Submitted:
04 July 2025
Posted:
08 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Peptides
2.2. Peptide Characterization by Reverse Phase-High Performance Liquid Chromatography and Structural Assessment
2.3. Assessment of Antibacterial Activity of Epinecidin-1 and Its Variants
2.4. Acridine Orange (AO) Ethidium Bromide (EtBr) Staining
2.5. Combined Antibacterial Effect of Peptides with Antibiotics
2.6. Wound Healing Assay
3. Results
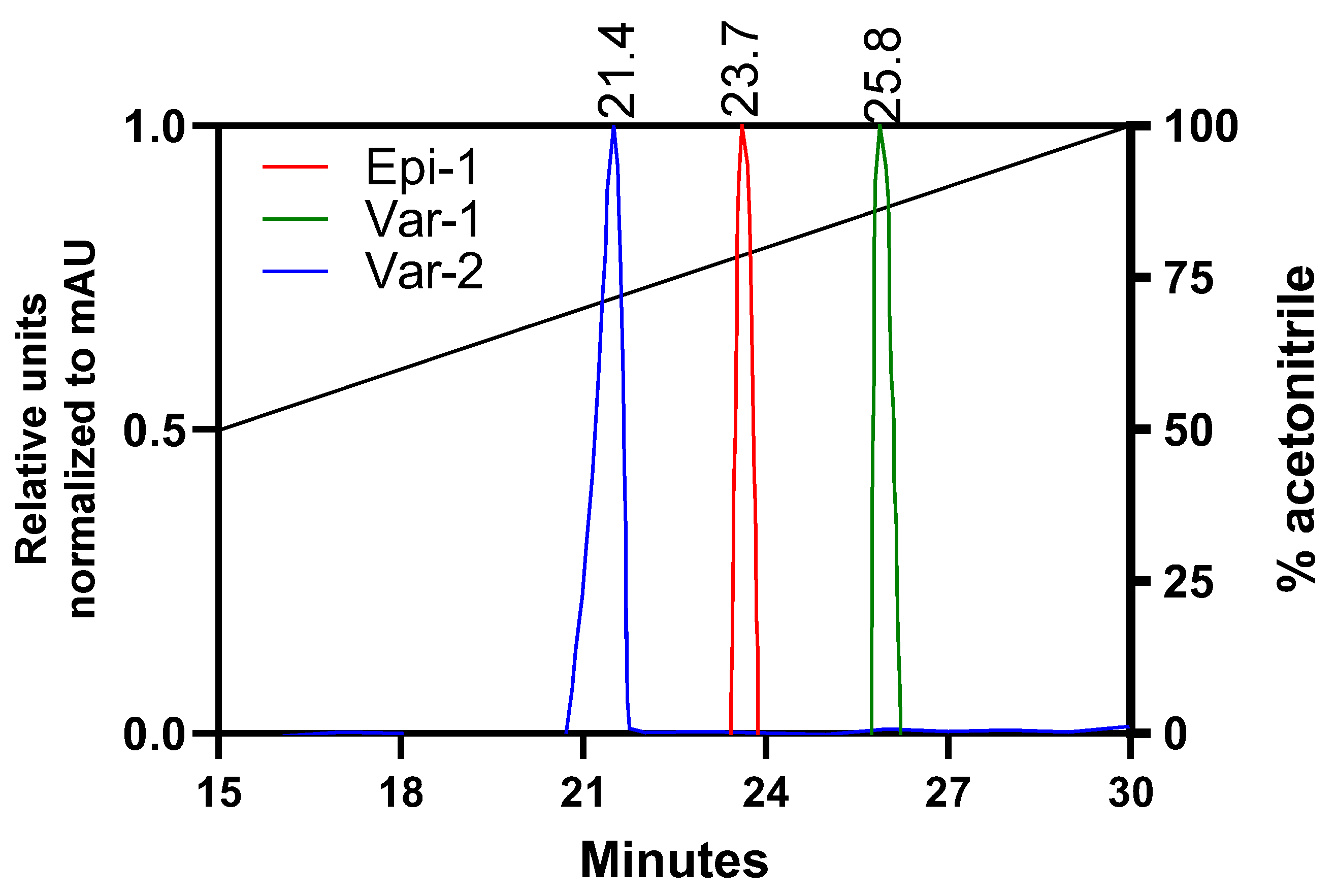
3.1. Characterization of Epi-1 and Its Variants
3.2. Electrostatic Surface Potential of Epi-1 and Its Variants
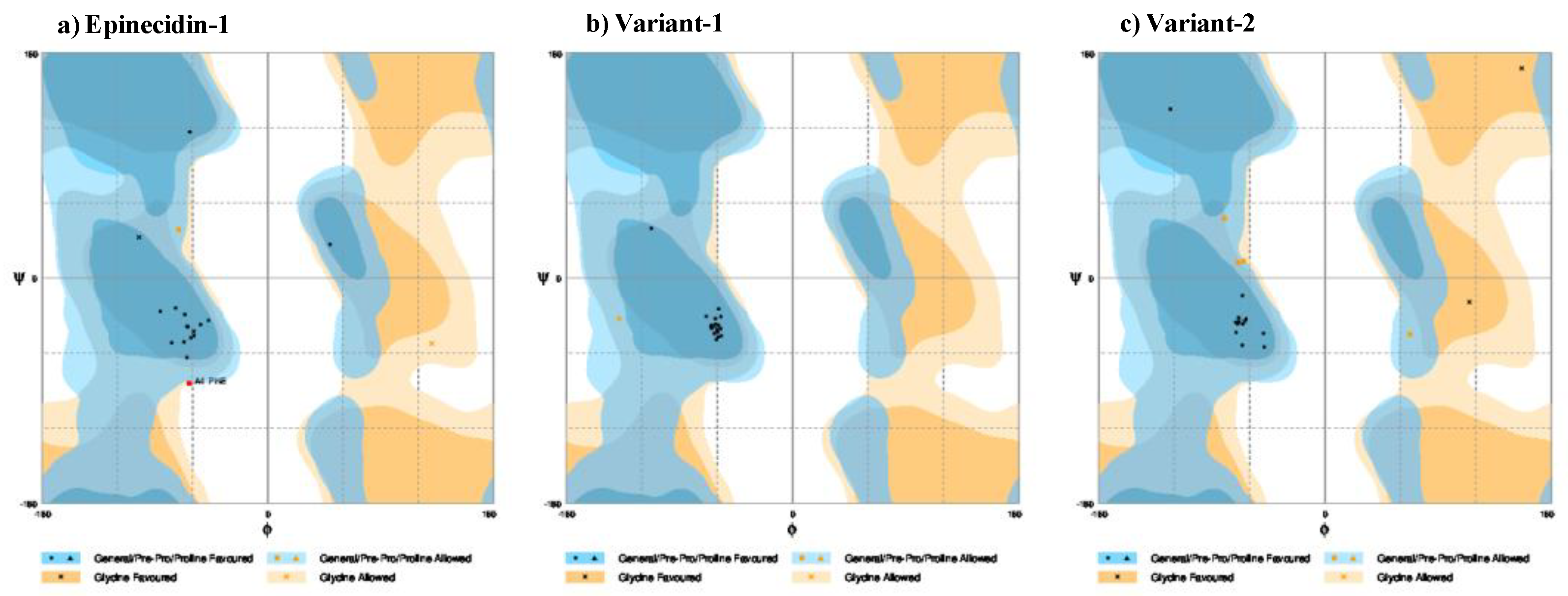
3.2. Ramachandran Plot of Epi-1 and Its Variants
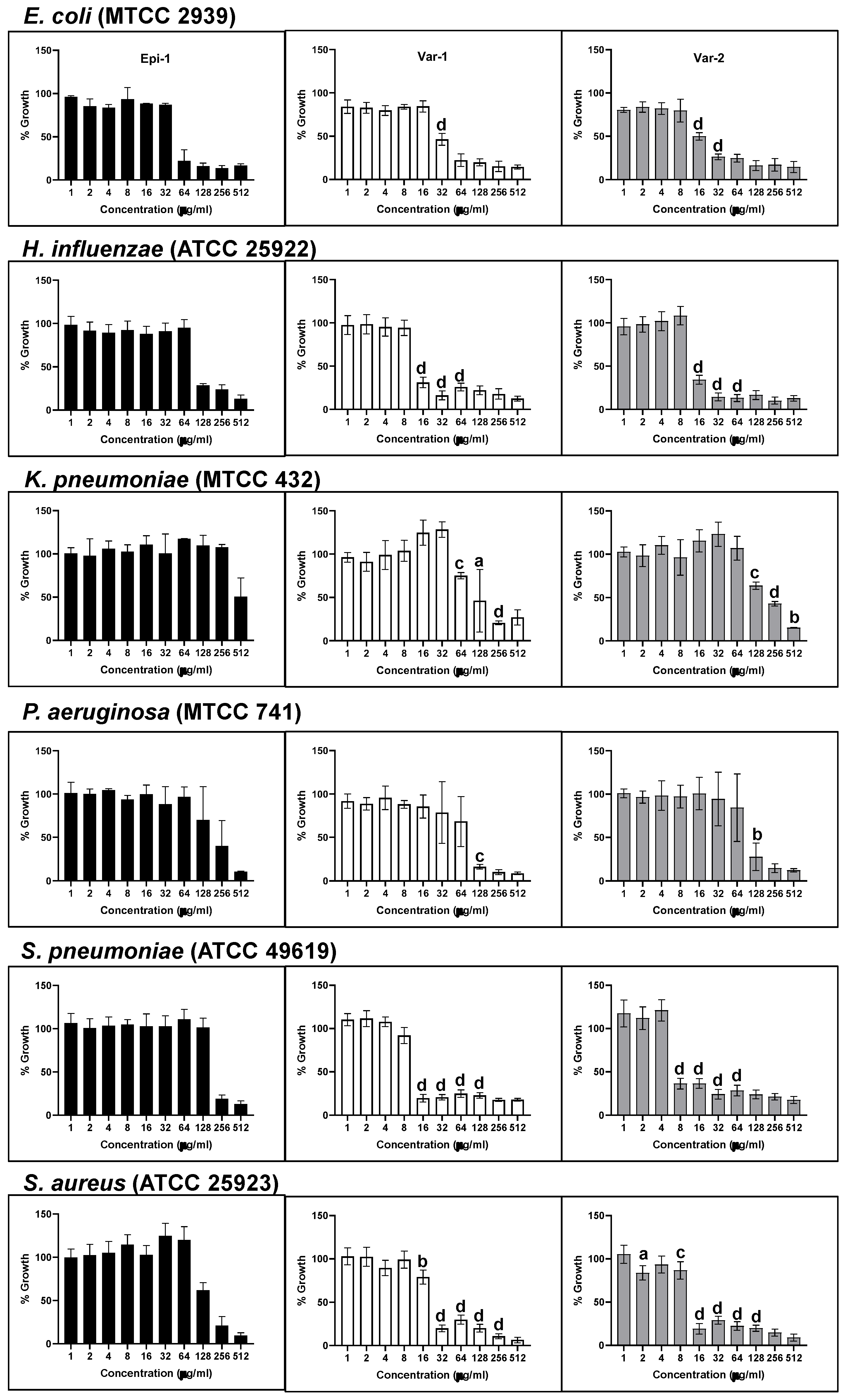
3.3. Bacteriostatic Effect of Epi-1 and Its Variants
| Name of the pathogen | Minimum Inhibition Concentration (μg/ml) |
||
|---|---|---|---|
| Epi-1 | Var-1 | Var-2 | |
| Gram negative | |||
| E. coli (MTCC 2939) | 64 | 64 | 32 |
| H. influenzae (ATCC 25922) | 128 | 16 | 16 |
| K. pneumoniae (MTCC 432) | 512 | 256 | 512 |
| P. aeruginosa (MTCC 741) | 512 | 128 | 128 |
| Gram positive | |||
| S. pneumoniae (ATCC 49619) | 256 | 16 | 8 |
| S. aureus (ATCC 25923) | 256 | 32 | 16 |
3.4. Impact of Epi-1 and Variants on Pathogenic Bacteria as Observed Through AO/EtBr Differential Staining
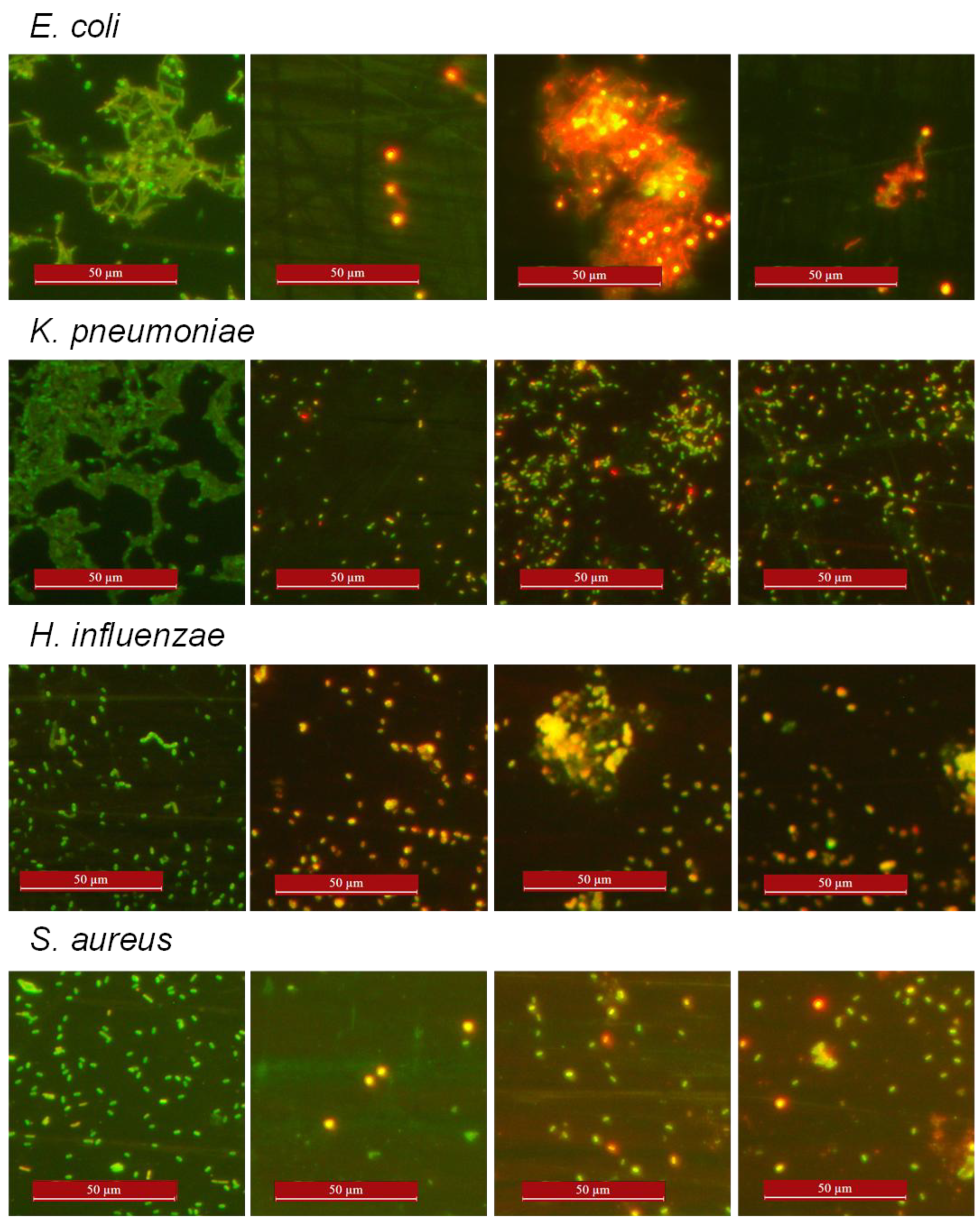
3.5. Combination Effect of Epi-1 Variants with Non Peptide Antibiotics on Antimicrobial Activity
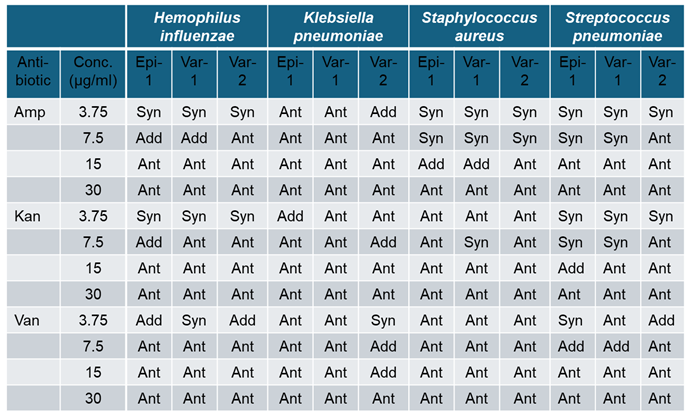 |
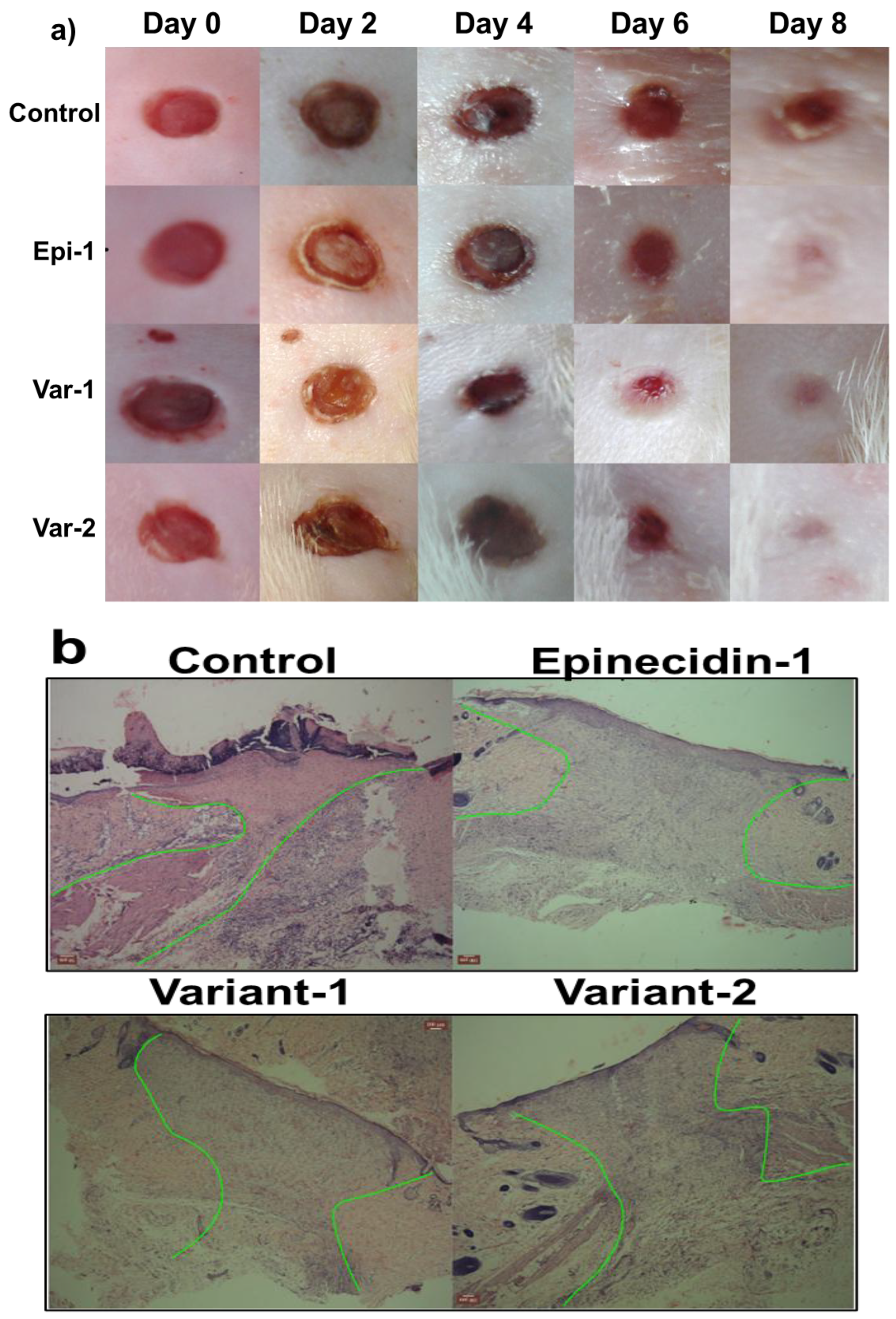
3.6. Epinecidin-1 and Its Variants Accelerated Wound Healing Activity in Rat Epidermis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
