Submitted:
20 July 2025
Posted:
21 July 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset Preparation
2.2. Compositional Analysis
2.3. Calculation of Physicochemical Properties
2.4. Use of Non-Parametric Methods for Antifungal Compound Identification
2.5. Machine Learning
2.5.1. Hyperparameter Tuning
2.5.2. Feature Selection
2.5.3. Neural Network
2.5.4. Model Training and Validation
2.5.5. Chemical Class Based Cross Validation
3. Results
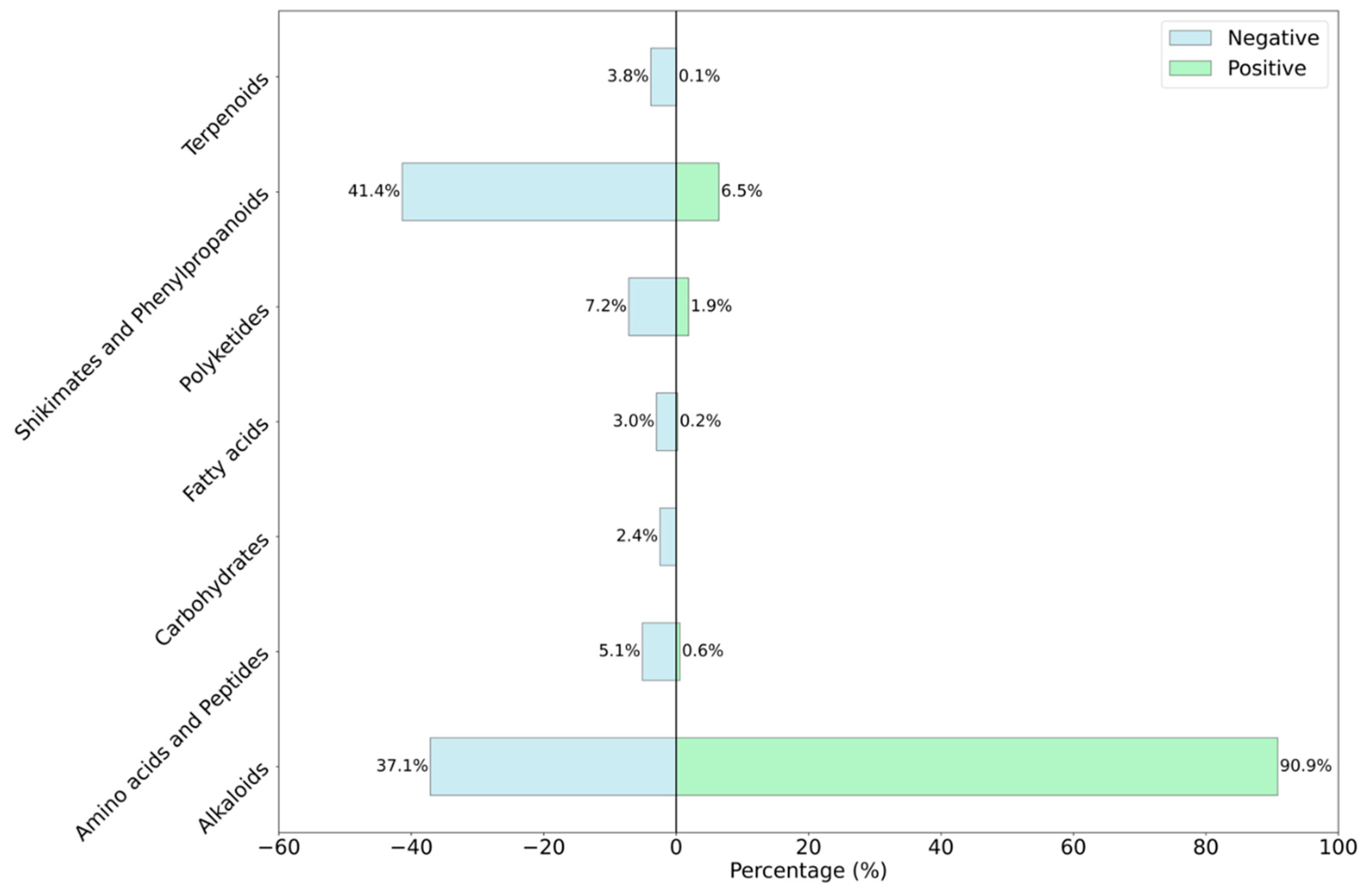
3.1. Compositional Analysis
3.2. Calculation of Physicochemical Properties
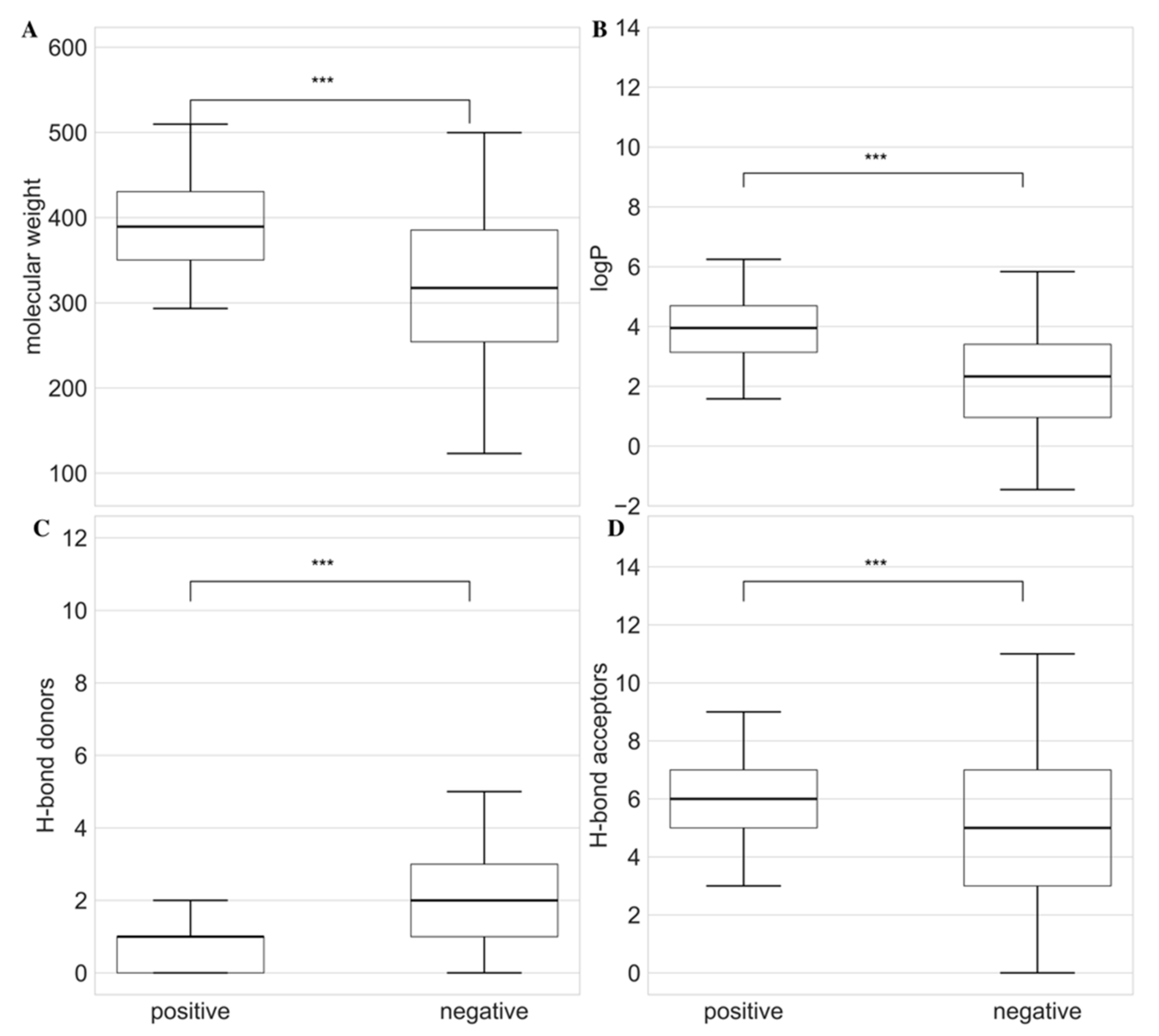
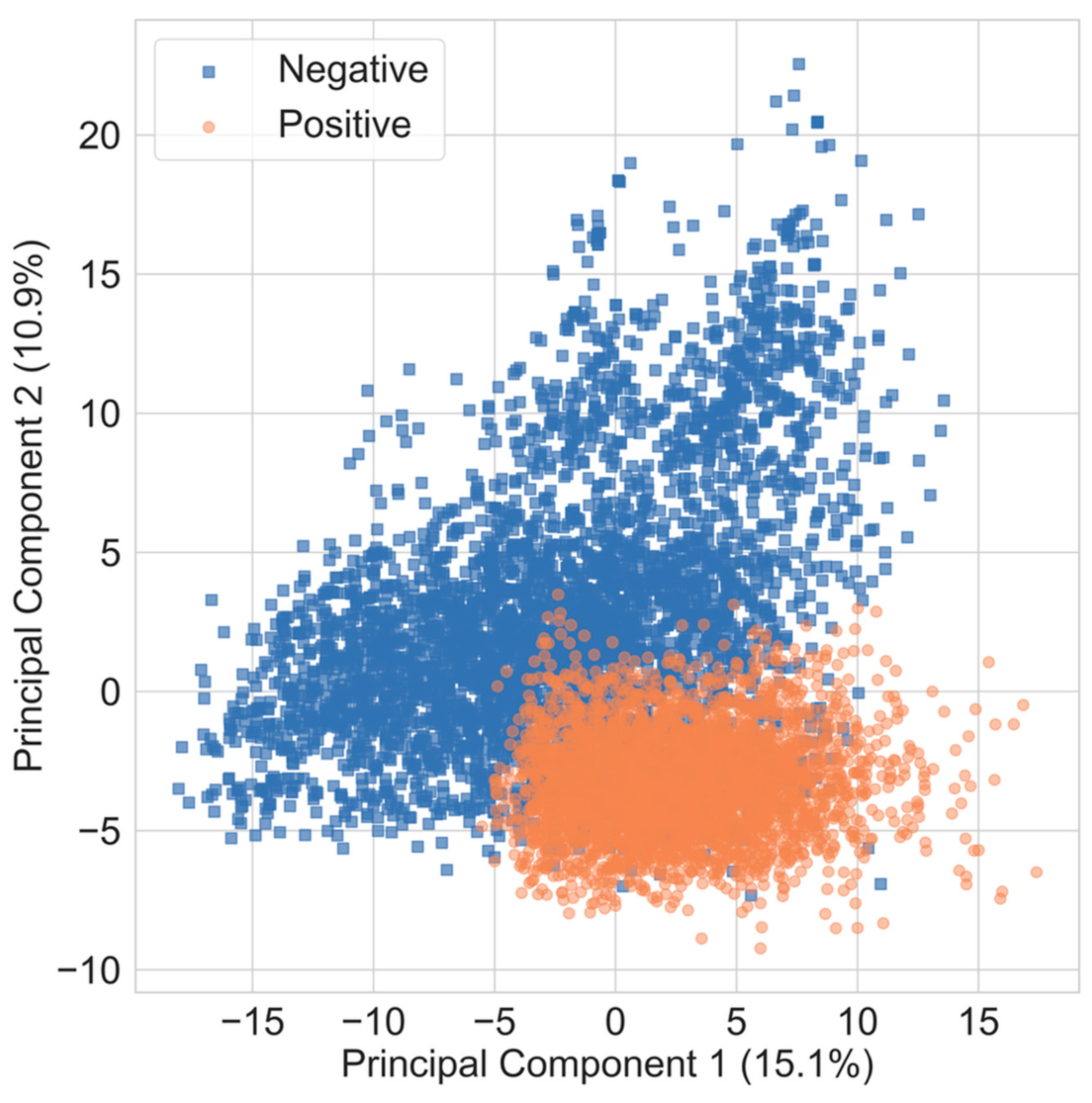
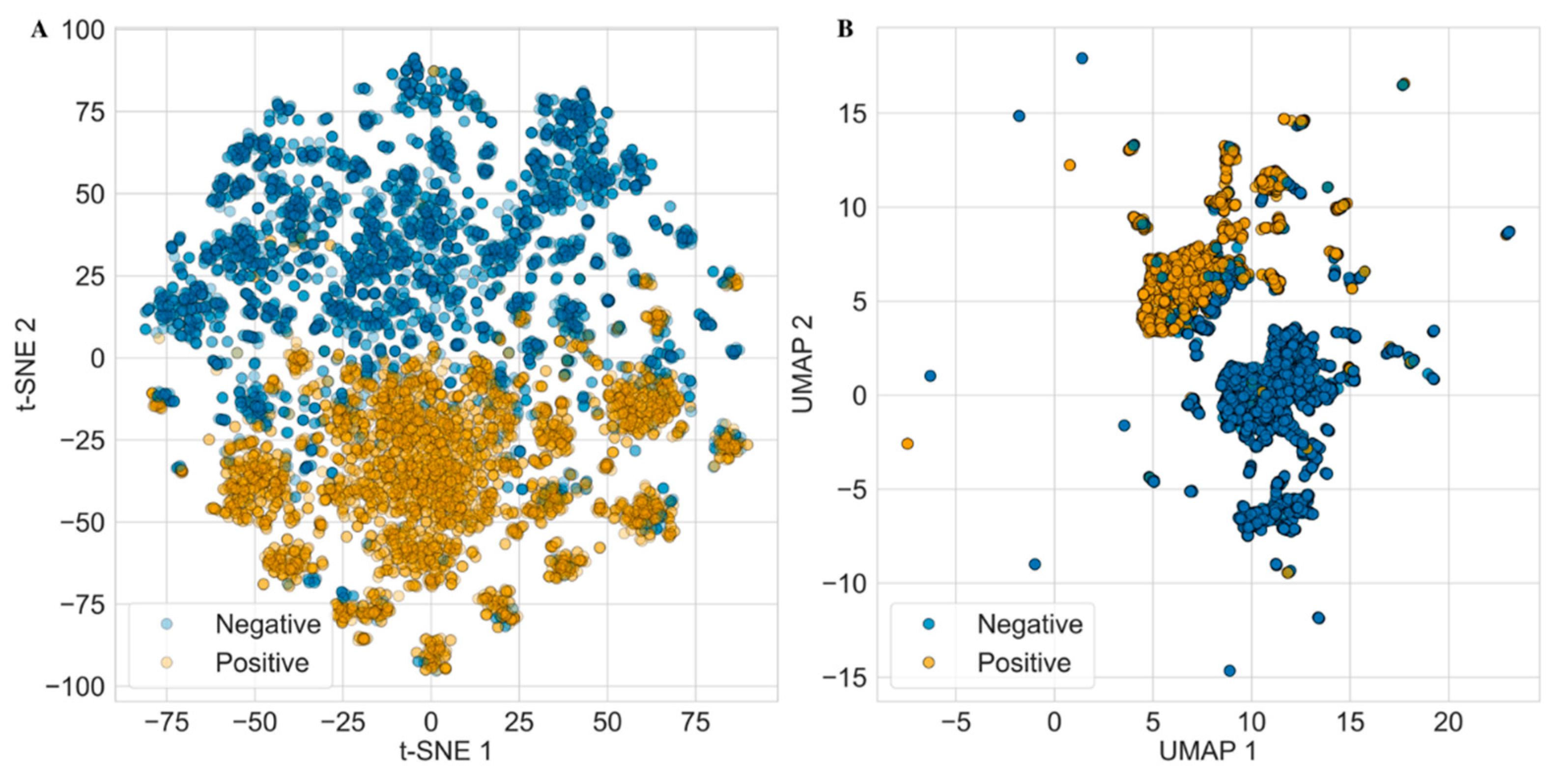
3.3. Use of Non-Parametric Methods for Antifungal Compound Identification
3.4. Machine Learning
3.4.1. Feature Selection
3.4.2. Model Training and Validation
Random Split Cross Validation
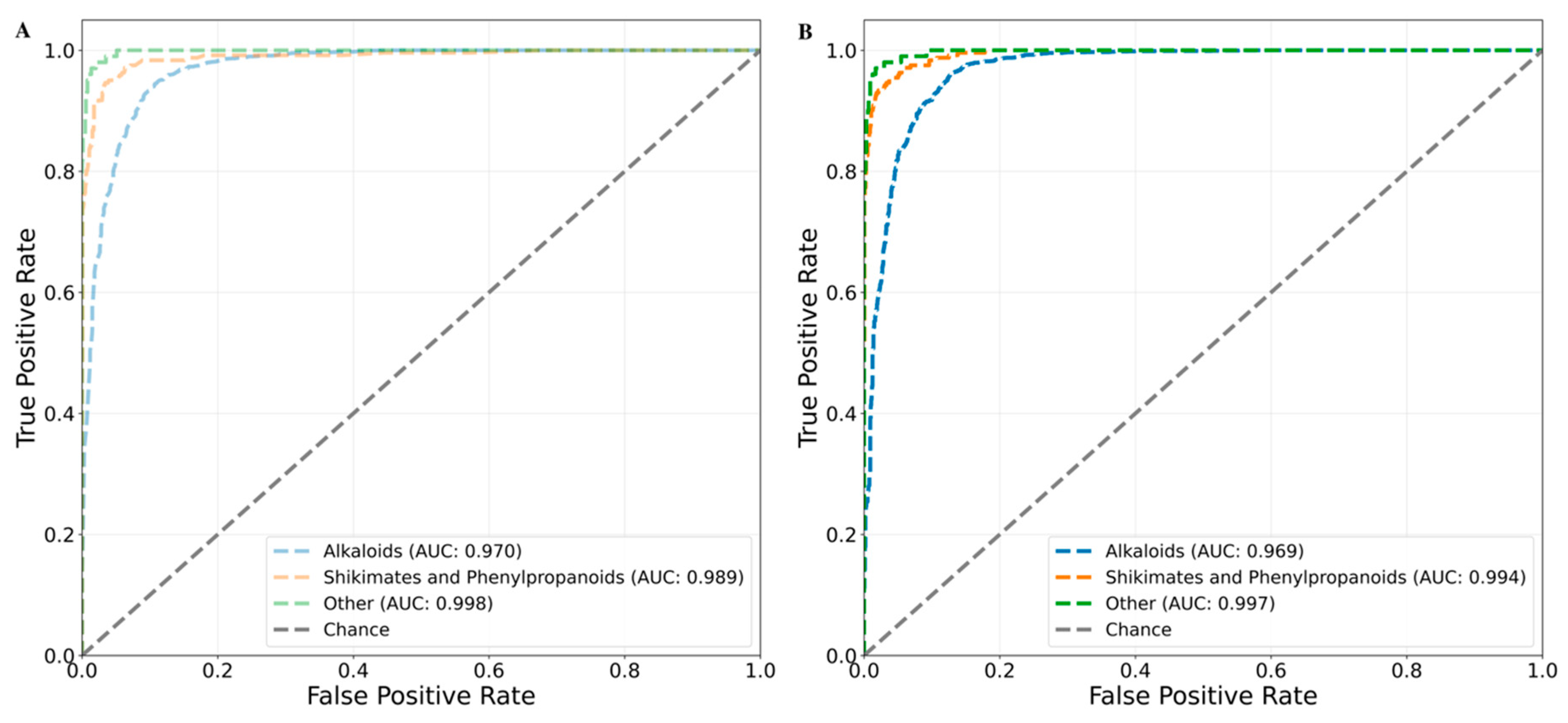
Chemical Class Based Cross Validation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgements
Conflicts of Interest
Abbreviations
| RF | Random Forest |
