Submitted:
15 July 2025
Posted:
16 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Works
3. Materials and Methods
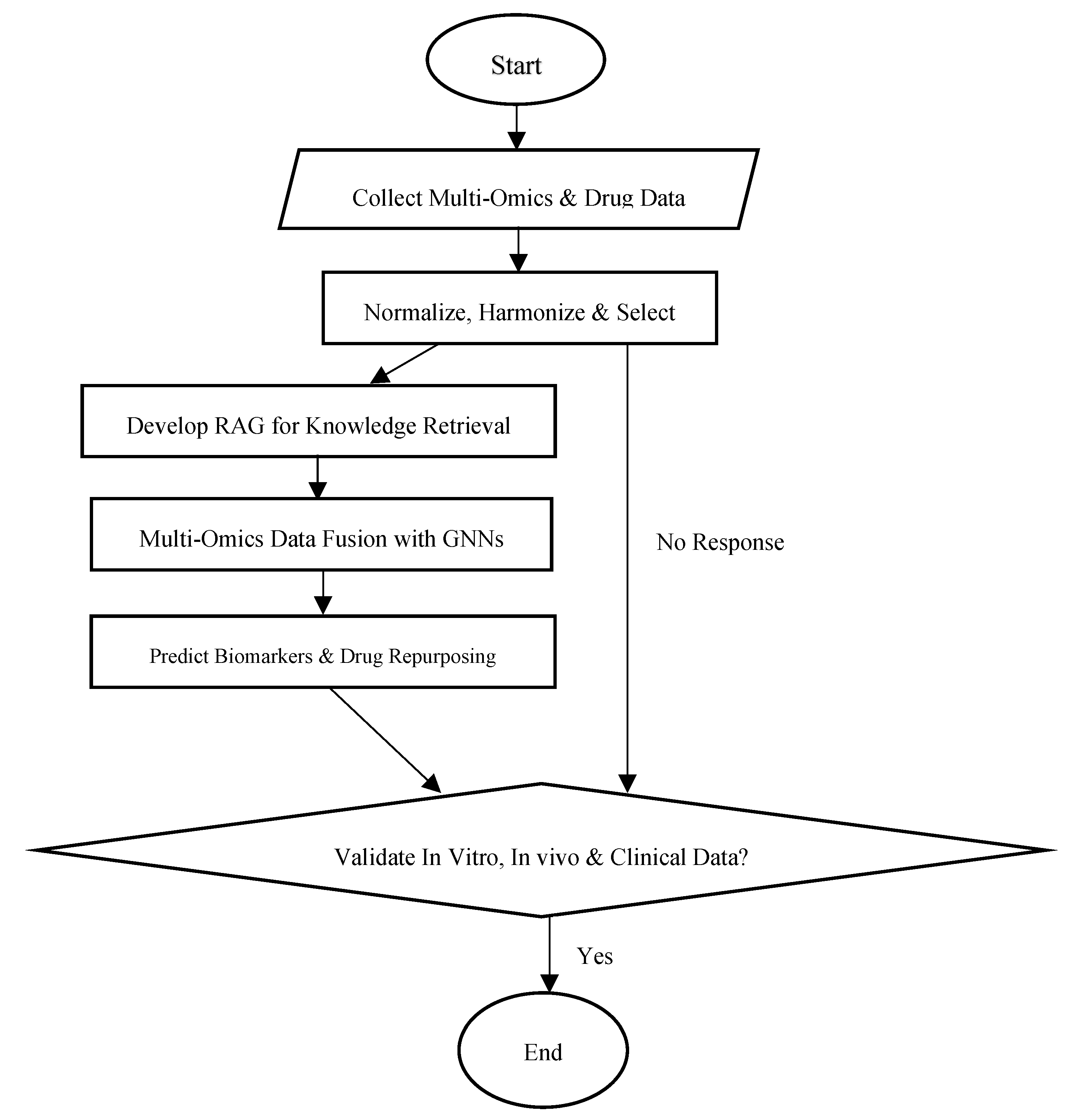
3.1. The ARMOA Framework
3.2. Data Collection and Preprocessing
3.3. Agentic RAG System Development
| Algorithm 1: Agentic RAG System pseudocode |
| specify knowledge_base, query, and agentic_rag_system: # Step 1: obtain pertinent papers documents = retrieve_documents(query, knowledge_base) # Step 2: Synthesize knowledge using LLM summary = llm_synthesize(documents). #Step 3: Update the knowledge base use knowledge_base.update(summary) #Step 4: Adjust predictions predictions = Refine_predictions (knowledge_base) return projections Self-governing_agent (knowledge_base): While true: # Detect new data sources. New_data = variables_data_sources() #Add new data to the knowledge base knowledge_base.update(new_data). # Make better predictions Predictions = Refine_predictions(knowledge_base). # Assessment and revision of agent policies predictions agent_policy.update |
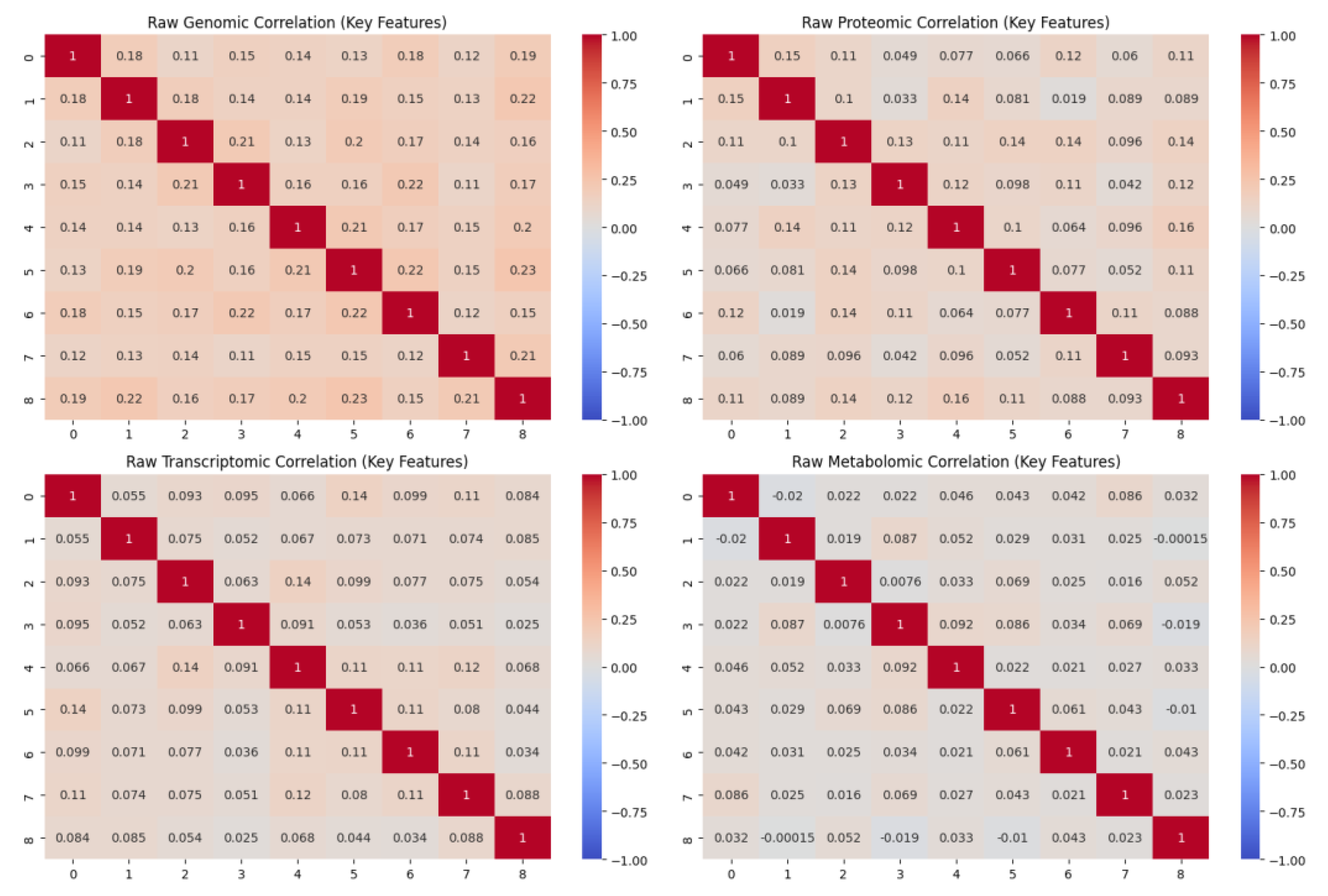
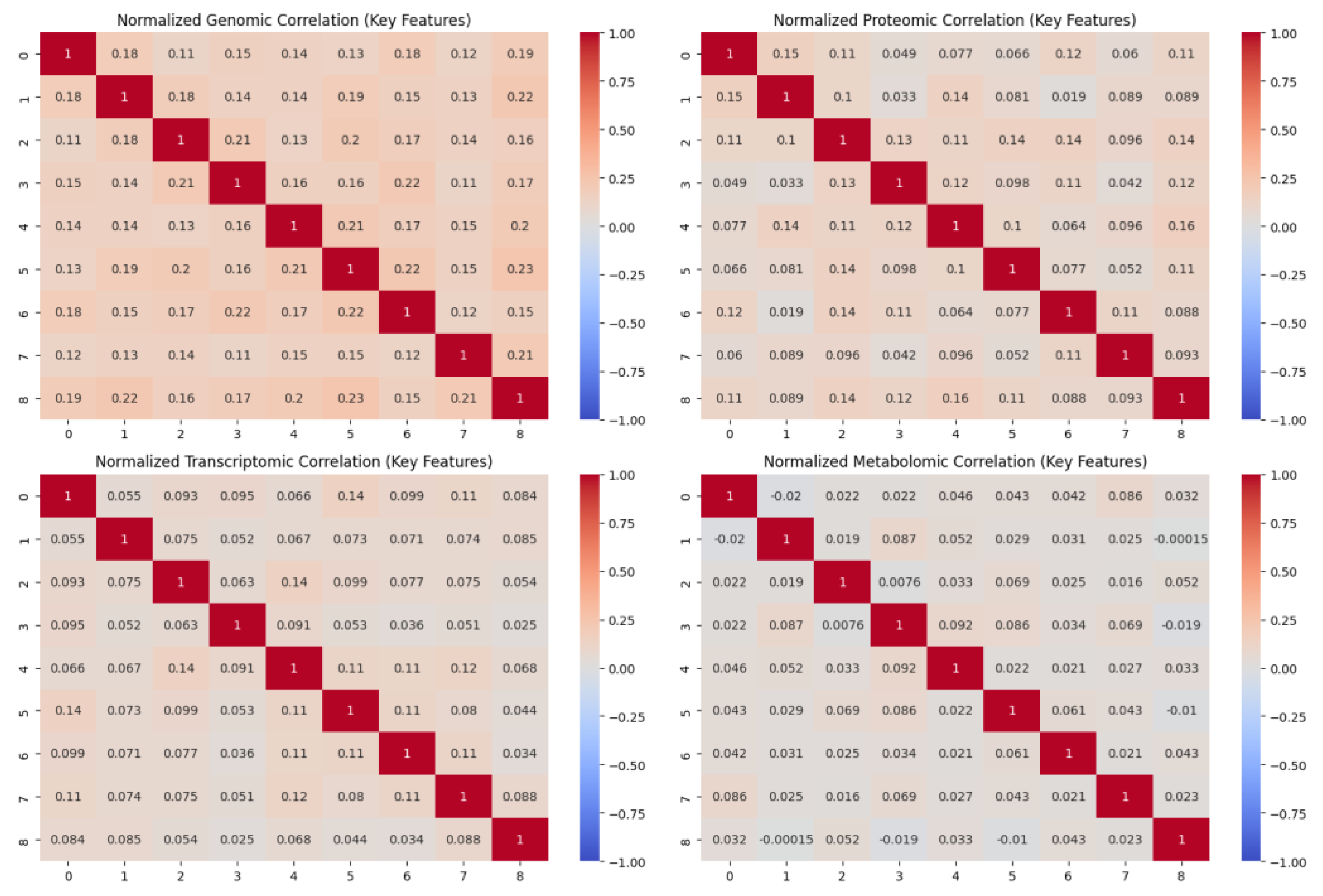
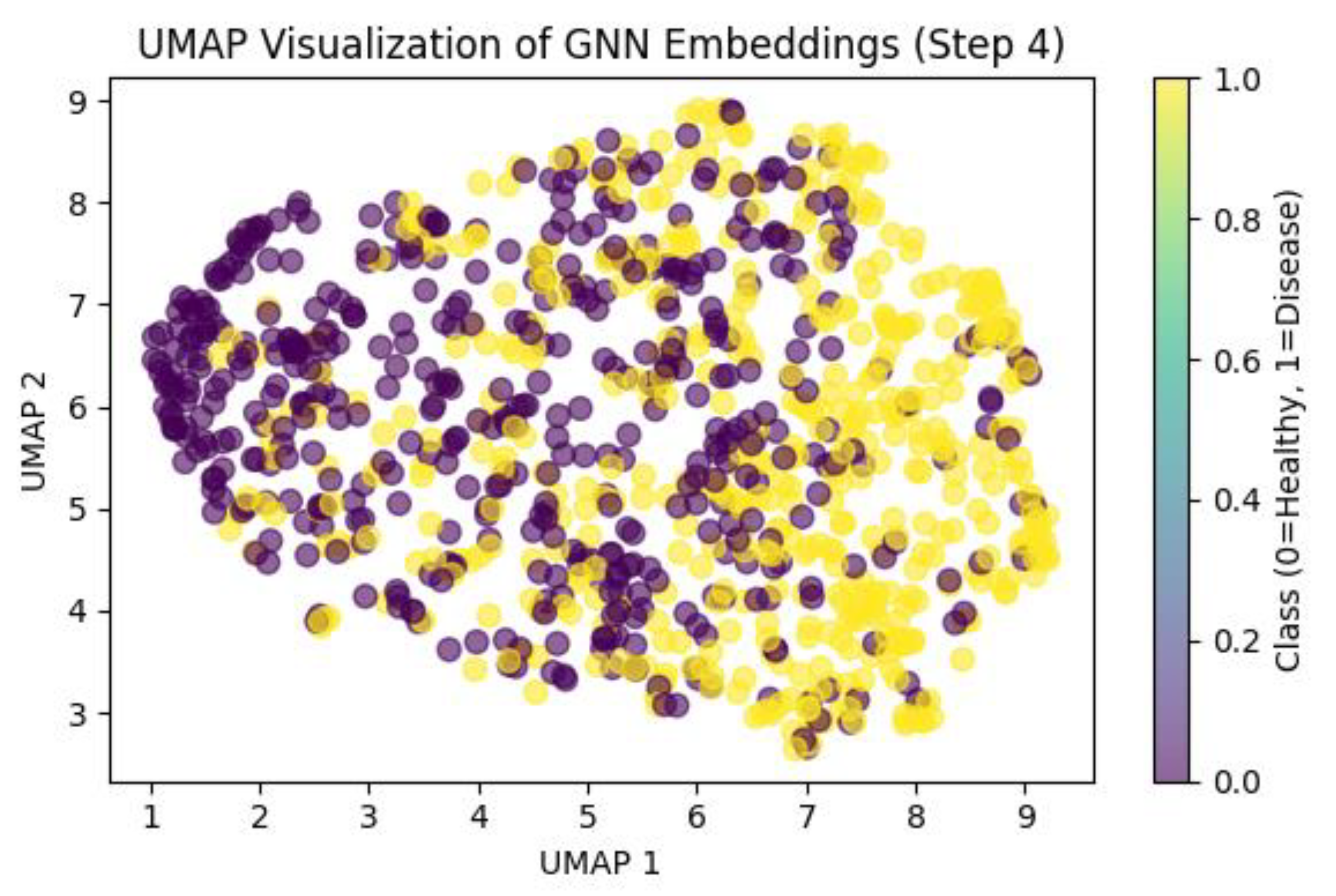
3.4. Multi-Omics Data Integration
| Algorithm 2: GNN-based pathway Pseudocode |
