Submitted:
30 July 2025
Posted:
06 August 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Methods

Results
| Neuroradiological Presentation | Disease Phenotype | |||
|---|---|---|---|---|
| MELAS (N=17) | LHON (N=10) | Leigh syndrome (N=7) | CPEO (N=14) | |
| Brain magnetic resonance imaging | ||||
| T2-Hyperintensity | 15 (88.2%) | 2 (20%) (bilateral optic nerves posterior segment) | 6 (85.71%) | NA |
| Bilateral-frontotemporal | 5 (33.3%) | NA | NA | NA |
| Frontoparietal areas | 5 (33.3%) | NA | NA | NA |
| Bilateral caudate | 4 (26.6%) | NA | NA | NA |
| Bilateral Putamen | 4 (26.6%) | NA | NA | NA |
| Bilateral caudate + bilateral Putamen | NA | NA | 4 | NA |
| Bilateral Thalamus | NA | NA | 3 (50%) | NA |
| Thalamic and periventricular hyperintensity | NA | NA | 2 | NA |
| Cerebellar Hyperintensity | 4 (26.6%) | NA | NA | NA |
| Bilateral fronto-parietal and occipital gliotic changes | 4(26.6%) | NA | NA | NA |
| Diffuse cortical atrophy | 4 (26.6%) | NA | NA | NA |
| Diffusion restriction | 5 (29.41%) | NA | NA | NA |
| Bilateral Thalamus | 2 (40%) | NA | NA | NA |
| Bilateral Caudate and putamen | 3 (60%) | NA | NA | NA |
| Magnetic resonance spectroscopy | 5 (29.41%) | NA | NA | NA |
| Lipid lactate peak (cortex of bilateral temporal, parietal, and occipital areas) | 3 (60%) | NA | NA | NA |
| N-acetylaspartate peak (frontal grey matter) | 2 (40%) | NA | NA | NA |
| N-acetylaspartate peak (over bilateral basal ganglia) | NA | NA | 1 | NA |
| Normal | 2 | 8 | 1 | 14 |
Discussion
Supplementary Materials
Acknowledgments
Authors’ Contributions
Competing Interests
References
- Suomalainen A, Battersby BJ. Mitochondrial diseases: the contribution of organelle stress responses to pathology. Nat Rev Mol Cell Biol. 2018;19(2):77-92. [CrossRef]
- Parikh S, Goldstein A, Koenig MK, et al. Diagnosis and management of mitochondrial disease: a consensus statement from the Mitochondrial Medicine Society. Genet Med. 2015;17(9):689-701. [CrossRef]
- Gorman GS, Chinnery PF, DiMauro S, et al. Mitochondrial diseases. Nat Rev Dis Primer. 2016;2(1). [CrossRef]
- Finsterer J. Overview on visceral manifestations of mitochondrial disorders. Neth J Med. 2006;64(3):61-71.
- Saneto RP, Sedensky MM. Mitochondrial disease in childhood: mtDNA encoded. Neurother J Am Soc Exp Neurother. 2013;10(2):199-211. [CrossRef]
- Gorman GS, Schaefer AM, Ng Y, et al. Prevalence of nuclear and mitochondrial DNA mutations related to adult mitochondrial disease. Ann Neurol. 2015;77(5):753-759. [CrossRef]
- Skladal D. Minimum birth prevalence of mitochondrial respiratory chain disorders in children. Brain. 2003;126(8):1905-1912. [CrossRef]
- Ibayashi K, Fujino Y, Mimaki M, Fujimoto K, Matsuda S, Goto YI. Estimation of the Number of Patients With Mitochondrial Diseases: A Descriptive Study Using a Nationwide Database in Japan. J Epidemiol. 2023;33(2):68-75. [CrossRef]
- Schaefer AM, McFarland R, Blakely EL, et al. Prevalence of mitochondrial DNA disease in adults. Ann Neurol. 2008;63(1):35-39. [CrossRef]
- Hettiarachchi D, Lakmal K, Dissanayake VHW. Mitochondrial diseases in South Asia – A systematic review. Mitochondrion. 2022;62:24-30. [CrossRef]
- Wolf NI, Smeitink JAM. Mitochondrial disorders: A proposal for consensus diagnostic criteria in infants and children. Neurology. 2002;59(9):1402-1405. [CrossRef]
- Rahman S, Blok RB, Dahl H -H. M, et al. Leigh syndrome: Clinical features and biochemical and DNA abnormalities. Ann Neurol. 1996;39(3):343-351. [CrossRef]
- Hirano M, Ricci E, Richard Koenigsberger M, et al. MELAS: An original case and clinical criteria for diagnosis. Neuromuscul Disord. 1992;2(2):125-135. [CrossRef]
- Nakamura M, Mimura O, Wakakura M, Inatani M, Nakazawa T, Shiraga F. [Designation criteria for Leber’s hereditary optic neuropathy]. Nippon Ganka Gakkai Zasshi. 2015;119(5):339-346.