| SVM | Support Vector Machine |
| XGBoost | Extreme gradient boosting tree |
| NN | Neural Network |
| PCA | Principal Component Analysis |
| t-SNE | t-Distributed Stochastic Neighbor Embedding |
| UMAP | Uniform Manifold Approximation and Projection |
| MCC | Matthew’s Correlation Coefficient |
| AUC | Area under the ROC curve |
References
- Bongomin, F.; et al. Global and Multi-National Prevalence of Fungal Diseases-Estimate Precision. J Fungi (Basel) 2017, 3. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; et al. Hidden killers: human fungal infections . Sci Transl Med 2012, 4, 165rv13. [Google Scholar] [CrossRef] [PubMed]
- Organization, W.H. WHO fungal priority pathogens list to guide research, development and public health action. 2022: World Health Organization.
- Perfect, J.R. ; The antifungal pipeline: a reality check . Nat Rev Drug Discov 2017, 16, 603–616. [Google Scholar] [CrossRef] [PubMed]
- Cowen, L.E.; et al. Harnessing Hsp90 function as a powerful, broadly effective therapeutic strategy for fungal infectious disease . Proc Natl Acad Sci U S A 2009, 106, 2818–23. [Google Scholar] [CrossRef]
- Satoh, K.; et al. Candida auris sp. nov., a novel ascomycetous yeast isolated from the external ear canal of an inpatient in a Japanese hospital. Microbiol Immunol 2009, 53, 41–4. [Google Scholar] [CrossRef]
- Vamathevan, J.; et al. Applications of machine learning in drug discovery and development . Nat Rev Drug Discov 2019, 18, 463–477. [Google Scholar] [CrossRef]
- Chen, H.; et al. The rise of deep learning in drug discovery . Drug Discov Today 2018, 23, 1241–1250. [Google Scholar] [CrossRef]
- Rifaioglu, A.S.; et al. Recent applications of deep learning and machine intelligence on in silico drug discovery: methods, tools and databases . Brief Bioinform 2019, 20, 1878–1912. [Google Scholar] [CrossRef]
- Tran, T.P.; et al. Prediction of kinase inhibitor response using activity profiling, in vitro screening, and elastic net regression . BMC Syst Biol 2014, 8, 74. [Google Scholar] [CrossRef]
- Wu, Z.; et al. MoleculeNet: a benchmark for molecular machine learning . Chem Sci 2018, 9, 513–530. [Google Scholar] [CrossRef]
