Submitted:
30 September 2025
Posted:
30 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Prostate Cancer Epidemiology
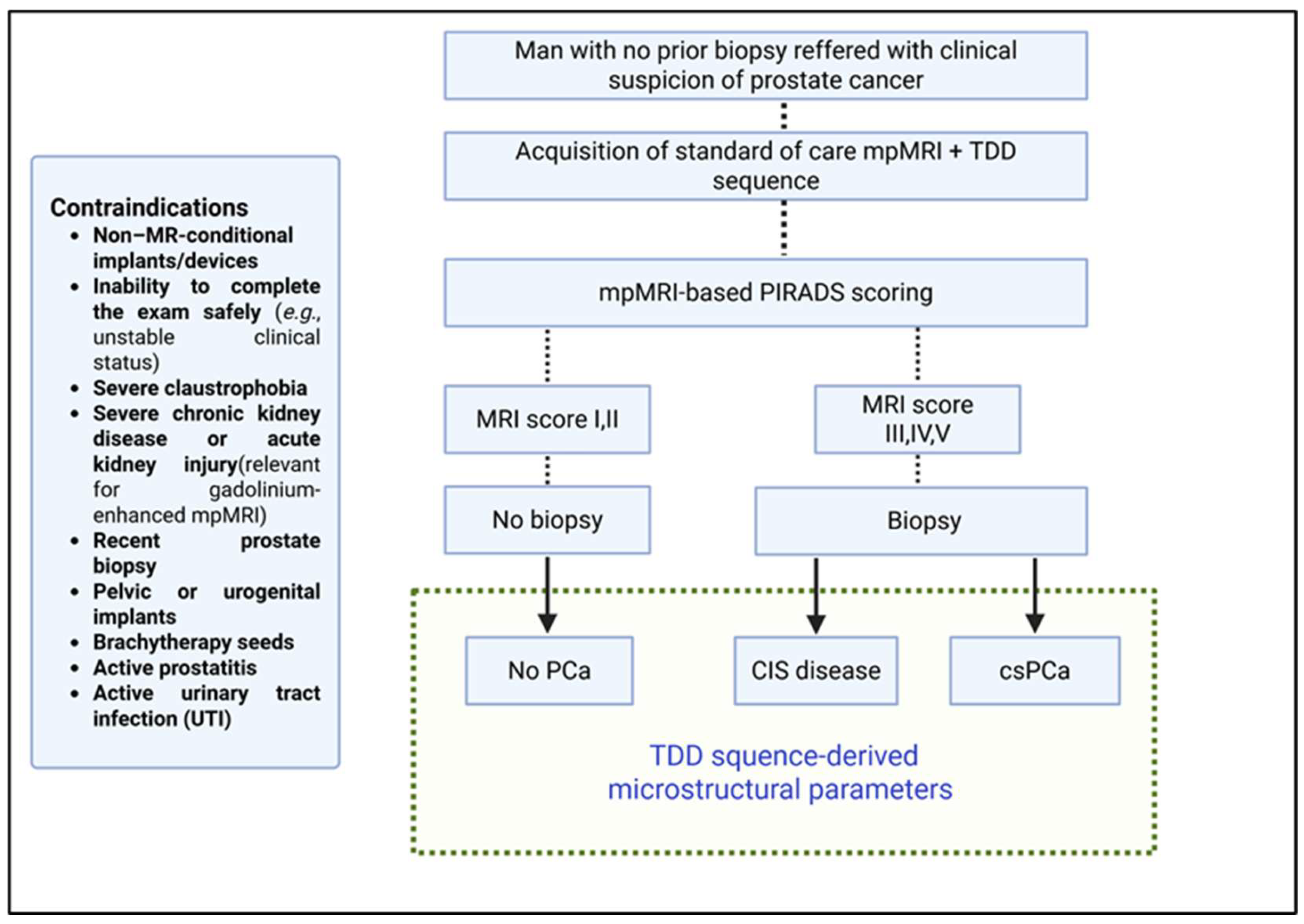
1.2. Overall Prostate Cancer Diagnostic Workflow
1.3. Relevance of Non-Invasive Evaluation by mpMRI
1.4. Diagnostic Limitations and Inter-Observer Variability of mpMRI
1.5. Time-Dependent Diffusion MRI
1.6. Further Possibilities with the TDD Sequence
1.7. Deep Learning-Based Interpretation
2. Materials and Methods
2.1. Study Design
2.2. Study Population
- PI-RADS 1: very low (clinically significant cancer highly unlikely)
- PI-RADS 2: low (clinically significant cancer unlikely)
- PI-RADS 3: intermediate (equivocal)
- PI-RADS 4: high (clinically significant cancer likely)
- PI-RADS 5: very high (clinically significant cancer highly likely)
2.3. Prostate Magnetic Resonance Imaging Data Acquisition and Fitting for Determination of Microstructural Parameters
2.4. Artificial Intelligence-Based Automatic Delineation of the Prostate Gland Zones
2.4.1. Training Dataset and Human-in-the-Loop Strategy
2.4.2. Segmentation Models
2.4.3. Integration with Microstructural Analysis
2.5. Radiological Imaging Analysis
2.6. Histopathologic Analysis
2.7. Statistical Analysis
3. Expected Results
3.1. Deep Learning Models
- Prostate Segmentation Model: Based on U-Net or ProGNet architectures, initially trained on the PROSTATEx dataset [38] and subsequently fine-tuned using multiparametric MRI (mpMRI) data from the Clinical Hospital of the University of Chile (HCUCH). This model is