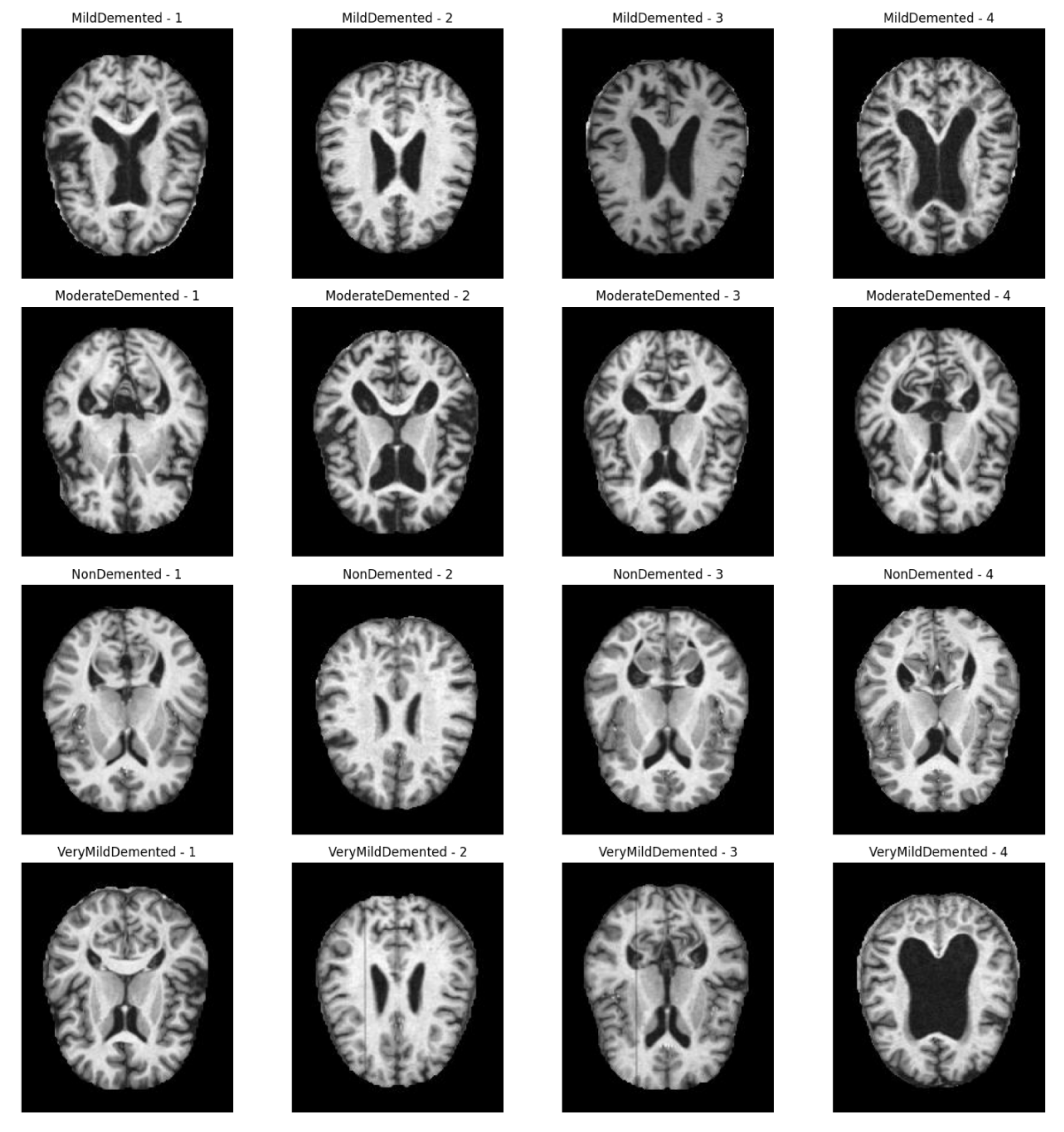
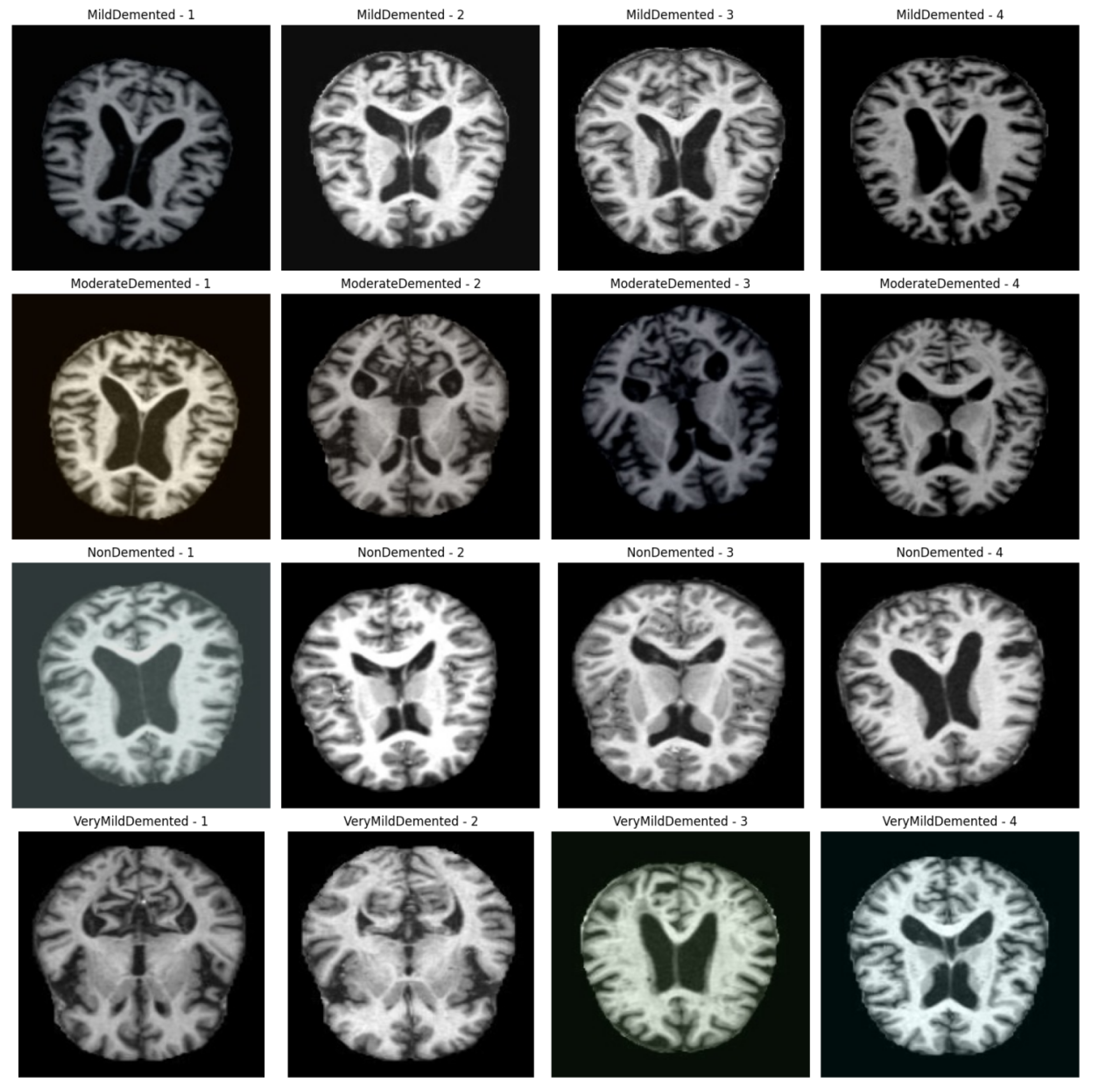
3.1. Methodological Processes
The WTRGANet method combines the wavelet transform and residual gated attention mechanism with the aim of improving the classification performance of Alzheimer’s disease MRI images.
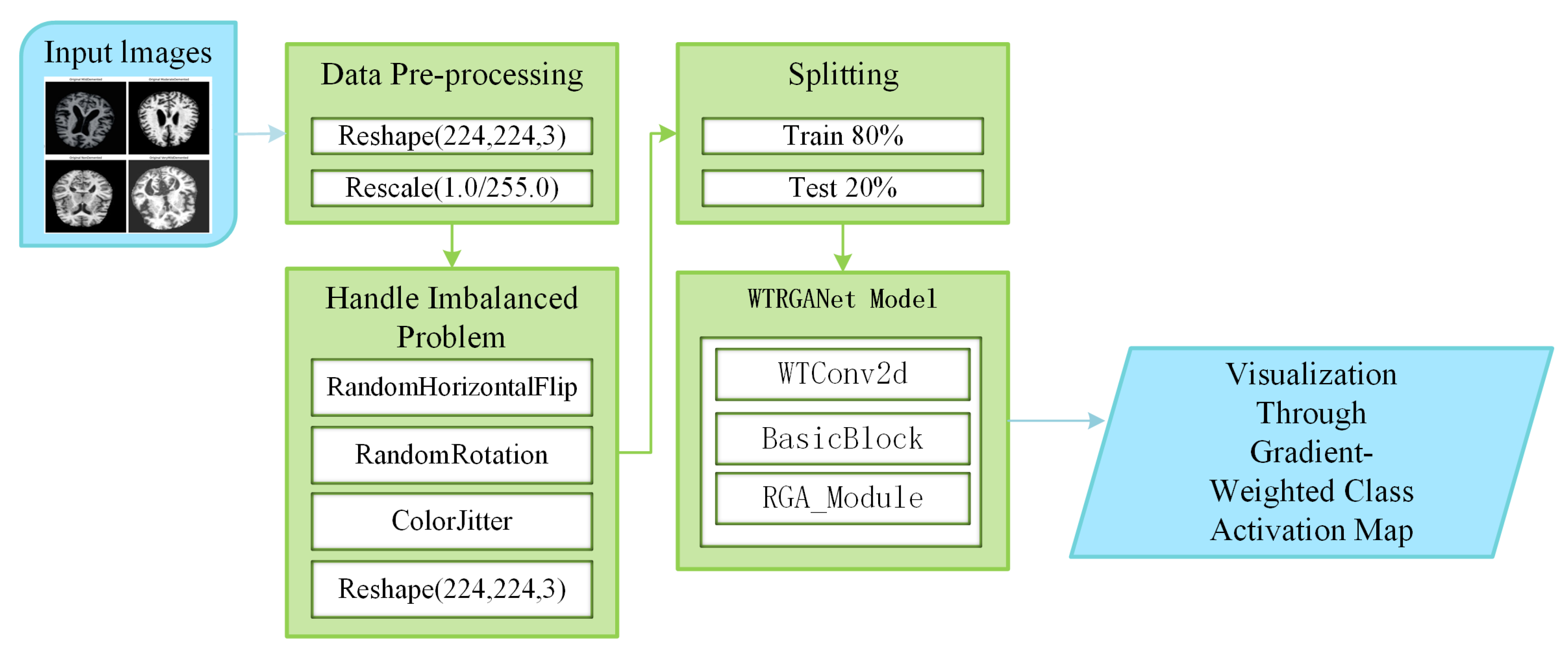
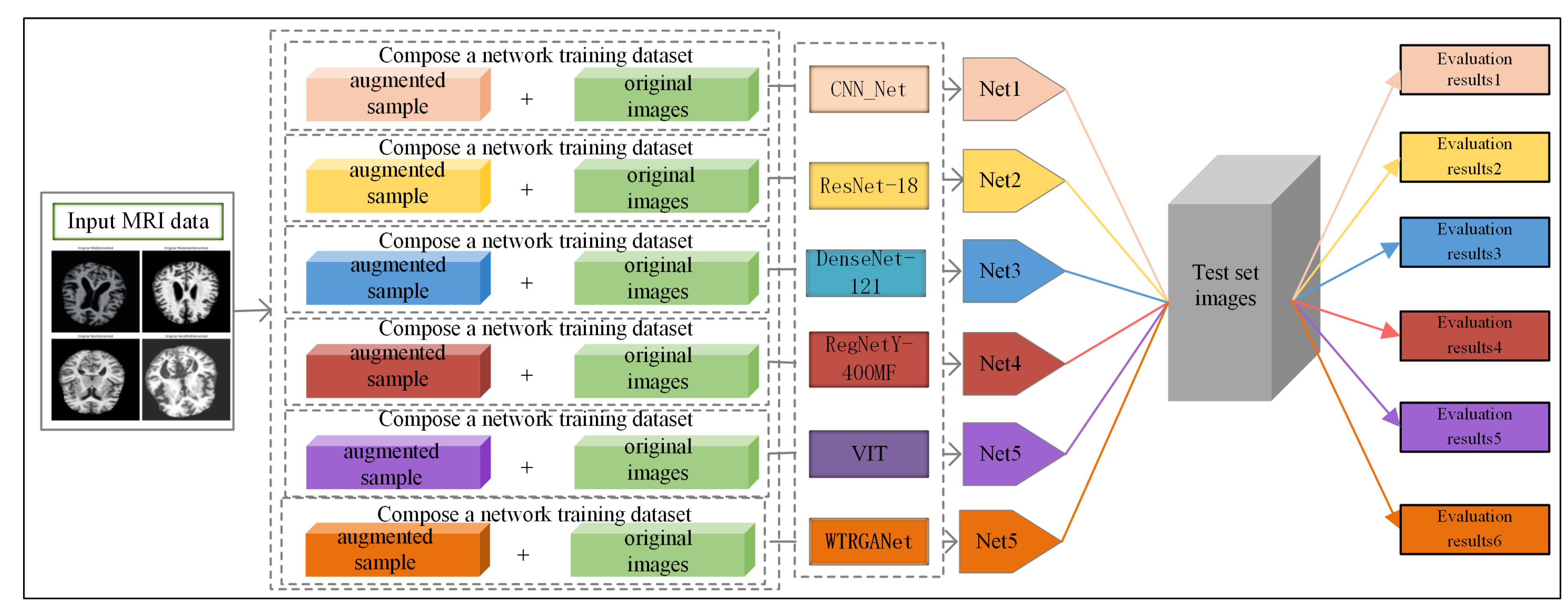
Figure 1 shows the processing flow of the algorithm. Firstly, the input MRI images are resized to 224x224x3 by a preprocessing step and normalised by pixel values in order to fit the input requirements of the neural network. The dataset is divided into a training set (80%) and a test set (20%) and data enhancement techniques (e.g., random level flipping, rotation and colour perturbation) are employed to cope with category imbalance and to improve the generalisation ability of the model.WTRGANet extracts