Submitted:
16 September 2025
Posted:
17 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methodology
Systematic Literature Review:
Selection Criteria:
- Used ML or DL for plant disease classification.
- Reported on publicly available or well-documented datasets.
- Provided measurable outcomes, such as accuracy, precision, or recall.
- Were peer-reviewed and published in reputable journals or conferences.
Exclusion Criteria Eliminated Studies That:
- Applied ML or DL techniques in non-agricultural domains.
- Lacked experimental validation or reproducibility.
- Did not provide sufficient details about their methodologies or datasets.
Technical Evaluation:
- Problem Addressed: The specific plant diseases or agricultural issues being tackled.
- Techniques Used: ML or DL algorithms and architectures employed.
- Data Sources: The origin, size, and diversity of the datasets used.
- Performance Metrics: Overall accuracy, robustness, and scalability of the models.
Focus on Performance:
3. Comprehensive Survey on Classification and Detection Techniques for Plant Leaf Diseases
4. Review Table
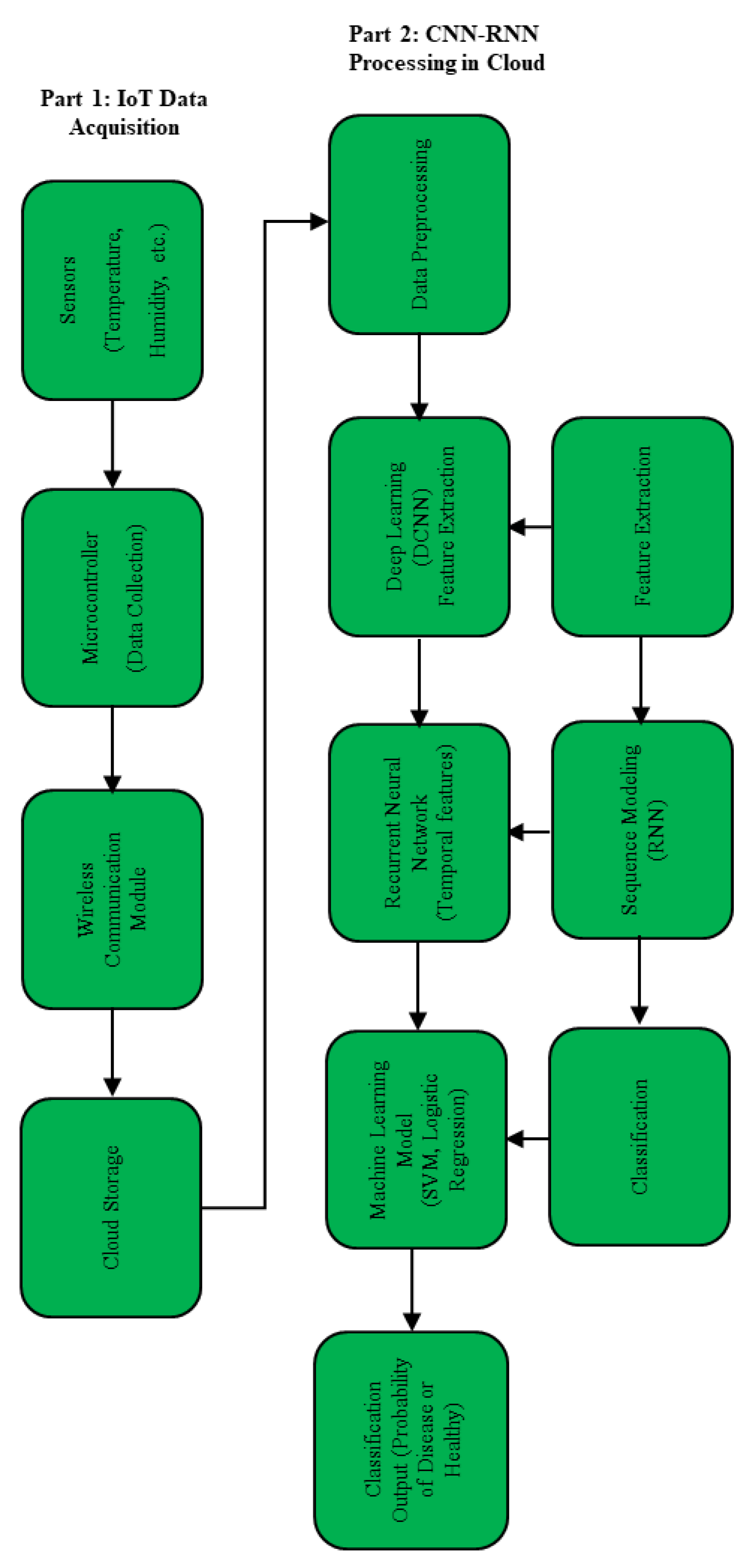
Part 1: IoT Data Acquisition
Part 2: CNN-RNN Processing in the Cloud
- Data Preprocessing: In this phase, both the sensor data and the plant images are subjected to preprocessing. It may be necessary to normalize or filter sensor data in order to ensure its quality. Similarly, images must be prepared for deep learning analysis.
- Feature Extraction: The convolutional neural network (CNN) is employed to analyze the preprocessed images, with the objective of extracting spatial features that indicate the presence of disease. The application of convolutional neural networks (CNNs) for pattern recognition in disease detection represents a crucial technology in precision agriculture, as observed by S. K. Javheri, who underscores the significance of artificial intelligence (AI) in crop health monitoring and disease detection [45].
- Sequence Modeling: Recurrent neural networks (RNNs) process the continuous data from sensors in order to capture temporal trends, which are crucial for understanding how plant health changes over time. This step is of particular importance for the early detection of diseases, which represents a central objective of the proposed architecture and is supported by the existing literature on the application of AI in agriculture.
- Feature Fusion and Classification: The features derived from the CNN and RNN are integrated to create a comprehensive representation, which is then classified using models such as SVM and Logistic Regression to determine the health status of the plant. This fusion exploits the spatial-temporal data to achieve accurate early detection and classification, thereby enhancing the precision of decision-making in agricultural contexts.
Core Projects and Literature Integration
5. Critical Analysis
6. Results and Findings
7. Future Directions
- Advanced Hybrid Architectures: Future models should prioritise hybrid frameworks (e.g., CNN-RNN, attention mechanisms) to integrate spatial-temporal data, enhancing early disease detection. This finding is consistent with studies such as Jiang et al. [16] and Zhang et al. [22], which underscore the efficacy of fused architectures for enhanced accuracy and real-time performance.
- Scalable and Diverse Datasets: Addressing dataset limitations, including class imbalance and a paucity of real-world variability, necessitates the curation of large-scale, multimodal datasets. The augmentation of synthetic data and the establishment of collaborative platforms for the dissemination of agricultural data in the public domain have the potential to mitigate biases and enhance the generalisability of models across a range of crops (e.g. millet, potatoes) and environments.
- Edge-AI and IoT Integration: The deployment of lightweight, energy-efficient models (e.g. MobileNetV3, EfficientNet) on edge devices will facilitate real-time, on-field diagnostics. This reduces reliance on cloud infrastructure, as seen in Nayak et al. [39], and supports IoT-enabled systems for continuous environmental monitoring and rapid response.
- Explainable AI (XAI): Bridging the trust gap between farmers and AI necessitates interpretable models. Techniques like Grad-CAM or attention visualization, as explored by Karthik et al. [20], can clarify decision-making processes, fostering adoption in practical settings.
- Cross-Domain Generalization: Advancing beyond the confines of a single crop (e.g., tomatoes), future research should prioritize the development of transferable models adaptable to diverse crops and geographies. This necessitates benchmarking against heterogeneous datasets and the incorporation of unsupervised and semi-supervised learning to mitigate the dependency on annotation.
- Sustainability-Driven Innovation: Developing resource-efficient algorithms (e.g. pruning, quantization) will democratize access for smallholder farmers. Moreover, the integration of artificial intelligence (AI) with climate-smart practices, such as the prediction of disease outbreaks under changing environmental conditions, has the potential to enhance global food security.
References
- Singh, Taranjeet, Krishna Kumar, and S. S. Bedi. A review on artificial intelligence techniques for disease recognition in plants. IOP Conference Series: Materials Science and Engineering 2021, 1022, 012032. [Google Scholar] [CrossRef]
- Jung, Minah, Jong Seob Song, Ah-Young Shin, Beomjo Choi, Sangjin Go, Suk-Yoon Kwon, Juhan Park, Sung Goo Park, and Yong-Min Kim. Construction of deep learning-based disease detection model in plants. Scientific reports 2023, 13, 7331. [Google Scholar] [CrossRef]
- Oliveira, Rosana Cavalcante de, and Rogério Diogne de Souza E. Silva. Artificial intelligence in agriculture: benefits, challenges, and trends. Applied Sciences 2023, 13, 7405. [Google Scholar] [CrossRef]
- Mkonyi, Lilian. Development of model for early identification of tomato plant damages caused by TUTA ABSOLUTA. PhD diss., NM-AIST, 2021. [CrossRef]
- Seth, Vishal, Rajeev Paulus, and Anil Kumar. Tomato leaf diseases detection using deep learning—a review. Intelligent Systems and Smart Infrastructure 2023, 118–131. [CrossRef]
- Gu, Jiuxiang, Zhenhua Wang, Jason Kuen, Lianyang Ma, Amir Shahroudy, Bing Shuai, Ting Liu et al. Recent advances in convolutional neural networks. Pattern recognition 2018, 77, 354–377. [Google Scholar] [CrossRef]
- Yamashita, Rikiya, Mizuho Nishio, Richard Kinh Gian Do, and Kaori Togashi. Convolutional neural networks: an overview and application in radiology. Insights into imaging 2018, 9, 611–629. [Google Scholar] [CrossRef] [PubMed]
- Halbouni, Asmaa, Teddy Surya Gunawan, Mohamed Hadi Habaebi, Murad Halbouni, Mira Kartiwi, and Robiah Ahmad. CNN-LSTM: hybrid deep neural network for network intrusion detection system. IEEE Access 2022, 10, 99837–99849. [Google Scholar] [CrossRef]
- Wang, Nan, Mingyue Cheng, and Kang Ning. Overcoming regional limitations: transfer learning for cross-regional microbial-based diagnosis of diseases. Gut 2023, 72, 2004–2006. [Google Scholar] [CrossRef] [PubMed]
- Corceiro, Ana, Khadijeh Alibabaei, Eduardo Assunção, Pedro D. Gaspar, and Nuno Pereira. Methods for detecting and classifying weeds, diseases and fruits using AI to improve the sustainability of agricultural crops: a review. Processes 2023, 11, 1263. [Google Scholar] [CrossRef]
- Xu, Zheng, and Cong Wu. Combination of Transfer Deep Learning and Classical Machine Learning Models for Multi-View Image Analysis. Computer Sciences & Mathematics Forum 2023, 7, 13. [Google Scholar] [CrossRef]
- B. Casella, A. B. Casella, A. Chisari, S. Battiato, and M. Giuffrida. Transfer Learning via Test-time Neural Networks Aggregation:. in Proceedings of the 17th International Joint Conference on Computer Vision, Imaging and Computer Graphics Theory and Applications, Online Streaming, --- Select a Country ---: SCITEPRESS - Science and Technology Publications, 2022, pp. 642–649. [CrossRef]
- N. K. E, K. N. K. E, K. M, P. P, A. R, and V. S. Tomato Leaf Disease Detection using Convolutional Neural Network with Data Augmentation. in 2020 5th International Conference on Communication and Electronics Systems (ICCES), COIMBATORE, India: IEEE, Jun. 2020, pp. 1125–1132. [CrossRef]
- Ashok, Surampalli, Gemini Kishore, Velpula Rajesh, S. Suchitra, SG Gino Sophia, and B. Pavithra. Tomato leaf disease detection using deep learning techniques. In 2020 5th International Conference on Communication and Electronics Systems (ICCES), pp. 979-983. IEEE, 2020. [CrossRef]
- Magsi, Aurangzeb, Javed Ahmed Mahar, Mirza Abdur Razzaq, and Sajid Habib Gill. Date palm disease identification using features extraction and deep learning approach. In 2020 IEEE 23rd International Multitopic Conference (INMIC), pp. 1-6. IEEE, 2020. [CrossRef]
- Jiang, Peng, Yuehan Chen, Bin Liu, Dongjian He, and Chunquan Liang. Real-time detection of apple leaf diseases using deep learning approach based on improved convolutional neural networks. Ieee Access 2019, 7, 59069–59080. [Google Scholar] [CrossRef]
- Wang, Qimei, Feng Qi, Minghe Sun, Jianhua Qu, and Jie Xue. Identification of tomato disease types and detection of infected areas based on deep convolutional neural networks and object detection techniques. Computational intelligence and neuroscience 2019, 2019, 9142753. [Google Scholar] [CrossRef]
- Kumar, Akshay, and M. Vani. Image based tomato leaf disease detection. In 2019 10th International Conference on Computing, Communication and Networking Technologies (ICCCNT), pp. 1-6. IEEE, 2019. [CrossRef]
- Ozguven, Mehmet Metin, and Kemal Adem. Automatic detection and classification of leaf spot disease in sugar beet using deep learning algorithms. Physica A: statistical mechanics and its applications 2019, 535, 122537. [Google Scholar] [CrossRef]
- Karthik, R., M. Hariharan, Sundar Anand, Priyanka Mathikshara, Annie Johnson, and R. Menaka. Attention embedded residual CNN for disease detection in tomato leaves. Applied Soft Computing 2020, 86, 105933. [Google Scholar] [CrossRef]
- Salih, Thair A. Deep learning convolution neural network to detect and classify tomato plant leaf diseases. Open Access Library Journal 2020, 7, 1. [Google Scholar] [CrossRef]
- Zhang, Yang, Chenglong Song, and Dongwen Zhang. Deep learning-based object detection improvement for tomato disease. IEEE access 2020, 8, 56607–56614. [Google Scholar] [CrossRef]
- Gangwar, Nitish, Divyansh Tiwari, Abhishek Sharma, Mritunjay Ashish, and Ankush Mittal. Grape leaf disease classification using transfer learning. International Research Journal of Engineering and Technology (IRJET) www.irjet.net. 2020.