- Wangkumhang P, Chaichoompu K, Ngamphiw C, et al. WASP: a Web-based Allele-Specific PCR assay designing tool for detecting SNPs and mutations. BMC Genomics. 2007;8(1):275. [CrossRef]
- El-Hattab AW, Adesina AM, Jones J, Scaglia F. MELAS syndrome: Clinical manifestations, pathogenesis, and treatment options. Mol Genet Metab. 2015;116(1-2):4-12. [CrossRef]
- Goto Y ichi, Nonaka I, Horai S. A mutation in the tRNALeu(UUR) gene associated with the MELAS subgroup of mitochondrial encephalomyopathies. Nature. 1990;348(6302):651-653. [CrossRef]
- Yu-Wai-Man P, Griffiths PG, Chinnery PF. Mitochondrial optic neuropathies – Disease mechanisms and therapeutic strategies. Prog Retin Eye Res. 2011;30(2):81-114. [CrossRef]
- Mancuso M, Orsucci D, Angelini C, et al. “Mitochondrial neuropathies”: A survey from the large cohort of the Italian Network. Neuromuscul Disord NMD. 2016;26(4-5):272-276. [CrossRef]
- Tuppen HAL, Blakely EL, Turnbull DM, Taylor RW. Mitochondrial DNA mutations and human disease. Biochim Biophys Acta BBA - Bioenerg. 2010;1797(2):113-128. [CrossRef]
- Chinnery PF, Turnbull DM. Epidemiology and treatment of mitochondrial disorders. Am J Med Genet. 2001;106(1):94-101. [CrossRef]
- Finsterer J. Leigh and Leigh-Like Syndrome in Children and Adults. Pediatr Neurol. 2008;39(4):223-235. [CrossRef]
- Kim JH, Lim MK, Jeon TY, et al. Diffusion and Perfusion Characteristics of MELAS (Mitochondrial Myopathy, Encephalopathy, Lactic Acidosis, and Stroke-Like Episode) in Thirteen Patients. Korean J Radiol. 2011;12(1):15. [CrossRef]
