Submitted:
19 September 2025
Posted:
24 September 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Material and Method
Quality and Eligibility Appraisal
Methodological Framework
Smoking-associated Respiratory Diseases Caused by Epigenetic Mechanisms
DNA Methylation
Histone Modifications
Non-Coding RNA Regulation
The Rehan et al. Study
Paternal Smoking and Epigenetic Transmission
AI in Epigenetics
The Molecular Memory of the Genome
Disease Prediction
Respiratory Disease Prediction
Neurological Disease Prediction
From Prediction to Prevention
Challenges and Ethical Considerations
Future Perspectives
Discussion
The Importance of DNA Methylation
The HUNT Study
The future with the GrimAge clock
Past, Present and Future Potential
Conclusions
Abbreviations
| ADAM33 | A Disintegrin And Metalloprotease 33 |
| AHRR | Aryl-Hydrocarbon Receptor Repressor |
| AI | Artificial Intelligence |
| BAL | Bronchoalveolar Lavage |
| cg05575921 | CpG site 05575921 [in AHRR gene] |
| COPD | Chronic Obstructive Pulmonary Disease |
| DNA | Deoxyribonucleic Acid |
| ECV | Electronic Cigarette Vapour |
| EPIC-Seq | Enhanced Pooled ImmunoCapture Sequencing |
| ETS | Environmental Tobacco Smoke |
| FosB | FBJ Murine Osteosarcoma Viral Oncogene Homolog B |
| H3 | Histone H3 |
| HEP | Human Epigenome Project |
| HGP | Human Genome Project |
| IL13 | Interleukin 13 |
| lncRNA | Long Non-Coding RNA |
| MCS | Maternal Cigarette Smoke |
| miR-21 | MicroRNA 21 |
| miR-223 | MicroRNA 223 |
| miRNA | MicroRNA |
| NAcc | Nucleus Accumbens |
| PAH | Polycyclic Aromatic Hydrocarbons |
| PPARγ | Peroxisome Proliferator-Activated Receptor-γ |
| RNA | Ribonucleic Acid |
| SERPINA1 | Serpin Peptidase Inhibitor, Clade A, Member 1 |
References
- WHO Report on the Global Tobacco Epidemic 2021 : Addressing New and Emerging Products. World Health Organization; 2021.
- Skinner, M.K. Environmental epigenomics and disease susceptibility. EMBO Rep. 2011, 12, 620–622. [Google Scholar] [CrossRef]
- Jones, P.A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012, 13, 484–492. [Google Scholar] [CrossRef]
- Reik, W.; Dean, W.; Walter, J. Epigenetic reprogramming in mammalian development. Science. 2001, 293, 1089–1093. [Google Scholar] [CrossRef]
- Breton, C.V.; Byun, H.M.; Wenten, M.; Pan, F.; Yang, A.; Gilliland, F.D. Prenatal tobacco smoke exposure affects global and gene-specific DNA methylation. Am J Respir Crit Care Med. 2009, 180, 462–467. [Google Scholar] [CrossRef]
- Joubert, B.R.; Felix, J.F.; Yousefi, P.; Bakulski, K.M.; Just, A.C.; Breton, C.; et al. DNA Methylation in Newborns and Maternal Smoking in Pregnancy: Genome-wide Consortium Meta-analysis. Am J Hum Genet. 2016, 98, 680–696. [Google Scholar] [CrossRef]