| def gnn_pathway_model(graph, attributes, layers): for node in graph.nodes: for layer in range(layers): neighbors(node) = graph.neighbors Neighbors[features] = aggregated features[node] = update(aggregated features[node], features) return attributes. |
3.5. Predictive Modeling and Validation
| Algorithm 3: drug repurposing |
| def drug_repurposing(omics_data, pathway_activity): train_random_forest(omics_data, pathway_activity) model Predict_drugs(model, omics_data) drug_candidates return drug candidates In_vitro results = test_cell_lines(drug_candidates) In_vivo results = test_mouse_models(drug_candidates) results of def validate_predictions(drug_candidates) In_vitro, in vivo, and clinical data return clinical_results = analyze_clinical_trials(drug_candidates). |
4. Results
5. Conclusions
References
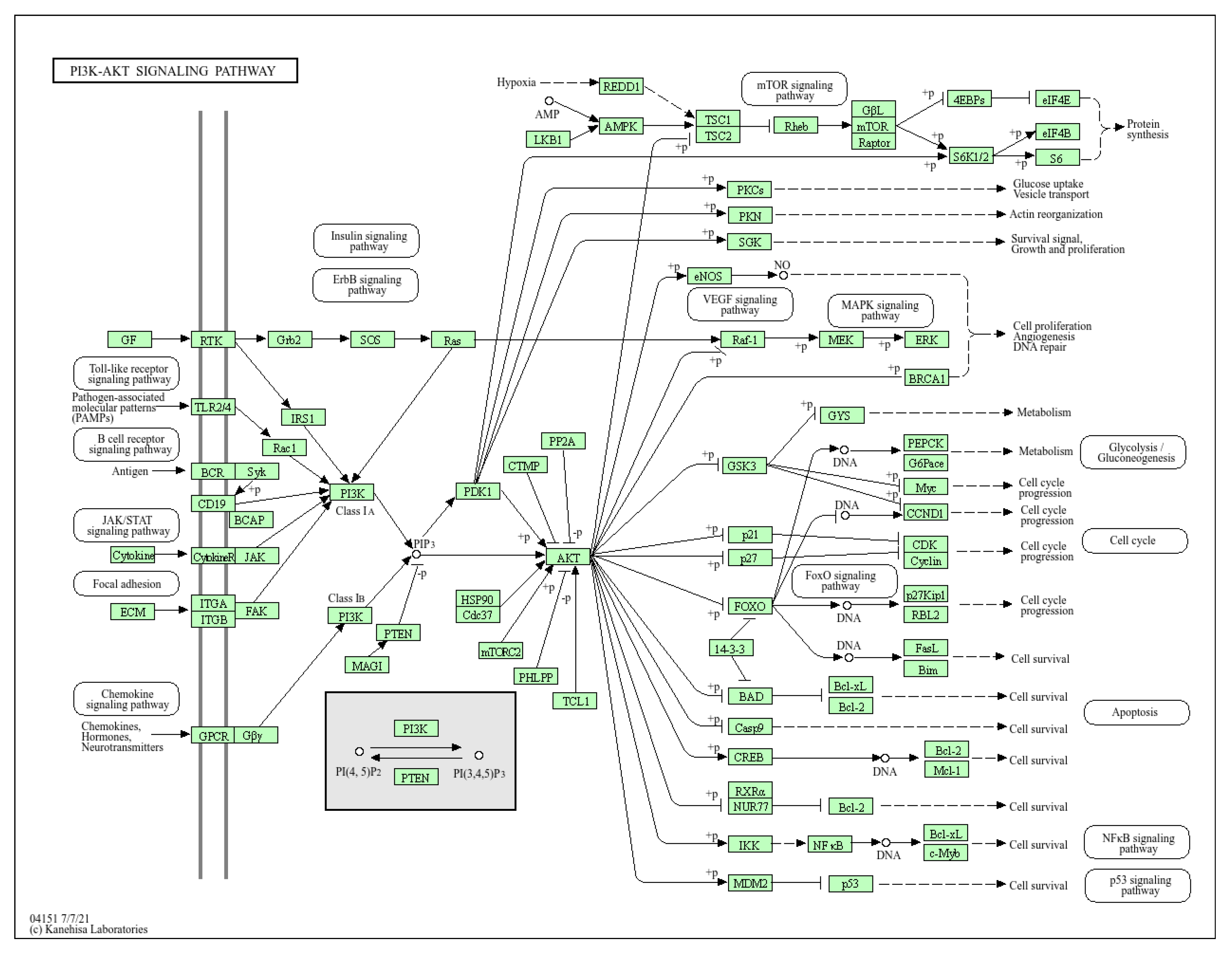
- He, Y.; Sun, M.M.; Zhang, G.G.; Yang, J.; Chen, K.S.; Xu, W.W.; Li, B. Targeting PI3K/Akt signal transduction for cancer therapy. Signal Transduct Target Ther 2021, 6, 425. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Geng, S.; Luo, H.; Wang, W.; Mo, Y.-Q.; Luo, Q.; Wang, L.; Song, G.-B.; Sheng, J.-P.; Xu, B. Signaling pathways involved in colorectal cancer: pathogenesis and targeted therapy. Signal Transduct Target Ther 2024, 9, 266. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; Peng, C.; Liu, Y. Regulation of ferroptosis by PI3K/Akt signaling pathway: a promising therapeutic axis in cancer. Front Cell Dev Biol 2024, 12. [Google Scholar] [CrossRef] [PubMed]