References
- Cižman, M.; Plankar Srovin, T. Antibiotic consumption and resistance of gram-negative pathogens (collateral damage). GMS Infect Dis 2018, 6, Doc05. [Google Scholar] [CrossRef] [PubMed]
- Woolhouse, M.; Waugh, C.; Perry, M.R.; Nair, H. Global disease burden due to antibiotic resistance - state of the evidence. J Glob Health 2016, 6, 010306. [Google Scholar] [CrossRef]
- Wang, C.H.; Hsieh, Y.H.; Powers, Z.M.; Kao, C.Y. Defeating Antibiotic-Resistant Bacteria: Exploring Alternative Therapies for a Post-Antibiotic Era. Int J Mol Sci 2020, 21. [Google Scholar] [CrossRef]
- Du Toit, A. Bacterial physiology: Efflux pumps, fitness and virulence. Nat Rev Microbiol 2017, 15, 512–513. [Google Scholar] [CrossRef] [PubMed]
- Pagès, J.M.; James, C.E.; Winterhalter, M. The porin and the permeating antibiotic: a selective diffusion barrier in Gram-negative bacteria. Nat Rev Microbiol 2008, 6, 893–903. [Google Scholar] [CrossRef]
- Masi, M.; Réfregiers, M.; Pos, K.M.; Pagès, J.M. Mechanisms of envelope permeability and antibiotic influx and efflux in Gram-negative bacteria. Nat Microbiol 2017, 2, 17001. [Google Scholar] [CrossRef]
- Reygaert, W.C. An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol 2018, 4, 482–501. [Google Scholar] [CrossRef]
- Mitchell, C.J.; Stone, T.A.; Deber, C.M. Peptide-Based Efflux Pump Inhibitors of the Small Multidrug Resistance Protein from Pseudomonas aeruginosa. Antimicrob Agents Chemother 2019, 63. [Google Scholar] [CrossRef] [PubMed]
- Rima, M.; Rima, M.; Fajloun, Z.; Sabatier, J.M.; Bechinger, B.; Naas, T. Antimicrobial Peptides: A Potent Alternative to Antibiotics. Antibiotics (Basel) 2021, 10. [Google Scholar] [CrossRef]