- Zhang, S.; Jin, J.; Xu, B.; Zheng, Q.; Mou, H. The relationship between epigenetic biomarkers and the risk of diabetes and cancer: a machine learning modelling approach. Front Public Health. 2025, 13, 1509458. [Google Scholar] [CrossRef]
- Rauschert, S.; Raubenheimer, K.; Melton, P.E. ; Huang, R,C. Machine learning and clinical epigenetics: a review of challenges for diagnosis and classification. Clin Epigenetics. 2020, 12, 51. [Google Scholar] [CrossRef] [PubMed]
- Burton, N.O.; Greer, E.L. Multigenerational epigenetic inheritance: Transmitting information across generations. Semin Cell Dev Biol. 2022, 127, 121–132. [Google Scholar] [CrossRef]
- Campagna, M.P.; Xavier, A.; Lechner-Scott, J.; Maltby, V.; Scott, R.J.; Butzkueven, H.; et al. Epigenome-wide association studies: current knowledge, strategies and recommendations. Clin Epigenetics. 2021, 13, 214. [Google Scholar] [CrossRef] [PubMed]
- Brasil, S.; Neves, C.J.; Rijoff, T.; Falcão, M.; Valadão, G.; Videira, P.A.; et al. Artificial Intelligence in Epigenetic Studies: Shedding Light on Rare Diseases. Front Mol Biosci. 2021, 8, 648012. [Google Scholar] [CrossRef] [PubMed]
- Nishitani, S.; Smith, A.K.; Tomoda, A.; Fujisawa, TX. Data science using the human epigenome for predicting multifactorial diseases and symptoms. Epigenomics. 2024, 16, 273–276. [Google Scholar] [CrossRef]
- Vinciguerra, M. The Potential for Artificial Intelligence Applied to Epigenetics. Mayo Clinic proceedings Digital health. 2023, 1, 476–479. [Google Scholar] [CrossRef]
- P5 eHealth: An Agenda for the Health Technologies of the Future. Springer; 2020.
- Tobacco smoking and involuntary smoking. World Health Organization; 2004.
- Keshawarz, A.; Joehanes, R.; Guan, W.; Huan, T.; DeMeo, D.L.; Grove, M.L.; et al. Longitudinal change in blood DNA epigenetic signature after smoking cessation. Epigenetics. 2022, 17, 1098–1109. [Google Scholar] [CrossRef]
- Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002, 16, 6–21. [Google Scholar] [CrossRef]
- Zeilinger, S.; Kühnel, B.; Klopp, N.; Baurecht, H.; Kleinschmidt, A.; Gieger, C.; et al. Tobacco smoking leads to extensive genome-wide changes in DNA methylation. PLoS One. 2013, 8, e63812. [Google Scholar] [CrossRef]
- Suter, M.A.; Abramovici, A.R.; Griffin, E.; Branch, D.W.; Lane, R.H.; Mastrobattista, J.; et al. In utero nicotine exposure epigenetically alters fetal chromatin structure and differentially regulates transcription of the glucocorticoid receptor in a rat model. Birth Defects Res A Clin Mol Teratol. 2015, 103, 583–588. [Google Scholar] [CrossRef] [PubMed]
- MALLARBHATTACHARYA. Airway architect Adam33 in asthma. Science Translational Medicine. 2016, 8, 130.