- Newman NJ, Biousse V. Hereditary optic neuropathies. Eye Lond Engl. 2004;18(11):1144-1160. [CrossRef]
- Chin HL, Lai PS, Tay SKH. A clinical approach to diagnosis and management of mitochondrial myopathies. Neurother J Am Soc Exp Neurother. 2024;21(1):e00304. [CrossRef]
- Benito-León J, Louis ED, Mato-Abad V, et al. A data mining approach for classification of orthostatic and essential tremor based on MRI-derived brain volume and cortical thickness. Ann Clin Transl Neurol. 2019;6(12):2531-2543. [CrossRef]
- Mato-Abad V, Labiano-Fontcuberta A, Rodríguez-Yáñez S, et al. Classification of radiologically isolated syndrome and clinically isolated syndrome with machine-learning techniques. Eur J Neurol. 2019;26(7):1000-1005. [CrossRef]
- Kuo CW, Chen HA, Hsu RH, et al. Machine learning to predict mitochondrial diseases by phenotypes. Mitochondrion. 2025;84:102061. [CrossRef]
- Shen L, Falk MJ, Gai X. MSeqDR Quick-Mitome (QM): Combining Phenotype-Guided Variant Interpretation and Machine Learning Classifiers to Aid Primary Mitochondrial Disease Genetic Diagnosis. Curr Protoc. 2024;4(1):e955. [CrossRef]
- Seaby EG, Ennis S. Challenges in the diagnosis and discovery of rare genetic disorders using contemporary sequencing technologies. Brief Funct Genomics. 2020;19(4):243-258. [CrossRef]
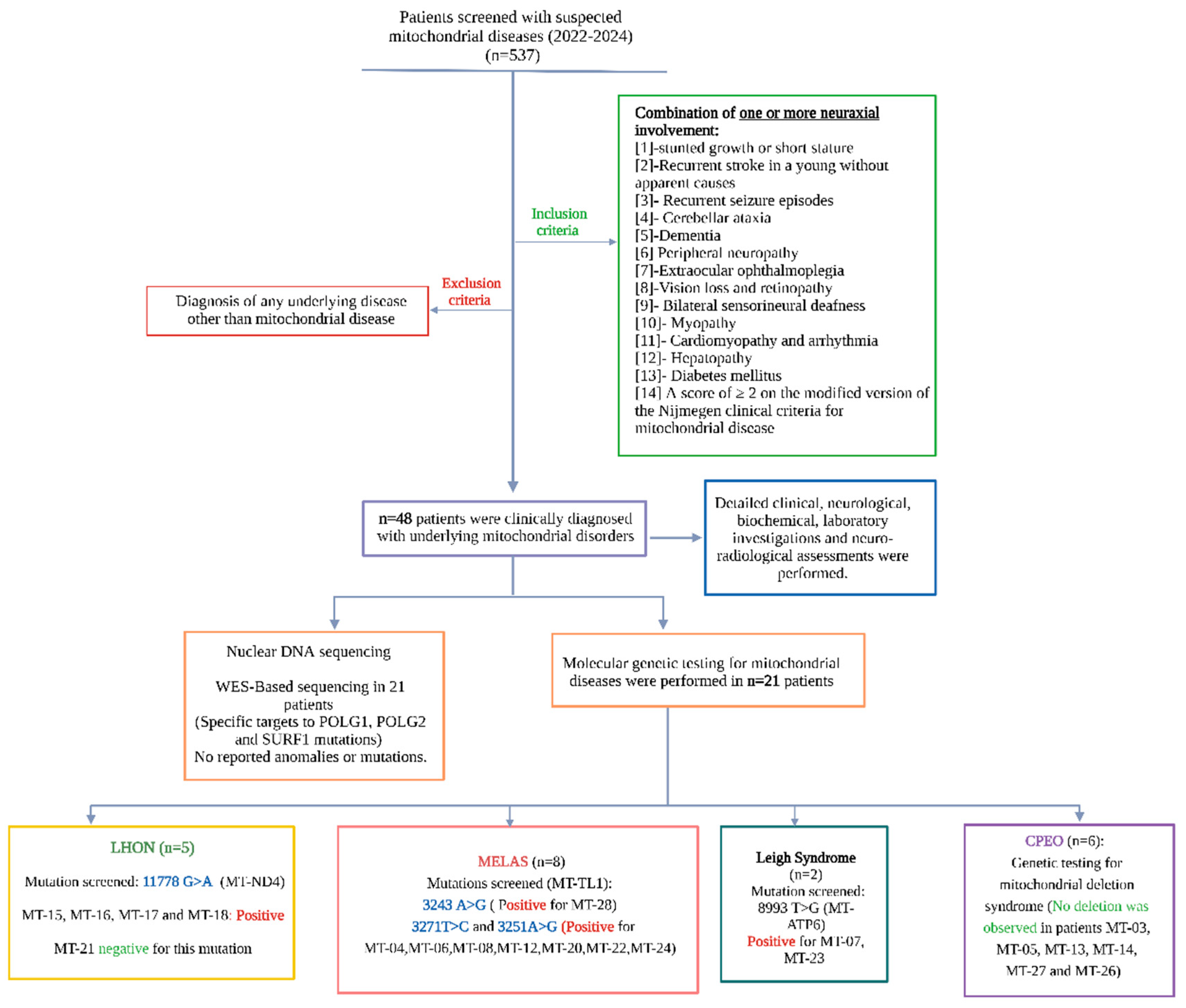
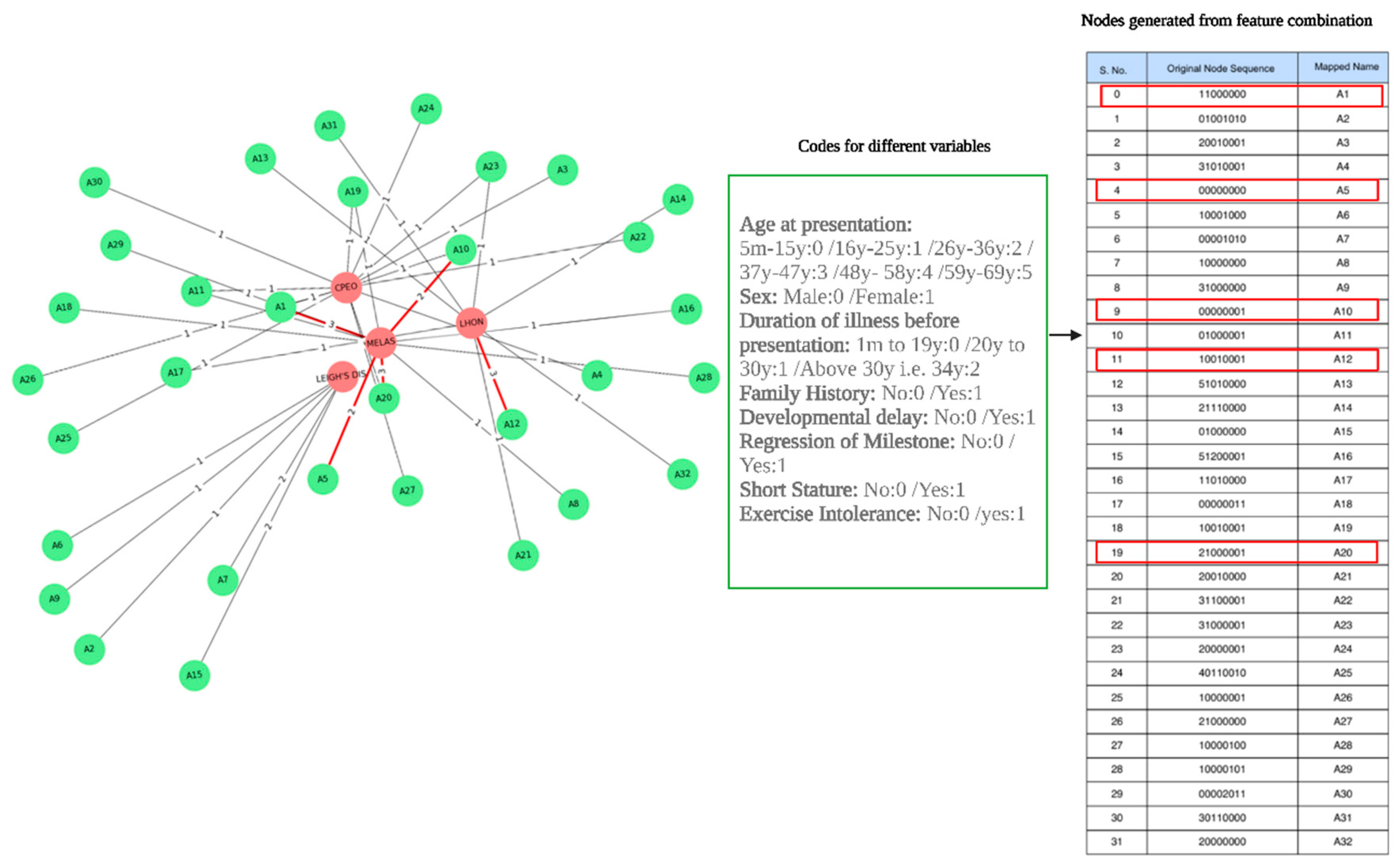
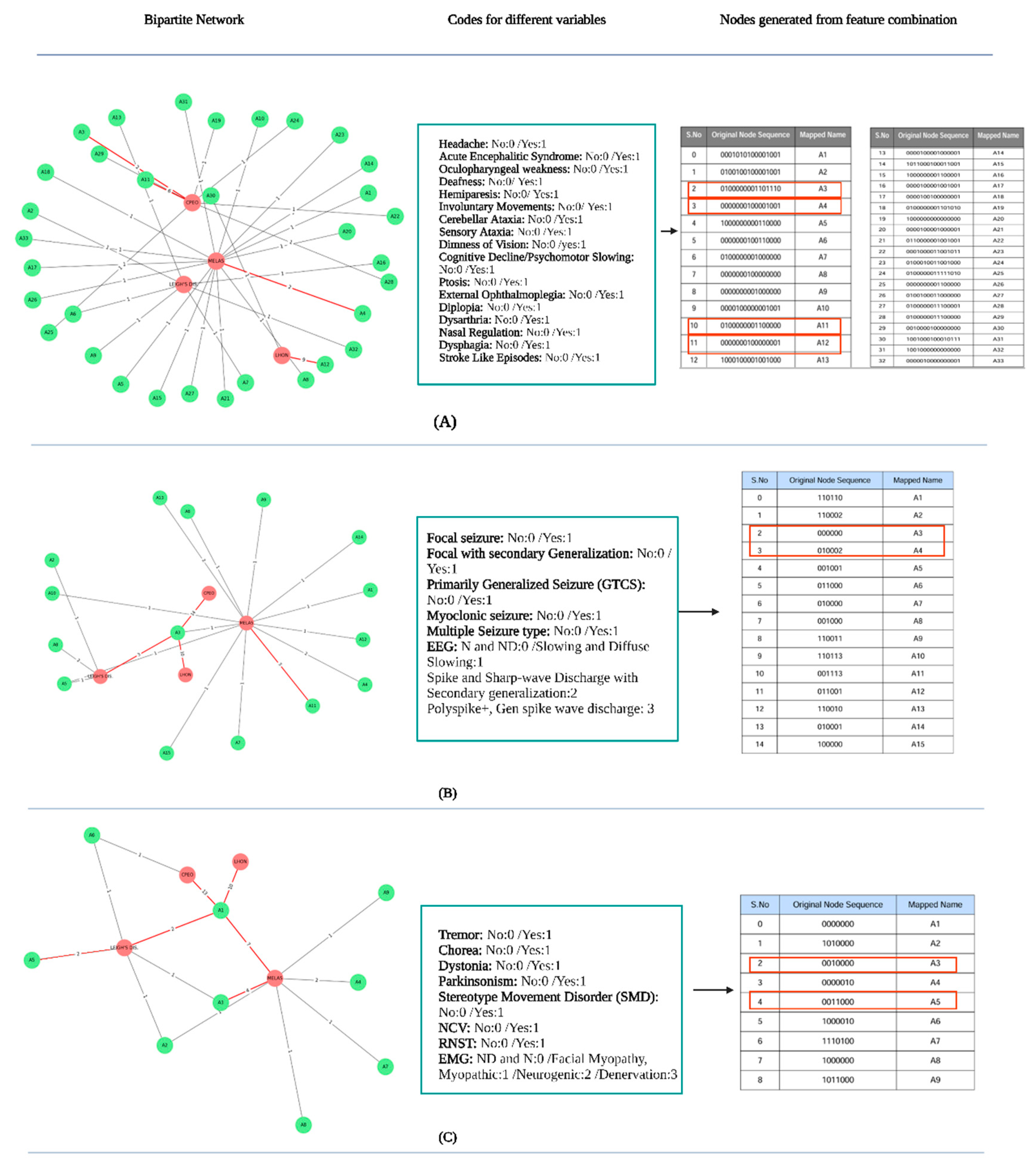
| Patient ID | Diagnosis | Patient ID | Diagnosis |
|---|---|---|---|
| MT-01 | MELAS | MT-25 | MELAS |
| MT-02 | LEIGH | MT-26 | CPEO |
| MT-03 * | CPEO | MT-27 | CPEO |
