Submitted:
21 September 2025
Posted:
22 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
| Butyrophilins | γδT-cell subset | Role of butyrophilins | References | |
| Peripheral blood | Mouse unidentified | Unidentified | Unidentified | NA |
| Alpaca BTN3 | Vγ9Vδ2+ T cells | No interaction has been identified | [35,36] | |
| Human BTN3A homodimers/ heterodimers and BTN2A1 homodimer |
Vγ9Vδ2+ T cells | Phosphoantigen mediated, CDR3-independent γδ T-cell activation | [30,33,37,38] | |
| Skin | Mouse Skint1 and Skint2 | Vγ5Vδ1+ DETC | Thymic selection, tissue homing of dendritic epidermal T cells to the skin | [27,28,32] |
| Human ? | Vδ1+ T cells | Unidentified, unknown if there is butyrophilin involvement | [39] | |
| Intestinal epithelium | Mouse Btnl1 and Btnl6 | Vγ7+ IEL | Phenotypic maintenance of the intestinal IEL compartment | [27,28] |
| Human BTNL3 and BTNL8 | Vγ4Vδ1+ IEL | Phenotypic maintenance of the intestinal IEL compartment | [21,27,29] |
| Population | Hom. for deletion | Het. for deletion | Hom. for full sequences | Deletion allele N | Deletion allele frequency | Group N | Carriers N |
Carriers % |
|
| HapMap | CEU | 17 | 56 | 68 | 90 | 0.319 | 141 | 73 | 51.8 |
| Toskani, Italia | 9 | 45 | 34 | 63 | 0.358 | 88 | 54 | 61.4 | |
| HGDP | France | 7 | 28 | 17 | 42 | 0.404 | 52 | 35 | 67.3 |
| Italy | 5 | 18 | 13 | 28 | 0.389 | 36 | 23 | 63.9 | |
| Italy (Bergamo) | 3 | 2 | 9 | 8 | 0.286 | 14 | 5 | 35.7 | |
| Orkney Islands | 1 | 11 | 3 | 13 | 0.433 | 15 | 12 | 80.0 | |
| Total | European ancestry | 42 | 160 | 144 | 244 | 0.353 | 346 | 202 | 58.4 |
2. Results
2.1. BTN2A1 SNPs Were Significantly Associated with CeD Risk in a Study of 94 Samples
2.1.1. Risk Associated HLA Genotypes Were Significantly More Frequent in CeD Patients
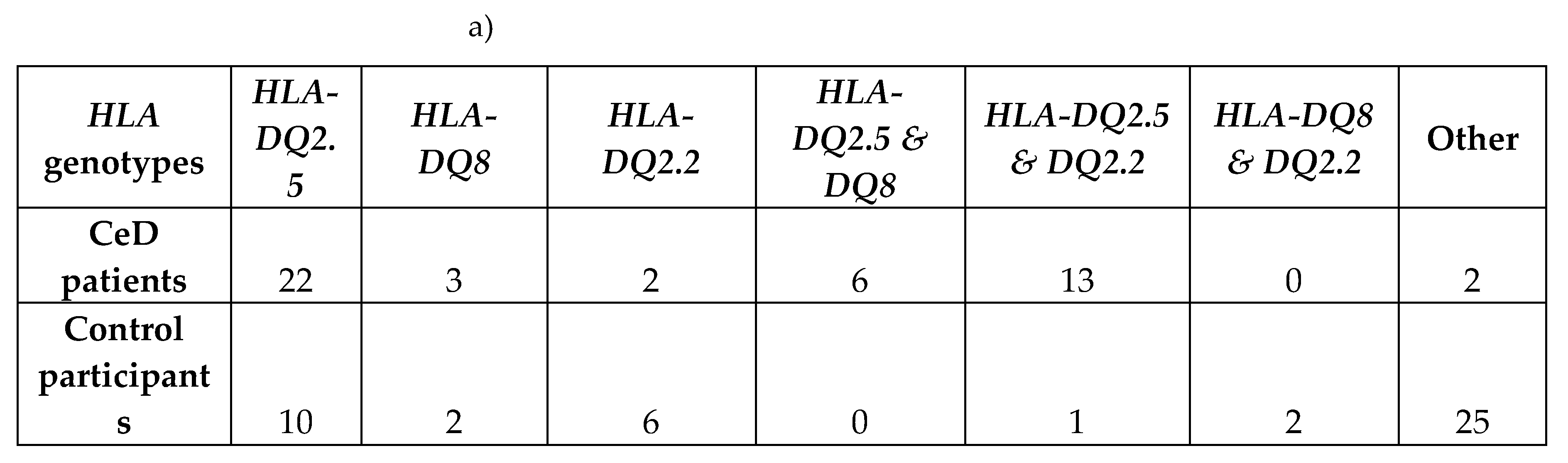 |
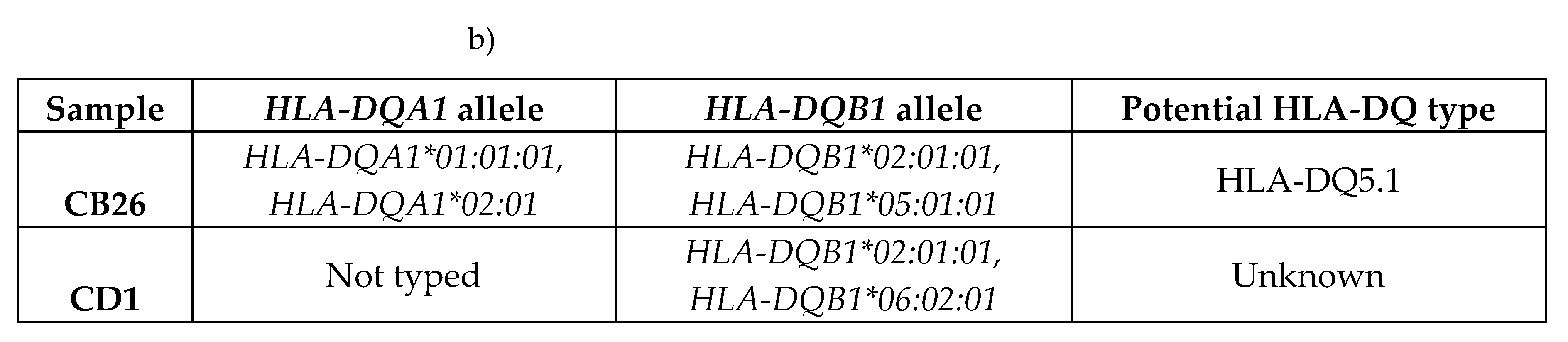 |
2.1.2. The BTNL8*BTNL3 Copy Number Variant Was not Associated with CeD
| rs72494581 genotype | TT | CT | CC |
| BTNL8-BTNL3genes | Homozygous for full length sequence | Heterozygous for BTNL8*BTNL3 deletion | Homozygous for BTNL8*BTNL3 deletion |
| Coeliac disease patients | 20 | 26 | 2 |
| Control participants | 24 | 17 | 5 |
2.1.3. BTN2A1 Gene Burden was Significantly Higher in CeD Patients
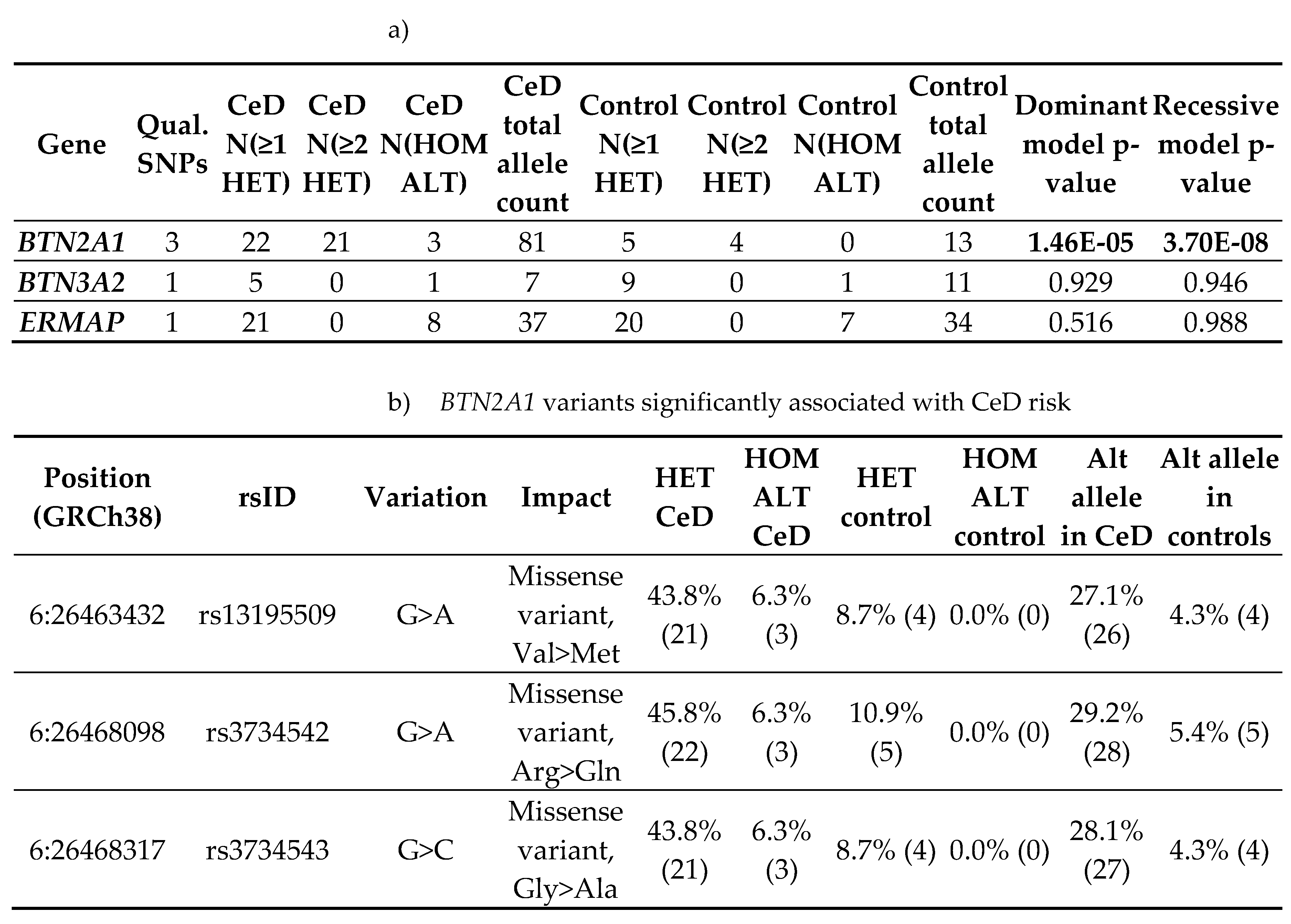 |
2.2. BTN3A1, BTN3A2, and BTN2A1 Genes Were significantly Associated with CeD in HLA-DQ2.5-Matched Participants of the UK Biobank Database
2.2.1. Risk Associated HLA Genotypes Were Significantly More frequent In CeD Patients of the UK Biobank
| HLA genotypes | HLA-DQ2.5 | HLA-DQ8 | HLA-DQ2.2 | HLA-DQ2.5 & DQ8 | HLA-DQ2.5 & DQ2.2 | HLA-DQ8 & DQ2.2 | Other |
| CeD participants | 1652 | 171 | 199 | 606 | 182 | 50 | 234 |
| Control participants | 6416 | 4203 | 4154 | 895 | 886 | 590 | 12618 |
2.2.2. BTN2A1, BTN3A1, and BTN3A2 SNPs Were Significantly Associated with CeD Status in the UK Biobank
| Gene | SNPs in NCBI | Unique SNPs in NCBI | SNPs in UK Biobank |
| BTN2A1 | 7912 | 7605 | 30 |
| BTN3A1 | 5348 | 5164 | 27 |
| BTN3A2 | 5905 | 5611 | 21 |
| BTNL3 | 6164 | 5929 | 10 |
| BTNL8 | 18 889 | 18 197 | 13 |
| Position (GRCh38) | SNP, reference allele | Gene | SNP consequence | CeD allele freq | Control allele freq | Total allele freq | ln(OR) | CeD risk | adjusted p-value |
| 6:26463347 | rs13195402 | BTN2A1 | STOP gained | 0.768 | 0.892 | 0.880 | -0.924 | decrease | 4.67E-158 |
| 6:26463432 | rs13195509 | BTN2A1 | missense | 0.754 | 0.879 | 0.867 | -0.857 | decrease | 1.61E-151 |
| 6:26475927 | rs1407045 | BTN2A1 | intronic | 0.584 | 0.516 | 0.522 | 0.273 | increase | 6.07E-22 |
| 6:26465807 | rs2273558 | BTN2A1 | intronic | 0.583 | 0.677 | 0.667 | -0.396 | decrease | 1.69E-41 |
| 6:26460493 | rs2893856 | BTN2A1 | intronic | 0.113 | 0.131 | 0.130 | -0.175 | decrease | 3.23E-03 |
| 6:26468098 | rs3734542 | BTN2A1 | missense | 0.753 | 0.878 | 0.867 | -0.855 | decrease | 8.59E-151 |
| 6:26468317 | rs3734543 | BTN2A1 | missense | 0.760 | 0.879 | 0.868 | -0.844 | decrease | 1.59E-140 |
| 6:26466954 | rs3799380 | BTN2A1 | intronic | 0.683 | 0.790 | 0.780 | -0.549 | decrease | 8.59E-77 |
| 6:26474343 | rs56296968 | BTN2A1 | intronic | 0.696 | 0.807 | 0.796 | -0.604 | decrease | 9.70E-89 |
| 6:26456215 | rs6456724 | BTN2A1 | 2kb upstream | 0.113 | 0.131 | 0.130 | -0.176 | decrease | 2.87E-03 |
| 6:26458037 | rs6929846 | BTN2A1 | 5’ UTR | 0.146 | 0.174 | 0.172 | -0.206 | decrease | 3.60E-06 |
| 6:26473816 | rs7773938 | BTN2A1 | intronic | 0.696 | 0.806 | 0.796 | -0.600 | decrease | 1.15E-87 |
| 6:26469647 | rs9358944 | BTN2A1 | intronic | 0.695 | 0.806 | 0.796 | -0.604 | decrease | 1.55E-89 |
| 6:26471886 | rs9358945 | BTN2A1 | intronic | 0.694 | 0.806 | 0.796 | -0.606 | decrease | 4.37E-90 |
| 6:26404730 | rs10456045 | BTN3A1 | intronic | 0.596 | 0.698 | 0.688 | -0.448 | decrease | 2.97E-57 |
| 6:26410572 | rs1796520 | BTN3A1 | intronic | 0.405 | 0.474 | 0.467 | -0.276 | decrease | 2.40E-22 |
| 6:26404146 | rs3799378 | BTN3A1 | intronic | 0.653 | 0.762 | 0.752 | -0.535 | decrease | 2.92E-75 |
| 6:26405825 | rs3857549 | BTN3A1 | intronic | 0.948 | 0.935 | 0.936 | 0.221 | increase | 1.53E-02 |
| 6:26409662 | rs41266839 | BTN3A1 | missense | 0.764 | 0.892 | 0.880 | -0.924 | decrease | 2.12E-168 |
| 6:26407180 | rs4609015 | BTN3A1 | intronic | 0.871 | 0.854 | 0.855 | 0.141 | increase | 3.82E-02 |
| 6:26412860 | rs6900725 | BTN3A1 | intronic | 0.870 | 0.853 | 0.855 | 0.139 | increase | 4.33E-02 |
| 6:26401210 | rs6912853 | BTN3A1 | 2kb upstream | 0.863 | 0.844 | 0.846 | 0.153 | increase | 7.85E-03 |
| 6:26413007 | rs6920986 | BTN3A1 | intronic | 0.870 | 0.854 | 0.856 | 0.138 | increase | 4.99E-02 |
| 6:26415409 | rs742090 | BTN3A1 | 500b downstream | 0.406 | 0.474 | 0.468 | -0.276 | decrease | 3.58E-22 |
| 6:26374321 | rs11758089 | BTN3A2 | intronic | 0.866 | 0.844 | 0.846 | 0.176 | increase | 6.30E-04 |
| 6:26372558 | rs12176317 | BTN3A2 | intronic | 0.744 | 0.867 | 0.856 | -0.809 | decrease | 6.72E-140 |
| 6:26366990 | rs12199613 | BTN3A2 | intronic | 0.514 | 0.612 | 0.602 | -0.400 | decrease | 1.76E-47 |
| 6:26377318 | rs1977 | BTN3A2 | 3’ UTR | 0.740 | 0.864 | 0.853 | -0.808 | decrease | 1.17E-136 |
| 6:26377363 | rs1979 | BTN3A2 | 3’ UTR | 0.743 | 0.867 | 0.855 | -0.809 | decrease | 8.23E-140 |
| 6:26375933 | rs1985732 | BTN3A2 | intronic | 0.595 | 0.698 | 0.688 | -0.457 | decrease | 2.87E-59 |
| 6:26374430 | rs2073526 | BTN3A2 | intronic | 0.370 | 0.442 | 0.435 | -0.295 | decrease | 9.15E-25 |
| 6:26363527 | rs9358934 | BTN3A2 | 2kb upstream | 0.744 | 0.866 | 0.855 | -0.803 | decrease | 2.34E-137 |
| 6:26364702 | rs9379855 | BTN3A2 | 2kb upstream | 0.743 | 0.866 | 0.855 | -0.804 | decrease | 8.85E-138 |
| 6:26367461 | rs9379858 | BTN3A2 | intronic | 0.743 | 0.866 | 0.855 | -0.802 | decrease | 3.19E-137 |
| 6:26369321 | rs9379859 | BTN3A2 | intronic | 0.744 | 0.867 | 0.855 | -0.803 | decrease | 5.37E-137 |
| 6:26373450 | rs9393713 | BTN3A2 | intronic | 0.743 | 0.868 | 0.856 | -0.814 | decrease | 1.07E-141 |
| 6:26373512 | rs9393714 | BTN3A2 | intronic | 0.743 | 0.868 | 0.856 | -0.813 | decrease | 6.99E-141 |
2.2.3. Twenty Butyrophilin SNPs from the UK Biobank Remained Significantly Associated with CeD Status When the Participants’ risk HLA Genotypes Were Taken into Account