expected to accurately delineate the prostate gland and serve as a pre-processing step for downstream microstructural analysis. For the segmentation task, we anticipate achieving a Dice Similarity Coefficient (DSC) of approximately 0.92, consistent with state-of-the-art literature.
- Tissue Microstructure Estimation Model: Employing a Transformer-based architecture inspired by sparse representation techniques; METSC [46], this model will estimate voxel-wise tissue microstructural parameters, including intracellular and extracellular volume fractions and diffusivities, from multi-shell diffusion MRI (dMRI) data. These microstructural parameters will then be used for tissue classification. For microstructure estimation and lesion classification, we anticipate an accuracy exceeding 80%.
3.2. Pipeline Integration
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PCa | Prostate cancer |
| PSA | Prostate-specific antigen |
| DRE | Digital rectal exam |
| mpMRI | multiparametric MRI |
| csPCa | Clinically significant prostate cancer |
| Gs | Gleason score |
| ISUP | International Society of Urological Pathology |
| CIS | Clinical insignificant prostate cancer |
| TDD | Time-dependent diffusion |
| GG | Gleason grade |
References
- International Agency for Research on Cancer. World: Cancer fact sheet (GLOBOCAN 2022, version 1.1) [Internet]. Lyon (FR): International Agency for Research on Cancer; 2024 [cited 2025 Jul 23]. Available from: https://gco.iarc.who.int/media/globocan/factsheets/populations/900-world-fact-sheet.pdf.