- Vallabhajosyula, Sasikala, Venkatramaphanikumar Sistla, and Venkata Krishna Kishore Kolli. Transfer learning-based deep ensemble neural network for plant leaf disease detection. Journal of Plant Diseases and Protection 2022, 129, 545–558. [Google Scholar] [CrossRef]
- Aversano, Lerina, Mario Luca Bernardi, Marta Cimitile, Martina Iammarino, and Stefano Rondinella. Tomato diseases classification based on VGG and transfer learning. In 2020 IEEE International Workshop on Metrology for Agriculture and Forestry (MetroAgriFor), pp. 129-133. IEEE, 2020. [CrossRef]
- Saleem, Muhammad Hammad, Johan Potgieter, and Khalid Mahmood Arif. Plant disease classification: A comparative evaluation of convolutional neural networks and deep learning optimizers. Plants 2020, 9, 1319. [Google Scholar] [CrossRef]
- Nawaz, Marriam, Tahira Nazir, Ali Javed, Momina Masood, Junaid Rashid, Jungeun Kim, and Amir Hussain. A robust deep learning approach for tomato plant leaf disease localization and classification. Scientific reports 2022, 12, 18568. [Google Scholar] [CrossRef]
- Agarwal, Mohit, Abhishek Singh, Siddhartha Arjaria, Amit Sinha, and Suneet Gupta. ToLeD: Tomato leaf disease detection using convolution neural network. Procedia Computer Science 2020, 167, 293–301. [Google Scholar] [CrossRef]
- Jeyalakshmi, S. , and R. Radha. CLASSIFICATION OF TOMATO DISEASES USING ENSEMBLE LEARNING. ICTACT Journal on Soft Computing 2021, 11. [Google Scholar] [CrossRef]
- Kibriya, Hareem, Rimsha Rafique, Wakeel Ahmad, and S. M. Adnan. Tomato leaf disease detection using convolution neural network. In 2021 International Bhurban Conference on Applied Sciences and Technologies (IBCAST), pp. 346-351. IEEE, 2021. [CrossRef]
- Parvez, Shamima, Md Ashraf Uddin, Md Manowarul Islam, Pallab Bharman, and Md Alamin Talukder. Tomato leaf disease detection using convolutional neural network. 2023. [CrossRef]
- Nagamani, H. S. , and H. Sarojadevi. Tomato leaf disease detection using deep learning techniques. International Journal of Advanced Computer Science and Applications 2022, 13. [Google Scholar] [CrossRef]
- Al-gaashani, Mehdhar SAM, Fengjun Shang, Mohammed SA Muthanna, Mashael Khayyat, and Ahmed A. Abd El-Latif. Tomato leaf disease classification by exploiting transfer learning and feature concatenation. IET Image Processing 2022, 16, 913–925. [Google Scholar] [CrossRef]
- Lakshmanarao, A. Supriya, and A. Arulmurugan. Plant disease prediction using transfer learning techniques. In 2022 Second International Conference on Advances in Electrical, Computing, Communication and Sustainable Technologies (ICAECT), pp. 1-5. IEEE, 2022. [CrossRef]
- Attallah, Omneya. Tomato leaf disease classification via compact convolutional neural networks with transfer learning and feature selection. Horticulturae 2023, 9, 149. [Google Scholar] [CrossRef]
- Borugadda, Premkumar, Ramasami Lakshmi, and Satyasangram Sahoo. Transfer Learning VGG16 Model for Classification of Tomato Plant Leaf Diseases: A Novel Approach for Multi-Level Dimensional Reduction. Pertanika Journal of Science & Technology 2023, 31. [Google Scholar] [CrossRef]
- Kaur, Prabhjot, Shilpi Harnal, Vinay Gautam, Mukund Pratap Singh, and Santar Pal Singh. A novel transfer deep learning method for detection and classification of plant leaf disease. Journal of Ambient Intelligence and Humanized Computing 2023, 14, 12407–12424. [Google Scholar] [CrossRef]
- Liu, Guangsheng, Jialiang Peng, and Ahmed A. Abd El-Latif. SK-MobileNet: a lightweight adaptive network based on complex deep transfer learning for plant disease recognition. Arabian journal for science and engineering 2023, 48, 1661–1675. [Google Scholar] [CrossRef]
- Nayak, Anshuman, Somsubhra Chakraborty, and Dillip Kumar Swain. Application of smartphone-image processing and transfer learning for rice disease and nutrient deficiency detection. Smart Agricultural Technology 2023, 4, 100195. [Google Scholar] [CrossRef]
- Nigam, Sapna, Rajni Jain, Sudeep Marwaha, Alka Arora, Md Ashraful Haque, Akshay Dheeraj, and Vaibhav Kumar Singh. Deep transfer learning model for disease identification in wheat crop. Ecological Informatics 2023, 75, 102068. [Google Scholar] [CrossRef]
- Jiang, Han, Zhi Peng Xue, and Yan Guo. Research on plant leaf disease identification based on transfer learning algorithm. Journal of Physics: Conference Series 2020, 1576, 012023. [Google Scholar] [CrossRef]
- El Massi, Ismail, Youssef Es-saady, Mostafa El Yassa, and Driss Mammass. Combination of multiple classifiers for automatic recognition of diseases and damages on plant leaves. Signal, Image and Video Processing 2021, 15, 789–796. [Google Scholar] [CrossRef]
- Marino, Sofia, Pierre Beauseroy, and André Smolarz. Deep Learning-based Method for Classifying and Localizing Potato Blemishes. ICPRAM 2019, 11996, 107–117. [Google Scholar] [CrossRef]
- Thotho, Doreen, and Paul Macheso. Comprehensive Survey on Applications of Internet of Things, Machine Learning and Artificial Intelligence in Precision Agriculture. Tanzania Journal of Engineering and Technology 2023, 42, 30–45. [Google Scholar] [CrossRef]
- S. K. Javheri. AGRICULTURE AND ARTIFICIAL INTELLIGENCE: A NEW RESEARCH ERA”. [CrossRef]
- Mohy-eddine, Mouaad, Azidine Guezzaz, Said Benkirane, and Mourade Azrour. IoT-enabled smart agriculture: security issues and applications. In The International Conference on Artificial Intelligence and Smart Environment, pp. 566-571. Cham: Springer International Publishing, 2023. [CrossRef]
- Mukti, Ishrat Zahan, and Dipayan Biswas. Transfer learning based plant diseases detection using ResNet50. In 2019 4th International conference on electrical information and communication technology (EICT), pp. 1-6. IEEE, 2019. [CrossRef]
- Militante, Sammy V., Bobby D. Gerardo, and Nanette V. Dionisio. Plant leaf detection and disease recognition using deep learning. In 2019 IEEE Eurasia conference on IOT, communication and engineering (ECICE), pp. 579-582. IEEE, 2019. [CrossRef]
- Hasan, Mosin, Bhavesh Tanawala, and Krina J. Patel. Deep learning precision farming: Tomato leaf disease detection by transfer learning. In Proceedings of 2nd international conference on advanced computing and software engineering (ICACSE). 2019. [CrossRef]
- Francis, Mercelin, and C. Deisy. Disease detection and classification in agricultural plants using convolutional neural networks—a visual understanding. In 2019 6th international conference on signal processing and integrated networks (SPIN), pp. 1063-1068. IEEE, 2019. [CrossRef]
- Jakjoud, F. Hatim, and A. Bouaddi. Detection of diseases on tomato leaves based on Sub–Classifiers Fuzzy Combination. International Journal of innovative Technology and Exploring Engineering (IJITEE), ISSN 2019, 2278–3075. [Google Scholar]
- Coulibaly, Solemane, Bernard Kamsu-Foguem, Dantouma Kamissoko, and Daouda Traore. Deep neural networks with transfer learning in millet crop images. Computers in industry 2019, 108, 115–120. [Google Scholar] [CrossRef]
- Zhang, Tao, Xiankun Zhu, Yiqing Liu, Kun Zhang, and Azhar Imran. Deep learning based classification for tomato diseases recognition. In IOP Conference Series: Earth and Environmental Science 2020, 474, 032014. [CrossRef]
- Mathulaprangsan, Seksan, Kitsana Lanthong, Duangpen Jetpipattanapong, Siwadol Sateanpattanakul, and Sujin Patarapuwadol. Rice diseases recognition using effective deep learning models. In 2020 Joint International Conference on Digital Arts, Media and Technology with ECTI Northern Section Conference on Electrical, Electronics, Computer and Telecommunications Engineering (ECTI DAMT & NCON), pp. 386-389. IEEE, 2020. [CrossRef]
- Ouhami, Maryam, Youssef Es-Saady, Mohamed El Hajji, Adel Hafiane, Raphael Canals, and Mostafa El Yassa. Deep transfer learning models for tomato disease detection. In Image and Signal Processing: 9th International Conference, ICISP 2020, Marrakesh, Morocco, June 4–6, 2020, Proceedings 9, pp. 65-73. Springer International Publishing, 2020. [CrossRef]
- Chen, Junde, Jinxiu Chen, Defu Zhang, Yuandong Sun, and Yaser Ahangari Nanehkaran. Using deep transfer learning for image-based plant disease identification. Computers and Electronics in Agriculture 2020, 173, 105393. [Google Scholar] [CrossRef]
- Chen, Junde, Defu Zhang, Yaser A. Nanehkaran, and Dele Li. Detection of rice plant diseases based on deep transfer learning. Journal of the Science of Food and Agriculture 2020, 100, 3246–3256. [Google Scholar] [CrossRef]
- Rosmala, D. , MR Prakha Anggara, and J. P. Sahat. Transfer learning with vgg16 and inceptionv3 model for classification of potato leaf disease. Journal of Theoretical and Applied Information Technology 2021, 99, 279–292. [Google Scholar]
- Wagle, S.A. ,. A Deep Learning-Based Approach in Classification and Validation of Tomato Leaf Disease. Traitement du signal 2021, 38. [Google Scholar] [CrossRef]
- Ashwinkumar, S., S. Rajagopal, V. Manimaran, and B. Jegajothi. Automated plant leaf disease detection and classification using optimal MobileNet based convolutional neural networks. Materials Today: Proceedings 2022, 51, 480–487. [Google Scholar] [CrossRef]
- Wagle, Shivali Amit, Jahariah Sampe, Faseehuddin Mohammad, and Sawal Hamid Md Ali. Effect of Data Augmentation in the Classification and Validation of Tomato Plant Disease with Deep Learning Methods. Traitement du Signal 2021, 38. [Google Scholar] [CrossRef]
- Hassan, Rondik J. , and Adnan Mohsin Abdulazeez. Plant Leaf Disease Detection by Using Different Classification Techniques: Comparative. Asian Journal of Research in Computer Science 2021, 8, 1–11. [Google Scholar] [CrossRef]
- Abbas, Amreen, Sweta Jain, Mahesh Gour, and Swetha Vankudothu. Tomato plant disease detection using transfer learning with C-GAN synthetic images. Computers and Electronics in Agriculture 2021, 187, 106279. [Google Scholar] [CrossRef]
- Chowdhury, Muhammad EH, Tawsifur Rahman, Amith Khandakar, Nabil Ibtehaz, Aftab Ullah Khan, Muhammad Salman Khan, Nasser Al-Emadi, Mamun Bin Ibne Reaz, Mohammad Tariqul Islam, and Sawal Hamid Md Ali. Tomato leaf diseases detection using deep learning technique. Technology in Agriculture 2021, 453. [Google Scholar] [CrossRef]
- Wang, Xuewei, and Jun Liu. Tomato anomalies detection in greenhouse scenarios based on YOLO-Dense. Frontiers in Plant Science 2021, 12, 634103. [Google Scholar] [CrossRef]
- Feng, Lei, Baohua Wu, Yong He, and Chu Zhang. Hyperspectral imaging combined with deep transfer learning for rice disease detection. Frontiers in Plant Science 2021, 12, 693521. [Google Scholar] [CrossRef]
- Khasawneh, Natheer, Esraa Faouri, and Mohammad Fraiwan. Automatic detection of tomato diseases using deep transfer learning. Applied Sciences 2022, 12, 8467. [Google Scholar] [CrossRef]
- Ahmed, Sabbir, Md Bakhtiar Hasan, Tasnim Ahmed, Md Redwan Karim Sony, and Md Hasanul Kabir. Less is more: Lighter and faster deep neural architecture for tomato leaf disease classification. IEEE Access 2022, 10, 68868–68884. [Google Scholar] [CrossRef]
- Nguyen, Thanh-Hai, Thanh-Nghia Nguyen, and Ba-Viet Ngo. A VGG-19 model with transfer learning and image segmentation for classification of tomato leaf disease. AgriEngineering 2022, 4, 871–887. [Google Scholar] [CrossRef]
- Al-Akkam, Reem Mohammed Jasim, and Mohammed Sahib Mahdi Altaei. Plants leaf diseases detection using deep learning. Iraqi JournalofScience 2022, 801816. [Google Scholar] [CrossRef]
- Boutalline, Mohammed, Adil Tannouche, Hassan Faouzi, Hamid Ouanan, and Malak Dargham. Automatic Detection and Classification of Apple Leaves Diseases Using MobileNet V2. Revue d'Intelligence Artificielle 2022, 36, 745. [Google Scholar] [CrossRef]
- Zhang, Liangji, Guoxiong Zhou, Chao Lu, Aibin Chen, Yanfeng Wang, Liujun Li, and Weiwei Cai. MMDGAN: A fusion data augmentation method for tomato-leaf disease identification. Applied Soft Computing 2022, 123, 108969. [Google Scholar] [CrossRef]
- Hajraoui, Noredine, Mourade Azrour, and Ahmad El Allaoui. Classification of diseases in tomato leaves with Deep Transfer Learning. In The International Conference on Artificial Intelligence and Smart Environment, pp. 607-612. Cham: Springer Nature Switzerland, 2023. [CrossRef]
- Hessane, Abdelaaziz, Ahmed El Youssefi, Yousef Farhaoui, Badraddine Aghoutane, and Fatima Amounas. A machine learning based framework for a stage-wise classification of date palm white scale disease. Big Data Mining and Analytics 2023, 6, 263–272. [Google Scholar] [CrossRef]
- Ur Rehman, Muhammad Zia, Fawad Ahmed, Muhammad Attique Khan, Usman Tariq, Sajjad Shaukat Jamal, Jawad Ahmad, and Iqtadar Hussain. Classification of Citrus Plant Diseases Using Deep Transfer Learning. Computers, Materials & Continua 2022, 70. [Google Scholar] [CrossRef]
- Bensaadi, Soumia, and Ahmed Louchene. Low-cost convolutional neural network for tomato plant diseases classifiation. IAES International Journal of Artificial Intelligence 2023, 12, 162. [Google Scholar] [CrossRef]
- Isnan, Mahmud, Alam Ahmad Hidayat, and Bens Pardamean. Indonesian agricultural-crops classification using transfer learning model. Procedia Computer Science 2023, 227, 128–136. [Google Scholar] [CrossRef]
- Ramya, R. , and P. Kumar. High-performance deep transfer learning model with batch normalization based on multiscale feature fusion for tomato plant disease identification and categorization. Environmental Research Communications 2023, 5, 125015. [Google Scholar] [CrossRef]
- Shahoveisi, F. , Taheri Gorji, H., Shahabi, S., Hosseinirad, S., Markell, S. and Vasefi, F.,. Application of image processing and transfer learning for the detection of rust disease. Scientific Reports 2023, 13, 5133. [Google Scholar] [CrossRef]
- Mimi, Afsana, Sayeda Fatema Tuj Zohura, Muhammad Ibrahim, Riddho Ridwanul Haque, Omar Farrok, Taskeed Jabid, and Md Sawkat Ali. Identifying selected diseases of leaves using deep learning and transfer learning models. Machine Graphics & Vision 2023, 32. [Google Scholar] [CrossRef]
- H. M. Zayani et al.. Deep Learning for Tomato Disease Detection with YOLOv8. Eng. Technol. Appl. Sci. Res. 2024, 14, 13584–13591. [Google Scholar] [CrossRef]
- U. Zahra, M. A. Khan, M. Alhaisoni, A. Alasiry, M. Marzougui, and A. Masood. An Integrated Framework of Two-Stream Deep Learning Models Optimal Information Fusion for Fruits Disease Recognition. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens, 2024, 17, 3038–3052. [Google Scholar] [CrossRef]
- S. A. F. Naqvi et al.. Fruit and vegetable leaf disease recognition based on a novel custom convolutional neural network and shallow classifier. Front. Plant Sci 2024, 15. [Google Scholar]