| SNP, reference allele | Gene | SNP consequence | ln(OR) | CeD risk | adjusted p-value |
| rs13195402 | BTN2A1 | STOP gained | -0.20727 | decrease | 8.15E-06 |
| rs13195509 | BTN2A1 | missense | -0.19239 | decrease | 1.62E-05 |
| rs3734542 | BTN2A1 | missense | -0.18831 | decrease | 2.94E-05 |
| rs3734543 | BTN2A1 | missense | -0.16744 | decrease | 8.23E-04 |
| rs56296968 | BTN2A1 | intronic | -0.11753 | decrease | 4.20E-02 |
| rs9358944 | BTN2A1 | intronic | -0.11786 | decrease | 3.83E-02 |
| rs9358945 | BTN2A1 | intronic | -0.12018 | decrease | 2.91E-02 |
| rs3799378 | BTN3A1 | intronic | -0.14327 | decrease | 7.04E-04 |
| rs41266839 | BTN3A1 | missense | -0.21469 | decrease | 1.06E-06 |
| rs12176317 | BTN3A2 | intronic | -0.1974 | decrease | 3.50E-06 |
| rs12199613 | BTN3A2 | intronic | -0.12296 | decrease | 3.31E-03 |
| rs1977 | BTN3A2 | 3’ UTR | -0.20238 | decrease | 2.06E-06 |
| rs1979 | BTN3A2 | 3’ UTR | -0.19756 | decrease | 3.40E-06 |
| rs1985732 | BTN3A2 | intronic | -0.10975 | decrease | 3.35E-02 |
| rs9358934 | BTN3A2 | 2kb upstream | -0.19286 | decrease | 7.53E-06 |
| rs9379855 | BTN3A2 | 2kb upstream | -0.19406 | decrease | 6.04E-06 |
| rs9379858 | BTN3A2 | intronic | -0.19156 | decrease | 8.99E-06 |
| rs9379859 | BTN3A2 | intronic | -0.19261 | decrease | 8.10E-06 |
| rs9393713 | BTN3A2 | intronic | -0.2056 | decrease | 9.27E-07 |
| rs9393714 | BTN3A2 | intronic | -0.20087 | decrease | 2.08E-06 |
2.2.4. Twenty-One Butyrophilin SNPs Were Significantly Associated with CeD Status in HLA-DQ2.5-Matched Case-Control Groups of UK Biobank Participants
| HLA genotype of individuals in model | Number of CeD participants | Number of controls | Number of significant SNPs |
| HLA-DQ2.2 | 199 | 4154 | 0 |
| HLA-DQ2.5 | 1652 | 6416 | 21 |
| HLA-DQ8 | 171 | 4203 | 0 |
| HLA-DQ2.2, HLA-DQ2.5 | 606 | 895 | 0 |
| HLA-DQ2.2, HLA-DQ8 | 50 | 590 | 0 |
| HLA-DQ2.5, HLA-DQ8 | 182 | 886 | 0 |
| Other | 234 | 12618 | 0 |
| SNP, reference allele | Gene | SNP consequence | CeD allele freq | Control allele freq | Total allele freq | ln(OR) | CeD risk | adjusted p-value |
| rs13195402 | BTN2A1 | STOP gained | 0.704 | 0.751 | 0.741 | -0.27812 | decrease | 5.10E-07 |
| rs13195509 | BTN2A1 | missense | 0.687 | 0.734 | 0.724 | -0.25542 | decrease | 1.75E-06 |
| rs3734542 | BTN2A1 | missense | 0.687 | 0.733 | 0.723 | -0.25235 | decrease | 2.52E-06 |
| rs3734543 | BTN2A1 | missense | 0.695 | 0.736 | 0.728 | -0.23279 | decrease | 5.43E-05 |
| rs56296968 | BTN2A1 | intronic | 0.640 | 0.673 | 0.666 | -0.15928 | decrease | 2.33E-02 |
| rs7773938 | BTN2A1 | intronic | 0.640 | 0.672 | 0.666 | -0.15755 | decrease | 2.66E-02 |
| rs9358944 | BTN2A1 | intronic | 0.638 | 0.671 | 0.664 | -0.16238 | decrease | 1.61E-02 |
| rs9358945 | BTN2A1 | intronic | 0.638 | 0.671 | 0.665 | -0.16499 | decrease | 1.26E-02 |
| rs3799378 | BTN3A1 | intronic | 0.596 | 0.637 | 0.628 | -0.18712 | decrease | 8.10E-04 |
| rs41266839 | BTN3A1 | missense | 0.697 | 0.747 | 0.737 | -0.28267 | decrease | 7.25E-08 |
| rs12176317 | BTN3A2 | intronic | 0.679 | 0.729 | 0.719 | -0.26575 | decrease | 2.82E-07 |
| rs12199613 | BTN3A2 | intronic | 0.459 | 0.500 | 0.491 | -0.1717 | decrease | 2.06E-03 |
| rs1977 | BTN3A2 | 3’ UTR | 0.676 | 0.726 | 0.716 | -0.268 | decrease | 2.99E-07 |
| rs1979 | BTN3A2 | 3’ UTR | 0.679 | 0.728 | 0.718 | -0.26376 | decrease | 3.63E-07 |
| rs1985732 | BTN3A2 | intronic | 0.539 | 0.574 | 0.567 | -0.15063 | decrease | 2.35E-02 |
| rs9358934 | BTN3A2 | 2kb upstream | 0.680 | 0.729 | 0.719 | -0.25825 | decrease | 8.85E-07 |
| rs9379855 | BTN3A2 | 2kb upstream | 0.680 | 0.728 | 0.718 | -0.25967 | decrease | 6.76E-07 |
| rs9379858 | BTN3A2 | intronic | 0.680 | 0.728 | 0.718 | -0.25566 | decrease | 1.18E-06 |
| rs9379859 | BTN3A2 | intronic | 0.681 | 0.729 | 0.719 | -0.26105 | decrease | 6.31E-07 |
| rs9393713 | BTN3A2 | intronic | 0.678 | 0.729 | 0.719 | -0.27313 | decrease | 1.06E-07 |
| rs9393714 | BTN3A2 | intronic | 0.679 | 0.729 | 0.719 | -0.26778 | decrease | 2.25E-07 |
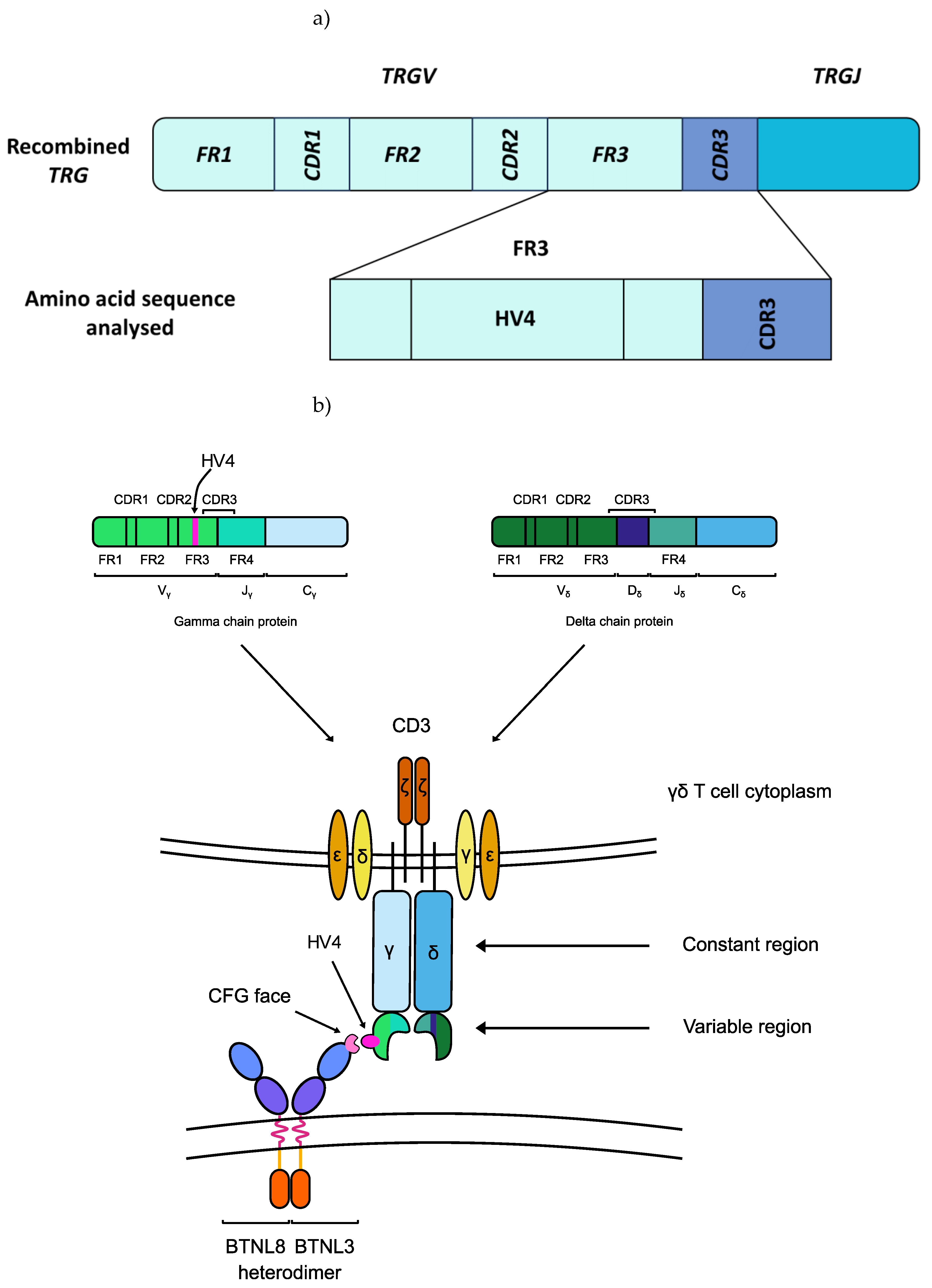
2.3. HV4 Variation was Not Significantly Associated with CeD Risk in a Study of 379 Samples
2.3.1. TRGV Usage Was not Significantly Different Between CeD and Control Samples
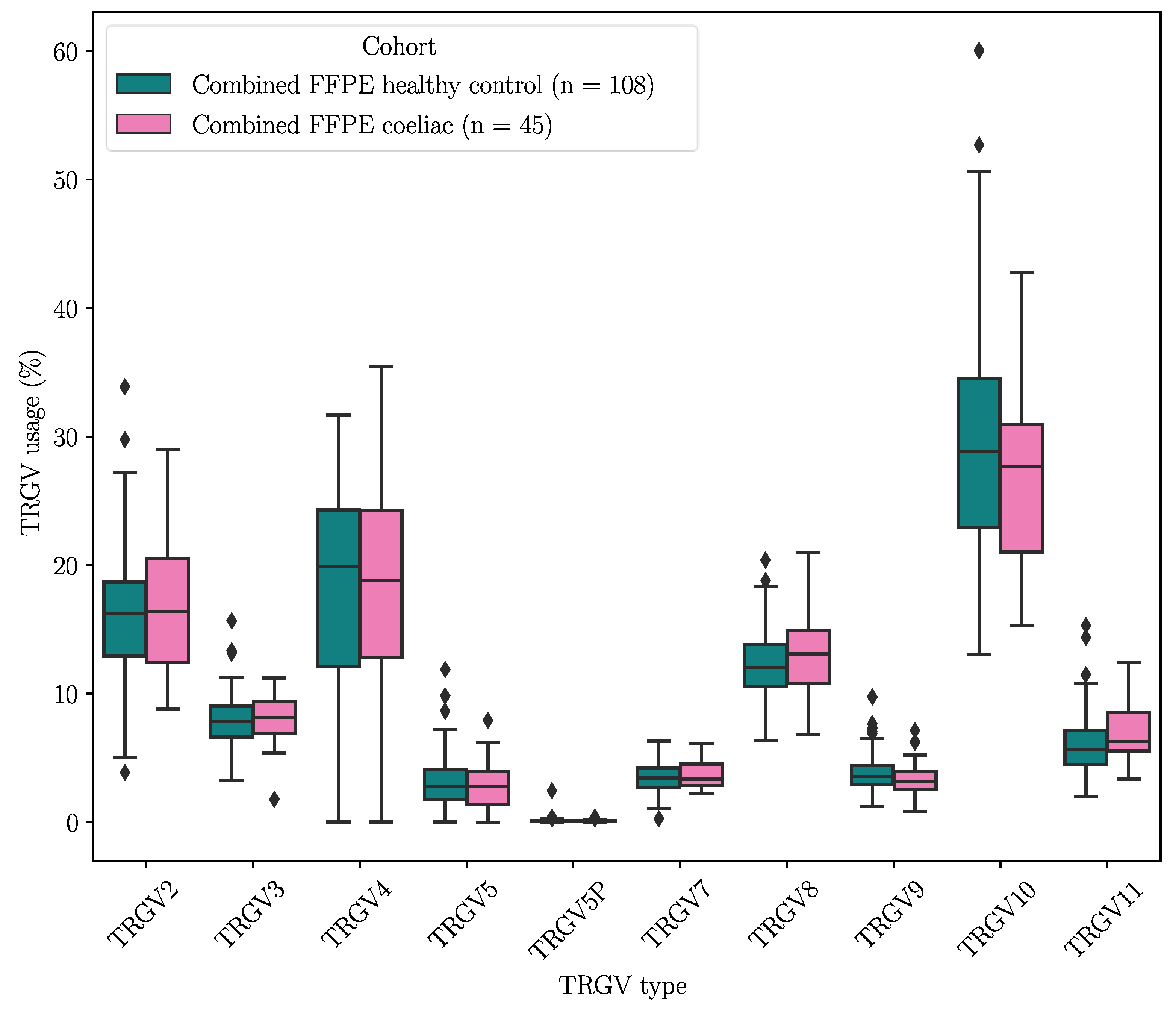
2.3.2. HV4 Sequence Variation Was not Significantly Associated with CeD Risk
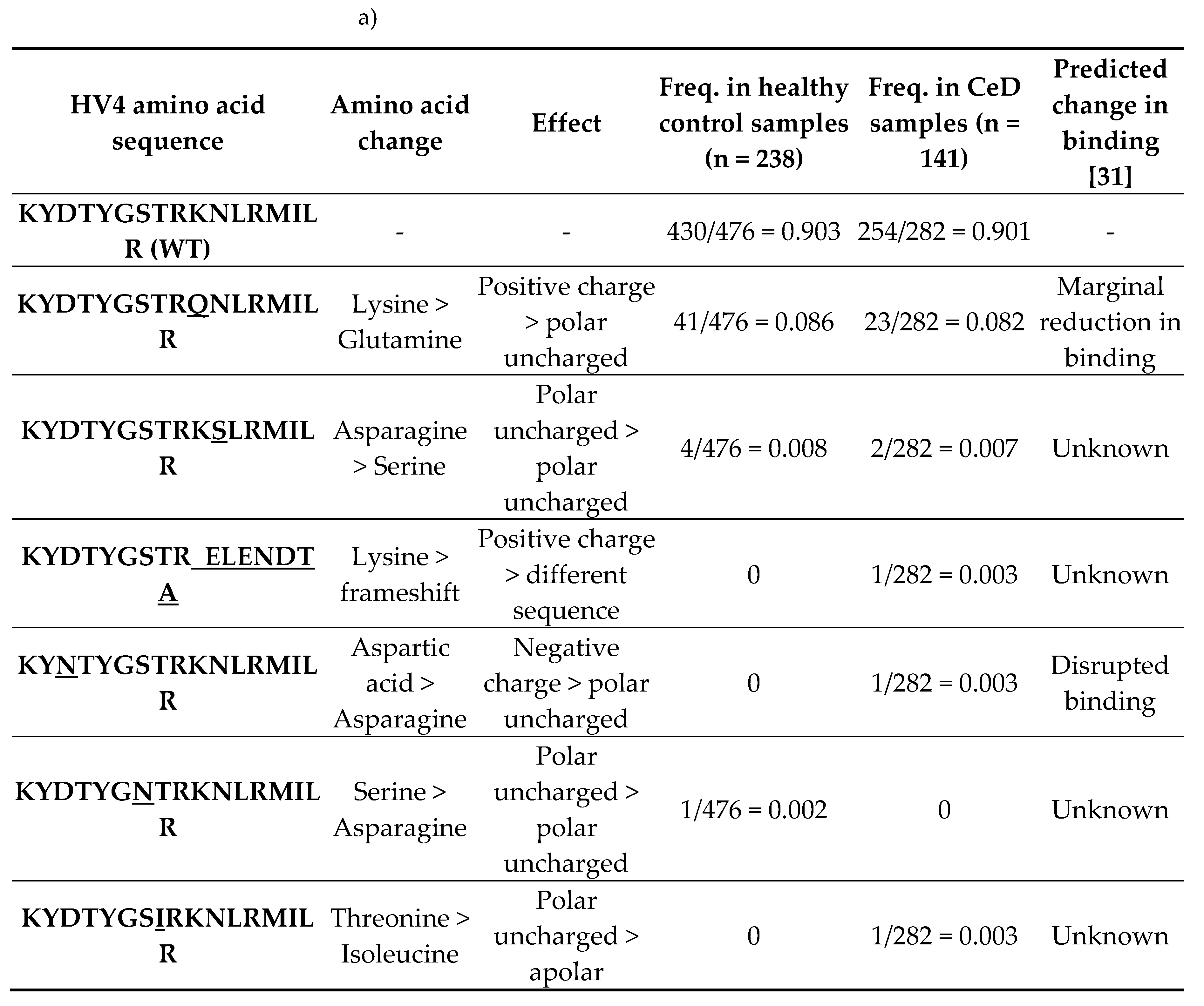 |
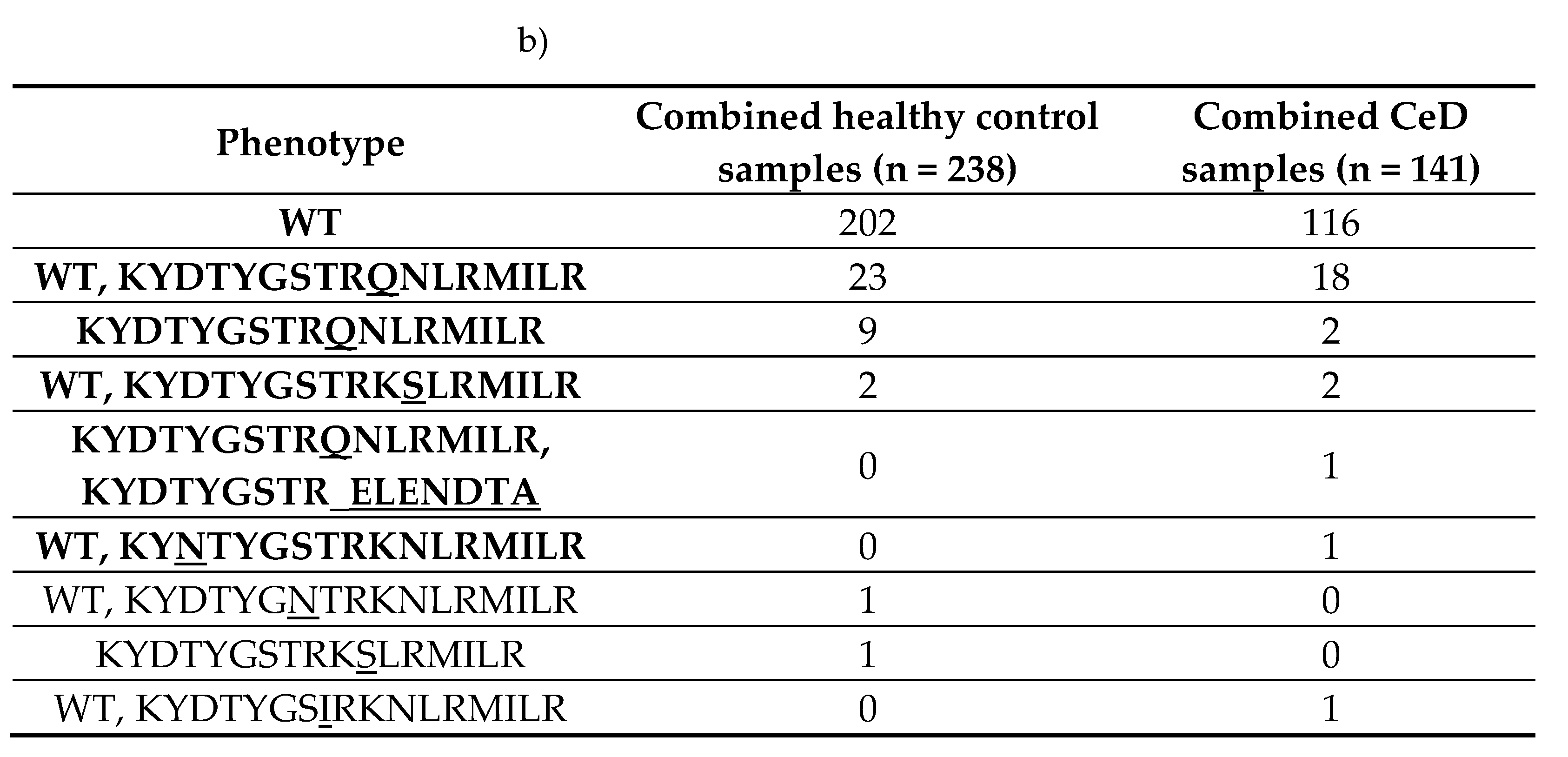 |
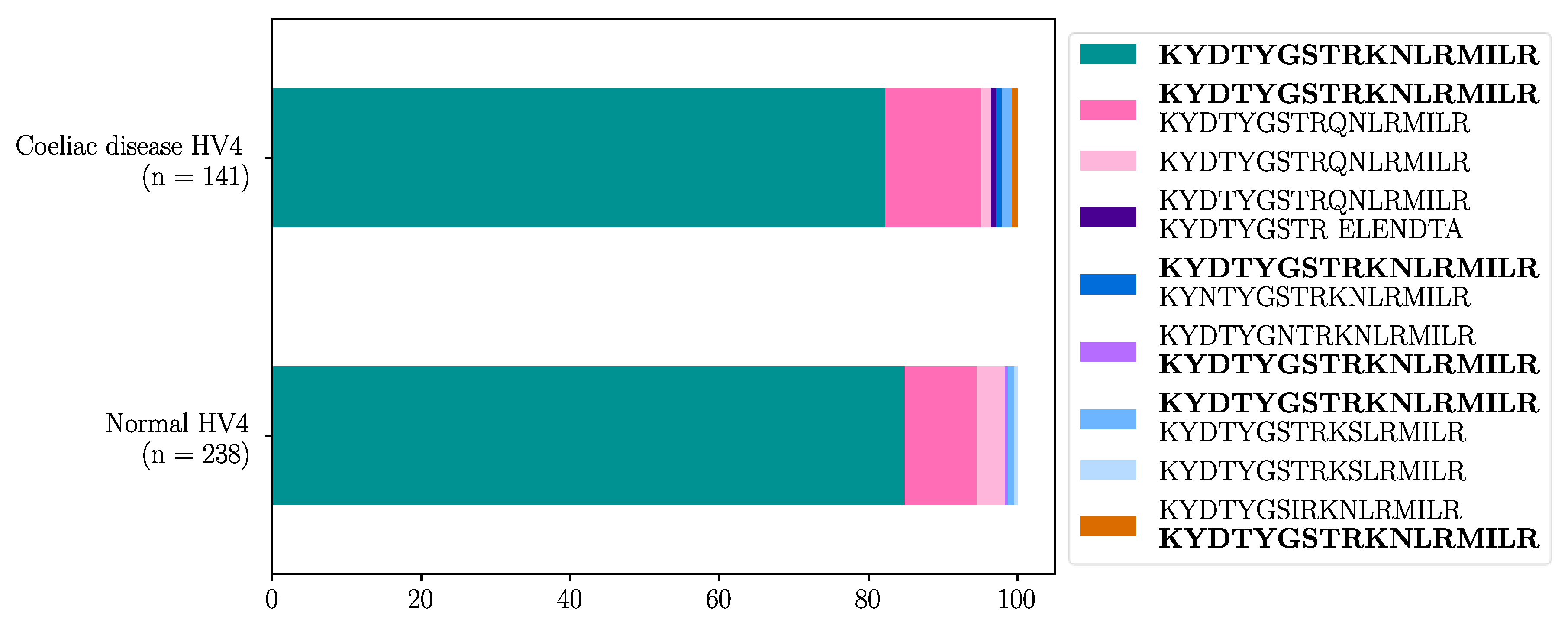
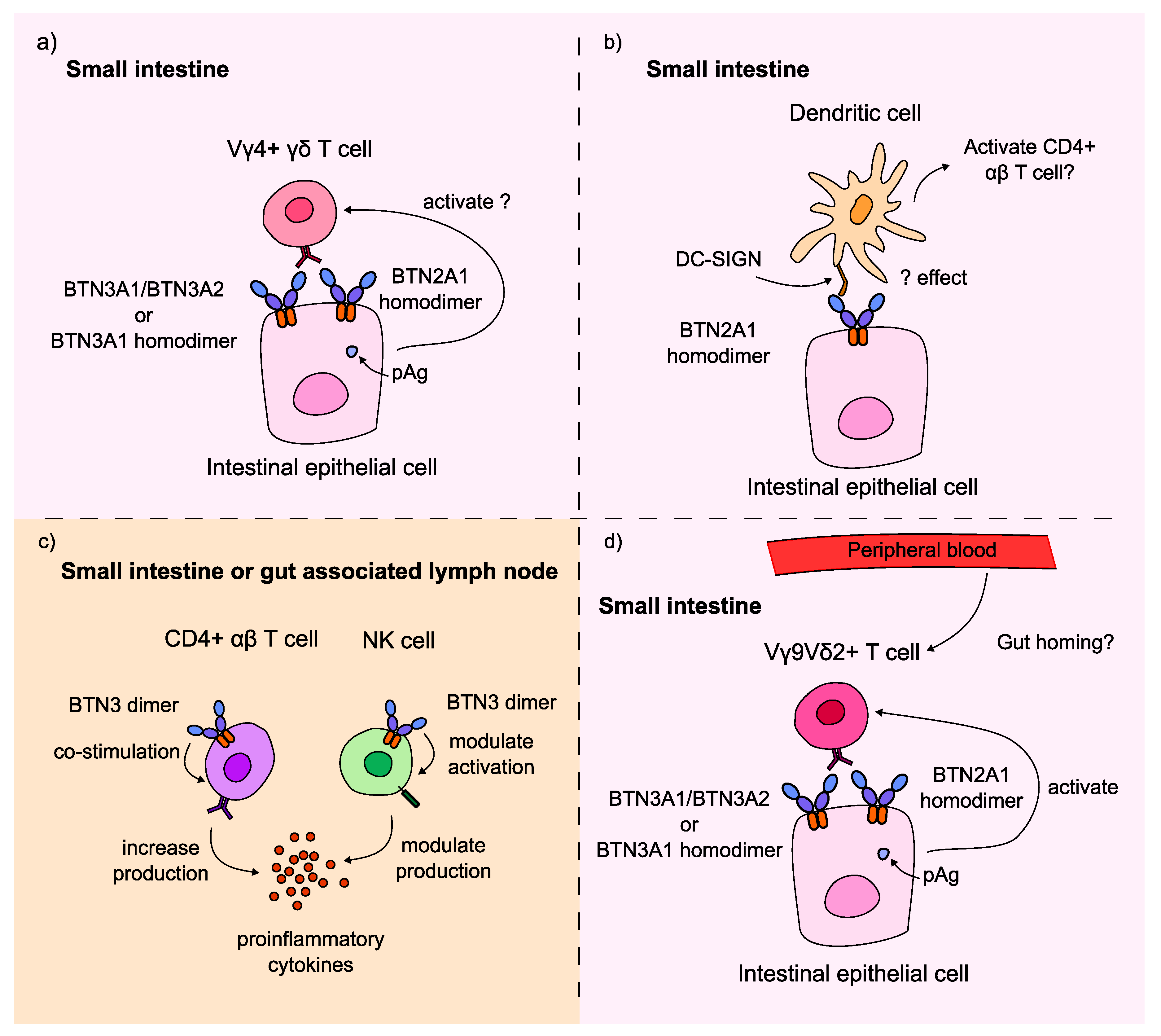
3. Discussion
4. Materials and Methods
4.1. Participant Selection Criteria
- Has CeD diagnosis
- Malabsorption
- Anaemia
- Lymphocytosis
- On a GFD
- Diarrhoea
4.1.1. Participant Selection for the Butyrophilin Family Gene Sequencing
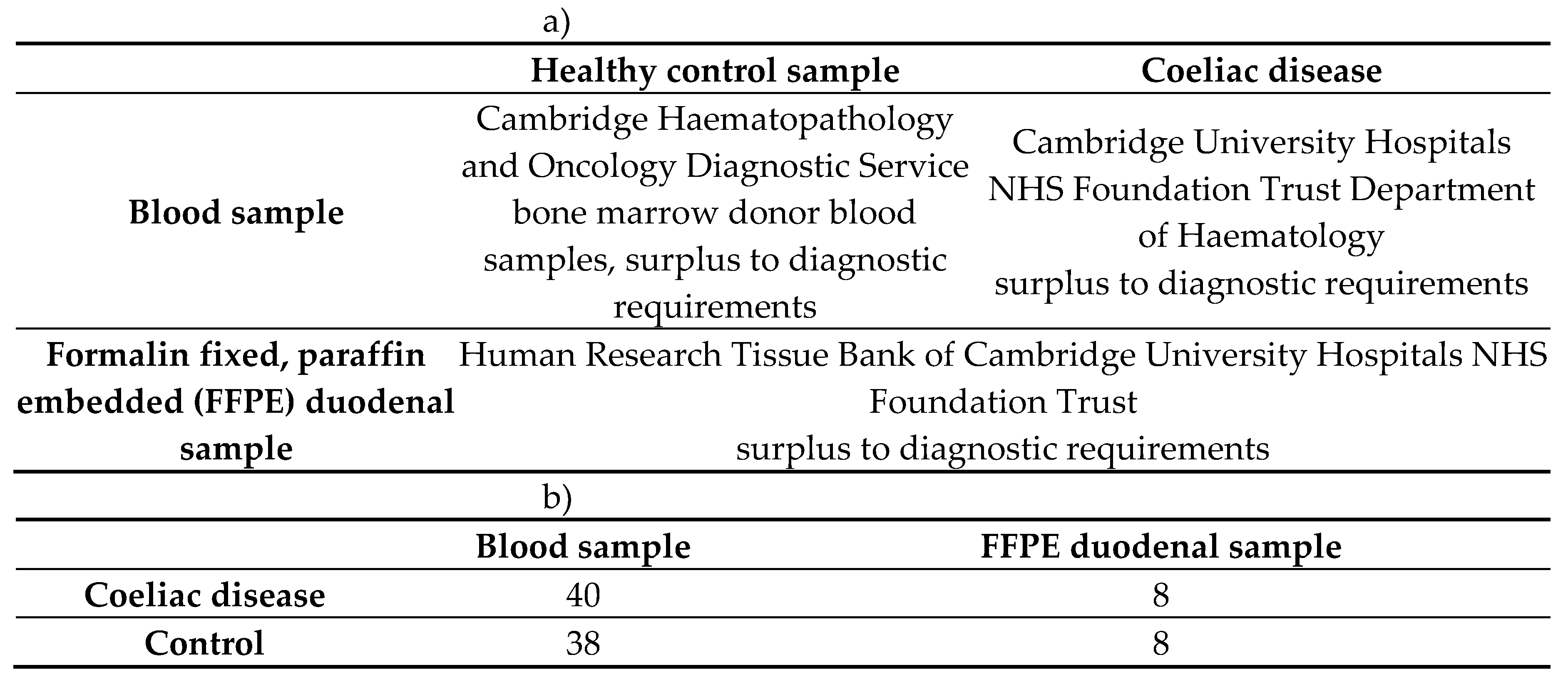 |
4.1.2. Participant Selection for the UK Biobank Single Variant Analysis
- Filled out CeD online questionnaire
- CeD mentioned in hospital inpatient record or death record
- Malabsorption
- Anaemia
- Lymphocytosis
- On a GFD
- Diarrhoea
- Hospital diagnosis record includes coeliac disease: ICD9 (5790), ICD10 (K90.0)
- Cause of death includes coeliac disease: ICD10 (K90.0)
4.1.3. Samples Selected for the HV4 Analysis
| Coeliac disease | Healthy control | Sequencing method | |
| Dataset 1 | 34 FFPE, 12 fresh frozen duodenal | 97 FFPE duodenal | Illumina Miseq micro |
| Dataset 2 | 11 FFPE duodenal | 11 FFPE duodenal | Illumina Miseq |
| Dataset 3 | 84 blood | 130 blood | Illumina NextSeq |
| Combined | 84 blood, 48 FFPE duodenal, 12 fresh frozen duodenal |
130 blood, 108 FFPE duodenal |
NA |
4.2. Analysis of Butyrophilin Family Variation in the Targeted Sequencing Cohort
4.2.1. Gene Selection and Hybridization Probe Design
| Gene of interest | Location (GRCh38.p12) |
| BTN2A1 | chr6:26,457,955-26,476,622 |
| BTN2A2 | chr6:26,382,893-26,394,874 |
| BTN3A1 | chr6:26,402,269-26,415,216 |
| BTN3A2 | chr6:26,365,170-26,378,320 |
| BTN3A3 | chr6:26,440,504-26,453,415 |
| BTNL2 | chr6:32,393,339-32,408,879 |
| BTNL3 | chr5:180,988,846-181,006,727 |
| BTNL8 | chr5:180,899,097-180,952,166 |
| ERMAP | chr1:42,817,122-42,844,991 |
| MOG | chr6:29,657,092-29,672,365 |
4.2.2. Library Capture and Sequencing of HLA Loci and Selected Butyrophilin Genes
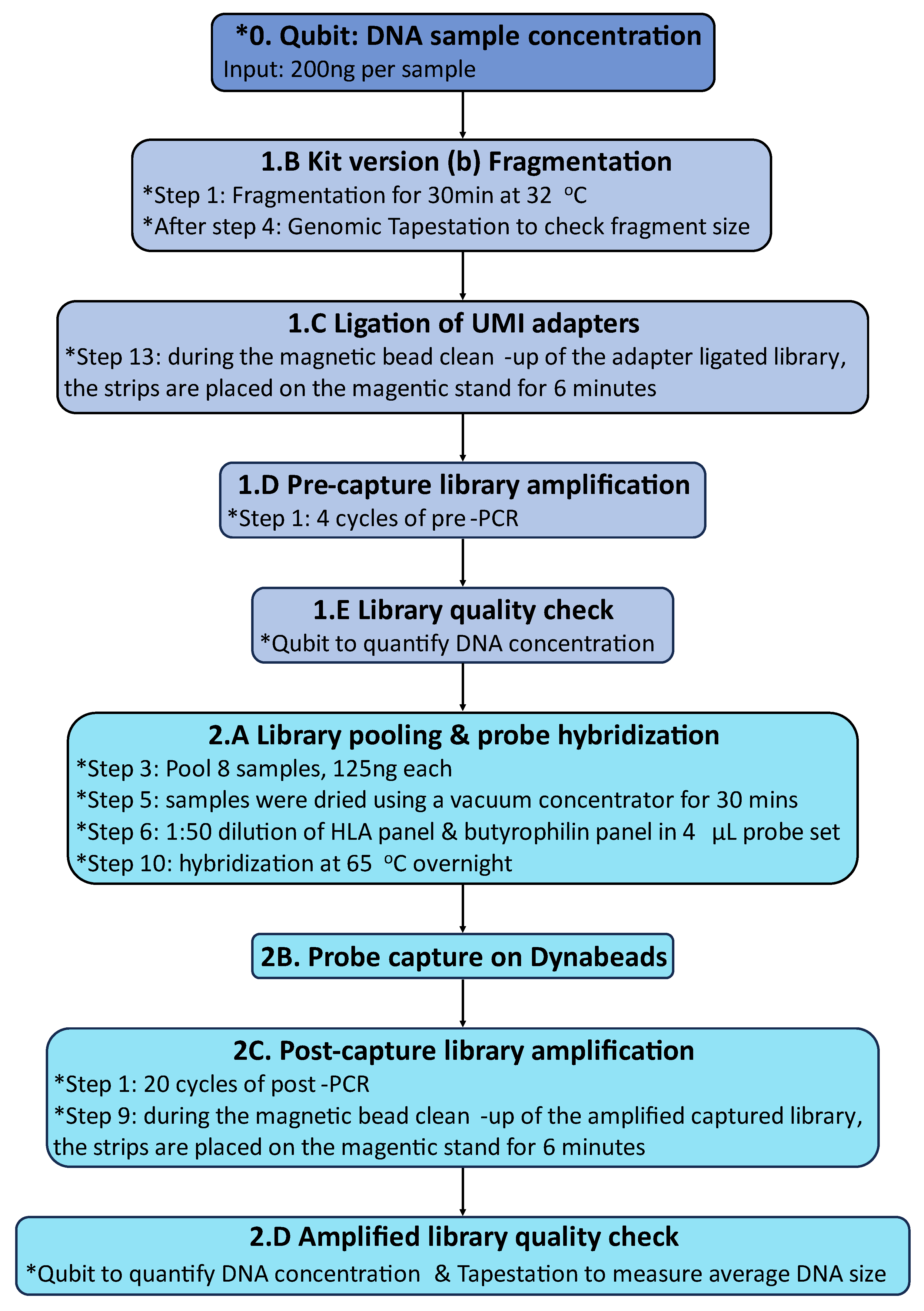
4.2.3. Germline Short Variant Discovery and HLA Typing
4.2.4. Copy Number Variation (CNV) Analysis of the BTNL8-BTNL3 Loci
| rs72494581 genotypes | Associated CNV at BTNL8-BTNL3 region of chromosome 5 | |
| TT | Homozygous for reference allele | Full length BTNL8-BTNL3 region on both copies of chromosome 5 |
| CT | Heterozygous | 1 copy has full length BTNL8-BTNL3 region 1 copy has BTNL8*BTNL3 deletion |
| CC | Homozygous for alternative allele | BTNL8*BTNL3 deletion on both copies of chromosome 5 |
4.2.5. Burden Testing Analysis
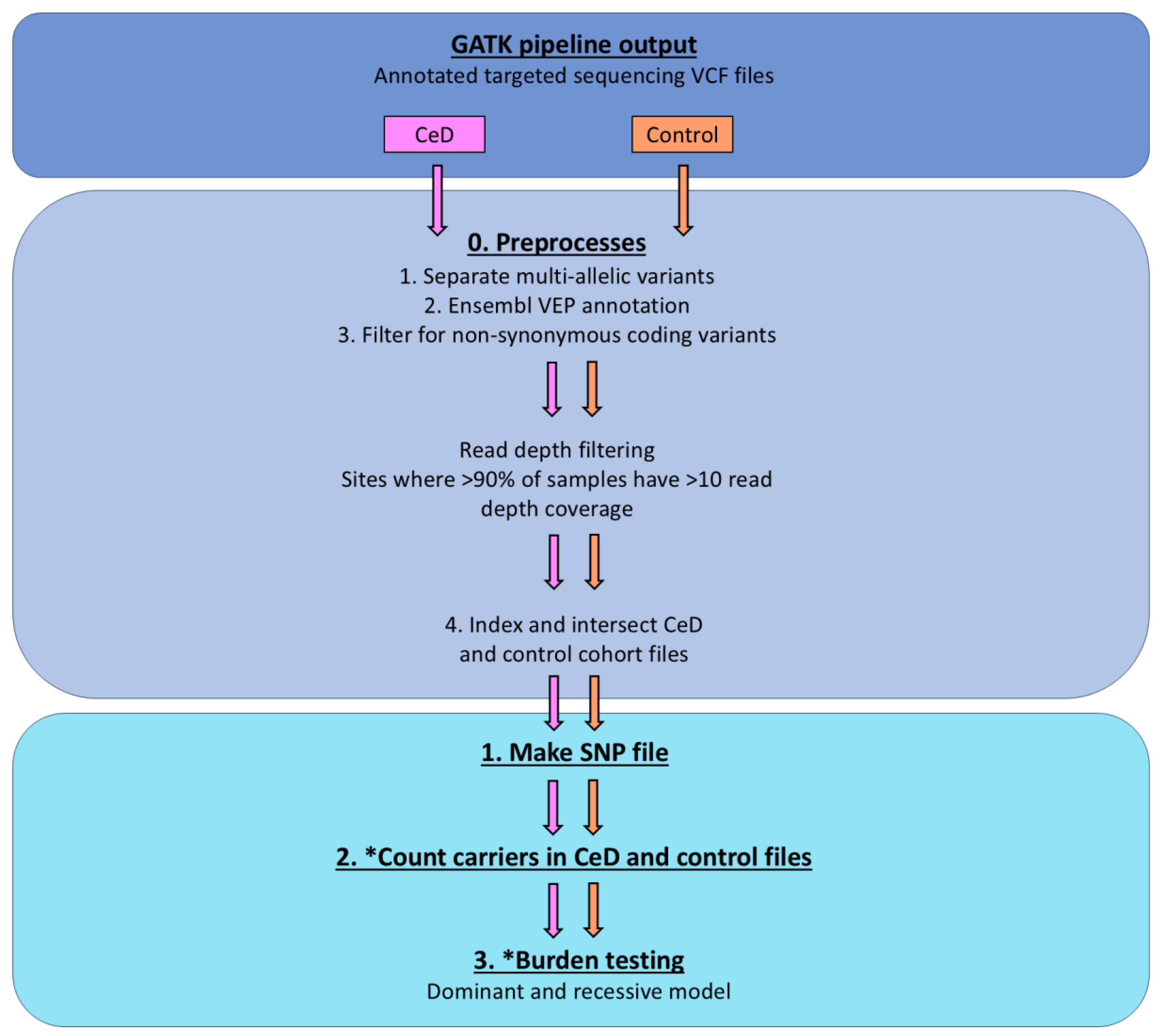
4.3. Single Variant Testing Butyrophilin Family Variance in the UK Biobank Database
4.3.1. CeD Risk Associated HLA Genotyping Using the HLA Imputation Values of the UK Biobank 500,000 Genome-Wide Genotyping Cohort
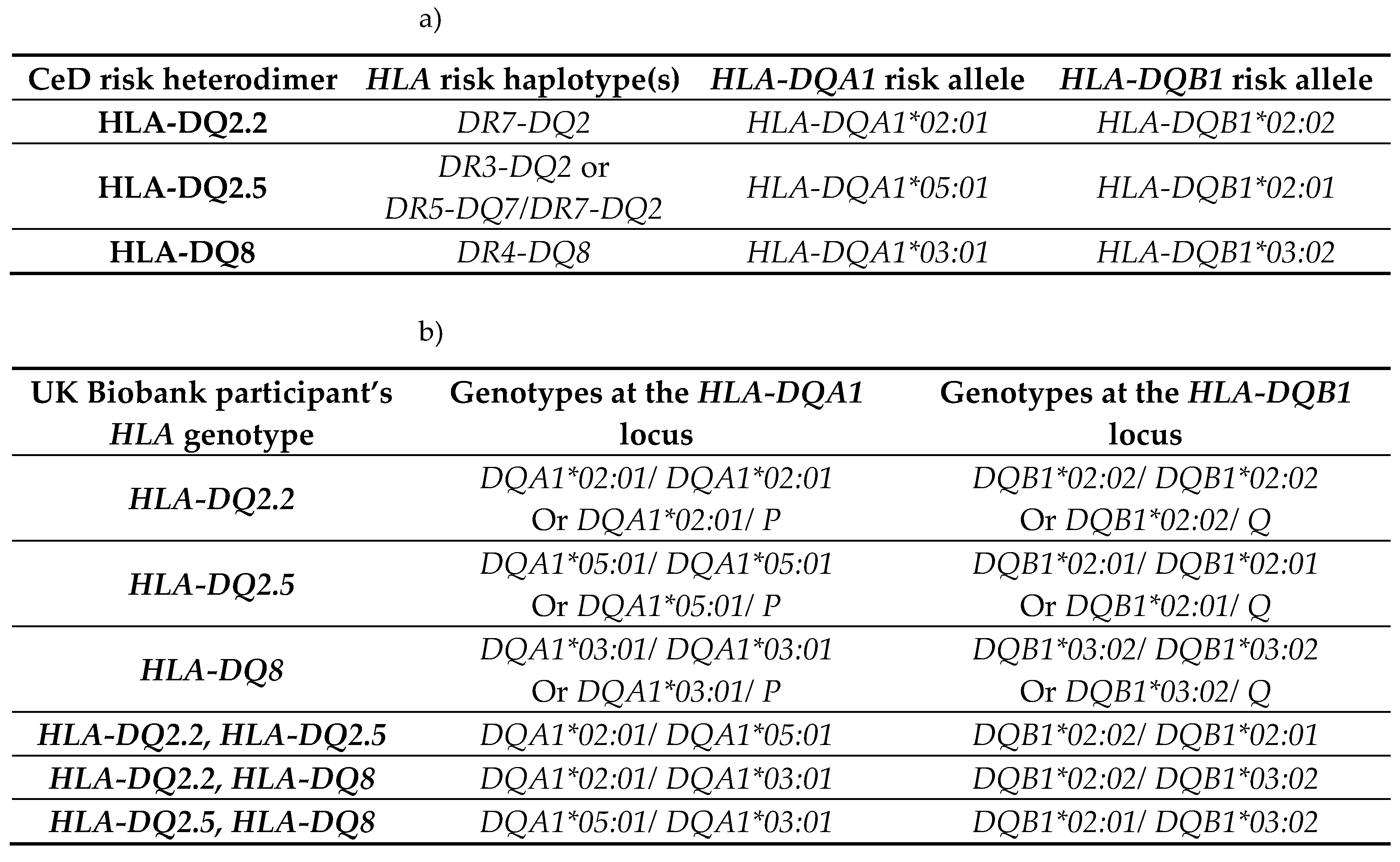 |
 |
4.3.2. Single Variant Testing Using Binomial Regression Models
| Test/group number | Association being tested | Predictor variable(s) | Response variable |
