Submitted:
13 July 2025
Posted:
29 July 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
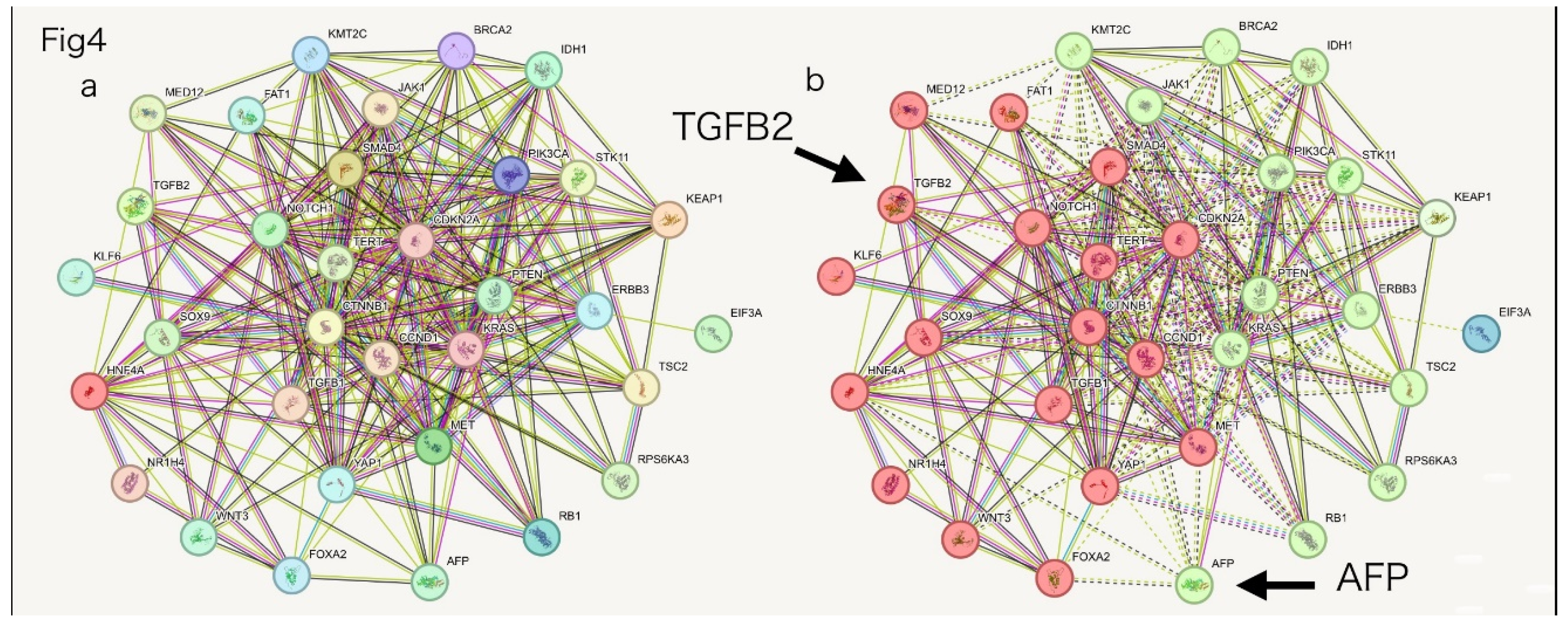
- Tools: ClustalW for evolutionary distance (dnd), STRING database for known and predicted PPIs, and AlphaFold3 (Colab or DeepMind server) for structural prediction and affinity scoring.