- Mohammadzadeh-Vardin, T.; Ghareyazi, A.; Gharizadeh, A.; Abbasi, K.; Rabiee, H.R. DeepDRA: Drug repurposing using multi-omics data integration with autoencoders. PLoS One 2024, 19, e0307649. [Google Scholar] [CrossRef] [PubMed]
- Caforio, M.; de Billy, E.; De Angelis, B.; Iacovelli, S.; Quintarelli, C.; Paganelli, V.; Folgiero, V. PI3K/Akt Pathway: The Indestructible Role of a Vintage Target as a Support to the Most Recent Immunotherapeutic Approaches. Cancers (Basel) 2021, 13, 4040. [Google Scholar] [CrossRef] [PubMed]
- Ager, C.; Reilley, M.; Nicholas, C.; Bartkowiak, T.; Jaiswal, A.; Curran, M.; Albershardt, T.C.; Bajaj, A.; Archer, J.F.; Reeves, R.S.; et al. 31st Annual Meeting and Associated Programs of the Society for Immunotherapy of Cancer (SITC 2016): part two. J Immunother Cancer 2016, 4, 73. [Google Scholar] [CrossRef]
- Delgado, F.M.; Gómez-Vela, F. Computational methods for Gene Regulatory Networks reconstruction and analysis: A review. Artif Intell Med 2019, 95, 133–145. [Google Scholar] [CrossRef] [PubMed]