multi-resolution features by integrating the wavelet transform, which enhances the response of the low-frequency information and extends the receptive field to improve the ability to capture the details of the image. The BasicBlock is used for local feature extraction, while the Residual Gated Attention Module combines channel and spatial attention mechanisms to dynamically optimise feature representation and suppress the effects of noise and blur. Eventually, the model is classified by the Fully Connected Layer,which outputs four categories: mildly demented, moderately demented, non-demented, and very mildly demented. The specific algorithm module design is described in detail below.
3.1.1. WTRGANet Network Architecture
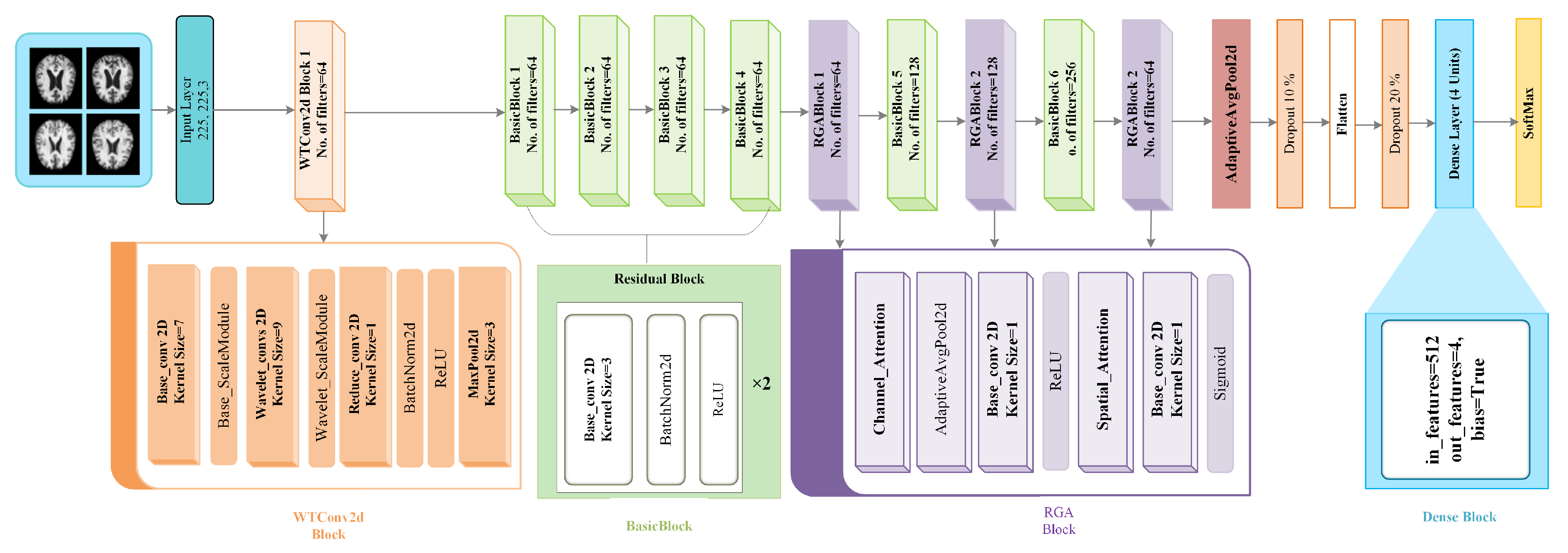
The network architecture of the WTRGANet model is shown in
Figure 2, combining four core modules: the WTConv2d, BasicBlock, RGABlock, and DenseBlock, each of which plays an important role at different stages of the network and works in concert to achieve improved recognition of MRI images of Alzheimer’s disease.First, the WTConv2d module serves as an input layer containing a convolutional layer and a wavelet transform convolutional layer.The standard convolutional layer is used to extract the initial features of the image, while the wavelet transform convolutional layer enhances the responsiveness of the low frequency information by performing a multi-resolution decomposition of the image .This module also includes a ScaleModule, which scales the image features to ensure that the different frequency components are efficiently passed on to subsequent layers.The ReduceConv layer further adjusts the dimensionality of the feature map to help the network extract more valuable features.