- M. S. A. M. Al-Gaashani et al.. Deep transfer learning with gravitational search algorithm for enhanced plant disease classification. Heliyon 2024, 10, e28967. [Google Scholar] [CrossRef]
- Abdul Aziz, Abdul Fadlil, and Tole Sutikno. Optimization of Convolutional Neural Network (CNN) Using Transfer Learning for Disease Identification in Rice Leaf Images. J. E-Komtek Elektro-Komput.-Tek. 2024, 8, 504–515. [Google Scholar] [CrossRef]
- W. Shafik, A. Tufail, C. De Silva Liyanage, and R. A. A. H. M. Apong. Using transfer learning-based plant disease classification and detection for sustainable agriculture. BMC Plant Biol. 2024, 24, 136. [Google Scholar] [CrossRef]
- Y. A. Bezabh, A. M. Ayalew, B. M. Abuhayi, T. N. Demlie, E. A. Awoke, and T. E. Mengistu. Classification of mango disease using ensemble convolutional neural network. Smart Agric. Technol. 2024, 8, 100476. [Google Scholar] [CrossRef]
- R. Gai, Y. Liu, and G. Xu. TL-YOLOv8: A Blueberry Fruit Detection Algorithm Based on Improved YOLOv8 and Transfer Learning. IEEE Access, 2024, 12, 86378–86390. [Google Scholar] [CrossRef]
- P. Buchke and A. V. R. Mayuri. Recognize and classify illnesses on tomato leaves using EfficientNet’s transfer learning approach with different size dataset. Signal Image Video Process 2024, 18, 731–746. [Google Scholar] [CrossRef]
- H. -T. Vo, K. C. Mui, N. N. Thien, P. P. Tien, and H. L. Le. Optimizing Grape Leaf Disease Identification Through Transfer Learning and Hyperparameter Tuning. Int. J. Adv. Comput. Sci. Appl 2024, 15. [Google Scholar] [CrossRef]
- D. Han and C. Guo. Automatic classification of ligneous leaf diseases via hierarchical vision transformer and transfer learning. Front. Plant Sci. 2024, 14, 1328952. [Google Scholar]
- P. Radočaj, D. Radočaj, and G. Martinović. Image-Based Leaf Disease Recognition Using Transfer Deep Learning with a Novel Versatile Optimization Module. Big Data Cogn. Comput 2024, 8. [Google Scholar]
| Author | Plant type (number of classes) | Model(s)/ Algorithms/ Technique(s)/ Methods |
Hyperparameters |
Performance /Accuracy | Advantages | Disadvantages / Challenges | Objective | Future Research Direction |
|---|---|---|---|---|---|---|---|---|
| 2019 | ||||||||
| Jiang et al. [16] | Apple (5 classes) | INAR-SSD (Improved SSD (single-shot multibox detector) with Inception modules and Rainbow concatenation) |
Learning Rate: 0.001. Optimizer: SGD. Momentum :0.9. Batch Size:32. Epochs: Not specified. |
78.80% mAP (mean Average Precision). | -The system is capable of real-time detection at high speed. -It is also capable of handling complex backgrounds and multiple diseases per image. -The system has been enhanced to improve small-object detection via Inception and Rainbow concatenation. -Robust data augmentation has been employed to reduce the risk of overfitting. |
-The lower accuracy observed in the identification of similar diseases is a notable finding. -It has been observed that the system struggles with extremely small lesions or noisy backgrounds. -Additionally, its performance lags that of standard SSD technology. |
The objective of this study is to develop a real-time, high-accuracy deep learning model for detecting five common apple leaf diseases. The model will aid early diagnosis and improve agricultural productivity. | The following improvements are proposed: -The detection of visually similar diseases is to be improved. -The performance on very small lesions is to be enhanced. -The speed for real-time field applications is to be optimised. -The scope is to be expanded to other crops or diseases. |
| Mukti et Biswas [47] | Various (38 classes). | ResNet50, VGG16, VGG19, AlexNet |
Learning Rate: Not specified. Optimizer: SGD. Batch Size: 32. Epochs: 25. |
99.80% | The Transfer Learning approach enabled the development of a deep CNN network cost-effectively for precise plant disease identification. | The study does not delve into the challenges of real-world deployment or scalability. | The main objective was to develop a CNN model based on transfer learning for the accurate identification of plant diseases. | Future studies could explore the possibility of studying transfer learning architectures or combining them with data augmentation strategies to further improve the model's performance. |
| Wang et al. [17] | Tomato (11 classes). | R-CNN with four Deep Convolutional Neural Networks (VGG-16, ResNet-50, ResNet-101, and MobileNet) |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: 500. |
99.64% mAP (mean Average Precision). | The model can accurately identify diseases and infected areas, thus enabling timely treatment and prevention measures. | May require large amounts of labeled data for training the models. | Develop a system for precise identification of types of tomato diseases and segmentation of infected areas using deep learning. | Explore incorporating more diverse datasets and potentially other object detection techniques for enhanced disease identification and segmentation. |
| Kumar et Vani [18] | Tomato (10 classes). | CNN-based architectures. |
Learning Rate: 0.0005. Momentum:0.9, Decay:0.0005. Optimizer: Adam. Batch Size: 30. Epochs: 30. |
99.25%. | The proposed model, despite some low losses, was able to maximize the accuracy. | VGGNet creates a time issue as it takes more time to train and requires sophisticated hardware to train. | -Develop a system for detecting tomato leaf diseases through image analysis. | -Test the system with a larger and more diverse dataset of tomato leaf images to improve its generalizability and robustness. - Exploring other transfer learning techniques to improve the accuracy and efficiency of disease detection. |
| Militante et al. [48] | Various: apple, corn, grapes, potato, sugar cane, and tomato. (32 classes) | -CNN with Adam optimizer using a categorical cross-entropy loss function. -Data augmentations techniques. |
Learning Rate: Not specified. Optimizer: Adam. Batch Size: 32. Epochs: 75. |
96.5% in training. 100% in testing. |
The model provided a highly accurate and effective solution for the early detection and identification of multiple plant diseases, which facilitates effective disease management and helps reduce the use of harmful chemicals. | The dataset and model have limitations on generalizability, due to being tested on a limited number of plant varieties and diseases. Unseen disease and real-world variability also negatively impact model performance. | Help farmers detect and accurately recognize plant diseases in various plant varieties to improve disease management by reducing chemical interventions, using deep learning techniques, specifically CNNs. | Research could focus on expanding the data to other plants, testing different CNN architectures, learning, and optimizers to improve performance and the model. |
| Ozguven et Adem [19] | Sugar beet (4 classes). | Modified Faster R-CNN architecture, a deep learning model for object detection. |
Learning Rate: Not specified. Optimizer: SGD, Momentum;0.85, Decay:0.001. Batch Size: The heap size for training was set to 64. Epochs: 150.00. |
95.48% | The Faster R-CNN model has a higher accuracy rate in detecting leaf spot disease in sugar beet compared to previous methods, thus providing a means of effective accurate diagnosis of disease in large production areas. | The sensitivity values of the proposed approach were lower than the specificity values, indicating a slight imbalance in the detection and classification. | Develop a method based on deep learning for automatic disease detection of leaf spots in sugar beet leaves, contributing to imaging-based expert systems for disease detection in agriculture. | To conduct further studies using deep learning algorithms trained with a larger amount of data to improve the accuracy of detection of sugar beet leaves. |
| Hasan et al. [49] | Tomato leaves (3 classes) | Transfer learning with an inception model. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: 250. |
99% | Transfer learning of the Inception model leverages pre-trained features to optimize the training of convolutional neural networks (CNN) for tomato leaf classification, reducing data and computational effort. | The dataset comprises images from the internet and local farms, which may introduce variability due to different lighting conditions, backgrounds, and quality. | Implement a precision agriculture system using drones for the detection of leaf diseases. The system aims to effectively identify areas of disease prevalence on the farm. |
To explore real-time disease severity assessment based on infection levels using the same drone-based precision farming system. |
| Francis et Deisy [50] | Apple and Tomato (2 classes) | Convolutional Neural Network (CNN) architecture. |
Learning Rate: Not specified. Optimizer: Adam. Batch Size: Not specified. Epochs: 8000 iterations. |
88.7% | The CNN model automatically identifies and stores features in the training dataset, thereby avoiding the need for manually designed features. | Training a CNN model from scratch can be a tedious process compared to existing deep-learning models. | The main objective of the article is to explore different learning architectures and their applications in agriculture, particularly in the classification of plant diseases. | Researchers could explore transfer learning techniques to improve model performance on other plant species to study multiclass disease classification. |
| Marino et al. [43] | Potatoes (6 classes) | -CNN models: AlexNet, VGG-16, and GoogLeNet. -An SVM classifier was used to classify the data further. |
Learning Rate: -Global rate:0.0001. -New fully connected layer rate: 0.002. Optimizer: Stochastic Gradient Descent (SGD). Momentum :0.9. Batch Size: 10. Epochs: 100. |
F1-score of 94%. | Deep learning methods automatically find representations for classification tasks without the need for manual features. | - Building pixel-labeled data sets using deep learning methods can be laborious and time-consuming. -Difficulty in designing a feature extractor for each pattern. |
-To efficiently classify and locate imperfections in potatoes using deep learning techniques. -Automate quality control of potatoes to overcome subjectivity and high labor costs. |
- Focus on enhancing the localization accuracy of potato blemishes using advanced deep learning architectures or incorporating multi-modal data for improved classification performance. -Investigate multi-modal fusion (e.g., combining visual and spectral information) for improved potato blemish detection and classification. |
| Jakjoud et al. [51] | Tomato (2 classes) | -Support Vector Machine (SVM) and K-Nearest Neighbors (KNN). -Co-occurrence matrix for extracting 14 Haralick features. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
KNN with Fuzzy Decision Maker: 98.38% on the validation set. |
- The use of KNN is its ability to store data without requiring a tedious training step. -The proposed approach combines subclassifiers using fuzzy logic, thus enhancing accuracy. |
- SVM may face hyperplane tuning issues due to the dependency between parameters. - The study does not explore multiclass classification beyond normal leave and sick leave. |
-The objective of the study is to develop a capable of detecting anomalies in leaves to improve agricultural productivity. -Automatically identify plant diseases, avoiding significant agricultural losses. |
Future research could focus on improving classification accuracy by exploring and integrating other advanced machine-learning techniques and developing feature extraction methods. |
| Coulibaly et al. [52] | Pearl millet (2 classes) | Transfer learning with VGG16. |
Learning Rate: 1 e-4. Optimizer: Stochastic Gradient Descent (SGD). Momentum:0.9. Batch Size: Not specified. Epochs: 100 epochs, with early stopping observed at the 30th epoch. |
95% | -transfer learning with pre-trained models such as VGG16 allows high accuracy in the classification of diseases with limited data. - The proposed approach facilitates rapid and interesting analysis of data in precision agriculture. |
- Deep learning algorithms may require datasets and computing resources for training. - Manually generating labeled data for small datasets can be difficult and expensive. |
- The objective is to improve the identification of diseases in millet crops using advanced deep-learning techniques. -To detect mildew disease in various crops and integrate the solution into digital devices for farmers to identify plant diseases. |
- Explore the application of transfer learning and deep neural networks for disease detection in a broader range of crops beyond millet, such as cotton and potatoes, to support smart farming initiatives. -Enhancing transfer learning techniques for even better disease detection in millet crops. |
| 2020 | ||||||||
| Zhang Tao [53] | Tomato (18 classes) | SE-ResNet |
Learning Rate: 0.001 (decays by 0.1× after 12 epochs). Optimizer: Adam. Batch Size: 32. Epochs: Early stopping (patience=12 epochs). |
Validation Accuracy = 88.83%. |
The model provides state-of-the-art results, demonstrating high accuracy and robustness. | The model effectively mitigates the risk of overfitting, although this is not explicitly stated. | To effectively identify various tomato diseases and their severity using deep learning approaches. | The future research direction includes studying disease identification methods when multiple diseases coexist. |
| Salih et al. [21] | Tomato (6 classes) | Convolutional Neural Network (CNN) |
Learning Rate: Initial rate: 0.001, reduced by a factor of 0.5 during training. Optimizer: Adam. Batch Size: 64. Epochs: 10. |
96.43% | The merits of this system include the ability to recognize and detect problems in a short time, as well as the capacity to identify plant diseases at an early stage. This, in turn, results in improved production and better quality. | -The duration of training is extensive. -The resolution of input images is a matter of some complexity. -The common diseases infecting tomato plants are similar, which poses a challenge to accurate classification. |
The application of modern techniques, in particular convolutional networks, is crucial for the early detection of diseases in tomato plants. | Further, it improves the accuracy of classification by addressing the challenges posed by the similarity of common diseases in tomato plants. |
| Karthik R.et al. [20] | Tomato (4 classes) | Attention Embedded based Residual Convolutional Neural Network (ResCNN) |
Learning Rate: Not specified. Optimizer: Adam (Adaptive Moment Estimation). Batch Size: Not specified. Epochs: 150. |