- Rao, J.; Wang, X.; Chen, X.; Liu, Y.; Jiang, J.; Wang, Z. Multi-omics analysis reveals that Cas13d contributes to PI3K-AKT signaling and facilitates cell proliferation via PFKFB4 upregulation. Gene 2024, 927, 148760. [Google Scholar] [CrossRef] [PubMed]
- Slobodyanyuk, M.; Bahcheli, A.T.; Klein, Z.P.; Bayati, M.; Strug, L.J.; Reimand, J. Directional integration and pathway enrichment analysis for multi-omics data. Nat Commun 2024, 15, 5690. [Google Scholar] [CrossRef] [PubMed]
- Karim, S.; Burzangi, A.S.; Ahmad, A.; Siddiqui, N.A.; Ibrahim, I.M.; Sharma, P.; Abualsunun, W.A.; Gabr, G.A. PI3K-AKT Pathway Modulation by Thymoquinone Limits Tumor Growth and Glycolytic Metabolism in Colorectal Cancer. Int J Mol Sci 2022, 23, 2305. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Sun, M.; Huang, H.; Jin, W.-L. Drug repurposing for cancer therapy. Signal Transduct Target Ther 2024, 9, 92. [Google Scholar] [CrossRef] [PubMed]
- Garg, P.; Ramisetty, S.; Nair, M.; Kulkarni, P.; Horne, D.; Salgia, R.; Singhal, S.S. Strategic advancements in targeting the PI3K/AKT/mTOR pathway for Breast cancer therapy. Biochem Pharmacol 2025, 236, 116850. [Google Scholar] [CrossRef] [PubMed]