- Cornford P, van den Bergh RCN, Briers E, Van den Broeck T, Brunckhorst O, Darraugh J, et al. EAU-EANM-ESTRO-ESUR-ISUP-SIOG Guidelines on Prostate Cancer – 2024 Update. Part I: Screening, Diagnosis, and Local Treatment with Curative Intent. Eur Urol. 2024;86(2):148-163. [CrossRef]
- Chatterjee A, Watson G, Myint E, Sved P, McEntee M, Bourne R. Changes in epithelium, stroma, and lumen space correlate more strongly with Gleason pattern and are stronger predictors of prostate ADC changes than cellularity metrics. Radiology. 2015;277(3):751-62. [CrossRef]
- Mortezavi A, Märzendorfer O, Donati OF, Rizzi G, Rupp NJ, Wettstein MS, et al. Diagnostic accuracy of multiparametric magnetic resonance imaging and fusion-guided targeted biopsy evaluated by transperineal template saturation prostate biopsy for the detection and characterization of prostate cancer. J Urol. 2018;200(2):309-18. [CrossRef]
- van der Leest MMG, Cornel EB, Israël B, Hendriks R, Padhani AR, Hoogenboom M, et al. Head-to-head comparison of transrectal ultrasound-guided prostate biopsy versus multiparametric prostate resonance imaging with subsequent magnetic resonance-guided biopsy in biopsy-naïve men with elevated prostate-specific antigen: a large prospective multicenter clinical study. Eur Urol. 2019;75(4):570-8. [CrossRef]
- Ahmed HU, El-Shater Bosaily A, Brown LC, Gabe R, Kaplan R, Parmar MK, et al. Diagnostic accuracy of multi-parametric MRI and TRUS biopsy in prostate cancer (PROMIS): a paired validating confirmatory study. Lancet. 2017;389(10071):815-22. [CrossRef]
- Pourvaziri A. Mass enhancement pattern at prostate MRI as a potential PI-RADS criterion. Radiol Imaging Cancer. 2022;4(2):e229006. [CrossRef]
- Park SY, Park BK, Kwon GY. Diagnostic performance of mass enhancement on dynamic contrast-enhanced MRI for predicting clinically significant peripheral zone prostate cancer. AJR Am J Roentgenol. 2020;214(4):792-799. [CrossRef]
- Agrotis G, Pais Pooch E, Marsitopoulos K, Vlychou M, Benndorf M, Beets-Tan RGH, Schoots IG. Detection rates for prostate cancer using PI-RADS 2.1 upgrading rules in transition zone lesions align with risk assessment categories: a systematic review and meta-analysis. Eur Radiol. 2025 Apr 27. [CrossRef]
- Wei CG, Zhang YY, Pan P, Chen T, Yu HC, Dai GC, et al. Diagnostic accuracy and interobserver agreement of PI-RADS Version 2 and Version 2.1 for the detection of transition zone prostate cancers. AJR Am J Roentgenol. 2021;216(5):1247-56. [CrossRef]
- Bhayana R, O’Shea A, Anderson MA, Bradley WR, Gottumukkala RV, Mojtahed A, et al. PI-RADS Versions 2 and 2.1: Interobserver agreement and diagnostic performance in peripheral and transition zone lesions among six radiologists. AJR Am J Roentgenol. 2021;217(1):141-51. [CrossRef]
- Wen J, Liu W, Shen X, Hu W. PI-RADS v2.1 and PSAD for the prediction of clinically significant prostate cancer among patients with PSA levels of 4–10 ng/ml. Sci Rep. 2024;14:6570. [CrossRef]
- Lee CH, Vellayappan B, Tan CH. Comparison of diagnostic performance and inter-reader agreement between PI-RADS v2.1 and PI-RADS v2: systematic review and meta-analysis. Br J Radiol. 2022;95(1131):20210509. [CrossRef]
- Wang W, Zhu M, Luo Z, Li F, Wan C, Zhu L. Diagnostic value analysis of PI-RADS v2.1 combined with ADC values in the risk stratification of prostate cancer Gleason scores: a retrospective study. Arch Esp Urol. 2024;77(8):889-896. [CrossRef]
- Wei X, Xu J, Zhong S, Zou J, Cheng Z, Ding Z, et al. Diagnostic value of combining PI-RADS v2.1 with PSAD in clinically significant prostate cancer. Abdom Radiol (NY). 2022;47(10):3574-3582. [CrossRef]
- Colvin DC, Yankeelov TE, Does MD, Yue Z, Quarles C, Gore JC. New insights into tumor microstructure using temporal diffusion spectroscopy. Cancer Res. 2008;68(14):5941-7. [CrossRef]
- Colvin DC, Loveless ME, Does MD, Yue Z, Yankeelov TE, Gore JC. Earlier detection of tumor treatment response using magnetic resonance diffusion imaging with oscillating gradients. Magn Reson Imaging. 2011;29(3):315-23. [CrossRef]