| MT-04 | MELAS | MT-28 | MELAS |
| MT-05 * | CPEO | MT-29 **** | LHON |
| MT-06 | MELAS | MT-30 **** | LHON |
| MT-07 | LEIGH | MT-31 | MELAS |
| MT-08 | MELAS | MT-32 | CPEO |
| MT-09 | LEIGH | MT-33 | LHON |
| MT-10 | MELAS | MT-34 | MELAS |
| MT-11 | LEIGH | MT-35 | CPEO |
| MT-12 | MELAS | MT-36 *** | CPEO |
| MT-13 | CPEO | MT-37 | CPEO |
| MT-14 | CPEO | MT-38 | CPEO |
| MT-15 ** | LHON | MT-39 | MELAS |
| MT-16 ** | LHON | MT-40 | MELAS |
| MT-17 ** | LHON | MT-41 | CPEO |
| MT-18 ** | LHON | MT-42 | CPEO |
| MT-19 | LEIGH | MT-43 | LHON |
| MT-20 | MELAS | MT-44 | MELAS |
| MT-21 *** | LHON | MT-45 | LEIGH |
| MT-22 | MELAS | MT-46 | CPEO |
| MT-23 | LEIGH | MT-47 | MELAS |
| MT-24 | MELAS | MT-48 | LHON |
| Variables | Disease Phenotype | |||
|---|---|---|---|---|
| MELAS (N=17) | LHON (N=10) | LEIGH Syndrome (N=7) | CPEO (N=14) | |
| Demographic Parameters | ||||
| Age (years) (mean±SD, range) | 20.4±12.3, 5–60 years | 32.3±12.1, 20–60 years | 12.2±11.9, 0.75-35 years | 28.1±12.4, 8–48 years |
| Age (years) at disease onset | 13.5±7.2 | 25.1±12.8 | 9.9±11.3; (SEM±4.6Y) | 20.6±10.7 |
| Duration of Illness (mean) | 6.9 years | 8.2 years | 2.3 years | 7.4 years |
| Positive Family History | 1 (5.9%) | 8 (80%) | 0 | 4 (28.6%) |
| Consanguinity | 3 (17.6%) | 0 | 2 (28.6%) | 1 (7.1%) |
| Sex | ||||
| Male | 9 (52.9%) | 6 (60%) | 3 (42.85%) | 7 (50%) |
| Female | 8 (47.05%) | 4 (40%) | 4 (57.14%) | 4 (50%) |
| Temporal Nature of Disease Progression | ||||
| Slowly Progressive | 8 (47.0%) | 4 (40%) | 3 (42.9%) | 14 (100%) |
| Relapsing /Progressive | 9 (52.9%) | 0 | 1 (14.2%) | 0 |