- Johnson, K.B.; Wei, W.; Weeraratne, D.; Frisse, M.E.; Misulis, K.; Rhee, K.; Zhao, J.; Snowdon, J.L. Precision Medicine, AI, and the Future of Personalized Health Care. Clin Transl Sci 2021, 14, 86–93. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-M.; Hsiao, T.-H.; Lin, C.-H.; Fann, Y.C. Unlocking precision medicine: clinical applications of integrating health records, genetics, and immunology through artificial intelligence. J Biomed Sci 2025, 32, 16. [Google Scholar] [CrossRef] [PubMed]
- Fu, C.; Chen, Q. The future of pharmaceuticals: Artificial intelligence in drug discovery and development. J Pharm Anal 2025, 101248. [Google Scholar] [CrossRef]
- Yunfan, G.; Yun, X.; Xinyu, G.; Kangxiang, J.; Jinliu, P.; Yuxi, B.; Yi, D.; Jiawei, S.; Haofen, W. Retrieval-Augmented Generation for Large Language Models: A Survey. Computer Science. Computer Science Computation and Language 2024, 11–21. [Google Scholar]
- Lin, X.; Deng, G.; Li, Y.; Ge, J.; Ho, J.W.K.; Liu, Y. GeneRAG: Enhancing Large Language Models with Gene-Related Task by Retrieval-Augmented Generation 2024.
- Li, M.; Kilicoglu, H.; Xu, H.; Zhang, R. BiomedRAG: A retrieval augmented large language model for biomedicine. J Biomed Inform 2025, 162, 104769. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Hein, M.Y.; Luber, C.A.; Paron, I.; Nagaraj, N.; Mann, M. Accurate Proteome-wide Label-free Quantification by Delayed Normalization and Maximal Peptide Ratio Extraction, Termed MaxLFQ. Molecular & Cellular Proteomics 2014, 13, 2513–2526. [Google Scholar]