Next, the four BasicBlock modules, which are used for in-depth processing of the image features, are used to operate with two layers of convolution, each of which is immediately followed by batch normalisation and the ReLU activation function.Through the residual connection, BasicBlock can effectively alleviate the problem of gradient vanishing, and make the information can be better passed to the subsequent layers to improve the expressive ability and training stability of the network.Subsequently, the feature representation is further optimised by the RGA Block (Residual Gated Attention Block) module with the BasicBlock module,Which incorporates the Channel Attention mechanism alongside the Spatial Attention mechanism.
Through adaptive average pooling and convolutional operations, the RGA module assigns weights to each channel and spatial location to strengthen key features and suppress irrelevant features.Next, features are aggregated through a Global Average Pooling layer to further reduce the number of parameters and enhance the representation of global information. The network also introduces a Dropout layer as a means of regularisation, Which randomly deactivates certain neuron activation values to mitigate overfitting and enhance the model’s generalization capability. Finally, The Dense Block module is designed to integrate high-level features within the network, facilitating more effective feature propagation and reuse,mapping the extracted high-level features to the final output layer using the Fully Connected Layer, and outputting a multi-category classification via SoftMax to complete the task of recognising mild, moderate, non-dementia and very mild dementia in Alzheimer’s disease.
Overall, the deep convolutional network backbone of WTRGANet consists of wavelet transform, multiple convolutional layers, residual units, channel attention mechanisms, and spatial attention mechanisms. It can be described as follows: First, the input image undergoes wavelet transform through a custom WTConv2d module, decomposing the image into low-frequency and high-frequency components. The low-frequency components are processed by a standard 7×7 convolutional layer for initial feature extraction, followed by batch normalization and ReLU activation to generate the initial feature map. The high-frequency components are processed by independent depthwise convolutional layers, which use grouped convolution (groups=9) to achieve efficient feature extraction and parameter sharing. Next, the network passes through four main BasicBlock stages. Each BasicBlock consists of two 3×3 convolutional layers, batch normalization layers, and ReLU activation functions, with skip connections adding the input feature map to the output feature map,is formulated as:
WTConv2d The feature maps after wavelet transformation are represented as follows, and the detailed transformation process will be described later.
.
two
convolutional layers and activation functions are represented as follows.
X is input feature map,
Y is output feature map.At the end of the stage, we integrate the Residual Gated Attention Module to recalibrate the output feature map through channel and spatial attention. Specifically, the RGA_Module dynamically adjusts the weights in the feature map using the following formula:
is channel attention weights, They are generated through global average pooling followed by two
convolutional layers.
is spatial attention weights, They are generated through a dimensionality-reduction convolution followed by two
convolutional layers, ⊙ denotes the element-wise multiplication operation. The channel attention mechanism strengthens the representation of significant channels, whereas the spatial attention mechanism emphasizes crucial features in key spatial regions, collectively improving the model’s discriminative power. As the network depth increases, the number of channels in the feature maps gradually increases to 64, 128 , and 256 , while the spatial dimensions of the feature maps are progressively reduced through convolutional operations with a stride of 2 .Finally, the feature maps processed through all convolutional layers and attention modules are compressed into fixed-size feature vectors via a global average pooling layer:
Subsequently, the feature vectors are mapped to a probability distribution over the target classes through a linear fully connected layer, achieving the final classification output:
and are the weight and bias parameters of the fully connected layer, respectively. The Softmax function transforms the output of the linear layer into a probability distribution over the classes. By integrating wavelet transform and residual gated attention mechanisms, the entire backbone network can dynamically adjust and optimize feature representations at different levels.
3.2. Wavelet Transform Convolutional Layer
In WTRGANet, the WTConv2d module is a core component responsible for multi-resolution feature extraction through wavelet transform. The module first performs wavelet decomposition on the input image, then conducts convolution operations in the wavelet domain, and finally reconstructs the feature maps via inverse wavelet transform. Specifically, the WTConv2d module consists of the following key parts: creation and initialization of wavelet filters, wavelet transform and inverse wavelet transform, basic convolution operations, convolution and scaling of high-frequency components, and feature reconstruction and output.
3.2.1. Creation and Initialisation of Wavelet Filters
The WTConv2d module utilizes Daubechies wavelets (e.g., db1) from the PyWavelets library to create wavelet filters for both forward and inverse transforms.Specifically, the create_wavelet_filter function generates low-pass
and high-pass filters
, which are defined as follows:
These filters form an orthonormal basis, enabling the wavelet transform and inverse wavelet transform to be implemented through standard convolution and transposed convolution. The low-pass filters are used to capture the main structural information of the image, while the high-pass filters are used to capture details and edge information.