- Sheridan, R.P. ; Time-split cross-validation as a method for estimating the goodness of prospective prediction . J Chem Inf Model 2013, 53, 783–90. [Google Scholar] [CrossRef] [PubMed]
- Wallach, I. and A. Heifets, Most Ligand-Based Classification Benchmarks Reward Memorization Rather than Generalization. . J Chem Inf Model 2018, 58, 916–932. [Google Scholar] [CrossRef] [PubMed]
- Campoy, S. and J.L. Adrio, Antifungals . Biochem Pharmacol 2017, 133, 86–96. [Google Scholar] [CrossRef]
- Wishart, D.S.; et al. HMDB 5.0: the Human Metabolome Database for 2022. . Nucleic Acids Res 2022, 50, D622–D631. [Google Scholar] [CrossRef]
- Chung, N.C.; et al. Jaccard/Tanimoto similarity test and estimation methods for biological presence-absence data . BMC Bioinformatics 2019, 20 (Suppl. S15), 644. [Google Scholar] [CrossRef]
- Kuwahara, H. and X. Gao, Analysis of the effects of related fingerprints on molecular similarity using an eigenvalue entropy approach. . J Cheminform 2021, 13, 27. [Google Scholar] [CrossRef]
- Kim, H.W.; et al. NPClassifier: A Deep Neural Network-Based Structural Classification Tool for Natural Products . J Nat Prod 2021, 84, 2795–2807. [Google Scholar] [CrossRef]
- Bento, A.P.; et al. An open source chemical structure curation pipeline using RDKit . J Cheminform 2020, 12, 51. [Google Scholar] [CrossRef]
- Mishra, P.; et al. Application of Student's t-test, Analysis of Variance, and Covariance . Ann Card Anaesth 2019, 22, 407–411. [Google Scholar] [CrossRef]
- Groth, D.; et al. Principal components analysis . Methods Mol Biol 2013, 930, 527–47. [Google Scholar]
- Jolliffe, I.T. and J. Cadima, Principal component analysis: A review and recent developments. . Philos Trans A Math Phys Eng Sci 2016, 374, 20150202. [Google Scholar]
- Cieslak, M.C.; et al. t-Distributed Stochastic Neighbor Embedding (t-SNE): A tool for eco-physiological transcriptomic analysis . Mar Genomics 2020, 51, 100723. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, G.; et al. Uniform Manifold Approximation and Projection (UMAP) Reveals Composite Patterns and Resolves Visualization Artifacts in Microbiome Data . mSystems 2021, 6, e0069121. [Google Scholar] [CrossRef]
