Submitted:
28 November 2025
Posted:
01 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
- A novel multitask network architecture that simultaneously performs staining normalization and nuclei segmentation, with the two tasks designed to mutually enhance performance, improving overall segmentation accuracy.
- A semi-supervised teacher-student paradigm that mitigates performance degradation in nuclei segmentation due to scarce pixel-level annotations, leveraging unlabeled data to boost accuracy.
- The integration of the multitask architecture and semi-supervised paradigm into 3SGAN, achieving superior staining normalization and nuclei segmentation performance with minimal annotated data on a large dataset of 1,408 WSIs from two medical institutions, encompassing 101 staining styles and only 5% nucleus-level annotations.
2. Related Work
2.1. Stain Normalization of Histopathological Images
2.2. Nuclei Segmentation
2.3. Multitask Strategy
2.4. Semisupervised Learning
3. Materials and Methods
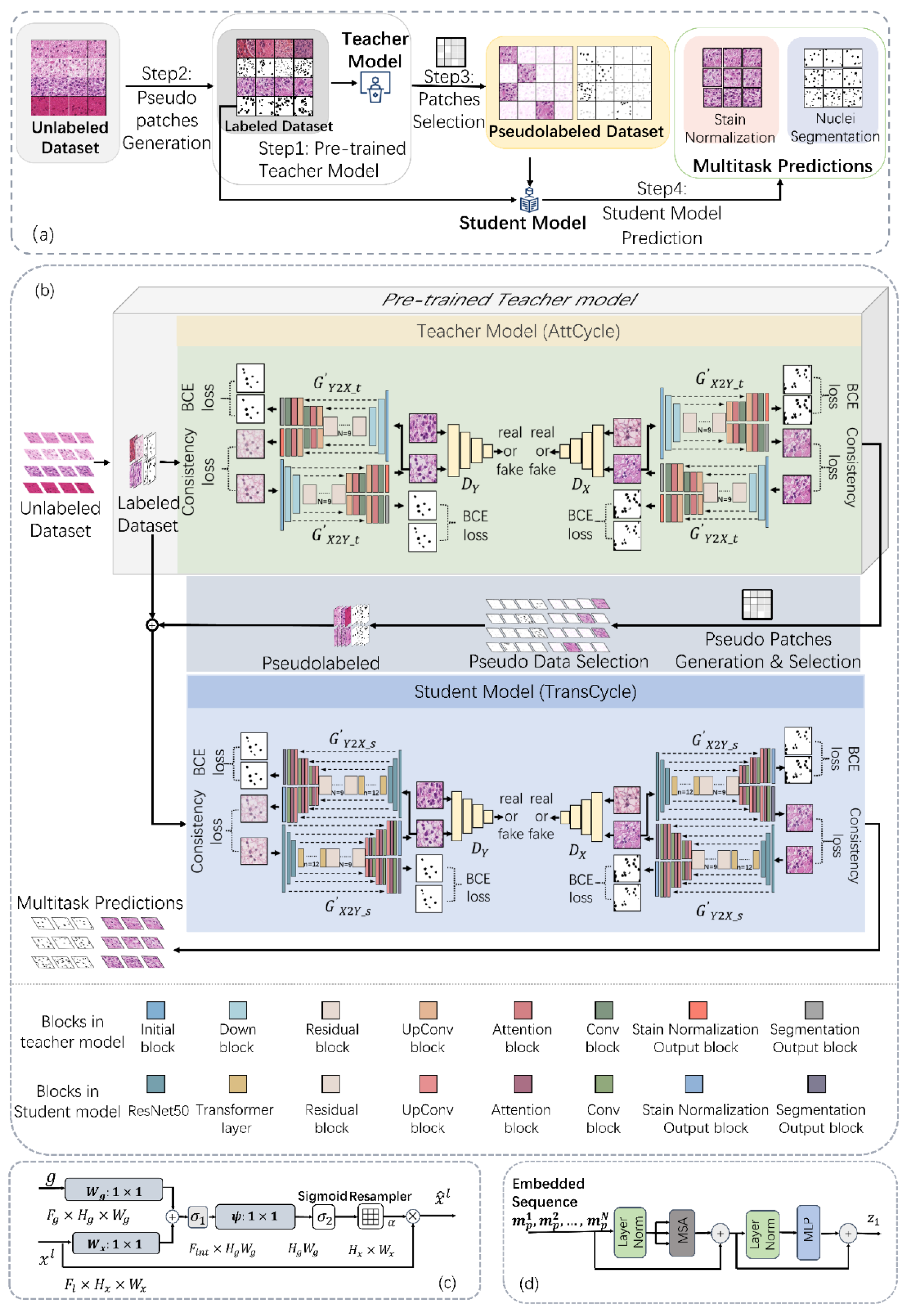
3.1. Methodological Overview
- The teacher model (AttCycle) is pretrained using a limited set of labeled samples.
- The pretrained teacher model generates nuclei segmentation and staining normalization results from large volumes of unlabeled data.
- A pseudolabeled dataset selection algorithm (described in Section 3.4) filters the teacher's outputs to create a high-quality pseudolabeled dataset, encompassing pseudolabeled nuclei and staining normalization data.
- The student model (TransCycle) is trained on both the pseudolabeled dataset and the original labeled dataset used for the teacher model.
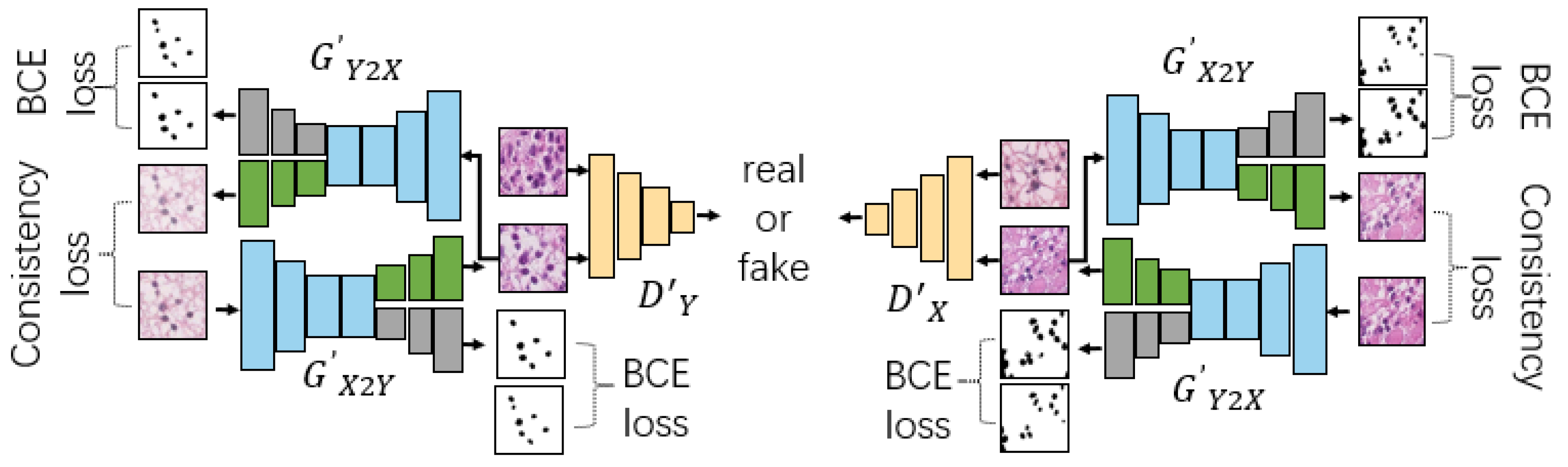
3.2. Stain Normalization and Segmentation Multitask Network
- Shared encoders (illustrated in blue)
- Stain normalization branches (depicted in green)
- Segmentation branches (represented in gray)
- PatchGAN-based discriminators (shown in yellow)
- Segmentation Terms: We utilized a binary cross entropy loss for the segmentation task in our approach. The use of this segmentation loss allowed for the effective weight updates of the segmentation task decoder within the multitask network. Through this approach, we were able to improve the stain normalization results by focusing specifically on the cell nucleus region with the aid of the segmentation loss. This resulted in more accurate segmentation of cell nuclei.