3.2.2. Wavelet Transform (math.)
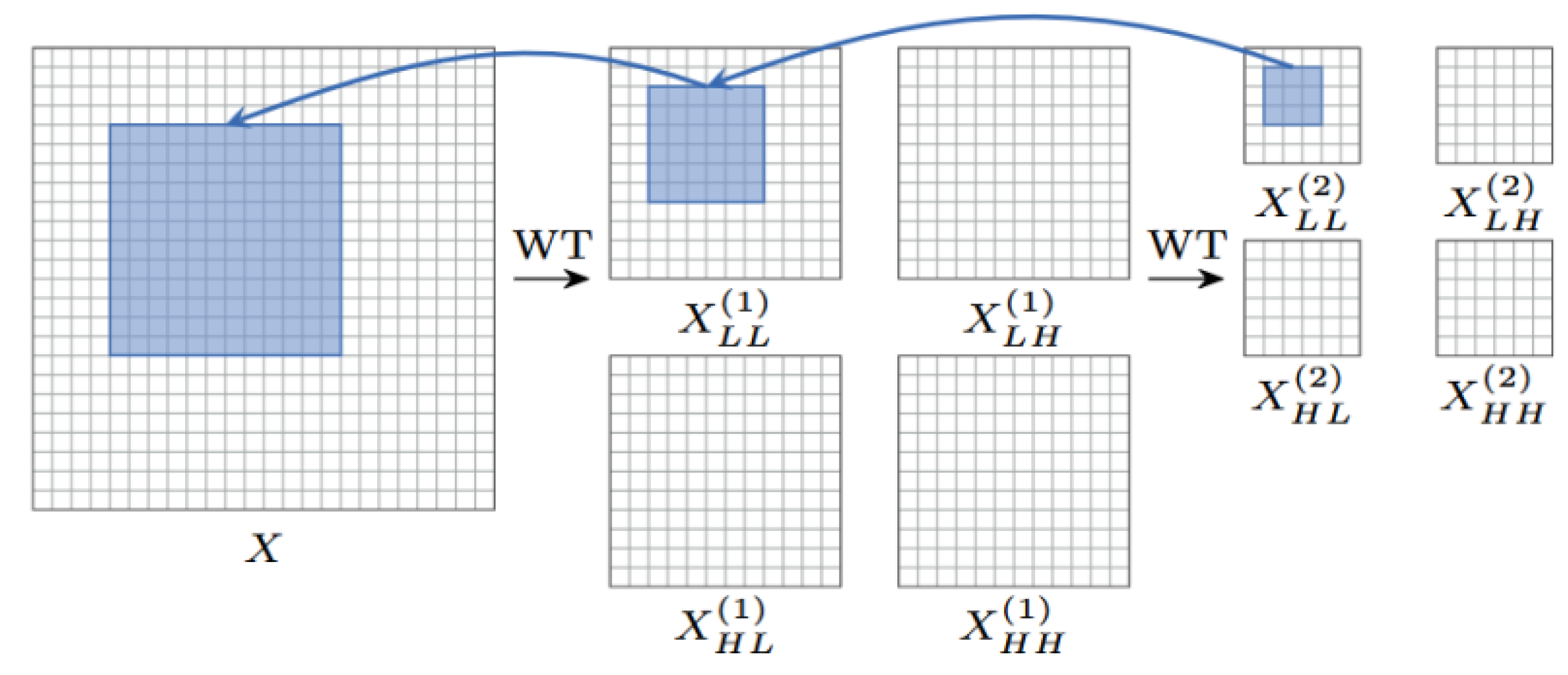
As shown in
Figure 3, during the forward propagation process, the input image
X is first decomposed into a low-frequency component
and high-frequency components
,
by the wavelet transform function
, whose mathematical expression is as follows:
Conv denotes the depthwise convolution operation with a stride of 2 , achieving downsampling. The low-frequency component retains the main structural information of the image, while the high-frequency component contains details and edge information.
3.2.3. Basic Convolutional Operations
The low-frequency component a undergoes further feature extraction through a standard
convolutional layer base_conv. This convolutional layer functions similarly to traditional convolution operations, but since the input features have already undergone multi-resolution processing via wavelet transform, it can more effectively capture the image’s details and structural information. The mathematical expression for the basic convolution operation is as follows:
Here, is the basic convolution kernel, and ScaleModule is a trainable scaling module used to dynamically adjust the scale of the feature maps.
3.2.4. Convolution and Scaling of High Frequency Components
The high-frequency component
is processed by a set of independent
convolutional layersConv
, which employ depth-wise convolution, meaning each input channel is convolved independently. The specific expression is as follows:
is the convolution kernel for the i-level wavelet decomposition, and ScaleModule i is the corresponding scaling module. The processed high-frequency features are dynamically adjusted through the scaling module, further enhancing their representational capability.
3.2.5. Feature Reconstruction and Export
The processed low-frequency and high-frequency features are recombined through the inverse wavelet transform function IWT to reconstruct the enhanced feature map Z. The specific steps are as follows:
- (1)
The low-frequency component
and the high-frequency component
are added together to obtain the fused feature map:
- (2)
The feature maps are reconstructed through the inverse wavelet transform function IWT :
Finally, the output feature map
Z combines the main structural features from the low-frequency information and the detailed features from the high-frequency information, forming the final output:
ReduceConv is a convolutional layer used to reduce the number of channels in the high-frequency components to match the output channels, ensuring consistent dimensionality of the feature maps.