- Xu J, Li K, Smith RA, Waterton JC, Zhao P, Chen H, et al. Characterizing tumor response to chemotherapy at various length scales using temporal diffusion spectroscopy. PLOS One. 2012;7(7):e41714. [CrossRef]
- Reynaud O, Winters KV, Hoang DM, Wadghiri YZ, Novikov DS, Kim SG. Surface-to-volume ratio mapping of tumor microstructure using oscillating gradient diffusion-weighted imaging. Magn Reson Med. 2016;76(1):237-47. [CrossRef]
- Baron CA, Beaulieu C. Oscillating gradient spin-echo (OGSE) diffusion tensor imaging of the human brain. Magn Reson Med. 2014;72(3):726-736. [CrossRef]
- Van AT, Holdsworth SJ, Bammer R. In vivo investigation of restricted diffusion in the human brain with optimized oscillating diffusion gradient encoding. Magn Reson Med. 2014;71(1):83-94. [CrossRef]
- Iima M, Yamamoto A, Kataoka M, Yamada Y, Omori K, Feiweier T, et al. Time-dependent diffusion MRI to distinguish malignant from benign head and neck tumors. J Magn Reson Imaging. 2019;50(1):88-95. [CrossRef]
- Xu J, Jiang X, Li H, Arlinghaus LR, McKinley ET, Devan SP, et al. Magnetic resonance imaging of mean cell size in human breast tumors. Magn Reson Med. 2020;83(6):2002-14. [CrossRef]
- Lemberskiy G, Rosenkrantz AB, Veraart J, Taneja SS, Novikov DS, Fieremans E. Time-dependent diffusion in prostate cancer. Invest Radiol. 2017;52(7):405-11. [CrossRef]
- Xu J, Jiang X, Devan SP, Arlinghaus LR, McKinley ET, Xie J, et al. MRI-cytometry: Mapping non-parametric cell size distributions using diffusion MRI. Magn Reson Med. 2021;85(2):748-61. [CrossRef]
- Jiang X, Li H, Xie J, McKinley ET, Zhao P, Gore JC, et al. In vivo imaging of cancer cell size and cellularity using temporal diffusion spectroscopy. Magn Reson Med. 2017;78(1):156-164. [CrossRef]
- Wu D, Jiang K, Li H, Zhang Z, Ba R, Zhang Y, et al. Time-dependent diffusion MRI for quantitative microstructural mapping of prostate cancer. Radiology. 2022;303(3):578-87. [CrossRef]
- Chatterjee A, Oto A. Prostate tissue microstructural estimates using time-dependent diffusion MRI. Radiology. 2022;303(3):588-589. [CrossRef]
- Fryback DG, Thornbury JR. The efficacy of diagnostic imaging. Med Decis Making. 1991;11(2):88-94. [CrossRef]
- Purysko AS, Rosenkrantz AB, Barentsz JO, Weinreb JC, Macura KJ. PI-RADS Version 2: A pictorial update. Radiographics. 2016;36(5):1354-1372. [CrossRef]
- van Sloun RJG, Wildeboer RR, Mannaerts CK, Postema AW, Gayet M, Beerlage HP, et al. Deep learning for real-time, automatic, and scanner-adapted prostate (zone) segmentation of transrectal ultrasound, for example, magnetic resonance imaging–transrectal ultrasound fusion prostate biopsy. Eur Urol Focus. 2021;7(1):78-85. [CrossRef]
- Bardis M, Houshyar R, Chantaduly C, Tran-Harding K, Ushinsky A, Chahine C, et al. Segmentation of the prostate transition zone and peripheral zone on MR images with deep learning. Radiol Imaging Cancer. 2021;3(3):e200024. [CrossRef]
- Glazer DI, Mayo-Smith WW, Sainani NI, Sadow CA, Vangel MG, Tempany CM, et al. Interreader agreement of Prostate Imaging Reporting and Data System version 2 using an in-bore MRI-guided prostate biopsy cohort: a single institution’s initial experience. AJR Am J Roentgenol. 2017;209(3):W145-W151. [CrossRef]
- Hectors SJ, Chen C, Chen J, Wang J, Gordon S, Yu M, et al. Magnetic resonance imaging radiomics-based machine learning prediction of clinically significant prostate cancer in equivocal PI-RADS 3 lesions. J Magn Reson Imaging. 2021;54(5):1466-1473. [CrossRef]
- Hansen NL, Patruno G, Wadhwa K, Gaziev G, Miano R, Barrett T, et al. Magnetic resonance and ultrasound image fusion supported transperineal prostate biopsy using the Ginsburg protocol: technique, learning points, and biopsy results. Eur Urol. 2016;70(2):332-340. [CrossRef]
- de Rooij M, Israël B, Tummers M, et al. ESUR/ESUI consensus statements on mpMRI quality requirements. Eur Radiol. 2020;30(10):5404-5416. [CrossRef]
- Emoryzzl. Prostate_impulse_MRI [Internet]. GitHub; [cited 2025 Aug 26]. Available from: https://github.com/Emoryzzl/Prostate_impulse_MRI.