| First test | Association between HLA risk genotypes and CeD risk |
HLA risk genotype: See Table 17 |
CeD status: CeD or control |
| Second group of tests (101 models) | Association between individual butyrophilin SNPs and CeD risk | Butyrophilin SNP: 2 reference alleles 1 reference allele 0 reference allele |
CeD status: CeD or control |
| Third group of tests (101 models) | Association between the combined effect of HLA genotypes and butyrophilin SNPs and CeD risk |
HLA risk genotype: See Table 17 Butyrophilin SNP: 2 reference alleles 1 reference allele 0 reference allele |
CeD status: CeD or control |
| Fourth group of tests | Association between butyrophilin SNPs and CeD risk in HLA-matched groups | Butyrophilin SNP: 2 reference alleles 1 reference allele 0 reference allele |
CeD status: CeD or control |
4.4. Analysis of TRGV Usage and HV4 Variation in CeD and Control Samples
4.4.1. Processing Samples and TCR Sequencing
4.4.2. TRGV and HV4 Analysis Pipeline
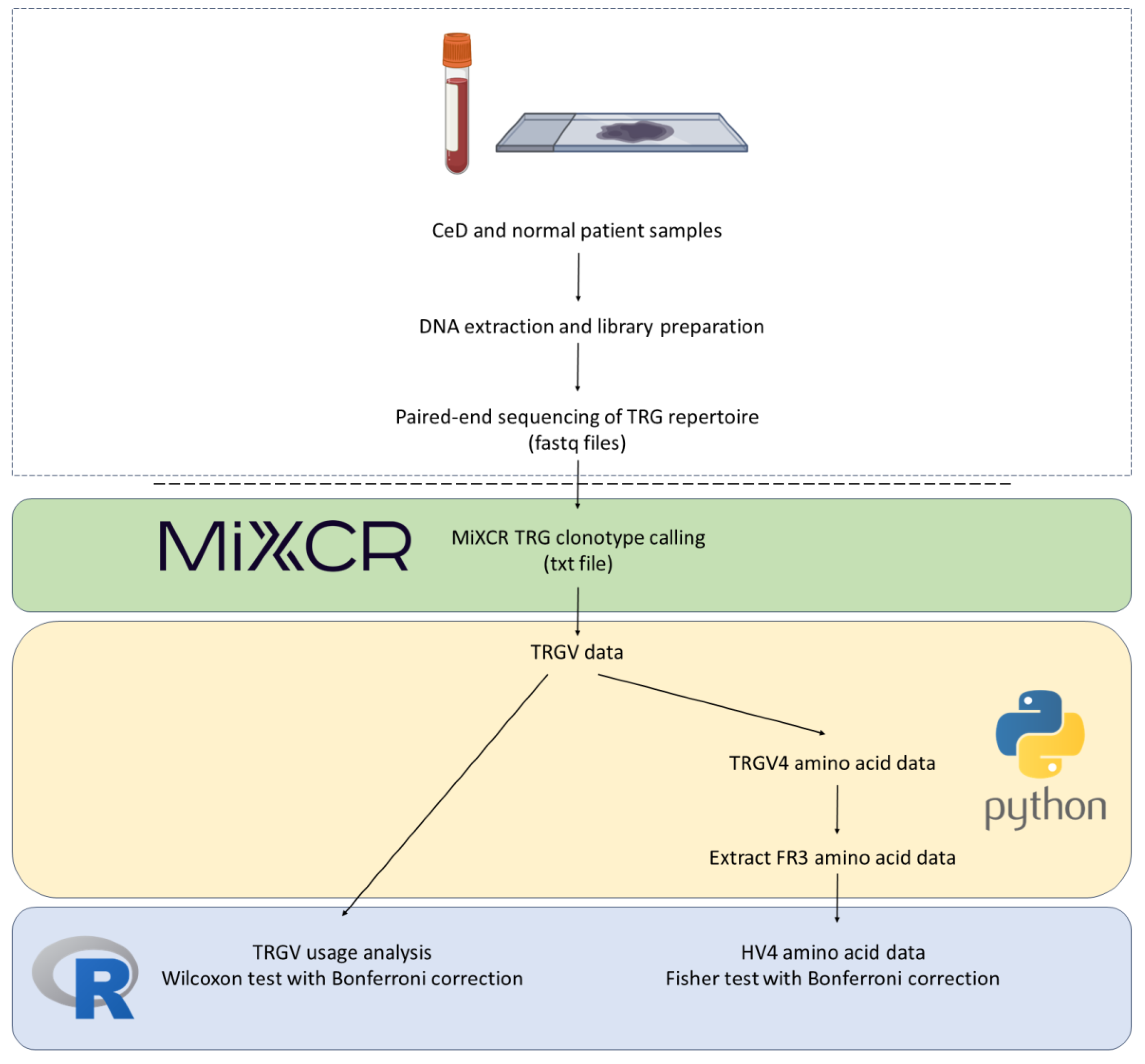
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BTN/BTNL | Butyrophilin/butyrophilin-like |
| CeD | Coeliac disease |
| CNV | Copy number variation |
| DC | Dendritic cell |
| GFD | Gluten-free diet |
| FFPE | Formalin fixed, paraffin embedded |
| HLA | Human leukocyte antigen |
| HPA | Human Protein Atlas |
| HV4 | Hypervariable region 4 |
| IEL | Intraepithelial lymphocyte |
| NK cell | Natural Killer cell |
| TRGV | T cell receptor γ variable region |
| SNP/SNV | Single nucleotide polymorphism/variation |
| WT | Wild type |
Appendix A – Supplementary Materials for Results Section 2.1
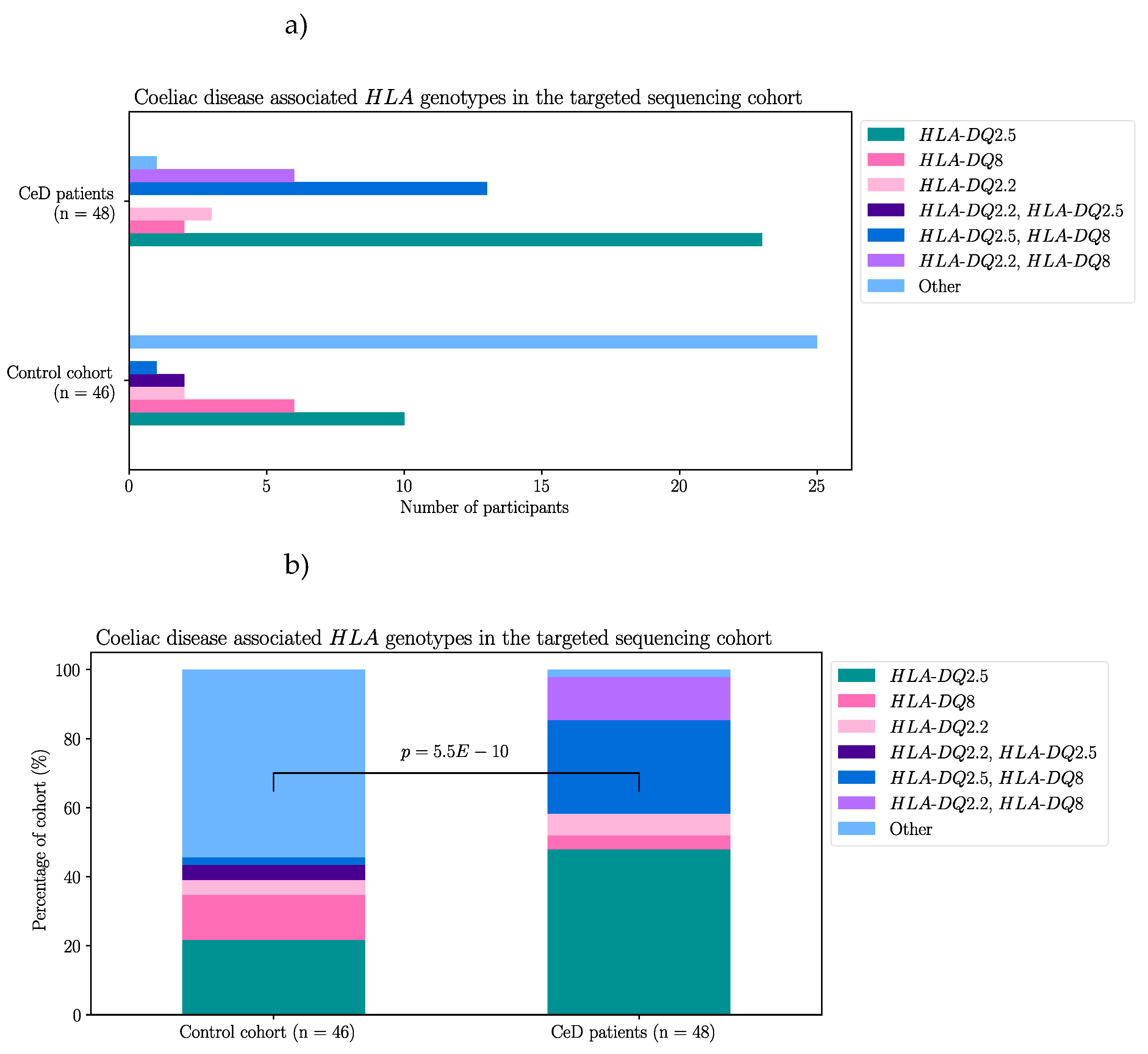
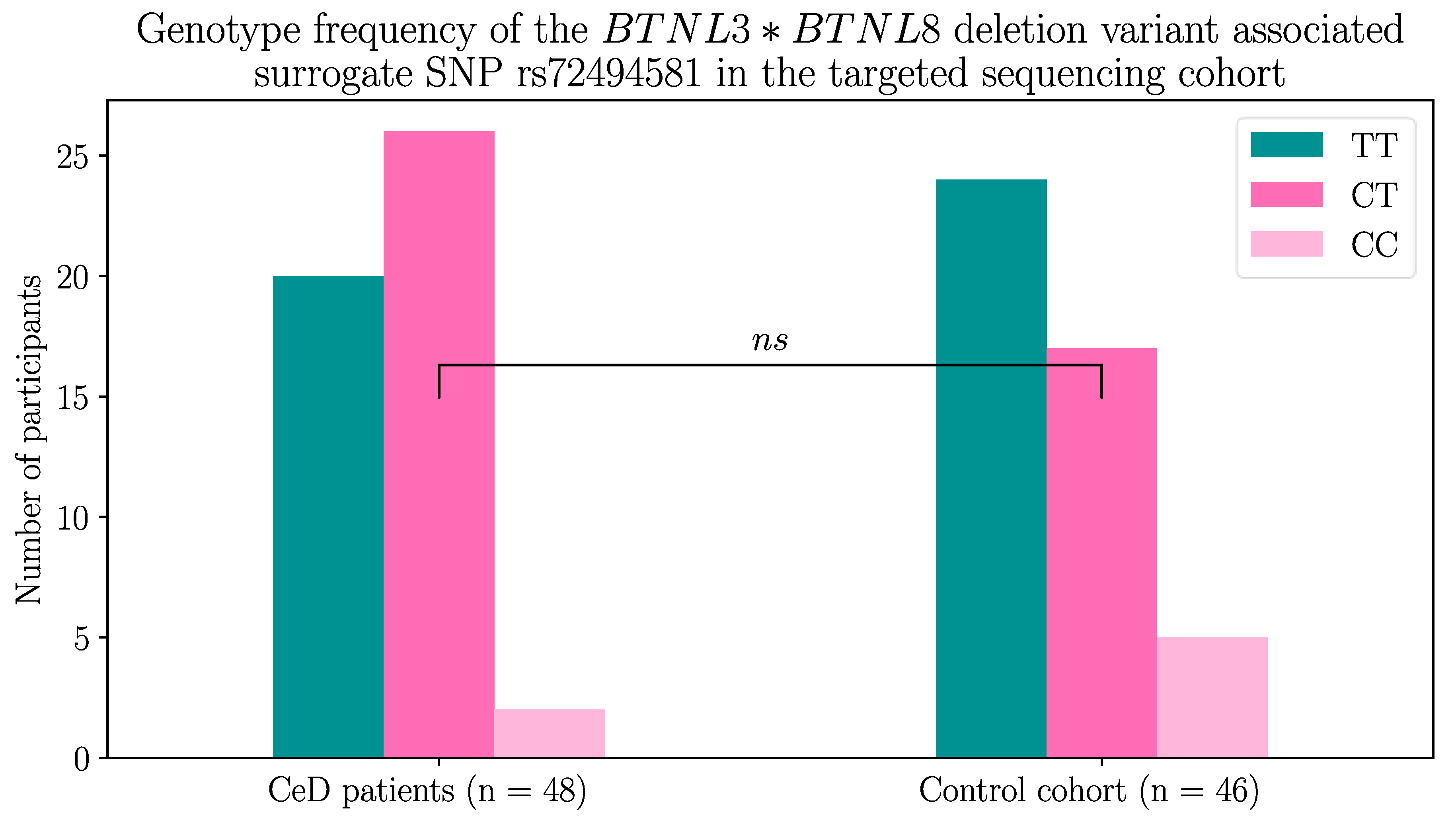
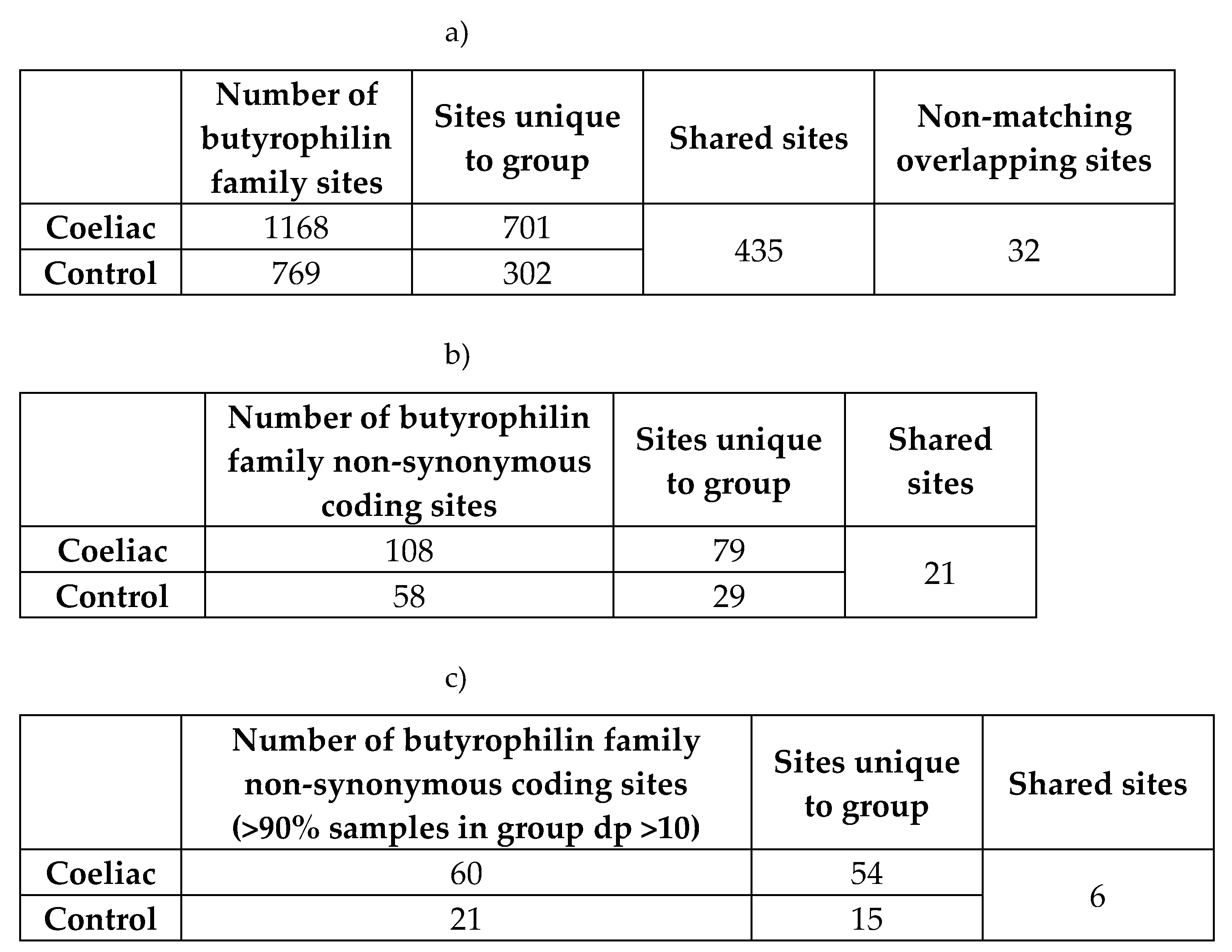 |
| Gene | Qual. SNPs | CeD %(≥1 HET) | CeD %(≥2 HET) | CeD %(HOM ALT) | CeD total qual. allele freq | Control %(≥1 HET) | Control %(≥2 HET) | Control %(HOM ALT) | Control total qual. allele freq | Dominant model p-value | Recessive model p-value |
| BTN2A1 | 3 | 45.8 | 43.8 | 6.3 | 0.281 | 10.9 | 8.7 | 0.0 | 0.047 | 1.46E-05 | 3.70E-08 |
| BTN3A2 | 1 | 10.4 | 0.0 | 2.1 | 0.073 | 19.6 | 0.0 | 2.2 | 0.120 | 0.929 | 0.946 |
| ERMAP | 1 | 43.8 | 0.0 | 16.7 | 0.385 | 43.5 | 0.0 | 15.2 | 0.370 | 0.516 | 0.988 |
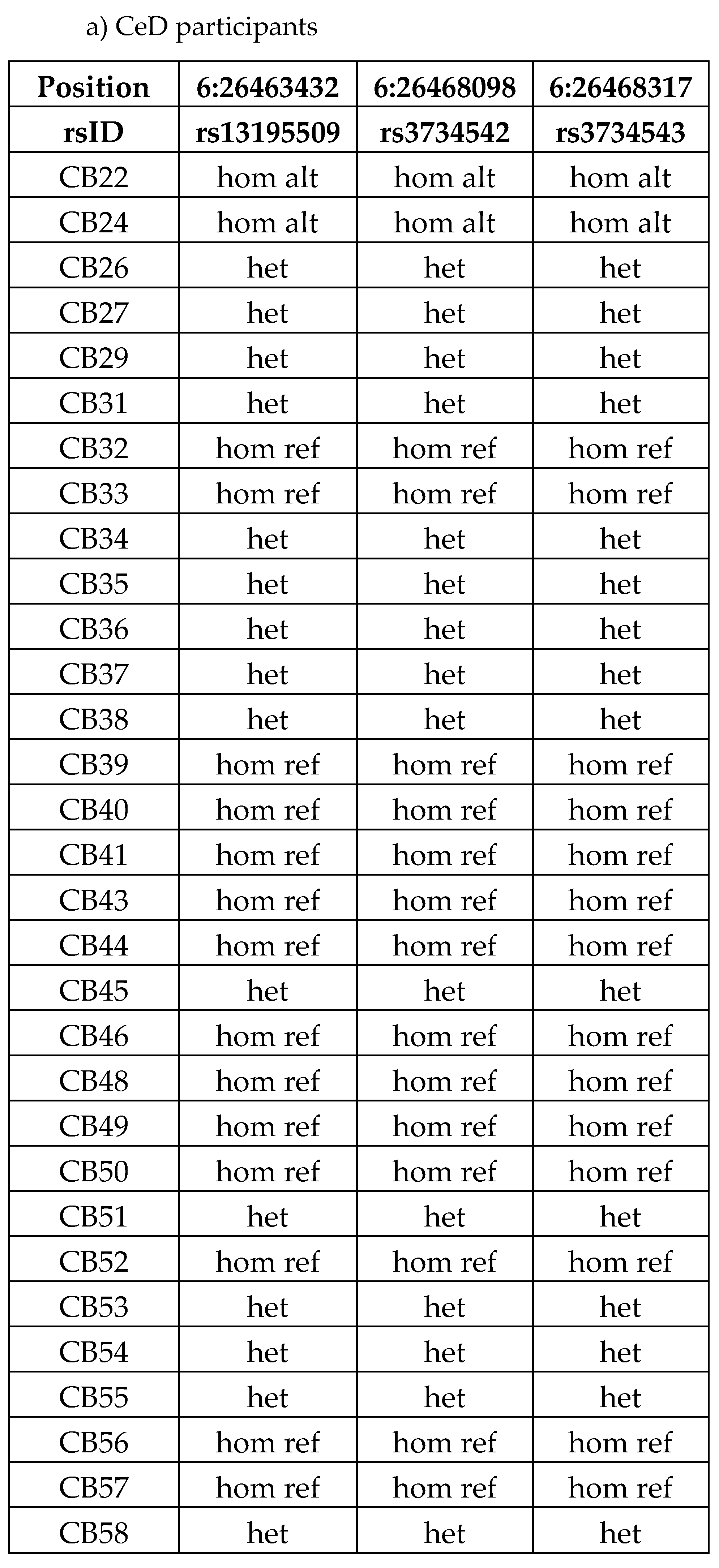 |
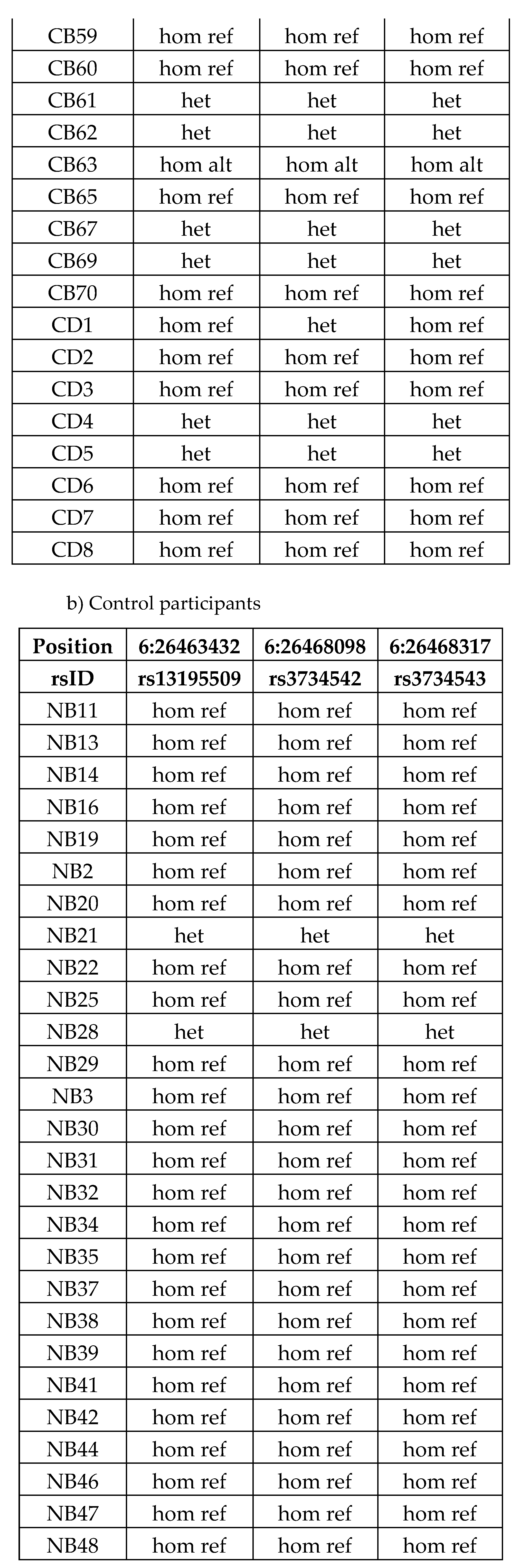 |
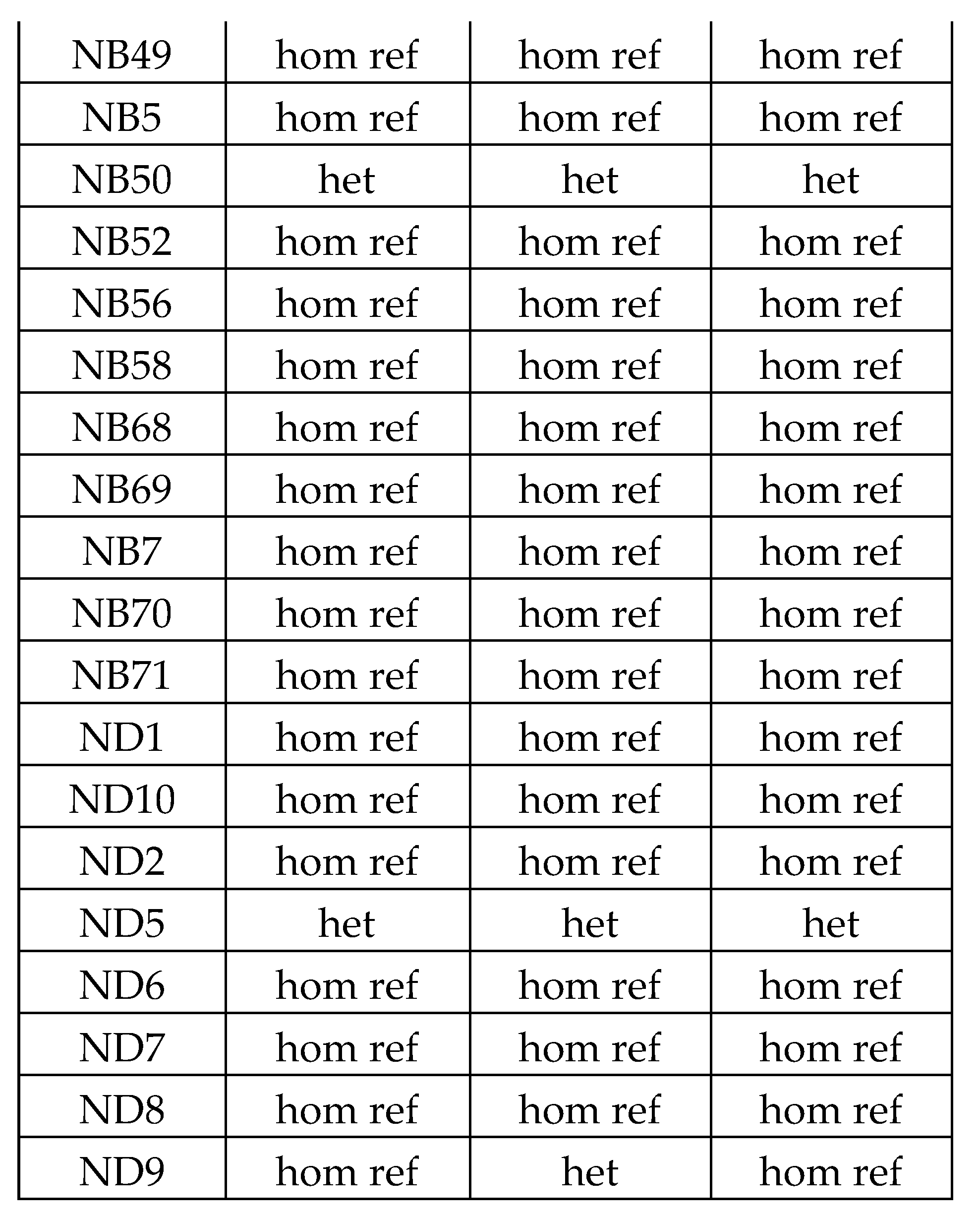 |
Appendix B – Demographic Data of the Selected UK Biobank Participants
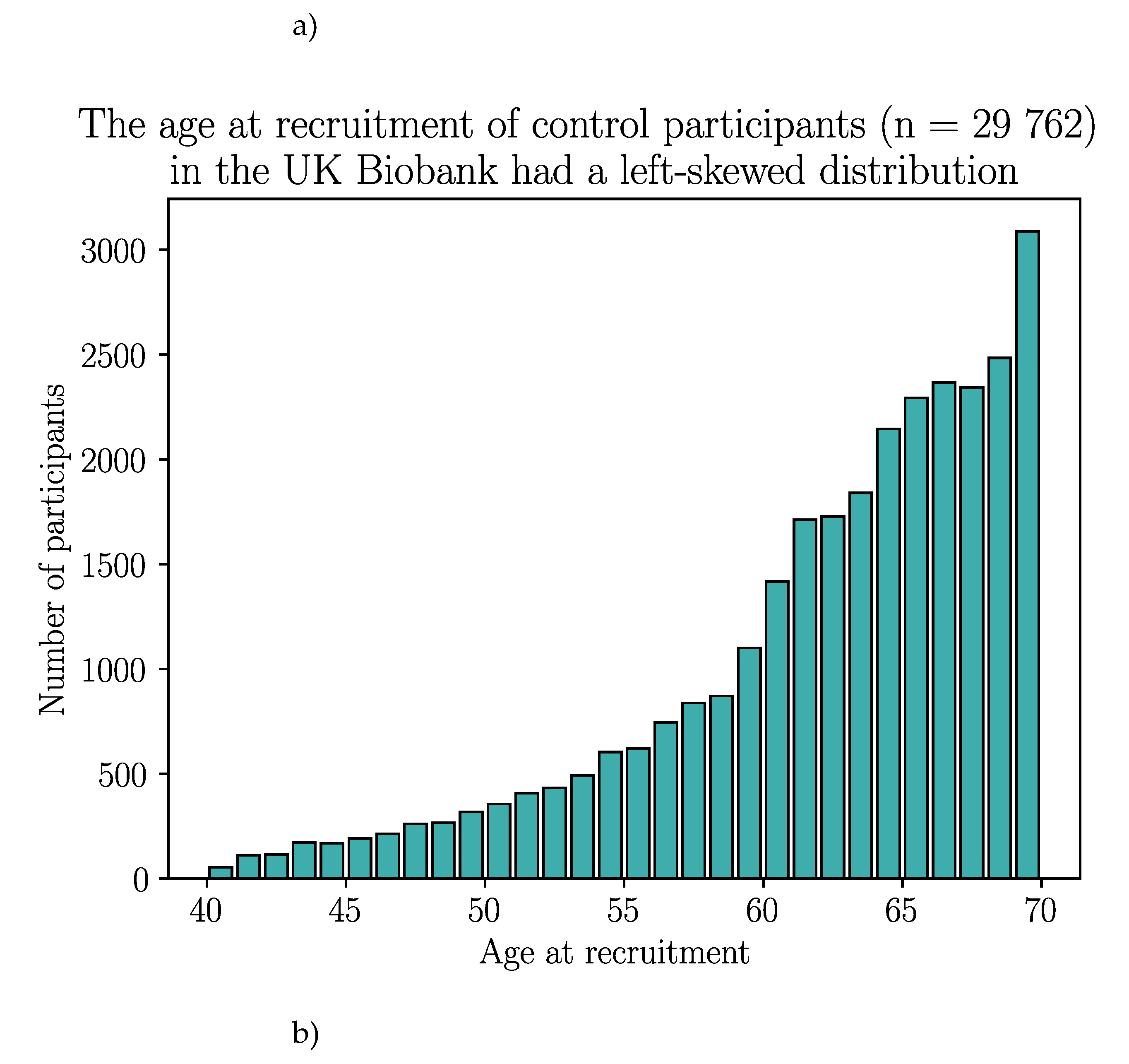
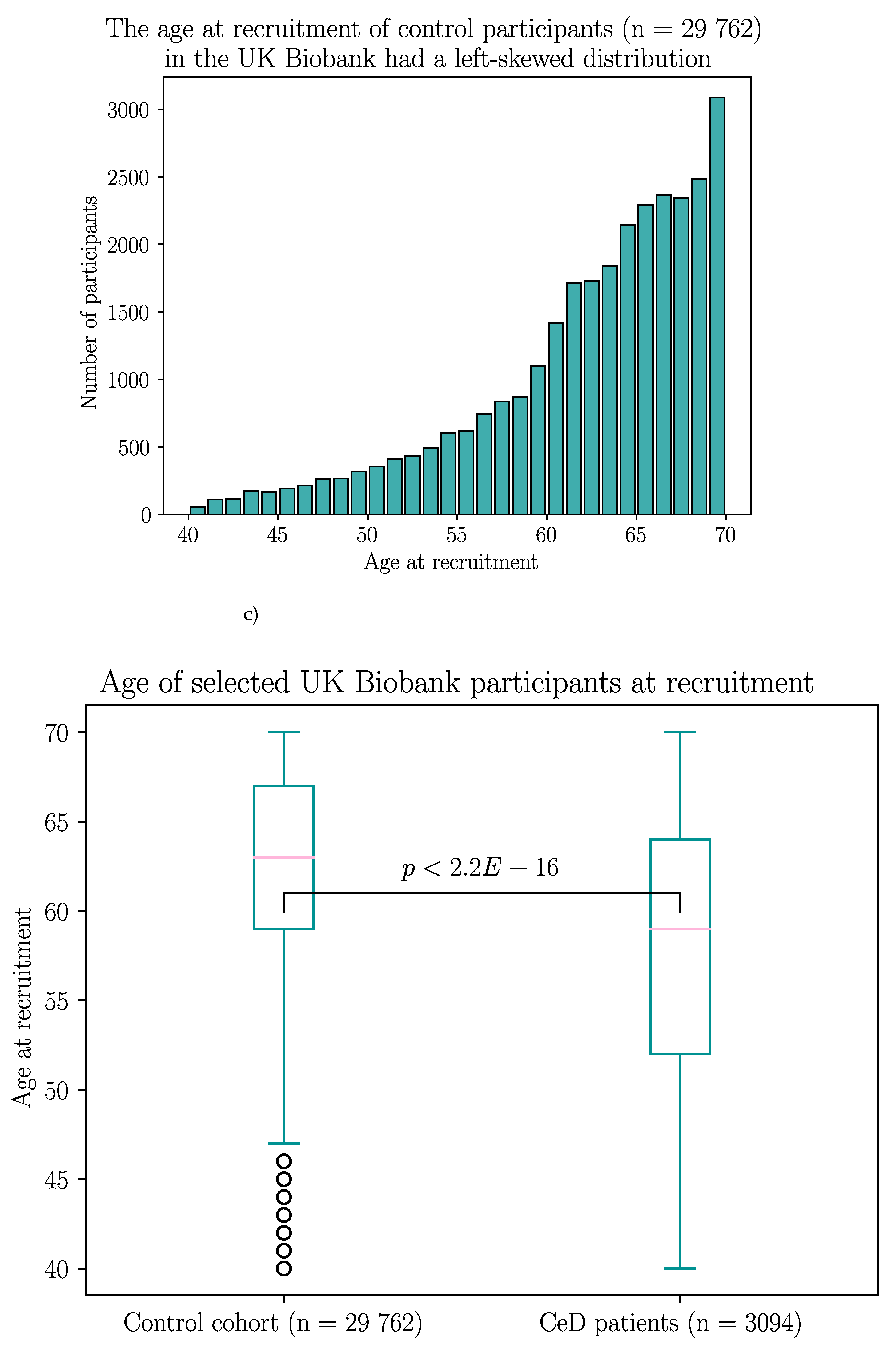
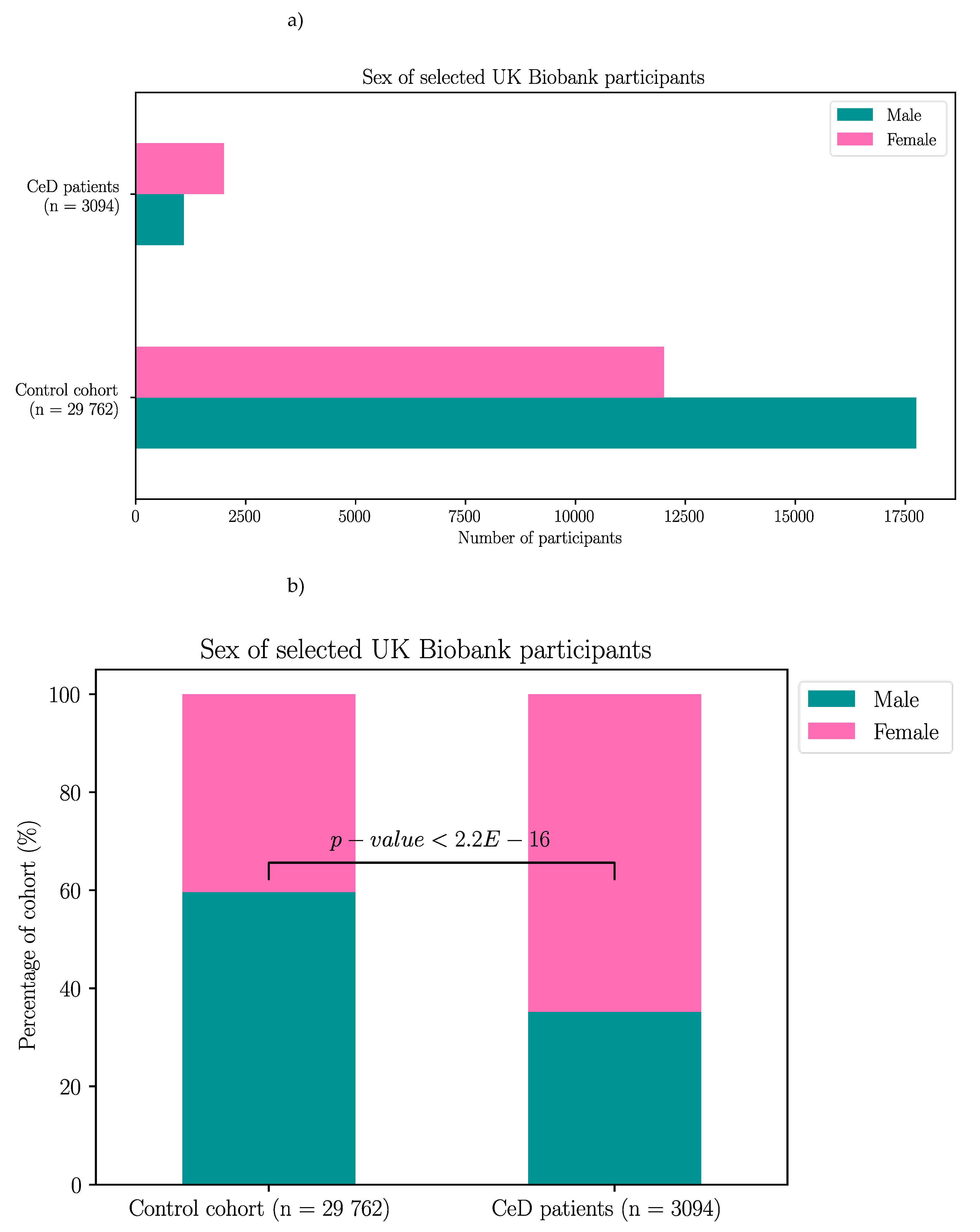
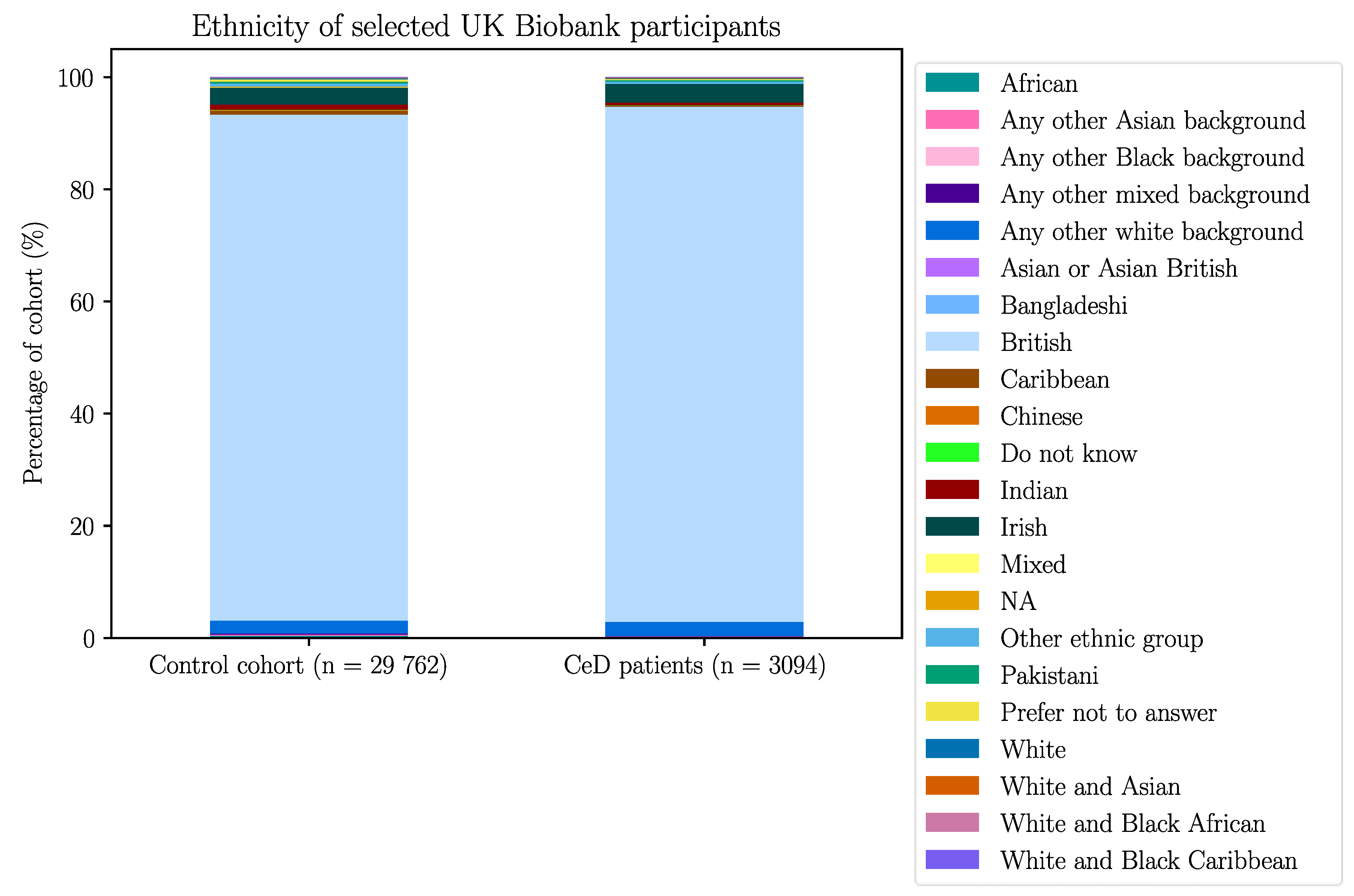
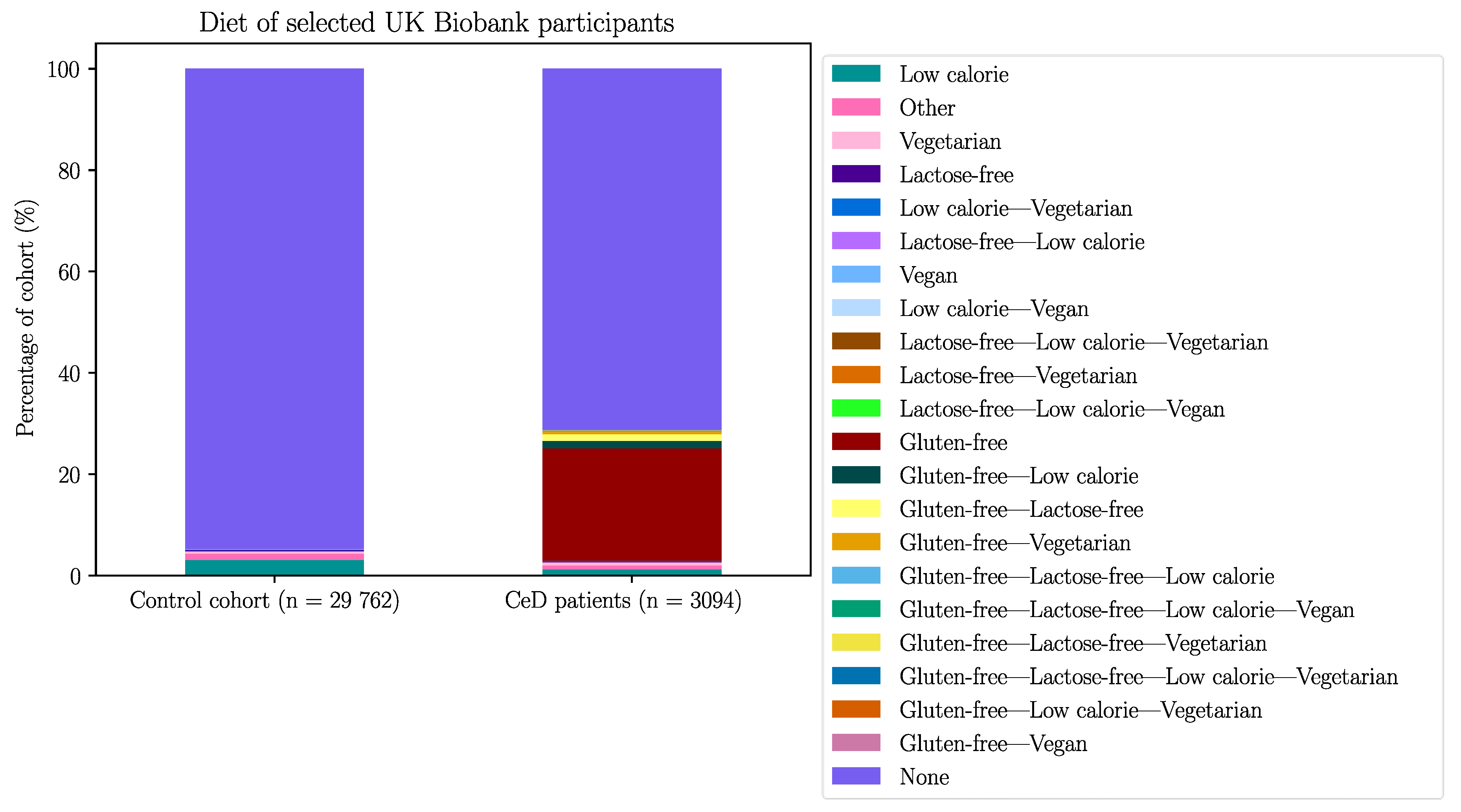
| Without GFD | With GFD | |||||
| Diet | N controls on diet | % Controls on diet | N CeD on diet | % CeD on diet (out of 2296) | N CeD on diet | % CeD on diet (out of 3094) |
| Lactose-free–low calorie | 20 | 0.067 | 0 | 0 | 5 | 0.162 |
| Vegan | 15 | 0.050 | 0 | 0 | 1 | 0.032 |
| Lactose-free–low calorie–vegetarian | 2 | 0.007 | 0 | 0 | 1 | 0.032 |
| Lactose-free–vegetarian | 2 | 0.007 | 0 | 0 | 1 | 0.032 |
| Lactose-free–low calorie–vegan | 1 | 0.003 | 0 | 0 | 2 | 0.065 |
Appendix C – Supplementary Materials for Results Section 2.2
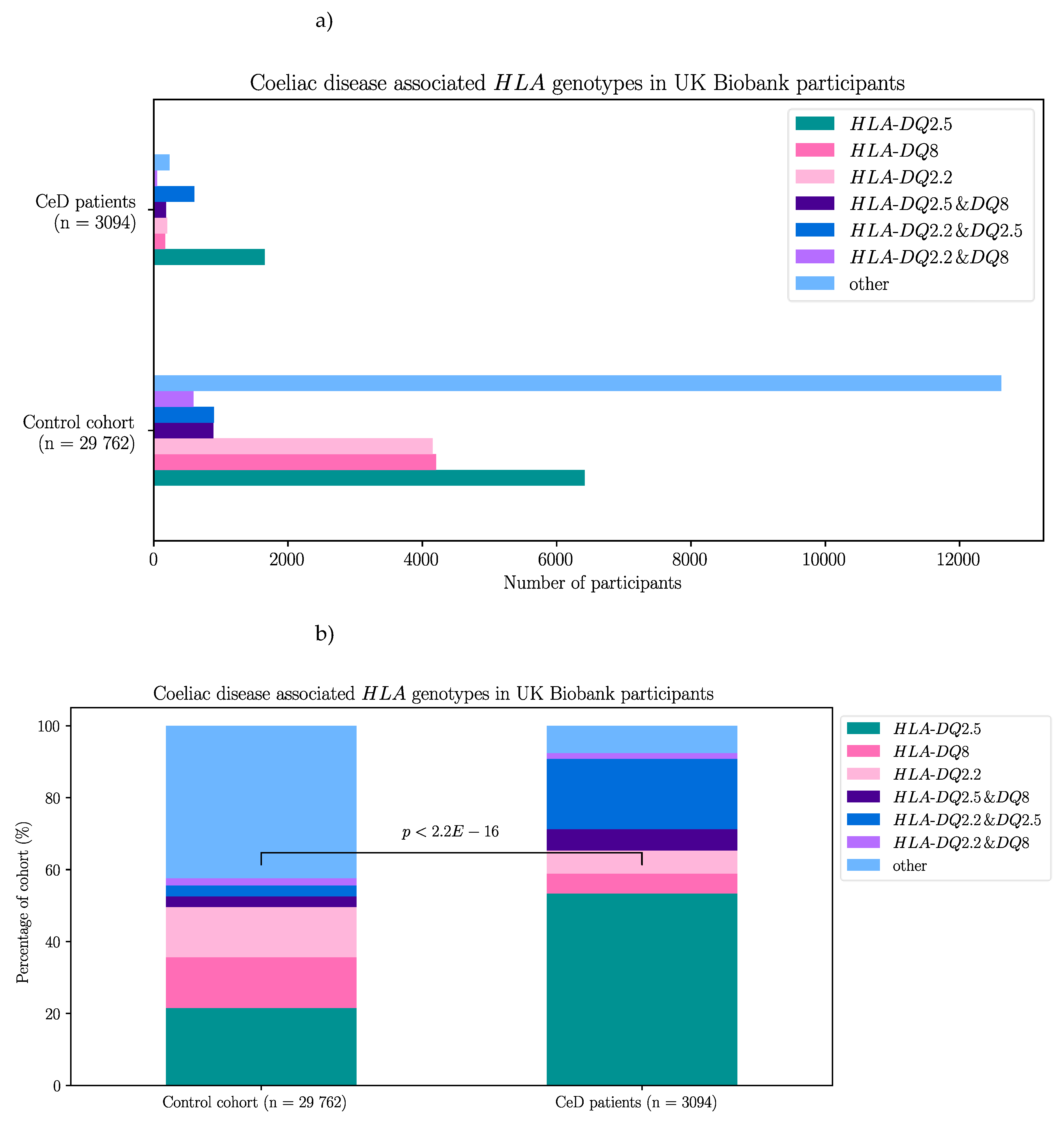
| HLAgenotype | Coefficient estimate | Standard error | z value | p-value | CeD risk |
| HLA-DQ2.2, HLA-DQ2.5 | 2.649 | 0.090 | 29.550 | < 2E-16 | Increase |
| HLA-DQ2.2, HLA-DQ8 | 0.570 | 0.164 | 3.474 | 5.13E-04 | Increase |
| HLA-DQ2.5 | 1.682 | 0.078 | 21.662 | < 2E-16 | Increase |
| HLA-DQ2.5, HLA-DQ8 | 1.456 | 0.109 | 13.352 | < 2E-16 | Increase |
| HLA-DQ8 | -0.163 | 0.107 | -1.533 | 0.125 | ns |
| Other HLA genotype | -0.949 | 0.098 | -9.677 | < 2E-16 | Decrease |
| Constant | -3.039 | 0.073 | -41.872 | < 2E-16 | NA |
| SNP name | Gene | OR | upper | lower | adjusted p-value | ln(OR) |
| rs10484441_G | BTN2A1 | 1.138386 | 1.249955 | 1.038835 | 0.60789 | 0.129612 |
| rs12660069_C | BTN2A1 | 1.106915 | 1.299592 | 0.948994 | 1 | 0.101577 |
| rs13195402_G | BTN2A1 | 0.397002 | 0.424644 | 0.371261 | 4.67E-158 | -0.92381 |
| rs13195509_G | BTN2A1 | 0.424358 | 0.45231 | 0.398239 | 1.61E-151 | -0.85718 |
| rs13437351_G | BTN2A1 | 1.485538 | 1.91326 | 1.176198 | 0.14151 | 0.395777 |
| rs1407045_A | BTN2A1 | 1.314112 | 1.385827 | 1.246309 | 6.07E-22 | 0.273161 |
| rs142951857_A | BTN2A1 | 1.391065 | 3.107641 | 0.723169 | 1 | 0.33007 |
| rs143104579_G | BTN2A1 | 1.14068 | 1.405753 | 0.935552 | 1 | 0.131625 |
| rs146399224_T | BTN2A1 | 10963.24 | NA | 0.003938 | 1 | 9.302303 |
| rs148111655_G | BTN2A1 | 1.130613 | 2.537072 | 0.583563 | 1 | 0.12276 |
| rs2273558_A | BTN2A1 | 0.673125 | 0.711994 | 0.636428 | 1.69E-41 | -0.39582 |
| rs2893856_T | BTN2A1 | 0.839708 | 0.911101 | 0.772766 | 0.0032259 | -0.1747 |
| rs2893857_C | BTN2A1 | 1.142895 | 1.255236 | 1.042695 | 0.48083 | 0.133565 |
| rs3734539_C | BTN2A1 | 4032.285 | NA | 0.007 | 1 | 8.302088 |
| rs3734542_G | BTN2A1 | 0.425283 | 0.453292 | 0.39911 | 8.59E-151 | -0.855 |
| rs3734543_G | BTN2A1 | 0.430012 | 0.458993 | 0.402974 | 1.59E-140 | -0.84394 |
| rs3799380_T | BTN2A1 | 0.577632 | 0.611726 | 0.545555 | 8.59E-77 | -0.54882 |
| rs56296968_C | BTN2A1 | 0.54644 | 0.579504 | 0.515375 | 9.70E-89 | -0.60433 |
| rs6456724_T | BTN2A1 | 0.838697 | 0.91004 | 0.771803 | 0.00287 | -0.17591 |
| rs6907857_T | BTN2A1 | 1.433106 | 1.834039 | 1.140956 | 0.29414 | 0.359844 |
| rs6911470_C | BTN2A1 | 1.472809 | 1.963726 | 1.132407 | 0.57829 | 0.387171 |
| rs6929846_T | BTN2A1 | 0.813898 | 0.875261 | 0.756001 | 3.60E-06 | -0.20592 |
| rs7773913_C | BTN2A1 | 1.433061 | 1.833981 | 1.14092 | 0.29439 | 0.359813 |
| rs7773938_C | BTN2A1 | 0.548927 | 0.582092 | 0.517765 | 1.15E-87 | -0.59979 |
| rs77870445_T | BTN2A1 | 1.219723 | 1.609943 | 0.942473 | 1 | 0.198624 |
| rs9348718_A | BTN2A1 | 1.267016 | 1.454243 | 1.109411 | 0.06114 | 0.236664 |
| rs9358943_C | BTN2A1 | 1.768069 | 31.86429 | 0.363137 | 1 | 0.569888 |
| rs9358944_A | BTN2A1 | 0.546443 | 0.579355 | 0.515512 | 1.55E-89 | -0.60432 |
| rs9358945_A | BTN2A1 | 0.54552 | 0.578367 | 0.514649 | 4.37E-90 | -0.60602 |
| rs9461254_G | BTN2A1 | 1.464419 | 1.953756 | 1.125151 | 0.66799 | 0.381458 |
| rs10456045_G | BTN3A1 | 0.638826 | 0.674371 | 0.605202 | 2.97E-57 | -0.44812 |
| rs10807008_G | BTN3A1 | 1.091584 | 1.192289 | 1.001092 | 1 | 0.087629 |
| rs12200782_C | BTN3A1 | 1.138999 | 1.250121 | 1.039809 | 0.56581 | 0.13015 |
| rs12207930_C | BTN3A1 | 1.147602 | 1.241418 | 1.062201 | 0.05418 | 0.137674 |
| rs12208447_C | BTN3A1 | 1.284779 | 1.624554 | 1.030908 | 1 | 0.250586 |
| rs12214924_T | BTN3A1 | 1.147514 | 1.241076 | 1.062324 | 0.05283 | 0.137597 |
| rs143476765_A | BTN3A1 | 1.108868 | 4.616795 | 0.396888 | 1 | 0.10334 |
| rs144114619_T | BTN3A1 | 1.212854 | 2.147941 | 0.740604 | 1 | 0.192976 |
| rs145059723_A | BTN3A1 | 1.412767 | 4.034311 | 0.629895 | 1 | 0.34555 |
| rs1741738_A | BTN3A1 | 1.144367 | 1.242176 | 1.05573 | 0.11623 | 0.134851 |
| rs17610161_G | BTN3A1 | 1.097034 | 1.199223 | 1.005306 | 1 | 0.09261 |
| rs1796520_C | BTN3A1 | 0.758646 | 0.800004 | 0.719297 | 2.40E-22 | -0.27622 |
| rs3799378_A | BTN3A1 | 0.585559 | 0.61957 | 0.553499 | 2.92E-75 | -0.53519 |
| rs3857549_C | BTN3A1 | 1.247408 | 1.401099 | 1.114594 | 0.01526 | 0.221068 |
| rs3902051_A | BTN3A1 | 1.090365 | 1.187959 | 1.002366 | 1 | 0.086513 |