| Acute onset/Non progressive | 0 | 6 (60%) | 0 | 0 |
| Acute onset/Progressive | 0 | 0 | 3 (42.9%) | 0 |
| Baseline Findings from General Survey | ||||
| Developmental delay | NA | NA | 4 | 1 |
| Regression of Milestone | 2 | NA | NA | NA |
| Short stature | 1 | NA | 3 | 2 |
| Hypertelorism | 1 | NA | 1 | NA |
| Short metatarsal | NA | 1 | NA | NA |
| Exercise intolerance | 8 | 1 | NA | 11 |
| Arterial hypertension | 1 | NA | NA | NA |
| Type-2 diabetes mellitus | 1 | 1 | NA | NA |
| Left bundle branch block | NA | NA | NA | 1 |
| Bifascicular Block | NA | NA | NA | 1 |
| Sexual infantilism | NA | NA | NA | 1 |
| Pregnancy-induced arterial hypertension | 1 | NA | NA | NA |
| Cryptogenic hepatic cirrhosis | 1 | 1 | NA | NA |
| Major depressive disorder | NA | 1 | NA | 1 |
| Variables | Disease Phenotype | |||
|---|---|---|---|---|
| MELAS (N=17) | LHON (N=10) | Leigh syndrome (N=7) | CPEO (N=14) | |
| Neurological presentations | ||||
| Headache | 7 (41.2%) | NA | NA | NA |
| Hemiparesis | 7 (41.2%) | NA | 1 (12.3%) | NA |
| Dysarthria | 10 (58.8%) | NA | 2 (28.6%) | 5 (35.7%) |
| Dysphagia | 3 (17.7%) | NA | NA | 4 (28.6%) |
| Diplopia | 1 (5.9%) | NA | NA | NA |
| Deafness | 1 (5.9%) | NA | 1 (12.3%) | 1 (7.1%) |
| Dimness of Vision | 1 (5.9%) | 10 (100%) | 2 (28.6%) | NA |
| Nasal Regurgitation | NA | NA | NA | 3 (21.4%) |
| External Ophthalmoplegia | 1 (5.9%) | NA | NA | 13 (92.6%) |
| Ptosis and impaired eye movement | 2 (11.8%) | NA | 1 (12.3%) | 13 (92.6%) |
| Oculopharyngeal weakness | NA | NA | 1 (12.3%) | 12 (85.7%) |