- Litjens G, Debats O, Barentsz J, Karssemeijer N, Huisman H. SPIE-AAPM PROSTATEx Challenge Data (Version 2) [dataset]. The Cancer Imaging Archive; 2017. [CrossRef]
- Cuocolo R, Stanzione A, Castaldo A, De Lucia DR, Imbriaco M. Quality control and whole-gland, zonal and lesion annotations for the PROSTATEx challenge public dataset. Eur J Radiol. 2021;138:109647. [CrossRef]
- Vásquez-Venegas C, Sotomayor CG, Ramos B, Castañeda V, Pereira G, Cabrera-Vives G, Härtel S. Human-in-the-Loop — A Deep Learning Strategy in Combination with a Patient-Specific Gaussian Mixture Model Leads to the Fast Characterization of Volumetric Ground-Glass Opacity and Consolidation in the Computed Tomography Scans of COVID-19 Patients. J Clin Med. 2024;13(17):5231. [CrossRef]
- Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH. nnU-Net: Self-adapting framework for U-Net-based medical image segmentation [Internet]. GitHub; 2025 [cited 2025 Sep 28]. Available from: https://github.com/MIC-DKFZ/nnUNet.
- Project MONAI. MONAI—Medical Open Network for AI [Internet]. [cited 2025 Aug 26]. Available from: https://monai.io.
- Ma J, Wang B, An L, Zhang Y, Wang H, et al. MedSAM: Segment Anything in Medical Images [Internet]. GitHub; 2025 [cited 2025 Sep 28]. Available from: https://github.com/bowang-lab/MedSAM.
- Soerensen SJC. ProGNet: Deep learning for automatic prostate segmentation [Internet]. GitHub; 2021 [cited 2025 Sep 28]. Available from: https://github.com/simonjcs/ProGNet.
- Soerensen SJC, Fan RE, Seetharaman A, Chen L, Shao W, Bhattacharya I, Kim YH, Sood R, Borre M, Chung BI, To’o KJ, Rusu M, Sonn GA. Deep learning improves speed and accuracy of prostate gland segmentations on magnetic resonance imaging for targeted biopsy. J Urol. 2021;206(3):604-612. [CrossRef]
- Zheng T, Yan G, Li H, Zheng W, Shi W, Zhang Y, et al. A microstructure estimation Transformer inspired by sparse representation for diffusion MRI. Med Image Anal. 2023;86:102788. [CrossRef]
- Epstein JI, Egevad L, Amin MB, Delahunt B, Srigley JR, Humphrey PA; Grading Committee. The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma: definition of grading patterns and proposal for a new grading system. Am J Surg Pathol. 2016;40(2):244–252. [CrossRef]
- Tukey JW. Comparing individual means in the analysis of variance. Biometrics. 1949;5(2):99–114. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