98% | -The detection rate is higher than that of existing methods. -The number of parameters is reduced (600K vs. millions in existing architectures). -The method is extensible to any input size. -The attention mechanism improves feature weighting and contextual learning. |
-The most recent CNN technique for prior results has not been discussed. -There has been no discussion about real-time deployment or computational efficiency. The training process requires substantial resources, requiring a duration of 10 hours on an NVIDIA Tesla P100 GPU. Furthermore, the augmented data dependency has the potential to result in overfitting. |
To develop a deep learning model for automated disease detection in tomato leaves. The model will be both computationally efficient and accurate, and it will be based on an attention-embedded residual CNN. | -Implementation for real-time field applications. -Expansion to encompass other crops or multi-disease detection. -Optimization for edge devices (e.g., drones, mobile applications). -Addressing class imbalance in datasets. -Investigate enhancing the attention-based mechanisms to improve disease detection accuracy. |
| Mathulaprangsan et al. [54] | Rice (5 classes) | ResNet50, ResNet101, DenseNet161, and DenseNet169 |
Learning Rate: 0.0001. Optimizer: Not specified. Batch Size: 64. Epochs: 15. |
95.74% | The DenseNet architecture effectively deals with the issue of fading gradients, allowing the network to be more parameter-efficient and achieve high performance. | -The complexity of deep learning models requires significant resources. - The general CNN models had difficulty functioning properly due to the high level of detail in rice disease images. |
Create a full-field rice disease image dataset and apply efficient deep learning models to classify devastating diseases of rice Thailand. | Focus on improving and scalability of deep learning models for broader agricultural applications beyond rice diseases. |
| Ashok et al. [14] | Tomato (4 classes) | -Convolutional Neural Network (CNN). -DWT (Discrete Wavelet Transform) and GLCM (Gray Level Co-occurrence Matrix). |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
98,12% | -Facilitation of early disease detection. -Integration of the Discrete Wavelet Transform (DWT) and the Gray Level Co-occurrence Matrix (GLCM) to ensure the extraction of robust features. -Offers efficient computational performance and the potential to automate the process, thereby facilitating faster and more consistent disease identification in comparison to manual methods. |
-The dataset size and diversity have not been specified. -No hyperparameter details have been provided. -Limited real-time testing has been mentioned. -A greater number of samples is required for broader disease classification. -A large dataset is necessary to effectively train the model. |
To develop a system capable of detecting tomato leaf diseases in their early stages using convolutional neural networks (CNNs) and image processing techniques, including discrete wavelet transform (DWT) and gray level co-occurrence matrix (GLCM), to assist farmers in implementing preventative measures. The proposed approach employs deep learning techniques for the purpose of disease detection in tomato leaves. | -Extension to other algorithms (e.g. artificial neural networks, fuzzy logic) -Implementation of real-time applications -improvement of disease categorization -Testing on larger and more diverse datasets -Exploration of different deep-learning architectures or incorporation of data to improve the accuracy of disease detection. |
| Nithish kannan et al. [13] | Tomato crop (6 classes) | Convolutional neural networks for classification and data augmentation techniques to augment the training dataset (Resnet50). |
Learning Rate: 0.001. Optimizer: Adam. Momentum;0.1. Batch Size: Not specified. Epochs: 20. |
97% | High accuracy (97%) of the multi-class disease detection is a notable strength of the system. Transfer learning has been employed to reduce the time taken for training, while data augmentation has been implemented to prevent overfitting. The system also generalizes well to diverse leaf conditions. | -This process requires advanced configuration, including high-end hardware such as the NVIDIA GTX 1050 Ti GPU and 16GB of RAM. -It should be noted that the training process is extensive due to the complexity of ResNet-50. -Furthermore, the system is constrained to the identification of only six tomato diseases. |
The objective of this study is to utilize deep learning as a tool to facilitate farmers in the timely identification of six tomato leaf diseases, contributing to the enhancement of their agricultural practices. | -The model should be extended to encompass other crops, such as apples, potatoes and aubergines. -Hyperparameters should be optimized to reduce the time taken for training. -Hardware efficiency should be improved for deployment in settings where resources are limited. |
| Zhang et al. [22] | Tomato (5 classes) | Faster RCNN-res101 with k-means clustering. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
98.54% mAP | The proposed technique for disease detection demonstrates faster detection speed than the original R-CNN. | - The approach may require significant computing resources and extensive training to achieve optimal results. - The given image reveals the detection of a disease on a leaf. |
To improve the accuracy of tomato disease identification and position detection using deep learning techniques. | - Future research could focus on using more sophisticated deep learning techniques and data enlargement to achieve greater accuracy in detecting and classifying plant diseases. - Explore the integration of additional data sources to improve disease detection accuracy. |
| Gangwar et al. [23] | Grape Crops (4 classes) | Transfer learning with the Inceptionv3 model followed by classifiers like logistic regression, SVM, and neural networks. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
99.4% | -Transfer learning allows leveraging pre-trained models, reducing training time and resource requirements. -The automated system enables accurate effective detection of grape diseases, thereby facilitating timely treatment efforts. |
- The approach does not include the location of the diseases on the vine leaves, which limits the analysis. - Dependence on the quality and relevance of the pre-trained model. |
- The objective is to develop a solution for the classification of vine diseases using transfer learning and various classifiers. | - Future research could explore segmentation techniques to improve disease localization on grape leaves. - Learn how these recent CNN architectures can be applied to specific tasks such as plant disease detection and further improve their performance. |
| Agarwal et al. [28] | Tomato (10 classes) | Convolutional Neural Network (CNN) |
Learning Rate: 0.001. Momentum :0.999. Batch Size: 64. Epochs: 1000. |
91,2% | -The proposed method offers an original approach to effectively manage diseases in tomato crops through image analysis, potentially helping farmers manage them promptly. -The model provides accurate and efficient disease identification. |
-The study may be faced with limitations in generalizing the results to different environmental or disease conditions outside of the dataset used. -Deep learning models can be computationally intensive. |
-To provide practice for farmers to detect and control diseases of tomato plants, thereby improving crop quality and yield. -Detect and predict diseases in tomato plant leaves using a deep learning-based approach. |
-Researchers could explore hybrid architectures combining them with other types of neural networks for better performance. -Explore the integration of real-time disease detection systems using CNN for field application. |
| Magsi et al. [15] | Palm (4 classes(stages) | -Convolutional Neural Networks (CNN). -Texture and color extraction methods. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: stopped at epoch 6 (out of a maximum of 1000). |
89,4% | The system has achieved accuracy rates in identifying disease of date palm, in advanced stages, which can facilitate timely intervention and disease management. | -Image processing may be computationally intensive and resource-consuming. - The early stages of identification showed lower success rates, indicating the need to further refine the detection capabilities. |
The main objective of the research was to develop an automatic disease identification system for the date palm to address losses related to date palm cultivation. | The authors could explore transfer learning to enhance disease identification accuracy or investigate the impact of different preprocessing techniques on model performance. |
| Aversano et al. [25] | Tomato (10 classes) | VGG-19, Xception, and ResNet-50. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: 50. |
VGG-19 : 97%. Xception : 95%. ResNet-50 : 60%. |
The use of CNNs and learning in the study allowed the detection and classification of tomato leaf diseases, thus contributing to the early treatment of plant pathologies. | - One of the models used, ResNet-50, does not perform as well as the others in terms of accuracy, requiring different optimization or further exploration models. -Constantly monitoring plants manually is time-consuming. |
The main objective of the study was to demonstrate the effectiveness of CNN and transfer learning in the automatic detection and classification of tomato diseases, improving thus food security and reducing crop losses. | In the future, it would be interesting to extend the dataset used in the study to include a larger number of classes and to improve the precision models for the detection and classification of plant diseases. |
| Ouhami et al. [55] | Tomato (6 classes) | -DensNet161. -DensNet121. -VGG16. -Transfer learning |
Learning Rate: 0.005. Optimizer: Stochastic Gradient Descent (SGD). Batch Size: Not specified. Epochs: 20. |
DenseNet161 : 95.65%, DenseNet121 : 94.93%, VGG16 : 90.58%. | The DenseNet models required fewer parameters to achieve better performance. DenseNet161 showed superior accuracy in classifying leafminers and powdery mildew (100%). | -The limited size of the data set, which consists of 666 images, could potentially compromise the generalizability of the results. -Misclassification due to the similarity of symptoms (e.g., confusion between early blight and late blight for DenseNet161). |
-To enhance crop protection through the precise identification and classification of tomato diseases using machine learning. -To evaluate and compare deep learning models for detecting tomato diseases in standard RGB images and determine the optimal performance of these models. |
Firstly, the augmentation of the dataset to ensure a more substantial sample size; and secondly, the identification and resolution of more challenging disease detection problems. |
| Chen et al. [56] | Rice (5 classes). Maize (4 classes). | VGGNet, Transfer Learning |
Learning Rate: Not specified. Optimizer: Stochastic Gradient Descent (SGD). Batch Size: Not specified. Epochs: 30. |
Public Dataset (PlantVillage - Maize): 84.25% average prediction. Collected Dataset (Maize): 80.38% average prediction. Collected Dataset (Rice): 92.00% average prediction. |
Using transfer learning from pre-trained models helps improve plant disease identification performance, particularly with limited training data. | -A potential limitation could be the need for significant computing resources to train learning models. - Classical approaches heavily rely on hand-designed features, which can be expensive and require expert knowledge. |
- The main objective is to develop a system for monitoring and identifying plant diseases for agricultural productivity. -To enhance the learning ability of tiny lesion symptoms while decreasing computational complexity using transfer learning for deep CNNs. |
- Involve extending the application of the developed model to other plant diseases and diseases for greater agricultural impact. -Focus on improving the learnability of deep learning algorithms for detecting plant diseases under various field conditions. |
| Chen et al. [57] | Rice (3 classes for Public Dataset (UCI), 13 classes for Collected Dataset) | DenseNet with the Inception module + transfer learning, |
Learning Rate: Not specified. Optimizer: Stochastic Gradient Descent (SGD). Batch Size: Not specified. Epochs: 30. |
The proposed approach has achieved a prediction accuracy of at least 94.07% in the public data set and an average accuracy of 98.63% for image class prediction of rice diseases. | - superior performance in detecting diseases with high accuracy rates. -The deep learning approach demonstrates superior performance compared to other state-of-the-art methods. |
- Limited discussion of the generalizability of the model to various environmental conditions. - Conventional visual-based disease identification of rice by experts can be costly and time-consuming. |
-Develops a rapid, automatic, accurate, and cost-effective method for detecting rice diseases using deep techniques. -Provides a rapid, automatic, cost-effective, and accurate method for the detection of rice diseases in the field of agriculture. |
- Study of integration of real-time monitoring systems and technology for early detection and management of rice plant diseases. -Future research could explore improving model robustness to variations in environmental conditions and rice differences. |
| 2021 | ||||||||
| Jeyalakshmi et Radha [29] | Tomato (4 classes) | -Support Vector Machines (SVM). -Random Forest (RF) -Multilayer Perceptron Neural Networks (MPNN) -Soft Voting |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
The highest accuracy achieved in this study was 93.13% using the Soft Voting Classifier. | - The ensemble learning approach combining multiple classifiers enabled the accuracy of disease classification. | The study may have limitations in terms of larger data sets or real-time ones. | - Develop an accurate and robust classification system for tomato diseases using ensemble learning techniques. -To accurately classify various tomato diseases to facilitate early detection and management. |
- Explore the application of ensemble learning techniques to classify diseases in other plant species such as corn, corn, and apples. -Investigate data augmentation, new learning techniques, or integration of additional data sources to improve the robustness of the classification of diseases of the tomatoes. |
| Kibriya et al. [30] | Tomato (4 classes) | -VGG16 -GoogleNet |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
-98% for VGG16. -99.23% for GoogleNet |