- Nicodemus-Johnson, J.; Naughton, K.A.; Sudi, J.; Hogarth, K.; Naurekas, E.T.; Nicolae, D.L.; et al. Genome-Wide Methylation Study Identifies an IL-13-induced Epigenetic Signature in Asthmatic Airways. Am J Respir Crit Care Med. 2016, 193, 376–385. [Google Scholar] [CrossRef]
- Qiu, W.; Baccarelli, A.; Carey, V.J.; Boutaoui, N.; Bacherman, H.; Klanderman, B.; et al. Variable DNA methylation is associated with chronic obstructive pulmonary disease and lung function. Am J Respir Crit Care Med. 2012, 185, 373–381. [Google Scholar] [CrossRef] [PubMed]
- Yang, I.V.; Schwartz, DA. Epigenetic mechanisms and the development of asthma. J Allergy Clin Immunol. 2012, 130, 1243–1255. [Google Scholar] [CrossRef] [PubMed]
- Breton, C.V.; Byun, H.M.; Wenten, M.; Pan, F.; Yang, A.; Gilliland, FD. Prenatal tobacco smoke exposure affects global and gene-specific DNA methylation. Am J Respir Crit Care Med. 2009, 180, 462–467. [Google Scholar] [CrossRef]
- Mahon, G.M.; Koppelman, G.H.; Vonk, JM. Grandmaternal smoking, asthma and lung function in the offspring: the Lifelines cohort study. Thorax. 2021, 76, 441–447. [Google Scholar] [CrossRef]
- Bannister, A.J.; Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef]
- Sundar, I.K.; Nevid, M.Z.; Friedman, A.E.; Rahman, I. Cigarette smoke induces distinct histone modifications in lung cells: implications for the pathogenesis of COPD and lung cancer. J Proteome Res. 2014, 13, 982–996. [Google Scholar] [CrossRef]
- Nilsson, E.E.; Sadler-Riggleman, I.; Skinner, MK. Environmentally induced epigenetic transgenerational inheritance of disease. Environ Epigenet. 2018, 4, dvy016. [Google Scholar] [CrossRef]
- Russ, R.; Slack, FJ. Cigarette-Smoke-Induced Dysregulation of MicroRNA Expression and Its Role in Lung Carcinogenesis. Pulm Med. 2012, 2012, 791234. [Google Scholar] [CrossRef]
- Graff, J.W.; Powers, L.S.; Dickson, A.M.; Kim, J.; Reisetter, A.C.; Hassan, I.H.; et al. Cigarette smoking decreases global microRNA expression in human alveolar macrophages. PLoS One. 2012, 7, e44066. [Google Scholar] [CrossRef]
- Tando, Y.; Matsui, Y. Inheritance of environment-induced phenotypic changes through epigenetic mechanisms. Environ Epigenet. 2023, 9, dvad008. [Google Scholar] [CrossRef] [PubMed]
- Di, H.K.; Gan, Y.; Lu, K.; Wang, C.; Zhu, Y.; Meng, X.; et al. Maternal smoking status during pregnancy and low birth weight in offspring: systematic review and meta-analysis of 55 cohort studies published from 1986 to 2020. World J Pediatr. 18, 176–185. [CrossRef] [PubMed]
- Küpers, L.K.; Xu, X.; Jankipersadsing, S.A.; Vaez, A.; la Bastide-van Gemert, S.; Scholtens, S.; et al. DNA methylation mediates the effect of maternal smoking during pregnancy on the birthweight of the offspring. Int J Epidemiol. 2015, 44, 1224–1237. [Google Scholar] [CrossRef] [PubMed]