- Gambato, S.; Bellotto, O.; Mardirossian, M.; Di Stasi, A.; Gennaro, R.; Pacor, S.; Caporale, A.; Berti, F.; Scocchi, M.; Tossi, A. Designing New Hybrid Antibiotics: Proline-Rich Antimicrobial Peptides Conjugated to the Aminoglycoside Tobramycin. Bioconjug Chem 2023, 34, 1212–1220. [Google Scholar] [CrossRef]
- Taheri-Araghi, S. Synergistic action of antimicrobial peptides and antibiotics: current understanding and future directions. Front Microbiol 2024, 15, 1390765. [Google Scholar] [CrossRef]
- Panjla, A.; Kaul, G.; Chopra, S.; Titz, A.; Verma, S. Short Peptides and Their Mimetics as Potent Antibacterial Agents and Antibiotic Adjuvants. ACS Chem Biol 2021, 16, 2731–2745. [Google Scholar] [CrossRef]
- Biswal, S.; Caetano, K.; Jain, D.; Sarrila, A.; Munshi, T.; Dickman, R.; Tabor, A.B.; Rath, S.N.; Bhakta, S.; Ghosh, A.S. Antimicrobial Peptides Designed against the Ω-Loop of Class A β-Lactamases to Potentiate the Efficacy of β-Lactam Antibiotics. Antibiotics (Basel) 2023, 12. [Google Scholar] [CrossRef] [PubMed]
- Boehr, D.D.; Draker, K.A.; Koteva, K.; Bains, M.; Hancock, R.E.; Wright, G.D. Broad-spectrum peptide inhibitors of aminoglycoside antibiotic resistance enzymes. Chem Biol 2003, 10, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Macyszyn, J.; Burmistrz, M.; Mieczkowski, A.; Wojciechowska, M.; Trylska, J. Conjugates of Aminoglycosides with Stapled Peptides as a Way to Target Antibiotic-Resistant Bacteria. ACS Omega 2023, 8, 19047–19056. [Google Scholar] [CrossRef]