| Acute encephalitic syndrome | 4 (23.5%) | NA | 3 (42.9%) | NA |
| Stroke-like episode | 10 (58.8%) | NA | NA | NA |
| Cognitive decline/ Psychomotor slowing | 7 (41.2%) | NA | 4 (57.1%) | 2 (14.3%) |
| Encephalopathy | 10 (58.8%) | NA | 3 (42.9%) | NA |
| Sensorineural hearing loss | 1 (5.9%) | NA | 1 (12.3%) | 2 (14.3%) |
| Pattern of Movement Disorder | ||||
| Tremor | 4 (36.4%) | NA | 2 (40%) | 1 (7.14%) |
| Chorea | 1 (9.1%) | NA | NA | NA |
| Dystonia | 7 (41.2%) | NA | 4 (80%) | NA |
| Parkinsonism | 1 (5.9%) | NA | 1 (20%) | NA |
| Cerebellar Ataxia | 6 (35.3%) | NA | 1 (20%) | 1 (7.14%) |
| Involuntary Movement | 1 (5.9%) | NA | 4 (80%) | NA |
| Axonal Neuropathic Changes | 1 (5.9%) | NA | NA | NA |
| Seizure Types | ||||
| Focal Seizure with Secondary Generalisation | 12 (75%) | NA | ||
| Generalized tonic-clonic seizure | 6 (54.5%) | NA | 3 (42.8%) | NA |
| Myoclonic Seizure | 8 (72.7%) | NA | ||
| Variables | Disease Phenotype | |||
|---|---|---|---|---|
| MELAS (N=17) | LHON (N=10) | Leigh syndrome (N=7) | CPEO (N=14) | |
| Biochemical Parameters | ||||
| Cerebrospinal fluid | ||||
| Increased cell count | Normal | 3 (33.3%) | 3 (42.9%) | 4 (28.57%) |
| Elevated glucose (n, %) Mean±SD (mg/dL) Range (mg/dL) |
2 (11.76%) 56.5±14.1 31-92 |
3 (33.3%) 69.4±17.9 45-108 |
NA 57.4±8.8 44-73 |
NA 55.6±11.3 34-78 |
| Elevated protein (n, %) Mean±SD (mg/dL) Range (mg/dL) |
Normal | 3 (33.3%) 41.9±12.8 21.2-65 |
Normal | 5 (35.7%) 49.1±17.4 23-86 |
| Elevated lactate (n, %) Mean±SD (mg/dL) Range (mg/dL) |
4 (23.5%) 17.9±4.7 10.3-29 |
