Submitted:
27 August 2025
Posted:
01 September 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Results
Hyperpathway
Hyperpathway Visualization of Genomic Data
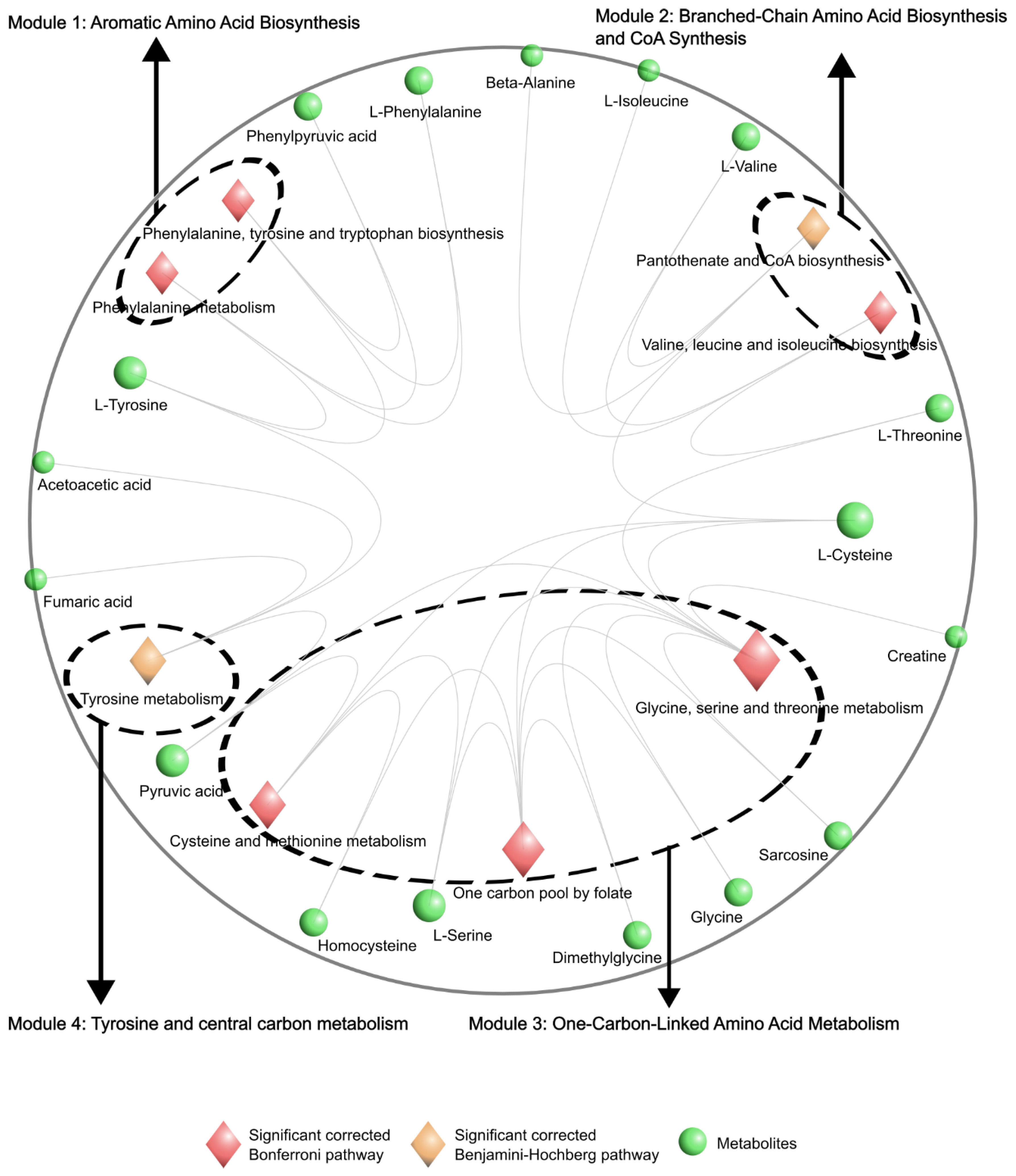
Hyperpathway Visualization of Metabolomics Data
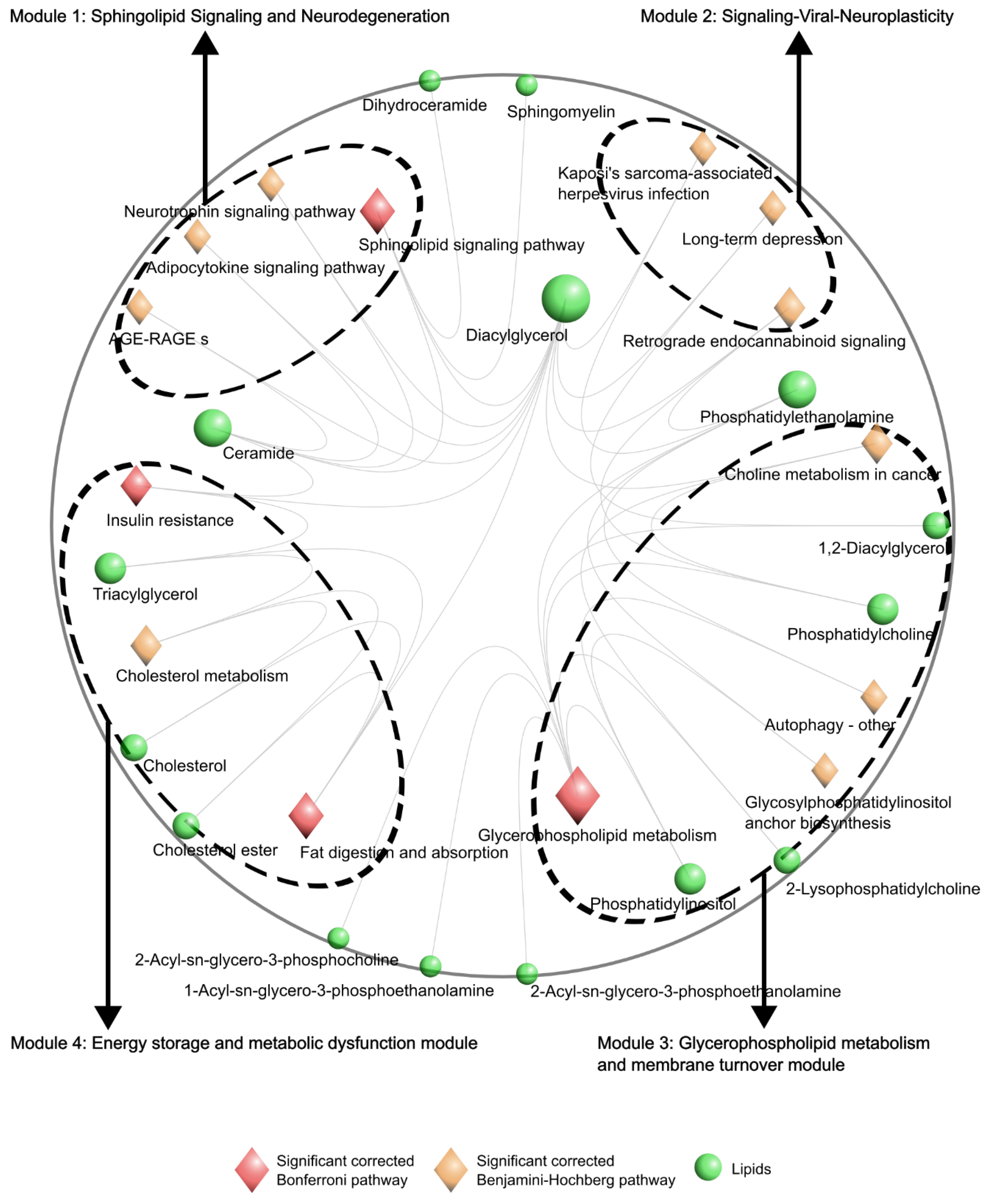
Hyperpathway Visualization of Lipidomic Data
Discussion
Methods
Genomic Data
Metabolomic Data
Lipidomic data
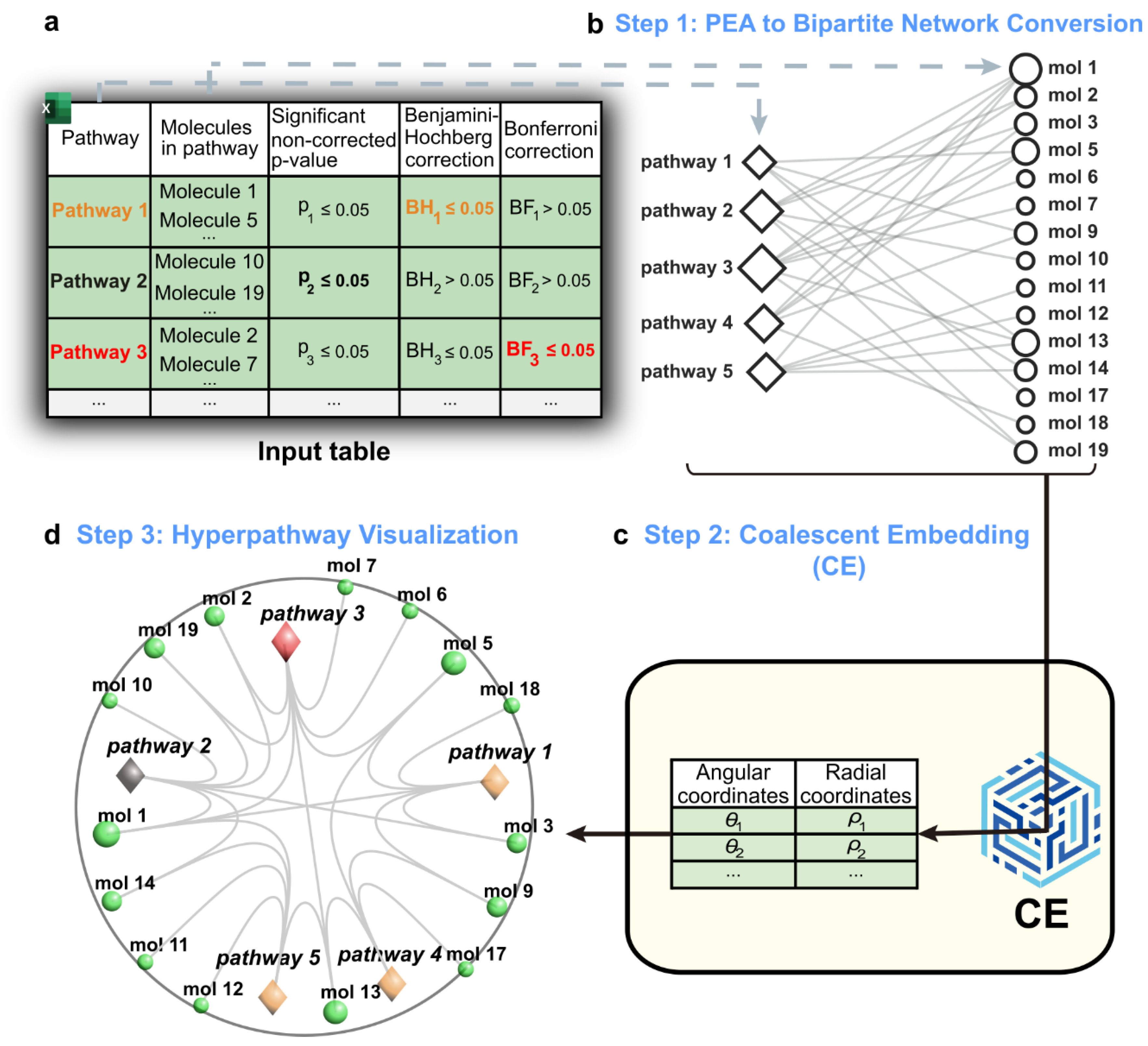
Hyperpathway
Coalescent Embedding of the PEA Bipartite Network
Artificial Linking Strategy Between Separated components
Visualization of the Embedded Network in the 2D Native Hyperbolic Disk
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
References
- Reimand, J.; et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat Protoc, 2019; 14, 482–517. [Google Scholar] [CrossRef]
- Ciucci, S.; et al. Enlightening discriminative network functional modules behind Principal Component Analysis separation in differential-omic science studies. Sci Rep, 2017; 7, 43946. [Google Scholar] [CrossRef]
- Huang, D. W. , Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res, 2009; 37, 1–13. [Google Scholar] [CrossRef]
- Chagoyen, M. & Pazos, F. MBRole: enrichment analysis of metabolomic data. Bioinformatics, 2011; 27, 730–731. [Google Scholar] [CrossRef]
- Huang, D. W. , Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc, 2009; 4, 44–57. [Google Scholar] [CrossRef]
- Subramanian, A.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences, 2005; 102, 15545–15550. [Google Scholar] [CrossRef]
- Xie, C. , Jauhari, S. & Mora, A. Popularity and performance of bioinformatics software: the case of gene set analysis. BMC Bioinformatics, 2021; 22, 191. [Google Scholar] [CrossRef]