- Cai, Z.; Poulos, R.C.; Liu, J.; Zhong, Q. Machine learning for multi-omics data integration in cancer. iScience 2022, 25, 103798. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Parmigiani, G.; Johnson, W.E. ComBat-seq: batch effect adjustment for RNA-seq count data. NAR Genom Bioinform 2020, 2. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Safronova, N.; Junghans, L.; Saenz, J.P. Temperature change elicits lipidome adaptation in the simple organisms Mycoplasma mycoides and JCVI-syn3B. Cell Rep 2024, 43, 114435. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Chen, S.; Bei, Y.; Yuan, Z.; Zhou, H.; Hong, Z.; Dong, J.; Chen, H.; Chang, Y.; Huang, X. A Survey of Graph Retrieval-Augmented Generation for Customized Large Language Models. 2025.
- Wang, Y.; Sun, Z.; He, Q.; Li, J.; Ni, M.; Yang, M. Self-supervised graph representation learning integrates multiple molecular networks and decodes gene-disease relationships. Patterns 2023, 4, 100651. [Google Scholar] [CrossRef] [PubMed]
- Shyam, P. In Silico Strategies for Cancer Model Development and Anticancer Drug Testing. In Preclinical cancer models for translational research and drug development; Springer Nature Singapore: Singapore, 2025; pp. 153–168.