- The deep learning approach using CNN models provides high accuracy rates for the detection of tomato leaves. |
- Limited information on model scalability and generalizability. - CNN models require large datasets. Additionally, system implementation and maintenance may require significant resources and expertise. |
- The main objective of the study is to develop a reliable solution for the early detection of tomato diseases to prevent losses of production. | - Exploring the integration of real-time monitoring and automated treatment recommendations based on disease detection results. -To develop an effective solution for tomato leaf disease detection using deep learning approaches. -To explore transfer learning with larger and more diverse datasets could enhance the accuracy. |
| El Massi et al. [42] | Tomato (6 classes) | Firstly, the Serial Combination (SC) method is employed. - Two SVM (Support Vector Machine) classifiers (color → texture/shape). 2. Hybrid Combination (HC): - Three SVMs (serial + parallel). Other techniques that may be employed include: - k-means clustering (segmentation). The features employed included colour moments (RGB/HSV), GLCM texture, and shape descriptors. -CNN. |
Learning Rate: 0.001. Optimizer: Adam. Momentum :0.1. Batch Size: Not specified. Epochs: 20. |
91.11%. | -The hybrid method has been developed for the purpose of handling class similarity, for example in cases of color overlap. It combines multiple features, including but not limited to colour, texture and shape. In addition, it has been designed to reduce the limitations of individual classifiers. | -The complexity of the thrips class is attributable to the varying damage characteristics exhibited by the organism. -The accuracy of the segmentation process is a prerequisite for effective classification. -The limited size of the dataset has a detrimental effect on the performance of convolutional neural networks (CNNs). |
The automatic recognition of plant diseases and damage is facilitated by the utilization of classifier combinations, which serve to address issues pertaining to class similarity. | -The following improvements are recommended for the hybrid method for complex classes: -The method should be improved, for example of thrips. -The dataset should be expanded. -Additional features should be incorporated, for example spectral data. |
| Rosmala et al. [58] | Potatoes (3 classes) | -VGG16 -InceptionV3 -Transfer learning |
Learning Rate: 0.0001. Optimizer: Stochastic Gradient Descent (SGD). Batch Size: 32. Epochs: 100. |
The VGG16 model demonstrated exceptional performance, achieving average precision, recall, and F1 score of 97% as well as a perfect precision rate of 100% on test data. | -The effectiveness of deep models to accurately classify plant diseases. -The VGG16 model showed better generalization of data compared to InceptionV3. |
-The need for training data for robust performance. -InceptionV3 had a slightly lower accuracy compared to VGG16. |
-To revolutionize the detection of diseases in agriculture. -To classify potato leaf diseases efficiently using deep learning models. |
Future research directions may involve expanding data to include other types of vegetable diseases to further support the agricultural industry in vegetable crops. |
| Wagle et R [59] | Tomato (9 classes) | -Transfer learning -AlexNet -VGG16 -GoogLeNet -MobileNetv2 -SqueezeNet |
Learning Rate: 0.0001. Optimizer: Not specified. Batch Size: 10. Epochs: Not specified. |
AlexNet 97.69%. VGG16 98.77%. GoogLeNet 93.73% MobileNetv2 95.25%. SqueezeNet 90.86%. | High accuracy with VGG16 using transfer learning and data augmentation, reducing overfitting. Deep learning enables efficient and accurate tomato leaf disease classification, minimizing manual effort and saving time. |
-A lengthy training period is required, particularly when utilising a restricted dataset. -The focus is exclusively on pre-trained models, neglecting the development of bespoke architecture. -VGG16 exhibits a higher execution time in comparison to models such as AlexNet. |
To employ deep learning models for the classification and validation of tomato leaf diseases, with the aim of achieving real-field data validation. | Future research will concentrate on extending the dataset, optimising model complexity while maintaining accuracy, and validating deep learning models for tomato leaf disease classification using real field data. |
| Ashwinkumar et al. [60] | Various (5 classes) | - OMNCNN (optimal mobile network-based convolutional neural network). -bilateral filtering-based preprocessing. -Kapur’s thresholding-based image segmentation. -MobileNet-based feature extraction. -extreme learning machine-based classification. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
The paper reports an accuracy of 98.7% for the proposed OMNCNN model. | The automated plant leaf disease detection model using OMNCNN offers superior performance with high precision, recall, accuracy, F-score, and kappa values | - The proposed model can require further validation on larger and larger datasets to assess generalizability. -This article does not explicitly mention the potential challenges to real-time implementation or the scalability of its OMNCNN model in an agricultural environment. |
-Develop an optimal model of MobileNet-based convolutional neural network for automated detection and classification of plant leaves. -Simplify and streamline the detection of plant diseases for farmers, because plant diseases constitute an important factor for the global economy. |
-Could focus on improving the detection efficiency of the OMNCNN method using advanced deep learning-based image segmentation techniques. -Investigate the integration of distinct image preprocessing techniques for more efficient extraction of features in plant disease models. |
| Wagle et al. [61] | Tomato (9 classes) | -ResNet50 -ResNet18 -ResNet101 -Transfer learning. |
Learning Rate: 0.0001. Optimizer: Not specified. Batch Size: 10. Epochs: 2. |
ResNet101 achieved an accuracy of 99.99% in testing and 95.83% in validation. | -The use of deep learning models such as ResNet50, ResNet18 ResNet101 allows very accurate classification of tomato diseases. -By augmenting the noise, blur, and color data, the dataset becomes very robust, which can improve the classification accuracy. |
The study does not discuss in detail the complexity or training time associated with the learning models used. | -The main objective of the study is to investigate the impact of increased data on the classification and validation of tomato plant diseases using deep learning methods. - Confirm the accuracy of a model designed for plant disease identification. |
- Future research could explore the scalability and generalizability of proposed deep learning models and augmentation techniques on different plant species for disease detection. -The exploration of new data augmentation techniques or the exploration of deep learning architectures could further improve the identification and classification of plant diseases. |
| Hassan et Abdulazeez [62] | Various: apple, rice, grapes, potato, sugarcane, and tomato. | -Naive Bayes -Decision Trees -Support Vector Machines (SVM) -Random Forest |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
SVM has the highest accuracy of 82.3%. | The study uses a range of classification techniques to detect plant leaf diseases, thus providing comprehensive analysis of different methods to improve the identification of diseases. | The time required to calculate the disease in the infected leaves is minimized, but memory consumption remains a problem. | Assist in the early identification of plant diseases and implement preventive measures to increase crop yield. | -Research could involve exploring the integration of deep learning techniques, such as convolutional neural networks, more accurate detection and more effective from plant leaf diseases. -Future research could focus on improving memory consumption while maintaining accuracy in disease detection systems. |
| Abbas et al. [63] | Tomato (5, 7, 10 classes) | -Conditional Generative Adversarial Network (C-GAN). - DenseNet121. |
Learning Rate: 0.0001. Optimizer: Adam. Batch Size: 32. Epochs: 100. |
-99.51% for 5 classes. -98.65% for 7 classes. -97.11% for 10 classes. |
The use of synthetic images generated by C-GAN improves generalizability of the network and avoids overfitting, leading to increased accuracy in disease classification. | Using deep learning models can require significant computing resources and expertise for implementation and training. | The objective of the study is to develop a deep learning-based method for accurate and early detection disease of tomato plants, outperforming existing methods. | Aims to involve extending the proposed method to identify diseases in various parts of the plant beyond just leaves, such as fruits, stems, and branches, as well as to explore the identification of different phases of the plant diseases. |
| E.H. Chowdhury et al. [64] | Tomato (2,6,10 classes) | -ResNet18. -DenseNet201. -InceptionV3. |
Learning Rate: 0.001. Optimizer: Adam. Batch Size: 16. Epochs: 15. |
-99.2% for binary classification. -97.99% for classification in six classes. -98.05% for classification ten classes. |
The study surpasses existing cutting-edge work in the field of plant disease detection using deep learning techniques. | Despite high accuracy rates, there were cases of misclassification, as indicated by the confusion matrix analysis. | The main objective is to study the effectiveness of CNN architecture in classifying images of tomato leaves for disease detection. | - Explore the reliability of leaf images across diverse classes of extended images to improve disease detection systems. -Explore the integration of real-time monitoring systems for early detection of tomato plants. |
| Wang et Liu [65] | Tomato (12 classes) | YOLO-Dense. |
Learning Rate: 0.0026. Optimizer: Not specified. Batch Size: 64. Epochs: Not specified. Momentum :0.9. Decay:0.0054. Factor:0.1. |
The model achieved a Mean Average Precision (mAP) of 96.41%. | YOLO-Dense offers rapid and accurate detection of tomato anomalies in complex environments. | Discussion on the scalability of the YOLO-Dense model to larger data sets or different data types is limited, which could affect its generalizability. | To enable precise and real-time identification of tomatoes anomalies to improve crop quality and yield. | Exploring the scalability and adaptability of the YOLO-Dense algorithm to detect anomalies in various species beyond tomatoes. |
| Feng et al.[66] | Rice (4 classes) | -Transfer learning. -Fine-Tuning. -Deep CORrelation ALignment (CORAL). - Deep Domain Confusion (DDC) |
Learning Rate: 0.0001. Optimizer: Not specified. Batch Size: 40. Epochs: Not specified. |
88%. | Deep transfer learning methods have shown promising results in effectively and cost-effectively detecting rice diseases in various rice varieties in the field. By combining hyperspectral imaging with deep transfer learning, rice diseases can be accurately and effectively classified, providing a potential tool for early disease detection on farms. | The development of classifiers for each rice variety of time requires many resources, while limited variability of data and cultivars could restrict generalizability and performance of transfer learning models | This study examines the feasibility of using data and deep transfer learning for accurate detection of rice diseases, with the aim of improving the performance of model thanks to the expansion of data and to facilitate prevention and precise disease. | Future studies could focus on two areas: to improve deep transfer learning models to detect rice diseases by including a wider range of rice types and diseases and use this transfer learning advances in conjunction with hyperspectral imaging detect diseases in various plant species, extending their agricultural. |
| 2022 | ||||||||
| Nawaz et al. [27] | Tomato (10 classes) | ResNet-34 based Faster-RCNN |
Learning Rate: 0.001. Optimizer: Not specified. Batch Size: 8. Epochs: 20. Threshold for matched region: 0.2. Threshold for unmatched areas: 0.5. |
99.97%. | The proposed deep learning (DL)-based approach, specifically the ResNet-34-based Faster-RCNN, achieves high accuracy in disease detection and localization. | Deep learning models require large amounts of labeled data for training, which can be difficult to obtain for some plant diseases. This may further limit their effectiveness in handling noisy or complex scenarios. | Develop a robust deep learning approach for accurate localization and classification of leaf diseases of tomato plants. | explore the integration of multi-sensor data fusion techniques with transfer or multispectral learning to improve model robustness, generalization capabilities and detection accuracy diseases various environments. |
| Nagamani et Sarojadevi. [32] |
Tomato (7 classes) | -Fuzzy Support Vector Machine (Fuzzy-SVM). -Convolution Neural Network (CNN). -Region-based Convolution Neural Network (R-CNN). |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: Not specified. |
96.735%. | The application of R-CNN, a deep learning technique, this study allowed us to obtain high precision in the detection of tomato leaf diseases. This could potentially improve crop productivity by earlier identification of diseases, which could reduce future losses, pesticide use and pollution. | Traditional manual methods are not suitable for large-scale agriculture, mainly due to difficulty in identifying and controlling plant diseases. However, deep learning models also require considerable resources for training and deployment. | The objective of the study was to detect leaves of tomato plants at one stage using machine learning techniques, thereby increasing agricultural production, and minimizing losses. | Future work could focus on expanding the data set for wider applicability and exploring improvements to learning models in depth to achieve greater accuracy in disease prediction. |
| Al-Gaashani et al. [33] | Tomato leaf (6 classes) | -MobileNetV2. -NASNetMobile -Multinomial logistic regression (MI.R). |
Support Vector Machine (SVM): - Penalty parameter C: 0.1. - Gamma parameter: 0.001. - Kernel: Linear Random Forest (RF): - Number of decision trees: 400. - Depth of each decision tree: 70. Multinomial Logistic Regression (MLR): - C parameter: 0.1. - Penalty: 12. - Optimiser: 'lbfgs'. |