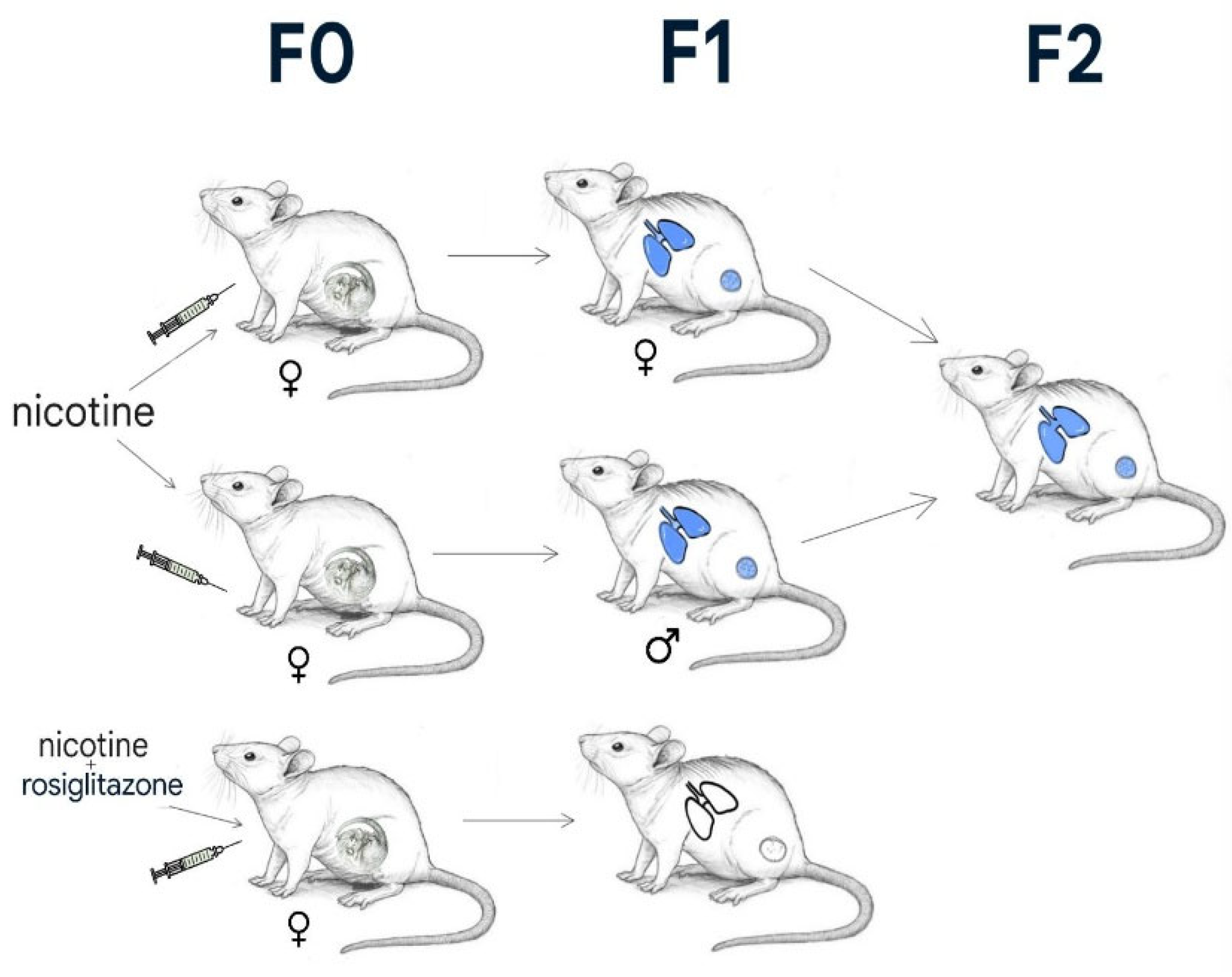
- Rehan, V.K.; Liu, J.; Naeem, E.; Tian, J.; Sakurai, R.; Kwong, K.; et al. Perinatal nicotine exposure induces asthma in second-generation offspring. BMC Med. 2012, 10, 129. [Google Scholar] [CrossRef] [PubMed]
- Leslie, FM. Multigenerational epigenetic effects of nicotine on lung function. BMC Med. 2013, 11, 27. [Google Scholar] [CrossRef]
- Bhadsavle, S.S.; Golding, MC. Paternal epigenetic influences on placental health and their impacts on offspring development and disease. Front Genet. 2022, 13, 1068408. [Google Scholar] [CrossRef] [PubMed]
- Vlachou, M.; Kyrkou, G.; Georgakopoulou, V.E.; Kapetanaki, A.; Vivilaki, V.; Spandidos, D.A.; et al. Smoke signals in the genome: Epigenetic consequences of parental tobacco exposure [Review]. Biomed Rep. 2025, 23, 146. [Google Scholar] [CrossRef]
- Holder, L.B.; Haque, M.M.; Skinner, MK. Machine learning for epigenetics and future medical applications. Epigenetics. 2017, 12, 505–514. [Google Scholar] [CrossRef]
- Liu, J.; Li, J.; Wang, H.; Yan, J. Application of deep learning in genomics. Sci China Life Sci. 2020, 63, 1860–1878. [Google Scholar] [CrossRef]
- Tahir, M.; Norouzi, M.; Khan, S.S.; Davie, J.R.; Yamanaka, S.; Ashraf, A. Artificial intelligence and deep learning algorithms for epigenetic sequence analysis: A review for epigeneticists and AI experts. Comput Biol Med. 2024, 183, 109302. [Google Scholar] [CrossRef]
- Ritzmann, F.; Brand, M.; Bals, R.; Wegmann, M.; Beisswenger, C. Role of Epigenetics in Chronic Lung Disease. Cells. 2025, 14. [Google Scholar] [CrossRef]
- Bannister, A.J.; Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef]
- Li, D.D.; Chen, T.; Ling, Y.L.; Jiang, Y.; Li, QG. A Methylation Diagnostic Model Based on Random Forests and Neural Networks for Asthma Identification. Comput Math Methods Med. 2022, 2022, 2679050. [Google Scholar] [CrossRef] [PubMed]
- Vinciguerra, M. The Potential for Artificial Intelligence Applied to Epigenetics. Mayo Clinic proceedings Digital health. 2023, 1, 476–479. [Google Scholar] [CrossRef] [PubMed]
- Gomes, B.; Ashley, EA. Artificial Intelligence in Molecular Medicine. N Engl J Med. 2023, 388, 2456–2465. [Google Scholar] [CrossRef]
- Kitaba, N.T.; Knudsen, G.T.M.; Johannessen, A.; Rezwan, F.I.; Malinovschi, A.; Oudin, A.; et al. Fathers’ preconception of smoking and offspring DNA methylation. Clin Epigenetics. 2023, 15, 131. [Google Scholar] [CrossRef]
- Natalie Crowley. AI Detects Disease Clues from DNA Methyation Data. What is Epigenetics. 2023 Oct 1.