- Khan, H. ; M.S. Mubarak, and S. Amin, Antifungal Potential of Alkaloids As An Emerging Therapeutic Target. Curr Drug Targets 2017, 18, 1825–1835. [Google Scholar] [CrossRef] [PubMed]
- Thawabteh, A.M.; et al. Antibacterial Activity and Antifungal Activity of Monomeric Alkaloids. Toxins (Basel) 2024, 16. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; et al. In vivo and in vitro antifungal activities of five alkaloid compounds isolated from Picrasma quassioides (D. Don) Benn against plant pathogenic fungi. Pestic Biochem Physiol 2022, 188, 105246. [Google Scholar] [CrossRef]
- Cushnie, T.P.B. Cushnie, and A.J. Lamb, Alkaloids: an overview of their antibacterial, antibiotic-enhancing and antivirulence activities. Int J Antimicrob Agents 2014, 44, 377–86. [Google Scholar] [CrossRef]
- Aniszewski, T. ; Alkaloids: Chemistry, biology, ecology, and applications. 2015: Elsevier.
- Kittakoop, P.C. Mahidol, and S. Ruchirawat, Alkaloids as important scaffolds in therapeutic drugs for the treatments of cancer, tuberculosis, and smoking cessation. Curr Top Med Chem 2014, 14, 239–52. [Google Scholar] [CrossRef]