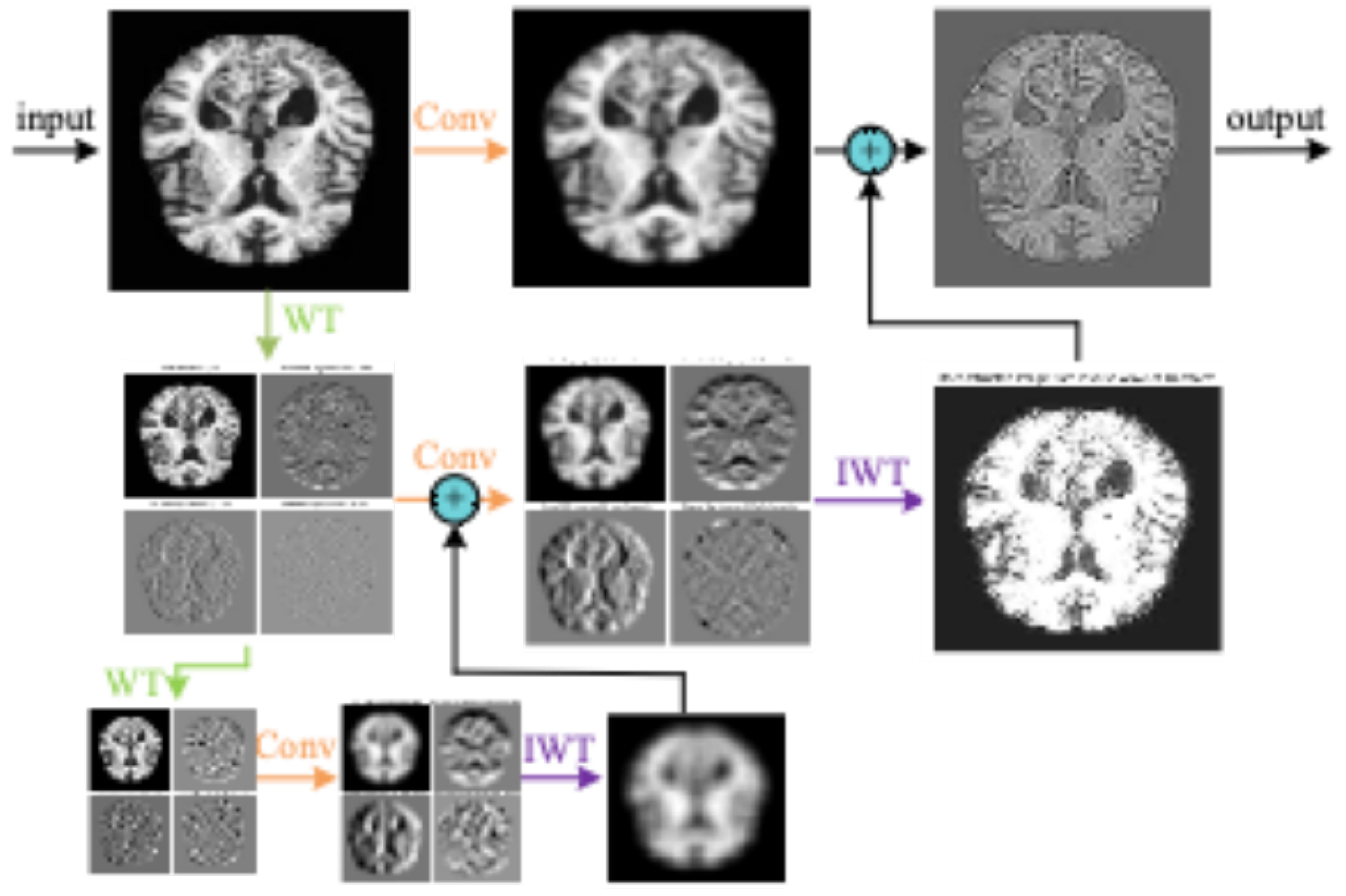
As shown in
Figure 4, the diagram illustrates an image processing pipeline that combines convolution, wavelet transform, and inverse wavelet transform. First, the original MRI image undergoes convolution operations to extract local features such as edges and textures. Next, wavelet transform is applied to decompose the image into components of different frequencies, capturing low-frequency smooth information and high-frequency detail information separately. Finally, the inverse wavelet transform reconstructs the image from the decomposed components, merging the frequency components to produce an image with enhanced details and structures. Through this multi-level, multi-scale learning process, the model can more effectively extract useful information from the image, improving its ability to understand and process image content.
3.2.6. Theoretical Advantages of WTConv2d
The WTConv2d module demonstrates its theoretical strengths in the following ways:
- (1)
Multi-resolution feature extraction:With the wavelet transform, WTConv2d is able to capture image features at different frequency levels, both extracting local details and preserving global structural information.
- (2)
Feel the Wild Expansion:Applying independent small convolutional kernel operations on the high frequency components, WTConv2d effectively extends the receptive field and improves the model’s ability to capture the global information of the image
- (3)
Parametric efficiency:Using deep convolution, WTConv2d avoids excessive growth in the number of parameters while maintaining efficient feature extraction capability.In concrete terms,Using multi-level wavelet decomposition level WT) The number of parameters increases only linearly as , and the sensory field grows exponentially as . k.
WTConv2d effectively controls the number of parameters and the computational cost by combining the wavelet transform and deep convolution while maintaining the feeling field expansion. The computational costs (FLOPs) of deep convolution can be expressed as:
C is number of input channels,
is Spatial dimension of input,
is Convolutional kernel size,
is stride.For example, for a single-channel input with a spatial size of
, the FLOPs for a
convolution kernel are 12.8 M , while the FLOPs for a
convolution kernel are 252 M .Considering a set of convolutional operations in WTConv2d, each wavelet-domain convolution reduces the spatial dimension by a factor of 2 but increases the number of channels by a factor of 4 . Therefore, its FLOPs count is:
ℓ is wavelet decomposition level,For a input and a 3 -level WTConv, the FLOPs using a convolution kernel are 15.1 M , which still demonstrates a significant advantage compared to the computational cost of standard depthwise convolution ( 17.9 M FLOPs). The WTConv2d module achieves the goals of multi-resolution feature extraction and receptive field expansion by combining wavelet transform with depthwise convolution, while effectively controlling the number of parameters and computational cost.