97%. | Using pre-trained models not only improves accuracy, also reduces the need for extensive training data. Additionally, integrating feature fusion with transfer learning models with reduction via Kernel PCA can potentially further improve classification accuracy. | - The study recognizes the potential impact of image acquisition conditions and generalizability limits of pre-trained models which may not cover all disease variants. Additionally, traditional machine learning methods can outperform deep in situations where training data is scarce. | This study aimed to develop a model that automates the process of classifying widespread leaf diseases thereby helping farmers to make an effective accurate diagnosis. | Future work could focus on expanding the disease detection capabilities of model to encompass new types and explore techniques aimed at improving the robustness of real-world agricultural applications. |
| Khasawneh et al. [67] | Tomato (10 classes) | Deep transfer learning: DenseNet-201, SqueezeNet, GooglLeNet, Inceptionv3, MobileNetv2, ResNet-101, ResNet-50, ResNet-18, Xception, ShuffleNet and DarkNet-53. |
Learning Rate: 3×10−4 Optimizer: Stochastic Gradient Descent with Momentum (SGDM). Batch Size:16. Epochs: 5. |
99.4%. | Deep transfer learning models simplify the process of disease detection and classification in tomatoes. By bypassing the need for explicit feature extraction and image preprocessing, these models facilitate rapid diagnostics and potentially mitigating economic losses in tomato cultivation. | Deep transfer learning models offer advantages in disease detection, but their resource-intensive nature poses challenges in resource-constrained environments | The research focuses on creating a system for automated detection and classification of tomato diseases using deep transfer learning. This system is designed to help farmers quickly identify diseases by streamlining the identification process. | Future avenues of research could focus on the real-time disease detection systems with smart agricultural devices for immediate deployment and implementation in the field. This could include the creation of smartphone applications allowing plant pathologists and farmers to detect and manage diseases in real time. |
| Lakshmanarao et al. [34] | Tomato, Potato, and Pepper bell (15 classes) | VGG16, RESNET50, and Inception. |
Learning Rate: 0.0001. Optimizer: Adam. Batch Size: 64. Epochs: Not specified. |
99%. | The proposed model achieved higher accuracy rates than traditional models, demonstrating the effectiveness of transfer learning for plant disease prediction. This technique exploits pre-trained models, allowing training even with limited data. | A salient disadvantage is the overreliance on numerous labelled data points to facilitate the efficient training of deep learning models. Moreover, the efficacy of transfer learning in adapting to variations in plant diseases may be inconsistent. | The primary objective of this study is to demonstrate the application of transfer learning techniques for accurate plant disease prediction, underscoring the significant advantages it offers to the agricultural sector. | Future research directions could include the study of and generalizability of transfer learning techniques to a wider range of plant species, which could increase the applicability of the model in various agricultural contexts. Additionally, the use of ensemble techniques could further improve disease prediction accuracy. |
| Vallabhajosyula et al. [24] | 14 different crops: -Tomato, Potato, Grape… (38 classes) |
-Deep Ensemble Neural Networks (DENN): - ResNet 50 & 101, InceptionV3, DenseNet 121 & 201, MobileNetV3, and NasNet. -Transfer learning, data augmentation |
Learning Rate: 0.001. Optimizers: Adam, Adamax, Adagrad, SGD (Stochastic Gradient Descent), Nadam, and RMSprop. Batch Size: 8. Epochs: 30. Momentum:0.9. Regulazer: L2 with factor 0.01. |
The suggested DENN can achieve 100% accuracy. Here are the baselines: -InceptionV3: 98.33% -VGG16: 99% -GoogLeNet: 99.35% -DenseNet121: 99.75% |
The implementation of a deep ensemble neural network (DENN) with transfer learning has been demonstrated to enhance the accuracy of categorizing various plant species to a significant degree, surpassing the capabilities of state-of-the-art, pre-trained individual models. -The approach is capable of handling overfitting via means of data augmentation and regularization. |
-Deep transfer learning (DTL) and deep ensemble models (DEM) necessitate substantial computing resources and voluminous datasets for efficacious training. -The substantial computational and data requirements of these methods present significant challenges for implementation in environments with limited resources. |
Using deep clustering networks together with transformative learning means that leaf disease can be detected more quickly and efficiently. This means that diseases can be detected earlier, which means we can produce more crops. |
-Combine different types of data, like spectral imaging and drone monitoring, to better detect plant diseases. -Develop systems that work in real-time for practical farming applications. -Implement solutions in real-world scenarios, like mobile apps. -Extend the system to cover more plant species. -Improve the system's computing performance. - Explore using lightweight structures to make it more efficient. |
| Ahmed, et al. [68] | Tomato (10 classes) | Transfer learning with pre-trained model MobileNetV2. |
Learning Rate: 10−5. . Optimizer: Adam. Batch Size: 16. Epochs: 1000 (with early stopping). |
99,30%. | The proposed architecture achieves high classification accuracy 99.30% with small model size and computational cost, which makes it suitable for low-end devices. | -The model had misclassifications, particularly diseases like downy mildew and powdery mildew. The need for further refinement of the model, particularly considering the challenges faced by previous methods of classifying tomato leaf diseases when dealing with larger data and complex pre-processing steps. |
-Develops a lightweight and efficient neural network architecture for transfer learning for the classification of tomato diseases. -Reduce crop yield loss, eliminate manual monitoring and minimize human effort in disease control while ensuring calculation efficiency and effectiveness. |
Future directions of research could include the study of ensemble learning to improve the classification performance and robustness of the architecture for complex disease scenarios, as well as optimization of lightweight models for efficient disease classification on low-end devices. |
| Nguyen et al. [69] | Tomato (10 classes) | VGG-19 model with transfer learning and image segmentation using the HSV color space. |
Learning Rate: 0.00001. Optimizer: Not specified. Batch Size: 60. Epochs: 300. |
99,72%. | The proposed model excels in classifying tomato leaf diseases due to its high accuracy, which is further enhanced by the segmentation of leaf images. This not only makes it effective for disease detection and classification but also optimizes the training time. | The mosaic virus disease type had lower disease, which can be attributed to the limited dataset. This indicates that implementing techniques such as oversampling or balanced weighting of classes improve the accuracy of the results. | The research aimed to develop a classification model using image segmentation and transfer learning techniques to accurately identify tomato leaf diseases, thereby improving performance and overall effectiveness of disease classification. | Future research around disease classification in tomato plants may involve advances in image processing, and incorporation of larger data sets for training and optimization model parameters. Additionally, the use of strategies such as oversampling or balanced class weighting methods improve accuracy, particularly for diseases represented in smaller datasets. |
| Al-Akkam et Altaei [70] | Various: Tomatoes, potatoes, Pepper bell … (15 classes) |
Image processing techniques with Convolutional Neural Network (CNN). |
Learning Rate: 0.001. Optimizer: Adam. Batch Size: 32 and reduced to 16. Epochs: 120. |
98.34%. | The study highlighted the effectiveness of learning techniques in agriculture by demonstrating higher accuracy rates in the detection and classification of plant leaf diseases, surpassing those of previous studies. | The dataset does not explicitly mention the specific types of plants or diseases it includes, potentially limiting the results' generalizability. | The main objective of this research is to establish a methodology to identify and classify leaf diseases as well as to accurately predict these diseases using deep learning techniques, which will facilitate treatment and control economic losses. | -Future research could integrate real-time monitoring systems with mobile applications for immediate detection and management of plant diseases. -Experiments with different learning rates, data variations expert systems could further improve the practical application of these for the identification and classification of plant diseases. |
| Zia Ur Rehman et al. [75] | Citrus (6 classes) | -MobileNetv2, DenseNet201, Whale Optimization Algorithm (WOA), transfer learning, SVM. |
Learning Rate0.001. Optimizer: Not specified. Batch Size: 64. Epochs: Not specified. Momentum: 0.93. |
95.70%. | -The proposed method outperforms recent techniques in terms of classification accuracy of citrus diseases. -This success is attributed to the use of models for learning efficient optimization algorithms. |
A notable disadvantage of deep learning techniques is the need to have a substantial volume of labeled data to train efficiently model. | -The study uses deep learning and transfer learning to improve the accuracy and efficiency of citrus fruit classification. -By classifying six diseases of citrus fruits, the model aims to optimize overall production and disease management practices. |
Future research directions may involve expanding the data set to encompass a wider variety of citrus diseases and potentially include multiple fruits. Particular attention could also be paid to the development of real-time disease detection systems for use in agricultural fields. This would help refine the classification of diseases. |
| Boutalline et al. [71] | Apple (9 classes) | MobileNet V2 |
Learning Rate: 0.001. Optimizer: Adagrad. Batch Size: 16. Epochs: Not specified. |
98%. | The application of MobileNet V2 and CNNs has significantly improved the accuracy of leaf disease identification, reaching a performance rate above 98% while also reducing the need for image preprocessing. | Limitations of the study include the focus on specific diseases, which may require application to other areas or diseases. Additionally, it does not consider different environmental conditions on disease detection. | The research aimed to equip farmers with a system for early detection and classification of apple leaf diseases, thereby improving crop quality, use of chemicals and minimizing impacts environmental. | -Use a larger dataset, apply it to various regions, and use advanced deep learning techniques to improve accuracy. -Explore the integration of temporal surveillance systems for proactive disease control. |
| Zhang et al. [72] | Tomato (4 classes) | MMDGAN (Multi-Feature Extraction Convolution GAN with Mixed Attention and Markovian Discriminator). |
Learning Rate: 0.00005. Optimizer: ADAM. Batch Size: 64. Number of Epochs: 1500. |
97.12%. | The MMDGAN method presents a new approach for the identification of diseases of leaves of tomato, which not only improves the quality of the data set, also shows superiority in terms accuracy compared to existing methodologies. | A notable drawback is the limited discussion of the method proposed for other plant disease datasets, coupled with the limitations inherent in data augmentation methods that require a substantial collection effort. | The study aimed to develop a robust method of augmentation using MMDGAN to improve the identification of tomato leaf diseases. | Future research in this area could benefit from applying MMDGAN to various plant disease datasets, as well as exploring transfer techniques to improve classification. diseases, evaluating its effectiveness in a broader agricultural context. |
| 2023 | ||||||||
| Hajraoui et al. [73] | Tomato (5 classes) | VGG16 and ResNet152v2 with transfer learning |
Optimizer: Adam. Learning Rate: 1 e-4. Batch Size: 32. Epochs: 175 |
99.0234% | The proposed model achieves high classification accuracy of tomato leaf diseases through deep learning and transfer learning. Additionally, the model demonstrates its effectiveness in training and testing. | -Limited dataset size. -The need for careful tuning of hyperparameters, such as learning rate, to solve the problem of overfitting, which requires a significant amount of trial and error. |
Develop a deep learning model that improves disease control in tomato plants, thereby helping maintain high yields and quality through accurate classification of diseases on tomato leaves. | -Study the scalability of the model to a wider range of plants species. - Solve the problems of classifying more complex diseases and apply the model to a wider range of patients. |
| Hessane et al. [74] | Palm (4 classes) | Machine Learning Methods like: -Support Vector Machine (SVM). -k-Nearest Neighbors (KNN). - Random Forest (RF). -Light Gradient Boosting Machine (LightGBM). |
Not specified | 98.29% | The framework uses data augmentation and combines texture and color features to improve palm disease classification. This allows early detection of mealybugs, essential for protecting date crops. | Reliance on image data limits the accuracy of disease detection. Additionally, deep learning methods are hampered by limited or imbalanced datasets, which hampers training and identification. | The objective is to develop a reliable machine learning tool for the detection and classification of white scale insect diseases in palm trees. | -Explore deep learning techniques such as convolutional neural networks for improved feature extraction and disease classification. -Expand the palm disease data set and integrate more sophisticated learning methods to improve detection accuracy. |
| Parvez et al. [31] | Tomato (3 classes) | Convolutional Neural Network (CNN). |
Learning Rate: Not specified. Optimizer: Adam. Batch Size: Not specified. Epochs: 50. |