2 (20%) 17.4±3.8 13-25 |
3 (42.9%) 21.5±6.5 15-34 |
9 (64.3%) 22.6±8.1 12.6-47 |
| Serum | ||||
| Elevated lactate (n, %) Mean±SD (mg/dL) |
13 (76.4%) 24.4±5.8 |
4 (40%) 24±16.6 |
4 (57.1%) 22.1±7 |
8 (57.1%) 27.9±19 |
| Electroencephalographic findings | ||||
| Diffuse Slowing | 4 (23.5%) | NA | 1 (14.3%) | NA |
| Spike and wave discharge | 5 (29.41%) | 3 (42.9%) | ||
| Left-sided spike and sharp wave discharge with secondary generalization | 6 (35.3%) | 1 (14.3%) | ||
| Not Significant | 1 | 10 | 3 | 14 |
| Patient ID | Type of Disease | Gene | Nucleotide Position | Nucleotide change | Condition | Conservation Score |
|---|---|---|---|---|---|---|
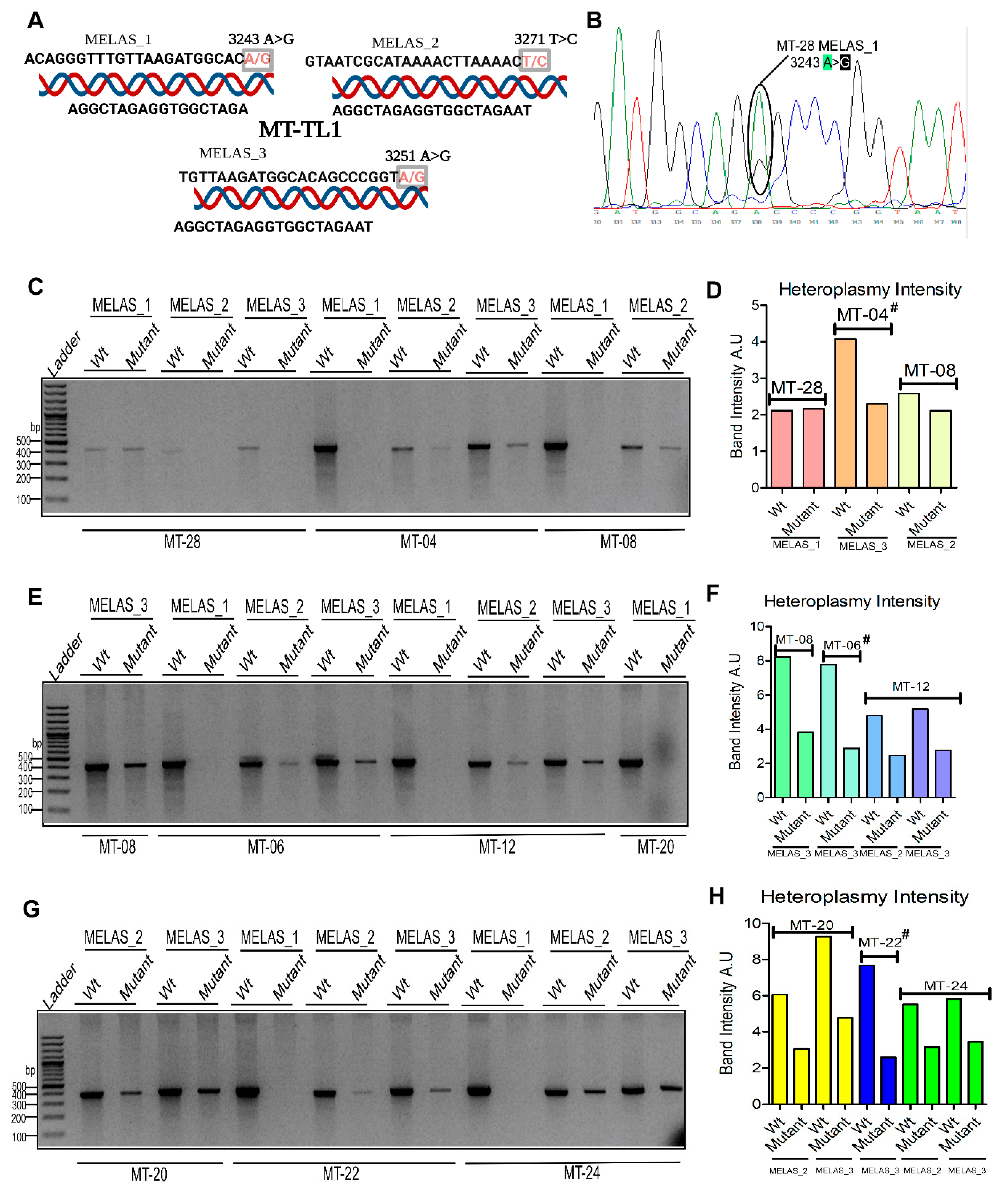
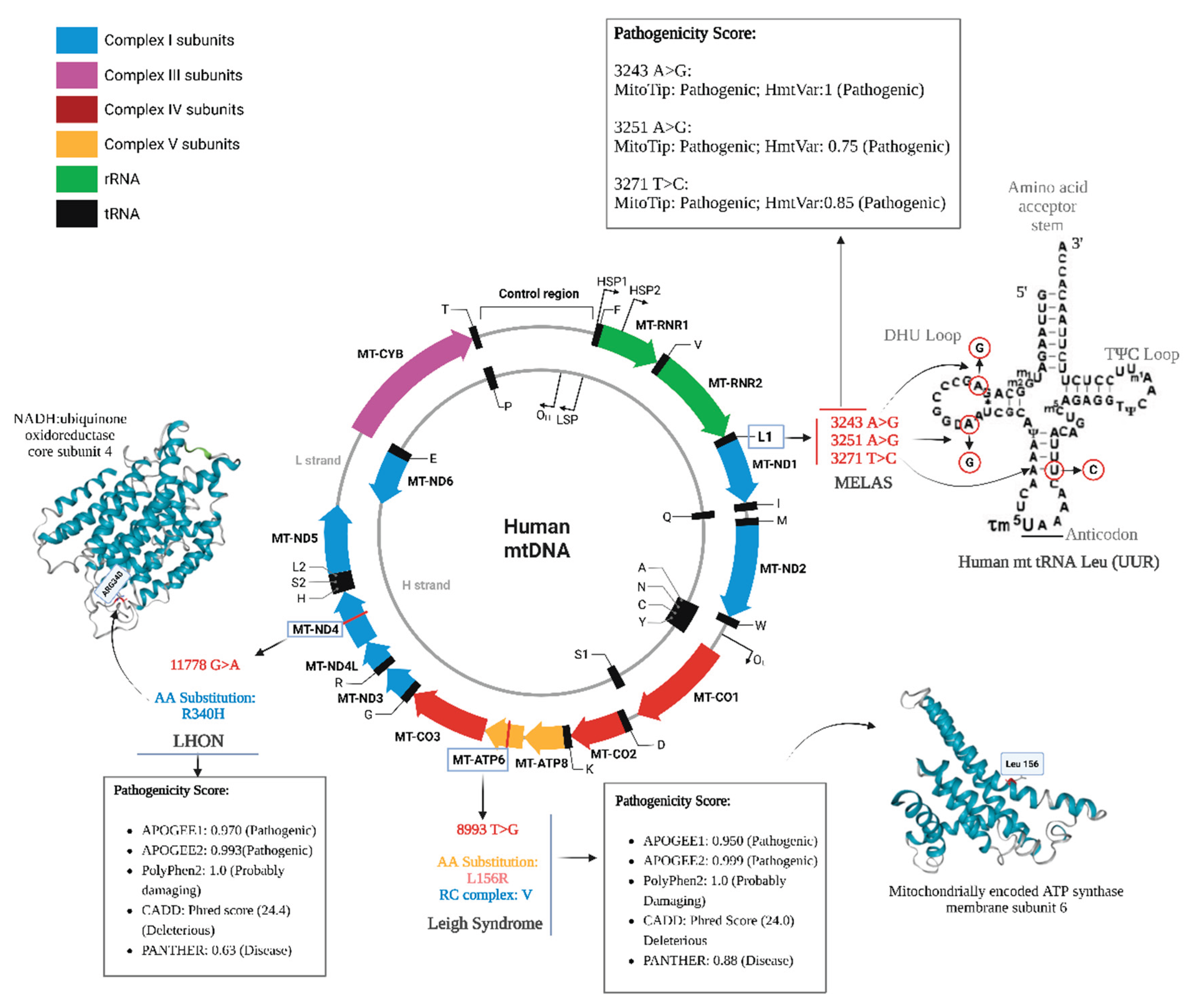
| MT-04 | MELAS | MT-TL1 (tRNA) | 3271 3251 |
T>C A>G |
Heteroplasmy for both nucleotide positions | 82.2% 93.3% |
| MT-06 | MT-TL1 (tRNA) | 3271 3251 |
T>C A>G |
Heteroplasmy for both nucleotide positions | 82.2% 93.3% |
|
| MT-08 | MT-TL1 (tRNA) | 3271 3251 |
T>C A>G |
Heteroplasmy for both nucleotide positions | 82.2% 93.3% |
|
| MT-12 | MT-TL1 (tRNA) | 3271 3251 |
T>C A>G |
Heteroplasmy for both nucleotide positions | 82.2% 93.3% |
|
| MT-20 | MT-TL1 (tRNA) | 3271 3251 |
T>C A>G |
Heteroplasmy for both nucleotide positions | 82.2% 93.3% |
|
| MT-22 | MT-TL1 (tRNA) | 3271 3251 |
T>C A>G |
Heteroplasmy for both nucleotide positions | 82.2% 93.3% |
|
| MT-24 | MT-TL1 (tRNA) | 3271 3251 |
T>C A>G |
Heteroplasmy for both nucleotide positions | 82.2% 93.3% |
|
| MT-28 | MT-TL1 (tRNA) | 3243 | A>G | Heteroplasmy for both nucleotide positions | 97.8% | |