- García-Campos, M. A. , Espinal-Enríquez, J. & Hernández-Lemus, E. Pathway Analysis: State of the Art. Front Physiol, 2015; 6. [Google Scholar] [CrossRef]
- Pavlidis, P. , Qin, J., Arango, V., Mann, J. J. & Sibille, E. Using the Gene Ontology for Microarray Data Mining: A Comparison of Methods and Application to Age Effects in Human Prefrontal Cortex. Neurochem Res, 2004; 29, 1213–1222. [Google Scholar] [CrossRef]
- Khatri, P. , Sirota, M. & Butte, A. J. Ten Years of Pathway Analysis: Current Approaches and Outstanding Challenges. PLoS Comput Biol, 2012; 8, e1002375. [Google Scholar] [CrossRef]
- Drier, Y. , Sheffer, M. & Domany, E. Pathway-based personalized analysis of cancer. Proceedings of the National Academy of Sciences, 2013; 110, 6388–6393. [Google Scholar] [CrossRef]
- Kankainen, M. , Gopalacharyulu, P., Holm, L. & Orešič, M. MPEA—metabolite pathway enrichment analysis. Bioinformatics, 2011; 27, 1878–1879. [Google Scholar]
- Stöckel, D.; et al. Multi-omics enrichment analysis using the GeneTrail2 web service. Bioinformatics, 2016; 32, 1502–1508. [Google Scholar] [CrossRef]
- Acevedo, A. , Durán, C., Ciucci, S., Gerl, M. & Cannistraci, C. V. LIPEA: Lipid Pathway Enrichment Analysis. Preprint, 2018. [Google Scholar] [CrossRef]
- Muscoloni, A. , Thomas, J. M., Ciucci, S., Bianconi, G. & Cannistraci, C. V. Machine learning meets complex networks via coalescent embedding in the hyperbolic space. Nat Commun, 2017; 8, 1615. [Google Scholar] [CrossRef]
- Boguñá, M. , Papadopoulos, F. & Krioukov, D. Sustaining the Internet with hyperbolic mapping. Nat Commun, 2010; 1, 62. [Google Scholar] [CrossRef]