- Turner, J.; Muraoka, A.; Bedenbaugh, M.; Childress, B.; Pernot, L.; Wiencek, M.; Peterson, Y.K. The Chemical Relationship Among Beta-Lactam Antibiotics and Potential Impacts on Reactivity and Decomposition. Front Microbiol 2022, 13, 807955. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, A. Advances in antibiotic measurement. Adv Clin Chem 2012, 56, 75–104. [Google Scholar] [CrossRef]
- Jeyarajan, S.; Peter, A.S.; Ranjith, S.; Sathyan, A.; Duraisamy, S.; Kandasamy, I.; Chidambaram, P.; Kumarasamy, A. Glycine-replaced epinecidin-1 variant bestows better stability and stronger antimicrobial activity against a range of nosocomial pathogenic bacteria. Biotechnol Appl Biochem 2024, 71, 1384–1404. [Google Scholar] [CrossRef]
- Jeyarajan, S.; Peter, A.S.; Sathyan, A.; Ranjith, S.; Kandasamy, I.; Duraisamy, S.; Chidambaram, P.; Kumarasamy, A. Expression and purification of epinecidin-1 variant (Ac-Var-1) by acid cleavage. Applied Microbiology and Biotechnology 2024, 108, 176–176. [Google Scholar] [CrossRef]
- Jeyarajan, S.; Sathyan, A.; Peter, A.S.; Ranjith, S.; Duraisamy, S.; Natarajaseenivasan, S.M.; Chidambaram, P.; Kumarasamy, A. Bioproduction and Characterization of Epinecidin-1 and Its Variants Against Multi Drug Resistant Bacteria Through In Silico and In Vitro Studies. International Journal of Peptide Research and Therapeutics 2023, 29, 66–66. [Google Scholar] [CrossRef]
- Gautier, R.; Douguet, D.; Antonny, B.; Drin, G. HELIQUEST: a web server to screen sequences with specific alpha-helical properties. Bioinformatics 2008, 24, 2101–2102. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Zheng, W.; Li, Y.; Pearce, R.; Zhang, C.; Bell, E.W.; Zhang, G.; Zhang, Y. I-TASSER-MTD: a deep-learning-based platform for multi-domain protein structure and function prediction. Nature protocols 2022, 17, 2326–2353. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, G.N.; Ramakrishnan, C.; Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J Mol Biol 1963, 7, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Lovell, S.C.; Davis, I.W.; Arendall, W.B., 3rd; de Bakker, P.I.; Word, J.M.; Prisant, M.G.; Richardson, J.S.; Richardson, D.C. Structure validation by Calpha geometry: phi,psi and Cbeta deviation. Proteins 2003, 50, 437–450. [Google Scholar] [CrossRef]
- Willard, L.; Ranjan, A.; Zhang, H.; Monzavi, H.; Boyko, R.F.; Sykes, B.D.; Wishart, D.S. VADAR: a web server for quantitative evaluation of protein structure quality. Nucleic Acids Res 2003, 31, 3316–3319. [Google Scholar] [CrossRef] [PubMed]
- Thankappan, B.; Jeyarajan, S.; Hiroaki, S.; Anbarasu, K.; Natarajaseenivasan, K.; Fujii, N. Antimicrobial and antibiofilm activity of designed and synthesized antimicrobial peptide, KABT-AMP. Applied biochemistry and biotechnology 2013, 170, 1184–1193. [Google Scholar] [CrossRef]
- Duarte, D.; Vale, N. Evaluation of synergism in drug combinations and reference models for future orientations in oncology. Curr Res Pharmacol Drug Discov 2022, 3, 100110. [Google Scholar] [CrossRef]
- Feldman, A.T.; Wolfe, D. Tissue processing and hematoxylin and eosin staining. Methods Mol Biol 2014, 1180, 31–43. [Google Scholar] [CrossRef]
- Thankappan, B.; Sivakumar, J.; Asokan, S.; Ramasamy, M.; Pillai, M.M.; Selvakumar, R.; Angayarkanni, J. Dual antimicrobial and anticancer activity of a novel synthetic α-helical antimicrobial peptide. European Journal of Pharmaceutical Sciences 2021, 161, 105784–105784. [Google Scholar] [CrossRef]