| rs41266839_G | BTN3A1 | 0.396746 | 0.423498 | 0.37178 | 2.12E-168 | -0.92446 |
| rs4609015_T | BTN3A1 | 1.151594 | 1.245578 | 1.066033 | 0.03817 | 0.141147 |
| rs4712990_C | BTN3A1 | 1.101764 | 1.203269 | 1.010539 | 1 | 0.096912 |
| rs55676749_T | BTN3A1 | 1.13993 | 1.38368 | 0.948273 | 1 | 0.130967 |
| rs56161420_G | BTN3A1 | 1.138909 | 1.230815 | 1.055142 | 0.09381 | 0.130071 |
| rs6900725_T | BTN3A1 | 1.149401 | 1.242804 | 1.064341 | 0.04333 | 0.139241 |
| rs6912853_C | BTN3A1 | 1.16488 | 1.257236 | 1.080577 | 0.00785 | 0.152618 |
| rs6920986_C | BTN3A1 | 1.148238 | 1.241884 | 1.062975 | 0.04995 | 0.138228 |
| rs6921148_T | BTN3A1 | 1.165176 | 1.411589 | 0.971033 | 1 | 0.152872 |
| rs742090_A | BTN3A1 | 0.759078 | 0.800536 | 0.719639 | 3.58E-22 | -0.27565 |
| rs7770214_G | BTN3A1 | 1.145779 | 1.239155 | 1.060758 | 0.06048 | 0.136085 |
| rs80153343_G | BTN3A1 | 1.136899 | 1.439622 | 0.910733 | 1 | 0.128305 |
| rs11758089_T | BTN3A2 | 1.192878 | 1.288514 | 1.105674 | 0.00063 | 0.176369 |
| rs12176317_A | BTN3A2 | 0.445307 | 0.474099 | 0.41837 | 6.72E-140 | -0.80899 |
| rs12194095_C | BTN3A2 | 1.118798 | 1.235914 | 1.015164 | 1 | 0.112255 |
| rs12199613_C | BTN3A2 | 0.67037 | 0.706851 | 0.635751 | 1.76E-47 | -0.39993 |
| rs12205731_G | BTN3A2 | 1.114755 | 1.232063 | 1.011029 | 1 | 0.108634 |
| rs144016445_G | BTN3A2 | 81101.36 | NA | 217.0251 | 1 | 11.30346 |
| rs1977_A | BTN3A2 | 0.445697 | 0.474831 | 0.418455 | 1.17E-136 | -0.80811 |
| rs1979_G | BTN3A2 | 0.44537 | 0.474171 | 0.418426 | 8.23E-140 | -0.80885 |
| rs1985732_A | BTN3A2 | 0.63289 | 0.668226 | 0.599467 | 2.87E-59 | -0.45746 |
| rs2073526_G | BTN3A2 | 0.74435 | 0.785579 | 0.705115 | 9.15E-25 | -0.29524 |
| rs35183513_G | BTN3A2 | 1.102861 | 1.203659 | 1.012184 | 1 | 0.097908 |
| rs58367598_T | BTN3A2 | 1.234256 | 1.449383 | 1.057523 | 0.89091 | 0.210469 |
| rs7765566_G | BTN3A2 | 1.269787 | 1.499163 | 1.083157 | 0.39847 | 0.23885 |
| rs9104_G | BTN3A2 | 1.092904 | 1.187567 | 1.007255 | 1 | 0.088838 |
| rs9358934_G | BTN3A2 | 0.447824 | 0.476845 | 0.420678 | 2.34E-137 | -0.80335 |
| rs9379855_T | BTN3A2 | 0.447682 | 0.47666 | 0.420575 | 8.85E-138 | -0.80367 |
| rs9379858_T | BTN3A2 | 0.448449 | 0.477474 | 0.421299 | 3.19E-137 | -0.80196 |
| rs9379859_C | BTN3A2 | 0.447843 | 0.476903 | 0.420662 | 5.37E-137 | -0.80331 |
| rs9379861_G | BTN3A2 | 1.225507 | 1.619363 | 0.945713 | 1 | 0.203355 |
| rs9393713_G | BTN3A2 | 0.442958 | 0.471602 | 0.416159 | 1.07E-141 | -0.81428 |
| rs9393714_G | BTN3A2 | 0.443419 | 0.47214 | 0.416551 | 6.99E-141 | -0.81324 |
| rs186813312_C | BTNL3 | 0.103966 | NA | NA | NA | -2.26369 |
| rs199970076_G | BTNL3 | 0.544013 | 10.42476 | 0.087697 | 1 | -0.60878 |
| rs201534771_G | BTNL3 | 0.108957 | NA | NA | NA | -2.2168 |
| rs201813197_C | BTNL3 | 1.141842 | 4.74926 | 0.409599 | 1 | 0.132642 |
| rs35157246_C | BTNL3 | 1.069831 | 1.242264 | 0.926052 | 1 | 0.067501 |
| rs4700774_G | BTNL3 | 0.943446 | 0.999778 | 0.89061 | 1 | -0.05822 |
| rs59220426_C | BTNL3 | 1.006054 | 1.132599 | 0.896669 | 1 | 0.006036 |
| rs73815153_G | BTNL3 | 1.009988 | 1.138564 | 0.899028 | 1 | 0.009938 |
| rs7713324_A | BTNL3 | 1.004294 | 1.130736 | 0.895001 | 1 | 0.004284 |
| rs7726604_C | BTNL3 | 1.004751 | 1.13121 | 0.895444 | 1 | 0.00474 |
| rs112469887_G | BTNL8 | 1.084234 | 1.330225 | 0.893133 | 1 | 0.080874 |
| rs113071395_G | BTNL8 | 0.867372 | 1.006007 | 0.751681 | 1 | -0.14229 |
| rs113534626_A | BTNL8 | 1.019956 | 1.206531 | 0.868252 | 1 | 0.019759 |
| rs141492316_T | BTNL8 | 0.891806 | 1.095588 | 0.733483 | 1 | -0.11451 |
| rs145199317_A | BTNL8 | 0.907765 | 1.254932 | 0.673198 | 1 | -0.09677 |
| rs151174174_C | BTNL8 | 0.770441 | 0.932722 | 0.641625 | 0.63031 | -0.26079 |
| rs17704291_C | BTNL8 | 0.940216 | 0.996283 | 0.88763 | 1 | -0.06165 |
| rs200633883_C | BTNL8 | 0.311552 | 1.114968 | 0.108457 | 1 | -1.16619 |
| rs201214790_T | BTNL8 | 4044.713 | NA | 1.08E-07 | 1 | 8.305166 |
| rs201891387_G | BTNL8 | 0.62355 | 2.663098 | 0.210953 | 1 | -0.47233 |
| rs2276995_A | BTNL8 | 0.983681 | 1.038422 | 0.931987 | 1 | -0.01645 |
| rs2619739_C | BTNL8 | 1.101246 | 1.212402 | 1.002386 | 1 | 0.096442 |
| rs7724813_G | BTNL8 | 1.078621 | 1.169543 | 0.996169 | 1 | 0.075683 |
| SNP, reference allele | Gene | Number of the SNP in control | Number of the SNP in CeD | Total allele count in control | Total allele count in CeD | Total number of the SNP in the UK Biobank | Total allele count in UK Biobank |
| rs13195402_G | BTN2A1 | 52060 | 4631 | 58392 | 6028 | 56691 | 64420 |
| rs13195509_G | BTN2A1 | 52271 | 4654 | 59474 | 6176 | 56925 | 65650 |
| rs1407045_A | BTN2A1 | 30596 | 3603 | 59306 | 6172 | 34199 | 65478 |
| rs2273558_A | BTN2A1 | 34599 | 3248 | 51134 | 5570 | 37847 | 56704 |
| rs2893856_T | BTN2A1 | 7815 | 697 | 59452 | 6182 | 8512 | 65634 |
| rs3734542_G | BTN2A1 | 52209 | 4656 | 59432 | 6180 | 56865 | 65612 |
| rs3734543_G | BTN2A1 | 52002 | 4644 | 59146 | 6114 | 56646 | 65260 |
| rs3799380_T | BTN2A1 | 46887 | 4213 | 59354 | 6164 | 51100 | 65518 |
| rs56296968_C | BTN2A1 | 47941 | 4295 | 59422 | 6170 | 52236 | 65592 |
| rs6456724_T | BTN2A1 | 7813 | 696 | 59418 | 6178 | 8509 | 65596 |
| rs6929846_T | BTN2A1 | 10355 | 903 | 59460 | 6180 | 11258 | 65640 |
| rs7773938_C | BTN2A1 | 47953 | 4289 | 59472 | 6162 | 52242 | 65634 |
| rs9358944_A | BTN2A1 | 47929 | 4294 | 59462 | 6182 | 52223 | 65644 |
| rs9358945_A | BTN2A1 | 47944 | 4292 | 59478 | 6182 | 52236 | 65660 |
| rs10456045_G | BTN3A1 | 41458 | 3680 | 59434 | 6174 | 45138 | 65608 |
| rs1796520_C | BTN3A1 | 28075 | 2504 | 59240 | 6176 | 30579 | 65416 |
| rs3799378_A | BTN3A1 | 45113 | 4017 | 59206 | 6152 | 49130 | 65358 |
| rs3857549_C | BTN3A1 | 55572 | 5848 | 59430 | 6170 | 61420 | 65600 |
| rs41266839_G | BTN3A1 | 52985 | 4720 | 59430 | 6178 | 57705 | 65608 |
| rs4609015_T | BTN3A1 | 50759 | 5371 | 59452 | 6170 | 56130 | 65622 |
| rs6900725_T | BTN3A1 | 50682 | 5378 | 59392 | 6182 | 56060 | 65574 |
| rs6912853_C | BTN3A1 | 50145 | 5329 | 59434 | 6176 | 55474 | 65610 |
| rs6920986_C | BTN3A1 | 50787 | 5381 | 59464 | 6182 | 56168 | 65646 |
| rs742090_A | BTN3A1 | 28172 | 2506 | 59438 | 6174 | 30678 | 65612 |
| rs11758089_T | BTN3A2 | 50176 | 5354 | 59432 | 6182 | 55530 | 65614 |
| rs12176317_A | BTN3A2 | 51604 | 4597 | 59488 | 6182 | 56201 | 65670 |
| rs12199613_C | BTN3A2 | 36321 | 3178 | 59396 | 6178 | 39499 | 65574 |
| rs1977_A | BTN3A2 | 50506 | 4497 | 58436 | 6074 | 55003 | 64510 |
| rs1979_G | BTN3A2 | 51551 | 4590 | 59448 | 6176 | 56141 | 65624 |
| rs1985732_A | BTN3A2 | 41492 | 3674 | 59442 | 6174 | 45166 | 65616 |
| rs2073526_G | BTN3A2 | 26272 | 2288 | 59434 | 6176 | 28560 | 65610 |
| rs9358934_G | BTN3A2 | 51477 | 4591 | 59412 | 6174 | 56068 | 65586 |
| rs9379855_T | BTN3A2 | 51458 | 4579 | 59392 | 6162 | 56037 | 65554 |
| rs9379858_T | BTN3A2 | 51474 | 4586 | 59422 | 6170 | 56060 | 65592 |
| rs9379859_C | BTN3A2 | 51530 | 4595 | 59452 | 6174 | 56125 | 65626 |
| rs9393713_G | BTN3A2 | 51601 | 4590 | 59472 | 6178 | 56191 | 65650 |
| rs9393714_G | BTN3A2 | 51581 | 4594 | 59456 | 6180 | 56175 | 65636 |
| SNP name | Gene | OR | upper | lower | adjusted p-value | ln(OR) |
| rs10484441_G | BTN2A1 | 0.979307 | 1.082186 | 0.887707 | 1 | -0.02091 |