- Cycle-Consistency Terms: The cycle-consistency loss is used to enforce the consistency of histopathological structures between X and :
- Total objective: This total loss function establishes a connection between the segmentation task and the stain normalization task by facilitating the hard parameter sharing of the common encoder during network training. This approach yields enhanced results for both tasks, leading to improved cell nucleus segmentation and stain normalization. The complete objective function of this total loss to minimize is summarized as following.
3.3. Semisupervised Teacher-Student Pipeline
- Advantages of CNNs in the Teacher Model: CNNs require fewer annotations for training due to their relatively small parameter count compared to Transformers. This is particularly beneficial in medical image analysis, where annotating pathological cell nuclei is labor-intensive and prone to errors. Complex teacher models often struggle to produce stable, high-quality pseudolabels under such conditions. In contrast, our CNN-based teacher model, AttCycle, excels at generating reliable pseudolabeled data even with minimal annotated samples.
- Benefits of a High-Capacity Student Model: Large models generally outperform smaller ones when trained on abundant data. Thus, we employ a student model, TransCycle, which integrates CNN and Transformer architectures. This hybrid design harnesses CNNs' prowess in preserving fine-grained details and Transformers' strength in modeling global contextual information, aiming to elevate overall performance.
- Training the Teacher Model (AttCycle):
- We begin by training AttCycle, a low-capacity CNN-based teacher model, on a small local dataset. AttCycle enhances its focus on cell nucleus regions by incorporating attention gates from Attention U-Net into a multitask network, improving both stain normalization and cell nucleus segmentation.
- Owing to its CNN architecture, AttCycle reliably generates high-quality pseudolabels with limited annotated data, outperforming Transformer-based alternatives in this constrained setting.
- To ensure pseudolabel quality, we apply a sigmoid probability method: only pseudolabels where each confidential pixel's sigmoid probability exceeds a predefined threshold, and the proportion of such pixels surpasses a preset confidence ratio, are retained.
- 2.
- Training the Student Model (TransCycle):
- The augmented dataset, enriched with high-quality pseudolabels from AttCycle, is then used to train TransCycle, our high-capacity student model. TransCycle employs a hybrid CNN-Transformer encoder within the dual-decoder generators of a multitask network.
- With a greater capacity than AttCycle, TransCycle excels at recovering localized spatial details and enhancing finer features when trained on large datasets, leveraging the complementary strengths of CNNs and Transformers.
3.4. Architecture and Design Details
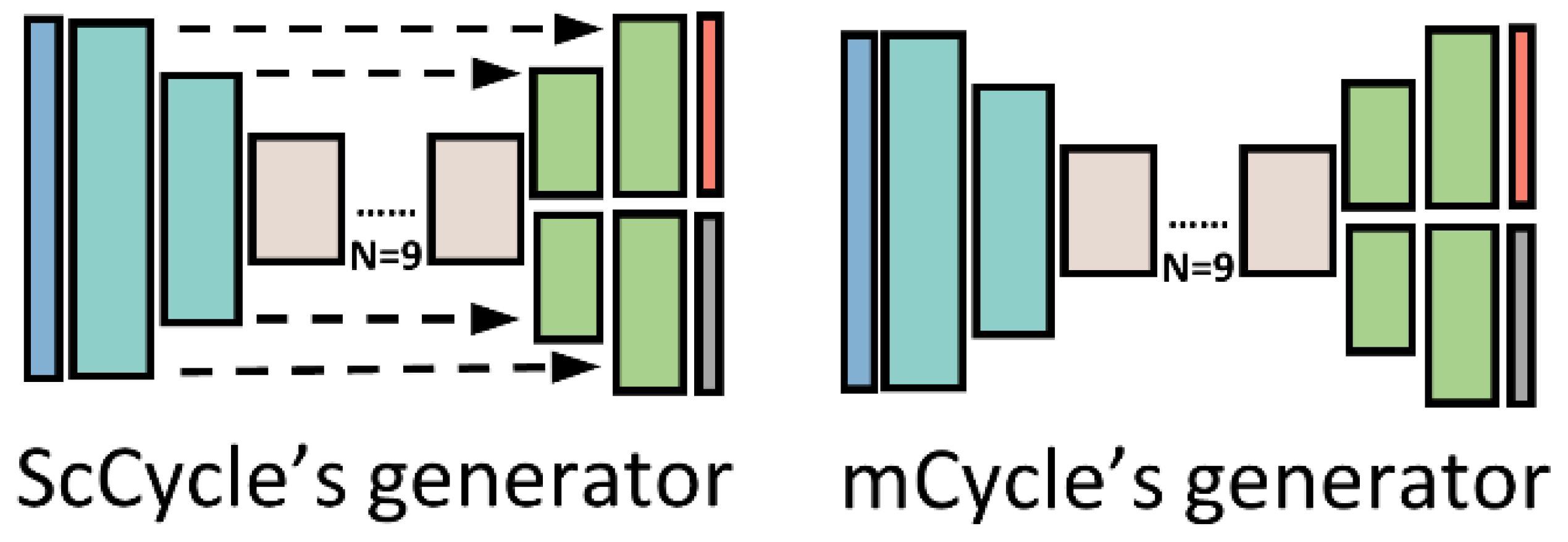
3.4.1. Teacher Model Generators
3.4.2. Student Model Generators
3.4.3. Pseudolabeled Dataset Selection
| Algorithm 1. Teacher-student pseudolabel selection with confidence thresholding. |