- Tossi, A.; Sandri, L.; Giangaspero, A. Amphipathic, alpha-helical antimicrobial peptides. Biopolymers 2000, 55, 4–30. [Google Scholar] [CrossRef]
- Bognár, B.; Spohn, R.; Lázár, V. Drug combinations targeting antibiotic resistance. NPJ Antimicrob Resist 2024, 2, 29. [Google Scholar] [CrossRef] [PubMed]
- Yadav, P.K.; Su, M.; Jeyarajan, S.; Giblin, F.J.; Ohi, M.D. Structural Organization of the Guinea Pig αA-Crystallin and αA66−80 Peptide Complex. Microscopy and Microanalysis 2019, 25, 1318–1319. [Google Scholar] [CrossRef]
- Kumarasamy, A.; Jeyarajan, S.; Kimler, V.A.; Premceski, A.; Cheon, J.; Mishra, V.; Giblin, F.J. In vitro studies on the interaction of guinea pig αA crystallin and αA crystallin (66-80) peptide using fluorescence polarization and transmission electron microscopy. Investigative Ophthalmology & Visual Science 2017, 58, 5301–5301. [Google Scholar]
- Anbarasu, K.; Sivakumar, J. Multidimensional significance of crystallin protein–protein interactions and their implications in various human diseases. Biochimica et Biophysica Acta (BBA)-General Subjects 2016, 1860, 222–233. [Google Scholar] [CrossRef] [PubMed]
- Jeyarajan, S.; Kumarasamy, A.; Cheon, J.; Premceski, A.; Seidel, E.; Kimler, V.A.; Giblin, F.J. TEM analysis of αA66-80 peptide-induced protein aggregates and amyloid fibrils in human and guinea pig αA-crystallins. Investigative Ophthalmology & Visual Science 2018, 59, 3043–3043. [Google Scholar]
- Jeyarajan, S.; Xavier, J.; Rao, N.M.; Gopal, V. Plasmid DNA delivery into MDA-MB-453 cells mediated by recombinant Her-NLS fusion protein. International Journal of Nanomedicine 2010, 5, 725–733. [Google Scholar] [CrossRef]