98.39%. | Convolutional neural networks enable accurate automated prediction of tomato plant diseases. This technology facilitates early detection, thereby avoiding substantial harvest costs and reducing the need for manual inspection. | A significant limitation of this approach is that training data can hinder the model's ability to generalize, highlighting the need to resort to data augmentation techniques to improve performance. | The objective of this research is to develop a comprehensive approach to the early detection and effective treatment of plant diseases, with a particular focus on leaf diseases affecting tomato crops. The objective is to enhance productivity and ensure the production of high-quality tomatoes, thereby increasing profitability. | Future research directions include the integration of real-time disease detection systems into practices for the rapid treatment of diseased plants, the exploration of convolutional neural network (CNN) architectures and techniques, data augmentation, and the application of CNN models to detect diseases in different crops, with the aim of increasing agricultural productivity. |
| Attallah O. [35] | Tomato (10 classes) | -KNN (Knearest neighbor) +Fully connected layer (MobileNet + ShuffleNet + ResNet-18) + hybrid FS. |
Learning Rate: 0.001. Optimizer: stochastic gradient descent with momentum. Batch Size: 10. Epochs: 20. |
99,92 % |
The advantage of the proposed pipeline lies in its CNN structures and feature selection, which simplifies the model compromising the high accuracy rates in tomato leaf diseases classification. | Limitations of the study include the reliance on laboratory capture, which may affect the applicability of the model to real-world scenarios, and the lack of exploration of the impact of various CNN architectures on beyond the three selected compact CNNs. | The objective was to develop an accurate and robust deep learning pipeline for the automated detection and classification of tomatoes. | Explore the application of field data in real-time disease detection and expand the pipeline for a greater variety of plant diseases for classification purposes. |
| Borugadda et al. [36] | Tomato (10 classes) | -Transfer learning with the VGG16 architecture. - Filter methods, Principal Components Analysis (PCA), and the Boruta feature selection method. |
*For the VGG16: - Learning Rate: 0.0001 - Optimizer: Stochastic Gradient Descent (SGD) - Batch Size: 8. - Epochs: 97 *For the Multi-Layer Algorithm (MLA) such as Support Vector Classifier (SVC): - C: 10 - Gamma: 0.0001. - Kernel: rbf’. |
95.79% | The new algorithm effectively solves problems such as overfitting and long training, thereby improving the diagnostic accuracy of leaf diseases of tomato plants. | Implementing multi-level dimensional reduction techniques can be complex and resource-intensive, there is a risk of overfitting with high-dimensional data. | This research explored the use of VGG16 transfer learning for leaf disease classification. They aimed to optimize feature extraction using a dimensionality reduction algorithm to improve disease detection. | Future work could explore the scalability of the proposed model for larger datasets and investigate its generalizability to other plant species for disease classification. |
| Kaur et al. [37] | Tomato (8 classes) | - Modified InceptionResNet-V2 (MIR-V2) with transfer learning |
Learning Rate: 0.0001. Optimizer: Adam. Batch Size: 32. Epochs: 50. |
98,92%. |
-high accuracy in detecting tomato leaf diseases | The approach relies on deep learning techniques, which may require substantial computational | To develop an effective computer-aided disease detection system for plant leaves. | Exploring the extension of this approach to other crop types and expanding the dataset could enhance disease detection accuracy. |
| Liu et al. [38] | Various: tomatoes, peppers, potatoes… (15 classes) | - Selective Kernel MobileNet (SK-MobileNet) |
Learning Rate: 00.0001. Optimizer: RMSProp with decay and momentum set at 0.8, and the Adam algorithm, with and set at 0.9 and 0.999 respectively. Batch Size: 32. Epochs: 50 for the model, with an early stop mechanism selecting 100 epochs. |
99,28 % | SK-MobileNet combines efficiency and accuracy in a model thus reducing costs and complexity. | Complex backgrounds decrease the accuracy of SK-MobileNet and DCGAN. Additionally, the method requires a lot of calculations. | The research aims to improve the recognition of plant diseases by developing a lightweight adaptive network that uses deep transfer learning and convolution techniques. | Improve SK-MobileNet for robust disease detection in complex agricultural settings; focus on the scalability and generalization of the model. |
| Nigam et al. [40] | Wheat (Triticum aestivum) (4 classes) | EfficientNet, EfficientNet B4, VGG19, ResNet152, DenseNet169, InceptionNetV3, MobileNetV2. |
Learning Rate: 0.001. Optimizer: Adam. Batch Size: 32 for B0-B5 models, 8 for B6-B7 models. Epochs: Ranged from 15 to 25, with a maximum defined epoch of 50 |
99.35%. | The model provides high-level wheat disease identification and actionable information for agricultural professionals, supported by an extensive set of innovative architecture. | The deployment of expertise and the utilization of high-performance graphics processing units (GPUs) are indispensable prerequisites for the successful execution of this task. However, the constraints imposed by limited resources establish a maximum size and scalability for the model. | The study develops a transfer learning model to identify rust diseases of wheat, thereby contributing to deep learning for agricultural disease diagnosis. | The model identifies diseases of wheat rust. Future work aims to predict the severity of the optimized use of pesticides and the integration of mobile applications for field diagnosis. |
| Nayak et al. [39] | Rice (Oryza sativa) (13 classes) | DenseNet 201, EfficientNetB0, InceptionV3, MobileNet, MobileNetV2, NASNetMobile, ResNet 101, ResNet 50, VGG16, VGG19, and Xception. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: Not specified. Epochs: 50. |
-DenseNet201 : 98.03%. | The research presents a cost-effective smartphone-based solution for immediate detection of diseases thereby improving accessibility for rural communities. | The effectiveness of the method is hampered by variable smartphone configurations, slow processing image segmentation, and insufficient training data for symptom detection. | The smartphone application for real-time detection of rice diseases and deficiencies targets intervention to improve crop health. | Optimize the application for various smartphones, expand plant analysis for health assessment and identify overlooked micronutrient deficiencies. |
| Bensaadi et Louchene [76] | Tomato (9 classes) | Convolutional neural network (CNN). -Data augmentation, -Stochastic gradient descent (SGD) with momentum. |
Learning Rate: η=7×10−1, η=7×10−3, η=7×10−4, and η=7×10−5. Optimizer: Stochastic Gradient Descent (SGD) algorithm with momentum. Batch Size: Not specified. Epochs: Not specified. |
97,04% | The study proposes an inexpensive and complex CNN architecture for a faster online classification of diseases of the plant. | The study addresses overfitting in a plant disease classification model using data augmentation and hyperparameter tuning; however, the scalability of the model has not yet been tested. | The aim is creation of a machine learning tool for the precise identification of plant diseases of tomatoes for farmers. | Improve model scalability and disease coverage, for real-time use, and incorporate advances to improve the efficiency of agricultural deep learning. |
| Isnan et al. [77] | Arbres, fruits et fleurs. (29 classes | Transfer learning with pre-trained CNN: EfficientNetB0, ResNet18, VGG19, and AlexNet |
Learning Rate: 0.0001. Optimizer: Adam. Batch Size: 16. Epochs: 50. |
EfficientNet-B0: 82.55%. | EfficientNet-B0 excels in agricultural crop classification thanks to its accuracy and computational efficiency ensured by transfer learning with minimal data. | The model had difficulty classifying crops within the same family due to their similar characteristics. | Research has studied the application of transfer learning to classify crops in Indonesia, recognizing its limitations and the need for further refinement. Its goal was to aid in the progress of identifying crops. | Future research will explore sophisticated unsupervised algorithms (SwAV) that exchange assignments between multiple views, to improve the accuracy. |
| Ramya et Kumar [78] | Tomato (10 classes) | -Deep transfer AlexNet CNN. -Batch normalization |
Learning Rate :0.001. Optimizer: Adam. Batch Size: Not specified. Epochs: 15. |
99.8% | The study uses deep learning techniques to accurately identify and categorize tomato leaf diseases. This approach allows us to obtain a high accuracy rate in disease detection. | This study could be limited by the need for a large volume of data to effectively train deep learning techniques. The accessibility of the data used could also be a problem. | The paper proposes a framework for continuous disease surveillance in agriculture using deep transfer learning. Thus, the objective is to identify plant diseases early to improve crop yield. | Future research could expand the application of learning models to encompass a wider variety of diseases in various crop species, while exploring detection and classification in real-time for wider agricultural use. |
| Shahoveisi et al. [79] | Various: sunflower, dry bean, and field pea. (3 classes) | ResNet50, Xception, EFficientNetB4, MobileNet |
Learning Rate: 0.001. Optimizer: Adam, Stochastic Gradient Descent (SGD), Root Mean Square Propagation (RMSprop), et Follow the Regularized Leader (Ftrl). Batch Size: 32. Epochs :100. |
94.29% |
Deep learning models were evaluated for plant rust disease accuracy. This approach could lead to more precise control of diseases and a reduction in the use of pesticides. | This study highlights the need for large-scale training data and sophisticated data equipment, which may limit its generalizability to other plant diseases. | The study aims to evaluate deep learning models for rust disease spraying and develop a practical solution for precision spraying using drones and handhelds. | Expand datasets: include various images (wheat, corn) to validate architectures and develop effective tools. Explore transferability: study model performance in agricultural environments for real-world use in precision agriculture. |
| Mimi et al. [80] | Bright Eyes (Catharanthus roseus) and Strawberry (Fragaria ×ananassa). | -Vanilla CNN model -CNN-SVM hybrid model. -MobileNetV2. -Transfer learning and data augmentation. |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: 64. Epochs: 10. |
97.35% | This research introduces a new set of data and Android applications for real-time health monitoring. The deep transfer learning model achieved high accuracy in disease identification, opening the door to automated disease detection. | The deep learning model for time-based monitoring of plant diseases does not consider lighting variations or image quality. Additionally, an unbalanced distribution of classes in the dataset will affect the accuracy. | The purpose of this study is to develop a computer vision using deep learning models to automatically classify and easily the leaves of diseased plants, thus effectively the gaps related to the use of such to detect plant diseases. | -Explore enhanced learning methods and study models such as ResNet, GoogLeNet, and EfficientNet. -Resolve class imbalance using resampling techniques to improve the accuracy of disease identification extending the applicability of the model to various crops. |
| Zayani et al. [81] | Tomato (3 classes) | YOLOv8 (You Only Look Once version 8), Data augmentation (mosaic augmentation) |
Learning Rate: Not specified. Optimizer: Not specified. Batch Size: 3. Epochs: 50. |
66.67% | The YOLOv8 model has been demonstrated to offer enhanced efficiency and flexibility in detecting tomato diseases, with the potential to improve crop yields. The model has been shown to exhibit efficient multiscale object detection, an anchor-free approach that improves adaptability, and high precision in confident detections | The dataset employed in the study demonstrates an imbalanced class distribution, a factor that has the potential to diminish the efficacy of learning systems. The presence of visual similarities between diseases, such as blossom end rot and sun scald rot, and the limitation of dataset diversity further complicate the process of accurate classification. The present accuracy of 66.67% indicates a substantial potential for enhancement, particularly with respect to differentiating among various disease types. | The utilization of YOLOv8, a deep-learning-based convolutional neural network, facilitates the automation of tomato disease detection, thereby enhancing crop yield and promoting sustainable agricultural practices in Saudi Arabia. | In addressing the issue of data imbalance, augmentation is a key strategy that should be employed. Moreover, the integration of a healthy class for the purposes of comparison is imperative. Exploration of alternative architectures, such as attention mechanisms, is also crucial. Finally, the development of disease-specific metrics is paramount. |
| 2024 | ||||||||
| Zahra et al. [82]& | Apple & Grape (8 classes (4 per fruit))& | Framework: Inception-ResNet-V2 (fine-tuned), DnCNN, Top-Bottom Hat Filtering, Entropy-based selection, Tree Growth Optimization, SVM Data Augmentation: Horizontal/Vertical Flip Feature Fusion: Serial entropy threshold.& |
Learning rate: 0.0001 Optimizer: SGD. Momentum: 0.6 Minibatch size: 16 Epochs: 100& |
Apple: 99.4\% Grape: 99.9\%& |
The system demonstrates high levels of accuracy, with automated disease detection and reduced computational time via feature selection. Furthermore, the incorporation of hybrid contrast enhancement has been shown to improve image quality.& | The fusion process has been shown to increase computational time, and thus requires the undertaking of preliminary processing steps, including, for example, augmentation and denoising.& | The objective of this study is to employ a combination of deep learning, feature optimization and fusion to achieve the classification of diseases affecting apple and grape leaves.& | The following techniques are worthy of note: -Intelligent fusion techniques -Encoder-decoder networks for feature extraction -Improved optimization algorithms.\\\hline |
| Naqvi et al. [83]& | Apple (4 classes)& | - Hybrid contrast enhancement (Bi-LSTM + Haze reduction) - Custom CNN models (BRwSA, IBRwSA) - Feature fusion + HLO optimization - SWNN classifier + LIME.& |
- Learning rate: 0.0002 - Optimizer: SGD. -Momentum: 0.702 - Mini-batch size: 64 - epochs :100 & |