| Algorithm 1 Pseudolabeled data selection Input: Labeled data , pixel-wise label = , teacher model segmentation output =, unlabeled data}, confidence threshold θ, confidence pixel ratio μ. 1: 2: For i = 1 to n do 3: , = teacher() 4: = sigmoid() 5: If ≥ θ then 6: number += 1 7: EndIf 8: = 9: EndFor 10: Training dataset of student model: S = X ∪ U( ≥ ) ∪ N( ≥ ), = ∪ P( ≥ |
4. Results
4.1. Experimental Dataset Composition
4.2. Evaluation Metrics and Comparison Methods
4.3. Implementation Details
4.4. Experimental Design and Results
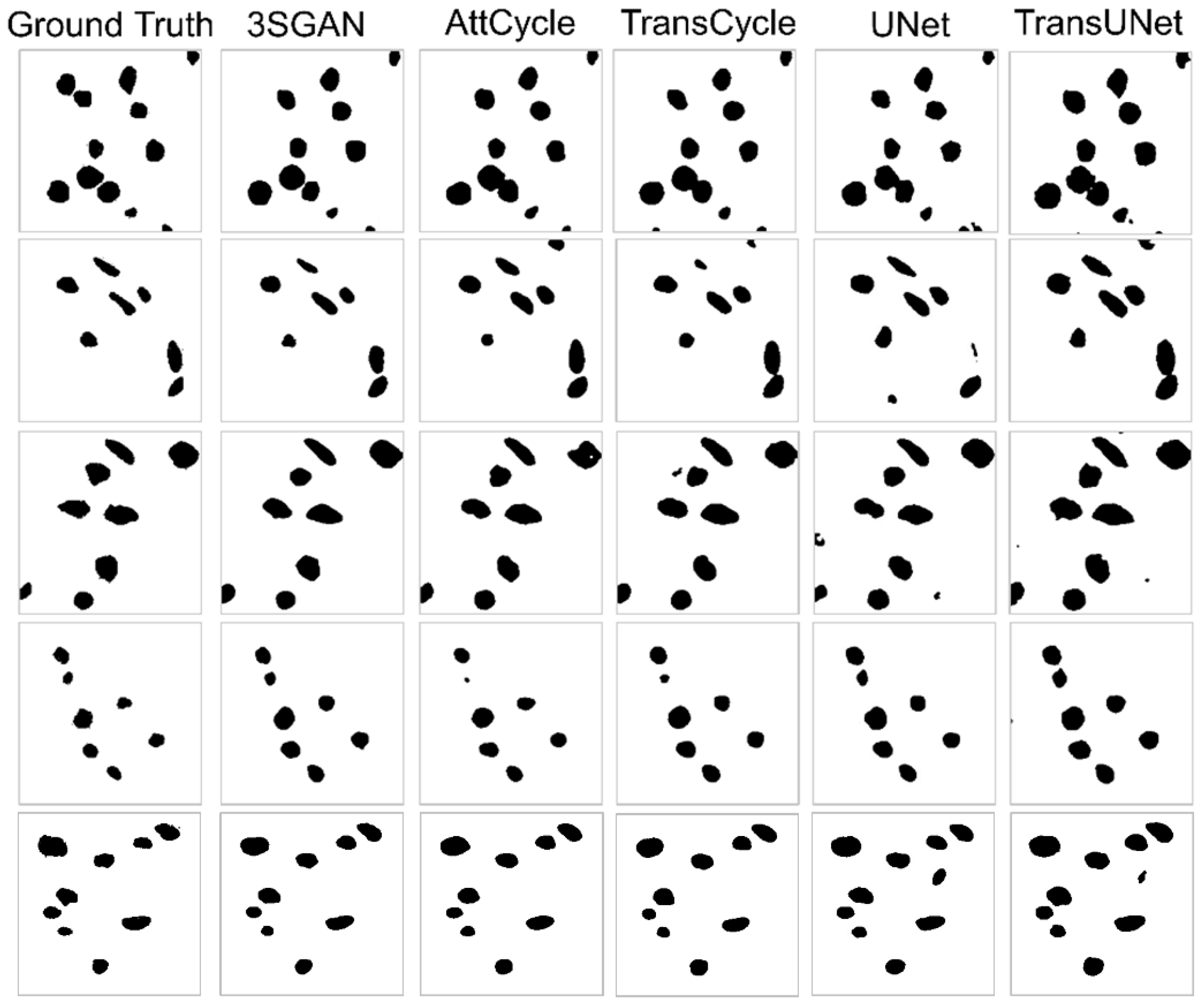
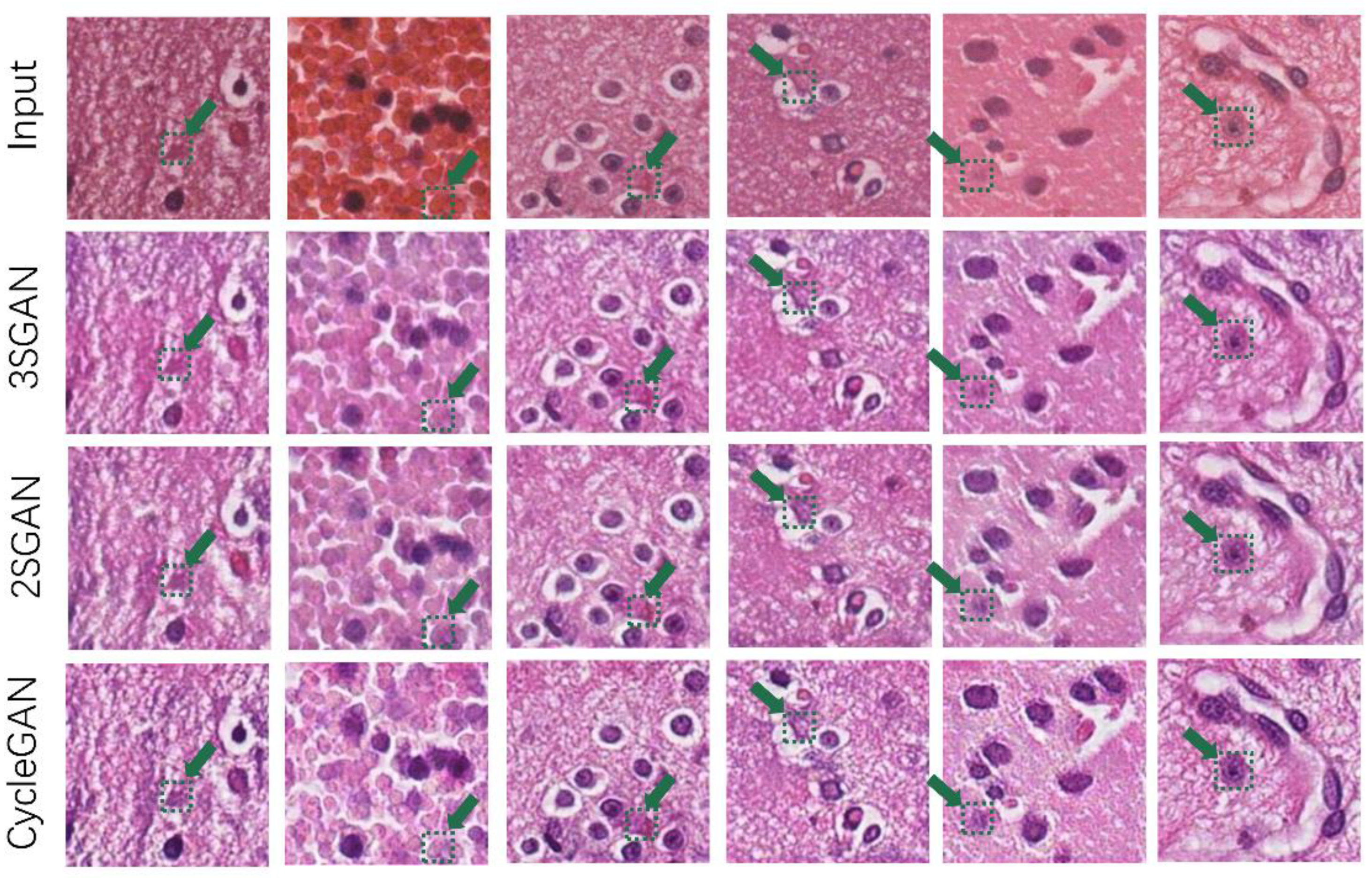
4.4.1. Superiority of the Proposed Multitask Network
4.4.2. Further Improvement Through the Teacher-Student Paradigm
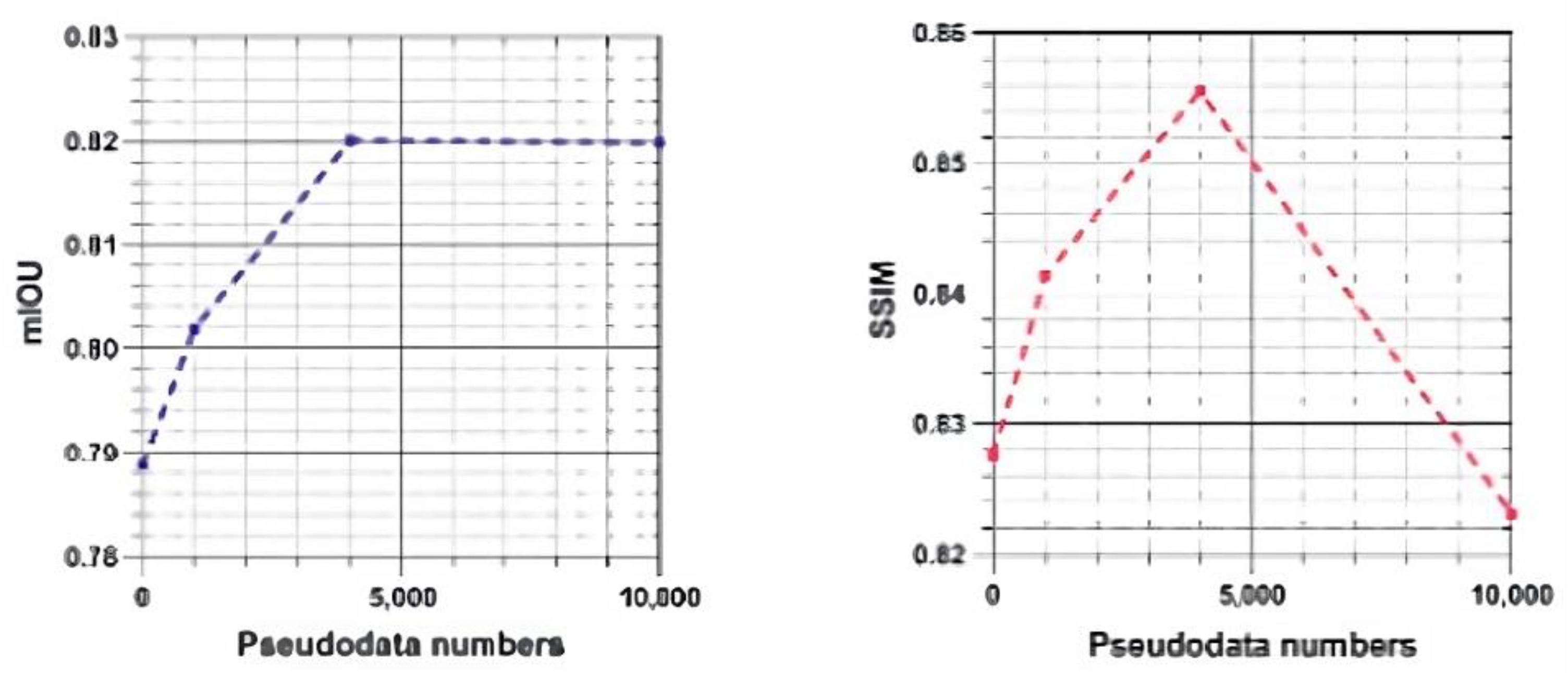
4.4.3. Effect of Different Quantities of Pseudolabeled Data
4.5. External Validation
4.5.1. External Validation on MoNuSeg and PanNuke
4.5.2. Generalization to Non-ROI Regions on In-House WSIs
5. Discussion
5.1. Teacher and Student Model Selection
5.2. Pseudolabeled Dataset Size Saturation Point.
5.3. Self-Training Ablation Experiments on Teacher and Student Models
5.4. Relevance of Stain Normalization in the Era of Foundation Models
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Madabhushi, A.; Lee, G. Image analysis and machine learning in digital pathology: Challenges and opportunities. Med. Image Anal. 2016, 33, 170–175. [Google Scholar] [CrossRef]
- Mahapatra, D.; Bozorgtabar, B.; Thiran, J.P.; Shao, L. Structure preserving stain normalization of histopathology images using self supervised semantic guidance. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2020; Martel, A.L., Abolmaesumi, P., Stoyanov, D., Mateus, D., Zuluaga, M.A., Zhou, S.K., Racoceanu, D., Joskowicz, L., Eds.; Springer: Cham, Switzerland, 2020; pp. 309–319. [Google Scholar] [CrossRef]
- Srinidhi, C.L.; Ciga, O.; Martel, A.L. Deep neural network models for computational histopathology: A survey. Med. Image Anal. 2021, 67, 101813. [Google Scholar] [CrossRef] [PubMed]
- Lyon, H.O.; De Leenheer, A.; Horobin, R.; Lambert, W.; Schulte, E.; Van Liedekerke, B.; Wittekind, D. Standardization of reagents and methods used in cytological and histological practice with emphasis on dyes, stains and chromogenic reagents. Histochem. J. 1994, 26, 533–544. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, I.; Xia, Y.; Cui, H.; Islam, Z.U. Dan-nucnet: A dual attention based framework for nuclei segmentation in cancer histology images under wild clinical conditions. Expert Syst. Appl. 2023, 213, 118945. [Google Scholar] [CrossRef]
- Majumdar, S.; Pramanik, P.; Sarkar, R. Gamma function based ensemble of cnn models for breast cancer detection in histopathology images. Expert Syst. Appl. 2023, 213, 119022. [Google Scholar] [CrossRef]
- Sethanan, K.; Pitakaso, R.; Srichok, T.; Khonjun, S.; Thannipat, P.; Wanram, S.; Boonmee, C.; Gonwirat, S.; Enkvetchakul, P.; Kaewta, C.; et al. Double amis-ensemble deep learning for skin cancer classification. Expert Syst. Appl. 2023, 234, 121047. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015; Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F., Eds.; Springer: Cham, Switzerland, 2015; pp. 234–241. [Google Scholar] [CrossRef]