| rs12660069_C | BTN2A1 | 0.987937 | 1.171444 | 0.837975 | 1 | -0.01214 |
| rs13195402_G | BTN2A1 | 0.812801 | 0.876887 | 0.753657 | 8.15E-06 | -0.20727 |
| rs13195509_G | BTN2A1 | 0.824983 | 0.886694 | 0.767819 | 1.62E-05 | -0.19239 |
| rs13437351_G | BTN2A1 | 1.359939 | 1.781839 | 1.056236 | 1 | 0.30744 |
| rs1407045_A | BTN2A1 | 1.061681 | 1.125065 | 1.001972 | 1 | 0.059854 |
| rs142951857_A | BTN2A1 | 1.189393 | 2.744421 | 0.589877 | 1 | 0.173443 |
| rs143104579_G | BTN2A1 | 1.045598 | 1.306706 | 0.844534 | 1 | 0.044589 |
| rs146399224_T | BTN2A1 | 2678.29 | NA | 0.001107 | 1 | 7.892934 |
| rs148111655_G | BTN2A1 | 1.06888 | 2.495456 | 0.522042 | 1 | 0.066611 |
| rs2273558_A | BTN2A1 | 0.924046 | 0.983637 | 0.868186 | 1 | -0.07899 |
| rs2893856_T | BTN2A1 | 0.978844 | 1.068109 | 0.895906 | 1 | -0.02138 |
| rs2893857_C | BTN2A1 | 0.983276 | 1.0868 | 0.891136 | 1 | -0.01687 |
| rs3734539_C | BTN2A1 | 12320.17 | NA | 2.26E-05 | 1 | 9.418993 |
| rs3734542_G | BTN2A1 | 0.828358 | 0.890318 | 0.770964 | 2.94E-05 | -0.18831 |
| rs3734543_G | BTN2A1 | 0.845824 | 0.910556 | 0.785966 | 0.000823 | -0.16744 |
| rs3799380_T | BTN2A1 | 0.906299 | 0.966317 | 0.850232 | 0.260718 | -0.09839 |
| rs56296968_C | BTN2A1 | 0.889118 | 0.949206 | 0.833053 | 0.042016 | -0.11753 |
| rs6456724_T | BTN2A1 | 0.975504 | 1.064451 | 0.892856 | 1 | -0.0248 |
| rs6907857_T | BTN2A1 | 1.296362 | 1.68837 | 1.012123 | 1 | 0.259562 |
| rs6911470_C | BTN2A1 | 1.225626 | 1.666065 | 0.92177 | 1 | 0.203452 |
| rs6929846_T | BTN2A1 | 0.956691 | 1.034347 | 0.884033 | 1 | -0.04427 |
| rs7773913_C | BTN2A1 | 1.29982 | 1.692843 | 1.014844 | 1 | 0.262226 |
| rs7773938_C | BTN2A1 | 0.892443 | 0.95269 | 0.83623 | 0.062934 | -0.11379 |
| rs77870445_T | BTN2A1 | 0.971098 | 1.300178 | 0.738108 | 1 | -0.02933 |
| rs9348718_A | BTN2A1 | 1.100624 | 1.274443 | 0.954703 | 1 | 0.095877 |
| rs9358943_C | BTN2A1 | 0.455962 | 8.266602 | 0.091903 | 1 | -0.78535 |
| rs9358944_A | BTN2A1 | 0.888825 | 0.948639 | 0.833001 | 0.038293 | -0.11786 |
| rs9358945_A | BTN2A1 | 0.886765 | 0.946407 | 0.831099 | 0.02906 | -0.12018 |
| rs9461254_G | BTN2A1 | 1.206876 | 1.642976 | 0.90642 | 1 | 0.188035 |
| rs10456045_G | BTN3A1 | 0.918688 | 0.975087 | 0.865666 | 0.527112 | -0.08481 |
| rs10807008_G | BTN3A1 | 0.94089 | 1.033981 | 0.857412 | 1 | -0.06093 |
| rs12200782_C | BTN3A1 | 0.987553 | 1.090084 | 0.896183 | 1 | -0.01253 |
| rs12207930_C | BTN3A1 | 0.994811 | 1.081946 | 0.91566 | 1 | -0.0052 |
| rs12208447_C | BTN3A1 | 0.971864 | 1.246094 | 0.767755 | 1 | -0.02854 |
| rs12214924_T | BTN3A1 | 0.997777 | 1.08479 | 0.918712 | 1 | -0.00223 |
| rs143476765_A | BTN3A1 | 0.541997 | 2.391824 | 0.175983 | 1 | -0.61249 |
| rs144114619_T | BTN3A1 | 0.937876 | 1.708275 | 0.552209 | 1 | -0.06414 |
| rs145059723_A | BTN3A1 | 0.851224 | 2.534279 | 0.356178 | 1 | -0.16108 |
| rs1741738_A | BTN3A1 | 1.055187 | 1.152999 | 0.966854 | 1 | 0.053718 |
| rs17610161_G | BTN3A1 | 0.948685 | 1.04337 | 0.863865 | 1 | -0.05268 |
| rs1796520_C | BTN3A1 | 0.925332 | 0.980491 | 0.873166 | 0.877114 | -0.0776 |
| rs3799378_A | BTN3A1 | 0.866517 | 0.922476 | 0.814111 | 0.000704 | -0.14327 |
| rs3857549_C | BTN3A1 | 1.20727 | 1.366146 | 1.0703 | 0.249929 | 0.188361 |
| rs3902051_A | BTN3A1 | 0.947893 | 1.038919 | 0.865991 | 1 | -0.05351 |
| rs41266839_G | BTN3A1 | 0.806793 | 0.868508 | 0.749711 | 1.06E-06 | -0.21469 |
| rs4609015_T | BTN3A1 | 0.999771 | 1.087084 | 0.920447 | 1 | -0.00023 |
| rs4712990_C | BTN3A1 | 0.953022 | 1.047148 | 0.8686 | 1 | -0.04812 |
| rs55676749_T | BTN3A1 | 1.055136 | 1.295119 | 0.867029 | 1 | 0.05367 |
| rs56161420_G | BTN3A1 | 1.003319 | 1.090026 | 0.92446 | 1 | 0.003313 |
| rs6900725_T | BTN3A1 | 0.998519 | 1.085343 | 0.919613 | 1 | -0.00148 |
| rs6912853_C | BTN3A1 | 1.060948 | 1.150125 | 0.979684 | 1 | 0.059162 |
| rs6920986_C | BTN3A1 | 0.997232 | 1.08425 | 0.918167 | 1 | -0.00277 |
| rs6921148_T | BTN3A1 | 1.075901 | 1.317605 | 0.885986 | 1 | 0.073159 |
| rs742090_A | BTN3A1 | 0.927391 | 0.982792 | 0.875004 | 1 | -0.07538 |
| rs7770214_G | BTN3A1 | 0.992737 | 1.079363 | 0.914026 | 1 | -0.00729 |
| rs80153343_G | BTN3A1 | 1.144379 | 1.465945 | 0.904679 | 1 | 0.134862 |
| rs11758089_T | BTN3A2 | 1.093163 | 1.188249 | 1.00675 | 1 | 0.089075 |
| rs12176317_A | BTN3A2 | 0.820862 | 0.880647 | 0.765377 | 3.50E-06 | -0.1974 |
| rs12194095_C | BTN3A2 | 0.971352 | 1.079478 | 0.875824 | 1 | -0.02907 |
| rs12199613_C | BTN3A2 | 0.884297 | 0.93714 | 0.834446 | 0.003312 | -0.12296 |
| rs12205731_G | BTN3A2 | 0.970415 | 1.079025 | 0.874528 | 1 | -0.03003 |
| rs144016445_G | BTN3A2 | 38750.66 | NA | 151.7424 | 1 | 10.5649 |
| rs1977_A | BTN3A2 | 0.816781 | 0.87678 | 0.761121 | 2.06E-06 | -0.20238 |
| rs1979_G | BTN3A2 | 0.820729 | 0.880499 | 0.765255 | 3.40E-06 | -0.19756 |
| rs1985732_A | BTN3A2 | 0.896055 | 0.951463 | 0.84398 | 0.033488 | -0.10975 |
| rs2073526_G | BTN3A2 | 0.925312 | 0.980986 | 0.872648 | 0.940603 | -0.07762 |
| rs35183513_G | BTN3A2 | 0.951537 | 1.044843 | 0.867784 | 1 | -0.04968 |
| rs58367598_T | BTN3A2 | 1.089293 | 1.290835 | 0.924221 | 1 | 0.085529 |
| rs7765566_G | BTN3A2 | 1.145085 | 1.363507 | 0.967785 | 1 | 0.135479 |
| rs9104_G | BTN3A2 | 0.945089 | 1.032973 | 0.86575 | 1 | -0.05648 |
| rs9358934_G | BTN3A2 | 0.824601 | 0.884767 | 0.76877 | 7.53E-06 | -0.19286 |
| rs9379855_T | BTN3A2 | 0.82361 | 0.883636 | 0.767905 | 6.04E-06 | -0.19406 |
| rs9379858_T | BTN3A2 | 0.825671 | 0.885867 | 0.769811 | 8.99E-06 | -0.19156 |
| rs9379859_C | BTN3A2 | 0.824802 | 0.885058 | 0.768893 | 8.10E-06 | -0.19261 |
| rs9379861_G | BTN3A2 | 1.039521 | 1.399242 | 0.785634 | 1 | 0.03876 |
| rs9393713_G | BTN3A2 | 0.814157 | 0.873453 | 0.759123 | 9.27E-07 | -0.2056 |
| rs9393714_G | BTN3A2 | 0.818022 | 0.877663 | 0.762671 | 2.08E-06 | -0.20087 |
| rs186813312_C | BTNL3 | 0.387143 | NA | NA | NA | -0.94896 |
| rs199970076_G | BTNL3 | 0.890096 | 19.02105 | 0.105649 | 1 | -0.11643 |
| rs201534771_G | BTNL3 | 0.373628 | NA | NA | NA | -0.98449 |
| rs201813197_C | BTNL3 | 0.906025 | 3.98319 | 0.295074 | 1 | -0.09869 |
| rs35157246_C | BTNL3 | 1.061269 | 1.243749 | 0.909647 | 1 | 0.059466 |
| rs4700774_G | BTNL3 | 0.953121 | 1.014114 | 0.896074 | 1 | -0.04801 |
| rs59220426_C | BTNL3 | 0.964396 | 1.094724 | 0.852069 | 1 | -0.03625 |
| rs73815153_G | BTNL3 | 0.979858 | 1.113514 | 0.864825 | 1 | -0.02035 |
| rs7713324_A | BTNL3 | 0.962434 | 1.092563 | 0.850278 | 1 | -0.03829 |
| rs7726604_C | BTNL3 | 0.963325 | 1.093541 | 0.851096 | 1 | -0.03736 |
| rs112469887_G | BTNL8 | 1.04641 | 1.299262 | 0.850631 | 1 | 0.045365 |
| rs113071395_G | BTNL8 | 0.908391 | 1.065436 | 0.777899 | 1 | -0.09608 |
| rs113534626_A | BTNL8 | 1.008072 | 1.204841 | 0.848563 | 1 | 0.00804 |
| rs141492316_T | BTNL8 | 0.930335 | 1.160408 | 0.752616 | 1 | -0.07221 |
| rs145199317_A | BTNL8 | 0.884686 | 1.250181 | 0.639686 | 1 | -0.12252 |
| rs151174174_C | BTNL8 | 0.828069 | 1.016412 | 0.679378 | 1 | -0.18866 |
| rs17704291_C | BTNL8 | 0.952291 | 1.013025 | 0.895481 | 1 | -0.04888 |
| rs200633883_C | BTNL8 | 0.419543 | 1.675223 | 0.123618 | 1 | -0.86859 |
| rs201214790_T | BTNL8 | 740.6942 | NA | 1.97E-08 | 1 | 6.607588 |
| rs201891387_G | BTNL8 | 0.495896 | 2.269513 | 0.147322 | 1 | -0.70139 |
| rs2276995_A | BTNL8 | 0.984754 | 1.043162 | 0.929755 | 1 | -0.01536 |