- Grezenko, H.; Ekhator, C.; Nwabugwu, N.U.; Ganga, H.; Affaf, M.; Abdelaziz, A.M.; et al. Epigenetics in Neurological and Psychiatric Disorders: A Comprehensive Review of Current Understanding and Future Perspectives. Cureus. 2023, 15, e43960. [Google Scholar] [CrossRef]
- Kuehner, J.N.; Bruggeman, E.C.; Wen, Z.; Yao, B. Epigenetic Regulations in Neuropsychiatric Disorders. Front Genet. 2019, 10, 268. [Google Scholar] [CrossRef]
- Kandel, D.B.; Kandel, ER. A molecular basis for nicotine as a gateway drug. N Engl J Med. 2014, 371, 2038–2039. [Google Scholar] [CrossRef]
- Dao, J.M.; McQuown, S.C.; Loughlin, S.E.; Belluzzi, J.D.; Leslie, FM. Nicotine alters limbic function in adolescent rat by a 5-HT1A receptor mechanism. Neuropsychopharmacology. 2011, 36, 1319–1331. [Google Scholar] [CrossRef]
- Rauschert, S.; Melton, P.E.; Heiskala, A.; Karhunen, V.; Burdge, G.; Craig, J.M.; et al. Machine Learning-Based DNA Methylation Score for Fetal Exposure to Maternal Smoking: Development and Validation in Samples Collected from Adolescents and Adults. Environ Health Perspect. 2020, 128, 97003. [Google Scholar] [CrossRef] [PubMed]
- Zakarya, R.; Adcock, I.; Oliver, BG. Epigenetic impacts of maternal tobacco and e-vapour exposure on the offspring lung. Clin Epigenetics. 2019, 11, 32. [Google Scholar] [CrossRef]
- Fernando, A.; Kondrup, E.; Cheung, K.; Uberoi, D.; Joly, Y. Still using genetic data? A comparative review of Canadian life insurance application forms before and after the GNDA. FACETS. 2024, 9, 1–10. [Google Scholar]
- Esfahani, M.S.; Hamilton, E.G.; Mehrmohamadi, M.; Nabet, B.Y.; Alig, S.K.; King, D.A.; et al. Inferring gene expression from cell-free DNA fragmentation profiles. Nat Biotechnol. 2022, 40, 585–597. [Google Scholar] [CrossRef] [PubMed]
- Leroy, A.; Teh, A.L.; Dondelinger, F.; Alvarez, M.A.; Wang, D. Longitudinal prediction of DNA methylation to forecast epigenetic outcomes. EBioMedicine. 2025, 115, 105709. [Google Scholar]
- Oliva, A.; Kaphle, A.; Reguant, R.; Sng, L.M.F.; Twine, N.A.; Malakar, Y.; et al. Future-proofing genomic data and consent management: a comprehensive review of technology innovations. Gigascience. 2024, 2, 13. [Google Scholar] [CrossRef]
- Teare, H.J.A.; Prictor, M.; Kaye, J. Reflections on dynamic consent in biomedical research: the story so far. Eur J Hum Genet. 2021, 29, 649–656. [Google Scholar] [CrossRef]
- Brauneck, A.; Schmalhorst, L.; Weiss, S.; Baumbach, L.; Völker, U.; Ellinghaus, D.; et al. Legal aspects of privacy-enhancing technologies in genome-wide association studies and their impact on performance and feasibility. Genome Biol. 2024, 25, 154. [Google Scholar]
- The Genetic Information Nondiscrimination Act [GINA]. American Society of Human Genetics.