- Upadhyay, S.; et al. Subcellular Compartmentalization and Trafficking of the Biosynthetic Machinery for Fungal Melanin . Cell Rep 2016, 14, 2511–8. [Google Scholar] [CrossRef]
- Herrmann, K.M. and L.M. Weaver, THE SHIKIMATE PATHWAY . Annu Rev Plant Physiol Plant Mol Biol 1999, 50, 473–503. [Google Scholar] [CrossRef]
- Dixon, R.A. and N.L. Paiva, Stress-Induced Phenylpropanoid Metabolism . Plant Cell 1995, 7, 1085–1097. [Google Scholar] [CrossRef]
- Lazaridis, T. and G. Hummer, Classical Molecular Dynamics with Mobile Protons . J Chem Inf Model 2017, 57, 2833–2845. [Google Scholar] [CrossRef]
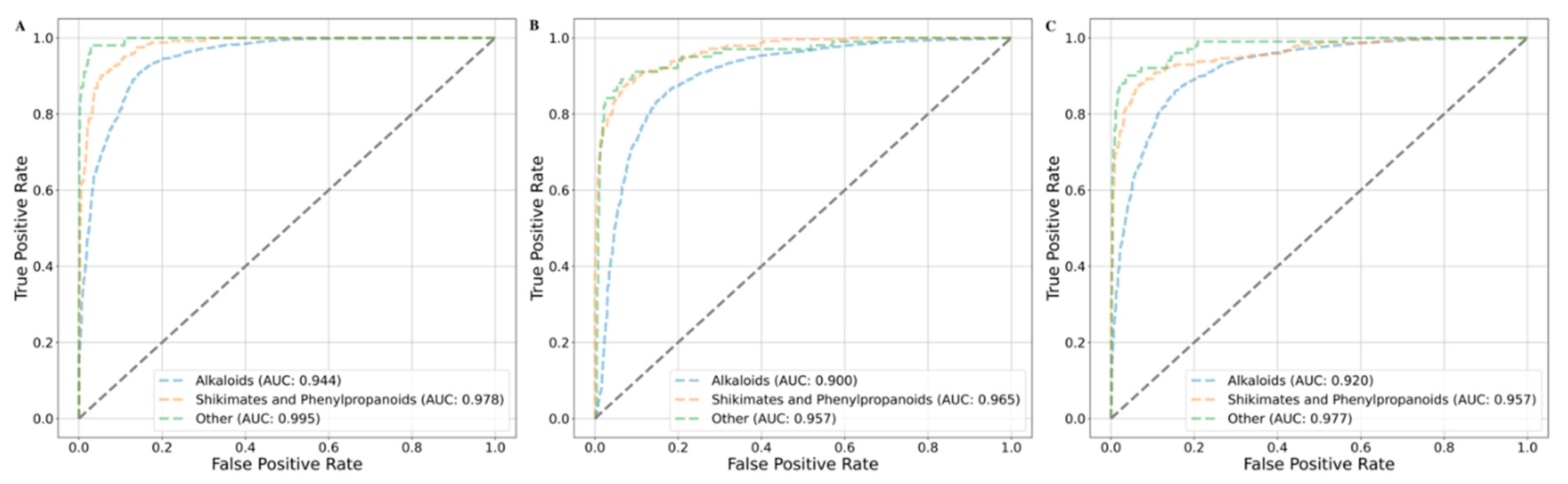
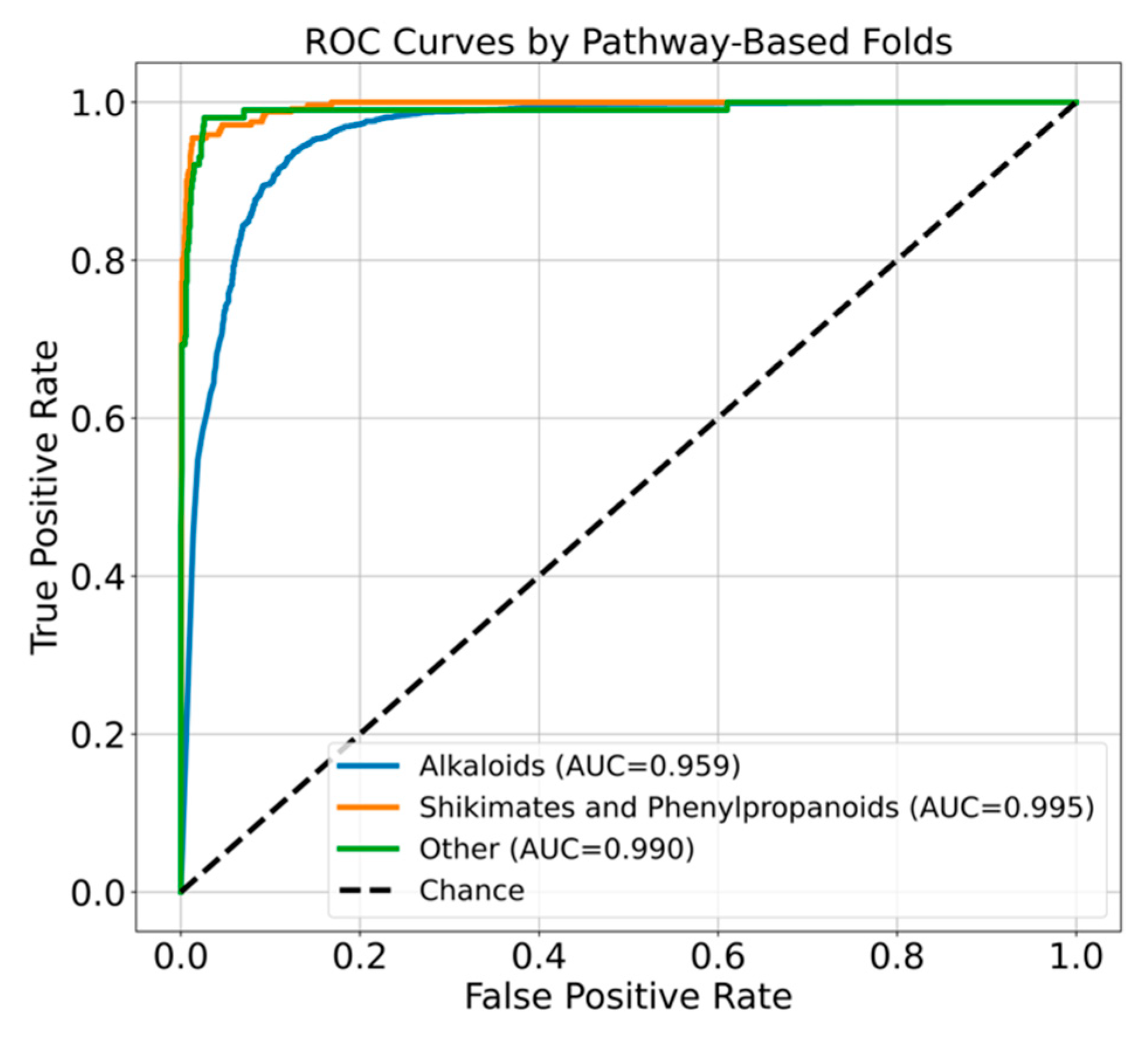
| Models | Balanced Accuracy | Precision | Recall | F1 | MCC | AUC |
|---|---|---|---|---|---|---|
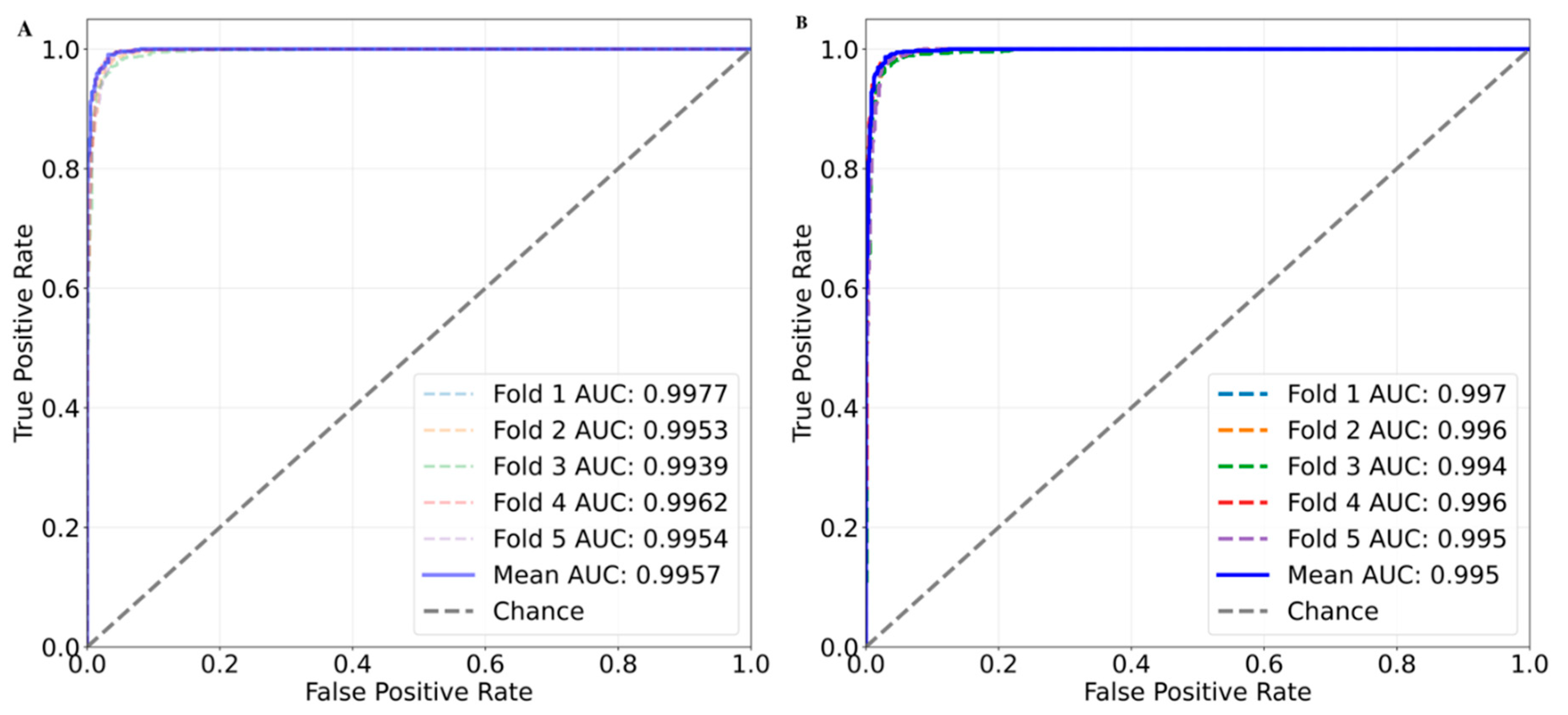
| Random Forest | 0.972±0.002 | 0.963±0.003 | 0.979±0.003 | 0.971±0.002 | 0.944±0.004 | 0.996±0.001 |