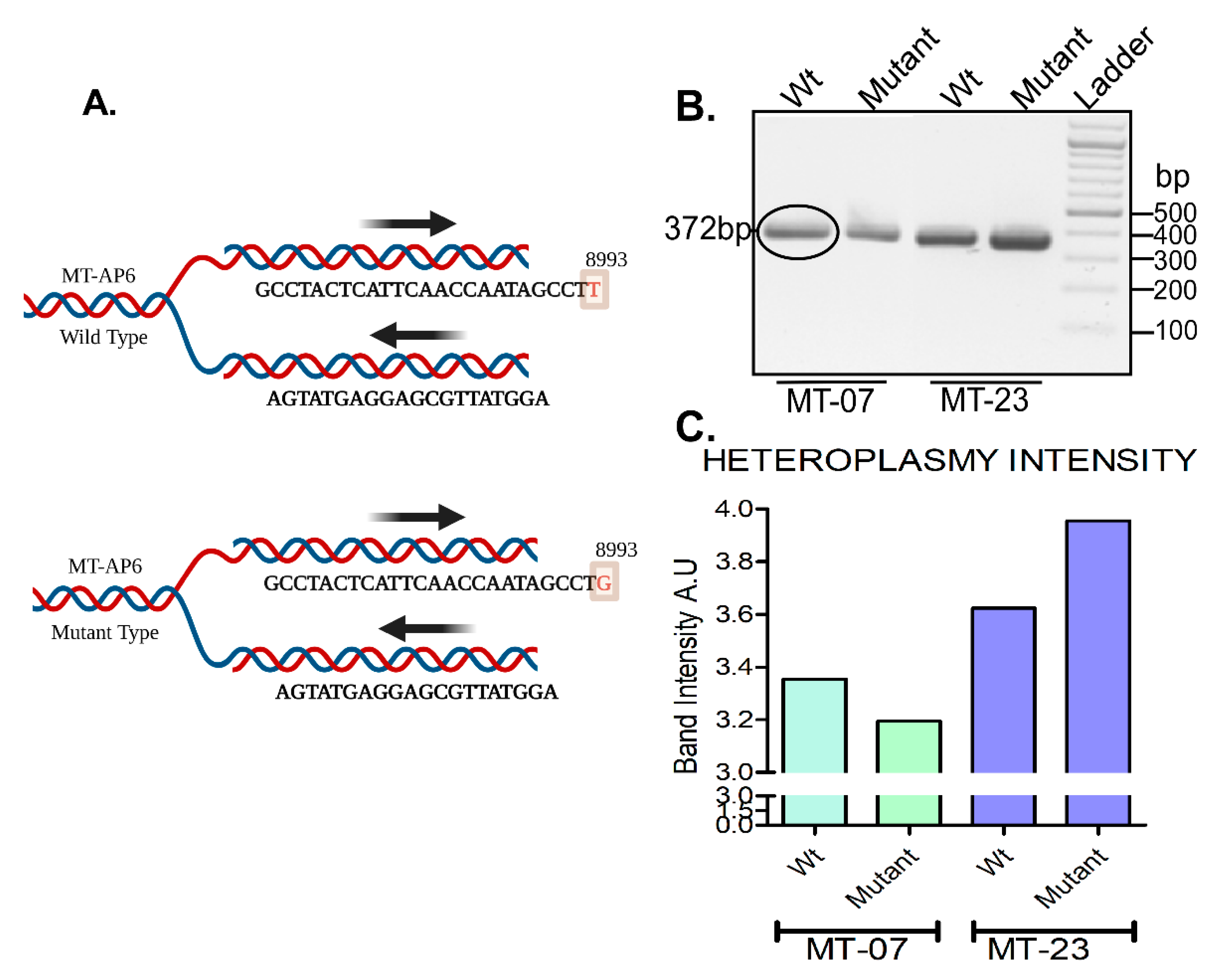
| MT-07 | LEIGH | MT-ATP6 | 8993 | T>G | Heteroplasmy for nucleotide position | 97.8% |
| MT-23 | MT-ATP6 | 8993 | T>G | Heteroplasmy for nucleotide position | 97.8% | |
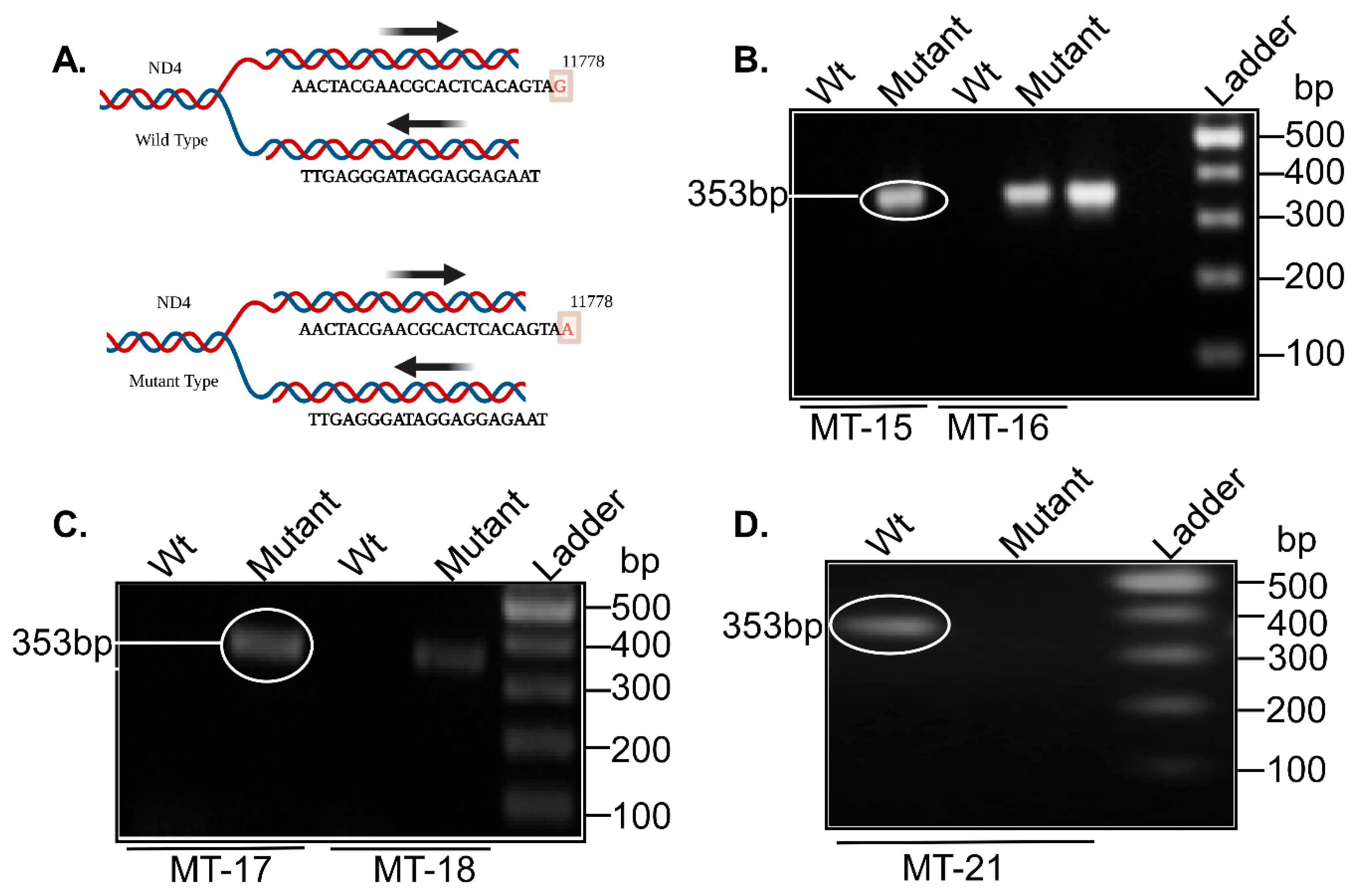
| MT-15 | LHON | MT-ND4 | 11778 | G>A | A mutation was found in the MT-ND4 gene | 100% |
| MT-16 | MT-ND4 | 11778 | G>A | A mutation was found in the MT-ND4 gene | 100% | |
| MT-17 | MT-ND4 | 11778 | G>A | A mutation was found in the MT-ND4 gene | 100% | |
| MT-18 | MT-ND4 | 11778 | G>A | A mutation was found in the MT-ND4 gene | 100% | |
| MT-21 | MT-ND4 | 11778 | G>A | Clinically diagnosed, and genetic testing was performed, but no mutation was found in MT-ND4 gene | 100% | |
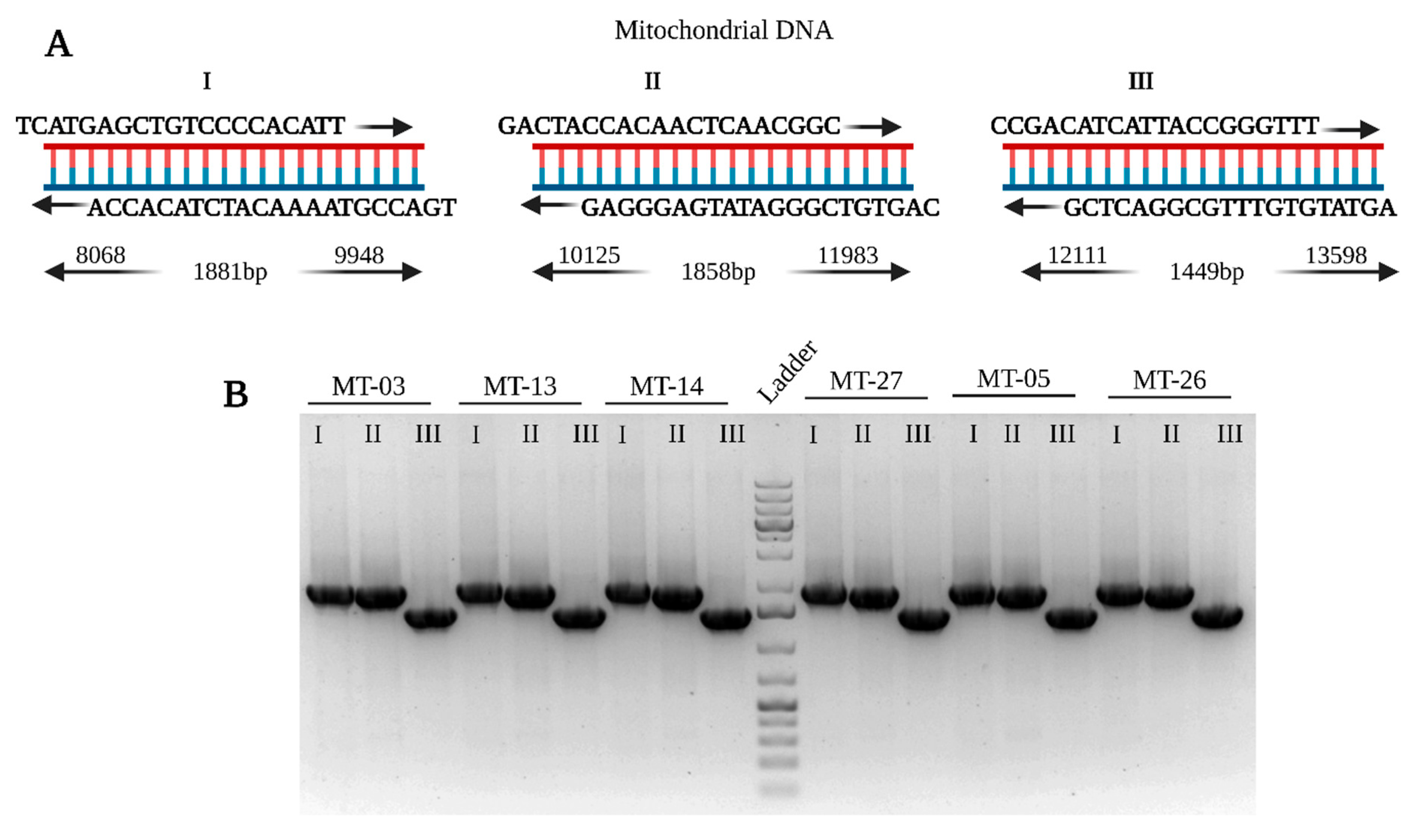
| MT-03 | CPEO | Not Applicable | Not Applicable | Not Applicable | No deletion was found between 8kb-14kb | Not Applicable |
| MT-05 | Not Applicable | Not Applicable | Not Applicable | No deletion was found between 8kb-14kb | Not Applicable | |
| MT-13 | Not Applicable | Not Applicable | Not Applicable | No deletion was found between 8kb-14kb | Not Applicable | |
| MT-14 | Not Applicable | Not Applicable | Not Applicable | No deletion was found between 8kb-14kb | Not Applicable | |
| MT-26 | Not Applicable | Not Applicable | Not Applicable | No deletion was found between 8kb-14kb | Not Applicable | |
| MT-27 | Not Applicable | Not Applicable | Not Applicable | No deletion was found between 8kb-14kb | Not Applicable |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