| rs2619739_C | BTNL8 | 1.078254 | 1.194765 | 0.974928 | 1 | 0.075343 |
| rs7724813_G | BTNL8 | 1.054983 | 1.150319 | 0.968751 | 1 | 0.053525 |
| SNP name | gene | OR | upper | lower | adjusted p-value | ln(OR) |
| rs10484441_G | BTN2A1 | 1.026879 | 1.182655 | 0.894513 | 1 | 0.026524 |
| rs12660069_C | BTN2A1 | 0.954382 | 1.207579 | 0.761803 | 1 | -0.04669 |
| rs13195402_G | BTN2A1 | 0.757206 | 0.831328 | 0.689864 | 5.10E-07 | -0.27812 |
| rs13195509_G | BTN2A1 | 0.77459 | 0.846648 | 0.708837 | 1.75E-06 | -0.25542 |
| rs13437351_G | BTN2A1 | 1.400742 | 2.090478 | 0.973921 | 1 | 0.337002 |
| rs1407045_A | BTN2A1 | 1.080193 | 1.170756 | 0.996977 | 1 | 0.07714 |
| rs142951857_A | BTN2A1 | 1.287748 | 4.431929 | 0.486536 | 1 | 0.252895 |
| rs143104579_G | BTN2A1 | 1.331616 | 1.885897 | 0.963027 | 1 | 0.286393 |
| rs146399224_T | BTN2A1 | 0.25741 | NA | NA | NA | -1.35708 |
| rs148111655_G | BTN2A1 | 0.944297 | 4.178925 | 0.294481 | 1 | -0.05731 |
| rs2273558_A | BTN2A1 | 0.892885 | 0.971388 | 0.820718 | 0.848901 | -0.1133 |
| rs2893856_T | BTN2A1 | 0.927404 | 1.048809 | 0.818043 | 1 | -0.07537 |
| rs2893857_C | BTN2A1 | 1.040565 | 1.199279 | 0.905854 | 1 | 0.039764 |
| rs3734539_C | BTN2A1 | 27166.76 | NA | 9.99E-14 | 1 | 10.20975 |
| rs3734542_G | BTN2A1 | 0.77697 | 0.849181 | 0.711073 | 2.52E-06 | -0.25235 |
| rs3734543_G | BTN2A1 | 0.792317 | 0.867952 | 0.723463 | 5.43E-05 | -0.23279 |
| rs3799380_T | BTN2A1 | 0.869548 | 0.945573 | 0.799787 | 0.107572 | -0.13978 |
| rs56296968_C | BTN2A1 | 0.852755 | 0.928301 | 0.783513 | 0.02331 | -0.15928 |
| rs6456724_T | BTN2A1 | 0.924581 | 1.045479 | 0.815654 | 1 | -0.07841 |
| rs6907857_T | BTN2A1 | 1.401132 | 2.091061 | 0.974192 | 1 | 0.33728 |
| rs6911470_C | BTN2A1 | 1.565436 | 2.679723 | 0.978088 | 1 | 0.448164 |
| rs6929846_T | BTN2A1 | 0.870873 | 0.973832 | 0.777255 | 1 | -0.13826 |
| rs7773913_C | BTN2A1 | 1.408409 | 2.10183 | 0.979326 | 1 | 0.34246 |
| rs7773938_C | BTN2A1 | 0.854233 | 0.929778 | 0.784985 | 0.026611 | -0.15755 |
| rs77870445_T | BTN2A1 | 1.022451 | 1.530146 | 0.704151 | 1 | 0.022203 |
| rs9348718_A | BTN2A1 | 1.308813 | 1.636793 | 1.057849 | 1 | 0.26912 |
| rs9358943_C | BTN2A1 | 0.257602 | NA | NA | NA | -1.35634 |
| rs9358944_A | BTN2A1 | 0.850121 | 0.924966 | 0.781482 | 0.016059 | -0.16238 |
| rs9358945_A | BTN2A1 | 0.847905 | 0.922571 | 0.77943 | 0.012607 | -0.16499 |
| rs9461254_G | BTN2A1 | 1.565001 | 2.678973 | 0.977818 | 1 | 0.447886 |
| rs10456045_G | BTN3A1 | 0.872024 | 0.944198 | 0.805371 | 0.074344 | -0.13694 |
| rs10807008_G | BTN3A1 | 0.988264 | 1.128972 | 0.867514 | 1 | -0.01181 |
| rs12200782_C | BTN3A1 | 0.969241 | 1.112166 | 0.84719 | 1 | -0.03124 |
| rs12207930_C | BTN3A1 | 1.059407 | 1.19361 | 0.94228 | 1 | 0.05771 |
| rs12208447_C | BTN3A1 | 1.183033 | 1.71485 | 0.838562 | 1 | 0.168081 |
| rs12214924_T | BTN3A1 | 1.059617 | 1.192843 | 0.943249 | 1 | 0.057908 |
| rs143476765_A | BTN3A1 | 0.385935 | 2.931895 | 0.063895 | 1 | -0.95209 |
| rs144114619_T | BTN3A1 | 0.800306 | 1.80142 | 0.391948 | 1 | -0.22276 |
| rs145059723_A | BTN3A1 | 1.546934 | 29.22579 | 0.264012 | 1 | 0.436275 |
| rs1741738_A | BTN3A1 | 1.126398 | 1.288465 | 0.987689 | 1 | 0.119025 |
| rs17610161_G | BTN3A1 | 0.999336 | 1.142965 | 0.876283 | 1 | -0.00066 |
| rs1796520_C | BTN3A1 | 0.901974 | 0.977599 | 0.831877 | 1 | -0.10317 |
| rs3799378_A | BTN3A1 | 0.829346 | 0.900397 | 0.763968 | 0.00081 | -0.18712 |
| rs3857549_C | BTN3A1 | 1.307821 | 1.557482 | 1.105147 | 0.217446 | 0.268362 |
| rs3902051_A | BTN3A1 | 0.979611 | 1.113404 | 0.864043 | 1 | -0.0206 |
| rs41266839_G | BTN3A1 | 0.753767 | 0.824792 | 0.68903 | 7.25E-08 | -0.28267 |
| rs4609015_T | BTN3A1 | 1.055122 | 1.187787 | 0.939241 | 1 | 0.053657 |
| rs4712990_C | BTN3A1 | 1.009758 | 1.153906 | 0.886126 | 1 | 0.00971 |
| rs55676749_T | BTN3A1 | 1.356015 | 1.867802 | 1.007293 | 1 | 0.30455 |
| rs56161420_G | BTN3A1 | 1.060354 | 1.193295 | 0.944184 | 1 | 0.058603 |
| rs6900725_T | BTN3A1 | 1.061349 | 1.194313 | 0.945175 | 1 | 0.059541 |
| rs6912853_C | BTN3A1 | 1.087423 | 1.216203 | 0.974151 | 1 | 0.08381 |
| rs6920986_C | BTN3A1 | 1.062734 | 1.196651 | 0.9458 | 1 | 0.060845 |
| rs6921148_T | BTN3A1 | 1.340282 | 1.834522 | 1.000858 | 1 | 0.29288 |
| rs742090_A | BTN3A1 | 0.903554 | 0.979423 | 0.833241 | 1 | -0.10142 |
| rs7770214_G | BTN3A1 | 1.053031 | 1.185369 | 0.937421 | 1 | 0.051673 |
| rs80153343_G | BTN3A1 | 0.979216 | 1.347528 | 0.724804 | 1 | -0.021 |
| rs11758089_T | BTN3A2 | 1.191327 | 1.352007 | 1.052499 | 0.618146 | 0.175068 |
| rs12176317_A | BTN3A2 | 0.766628 | 0.836949 | 0.702371 | 2.82E-07 | -0.26575 |
| rs12194095_C | BTN3A2 | 0.943672 | 1.092386 | 0.818034 | 1 | -0.05798 |
| rs12199613_C | BTN3A2 | 0.842231 | 0.911373 | 0.77819 | 0.002057 | -0.1717 |
| rs12205731_G | BTN3A2 | 0.933998 | 1.081909 | 0.809117 | 1 | -0.06828 |
| rs144016445_G | BTN3A2 | 73914.07 | NA | 1.75E-06 | 1 | 11.21066 |
| rs1977_A | BTN3A2 | 0.764904 | 0.835801 | 0.700168 | 2.99E-07 | -0.268 |
| rs1979_G | BTN3A2 | 0.76816 | 0.838582 | 0.70381 | 3.63E-07 | -0.26376 |
| rs1985732_A | BTN3A2 | 0.860169 | 0.932012 | 0.793857 | 0.023502 | -0.15063 |
| rs2073526_G | BTN3A2 | 0.90252 | 0.979048 | 0.831612 | 1 | -0.10256 |
| rs35183513_G | BTN3A2 | 1.000653 | 1.140268 | 0.880486 | 1 | 0.000652 |
| rs58367598_T | BTN3A2 | 1.153984 | 1.470445 | 0.915197 | 1 | 0.143221 |
| rs7765566_G | BTN3A2 | 1.171754 | 1.497136 | 0.928192 | 1 | 0.158502 |
| rs9104_G | BTN3A2 | 1.010674 | 1.145196 | 0.894134 | 1 | 0.010617 |
| rs9358934_G | BTN3A2 | 0.772404 | 0.843549 | 0.707423 | 8.85E-07 | -0.25825 |
| rs9379855_T | BTN3A2 | 0.771306 | 0.842174 | 0.706564 | 6.76E-07 | -0.25967 |
| rs9379858_T | BTN3A2 | 0.774407 | 0.845625 | 0.709352 | 1.18E-06 | -0.25566 |
| rs9379859_C | BTN3A2 | 0.770242 | 0.841254 | 0.705384 | 6.31E-07 | -0.26105 |
| rs9379861_G | BTN3A2 | 0.987622 | 1.586757 | 0.639198 | 1 | -0.01246 |
| rs9393713_G | BTN3A2 | 0.760996 | 0.830868 | 0.697149 | 1.06E-07 | -0.27313 |
| rs9393714_G | BTN3A2 | 0.765074 | 0.835356 | 0.700859 | 2.25E-07 | -0.26778 |
| rs186813312_C | BTNL3 | 0.257602 | NA | NA | NA | -1.35634 |
| rs199970076_G | BTNL3 | 0.272926 | 6.904492 | 0.010788 | 1 | -1.29856 |
| rs201534771_G | BTNL3 | 0.273671 | NA | NA | NA | -1.29583 |
| rs201813197_C | BTNL3 | 0.514697 | 3.715301 | 0.100372 | 1 | -0.66418 |
| rs35157246_C | BTNL3 | 1.03751 | 1.287524 | 0.842551 | 1 | 0.036824 |
| rs4700774_G | BTNL3 | 0.978837 | 1.065525 | 0.899706 | 1 | -0.02139 |
| rs59220426_C | BTNL3 | 1.017762 | 1.216405 | 0.856252 | 1 | 0.017606 |
| rs73815153_G | BTNL3 | 1.02474 | 1.225825 | 0.861442 | 1 | 0.024438 |
| rs7713324_A | BTNL3 | 1.018119 | 1.216841 | 0.856546 | 1 | 0.017957 |
| rs7726604_C | BTNL3 | 1.020763 | 1.219953 | 0.85881 | 1 | 0.02055 |
| rs112469887_G | BTNL8 | 1.002918 | 1.333287 | 0.765187 | 1 | 0.002914 |
| rs113071395_G | BTNL8 | 0.93155 | 1.162634 | 0.752338 | 1 | -0.07091 |
| rs113534626_A | BTNL8 | 0.9371 | 1.185234 | 0.747845 | 1 | -0.06496 |
| rs141492316_T | BTNL8 | 0.963798 | 1.314391 | 0.718601 | 1 | -0.03687 |
| rs145199317_A | BTNL8 | 1.12544 | 1.912393 | 0.697448 | 1 | 0.118174 |
| rs151174174_C | BTNL8 | 0.729764 | 0.953457 | 0.564223 | 1 | -0.31503 |
| rs17704291_C | BTNL8 | 0.969451 | 1.055155 | 0.891212 | 1 | -0.03103 |