- Marvin van Bekkum, Frederik Zuiderveen Borgesius, Tom Heskes. AI, insurance, discrimination and unfair differentiation: an overview and research agenda. Taylor&Francis. 2025, 177–204.
- Gorzynski, J.E.; Goenka, S.D.; Shafin, K.; Jensen, T.D.; Fisk, D.G.; Grove, M.E.; et al. Ultrarapid Nanopore Genome Sequencing in a Critical Care Setting. N Engl J Med. 2022, 386, 700–702. [Google Scholar] [CrossRef] [PubMed]
- Alsaedi, S.; Ogasawara, M.; Alarawi, M.; Gao, X.; Gojobori, T. AI-powered precision medicine: utilizing genetic risk factor optimization to revolutionize healthcare. NAR Genom Bioinform. 2025, 7, lqaf038. [Google Scholar] [CrossRef]
- Zhang, S.; Jin, J.; Xu, B.; Zheng, Q.; Mou, H. The relationship between epigenetic biomarkers and the risk of diabetes and cancer: a machine learning modelling approach. Front Public Health. 2025, 13, 1509458. [Google Scholar] [CrossRef]
- Rauschert, S.; Raubenheimer, K.; Melton, P.E.; Huang, RC. Machine learning and clinical epigenetics: a review of challenges for diagnosis and classification. Clin Epigenetics. 2020, 12, 51. [Google Scholar] [CrossRef]
- Burton, N.O.; Greer, EL. Multigenerational epigenetic inheritance: Transmitting information across generations. Semin Cell Dev Biol. 2022, 127, 121–132. [Google Scholar] [CrossRef]
- Arefeen, M.A.; Nimi, S.T.; Rahman, M.S.; Arshad, S.H.; Holloway, J.W.; Rezwan, FI. Prediction of Lung Function in Adolescence Using Epigenetic Aging: A Machine Learning Approach. Methods Protoc. 2020, 3. [Google Scholar] [CrossRef]
- Krauss-Etschmann, S.; Meyer, K.F.; Dehmel, S.; Hylkema, MN. Inter- and transgenerational epigenetic inheritance: evidence in asthma and COPD? Clin Epigenetics. 2015, 7, 53. [Google Scholar] [CrossRef]
- Jacobsen, K.K.; Schnohr, P.; Jensen, G.B.; Bojesen, SE. AHRR [cg05575921] Methylation Safely Improves Specificity of Lung Cancer Screening Eligibility Criteria: A Cohort Study. Cancer Epidemiol Biomarkers Prev. 2022, 31, 758–765. [Google Scholar] [CrossRef] [PubMed]
- Tsai, P.C.; Glastonbury, C.A.; Eliot, M.N.; Bollepalli, S.; Yet, I.; Castillo-Fernandez, J.E.; et al. Smoking induces coordinated DNA methylation and gene expression changes in adipose tissue with consequences for metabolic health. Clin Epigenetics. 2018, 10, 126. [Google Scholar] [CrossRef] [PubMed]
- Stueve, T.R.; Li, W.Q.; Shi, J.; Marconett, C.N.; Zhang, T.; Yang, C.; et al. Epigenome-wide analysis of DNA methylation in lung tissue shows concordance with blood studies and identifies tobacco smoke-inducible enhancers. Hum Mol Genet. 2017, 26, 3014–3027. [Google Scholar] [CrossRef] [PubMed]