- Oktay, O.; Schlemper, J.; Folgoc, L.L.; Lee, M.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.; Hammerla, N.Y.; Kainz, B.; et al. Attention u-net: Learning where to look for the pancreas. arXiv 2018, arXiv:1804.03999. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An image is worth 16 × 16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Yi, X.; Walia, E.; Babyn, P. Generative adversarial network in medical imaging: A review. Med. Image Anal. 2019, 58, 101552. [Google Scholar] [CrossRef]
- Reinhard, E.; Adhikhmin, M.; Gooch, B.; Shirley, P. Color transfer between images. IEEE Comput. Graph. Appl. 2001, 21, 34–41. [Google Scholar] [CrossRef]
- Onder, D.; Zengin, S.; Sarioglu, S. A review on color normalization and color deconvolution methods in histopathology. Appl. Immunohistochem. Mol. Morphol. 2014, 22, 713–719. [Google Scholar] [CrossRef]
- Khan, A.M.; Rajpoot, N.; Treanor, D.; Magee, D. A nonlinear mapping approach to stain normalization in digital histopathology images using image-specific color deconvolution. IEEE Trans. Biomed. Eng. 2014, 61, 1729–1738. [Google Scholar] [CrossRef] [PubMed]
- Vahadane, A.; Peng, T.; Sethi, A.; Albarqouni, S.; Wang, L.; Baust, M.; Steiger, K.; Schlitter, A.M.; Esposito, I.; Navab, N.; et al. Structure-preserving color normalization and sparse stain separation for histological images. IEEE Trans. Med. Imaging 2016, 35, 1962–1971. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Xiang, L.; Liu, Q.; Gilmore, H.; Wu, J.; Tang, J.; Madabhushi, A. Stacked sparse autoencoder (SSAE) for nuclei detection on breast cancer histopathology images. IEEE Trans. Med. Imaging 2015, 35, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. In Advances in Neural Information Processing Systems 27; Ghahramani, Z., Welling, M., Cortes, C., Lawrence, N., Weinberger, K.Q., Eds.; MIT Press: Cambridge, MA, USA, 2014. [Google Scholar]
- Isola, P.; Zhu, J.Y.; Zhou, T.; Efros, A.A. Image-to-image translation with conditional adversarial networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 1125–1134. [Google Scholar] [CrossRef]
- Zhu, J.Y.; Park, T.; Isola, P.; Efros, A.A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proceedings of the IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 2223–2232. [Google Scholar] [CrossRef]
- Choi, Y.; Choi, M.; Kim, M.; Ha, J.W.; Kim, S.; Choo, J. Stargan: Unified generative adversarial networks for multi-domain image-to-image translation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 8789–8797. [Google Scholar] [CrossRef]
- Karras, T.; Laine, S.; Aittala, M.; Hellsten, J.; Lehtinen, J.; Aila, T. Analyzing and improving the image quality of stylegan. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, 13–19 June 2020; pp. 8110–8119. [Google Scholar] [CrossRef]
- BenTaieb, A.; Hamarneh, G. Adversarial stain transfer for histopathology image analysis. IEEE Trans. Med. Imaging 2017, 37, 792–802. [Google Scholar] [CrossRef]
- Shaban, M.T.; Baur, C.; Navab, N.; Albarqouni, S. Staingan: Stain style transfer for digital histological images. In Proceedings of the 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), Venice, Italy, 8–11 April 2019; pp. 953–956. [Google Scholar] [CrossRef]
- Cong, C.; Liu, S.; Di Ieva, A.; Pagnucco, M.; Berkovsky, S.; Song, Y. Colour adaptive generative networks for stain normalisation of histopathology images. Med. Image Anal. 2022, 82, 102580. [Google Scholar] [CrossRef]