| XGBoost | 0.973±0.003 | 0.964±0.003 | 0.979±0.006 | 0.971±0.002 | 0.945±0.006 | 0.995±0.001 |
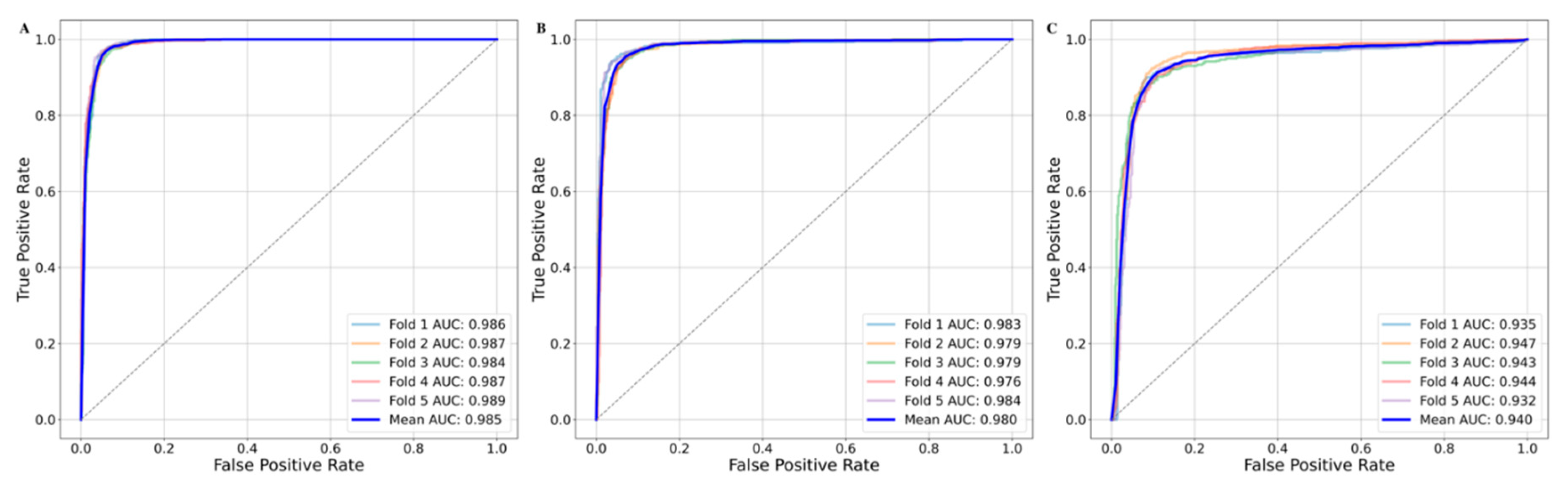
| SVM Polynomial | 0.942±0.006 | 0.938±0.012 | 0.942±0.008 | 0.940±0.007 | 0.885±0.013 | 0.980±0.003 |
| SVM RBF | 0.955±0.003 | 0.936±0.008 | 0.971±0.004 | 0.953±0.003 | 0.909±0.006 | 0.986±0.002 |
| SVM Sigmoid | 0.901±0.007 | 0.897±0.008 | 0.897±0.011 | 0.897±0.007 | 0.803±0.013 | 0.940±0.006 |
| Neural Network | 0.977±0.004 | 0.976±0.005 | 0.977±0.006 | 0.976±0.004 | 0.954±0.008 | 0.996±0.001 |
| Models | Balanced Accuracy | Precision | Recall | F1 | MCC | AUC |
|---|---|---|---|---|---|---|
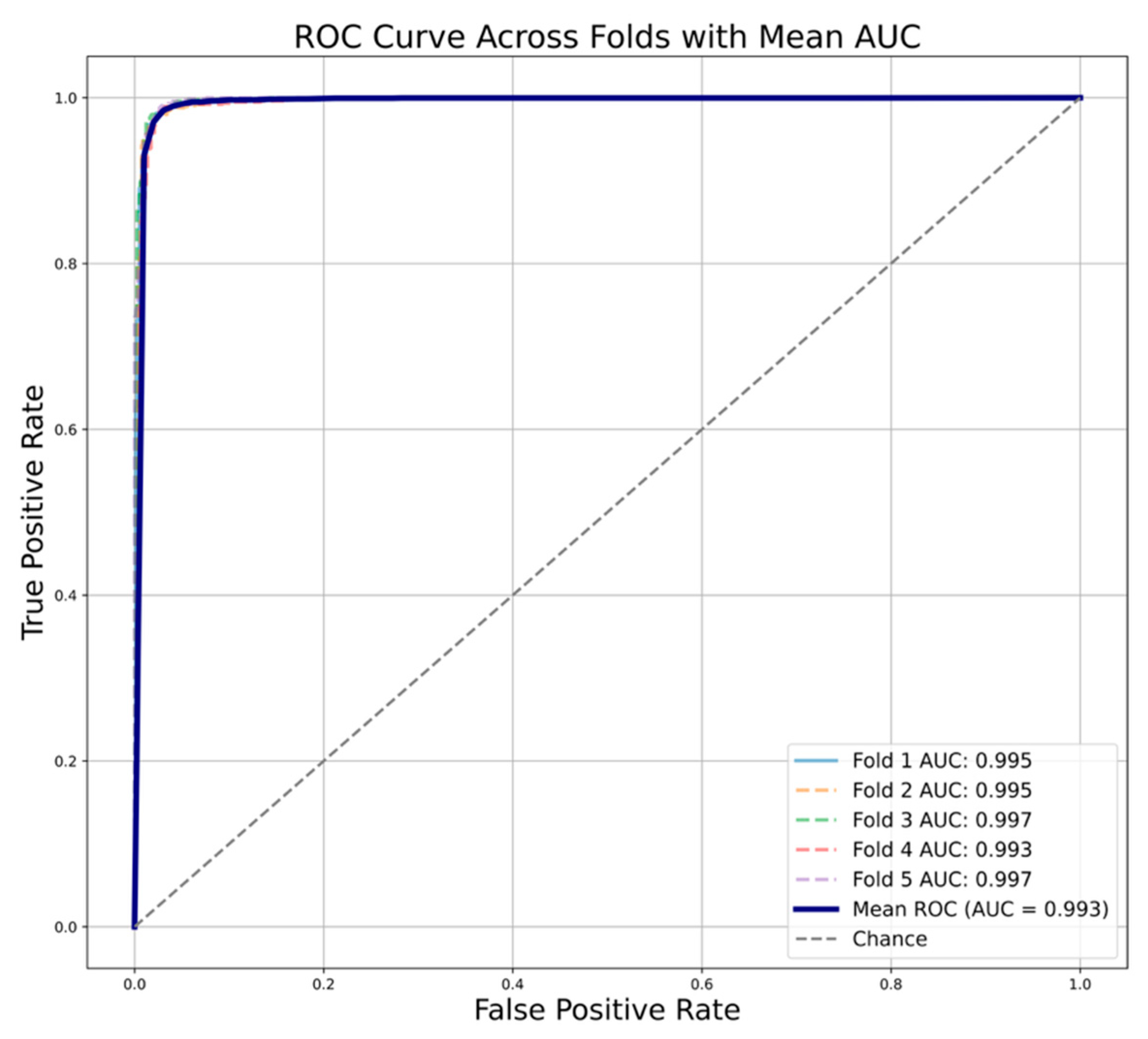
| Random Forest | 0.933 | 0.923 | 0.923 | 0.922 | 0.878 | 0.986 |
| XGBoost | 0.933 | 0.933 | 0.919 | 0.926 | 0.883 | 0.986 |
| SVM Polynomial | 0.880 | 0.708 | 0.861 | 0.764 | 0.669 | 0.941 |
| SVM RBF | 0.911 | 0.819 | 0.882 | 0.841 | 0.759 | 0.972 |
| SVM Sigmoid | 0.881 | 0.838 | 0.848 | 0.843 | 0.754 | 0.951 |
| Neural Network | 0.932 | 0.928 | 0.935 | 0.914 | 0.862 | 0.981 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