- Govindarajan, S.; Sivakumar, J.; Garimidi, P.; Rangaraj, N.; Kumar, J.M.; Rao, N.M.; Gopal, V. Targeting human epidermal growth factor receptor 2 by a cell-penetrating peptide–affibody bioconjugate. Biomaterials 2012, 33, 2570–2582. [Google Scholar] [CrossRef]
- Jeyarajan, S.; Ranjith, S.; Veerapandian, R.; Natarajaseenivasan, K.; Chidambaram, P.; Kumarasamy, A. Antibiofilm Activity of Epinecidin-1 and Its Variants Against Drug-Resistant Candida krusei and Candida tropicalis Isolates from Vaginal Candidiasis Patients. Infectious Disease Reports 2024, 16, 1214–1229. [Google Scholar] [CrossRef]
- Jeyarajan, S.; Ranjith, S.; Thankappan, B.; Kumarasamy, A. A Step-by-Step Guide for Biosynthesis of Recombinant Fusion Antimicrobial Peptide and Release of the Active Peptide from Its Fusion Partner by Formic Acid Cleavage. Methods Mol Biol 2025, 2931, 155–175. [Google Scholar] [CrossRef]
- Jeyarajan, S.; Kumarasamy, A. Tagging fluorescent reporter to epinecidin-1 antimicrobial peptide. bioRxiv 2006. [Google Scholar] [CrossRef]
- Jeyarajan, S.; Kandasamy, I.; Veerapandian, R.; Jayachandran, J.; Chandrashekar, S.; Natarajaseenivasan, K.; Chidambaram, P.; Kumarasamy, A. Antibiofilm Activity of Protamine Against the Vaginal Candidiasis Isolates of Candida albicans, Candida tropicalis and Candida krusei. Applied Biosciences 2025, 4, 5. [Google Scholar] [CrossRef]
- Li, Y.; Wang, Y.; Luo, Y.L.; Bai, D.Q.; Zhang, G.; Wang, J.R.; Wei, H.; Li, S. Epinecidin-1 and lactic acid synergistically inhibit Aeromonas hydrophila through membrane disruption. Microb Pathog 2024, 196, 106879. [Google Scholar] [CrossRef] [PubMed]