3.3. Residual Gated Attention Module
Residual Gated Attention Module The role of the residual gated attention module is to allow the model to focus on key information, such as important features. The residual gated attention module is not the same as a static convolution or attention mechanism; the residual gated attention module allows the network to focus on wherever it wants to focus, or it can ignore some unimportant features and focus on only the important ones. The model in this study combines the two mechanisms, the channel attention mechanism and the spatial attention mechanism, and then applies a residual link. In this way, the key information in the feature map can be passed on more efficiently, and the gradient back propagation will not get stuck. Channel attention is responsible for judging ‘those feature channels are more important’, while spatial attention is focusing on ‘that part of the graph is more worthy of attention’. Although these two mechanisms work independently of each other, they have many limitations when used alone. In practice, this design is particularly efficient, avoiding the problem of sometimes messing up the weights of the attention mechanism, but also relying on the residual structure to keep the deep network from getting stuck during training. Channel attention mechanism in doing image classification, some channels may specialize in texture, some focus on the colour. Channel attention will automatically determine what the basis of classification is, and then pull the weight of the corresponding channel full, and other weights will be reduced. This dynamic adjustment mechanism is more flexible than the fixed weight approach.
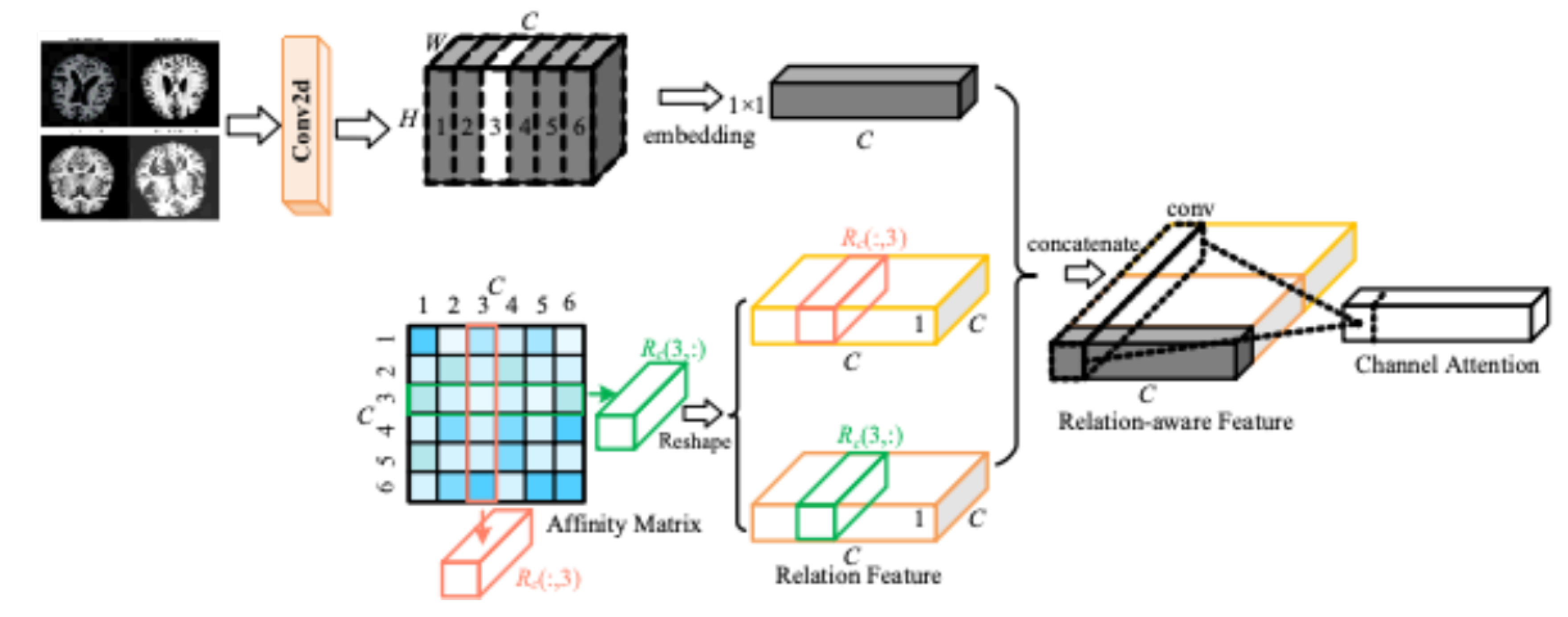
As shown in the
Figure 5, the input MRI image, the first step is the extraction of features by the operation of convolution. The convolution kernel is used in the input feature map to extract the local features in the MRI image. With this operation basic features like edges, texture etc. are extracted from the MRI image. The green, orange and red boxes in the figure indicate the attention computation for different channels. Each layer of the feature map sleeps through the attention mechanism to compute the importance of this channel. The second half of the figure represents the weighting operation for different channels. After this process, the weights of each channel are different and are adjusted so that the weights of the important channels are increased and the weights of the other unimportant ones are decreased. In this way, the attention mechanism will reduce or suppress the information of irrelevant channels by strengthening the important channels.
The spatial attention mechanism teaches the network to learn to ‘see the point’. In the same way that our human eyes unconsciously focus on the key parts of something, it allows the neural network to automatically find the real focus of attention in a picture. The mechanism works as follows: it scans every corner of the picture and then scores the different areas to determine where it is more important, and then ‘looks more’ at the important places and ‘glances’ at the ones that are not red. The advantage of doing so is: computer resources can be used on the knife edge, without wasting computing power in irrelevant areas. After the key features are enlarged, the recognition accuracy will naturally go up, but also can automatically adapt to different sizes and positions of the object. In MRI images, some regions can provide more useful information than others, the spatial attention mechanism by giving these fish a higher weight, yes the network can pay more attention to these regions, and then improve the performance.