- Guo, W.; Liu, S.; Zheng, X.; Xiao, Z.; Chen, H.; Sun, L.; Zhang, C.; Wang, Z.; Lin, L. Network Pharmacology/Metabolomics-Based Validation of AMPK and PI3K/AKT Signaling Pathway as a Central Role of Shengqi Fuzheng Injection Regulation of Mitochondrial Dysfunction in Cancer-Related Fatigue. Oxid Med Cell Longev 2021, 2021. [Google Scholar] [CrossRef] [PubMed]
- Richardson, E.; Trevizani, R.; Greenbaum, J.A.; Carter, H.; Nielsen, M.; Peters, B. The receiver operating characteristic curve accurately assesses imbalanced datasets. Patterns 2024, 5, 100994. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Wang, Z.; Wang, C.; Li, C.; Wang, B. Comparative Evaluation of Machine Learning Models for Subtyping Triple-Negative Breast Cancer: A Deep Learning-Based Multi-Omics Data Integration Approach. J Cancer 2024, 15, 3943–3957. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Liao, N.; Du, X.; Chen, Q.; Wei, B. A semi-supervised approach for the integration of multi-omics data based on transformer multi-head self-attention mechanism and graph convolutional networks. BMC Genomics 2024, 25, 86. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Zhang, W.; Lu, F.; Qin, T.; Gou, Y.; Guo, E.; Peng, D.; Zhang, L.; Yang, B.; Liu, S.; et al. Large language models completely understand molecular characteristics of squamous cervical cancer 2023.
- Asada, K.; Kobayashi, K.; Joutard, S.; Tubaki, M.; Takahashi, S.; Takasawa, K.; Komatsu, M.; Kaneko, S.; Sese, J.; Hamamoto, R. Uncovering Prognosis-Related Genes and Pathways by Multi-Omics Analysis in Lung Cancer. Biomolecules 2020, 10, 524. [Google Scholar] [CrossRef] [PubMed]

| Evaluation Measure | Value |
|---|---|
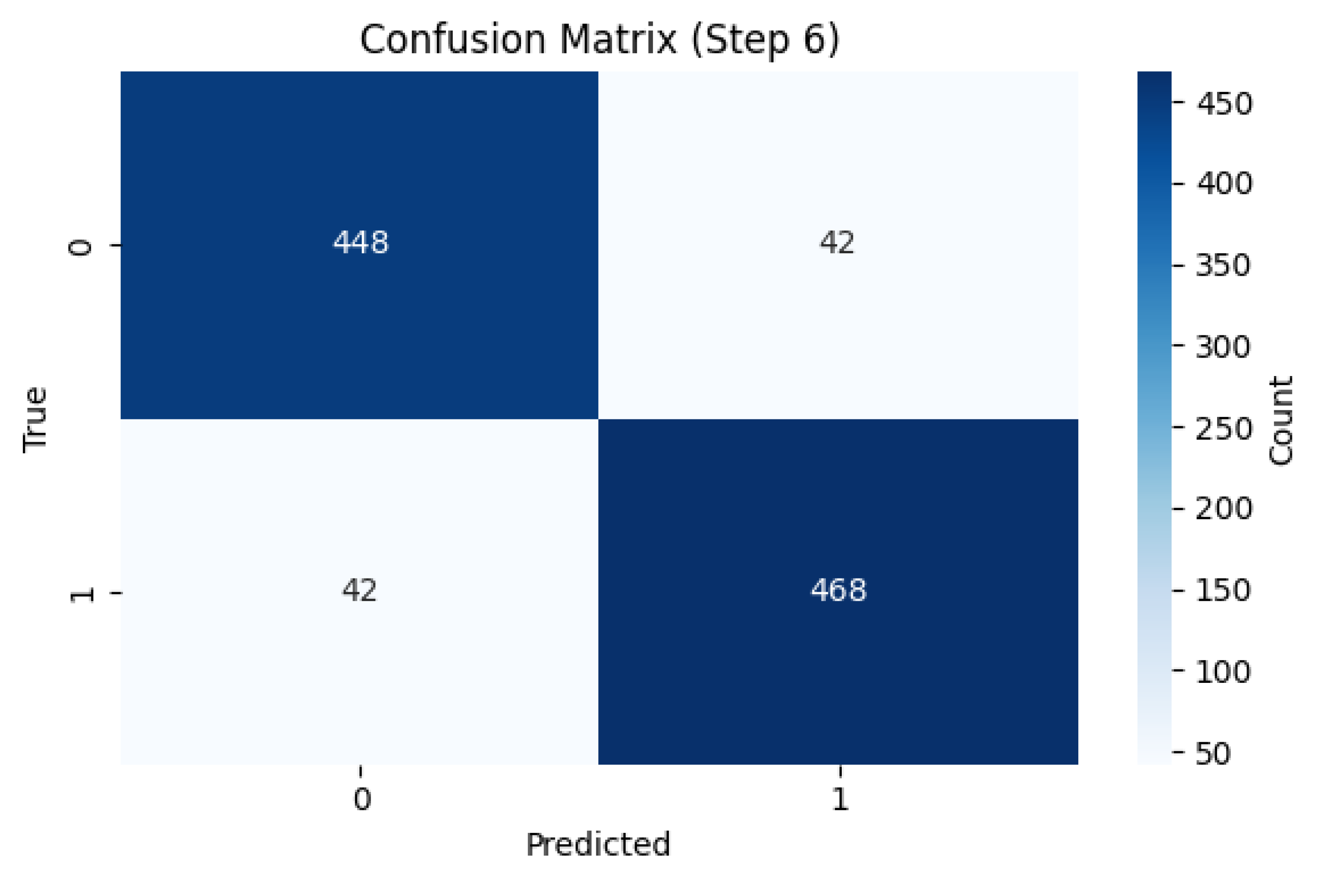
| Accuracy | 0.9200 |
| Sensitivity | 0.9176 |
| Specificity | 0.9143 |
| Precision | 0.9176 |
| Recall | 0.9176 |
| F1-Score | 0.9176 |
| Matthews Correlation Coefficient (MCC) | 0.8319 |
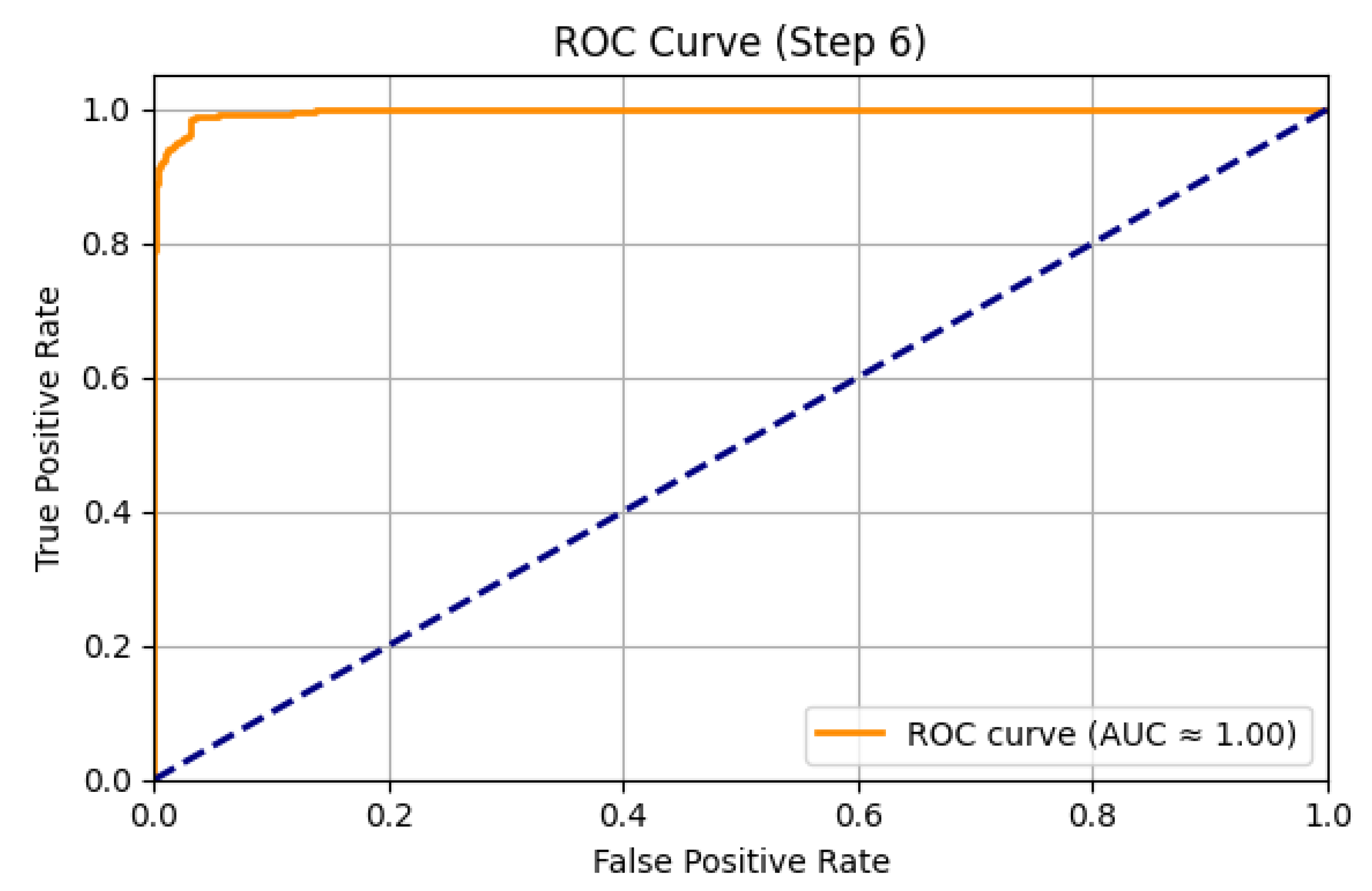
| ROC-AUC | 0.9000 |
| Novelty Detection Rate (NDR) | 0.8000 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