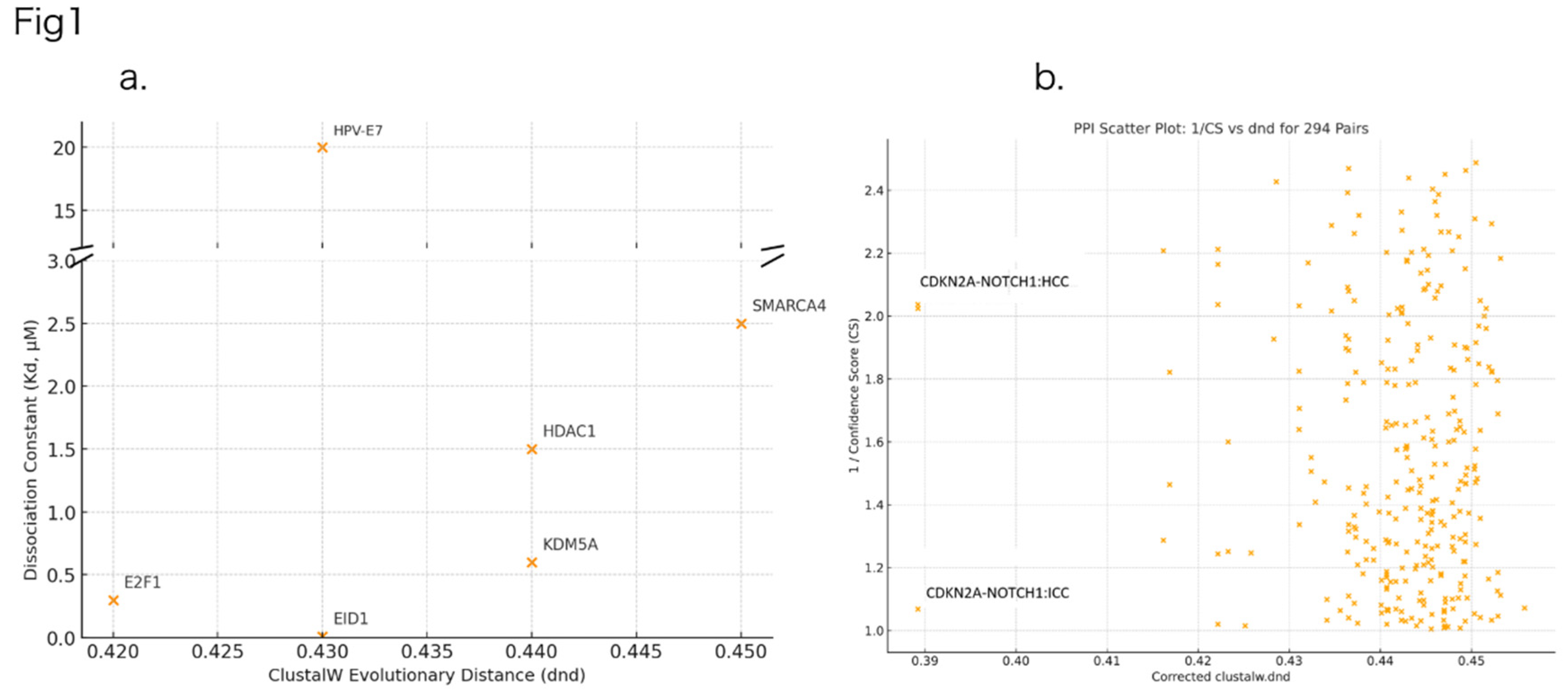
- Dataset: 27,000 cancer-related PPIs from the supplementary Table S2 of Computed Cancer Interactome (5), from which 294 literature-based pairs relevant to HCC and ICC were selected. Since AlphaFold-based structural confidence (CS) is highly dependent on the input amino acid sequences, we utilized canonical UniProt sequences. Database: UniProtKB/Swiss-Prot as of 2025-04-15. Variations in isoforms may affect structural interpretations.