- Cacciola, A.; et al. Coalescent embedding in the hyperbolic space unsupervisedly discloses the hidden geometry of the brain. arXiv, 2017; arXiv:1705.04192. [Google Scholar]
- Cannistraci, C. V. & Muscoloni, A. Geometrical congruence, greedy navigability and myopic transfer in complex networks and brain connectomes. Nat Commun, 2022; 13, 7308. [Google Scholar] [CrossRef]
- Alanis-Lobato, G. , Mier, P. & Andrade-Navarro, M. The latent geometry of the human protein interaction network. Bioinformatics, 2018; 34, 2826–2834. [Google Scholar] [CrossRef]
- Zhang, Y. , Zhao, J., Wu, W. & Muscoloni, A. Epitopological learning and cannistraci-hebb network shape intelligence brain-inspired theory for ultra-sparse advantage in deep learning. in The Twelfth International Conference on Learning Representations (2024).
- Daminelli, S. , Thomas, J. M., Durán, C. & Vittorio Cannistraci, C. Common neighbours and the local-community-paradigm for topological link prediction in bipartite networks. New J Phys, 2015. [Google Scholar] [CrossRef]
- Kovács, I. A.; et al. Network-based prediction of protein interactions. Nat Commun, 2019; 10, 1–8. [Google Scholar]
- Muscoloni, A. , Abdelhamid, I. & Cannistraci, C. V. Local-community network automata modelling based on length-three-paths for prediction of complex network structures in protein interactomes, food webs and more. [CrossRef]
- Muscoloni, A. & Cannistraci, C. V. A nonuniform popularity-similarity optimization (nPSO) model to efficiently generate realistic complex networks with communities. New J Phys, 2018; 20, 052002. [Google Scholar] [CrossRef]