- Jeong, J.; Kim, K.D.; Nam, Y.; Cho, C.E.; Go, H.; Kim, N. Stain normalization using score-based diffusion model through stain separation and overlapped moving window patch strategies. Comput. Biol. Med. 2023, 152, 106335. [Google Scholar] [CrossRef]
- Yang, H.; Lyu, M.; Yan, S.; Zhong, T.; Li, J.; Xu, T.; Xie, H.; Liu, S. Sastaindiff: Self-supervised stain normalization by stain augmentation using denoising diffusion probabilistic models. Biomed. Signal Process. Control 2025, 107, 107861. [Google Scholar] [CrossRef]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar] [CrossRef]
- Chen, J.; Lu, Y.; Yu, Q.; Luo, X.; Adeli, E.; Wang, Y.; Lu, L.; Yuille, A.L.; Zhou, Y. Transunet: Transformers make strong encoders for medical image segmentation. arXiv 2021, arXiv:2102.04306. [Google Scholar] [CrossRef]
- Zhang, T.; Wei, Q.; Li, Z.; Meng, W.; Zhang, M.; Zhang, Z. Segmentation of paracentral acute middle maculopathy lesions in spectral-domain optical coherence tomography images through weakly supervised deep convolutional networks. Comput. Methods Programs Biomed. 2023, 240, 107632. [Google Scholar] [CrossRef]
- Han, C.; Yao, H.; Zhao, B.; Li, Z.; Shi, Z.; Wu, L.; Chen, X.; Qu, J.; Zhao, K.; Lan, R.; et al. Meta multi-task nuclei segmentation with fewer training samples. Med. Image Anal. 2022, 80, 102481. [Google Scholar] [CrossRef] [PubMed]
- Kirillov, A.; Mintun, E.; Ravi, N.; Mao, H.; Rolland, C.; Gustafson, L.; Xiao, T.; Whitehead, S.; Berg, A.C.; Lo, W.Y.; et al. Segment anything. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Paris, France, 2–6 October 2023; pp. 4015–4026. [Google Scholar] [CrossRef]
- Ma, J.; He, Y.; Li, F.; Han, L.; You, C.; Wang, B. Segment anything in medical images. Nat. Commun. 2024, 15, 654. [Google Scholar] [CrossRef] [PubMed]
- Griebel, T.; Archit, A.; Pape, C. Segment anything for histopathology. arXiv 2025, arXiv:2502.00408. [Google Scholar] [CrossRef]
- Zhang, Z.; Yang, L.; Zheng, Y. Translating and segmenting multimodal medical volumes with cycle-and shape-consistency generative adversarial network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 9242–9251. [Google Scholar] [CrossRef]
- Ren, M.; Dey, N.; Fishbaugh, J.; Gerig, G. Segmentation-renormalized deep feature modulation for unpaired image harmonization. IEEE Trans. Med. Imaging 2021, 40, 1519–1530. [Google Scholar] [CrossRef]
- Tomczak, A.; Ilic, S.; Marquardt, G.; Engel, T.; Forster, F.; Navab, N.; Albarqouni, S. Multi-task multi-domain learning for digital staining and classification of leukocytes. IEEE Trans. Med. Imaging 2020, 40, 2897–2910. [Google Scholar] [CrossRef]
- Jin, S.; Yu, S.; Peng, J.; Wang, H.; Zhao, Y. A novel medical image segmentation approach by using multi-branch segmentation network based on local and global information synchronous learning. Sci. Rep. 2023, 13, 6762. [Google Scholar] [CrossRef]
- Yalniz, I.Z.; Jégou, H.; Chen, K.; Paluri, M.; Mahajan, D. Billion-scale semi-supervised learning for image classification. arXiv 2019, arXiv:1905.00546. [Google Scholar]