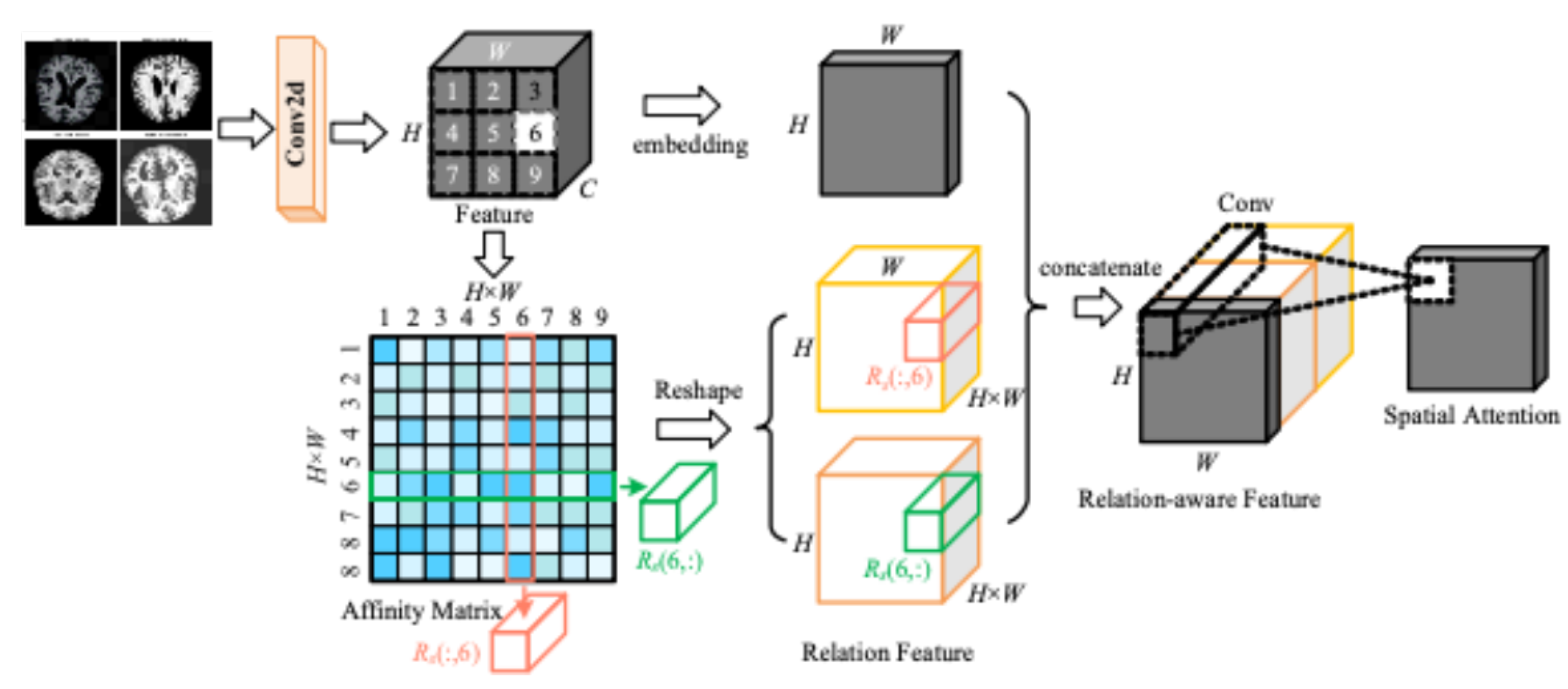
As shown in
Figure 6, the input MRI image is extracted by convolution operation to get the feature map. A convolution kernel of size 3 X 3 is used on the extracted feature map to extract features that are localised in the image. The spatial attention mechanism calculates different weights for the spatial regions and the model gets an enhanced feature map which contains more information from the important regions and helps the network to make more accurate predictions in subsequent tasks.
Specifically, the RGA_Module first receives the feature map
Y processed by the BasicBlock as input. To generate channel attention weights, the module applies Global Average Pooling to
Y, compressing the feature map
into a channel descriptor vector
, which is calculated as follows:
Subsequently, the channel descriptor vector
is processed through two fully connected layers. The first fully connected layer reduces the dimensionality from
C to
and introduces non-linearity via the ReLU activation function. The second fully connected layer restores the dimensionality to
C and generates the channel attention weights
through the Sigmoid activation function. The specific formulas are as follows:
Here,
and
are the weight matrices of the fully connected layers,
r is the reduction ratio (typically set to 16 ), and
denotes the Sigmoid function. By applying the channel attention weights
to the input feature map
Y, the channel-weighted feature map
is obtained:
Next, the RGA_Module computes spatial attention for the channel-weighted feature map
. First, the feature map undergoes dimensionality reduction through a
convolution to reduce computational complexity, resulting in the reduced-dimension feature map
:
Then,
is passed through a
convolutional layer, followed by the Sigmoid activation function, to generate the spatial attention weights
:
By applying the spatial attention weights
to the channel-weighted feature map
, the spatially weighted feature map O is obtained:
To further enhance feature representation, the RGA_Module introduces a residual connection, adding the original input feature map Y to the attention-weighted feature map O , resulting in the final output feature map
:
Through the above steps, the RGA_Module achieves dual attention to the feature maps in both channel and spatial dimensions. The channel attention mechanism enhances the representation of key features by emphasizing important channels, while the spatial attention mechanism improves the model’s ability to identify critical regions by highlighting key spatial areas. The introduction of residual connections not only preserves the original feature information but also facilitates effective gradient propagation, mitigating the vanishing gradient problem in deep networks.
In the overall architecture of WTRGANet, the RGA_Module is integrated at the end of the main BasicBlock stages. The specific process is as follows: First, after extracting preliminary features Y through the BasicBlock, the feature map Y is fed into the RGA_Module for attention weighting, generating the weighted feature map . Subsequently, is passed to the next stage for further processing. This modular design enables WTRGANet to fully leverage attention mechanisms at different levels, dynamically optimizing feature representation.
3.4. Loss Function
To effectively train the WTRGANet model, this study adopts the Cross-Entropy Loss as the primary loss function. Its formula is as follows:
Here, N represents the number of samples, C denotes the number of classes, is the ground truth label of sample i for class c (usually represented as one-hot encoding), and is the predicted probability of sample i for class c.
During the optimization process, this study employs the Adam Optimizer, and its parameter update formula is as follows:
Here, represents the model parameters, is the learning rate, and are the estimates of the first and second moments of the gradients, respectively, and is a small constant to prevent division by zero. The Adam optimizer accelerates the convergence speed and improves training stability by adaptively adjusting the learning rate for each parameter.