| rs200633883_C | BTNL8 | 0.171372 | 1.035119 | 0.022558 | 1 | -1.76392 |
| rs201214790_T | BTNL8 | 0.256248 | NA | NA | NA | -1.36161 |
| rs201891387_G | BTNL8 | 0.171476 | 1.035747 | 0.022572 | 1 | -1.76331 |
| rs2276995_A | BTNL8 | 1.002057 | 1.084332 | 0.926283 | 1 | 0.002055 |
| rs2619739_C | BTNL8 | 0.966968 | 1.107812 | 0.846435 | 1 | -0.03359 |
| rs7724813_G | BTNL8 | 1.041603 | 1.171988 | 0.927725 | 1 | 0.040761 |
| Participants with the HLA-DQ2.5 genotype | |||||||
| SNP, reference allele | Gene | Number of the SNP in controls with HLA-DQ2.5 | Number of the SNP in CeD with HLA-DQ2.5 | Total allele count in control with HLA-DQ2.5 | Total allele count in CeD with HLA-DQ2.5 | Total number of the SNP in the UK Biobank with HLA-DQ2.5 | Total allele count in UK Biobank with HLA-DQ2.5 |
| rs13195402_G | BTN2A1 | 9398 | 2256 | 12514 | 3206 | 11654 | 15720 |
| rs13195509_G | BTN2A1 | 9408 | 2265 | 12820 | 3296 | 11673 | 16116 |
| rs3734542_G | BTN2A1 | 9386 | 2266 | 12808 | 3300 | 11652 | 16108 |
| rs3734543_G | BTN2A1 | 9366 | 2265 | 12722 | 3258 | 11631 | 15980 |
| rs56296968_C | BTN2A1 | 8609 | 2107 | 12798 | 3290 | 10716 | 16088 |
| rs7773938_C | BTN2A1 | 8606 | 2106 | 12804 | 3290 | 10712 | 16094 |
| rs9358944_A | BTN2A1 | 8603 | 2108 | 12818 | 3304 | 10711 | 16122 |
| rs9358945_A | BTN2A1 | 8607 | 2106 | 12818 | 3302 | 10713 | 16120 |
| rs3799378_A | BTN3A1 | 8129 | 1959 | 12768 | 3286 | 10088 | 16054 |
| rs41266839_G | BTN3A1 | 9577 | 2298 | 12820 | 3298 | 11875 | 16118 |
| rs12176317_A | BTN3A2 | 9347 | 2242 | 12826 | 3302 | 11589 | 16128 |
| rs12199613_C | BTN3A2 | 6396 | 1515 | 12802 | 3300 | 7911 | 16102 |
| rs1977_A | BTN3A2 | 9135 | 2197 | 12574 | 3248 | 11332 | 15822 |
| rs1979_G | BTN3A2 | 9330 | 2242 | 12808 | 3302 | 11572 | 16110 |
| rs1985732_A | BTN3A2 | 7353 | 1777 | 12816 | 3294 | 9130 | 16110 |
| rs9358934_G | BTN3A2 | 9333 | 2240 | 12810 | 3292 | 11573 | 16102 |
| rs9379855_T | BTN3A2 | 9324 | 2239 | 12802 | 3294 | 11563 | 16096 |
| rs9379858_T | BTN3A2 | 9321 | 2242 | 12804 | 3296 | 11563 | 16100 |
| rs9379859_C | BTN3A2 | 9340 | 2244 | 12806 | 3296 | 11584 | 16102 |
| rs9393713_G | BTN3A2 | 9345 | 2237 | 12812 | 3298 | 11582 | 16110 |
| rs9393714_G | BTN3A2 | 9346 | 2241 | 12818 | 3300 | 11587 | 16118 |
Appendix D – Supplementary Materials for Results Section 2.3
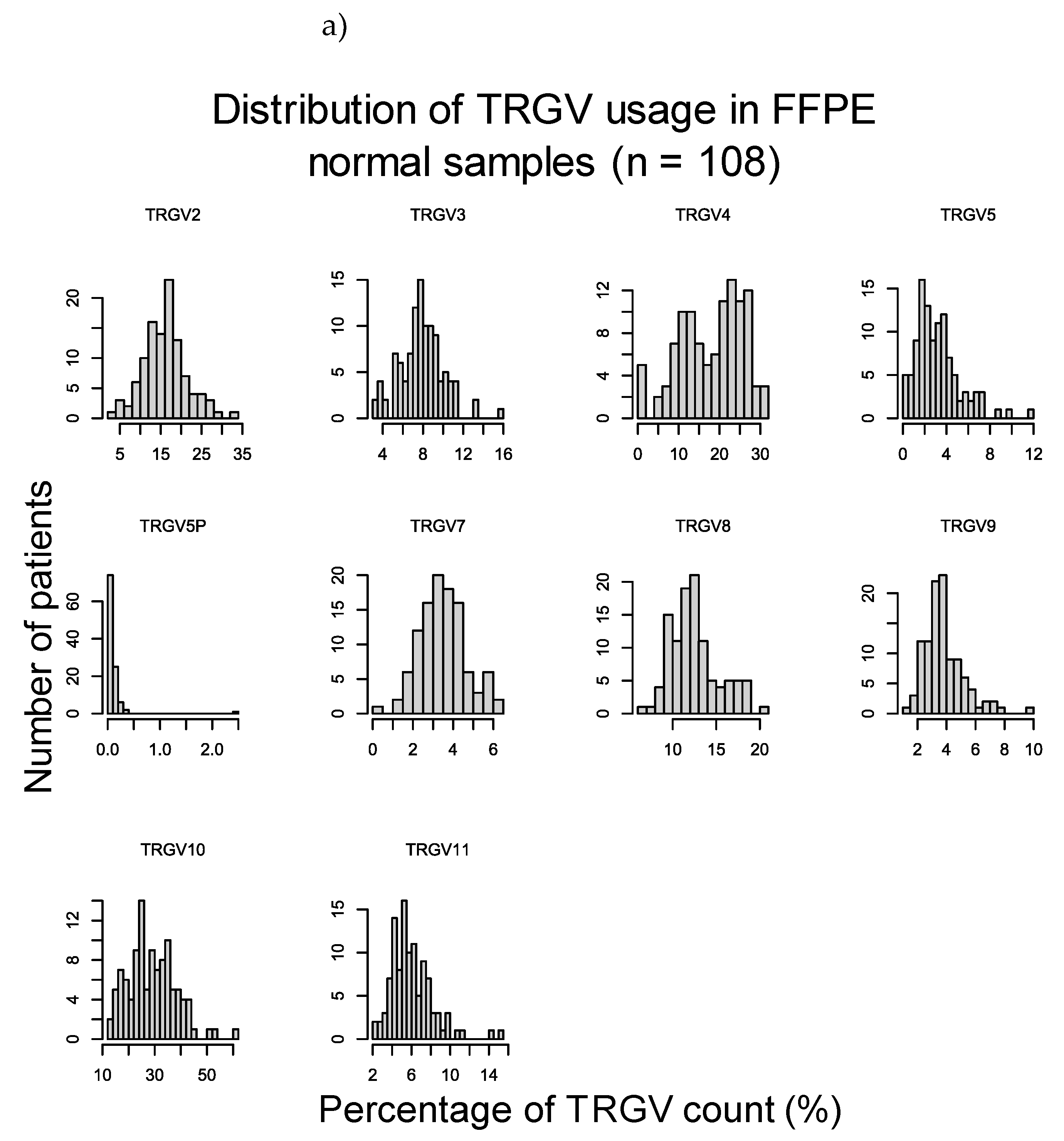
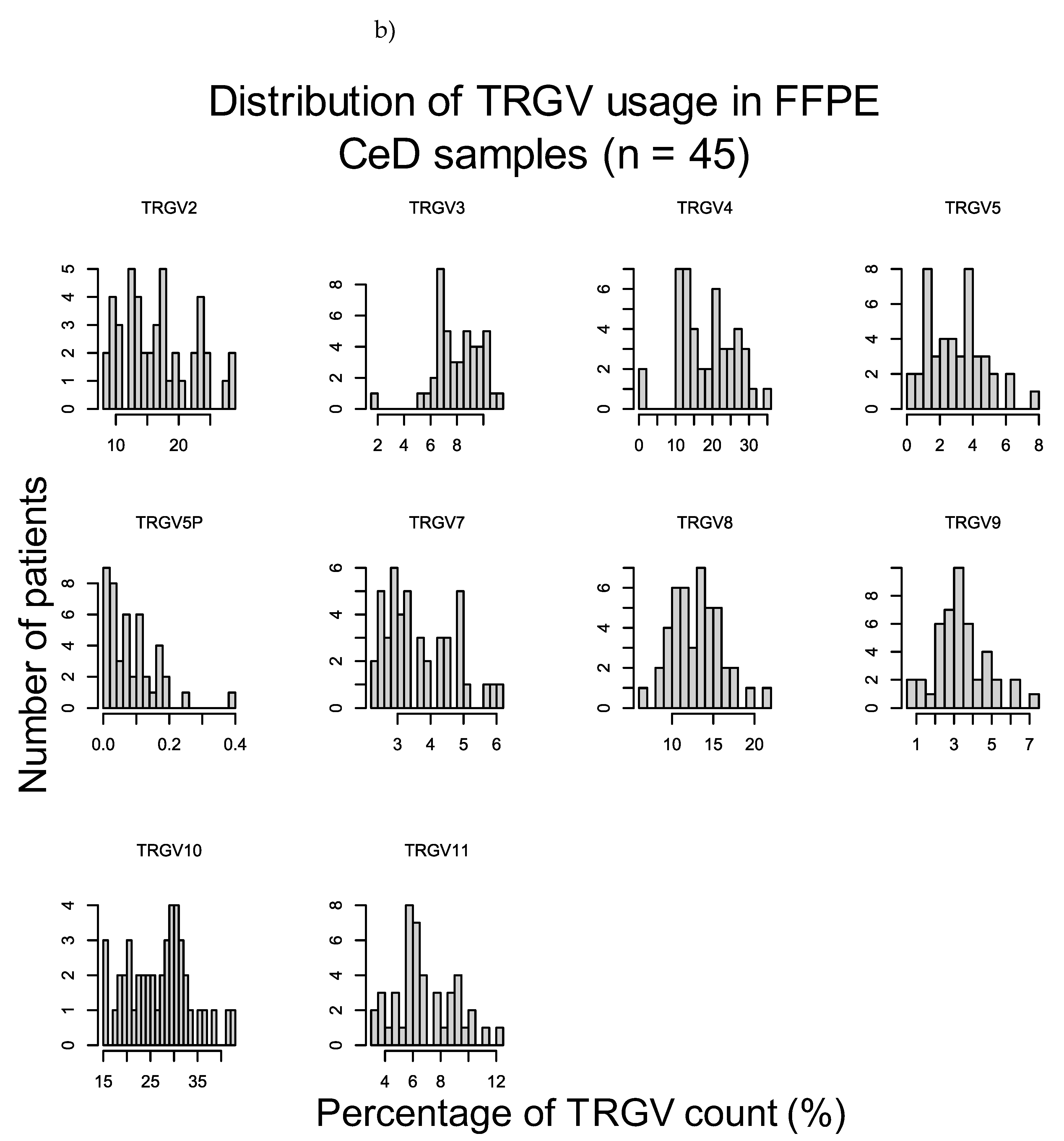
| FFPE CeD (n = 45) vs FFPE healthy control (n = 108) | ||
| Raw p-value (MWU) | Adjusted p-value | |
| TRGV2 | 0.775 | 1 |
| TRGV3 | 0.411 | 1 |
| TRGV4 | 0.812 | 1 |
| TRGV5 | 0.906 | 1 |
| TRGV5P | 0.684 | 1 |
| TRGV7 | 0.566 | 1 |
| TRGV8 | 0.382 | 1 |
| TRGV9 | 0.070 | 0.70 |
| TRGV10 | 0.248 | 1 |
| TRGV11 | 0.025 | 0.25 |
| 141 CeD vs 238 healthy control samples | ||
| Raw p-values (Fisher) | Adjusted p-values | |
| WT vs WT, KYDTYGSTRQNLRMIL | 0.3925 | 1 |
| WT vs KYDTYGSTRQNLRMILR | 0.3392 | 1 |
| WT vs KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA | 0.3668 | 1 |
| WT vs WT, KYNTYGSTRKNLRMILR | 0.3668 | 1 |
| WT vs KYDTYGNTRKNLRMILR, WT | 1 | 1 |
| WT vs WT, KYDTYGSTRKSLRMILR | 0.6258 | 1 |
| WT vs KYDTYGSTRKSLRMILR | 1 | 1 |
| WT vs WT, KYDTYGSIRKNLRMILR | 0.3668 | 1 |
| WT, KYDTYGSTRQNLRMILR vs KYDTYGSTRQNLRMILR | 0.1697 | 1 |
| WT, KYDTYGSTRQNLRMILR vs KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA | 0.4524 | 1 |
| WT, KYDTYGSTRQNLRMILR vs WT, KYNTYGSTRKNLRMILR | 0.4524 | 1 |
| WT, KYDTYGSTRQNLRMILR vs WT, KYDTYGNTRKNLRMILR | 1 | 1 |
| WT, KYDTYGSTRQNLRMILR vs WT, KYDTYGSTRKSLRMILR | 1 | 1 |
| WT, KYDTYGSTRQNLRMILR vs KYDTYGSTRKSLRMILR | 1 | 1 |
| WT, KYDTYGSTRQNLRMILR vs WT, KYDTYGSIRKNLRMILR | 0.4524 | 1 |
| KYDTYGSTRQNLRMILR vs KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA | 0.25 | 1 |
| KYDTYGSTRQNLRMILR vs WT, KYNTYGSTRKNLRMILR | 0.25 | 1 |
| KYDTYGSTRQNLRMILR vs WT, KYDTYGNTRKNLRMILR | 1 | 1 |
| KYDTYGSTRQNLRMILR vs WT, KYDTYGSTRKSLRMILR | 0.5165 | 1 |
| KYDTYGSTRQNLRMILR vs KYDTYGSTRKSLRMILR | 1 | 1 |
| KYDTYGSTRQNLRMILR vs WT, KYDTYGSIRKNLRMILR | 0.25 | 1 |
| KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA vs WT, KYNTYGSTRKNLRMILR | 1 | 1 |
| KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA vs WT, KYDTYGNTRKNLRMILR | 1 | 1 |
| KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA vs WT, KYDTYGSTRKSLRMILR | 1 | 1 |
| KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA vs KYDTYGSTRKSLRMILR | 1 | 1 |
| KYDTYGSTRQNLRMILR, KYDTYGSTR_ELENDTA vs WT, KYDTYGSIRKNLRMILR | 1 | 1 |
| WT, KYNTYGSTRKNLRMILR vs WT, KYDTYGNTRKNLRMILR | 1 | 1 |
| WT, KYNTYGSTRKNLRMILR vs WT, KYDTYGSTRKSLRMILR | 1 | 1 |
| WT, KYNTYGSTRKNLRMILR vs KYDTYGSTRKSLRMILR | 1 | 1 |
| WT, KYNTYGSTRKNLRMILR vs WT, KYDTYGSIRKNLRMILR | 1 | 1 |
| WT, KYDTYGNTRKNLRMILR vs WT, KYDTYGSTRKSLRMILR | 1 | 1 |
| WT, KYDTYGNTRKNLRMILR vs KYDTYGSTRKSLRMILR | 1 | 1 |
| WT, KYDTYGNTRKNLRMILR vs WT, KYDTYGSIRKNLRMILR | 1 | 1 |
| WT, KYDTYGSTRKSLRMILR vs KYDTYGSTRKSLRMILR | 1 | 1 |
| WT, KYDTYGSTRKSLRMILR vs WT, KYDTYGSIRKNLRMILR | 1 | 1 |
| KYDTYGSTRKSLRMILR vs WT, KYDTYGSIRKNLRMILR | 1 | 1 |
Appendix E – Selecting and Sequencing the Butyrophilin Genes of Interest
Appendix E.1: HPA Expression Profiles of Butyrophilin Family Genes
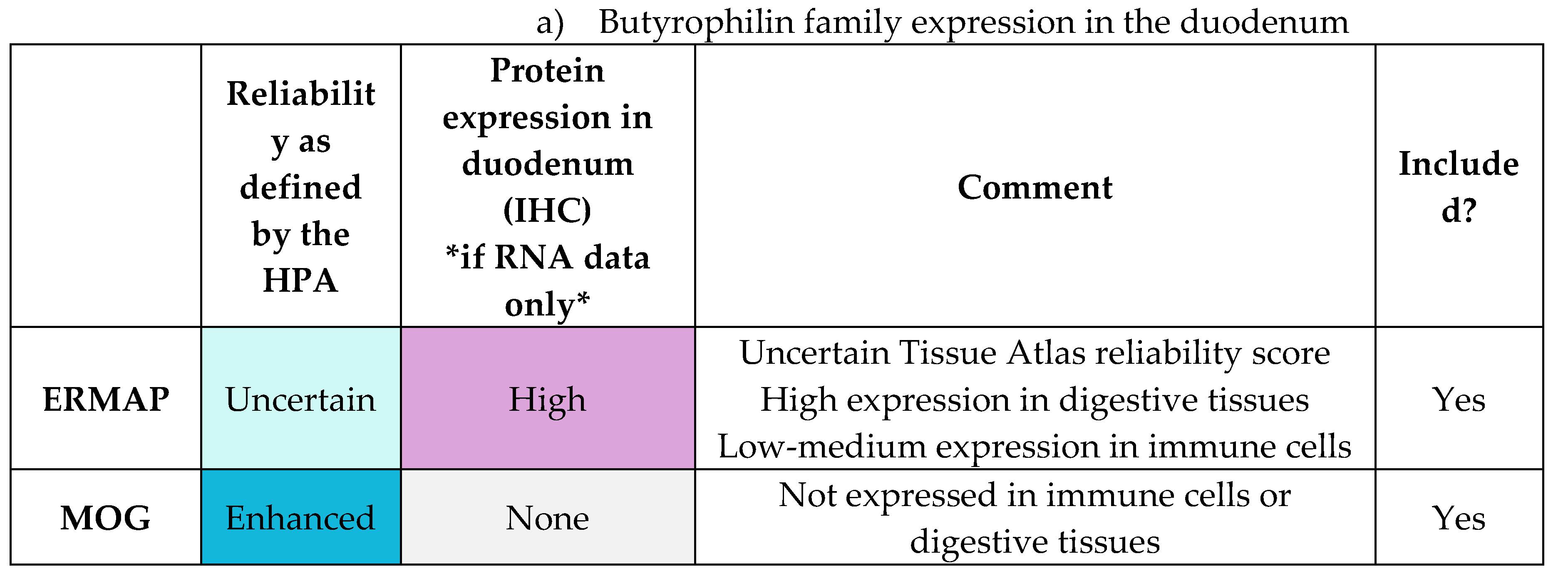 |
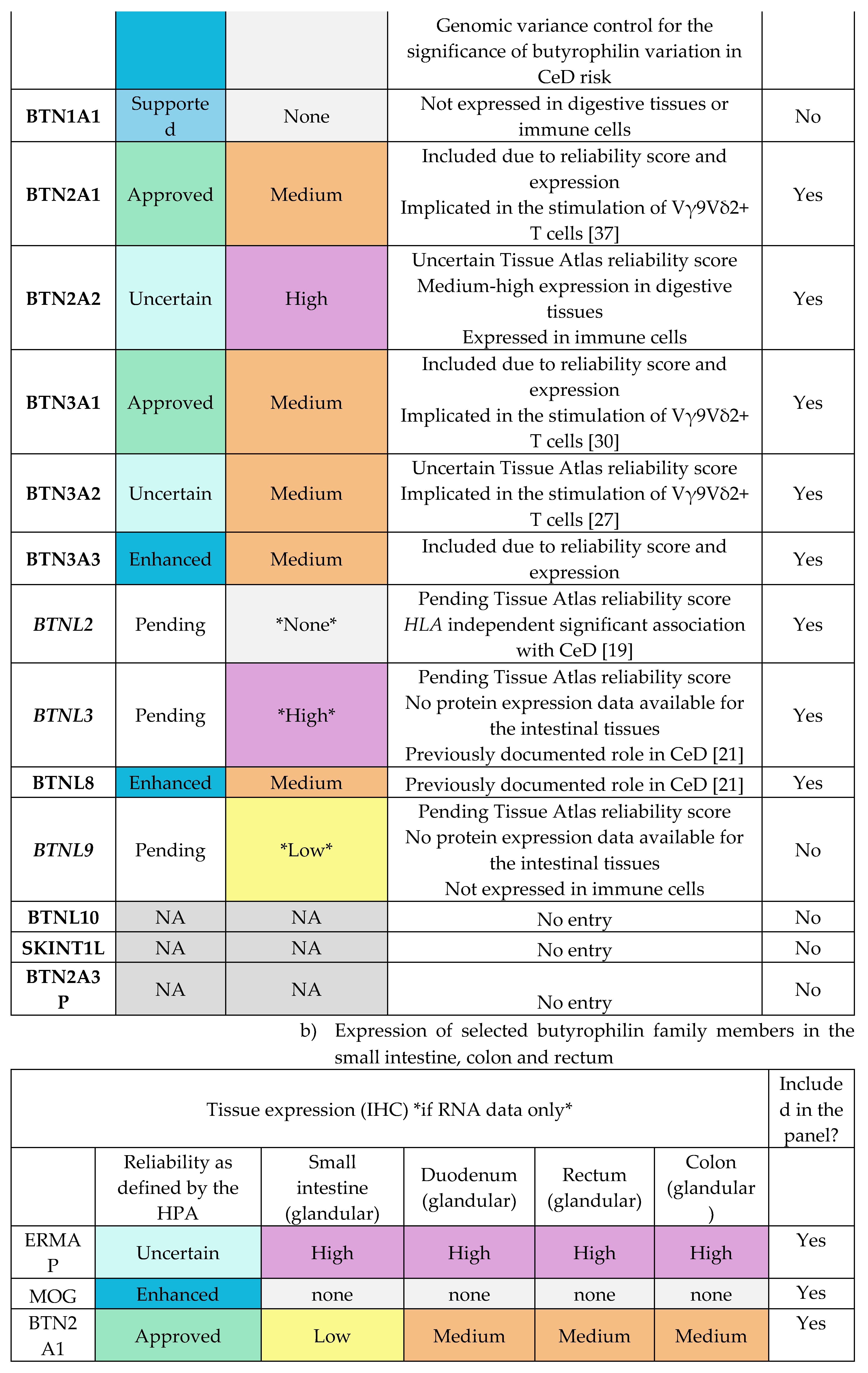 |
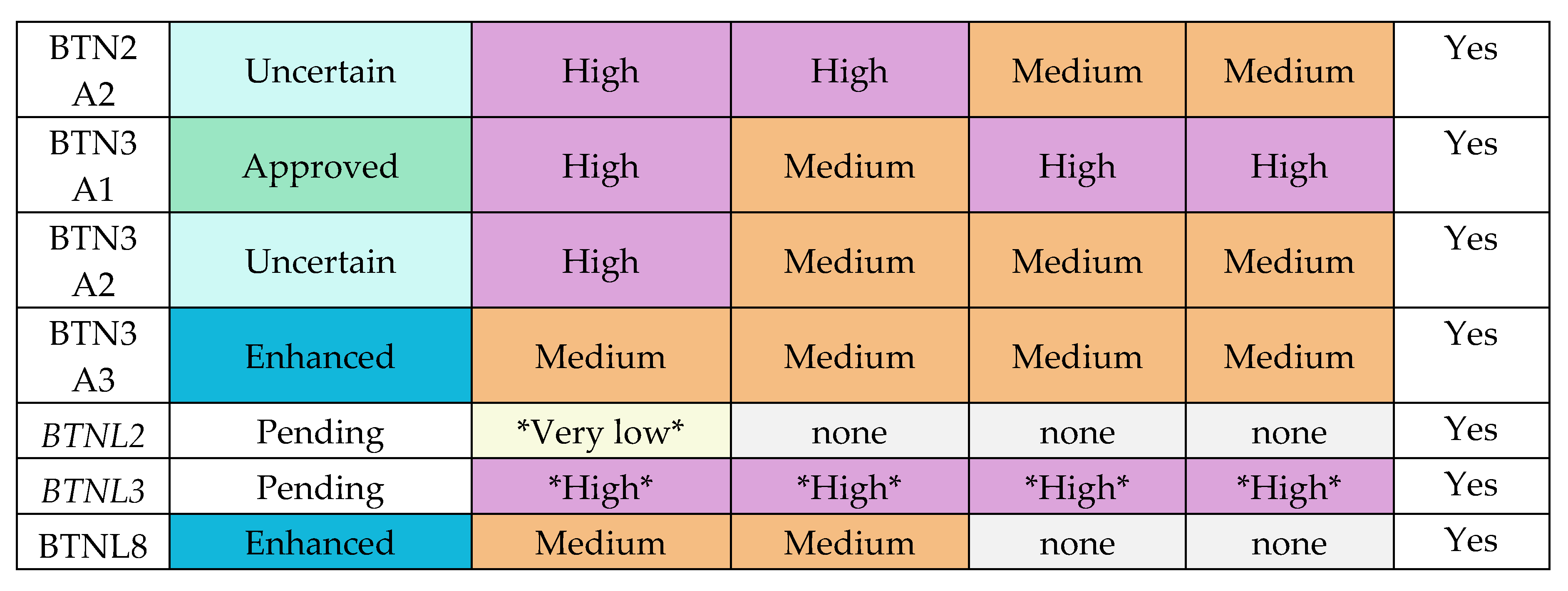 |
| Immune cell expression (RNA sequencing) | Included in the panel? | |||||||
| Reliability as defined by the HPA | γδ T cells | T cells | T-reg | DCs | Macrophages | NK cells | ||
| ERMAP | Uncertain | Low | Medium | Medium | Medium | Medium | Low | Yes |
| MOG | Enhanced | Very low | None | Very low | None | None | None | Yes |
| BTN2A1 | Approved | Medium | Medium | Medium | Medium | High | Low | Yes |
| BTN2A2 | Uncertain | Low | Medium | Medium | High | High | Medium | Yes |
| BTN3A1 | Approved | High | High | High | Low | Medium | High | Yes |
| BTN3A2 | Uncertain | High | High | High | Medium | High | High | Yes |
| BTN3A3 | Enhanced | High | High | High | Medium | Medium | High | Yes |
| BTNL2 | Pending | None | None | None | None | None | None | Yes |
| BTNL3 | Pending | None | None | None | None | None | None | Yes |
| BTNL8 | Enhanced | None | None | None | None | None | None | Yes |
Appendix E.2: Modified Nonacus Cell3 Hybridization Capture and Illumina Sequencing
Appendix E.3: Measuring DNA Quantity and Fragment Size
Appendix F – Detailed Germline Short Variant Discovery Protocol
Appendix F.1: Per Sample Preprocesses and Variant call Using GATK
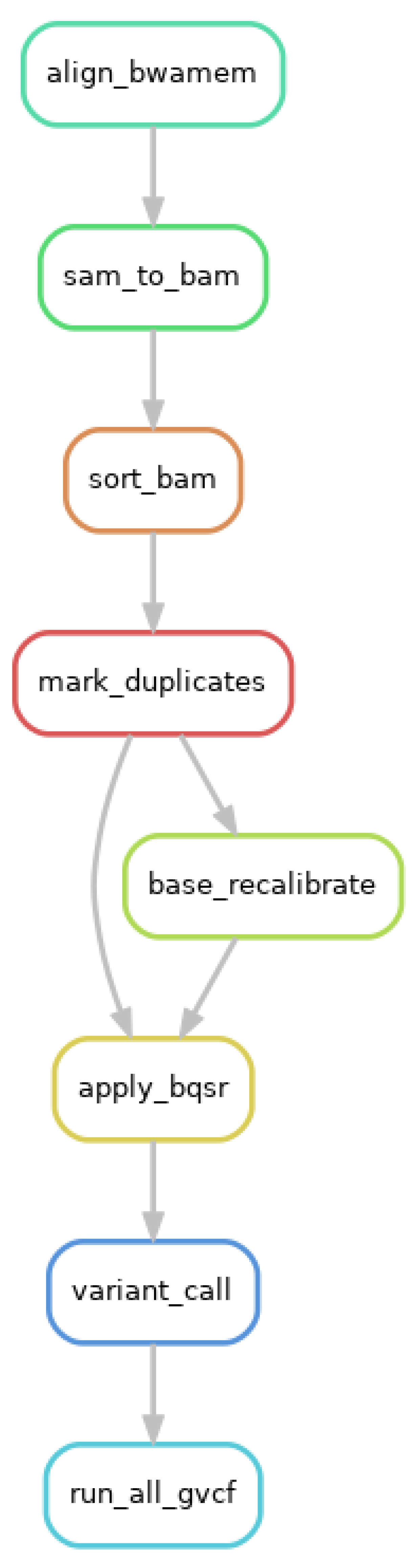
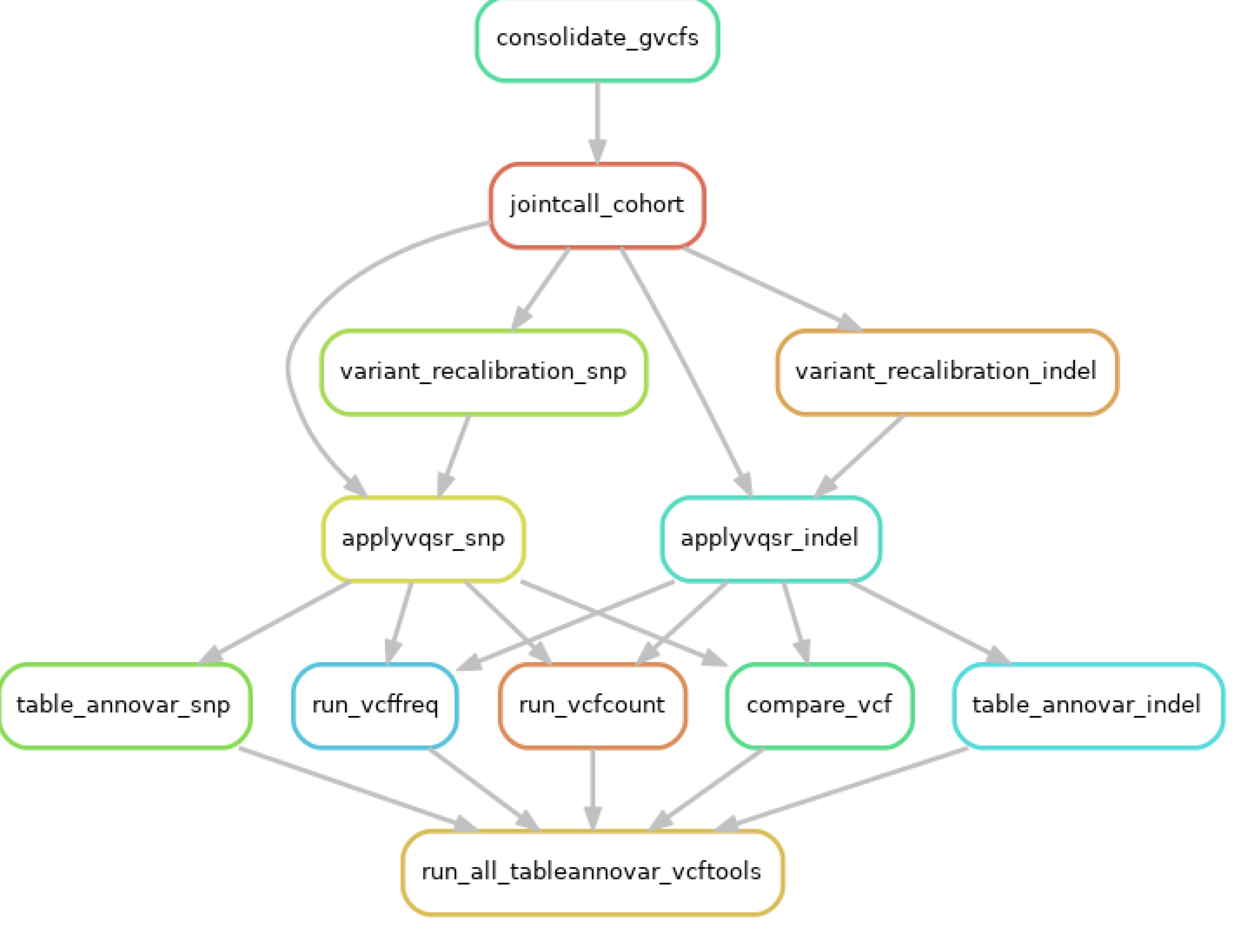
Appendix F.2: Joint Genotyping Using GATK and Variant Annotation Using Vcftools and ANNOVAR
References
- Abadie, V.; Sollid, L.M.; Barreiro, L.B.; Jabri, B. Integration of genetic and immunological insights into a model of celiac disease pathogenesis. Annual Review of Immunology 2011, 29, 493-525. [CrossRef]
- Jabri, B.; Sollid, L.M. Tissue-mediated control of immunopathology in coeliac disease. Nat Rev Immunol 2009, 9, 858-870. [CrossRef]
- Trier, J.S. Diagnosis of celiac sprue. Gastroenterology 1998, 115, 211-216. [CrossRef]
- NICE. Coeliac disease: recognition, assessment and management. Available online: https://www.nice.org.uk/guidance/ng20/chapter/Recommendations (accessed on 27th November 2020).