- Shaw, S.; Pajak, M.; Lisowska, A.; Tsaftaris, S.A.; O'Neil, A.Q. Teacher-student chain for efficient semi-supervised histology image classification. arXiv 2020, arXiv:2003.08797. [Google Scholar]
- Lee, E.J.E. Corrective feedback preferences and learner repair among advanced ESL students. System 2013, 41, 217–230. [Google Scholar] [CrossRef]
- Zhang, D.; Chen, B.; Chong, J.; Li, S. Weakly-supervised teacher-student network for liver tumor segmentation from non-enhanced images. Med. Image Anal. 2021, 70, 102005. [Google Scholar] [CrossRef]
- Xiao, Z.; Su, Y.; Deng, Z.; Zhang, W. Efficient combination of cnn and transformer for dual-teacher uncertainty-guided semi-supervised medical image segmentation. Comput. Methods Programs Biomed. 2022, 226, 107099. [Google Scholar] [CrossRef]
- Lu, S.; Zhang, Z.; Yan, Z.; Wang, Y.; Cheng, T.; Zhou, R.; Yang, G. Mutually aided uncertainty incorporated dual consistency regularization with pseudo label for semi-supervised medical image segmentation. Neurocomputing 2023, 548, 126411. [Google Scholar] [CrossRef]
- Sara, U.; Akter, M.; Uddin, M.S. Image quality assessment through fsim, ssim, mse and psnr—a comparative study. J. Comput. Commun. 2019, 7, 8–44. [Google Scholar] [CrossRef]
- Kumar, N.; Verma, R.; Sharma, S.; Bhargava, S.; Vahadane, A.; Sethi, A. A dataset and a technique for generalized nuclear segmentation for computational pathology. IEEE Trans. Med. Imaging 2017, 36, 1550–1560. [Google Scholar] [CrossRef]
- Zhao, B.; Han, C.; Pan, X.; Lin, J.; Yi, Z.; Liang, C.; Chen, X.; Li, B.; Qiu, W.; Li, D.; et al. Restainnet: a self-supervised digital re-stainer for stain normalization. Comput. Electr. Eng. 2022, 103, 108304. [Google Scholar] [CrossRef]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, F.F. Imagenet: A large-scale hierarchical image database. In Proceedings of the 2009 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR 2009), Miami, FL, USA, 20–25 June 2009; pp. 248–255. [Google Scholar] [CrossRef]
- Cho, J.H.; Hariharan, B. On the efficacy of knowledge distillation. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Seoul, Republic of Korea, 27 October–2 November 2019; pp. 4794–4802. [Google Scholar] [CrossRef]
- Kumar, N.; Verma, R.; Anand, D.; Zhou, Y.; Onder, O.F.; Tsougenis, E.; Chen, H.; Heng, P.A.; Li, J.; Hu, Z.; et al. A multi-organ nucleus segmentation challenge. IEEE Trans. Med. Imaging 2019, 39, 1380–1391. [Google Scholar] [CrossRef]
- Gamper, J.; Alemi Koohbanani, N.; Benet, K.; Khuram, A.; Rajpoot, N. Pannuke: An open pan-cancer histology dataset for nuclei instance segmentation and classification. In European Congress on Digital Pathology; Reyes-Aldasoro, C.C., Janowczyk, A., Veta, M., Bankhead, P., Sirinukunwattana, K., Eds.; Springer: Cham, Switzerland, 2019; pp. 11–19. [Google Scholar] [CrossRef]

| Metric | Value |
|---|---|
| Total WSIs | 1,408 |
| Total stain-style clusters (GMM, BIC-selected) | 101 |
| Median WSIs per cluster (IQR) | 11 (7–18) |
| Clusters present in both hospitals | 84 (83.2%) |
| Clusters enriched in Hospital A only | 9 (8.9%) |
| Clusters enriched in Hospital B only | 8 (7.9%) |
| Mean within-cluster ΔE2000 ± SD | 3.4 ± 0.9 |
| Mean between-cluster ΔE2000 ± SD | 8.7 ± 2.1 |
| Medical Centers | Train (X→Y) | Train (Y→X) | Validate (X→Y) | Validate (Y→X) | Test (X→Y) | Test (Y→X) | Total |