- Pang, Z.; et al. MetaboAnalyst 6.0: towards a unified platform for metabolomics data processing, analysis and interpretation. Nucleic Acids Res, 2024; 52, W398–W406. [Google Scholar] [CrossRef]
- Lionaki, E. , Ploumi, C. & Tavernarakis, N. One-Carbon Metabolism: Pulling the Strings behind Aging and Neurodegeneration. Cells, 2022; 11, 214. [Google Scholar] [CrossRef]
- Slieker, R. C.; et al. Identification of biomarkers for glycaemic deterioration in type 2 diabetes. Nat Commun, 2023; 14, 2533. [Google Scholar] [CrossRef]
- Lv, C.; et al. β- cell dynamics in type 2 diabetes and in dietary and exercise interventions. J Mol Cell Biol, 2022; 14. [Google Scholar] [CrossRef]
- Folli, F.; et al. Altered Insulin Receptor Signalling and β-Cell Cycle Dynamics in Type 2 Diabetes Mellitus. PLoS One, 2011; 6, e28050–e28050. [Google Scholar] [CrossRef]
- Yang, S. J. , Kwak, S.-Y., Jo, G., Song, T.-J. & Shin, M.-J. Serum metabolite profile associated with incident type 2 diabetes in Koreans: findings from the Korean Genome and Epidemiology Study. Sci Rep, 2018; 8, 8207. [Google Scholar] [CrossRef]
- Papadopoulos, F. , Kitsak, M., Serrano, M. Á., Boguñá, M. & Krioukov, D. Popularity versus similarity in growing networks. Nature, 2012; 489, 537–540. [Google Scholar] [CrossRef]
- Papadopoulos, F. , Kitsak, M., Serrano, M. Á., Boguñá, M. & Krioukov, D. Popularity versus similarity in growing networks. Nature, 2012; 489, 537–540. [Google Scholar] [CrossRef]
- Tenenbaum, J. B. , Silva, V. de & Langford, J. C. A Global Geometric Framework for Nonlinear Dimensionality Reduction. Science (1979), 2000; 290, 2319–2323. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