- Li, J.L.; Jain, N.; Tamayo, L.I.; Tong, L.; Jasmine, F.; Kibriya, M.G.; et al. The association of cigarette smoking with DNA methylation and gene expression in human tissue samples. Am J Hum Genet. 2024, 111, 636–653. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Nwozor, K.O.; van den Berge, M.; Slebos, D.J.; Faiz, A.; Jonker, M.R.; et al. From Differential DNA Methylation in COPD to Mitochondria: Regulation of AHRR Expression Affects Airway Epithelial Response to Cigarette Smoke. Cells. 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.Q.; Richmond, R.C.; Suderman, M.; Min, J.L.; Battram, T.; Flatberg, A.; et al. Assessing the role of genome-wide DNA methylation between smoking and risk of lung cancer using repeated measurements: the HUNT study. Int J Epidemiol. 2021, 50, 1482–1497. [Google Scholar] [CrossRef]
- Lu, A.T.; Quach, A.; Wilson, J.G.; Reiner, A.P.; Aviv, A.; Raj, K.; et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging. 2019, 11, 303–327. [Google Scholar] [CrossRef]
- Lu, A.T.; Binder, A.M.; Zhang, J.; Yan, Q.; Reiner, A.P.; Cox, S.R.; et al. DNA methylation GrimAge version 2. Aging. 2022, 14, 9484–9549. [Google Scholar] [CrossRef]
- Zhang, Q. An interpretable biological age. Lancet Healthy Longev. 2023, 4, e662–3. [Google Scholar] [CrossRef]
- Qiu, W.; Chen, H.; Kaeberlein, M.; Lee, SI. ExplaiNAble BioLogical Age [ENABL Age]: an artificial intelligence framework for interpretable biological age. Lancet Healthy Longev. 2023, 4, e711–e723. [Google Scholar] [CrossRef]
- Song, M.A.; Mori, K.M.; McElroy, J.P.; Freudenheim, J.L.; Weng, D.Y.; Reisinger, S.A.; et al. Accelerated epigenetic age, inflammation, and gene expression in lung: comparisons of smokers and vapers with non-smokers. Clin Epigenetics. 2023, 15, 160. [Google Scholar] [CrossRef]
- Klose, D.; Needhamsen, M.; Ringh, M.V.; Hagemann-Jensen, M.; Jagodic, M.; Kular, L. Smoking affects epigenetic ageing of lung bronchoalveolar lavage cells in Multiple Sclerosis. Mult Scler Relat Disord. 2023, 79, 104991. [Google Scholar] [CrossRef]
- Human Genome Project: The most important biomedical research undertaking of the 20th Century. National Human Genome Research Institute. 2024 Jun 13.
- Hood, L.; Rowen, L. The Human Genome Project: big science transforms biology and medicine. Genome Med. 2013, 5, 79. [Google Scholar] [CrossRef]
- Kabata, F.; Thaldar, D. The human genome as the common heritage of humanity. Front Genet. 2023, 14, 1282515. [Google Scholar] [CrossRef] [PubMed]
- Bradbury, J. Human epigenome project--up and running. PLoS Biol. 2003, 1, E82. [Google Scholar] [CrossRef] [PubMed]
- Eckhardt, F.; Beck, S.; Gut, I.G.; Berlin, K. Future potential of the Human Epigenome Project. Expert Rev Mol Diagn. 2004, 4, 609–618. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