|---|---|---|---|---|---|---|---|
| Huashan Hospital (HS) | 312 | 320 | 104 | 112 | 104 | 96 | 848 |
| CUHK | 198 | 190 | 66 | 58 | 66 | 82 | 560 |
| Total | 510 | 510 | 170 | 170 | 170 | 178 | 1,408 |
| Tasks | Methods | Dataset | Metrics | |||
|---|---|---|---|---|---|---|
| Labeled | Pseudo-labeled | F1↑ | mIOU↑ | AJI↑ | ||
|
Nuclei segmenta-tion |
3SGAN | ✓ | ✓(20002) | 0.8140±0.0042 | 0.8201±0.0040 | 0.6915±0.0045 |
| SAM [31] | 0.7325±0.0095 | 0.7518±0.0090 | 0.6011±0.0102 | |||
| MedSAM [32] | 0.7781±0.0080 | 0.7724±0.0078 | 0.6325±0.0088 | |||
| TransUNet [28] | ✓ | 0.7427±0.0088 | 0.7554±0.0085 | 0.5588±0.0105 | ||
| Attention-UNet [9] | ✓ | 0.6602±0.0120 | 0.6947±0.0115 | 0.5971±0.0110 | ||
| U-Net [8] | ✓ | 0.7115±0.0105 | 0.7358±0.0100 | 0.5728±0.0112 | ||
|
Stain normaliza-tion |
RMSE↓ | PSNR↓ | SSIM↑ | |||
| 3SGAN | ✓ | ✓(20002) | 0.0908±0.0020 | 21.0615±0.1650 | 0.8556±0.0042 | |
| 2SGAN | ✓ | ✓(20002) | 0.1008±0.0035 | 20.1358±0.2800 | 0.8474±0.0065 | |
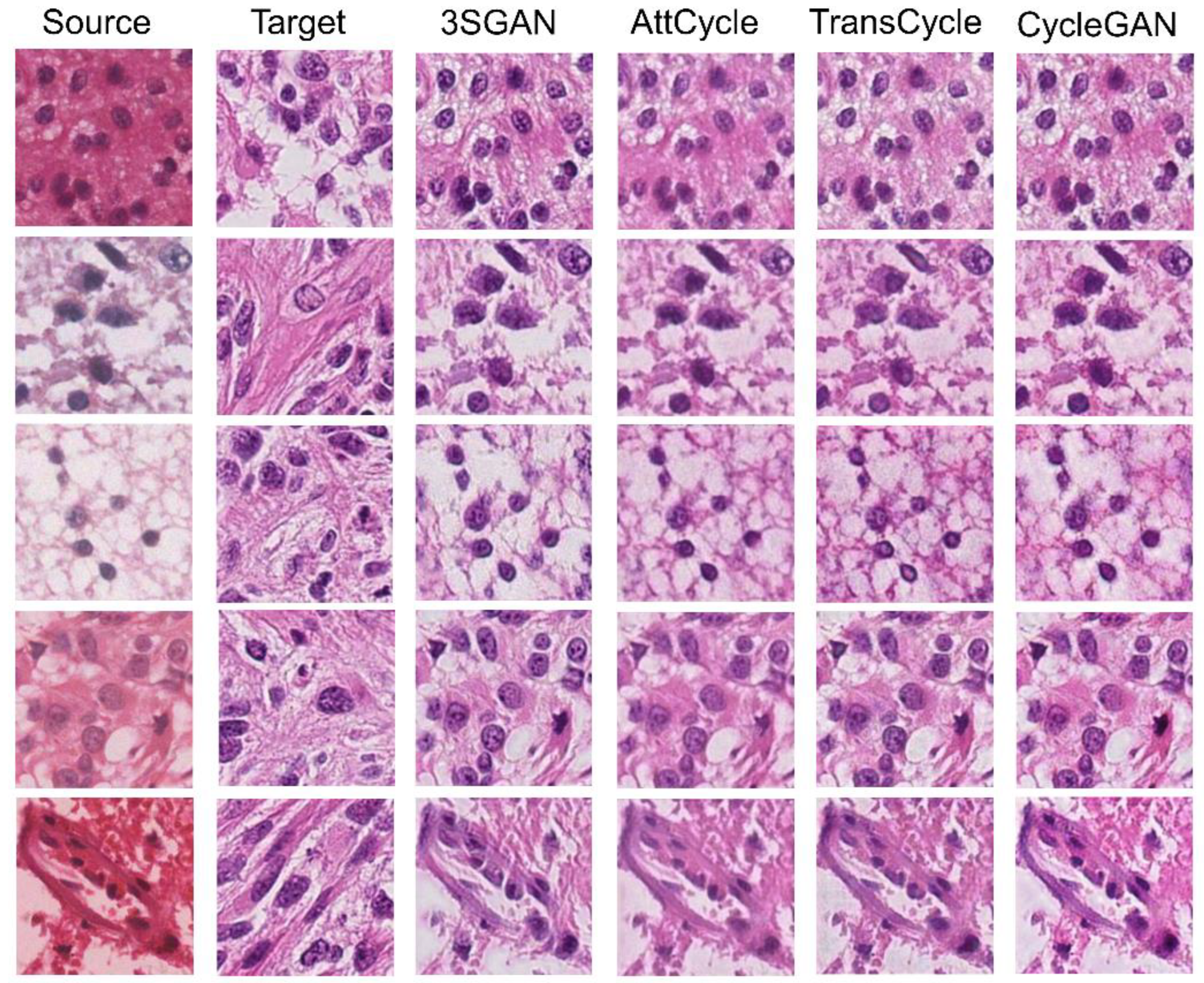
| CycleGAN [19] | ✓ | 0.1018±0.0040 | 20.1150±0.3200 | 0.8140±0.0095 | ||
| Models | Designs | Parameter Amounts | |||
|---|---|---|---|---|---|
| DD | AT | SC | TR | ||
| TransCycle | ✓ | ✓ | 115,938,852 | ||
| AttCycle | ✓ | ✓ | 23,519,248 | ||
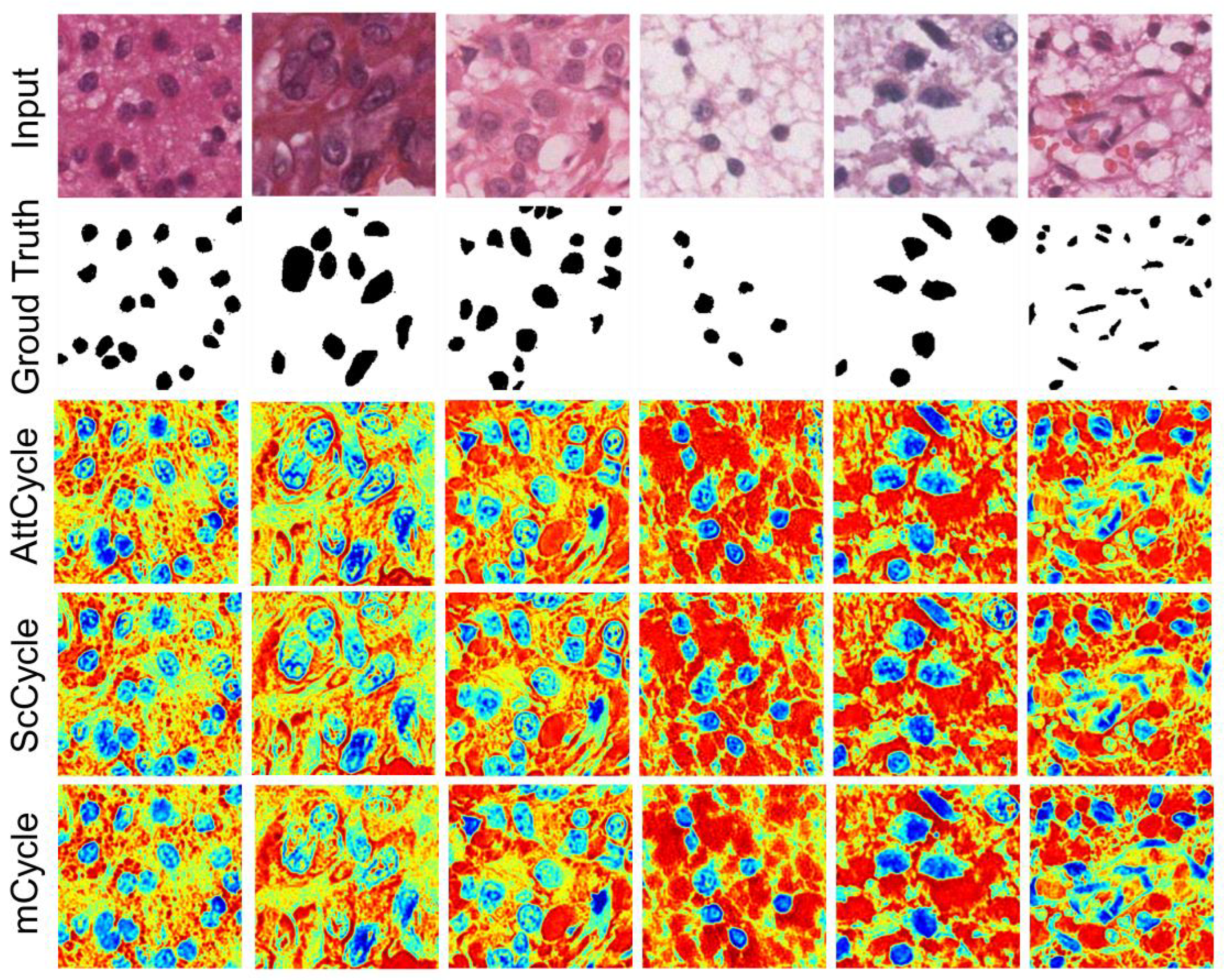
| ScCycle | ✓ | ✓ | 22,740,228 | ||
| mCycle | ✓ | 22,371,588 | |||
| Tasks | Methods | Dataset | Metrics | |||
|---|---|---|---|---|---|---|
| Labeled | Pseudo-labeled | F1↑ | mIOU↑ | AJI↑ | ||
|
Nuclei Segmenta-tion |
3SGAN | ✓ | ✓(20002) | 0.8140±0.0042 | 0.8201±0.0040 | 0.6915±0.0045 |
| AttCycle | ✓ | 0.8095±0.0065 | 0.8156±0.0062 | 0.6835±0.0070 | ||
| TransCycle | ✓ | 0.7733±0.0088 | 0.7888±0.0085 | 0.6382±0.0092 | ||
| ScCycle | ✓ | 0.8089±0.0068 | 0.8154±0.0065 | 0.6834±0.0072 | ||
| mCycle | ✓ | 0.8008±0.0072 | 0.8081±0.0070 | 0.6770±0.0078 | ||
| SAM [31] | ✓ | 0.7325±0.0095 | 0.7518±0.0090 | 0.6011±0.0102 | ||
| MedSAM [32] | ✓ | 0.7781±0.0080 | 0.7724±0.0078 | 0.6325±0.0088 | ||
| TransUNet [28] | ✓ | 0.7427±0.0088 | 0.7554±0.0085 | 0.5588±0.0105 | ||
| Attention U-Net [9] | ✓ | 0.6602±0.0120 | 0.6947±0.0115 | 0.5971±0.0110 | ||
| U-Net [8] | ✓ | 0.7115±0.0105 | 0.7358±0.0100 | 0.5728±0.0112 | ||
|
Stain Normaliza-tion |
✓ | RMSE↓ | PSNR↑ | SSIM↑ | ||
| 3SGAN | ✓ | ✓(20002) | 0.0908±0.0020 | 21.0615±0.1650 | 0.8556±0.0042 | |
| AttCycle | ✓ | 0.1016±0.0038 | 20.0063±0.3100 | 0.8276±0.0085 | ||
| TransCycle | ✓ | 0.1054±0.0042 | 19.7959±0.3500 | 0.8275±0.0090 | ||
| ScCycle | ✓ | 0.0995±0.0035 | 20.2664±0.2900 | 0.8354±0.0080 | ||
| mCycle | ✓ | 0.0984±0.0032 | 20.3820±0.2700 | 0.8197±0.0088 | ||
| CycleGAN [19] | ✓ | 0.1018±0.0040 | 20.1150±0.3200 | 0.8140±0.0095 | ||
| Amount of Pseudo Labels | Metrics | ||||||
|---|---|---|---|---|---|---|---|
| Nuclei segmentation | Stain normalization | ||||||
| F1↑ | mIOU↑ | AJI↑ | RMSE↓ | PSNR↑ | SSIM↑ | ||
| 0 | 0.7733 ±0.0088 |
0.7888 ±0.0085 |
0.6382 ±0.0092 |
0.1054 ±0.0042 |
19.7959 ±0.3500 |
0.8275 ±0.0090 |
|
| 5002 | 0.7923 ±0.0065 |
0.8018 ±0.0062 |
0.6722 ±0.0070 |
0.1015 ±0.0035 |
20.1354 ±0.2800 |
0.8413 ±0.0075 |
|
| 20002 | 0.8140 ±0.0042 |
0.8201 ±0.0040 |
0.6915 ±0.0045 |
0.0908 ±0.0020 |
21.0615 ±0.1650 |
0.8556 ±0.0042 |
|