- Bolatchiev, A. Antimicrobial Peptides Epinecidin-1 and Beta-Defesin-3 Are Effective against a Broad Spectrum of Antibiotic-Resistant Bacterial Isolates and Increase Survival Rate in Experimental Sepsis. Antibiotics (Basel) 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Talapko, J.; Meštrović, T.; Juzbašić, M.; Tomas, M.; Erić, S.; Horvat Aleksijević, L.; Bekić, S.; Schwarz, D.; Matić, S.; Neuberg, M.; et al. Antimicrobial Peptides-Mechanisms of Action, Antimicrobial Effects and Clinical Applications. Antibiotics (Basel) 2022, 11. [Google Scholar] [CrossRef]
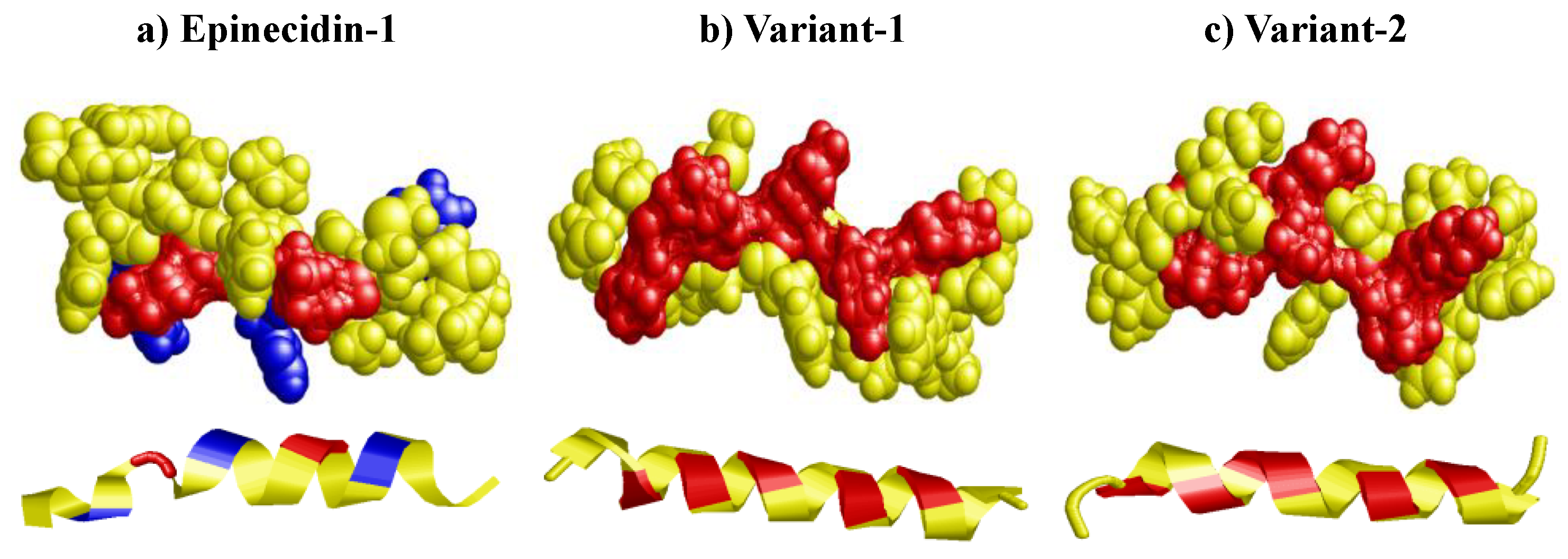
| Peptide name | Sequence |
| Epinecidin-1 | GFIFHIIKGLFHAGKMIHGLV |
| Variant 1- Replacement of H with K | *GFIFKIIKGLFKAGKMIKGLV |
| Variant 2- Replacement of A with K | *GFIFKIIKGLFKKGKMIKGLV |
| Peptide | Sequence | Molecular weight (Da) |
Net Charge |
pI | Hydrophobic Moment |
Elution time (min) |
|---|---|---|---|---|---|---|
| Epi-1 | GFIFHIIKGLFHAGKMIHGLV | 2335.8 | +2 | 10 | 0.432 | 23.7 |
| Var-1 | GFIFKIIKGLFKAGKMIKGLV | 2308.9 | +5 | 10.6 | 0.513 | 25.8 |
| Var-2 | GFIFKIIKGLFKKGKMIKGLV | 2366.0 | +6 | 10.7 | 0.477 | 21.45 |
| Secondary structure | Distribution statistics of aminoacids in the Ramachandran map | |||||||
|---|---|---|---|---|---|---|---|---|
| Helix (%) |
Beta (%) |
Coil (%) |
Turn (%) |
Favored Region (%) |
Allowed Region (%) |
Outlier Region (%) |
||
| Epi-1 | 47 | - | 52 | 1 | 84.2 | 10.5 | 5.3 | |
| Var-1 | 76 | - | 23 | - | 94.7 | 5.3 | - | |
| Var-2 | 76 | - | 23 | 1 | 89.5 | 10.5 | - | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