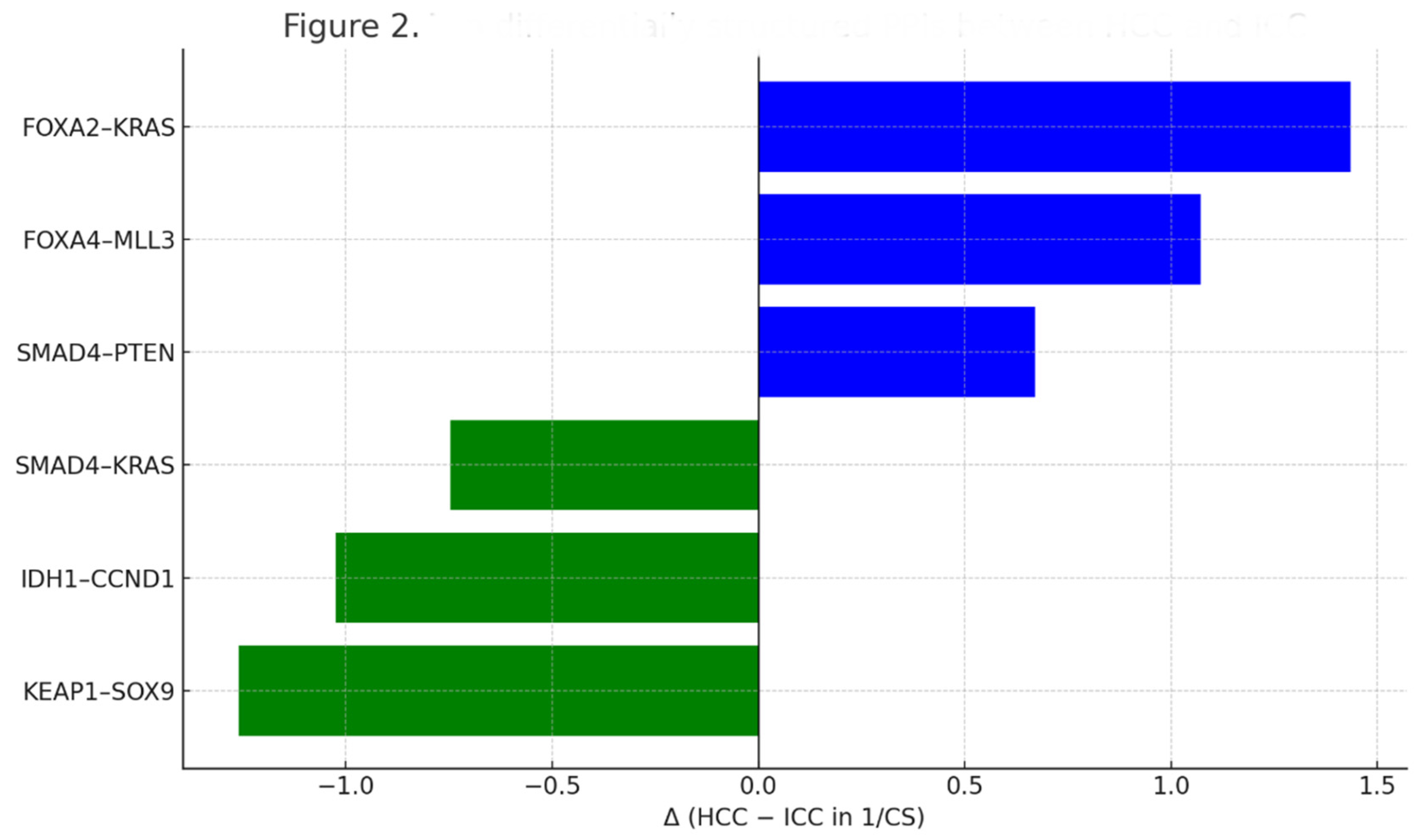
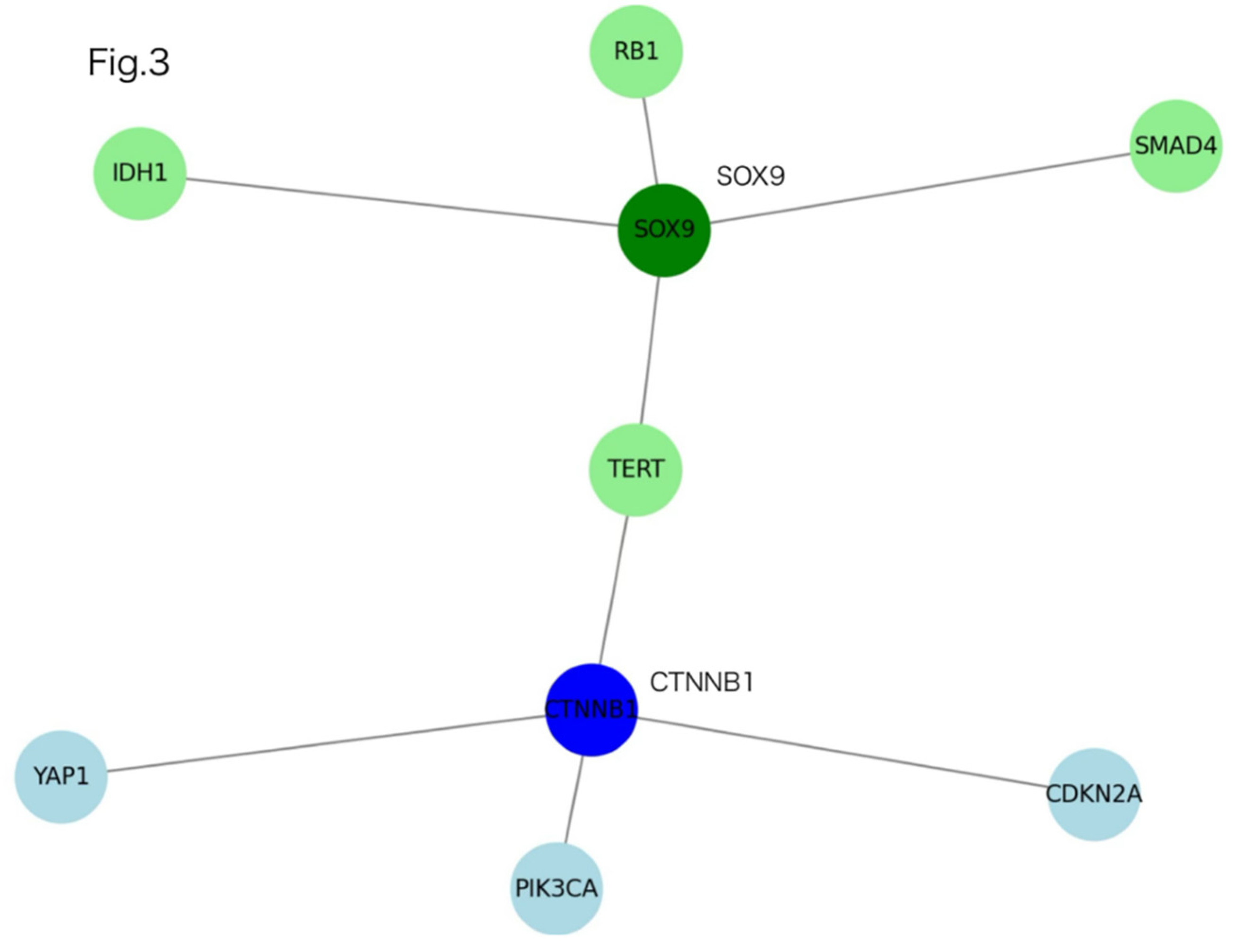
- Analyses: Evolutionary distances (ClustalW dnd), structural affinity scoring (CS), volcano plotting of differential PPI interactions, and STRING-based clustering with k-means.
Results
Discussion
Conclusion and Outlook
References
- Ma L, Wang L, Khatib SA, et al. Single-cell atlas of tumor cell evolution in response to therapy in hepatocellular carcinoma and intrahepatic cholangiocarcinoma. J Hepatol. 2021;75(6):1397–1408. [CrossRef]
- Wodarz D, Newell AC, Komarova NL. Passenger mutations can accelerate tumour suppressor gene inactivation in cancer evolution. J R Soc Interface. 2018;15(143):20170967. [CrossRef]
- Jumper J, Evans R, Pritzel A, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596(7873):583–589. [CrossRef]
- Varadi M, Anyango S, Deshpande M, et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 2022;50(D1):D439–D444. [CrossRef]
- Zhang J, et al. Computed cancer interactome explains the effects of somatic mutations in cancers. Protein Sci. 2022;31(12):e4479. [CrossRef]
- Dick FA et al. Structural basis for tunable affinity and specificity of LxCxE-dependent protein interactions with the retinoblastoma protein family. Cell Rep. 2018.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