| 50002 | 0.8153 ±0.0045 |
0.8199 ±0.0043 |
0.6928 ±0.0048 |
0.1150 ±0.0055 |
18.9607 ±0.4200 |
0.8231 ±0.0105 |
|
| Tasks | Methods | MoNuSeg | PanNuke | ||||
|---|---|---|---|---|---|---|---|
| F1↑ | mIOU↑ | AJI↑ | F1↑ | mIOU↑ | AJI↑ | ||
|
Nuclei Segmentation |
U-Net [8] | 0.8000 ±0.0080 |
0.6800 ±0.0085 |
0.5900 ±0.0092 |
0.7900 ±0.0085 |
0.6600 ±0.0090 |
0.5700 ±0.0098 |
| Attention U-Net [9] | 0.8200 ±0.0072 |
0.7000 ±0.0078 |
0.6100 ±0.0085 |
0.8100 ±0.0078 |
0.6800 ±0.0083 |
0.5900 ±0.0090 |
|
| TransUNet [28] | 0.8300 ±0.0065 |
0.7100 ±0.0070 |
0.6200 ±0.0076 |
0.8200 ±0.0070 |
0.6900 ±0.0075 |
0.6000 ±0.0082 |
|
| SAM [31] | 0.7900 ±0.0092 |
0.6600 ±0.0098 |
0.5700 ±0.0105 |
0.7800 ±0.0095 |
0.6500 ±0.0102 |
0.5600 ±0.0110 |
|
| MedSAM [32] | 0.8100 ±0.0078 |
0.6900 ±0.0082 |
0.6000 ±0.0090 |
0.8000 ±0.0080 |
0.6700 ±0.0087 |
0.5800 ±0.0095 |
|
| 3SGAN |
0.8500 ±0.0038 |
0.7300 ±0.0040 |
0.6500 ±0.0043 |
0.8400 ±0.0040 |
0.7200 ±0.0042 |
0.6400 ±0.0045 |
|
|
Stain Normalization |
Methods | MoNuSeg | PanNuke | ||||
| RMSE↓ | PSNR↑ | SSIM↑ | RMSE↓ | PSNR↑ | SSIM↑ | ||
| CycleGAN [19] | 0.1040 ±0.0048 |
19.6200±0.3800 | 0.8100 ±0.0115 |
0.1060 ±0.0052 |
19.4200±0.4100 | 0.8080 ±0.0120 |
|
| AttCycle | 0.0990 ±0.0040 |
20.1200±0.3200 | 0.8350 ±0.0098 |
0.1010 ±0.0045 |
19.9400±0.3600 | 0.8300 ±0.0105 |
|
| TransCycle | 0.1020 ±0.0042 |
19.8800±0.3500 | 0.8280 ±0.0100 |
0.1040 ±0.0048 |
19.7100±0.3700 | 0.8220 ±0.0108 |
|
| ScCycle | 0.0980 ±0.0038 |
20.3400±0.3000 | 0.8380 ±0.0095 |
0.1000 ±0.0042 |
20.0800±0.3400 | 0.8340 ±0.0102 |
|
| mCycle | 0.0970 ±0.0035 |
20.5100±0.2800 | 0.8420 ±0.0090 |
0.0990 ±0.0040 |
20.2700±0.3200 | 0.8400 ±0.0098 |
|
| 3SGAN |
0.0920 ±0.0020 |
21.0300±0.1600 |
0.8580 ±0.0040 |
0.0930 ±0.0023 |
20.9200±0.1900 |
0.8550 ±0.0045 |
|
| Tasks | Methods | Metrics | ||||
|---|---|---|---|---|---|---|
| F1↑ | mIOU↑ | AJI↑ | ||||
|
Nuclei Segmentation |
U-Net [8] | 0.7900±0.0082 | 0.6700±0.0088 | 0.5800±0.0095 | ||
| Attention U-Net [9] | 0.8100±0.0075 | 0.6900±0.0080 | 0.6000±0.0087 | |||
| TransUNet [28] | 0.8200±0.0068 | 0.7000±0.0072 | 0.6100±0.0078 | |||
| SAM [31] | 0.8000±0.0090 | 0.6800±0.0095 | 0.5900±0.0102 | |||
| MedSAM [32] | 0.8200±0.0070 | 0.7000±0.0075 | 0.6100±0.0080 | |||
| 3SGAN | 0.8700±0.0035 | 0.7400±0.0038 | 0.6600±0.0041 | |||
|
Stain Normalization |
RMSE↓ | PSNR↑ | SSIM↑ | |||
| CycleGAN [19] | 0.1050±0.0050 | 19.8000±0.4200 | 0.8120±0.0130 | |||
| AttCycle | 0.1010±0.0042 | 20.1800±0.3500 | 0.8320±0.0105 | |||
| TransCycle | 0.1040±0.0045 | 19.9300±0.3800 | 0.8250±0.0112 | |||
| ScCycle | 0.1000±0.0040 | 20.3000±0.3300 | 0.8370±0.0100 | |||
| mCycle | 0.0990±0.0038 | 20.4500±0.3100 | 0.8230±0.0108 | |||
| 3SGAN | 0.0930±0.0025 | 20.9500±0.2000 | 0.8570±0.0045 | |||
| Different training strategies | Metrics | ||||||
|---|---|---|---|---|---|---|---|
| Nuclei segmentation | Stain normalization | ||||||
| F1↑ | mIOU↑ | AJI↑ | RMSE↓ | PSNR↑ | SSIM↑ | ||
| Semisupervised | 0.0407 ±0.0031 |
0.0313 ±0.0028 |
0.0546 ±0.0040 |
−0.0150 ±0.0018 |
1.2656 ±0.1120 |
0.0281 ±0.0035 |
|
| Self-training | 0.0158 ±0.0042 |
0.0140 ±0.0035 |
0.0129 ±0.0048 |
−0.0127 ±0.0022 |
1.2403 ±0.1350 |
0.0471 ±0.0052 |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




