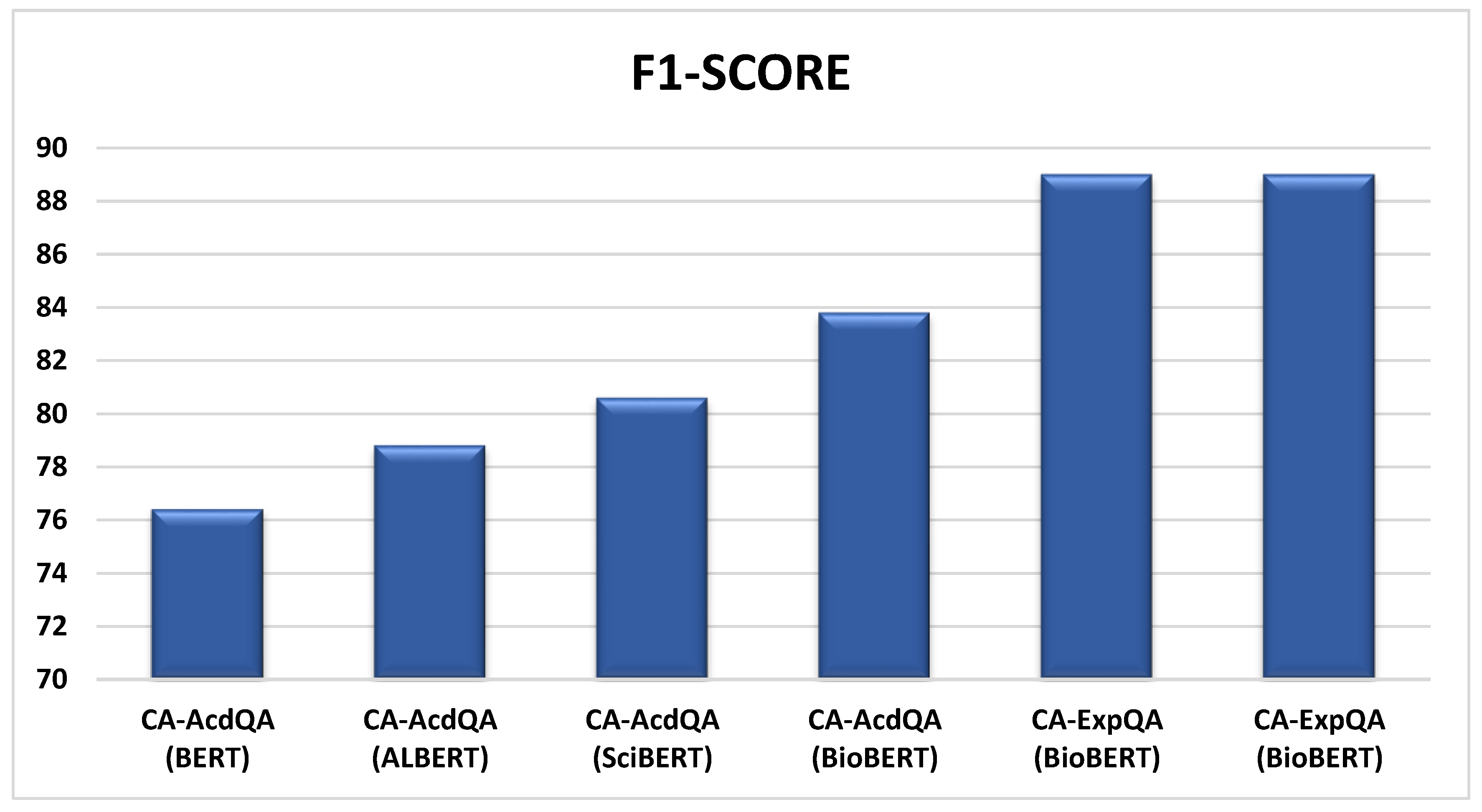
| CA-AcdQA* (BERT) | COVID-QA | 55.6 | 76.4 |
| CA-AcdQA* (ALBERT) | COVID-QA | 57.5 | 78.8 |
| CA-AcdQA* (SciBERT) | COVID-QA | 68.8 | 80.6 |
| CA-AcdQA (BioBERT) | COVID-QA | 73.2 | 83.8 |
| CA-ExpQA (BioBERT) | COVID-QA | 87.0 | 89.0 |
| CA-ExpQA (BioBERT) | MEDQUAD | 85.3 | 89.0 |
| MODEL | DATASET | F1 score for each question category | |||||
|---|---|---|---|---|---|---|---|
| What | Where | When | Why | Who | How | ||
|
CA-AcdQA (BERT) Cust. Giveme 5W1H CAI(English) |
COVID-QA | 75.3 | 76.7 | 76.9 | 74.9 | 76.6 | 77.7 |
|
CA-AcdQA (ALBERT) Cust. Giveme 5W1H CAI(English) |
COVID-QA | 78.9 | 78.6 | 79.7 | 77.2 | 78.5 | 79.9 |
|
CA-ExpQA (BioBERT) Cust. Giveme 5W1H CAI(Medical) |
COVID-QA | 87.5 | 96.0 | 96.0 | 94.0 | 98.8 | 80.5 |
|
CA-ExpQA (BioBERT) Cust. Giveme 5W1H CAI(Medical) |
MEDQUAD | 96.6 | 99.7 | 99.8 | 99.5 | 96.4 | 97.0 |
| *MODEL | DATASET | EVALUATION METRICES | |
|---|---|---|---|
| *EM | F1-SCORE | ||
|
CA-ACDQA (BERT) With CNN-Mechanism |
COVID-QA | 55.6 | 76.4 |
|
CA-ACDQA (ALBERT) With CNN-Mechanism |
COVID-QA | 57.5 | 78.8 |
|
CA-EXPQA (BioBERT) With RNN-Mechanism |
COVID-QA | 93.1 | 95.0 |
|
CA-EXPQA (BioBERT) With RNN-Mechanism |
MEDQUAD | 87.1 | 92.3 |
| MODEL | DATASET | EVALUATION METRICES | |
|---|---|---|---|
| EM | F1-SCORE | ||
|
CA-ACDQA (BERT) QE with English thesaurus |
COVID-QA | 55.6 | 76.4 |
|
CA-ACDQA (ALBERT) QE with English thesaurus |
COVID-QA | 57.5 | 78.8 |
|
CA-EXPQA (BioBERT) QE with medical thesaurus |
COVID-QA | 77.6 | 83.0 |
|
CA-EXPQA (BioBERT) QE with medical thesaurus |
MEDQUAD | 75.7 | 81.2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