94.8% & | The following improvements have been implemented: The enhancements have been designed to ensure enhanced accuracy in comparison to the SOTA (state of the art) benchmark, whilst concomitantly reducing the computational time required for post-optimization analysis. The second of these enhancements is intended to facilitate interpretability via the use of LIME (Limited Instance Machine Learning).& |
- Feature fusion increases computation time - Inverted bottleneck may lose critical features.& |
Develop a deep learning framework for accurate and efficient leaf disease recognition.& | - Lightweight vision transformers - Activation-based fusion - Dataset combination for robustness testing. \\\hline |
| Cucumber (5 classes) |
- HLO (Human Learning Optimization) parameters: 10 solutions, 100 interactions, validation ratio 0.3 - Same as Apple for training.& |
94.9\%& | The enhanced disease contrast facilitates improved feature extraction, while the optimized feature selection process ensures a more efficient utilization of resources.& | The dataset for cucumber is limited in size due to its private nature.& | Address the challenges associated with low-contrast disease recognition and high parameter pre-trained models.& | The following objectives are to be pursued: The generalizability of the system needs to be improved. Exploration is to be undertaken of the use of lightweight architecture. Additionally, there is a necessity to reduce the complexity of fusion.\\\hline |
||
| Al-Gaashani et al. [84]& | Multiple (Tomato, Coffee, Corn …) (18 classes)& | - MobileNetV2 - ResNet50V2 - Transfer Learning (TL) - Gravitational Search Algorithm (GSA) - MLR/KNN.& |
Not specified.& | - MLR + GSA: 99.2\%& | The advantages of this system are manifold. Primarily, evidence has demonstrated that it can reduce features by 50%. Secondly, it is highly accurate. Thirdly, it is both computationally efficient and capable of multi-level feature fusion.& | -There is a risk of overfitting. -The dataset is too homogeneous. -There is a dependency on specific pre-trained models. -The approach has not been tested on other plant species.& |
To develop a methodology for the early detection of plant diseases, with a view to enhancing food security. The proposed approach involves the implementation of an automated, resource-efficient classification system.& | -The dataset should be expanded in both diversity and size. -The efficacy of alternative pre-trained models should be tested. -A range of feature selection methods should be explored. -The model should be implemented in real-world agricultural settings.\\\hline |
| Abdul Aziz et al. [85]& | Rice (10 classes)& | - Convolutional Neural Network (CNN). - Transfer Learning (EfficientNetB0). - Data Augmentation - 5-fold Cross-Validation.& |
- Learning Rate: 0.1 (adjusted per epoch) - Optimizer: Stochastic Gradient Descent (SGD). Momentum:0.9. - Epochs: 100. - Batch Size: Not specified.& |
98.86\%& | -It exhibits high accuracy and low error rate. -It utilises parameters efficiently via EfficientNetB0. -It reduces training time and computational resources with transfer learning. -It handles dataset variability well.& |
-Relying on large, annotated datasets is suboptimal. -The testing data is limited (5% of the total dataset). -The potential computational costs for deep models are not discussed. -There is no explicit discussion of model interpretability.& |
to develop an automated rice leaf disease detection system using CNN (Convolutional Neural Network) and transfer learning to enhance the system's accuracy, efficiency, and applicability in agricultural technology.& | Future research will focus on: -The implementation of the system in real automated systems is imperative. -The study assumes that the concept under investigation should be extended to other cultures. -The implementation of test procedures on larger and more diverse data sets is imperative. -Integration of the system with mobile applications for use in the field is a crucial aspect that needs to be addressed.\\\hline |
| Shafik et al. [86]& | Multiple (Tomato, Maize, Apple, Potato, Strawberry…) (15 classes)& | The following are the algorithms under consideration: -PDDNet-AE (Early Fusion) -PDDNet-LVE (Lead Voting Ensemble) -Nine pre-trained CNNs: DenseNet201, ResNet101, ResNet50, GoogleNet, AlexNet, ResNet18, EfficientNetB7, NASNetMobile, ConvNeXtSmall -Logistic Regression (LR) classifier.& |
Learning rate: 0.1–0.001. - Optimizer: Adam. - Batch size: 10–100. - Epochs: 10. - Gradient threshold: 1 - Weight decay: 0.0001. - MB-SGD (Mini Batch Stochastic Gradient Descent) for optimization.& |
97.79\%& | Firstly, it is robust and generalizes well across diverse environments. Secondly, it is both computationally efficient and parsimonious in its use of parameters. Thirdly, it is capable of effective feature extraction using ensemble methods. Finally, it utilises natural background images (not controlled).& | -The presence of computational challenges on small devices has been noted. -The presence of class imbalance in datasets (mitigated by selecting 15 balanced classes) has been noted. -There is a dependency on hyperparameter tuning, which is to be expected in this field. -There is a limited capacity for real-world testing in natural settings, which is a limitation of the current study and will require further investigation in future.& |
The objective of this study is to develop efficient models for the detection and classification of plant diseases. These models will be developed using transfer learning and ensemble methods, with a view to enhancing agricultural sustainability.& | -The real-time collection of data and the detection of multiple objects (clusters of leaves) -The development of mobile and web-based applications for field deployment -The creation of lightweight models (quantization, vision transformers) -The addressing complex backgrounds and the challenges of localization.\\\hline |
| Bezabh et al. [87]& | Mango (6 classes)& | - Ensemble of GoogLeNet and VGG16-based CNN - Segmentation: K-Means, Mask R-CNN - Preprocessing: Image resizing, noise removal (GF, AMF), data augmentation - Classification: CNN + Softmax.& |
Learning rate: Not specified. - Optimizer: Not specified. - Batch Size: 64 - Epochs: 100.& |
99.21\%& | Primarily, it demonstrates a high level of classification accuracy. Secondly, it reduces computational complexity. Thirdly, it is effective in terms of segmentation (Mask R-CNN). Finally, it combines the strengths of GoogLeNet and VGG16.& | -There is a possibility of overfitting due to the high level of accuracy. -The methodology is limited to the identification of mango diseases (it is not generalizable). -The methodology requires a labelled dataset and preprocessing. -The real-time deployment of the methodology has not yet been tested.& |
The objective of this study is to utilize an ensemble CNN model for the classification of mango leaf and fruit diseases. This approach aims to facilitate early detection, thereby enhancing the efficacy of crop management practices.& | -The expansion of the present research to encompass other plant diseases. -The implementation of mobile-based real-time detection. -The integration of hybrid classifiers (e.g. Support Vector Machines, Random Forests). -The exploration of advanced segmentation techniques (e.g. deep learning).\\\hline |
| Gai et al. [88]& | Blueberry (2 classes).& | Enhanced YOLOv8 algorithm that incorporates a transfer learning approach (TL-YOLOv8).& |
- Initial Learning Rate: 0.01 (freezing phase). - Optimizer: Not specified. Momentum: 0.937. - Confidence Threshold: 0.25. - NMS Threshold: 0.7. - Batch Size: 16. - Epochs: 300.& |
mAP50 of 94.1\%& | -The feature extraction process is enhanced via the use of MPCA (Multiplexed Coordinated Attention). -The training process is accelerated through the implementation of OREPA (Online Convolutional Reparameterization). -The system demonstrates improved occlusion handling through the integration of MultiSEAM (Multi-scale Separation and Occlusion-Aware Module). -The model is designed to be compact, suitable for deployment at the edge. -The system exhibits high generalization capabilities using transfer learning.& |
-An increased computational complexity from MPCA/MultiSEAM. -A potential degradation in performance under varying lighting, weather, or seasonal conditions not included in the training data.& |
Enhance the accuracy of blueberry detection in complex agricultural environments, characterized by factors such as small size, dense distribution, leaf occlusion, and color similarity to leaves. This study employs an optimized YOLOv8 model and the strategy of transfer learning to achieve this objective.& | Prospective developments include the incorporation of additional lighting, weather and seasonal variations, and the exploration of model pruning for enhanced deployment on embedded systems.\\\hline |
| Buchke et Mayuri [89]& | Tomato leaves (10 classes)& | EfficientNet-B3 with Transfer Learning&. |
- Learning Rate: Controlled via callbacks (not explicitly stated) - Optimizer: Not specified. - Batch Size: Not specified. - Epochs: 12.& |
99.5\% with 10000 images& | Firstly, it is highly accurate, whilst requiring minimal hardware. Secondly, it makes efficient use of transfer learning and compound scaling. Thirdly, it is simple to implement, whilst reducing training time. Finally, it performs robustly across varying dataset sizes.& | -The performance of the system is dependent on the size of the dataset. -There is a limited exploration of the hyperparameters (e.g. optimizer, learning rate). -There is a potential for overfitting to occur with smaller datasets. -The resolution (200x200) may result in the loss of fine-grained details.& |
to develop an EfficientNet-based model for the early detection and classification of tomato leaf diseases using transfer learning, with a view to improving precision agriculture practices.& | -The experimental exploration of diverse optimizers and learning rates to ascertain their respective efficacies. -The investigation of datasets of augmented size and images of enhanced resolution to expand the scope of analysis. -The assessment of the model's resilience to real-world conditions, thereby ensuring its practical applicability. -The extension of the model to encompass other crops and diseases, fostering a comprehensive understanding of its generalisability.\\\hline |
| Vo et al. [90]& | Grape (4 classes)& | Transfer learning with ResNet50V2, ResNet152V2, MobileNetV2, Xception, InceptionV3; Hyperband optimization.& |
Learning Rate: 0.0001. Optimizer: Adamax. Batch Size: 32. Epochs: 30.& |
99.94\% & | The system has been demonstrated to achieve state-of-the-art accuracy. Furthermore, it employs transfer learning in an efficient manner, even when the available data is limited. Finally, Hyperband has been shown to reduce tuning time.& | -The search for optimal hyperparameters is computationally intensive. -There is a possibility of dataset bias (Kaggle-sourced).& |
to optimize the identification of grape leaf disease through the utilization of transfer learning and hyperparameter tuning.& | It is recommended that future research endeavours encompass the exploration of additional hyperparameters, such as weight decay, activation functions, and batch sizes, in addition to alternative optimization techniques.\\\hline |
| Han et Guo [91]& | Ligneous plants (Cherry Apple, Citrus, Grape, Peach) (22 classes)& | Hierarchical Vision Transformer (Swin Transformer) with Transfer Learning.& |
- Learning Rate: 1e-4. - Optimizer: Adam. -Batch Size: 8. - Depth (transformer blocks): 12 - Epochs: 100.& |
86.43\%& | -An improvement in accuracy -A better handling of dispersed disease regions than CNNs -A reduction in computational complexity compared to the original Vision Transformer -A reduction in training time thanks to transfer learning.& |
-The computational expense of the method in comparison with traditional CNNs is a notable issue. -The dataset is imbalanced (unbalanced classes). -A substantial amount of training data is required. -The dataset is limited and there are issues with class imbalance.& |
To develop an automated classification system for ligneous leaf diseases. This will be achieved by using a hierarchical Vision Transformer to improve accuracy and efficiency over existing methods.& | It is recommended that future research should include the exploration of multi-modal deep learning models. In addition, the issue of class imbalance should be addressed by collecting a more diverse set of samples. Finally, further optimization of transformer architectures is required to enhance their performance.\\\hline |
| Radočaj et al. [92]& | Tomato (6 classes)& | - Convolutional Neural Networks (CNNs) with transfer learning. - Pre-trained models: InceptionV3, InceptionResNetV2, MobileNetV2, DenseNet201. - Proposed IncMB module (Inception module, Mish activation function, Batch normalization). - Support Vector Machine (SVM) for comparison.& |
- Learning rate: Automatically determined and adjusted during training. - Optimizer: Adam. - Batch size: 32. - Epochs: 15 and 30.& |
InceptionV3 model, when utilizing the IncMB module, attains an accuracy of 97.78\%.& | -The ability to detect plant diseases at an early stage. -Greater accuracy compared to traditional methods. -A reduction in the time and labor required for disease detection. -The potential for real-time disease detection using mobile devices (MobileNetV2 with IncMB).& |
-The model demonstrates elevated computational complexity and hardware requirements. -The model is dependent on background features in images. -The model's performance may be affected by the limited size of the dataset. -The model's performance may be affected by the presence of overlapping symptoms of diseases, which can lead to misclassification.& |
To develop a versatile module (IncMB) for optimizing convolutional neural networks (CNNs) in the detection of tomato leaf diseases; and to compare the performance of CNNs, CNNs with support vector machines (SVMs), and CNNs with the IncMB module for the purpose of early disease detection.& | -The dataset should be expanded to include a greater number of tomato diseases, as well as healthy leaves. -The IncMB module should be tested on other plant disease datasets. -The model should be optimized for faster processing and real-time applications. -Hyperspectral imaging should be integrated for the early detection of diseases before visible symptoms appear.\\\hline |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