- Al-Toma, A.; Goerres, M.S.; Meijer, J.W.; Pena, A.S.; Crusius, J.B.; Mulder, C.J. Human leukocyte antigen-DQ2 homozygosity and the development of refractory celiac disease and enteropathy-associated T-cell lymphoma. Clin Gastroenterol Hepatol 2006, 4, 315-319. [CrossRef]
- Ayesh, B.M.; Zaqout, E.K.; Yassin, M.M. HLA-DQ2 and -DQ8 haplotypes frequency and diagnostic utility in celiac disease patients of Gaza strip, Palestine. Autoimmun Highlights 2017, 8, doi:ARTN 1110.1007/s13317-017-0099-0.
- Björck, S.; Brundin, C.; Lörinc, E.; Lynch, K.F.; Agardh, D. Screening detects a high proportion of celiac disease in young HLA-genotyped children. J Pediatr Gastr Nutr 2010, 50, 49-53. [CrossRef]
- Karell, K.; Louka, A.S.; Moodie, S.J.; Ascher, H.; Clot, F.; Greco, L.; Ciclitira, P.J.; Sollid, L.M.; Partanen, J.; European Genetics Cluster on Celiac, D. HLA types in celiac disease patients not carrying the DQA1*05-DQB1*02 (DQ2) heterodimer: results from the European Genetics Cluster on Celiac Disease. Hum Immunol 2003, 64, 469-477. [CrossRef]
- Karhus, L.L.; Thuesen, B.H.; Skaaby, T.; Rumessen, J.J.; Linneberg, A. The distribution of HLA DQ2 and DQ8 haplotypes and their association with health indicators in a general Danish population. United Eur Gastroent 2018, 6, 866-878. [CrossRef]
- Murad, H.; Jazairi, B.; Khansaa, I.; Olabi, D.; Khouri, L. HLA-DQ2 and -DQ8 genotype frequency in Syrian celiac disease children: HLA-DQ relative risks evaluation. BMC Gastroenterol 2018, 18, 70. [CrossRef]
- Sollid, L.M.; Thorsby, E. HLA susceptibility genes in celiac disease: genetic mapping and role in pathogenesis. Gastroenterology 1993, 105, 910-922. [CrossRef]
- Sollid, L.M. Molecular basis of celiac disease. Annu Rev Immunol 2000, 18, 53-81. [CrossRef]
- Sollid, L.M.; Markussen, G.; Ek, J.; Gjerde, H.; Vartdal, F.; Thorsby, E. Evidence for a primary association of celiac disease to a particular HLA-DQ alpha/beta heterodimer. J Exp Med 1989, 169, 345-350. [CrossRef]
- Sollid, L.M.; Thorsby, E. The primary association of celiac disease to a given HLA-DQ alpha/beta heterodimer explains the divergent HLA-DR associations observed in various Caucasian populations. Tissue Antigens 1990, 36, 136-137. [CrossRef]
- Rubio-Tapia, A.; Hill, I.D.; Kelly, C.P.; Calderwood, A.H.; Murray, J.A.; American College of, G. ACG clinical guidelines: diagnosis and management of celiac disease. Am J Gastroenterol 2013, 108, 656-676; quiz 677. [CrossRef]
- Djilali-Saiah, I.; Caillat-Zucman, S.; Schmitz, J.; Chaves-Vieira, M.L.; Bach, J.F. Polymorphism of antigen processing (TAP, LMP) and HLA class II genes in celiac disease. Hum Immunol 1994, 40, 8-16. [CrossRef]
- Hunt, K.A.; Zhernakova, A.; Turner, G.; Heap, G.A.; Franke, L.; Bruinenberg, M.; Romanos, J.; Dinesen, L.C.; Ryan, A.W.; Panesar, D.; et al. Newly identified genetic risk variants for celiac disease related to the immune response. Nat Genet 2008, 40, 395-402. [CrossRef]
- Dubois, P.C.; van Heel, D.A. Translational mini-review series on the immunogenetics of gut disease: immunogenetics of coeliac disease. Clin Exp Immunol 2008, 153, 162-173. [CrossRef]
- Goudey, B.; Abraham, G.; Kikianty, E.; Wang, Q.; Rawlinson, D.; Shi, F.; Haviv, I.; Stern, L.; Kowalczyk, A.; Inouye, M. Interactions within the MHC contribute to the genetic architecture of celiac disease. PLoS One 2017, 12, e0172826. [CrossRef]
- Pietz, G.; De, R.; Hedberg, M.; Sjoberg, V.; Sandstrom, O.; Hernell, O.; Hammarstrom, S.; Hammarstrom, M.L. Immunopathology of childhood celiac disease-Key role of intestinal epithelial cells. PLoS One 2017, 12, e0185025. [CrossRef]
- Mayassi, T.; Ladell, K.; Gudjonson, H.; McLaren, J.E.; Shaw, D.G.; Tran, M.T.; Rokicka, J.J.; Lawrence, I.; Grenier, J.C.; van Unen, V.; et al. Chronic inflammation permanently reshapes tissue-resident immunity in celiac disease. Cell 2019, 176, 967-981. [CrossRef]
- Rhodes, D.A.; Stammers, M.; Malcherek, G.; Beck, S.; Trowsdale, J. The cluster of BTN genes in the extended major histocompatibility complex. Genomics 2001, 71, 351-362. [CrossRef]
- Arnett, H.A.; Viney, J.L. Immune modulation by butyrophilins. Nat Rev Immunol 2014, 14, 559-569. [CrossRef]
- Rhodes, D.A.; Reith, W.; Trowsdale, J. Regulation of immunity by butyrophilins. Annu Rev Immunol 2016, 34, 151-172. [CrossRef]
- Malcherek, G.; Mayr, L.; Roda-Navarro, P.; Rhodes, D.; Miller, N.; Trowsdale, J. The B7 homolog butyrophilin BTN2A1 is a novel ligand for DC-SIGN. J Immunol 2007, 179, 3804-3811. [CrossRef]
- Messal, N.; Mamessier, E.; Sylvain, A.; Celis-Gutierrez, J.; Thibult, M.L.; Chetaille, B.; Firaguay, G.; Pastor, S.; Guillaume, Y.; Wang, Q.; et al. Differential role for CD277 as a co-regulator of the immune signal in T and NK cells. Eur J Immunol 2011, 41, 3443-3454. [CrossRef]
- Di Marco Barros, R.; Roberts, N.A.; Dart, R.J.; Vantourout, P.; Jandke, A.; Nussbaumer, O.; Deban, L.; Cipolat, S.; Hart, R.; Iannitto, M.L.; et al. Epithelia use butyrophilin-like molecules to shape organ-specific gamma delta T cell compartments. Cell 2016, 167, 203-218. [CrossRef]
- Jandke, A.; Melandri, D.; Monin, L.; Ushakov, D.S.; Laing, A.G.; Vantourout, P.; East, P.; Nitta, T.; Narita, T.; Takayanagi, H.; et al. Butyrophilin-like proteins display combinatorial diversity in selecting and maintaining signature intraepithelial gammadelta T cell compartments. Nat Commun 2020, 11, 3769. [CrossRef]
- Melandri, D.; Zlatareva, I.; Chaleil, R.A.G.; Dart, R.J.; Chancellor, A.; Nussbaumer, O.; Polyakova, O.; Roberts, N.A.; Wesch, D.; Kabelitz, D.; et al. The γδ TCR combines innate immunity with adaptive immunity by utilizing spatially distinct regions for agonist selection and antigen responsiveness. Nat Immunol 2018, 19, 1352-1365. [CrossRef]
- Vantourout, P.; Laing, A.; Woodward, M.J.; Zlatareva, I.; Apolonia, L.; Jones, A.W.; Snijders, A.P.; Malim, M.H.; Hayday, A.C. Heteromeric interactions regulate butyrophilin (BTN) and BTN-like molecules governing gammadelta T cell biology. Proc Natl Acad Sci U S A 2018, 115, 1039-1044. [CrossRef]
- Willcox, C.R.; Vantourout, P.; Salim, M.; Zlatareva, I.; Melandri, D.; Zanardo, L.; George, R.; Kjaer, S.; Jeeves, M.; Mohammed, F.; et al. Butyrophilin-like 3 Directly Binds a Human Vγ4(+) T Cell Receptor Using a Modality Distinct from Clonally-Restricted Antigen. Immunity 2019, 51, 813-825 e814. [CrossRef]
- Lewis, J.M.; Girardi, M.; Roberts, S.J.; Barbee, S.D.; Hayday, A.C.; Tigelaar, R.E. Selection of the cutaneous intraepithelial γδ+ T cell repertoire by a thymic stromal determinant. Nat Immunol 2006, 7, 843-850. [CrossRef]
- Cano, C.E.; Pasero, C.; De Gassart, A.; Kerneur, C.; Gabriac, M.; Fullana, M.; Granarolo, E.; Hoet, R.; Scotet, E.; Rafia, C.; et al. BTN2A1, an immune checkpoint targeting Vγ9Vδ2 T cell cytotoxicity against malignant cells. Cell Rep 2021, 36, 109359. [CrossRef]
- Hayday, A.C.; Vantourout, P. The innate biologies of adaptive antigen receptors. Annu Rev Immunol 2020, 38, 487-510. [CrossRef]
- Karunakaran, M.M.; Gobel, T.W.; Starick, L.; Walter, L.; Herrmann, T. Vγ9 and Vδ2 T cell antigen receptor genes and butyrophilin 3 (BTN3) emerged with placental mammals and are concomitantly preserved in selected species like alpaca (Vicugna pacos). Immunogenetics 2014, 66, 243-254. [CrossRef]
- Fichtner, A.S.; Karunakaran, M.M.; Gu, S.; Boughter, C.T.; Borowska, M.T.; Starick, L.; Nohren, A.; Gobel, T.W.; Adams, E.J.; Herrmann, T. Alpaca (Vicugna pacos), the first nonprimate species with a phosphoantigen-reactive Vγ9Vδ2 T cell subset. Proc Natl Acad Sci U S A 2020, 117, 6697-6707. [CrossRef]
- Rigau, M.; Ostrouska, S.; Fulford, T.S.; Johnson, D.N.; Woods, K.; Ruan, Z.; McWilliam, H.E.G.; Hudson, C.; Tutuka, C.; Wheatley, A.K.; et al. Butyrophilin 2A1 is essential for phosphoantigen reactivity by gammadelta T cells. Science 2020, 367. [CrossRef]
- Sandstrom, A.; Peigne, C.M.; Leger, A.; Crooks, J.E.; Konczak, F.; Gesnel, M.C.; Breathnach, R.; Bonneville, M.; Scotet, E.; Adams, E.J. The intracellular B30.2 domain of butyrophilin 3A1 binds phosphoantigens to mediate activation of human Vγ9Vδ2 T cells. Immunity 2014, 40, 490-500. [CrossRef]
- Hu, W.; Shang, R.; Yang, J.; Chen, C.; Liu, Z.; Liang, G.; He, W.; Luo, G. Skin γδ T tells and their function in wound healing. Front Immunol 2022, 13, 875076. [CrossRef]
- Han, A.; Newell, E.W.; Glanville, J.; Fernandez-Becker, N.; Khosla, C.; Chien, Y.H.; Davis, M.M. Dietary gluten triggers concomitant activation of CD4+ and CD8+ alphabeta T cells and gammadelta T cells in celiac disease. Proc Natl Acad Sci U S A 2013, 110, 13073-13078. [CrossRef]
- Aigner, J.; Villatoro, S.; Rabionet, R.; Roquer, J.; Jimenez-Conde, J.; Marti, E.; Estivill, X. A common 56-kilobase deletion in a primate-specific segmental duplication creates a novel butyrophilin-like protein. Bmc Genet 2013, 14, doi:Artn 6110.1186/1471-2156-14-61.
- Mitsunaga, S.; Hosomichi, K.; Okudaira, Y.; Nakaoka, H.; Kunii, N.; Suzuki, Y.; Kuwana, M.; Sato, S.; Kaneko, Y.; Homma, Y.; et al. Exome sequencing identifies novel rheumatoid arthritis-susceptible variants in the BTNL2. J Hum Genet 2013, 58, 210-215. [CrossRef]
- Sirota, M.; Schaub, M.A.; Batzoglou, S.; Robinson, W.H.; Butte, A.J. Autoimmune disease classification by inverse association with SNP alleles. PLoS Genet 2009, 5, e1000792. [CrossRef]
- Orozco, G.; Eerligh, P.; Sanchez, E.; Zhernakova, S.; Roep, B.O.; Gonzalez-Gay, M.A.; Lopez-Nevot, M.A.; Callejas, J.L.; Hidalgo, C.; Pascual-Salcedo, D.; et al. Analysis of a functional BTNL2 polymorphism in type 1 diabetes, rheumatoid arthritis, and systemic lupus erythematosus. Hum Immunol 2005, 66, 1235-1241. [CrossRef]
- Traherne, J.A.; Barcellos, L.F.; Sawcer, S.J.; Compston, A.; Ramsay, P.P.; Hauser, S.L.; Oksenberg, J.R.; Trowsdale, J. Association of the truncating splice site mutation in BTNL2 with multiple sclerosis is secondary to HLA-DRB1*15. Hum Mol Genet 2006, 15, 155-161. [CrossRef]
- Hippich, M.; Beyerlein, A.; Hagopian, W.A.; Krischer, J.P.; Vehik, K.; Knoop, J.; Winker, C.; Toppari, J.; Lernmark, A.; Rewers, M.J.; et al. Genetic contribution to the divergence in type 1 diabetes risk between children from the general population and children from affected families. Diabetes 2019, 68, 847-857. [CrossRef]
- Johnson, S.B.; Lynch, K.F.; Roth, R.; Lundgren, M.; Parikh, H.M.; Akolkar, B.; Hagopian, W.; Krischer, J.; Rewers, M.; She, J.X.; et al. First-appearing islet autoantibodies for type 1 diabetes in young children: maternal life events during pregnancy and the child’s genetic risk. Diabetologia 2021, 64, 591-602. [CrossRef]
- He, C.; Hamon, S.; Li, D.; Barral-Rodriguez, S.; Ott, J.; Diabetes Genetics, C. MHC fine mapping of human type 1 diabetes using the T1DGC data. Diabetes Obes Metab 2009, 11 Suppl 1, 53-59. [CrossRef]
- Boyle, A.P.; Hong, E.L.; Hariharan, M.; Cheng, Y.; Schaub, M.A.; Kasowski, M.; Karczewski, K.J.; Park, J.; Hitz, B.C.; Weng, S.; et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res 2012, 22, 1790-1797. [CrossRef]
- Dong, S.; Zhao, N.; Spragins, E.; Kagda, M.S.; Li, M.; Assis, P.; Jolanki, O.; Luo, Y.; Cherry, J.M.; Boyle, A.P.; et al. Annotating and prioritizing human non-coding variants with RegulomeDB. bioRxiv 2022. [CrossRef]
- Spurkland, A.; Sollid, L.M.; Polanco, I.; Vartdal, F.; Thorsby, E. HLA-DR and -DQ genotypes of celiac disease patients serologically typed to be non-DR3 or non-DR5/7. Hum Immunol 1992, 35, 188-192. [CrossRef]
- Kawaguchi, S.; Higasa, K.; Shimizu, M.; Yamada, R.; Matsuda, F. HLA-HD: An accurate HLA typing algorithm for next-generation sequencing data. Hum Mutat 2017, 38, 788-797. [CrossRef]
- Dart, R.J.; Zlatareva, I.; Vantourout, P.; Theodoridis, E.; Amar, A.; Kannambath, S.; East, P.; Recaldin, T.; Mansfield, J.C.; Lamb, C.A.; et al. Conserved gammadelta T cell selection by BTNL proteins limits progression of human inflammatory bowel disease. Science 2023, 381, eadh0301. [CrossRef]
- Guo, M.H.; Plummer, L.; Chan, Y.M.; Hirschhorn, J.N.; Lippincott, M.F. Burden Testing of Rare Variants Identified through Exome Sequencing via Publicly Available Control Data. Am J Hum Genet 2018, 103, 522-534. [CrossRef]
- Guo, M.H. Burden testing against public controls. Available online: https://github.com/mhguo1/TRAPD (accessed on.
- Viken, M.K.; Blomhoff, A.; Olsson, M.; Akselsen, H.E.; Pociot, F.; Nerup, J.; Kockum, I.; Cambon-Thomsen, A.; Thorsby, E.; Undlien, D.E.; et al. Reproducible association with type 1 diabetes in the extended class I region of the major histocompatibility complex. Genes and Immunity 2009, 10, 323-333. [CrossRef]
- Horton, R.; Wilming, L.; Rand, V.; Lovering, R.C.; Bruford, E.A.; Khodiyar, V.K.; Lush, M.J.; Povey, S.; Talbot, C.C., Jr.; Wright, M.W.; et al. Gene map of the extended human MHC. Nat Rev Genet 2004, 5, 889-899. [CrossRef]
- Bycroft, C.; Freeman, C.; Petkova, D.; Band, G.; Elliott, L.T.; Sharp, K.; Motyer, A.; Vukcevic, D.; Delaneau, O.; O’Connell, J.; et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 2018, 562, 203-209. [CrossRef]
- Foers, A.D.; Shoukat, M.S.; Welsh, O.E.; Donovan, K.; Petry, R.; Evans, S.C.; FitzPatrick, M.E.; Collins, N.; Klenerman, P.; Fowler, A.; et al. Classification of intestinal T-cell receptor repertoires using machine learning methods can identify patients with coeliac disease regardless of dietary gluten status. J Pathol 2021, 253, 279-291. [CrossRef]
- Lindfors, K.; Ciacci, C.; Kurppa, K.; Lundin, K.E.A.; Makharia, G.K.; Mearin, M.L.; Murray, J.A.; Verdu, E.F.; Kaukinen, K. Coeliac disease. Nat Rev Dis Primers 2019, 5, 3. [CrossRef]
- Falchuk, Z.M.; Rogentine, G.N.; Strober, W. Predominance of histocompatibility antigen HL-A8 in patients with gluten-sensitive enteropathy. J Clin Invest 1972, 51, 1602-1605. [CrossRef]
- Stokes, P.L.; Asquith, P.; Holmes, G.K.; Mackintosh, P.; Cooke, W.T. Histocompatibility antigens associated with adult coeliac disease. Lancet 1972, 2, 162-164. [CrossRef]
- Karunakaran, M.M.; Willcox, C.R.; Salim, M.; Paletta, D.; Fichtner, A.S.; Noll, A.; Starick, L.; Nohren, A.; Begley, C.R.; Berwick, K.A.; et al. Butyrophilin-2A1 directly binds germline-encoded regions of the Vγ9Vδ2 TCR and is essential for phosphoantigen sensing. Immunity 2020, 52, 487-498 e486. [CrossRef]
- Rhodes, D.A.; Chen, H.C.; Price, A.J.; Keeble, A.H.; Davey, M.S.; James, L.C.; Eberl, M.; Trowsdale, J. Activation of human gammadelta T cells by cytosolic interactions of BTN3A1 with soluble phosphoantigens and the cytoskeletal adaptor periplakin. J Immunol 2015, 194, 2390-2398. [CrossRef]
- Sudlow, C.; Gallacher, J.; Allen, N.; Beral, V.; Burton, P.; Danesh, J.; Downey, P.; Elliott, P.; Green, J.; Landray, M.; et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med 2015, 12, e1001779. [CrossRef]
- Sayers, E.W.; Bolton, E.E.; Brister, J.R.; Canese, K.; Chan, J.; Comeau, D.C.; Connor, R.; Funk, K.; Kelly, C.; Kim, S.; et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 2022, 50, D20-D26. [CrossRef]
- Nonacus. Nonacus Probe Design Tool. Available online: https://mynonacus.nonacus.com/view-panel-designs (accessed on 8th February 2021).
- Simms, V. 5 Tips for using the Nonacus Panel Design Tool. Available online: https://nonacus.com/blog-get-great-coverage-for-the-genes-you-care-about/ (accessed on.
- Nonacus. Custom NGS Panel Design Tool. Available online: https://nonacus.com/panel-design/ (accessed on.
- Andrews, S. FastQC: A quality control analysis tool for high throughput sequencing data. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on.
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114-2120. [CrossRef]
- Van der Auwera, G.A.; O’Connor, B.D. Genomics in the Cloud: Using Docker, GATK, and WDL in Terra, 1st ed.; O’Reilly Media: 2020.
- Zhao, S.; Agafonov, O.; Azab, A.; Stokowy, T.; Hovig, E. Accuracy and efficiency of germline variant calling pipelines for human genome data. Sci Rep 2020, 10, 20222. [CrossRef]
- Cucco, F.; Barrans, S.; Sha, C.; Clipson, A.; Crouch, S.; Dobson, R.; Chen, Z.; Thompson, J.S.; Care, M.A.; Cummin, T.; et al. Distinct genetic changes reveal evolutionary history and heterogeneous molecular grade of DLBCL with MYC/BCL2 double-hit. Leukemia 2020, 34, 1329-1341. [CrossRef]
- Cucco, F.; Clipson, A.; Kennedy, H.; Sneath Thompson, J.; Wang, M.; Barrans, S.; van Hoppe, M.; Ochoa Ruiz, E.; Caddy, J.; Hamid, D.; et al. Mutation screening using formalin-fixed paraffin-embedded tissues: a stratified approach according to DNA quality. Lab Invest 2018, 98, 1084-1092. [CrossRef]
- Matthews, J. A snakemake pipeline for analysing (cancer) DNA sequencing data. Available online: https://gitlab.com/jdm204/dnaseq_snakemake (accessed on.
- Kawaguchi, S. HLA-HD. Available online: https://w3.genome.med.kyoto-u.ac.jp/HLA-HD/ (accessed on.
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve years of SAMtools and BCFtools. Gigascience 2021, 10. [CrossRef]
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.; Thormann, A.; Flicek, P.; Cunningham, F. The Ensembl Variant Effect Predictor. Genome Biol 2016, 17, 122. [CrossRef]
- Dilthey, A.; Leslie, S.; Moutsianas, L.; Shen, J.; Cox, C.; Nelson, M.R.; McVean, G. Multi-population classical HLA type imputation. PLoS Comput Biol 2013, 9, e1002877. [CrossRef]
- Yu, Y.; Fedele, G.; Celardo, I.; Loh, S.H.Y.; Martins, L.M. Parp mutations protect from mitochondrial toxicity in Alzheimer’s disease. Cell Death Dis 2021, 12, 651. [CrossRef]
- Grueneberg, A.; de Los Campos, G. BGData - A Suite of R Packages for Genomic Analysis with Big Data. G3 (Bethesda) 2019, 9, 1377-1383. [CrossRef]
- NCBI. SNP. Available online: https://www.ncbi.nlm.nih.gov/snp (accessed on.
- Gustavsen, J.; Rüeger, S.; Chamberlain, S.; Ushey, K.; Zhu, H. rsnps: Get ‘SNP’ (‘Single-Nucleotide’ ‘Polymorphism’) Data on the Web, 2024.
- Lefranc, M.P.; Giudicelli, V.; Duroux, P.; Jabado-Michaloud, J.; Folch, G.; Aouinti, S.; Carillon, E.; Duvergey, H.; Houles, A.; Paysan-Lafosse, T.; et al. IMGT(R), the international ImMunoGeneTics information system(R) 25 years on. Nucleic Acids Res 2015, 43, D413-422. [CrossRef]
- Bolotin, D.A.; Poslavsky, S.; Mitrophanov, I.; Shugay, M.; Mamedov, I.Z.; Putintseva, E.V.; Chudakov, D.M. MiXCR: software for comprehensive adaptive immunity profiling. Nat Methods 2015, 12, 380-381. [CrossRef]
- Bolotin, D.A.; Poslavsky, S.; Davydov, A.N.; Frenkel, F.E.; Fanchi, L.; Zolotareva, O.I.; Hemmers, S.; Putintseva, E.V.; Obraztsova, A.S.; Shugay, M.; et al. Antigen receptor repertoire profiling from RNA-seq data. Nat Biotechnol 2017, 35, 908-911. [CrossRef]
- McDonald, J.H. Handbook of Biological Statistics, 3rd ed.; Sparky House Publishing: Baltimore, Maryland, 2014.
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156-2158. [CrossRef]
- Human Protein Atlas, P. Human Protein Atlas. Available online: http://www.proteinatlas.org (accessed on 20th April 2021).
- Uhlen, M.; Fagerberg, L.; Hallstrom, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, A.; Kampf, C.; Sjostedt, E.; Asplund, A.; et al. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [CrossRef]
- Mölder, F.; Jablonski, K.; Letcher, B.; Hall, M.; Tomkins-Tinch, C.; Sochat, V.; Forster, J.; Lee, S.; Twardziok, S.; Kanitz, A.; et al. Sustainable data analysis with Snakemake [version 1; peer review: 1 approved, 1 approved with reservations]. F1000Research 2021, 10. [CrossRef]
- Schneider, V.A.; Graves-Lindsay, T.; Howe, K.; Bouk, N.; Chen, H.C.; Kitts, P.A.; Murphy, T.D.; Pruitt, K.D.; Thibaud-Nissen, F.; Albracht, D.; et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res 2017, 27, 849-864. [CrossRef]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. 2013. [CrossRef]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 2010, 38, e164. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




